3.1. Origin of Bipartite Viruses in the Strand Model
The following standard values of parameters are used in all results with the strand model. We set , so that binding, copying and translation all occur on similar time scales. The time units used are arbitrary because only relative rates influence the numbers of different strand types produced. The number of viruses produced per infected cell is . The length of the full virus , and the length of the defective strands is , but only relative lengths influence the outcome.
Firstly, we consider the mean number of strands produced in one cell from the given initial numbers of infecting strands, as shown in
Table 1. The mean output numbers are determined from an average of
cells, all beginning from the same initial number of strands. When only A is present,
exactly. For other combinations, the total of A, D and E is 100. When A is present with either D or E, it is significantly suppressed. It can be seen in
Table 1 that E suppresses A more strongly than D does, even though D and E have the same length. It can also be seen that when D and E are in the same cell (either with or without A), more E than D is produced. E strands (which produce polymerases) have a slight advantage over D strands (that produce capsid proteins) in this model.
The difference between D and E is a somewhat surprising feature of the stochastic model. This difference does not arise in a deterministic model in which differential equations are written down for the mean numbers of strands per cell because the equations for the rates of change of D and E are the same if they have the same length. We attribute the difference observed between D and E in the stochastic model to the fact that the number of polymerases present in a cell is correlated with the number of polymerase genes, and hence the rate of replication is correlated with the number of E strands. In cells in which E strands are frequent, there are more polymerases produced and replication is faster, amplifying the number of Es. In cells in which D is frequent, there are fewer polymerases, and the replication of the D strands is slower. The net result is a higher average number of Es when averaged over many cells.
The numbers in
Table 1 are the mean numbers of viruses produced per cell. There can be substantial variation from one cell to another, even when cells begin with the same number of strands in the initial infection.
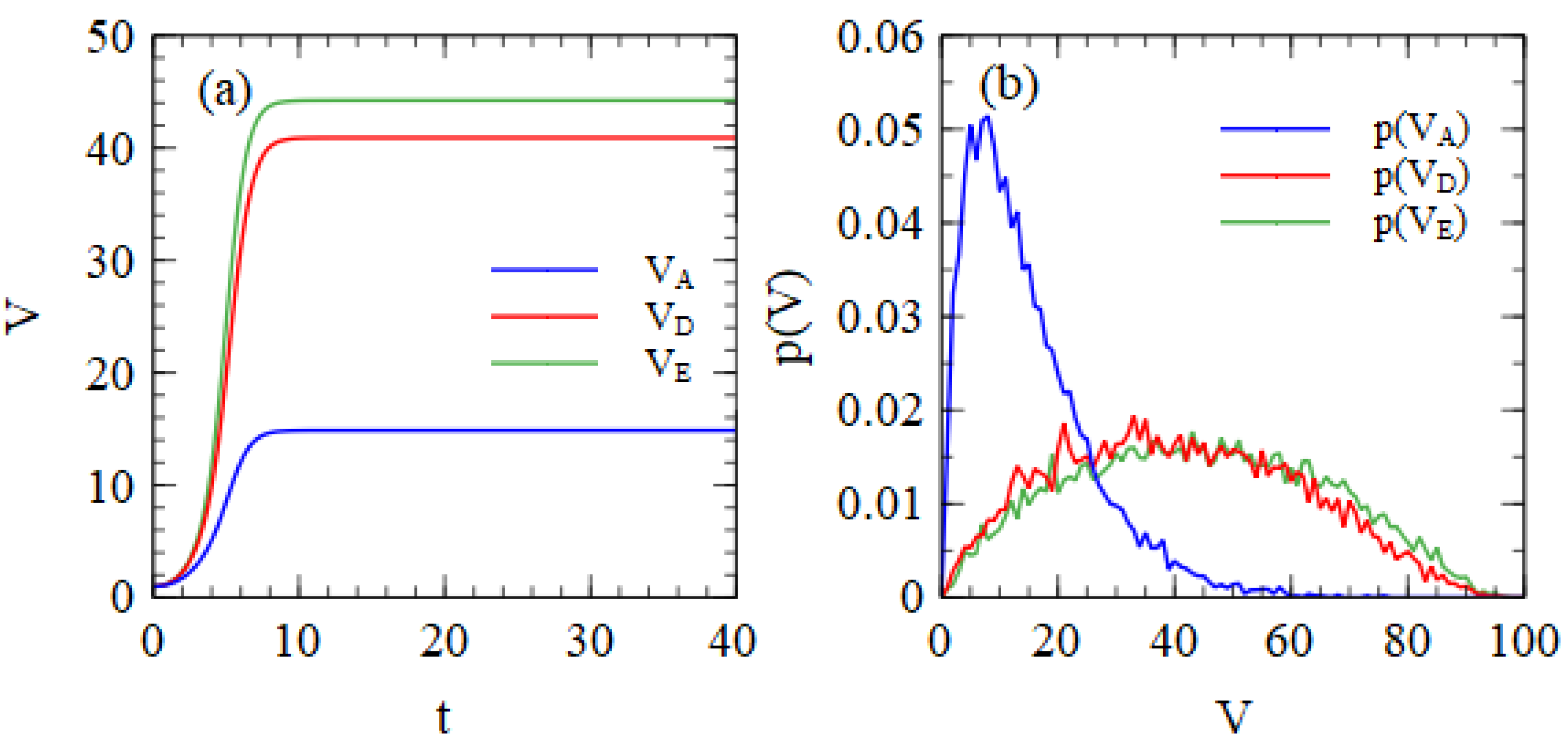
Figure 1a shows the mean numbers of strands of A, D and E per cell as a function of time, beginning from one copy of each. Replication in each cell is stopped when the total number of strands reaches 100. The time units are arbitrary, but all the cells have reached 100 strands by the time
t = 40.
Figure 1b shows the probability distribution of the number of output strands after all strands have reached 100 total strands. These distributions are very broad.
We now consider the transmission of strands over multiple cell generations. The population size is
N = 10,000. We begin with
= 1 for each of A, D and E, and determine λ in subsequent generations from the previous output numbers, as in Equations (1) and (2). Note that each cell in a generation has different numbers
, determined independently from the same Poisson distributions, and that the values of
are different for each type of strand because the output numbers of strands are different. The transmissibility α is the same for each kind of strand.
