Antigen-Specific Antibody Signature Is Associated with COVID-19 Outcome
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethics
2.2. Patient Recruitment and Sampling
2.3. SARS-CoV-2 Proteins
2.4. Measurement of IgA, IgG and IgG-Subclass Response
2.5. RBD-ACE2 Competitive Assay
2.6. SARS-CoV-2 Plaque Reduction Neutralization Test
2.7. Statistics and Data Analysis
3. Results
3.1. Patient Demographics
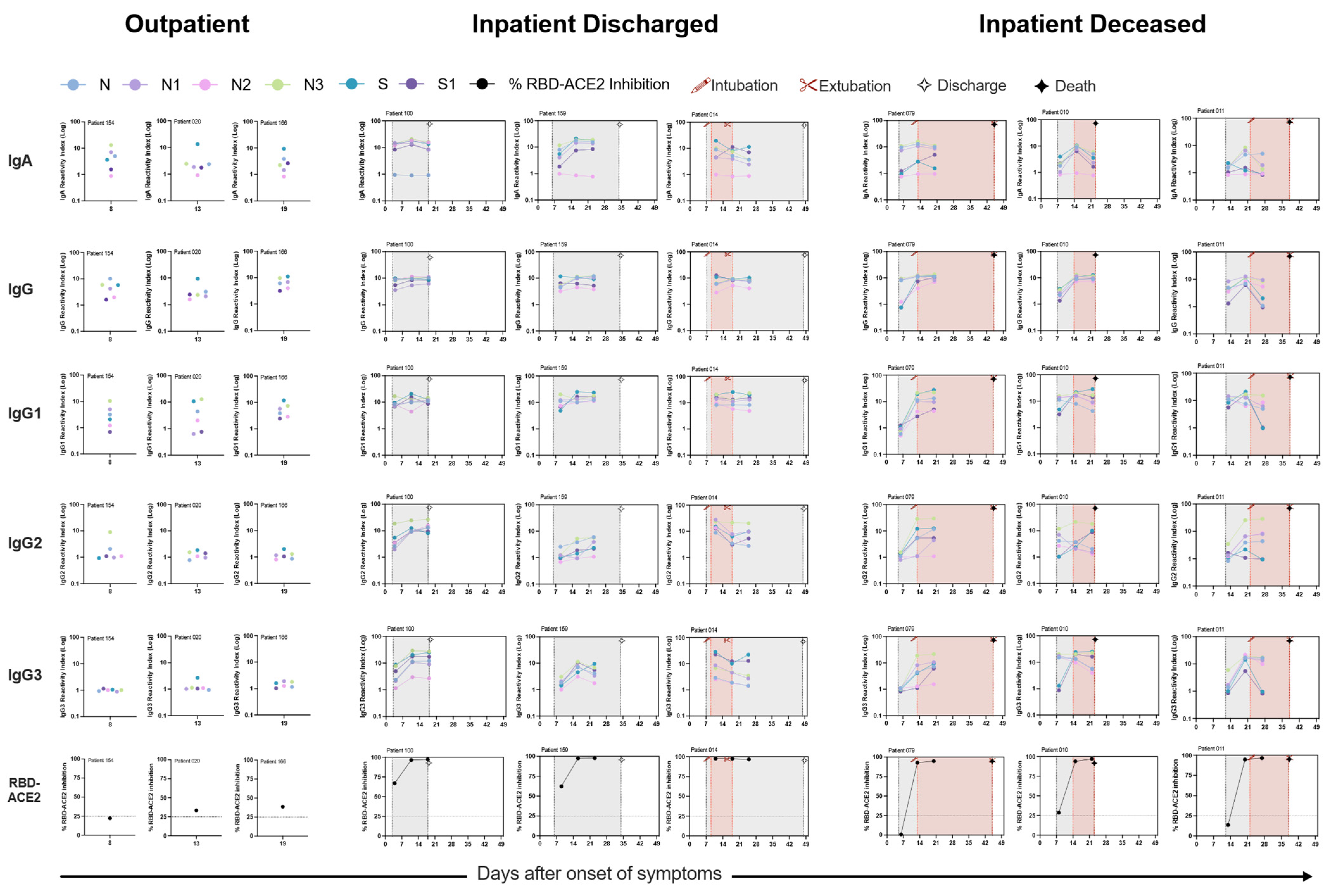
3.2. Robust Humoral Response against the Nucleocapsid and Spike Full-Length, and Truncated Proteins
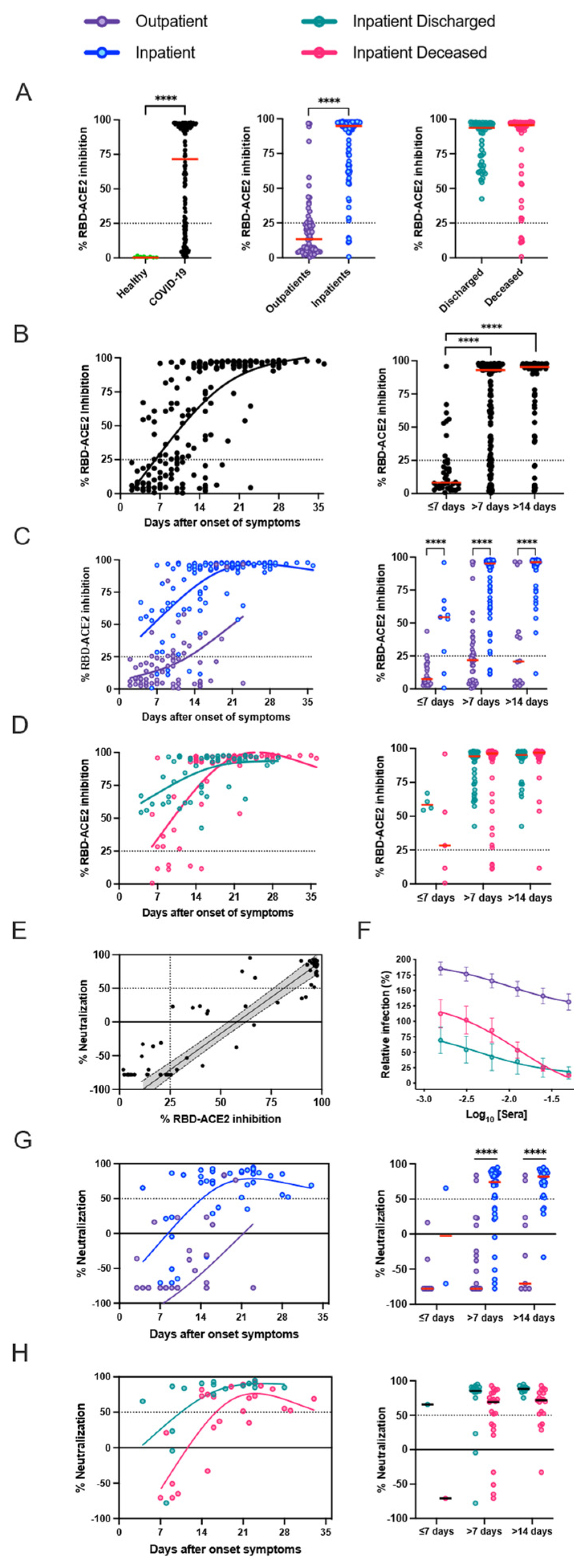
3.3. Presence of SARS-CoV-2 Neutralizing Antibodies Does Not Affect Disease Outcome