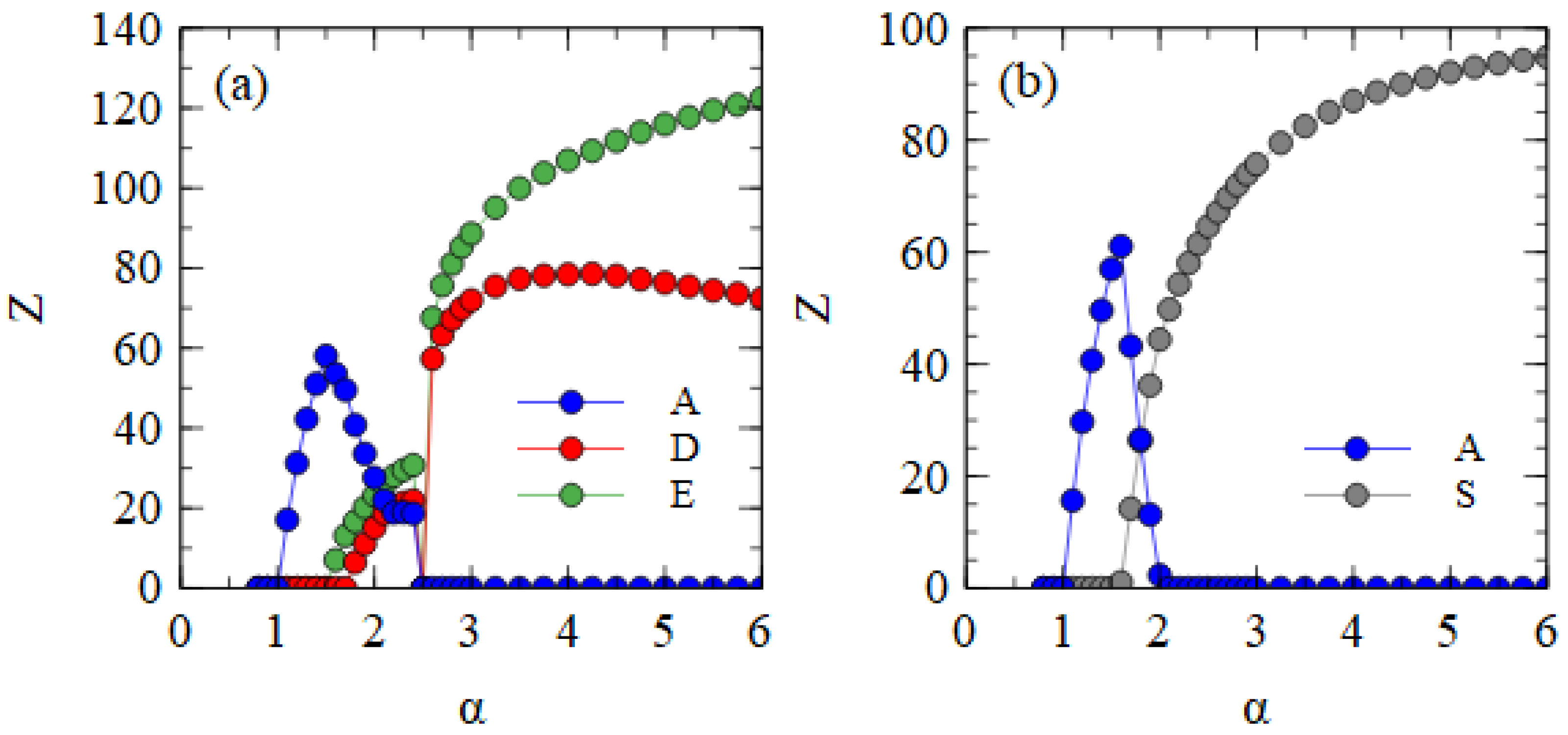
Figure 2a shows the mean output numbers of viruses per cell in the steady state as a function of α. Separate simulations are performed for each value of α, and the virus numbers are averaged over cells and over cell generations once steady-state numbers are reached.
Several regimes are visible in
Figure 2a. For α ≤ 1, all types of strands die out because they are not transmitted with sufficient frequency. For α > 1, the complete monopartite A virus can survive alone. As α increases, the fraction of cells infected by A increases and eventually becomes high enough to support the transmission of D and E as defective viruses. Since E suppresses A more strongly than D (as shown in
Table 1), E can survive with A at a lower value of α than D. In the range
, only A and E survive. For
all three strands survive. As α increases further, D and E suppress A to a greater extent until a point is reached where A is driven to extinction. For
, D and E survive as a bipartite virus in the absence of A.
The extinction of A only occurs because D and E have complementary genes. To demonstrate this, in
Figure 2b, we consider the case where D and E are not complementary. In this case, when D and E are in a cell with A, the model is the same as before, but when D and E are in the same cell without A, no replication occurs and there is no output of viruses from this cell. These simulations begin with A, D and E all present, but D is eliminated in all cases. Therefore, there is no curve for D in
Figure 2b. When the two defectives do not have complementary functions, E eliminates D because it is a better parasite of A. E is dependent on A; therefore, it cannot cause the extinction of A.
The strand model gives a simple explanation of why bipartite viruses arise. Some authors have looked for reasons as to why using separate particles might be advantageous. We suggest that the only advantage of D and E is that they are shorter and replicate faster than A when in the same cell. There is no advantage to being in separate particles. The two parts are packaged in separate particles by default because they are both derived from defective viruses of the same monopartite virus. The monopartite virus packages one complete strand per capsid; hence, it is likely that there should be one defective strand per capsid unless some new mechanism evolves that causes packaging of two strands in the same capsid (thus forming a segmented virus). Our model shows that the bipartite virus sometimes outcompetes the monopartite virus at sufficiently high α without the need to evolve a mechanism of segmented particle assembly.
The strand model does not yet consider the possibility of segmented viruses. In order to do this, it is necessary to allow for some possibility of packaging D and E in the same particle. Hence, we need a model that includes steps related to capsid production and assembly. The assumption of a fixed number of virus particles produced per cell seems too simple when we consider segmented viruses. Is the number of virus particles produced limited by the number of RNA strands or the number of capsids? This distinction does not matter if there is one strand per capsid because the number of strands is equal to the number of capsids. However, this is not true if we allow more than one strand per capsid. For these reasons, we find that the strand model is not sufficient to consider the origin of segmented viruses. As such, we now turn to the assembly model, which is able to address these issues.
3.2. Origin of Bipartite and Segmented Viruses in the Assembly Model
The following standard values of parameters are used in all results with the assembly model. We set , , and , as in the strand model. The assembly rate for particles containing A strands is always . The assembly rate for defective strands D and E is , and we begin with the case where this is also 1. The capsid number at which assembly becomes rapid is . We begin with the case where there is no association between D and E, ( A cell has a limiting amount of protein resources, , measured in units of number of virus capsids that can be synthesized, and a limiting amount of RNA resources, , measured in units of the number of full-length RNA strands that can be synthesized. Initially we set both limits to be equal, , with . We refer to the case where both limits are equal as ‘balanced resources’ (BR).
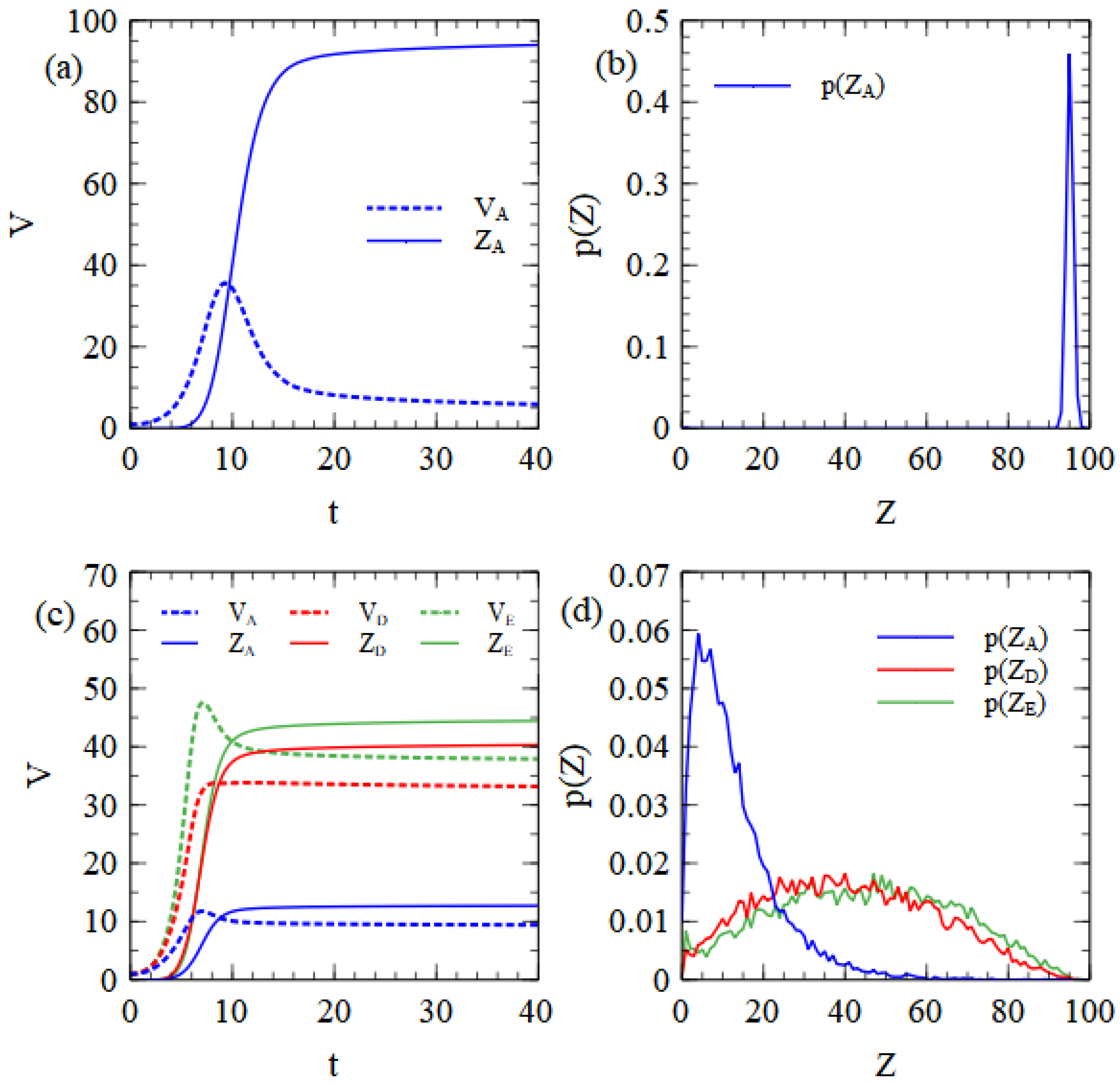
Figure 3a shows the mean number of unpackaged strands,
, and complete virus particles,
, per cell as functions of time. Unpackaged strands initially increase, but then decrease once packaging becomes faster than copying of new strands. After the resources are used, there is no further synthesis of capsids or RNA strands and the remaining capsids and strands are slowly assembled into virus particles. As the number of remaining capsids becomes low, the assembly rate drops greatly because it depends on
. A small number of strands therefore remains unpackaged when the maximum time
is reached. This seems to us to be a reasonable feature of the model that is likely to be true in real viruses.
We observed that if the fourth-power dependence is replaced by a linear dependence on C, there are fewer leftover unpackaged strands with long times, but that virus assembly is too rapid in the early stages of infection. There is a possibility that all virus strands become encapsulated when they are still few in number and that this prevents further replication before the resources of the cell are exhausted. This latter result does not seem realistic; therefore, we stick to the fourth-power dependence. The mean number of complete particles produced is 94.1 in this example. The probability distribution of the number of particles produced per cell (see
Figure 3b) is a sharp peak close to the mean. The monopartite virus is well adapted to these balanced resource conditions, since in most cells it uses both protein and RNA resources to the limit and manages to produce a number of particles that is very close to the limit given by the cell resources.
Figure 3c shows the mean numbers of unpackaged strands and virus particles as a function of time, beginning from a single strand of each of A, D and E. In this case, the RNA resource limit is 100, but the D and E strands are half-length, and only count for half a resource unit. Up to 200 D and E strands can be synthesized, which is a quantity larger than the limit on the capsids (which is still 100). A substantial number of unpackaged strands therefore remain in the cell when all the capsid proteins are turned into virus particles. The distribution of the number of particles produced of each type is shown in
Figure 3d, and this is very similar to the corresponding results for the strand model shown in
Figure 1b.
The mean numbers of virus particles produced from different combinations of starting strands are shown in
Table 2. The top five lines consider cases where
, meaning that no S complexes can be formed. These results are very similar to those obtained for the strand model in
Table 1, with the exception that the total number of particles produced is slightly less than 100 instead of exactly 100. We refer to the case where
and
as fully bipartite since only separate D and E particles are produced. The bottom five lines in
Table 2 consider cases where
and segmented particles can also form. In all cases, the assembly rate constant for segmented particles is
, the same as is obtained for monopartite A particles. We refer to the case with
and
as mostly bipartite. With this slow rate of complex formation and fast rate of assembly of defective particles, most of the D and E strands end up in separate particles and few segmented particles are produced.
We attempted to increase the number of segmented particles by increasing the rate of association between D and E to . However, unexpectedly, this leads to a slight reduction in the number of S particles and to a considerable reduction in the number of D and E particles. The problem is that D and E now form the complex S rapidly, which allows for rapid packaging into S particles but stops the replication of D and E. Additionally, the number of copies of D and E in a cell is not always balanced and, if the last copy of the rarer strand forms a complex, then no further replication of that strand is possible. Thus, the rapid formation of the S complex does not favor the production of more S particles. Increasing above 0.1 leads to the production of even fewer S particles.
In contrast, the number of S particles can be increased substantially if the rate of assembly of the single D and E particles is reduced to , while and the rate of formation of the complex remains low at . We refer to this parameter combination as mostly segmented because a large number of S particles is now produced, while there are fewer D and E particles. If , we have obtained the fully segmented case in which only S particles are produced and D and E strands cannot form separate particles.
The mean number of S particles in the fully segmented case is only 51.8. This value is much lower than the maximum of 100 that can be obtained from the available resources, and much lower than the number of monopartite viruses produced (94.1) when the monopartite virus is alone in the cell. The reason for this is that, even if a cell begins with , and even if the two strands replicate at the same average rate, the numbers of copies of the two strands do not remain equal. After one replication, we have a 2:1 ratio; if the next strand to be replicated is chosen randomly from the three, we are twice as likely to go a 3:1 ratio than to go to 2:2. By the time large numbers of strands are copied, it is likely that one of them is significantly more frequent than the other. If the RNA resource limit is 100, the total number of half-length strands produced is 200, meaning that there is an average number of 100 copies of each of D and E produced per cell. However, the rarer of the two strands is likely to have significantly fewer than 100 copies. In the fully segmented case, the maximum number of S particles possible is equal to the number of copies of the rarer of the two strands, which is usually much less than 100. This significant disadvantage of the segmented virus with respect to the monopartite virus arises from the stochastic replication process in the model, and this seems to be a realistic feature that will be experienced by real segmented viruses. It is not easy to see how a real virus could manage to achieve more balanced numbers of copies of the two strands than is created by random replication. The problem is significant if the D and E strands have equal length, as considered here, and it would be even worse if the length of the strands were unequal since the longer strand would usually be much rarer.
The final line of
Table 2 shows that when A, D and E are all in the same cell, more S particles are produced than A particles. For this reason, it is still possible for a segmented virus to outcompete a monopartite virus despite the fact that fewer S than A particles are produced when they are in separate cells, as we will now show.
We now consider transmission between cells using the assembly model.
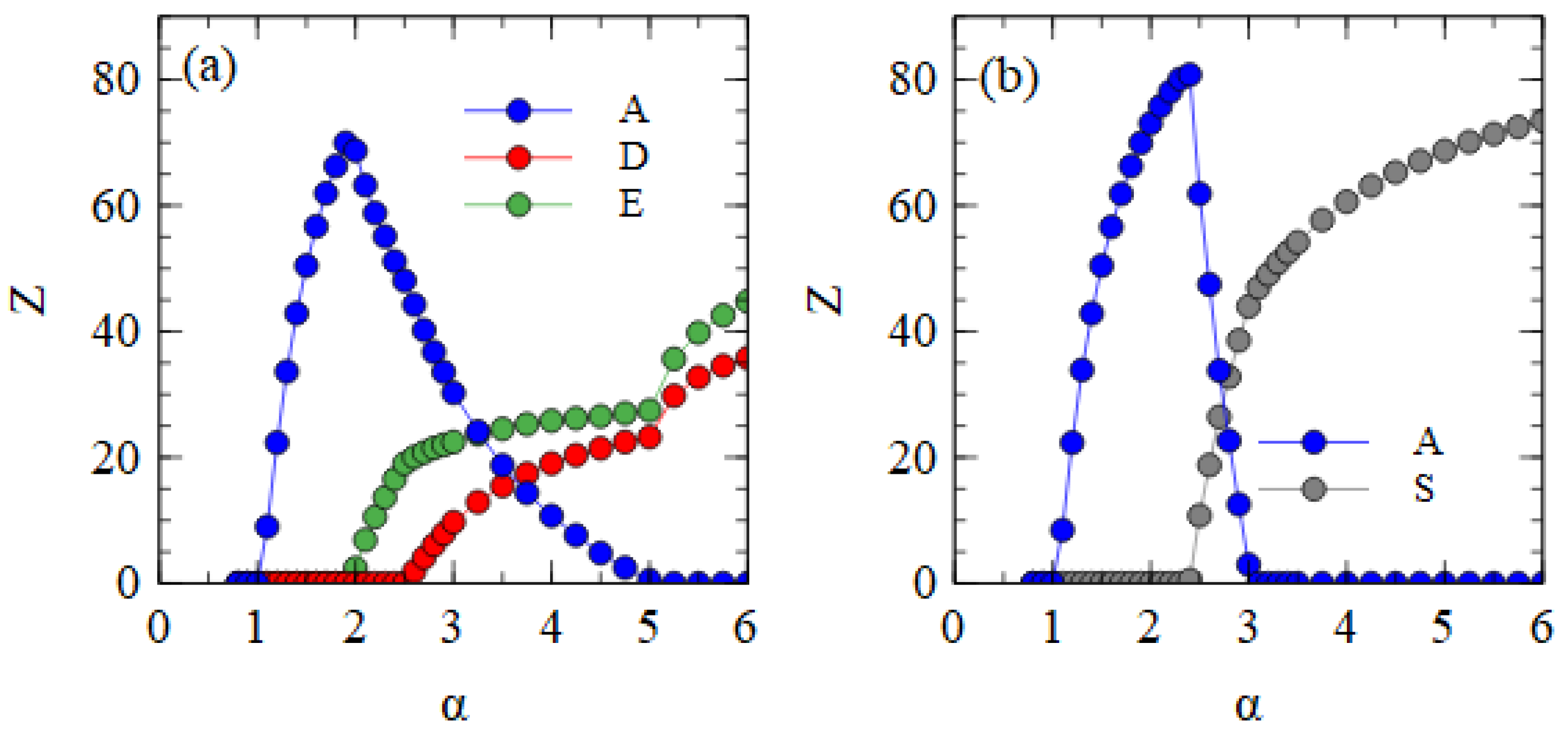
Figure 4a shows the mean numbers of A, D and E particles produced per cell, averaged over time, in the fully bipartite case where
and
. The bipartite case in the assembly model is very similar to the strand model (shown in
Figure 2a). There is a regime with only A at low α; this is followed by regimes of A + E, and A + D + E as α increases; these are followed by a regime with only D and E.
Figure 4b shows the fully segmented case where
and
. In this case, there is a regime of only A at low α, followed by a narrow regime where A and S coexist, followed by a regime at high α where the segmented virus eliminates the monopartite virus. This confirms that the advantage of S when in the same cell as A can outweigh the smaller production rate of S when they are in separate cells as long as transmissibility α is high enough.
3.3. Evolution of an Assembly Mechanism for the Segmented Virus
A comparison of the two cases in
Figure 4a,b shows that the minimum α required for the bipartite virus to eliminate the monopartite is 5, while the minimum α required for a segmented virus to eliminate the monopartite is 3. This may suggest that the using the segmented virus is a “better” strategy than relying on the bipartite virus, and moreover seems to confirm our intuition that packaging the two strands in the same particle makes more sense than packaging them separately. However, we are assuming that the D and E strands are defectives that arise via deletions in the A virus. As the A strands assemble into particles containing a single strand, it is likely that the D and E strands will assemble separately into particles containing a single strand unless a mechanism evolves that causes the association of D and E prior to assembly into a virus particle.
In this section, we suppose that D and E strands originally form without the ability to associate (), but that then a variant of D arises, which we call D*, that possesses some element of sequence or structure which allows it to associate with E. Both D and D* can form particles with a single strand, but only D* can form S particles. We wish to determine whether D* is selected relative to D.
Figure 5 shows the mostly bipartite case, with
for all short strands, i.e., D, D* and E, and
for the D* and E strands. As always,
. For each value of α, the simulation is initiated with A, D, D* and E strands present and allowed to proceed until a steady state is reached. E survives with A for α > 1.9. D and D* survive for α > 2.4. These limits are the same as those shown in
Figure 4a, which only has one D variant. Small numbers of S particles are also formed once α is large enough for D* to survive. However, we have already seen in
Table 2 that very few S particles form when
. This occurs because the D* strands are usually packaged separately before they can form the complex. Therefore, very few S particles are formed in the middle range of α in
Figure 5.
Given that few S particles are formed, there is little difference between D and D*. The data points for D and D* fluctuate up and down rather randomly, whereas the total of D and D* (dashed line) follows a smooth curve. This is a sign that the amounts of D and D* fluctuate by random drift and that there is no indication of selection in favor of D*. For α > 5, the bipartite virus eliminates the monopartite virus. At this point, D* is also eliminated, leaving only D and E. This shows that in cases where the bipartite virus does well, there is selection against the D* variant that forms the association with E. Thus, in conditions where D and E strands assemble efficiently into separate D and E particles, a mechanism that leads to the formation of S particles is not selected by evolution.
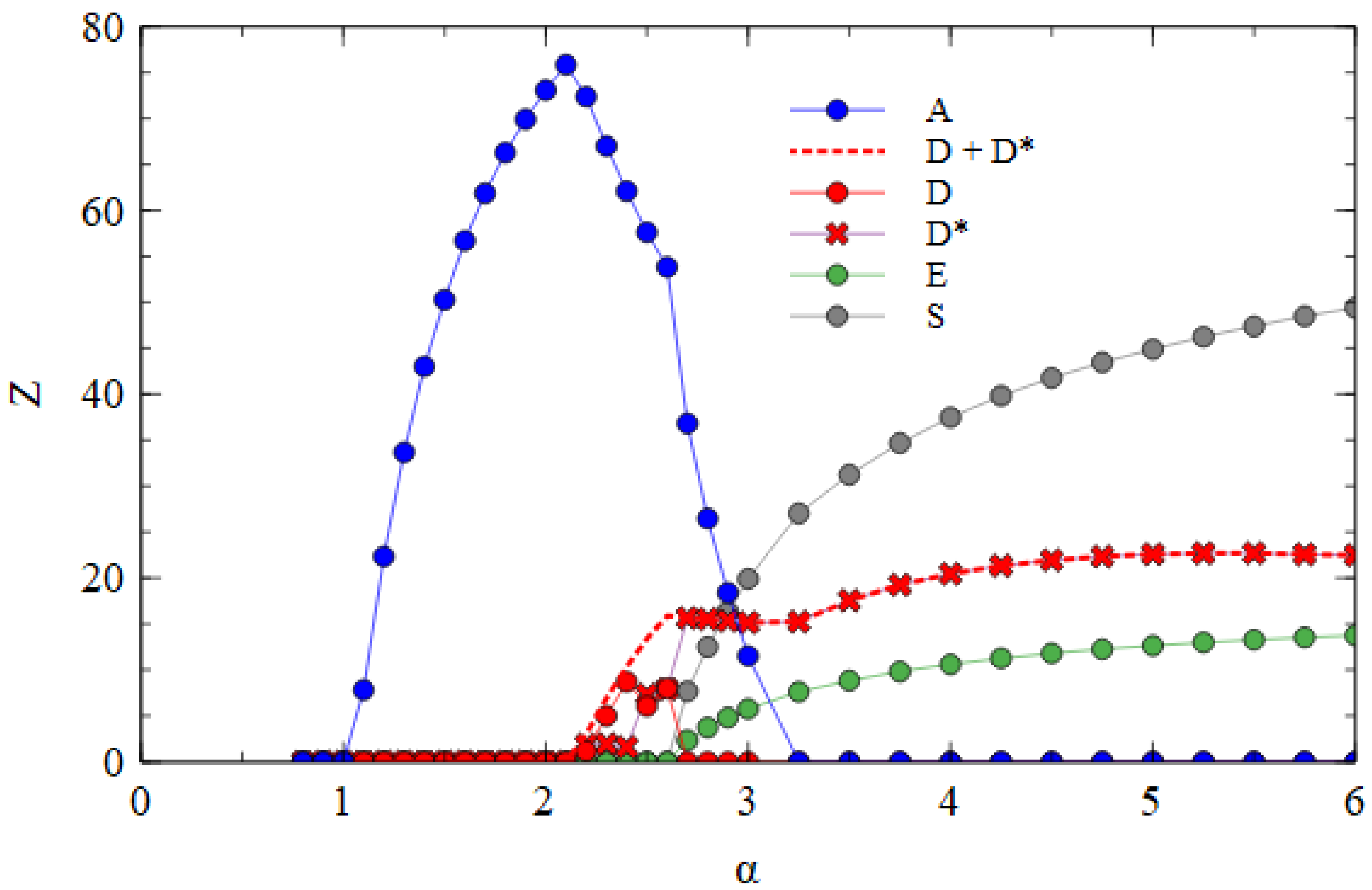
Figure 6 shows the mostly segmented case, where
for all short strands D, D* and E, and
for the D* and E strands. For these parameters, D survives at a lower α value than E. As D is the strand that produces capsid proteins, having a higher concentration of capsid proteins is an advantage when the packaging of defective strands is slow. The fact that E produces additional polymerases, which gives an advantage for E in previous cases, is less important when
is low. In the range 2.2 ≤ α ≤ 2.6, D and D* survive with A. As there is no E in this range, there is no difference between D and D*, and the frequencies of D and D* can fluctuate due to random drift, with only the total of D and D* being under selection. For α > 2.6, E also survives. At this point, D* and E start to form S complexes and S particles. We see in
Table 2 that substantial numbers of S particles form for these mostly segmented parameters. In
Figure 6, the number of S particles rises quickly with α. As soon as E survives (α > 2.6), D* is selected and D is eliminated. For α ≥ 3.25, A is also eliminated, leaving only D* and E strands which are transmitted mostly as S particles with smaller numbers of D* and E particles. Thus, if defective strands are not packaged efficiently into separate particles, a mechanism that causes the association of D* and E into segmented particles is selected by evolution. The resulting segmented particles compete successfully against the original monopartite virus and sometimes eliminate it. We have supposed that it is the D strand that evolves the new variant D*. However, we could equally well have considered E evolving into a variant E*, or into two variants D* and E*. We expect all these cases to be very similar.
3.4. Effect of Varying Resources
So far, we have considered balanced resources (BR) where
. Resource limits are a property of the host cell, and there is no reason why these should always be such that an equal number of capsids and RNA strands can be synthesized. Now, we consider the effect of varying the relative amounts of protein and RNA resources. We define excess protein resources (XPR) as cellular conditions where
. We simulate the case where
, and
, keeping
We define excess RNA resources (XRR) as cellular conditions where
. We simulate the case where
, and
, keeping
Table 3 shows mean numbers of viruses produced per cell under each resource condition for the monopartite, fully bipartite, and fully segmented viruses, scenarios in which each type of virus is alone in the cell.
The monopartite virus does well in BR conditions, producing 94.1 viruses, a number which approaches the maximum limit of 100.
Table 3 shows that the numbers of monopartite viruses in the XPR and XRR conditions are 99.9 and 98.0, and these values are only slightly higher than for BR conditions. Increasing the levels of one resource over the other does not make much difference to the monopartite virus because it is still limited by whichever of the two resources is the lower.
The bipartite virus also does well in BR conditions, producing 97.2 particles in total. However, for BR, up to 200 half-length strands can be produced, and only half of these can be packaged. For XPR conditions, up to 300 units of capsid proteins can be produced, which is enough to package all the 200 strands that are produced. Thus, the number of bipartite virus particles produced is roughly doubled in XPR conditions, which is a significant advantage. On the other hand, for XRR conditions, up to 600 half-length strands can be produced; however the maximum number of particles is still limited to 100 by the capsid proteins. Therefore, increasing RNA resources makes little difference to the bipartite virus.
The segmented virus has a significant disadvantage in BR conditions because the two strands tend to be produced in unequal numbers and as the number of segmented particles produced is limited by the rarer of the two. Up to 100 units of capsid proteins can be synthesized, but only 51.8 S particles are formed on average, leaving many capsid proteins remaining that do not end up in virus particles. In XPR conditions, up to 300 units of capsid proteins can be synthesized, but this simply increases the number of excess capsid proteins that do not form particles. Therefore, increasing protein resources does not benefit segmented viruses. On the other hand, for XRR conditions, up to 600 half-length strands are produced, and the distribution of the numbers of strands for each type is broad over the range from 0 to 600, with the mean being close to 300. In this case, there are more copies of the rarer strand than for the BR case, and there is a good chance that even the rarer strand has more than 100 copies. The mean number of S particles formed increases to 68.3, which is a big improvement from 51.8. Thus, increasing RNA resources gives a significant advantage to segmented viruses.
Figure 7a shows the mean number of viruses produced per cell in the steady state in the fully bipartite case with XPR. As expected, XPR gives an advantage to bipartite viruses. The curves are shifted to the left relative to those shown for BR in
Figure 4a. The minimum α at which the bipartite virus eliminates the monopartite is 2.5 for XPR conditions and 5.0 for BR conditions. With XRR conditions, there is no extra benefit provided to the bipartite virus. The minimum α at which the bipartite eliminates the monopartite remains at 5.0.
Figure 7b shows the mean number of viruses produced per cell in the fully segmented case with XRR. As expected, XRR gives an advantage to segmented viruses. The curves are shifted to the left relative to those obtained for BR shown in
Figure 4b. The minimum α at which the segmented virus eliminates the monopartite is 2.0 for XRR conditions and 3.1 for BR conditions. With XPR conditions, there is no extra benefit provided to the segmented virus. The minimum α at which the segmented eliminates the monopartite is 3.0, almost the same as the value for BR.
3.5. Effect of Deleterious Mutations
RNA viruses are known to have low-fidelity polymerases with high mutation rates. We therefore investigate the effect of deleterious mutations on the competition between virus types. The bipartite case is well described by the simpler strand model; therefore, we return to the strand model to look at deleterious mutations. An A virus with working versions of both capsid and polymerase genes is denoted as . Deleterious mutations are denoted as 0. Strands with deleterious mutations in one or both genes are denoted as , or . is a strand with a working capsid gene. is a strand with a deleterious mutation. is a strand with a working polymerase gene. is a strand with a deleterious mutation. Strands with mutant genes produce no proteins from the mutant genes, but they can still be replicated and transmitted between cells. Each time a sequence is copied, there is the probability of a deleterious mutation occurring in each gene, and a probability of correctly copying the gene. Thus, copying a gives a with probability of . Copying an gives an with probability . Copying an gives an or with probability or an with probability . The reverse mutation from a 0 to a functional gene is assumed to be negligible.
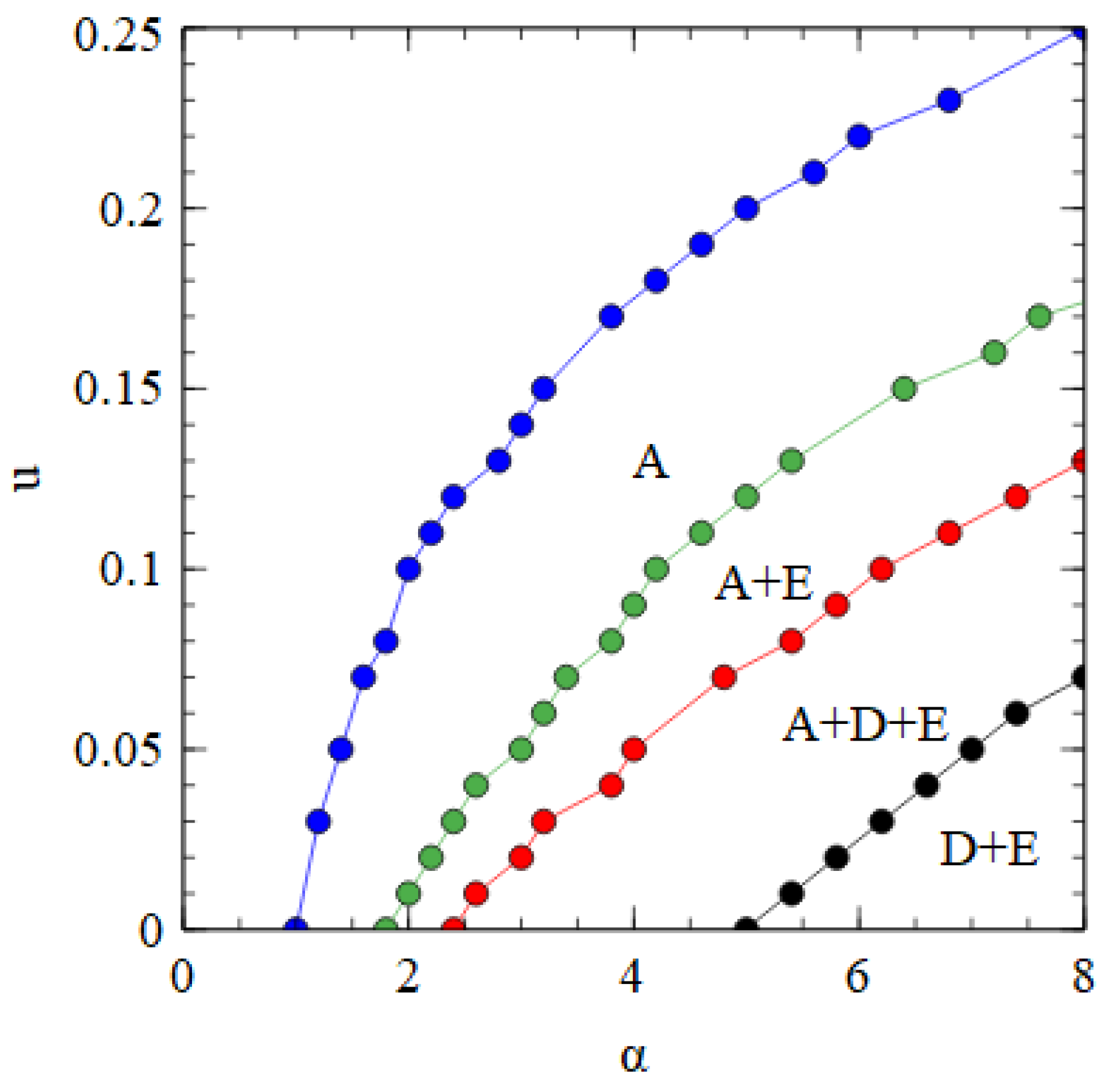
Figure 8 shows a phase diagram indicating which viruses survive in which regions of the
parameter space. The horizontal axis with
is equivalent to
Figure 2a. There is a region at high
and low
where only the bipartite virus D + E survives. By either decreasing
or increasing
, we move through regions of A + D + E, A + E, and only A, and eventually reach a region where no virus survives. The upper boundary line is the error threshold of the A virus. For mutation rates above the error threshold, the virus is eliminated by the accumulation of deleterious mutations.
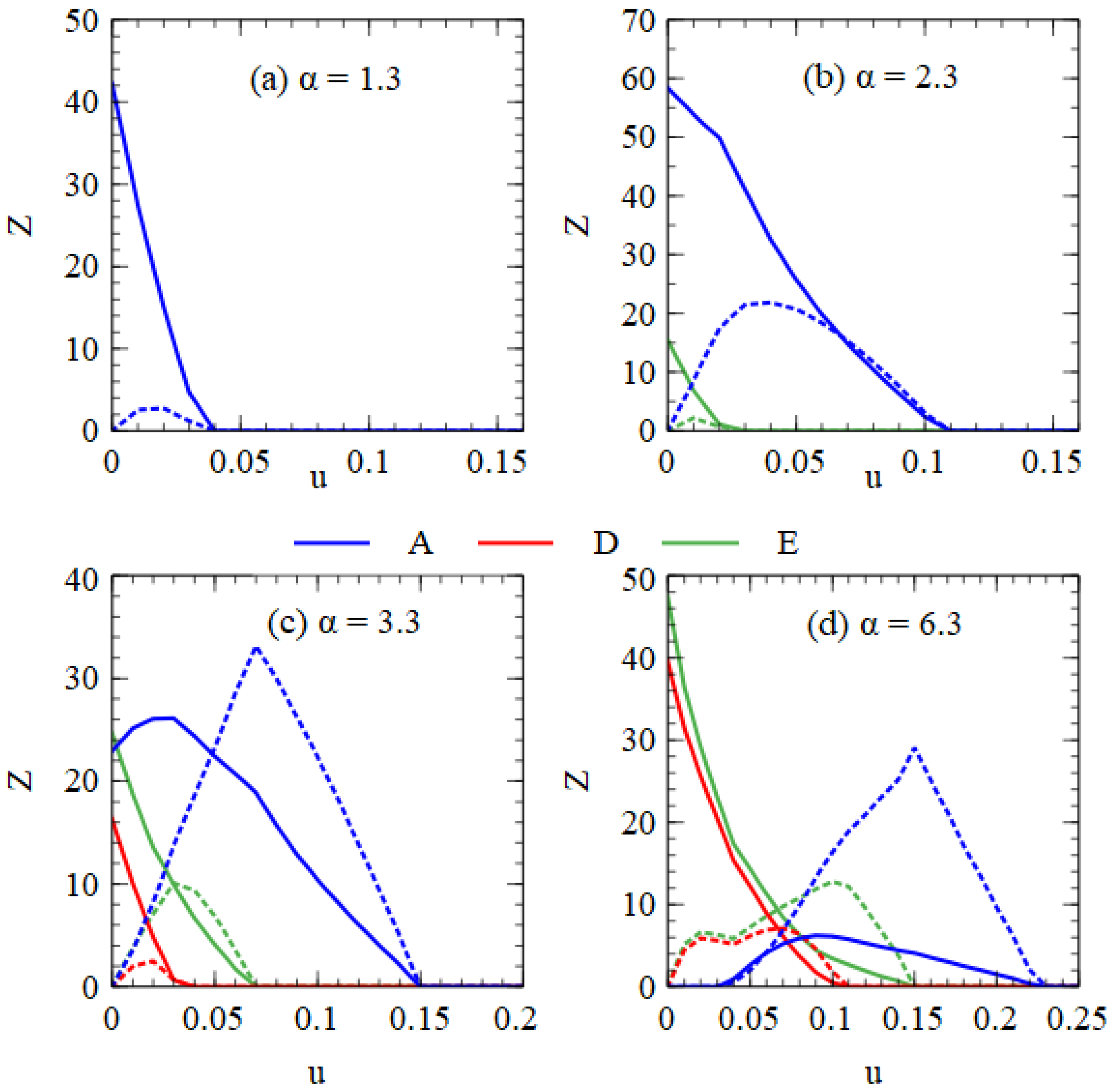
Figure 9 shows error threshold curves obtained by plotting virus numbers as a function of
u at fixed α values. In
Figure 9a for α = 1.3, we are in the regime of only A when
u = 0. As
u increases, we reach the error threshold for A. In
Figure 9b for α = 2.3, we are in the regime of A + E when
u = 0. As
u increases, we pass through the error thresholds for E and A. In
Figure 9c for α = 3.3, we are in the regime of A + D + E when
u = 0. As
u increases, we pass through the error thresholds for D, E and A. In
Figure 9d for α = 6.3, we are in the regime of D + E when
u = 0. As
u increases, we pass through the point where A first appears, followed by the error thresholds for D, E and A. In all these examples, increasing
u leads to A outcompeting D and E. Although both the monopartite and bipartite viruses are adversely affected by deleterious mutations, increasing the mutation rate favors the monopartite virus relative to the bipartite virus.
We also consider the effect of deleterious mutations in the assembly model. The phase diagram for the fully bipartite case within the assembly model is very similar to that shown in
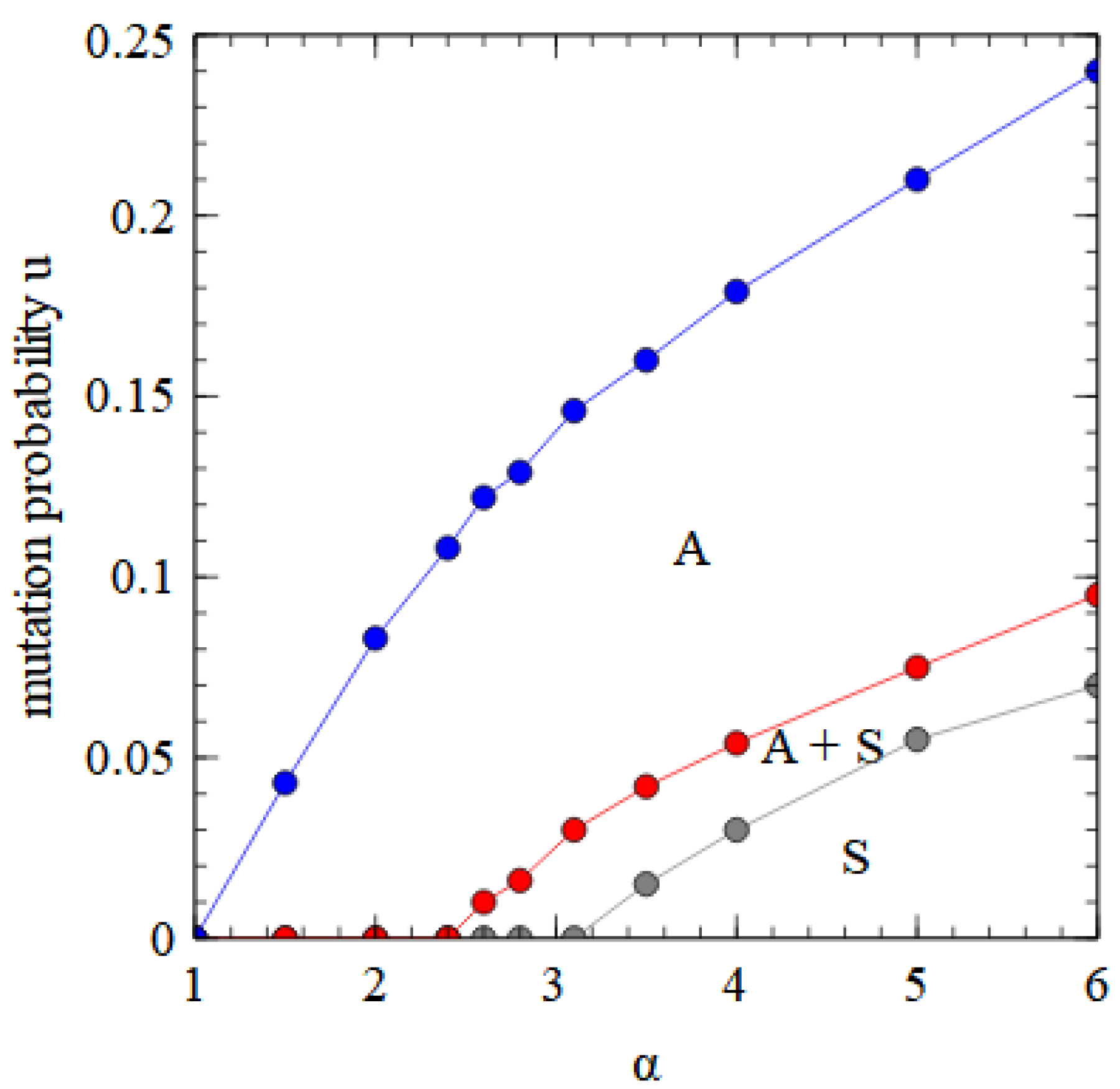
Figure 8 for the strand model. The phase diagram for the fully segmented case in the assembly model is shown in
Figure 10. The segmented virus eliminates the monopartite virus at high α and low
u.
In
Figure 8 and
Figure 10 we see that a bipartite or a segmented virus can eliminate a monopartite virus at high α and low
. As
increases, there is a transition to a state where D and E are dependent on A, and eventually to a state where only A survives. The fact that bipartite viruses are favored by high transmissibility (or high multiplicity of infection) is not surprising and has been seen in previous theories [
25]. However, the fact that bipartite viruses are favored by
low mutation rates in our models is somewhat surprising, as several previous theories [
27,
28,
29] have argued that bipartite viruses are favored by
high mutation rates and that the low fidelity of RNA replication in comparison to DNA replication provides an explanation for why bipartite viruses are more common in RNA than DNA viruses. We therefore wished to investigate why our models give qualitatively different results than those obtained from previous theories.
We will discuss our theory in comparison to that of Nee [
27]. The theories of [
28] and [
29] are similar in that bipartite viruses are favored by the low fidelity of RNA replication (however, they differ in other factors). Nee’s theory considers a complete virus C (equivalent to our A) that encodes both a coat protein gene and a polymerase and two types of incomplete sequences I (equivalent to our D and E) that encode either the coat protein or the polymerase. The lengths of C and I sequences are L and L/2. The per-base replication fidelity is
, meaning that the fidelities of replication of C and I are
and
. In our model, the fidelities are
, and
. These notations are equivalent if
Nee then writes the fitnesses of C and I as
and
, where
and
are the numbers of copies of a molecule produced per cell if replication occurs, and
is the probability that an incomplete molecule is complemented by co-infection with the other type of strand. He then goes on to say that the bipartite virus should evolve when
. However, we now argue that these formulae for
and
are oversimplified for several reasons, and that this appears to lead to incorrect conclusions.
Firstly, Nee’s theory uses single parameters for
and
. These are similar to the output numbers of viruses per cell in our models; however, we have shown that the numbers of viruses of each type produced per cell depends critically on the other types of viruses that infect the same cell (
Table 1 and
Table 2 above). The number of A strands produced is reduced substantially when either D or E is in the same cell. The number of D and E strands produced depends on whether D and E are working together as a bipartite virus or whether they are parasites in a cell with A. These factors are essential in our model, but are simply ignored in the theory of Nee [
27], and this is one reason why the conclusions of the earlier theory may have been misleading.
Secondly, the replication fidelities discussed above apply to a single round of replication from a functional sequence. However, we have assumed that there are multiple rounds of replication in a single cell. Infection by a single functional A (which we called Acp) produces a mixture of functional and mutant sequences: Acp, Ac0 and A00. Although mutant sequences cannot initiate virus replication unless they are complemented by other strands, mutant sequences produced from functional strands can continue to replicate many times in the same cell. We define the effective fidelity of A, , as the fraction of Acp strands relative to the total number of A produced within a cell, assuming that the infection begins with a functional Acp. Similarly, the effective fidelities and are the fractions of Dc and Ep relative to the total numbers of D and E, when the infections begin with functional strands.
We considered a case where , giving single replication fidelities of , and . We found that 0.479 when beginning from a single A, and that when beginning from one D and one E. The effective fidelities are much lower than the single replication fidelities. Thus, it cannot be assumed that the fitnesses and are proportional to the single replication fidelities. This is a second reason why the conclusions of the earlier theory may be unreliable.
Furthermore, the earlier theory does not clearly distinguish between deleterious mutations, which change the sequence without changing the length, and deletions, which reduce the sequence length but leave the remaining sequence intact. In our models, is the rate of deleterious mutations, not deletions. A mutation in Acp creates Ac0 or A0p, each of which have the same length as Acp. Deletions in Acp might create Dc or Ep, but these have a shorter length and are not equivalent to Ac0 and A0p. We have assumed that D and E were created originally by deletions, but are not produced continually by recurrent deletions, but that mutant A strands are created continually from functional A strands.
We note that the model of Iranzo and Manrubia [
25] includes a parameter ρ that they call “loss of segments through mutation and replication fidelity”. Despite its being referred to as mutation, it appears that this parameter represents recurrent deletions, and not deleterious point mutations. Thus, none of the previous models has considered deleterious mutations in the way we have here.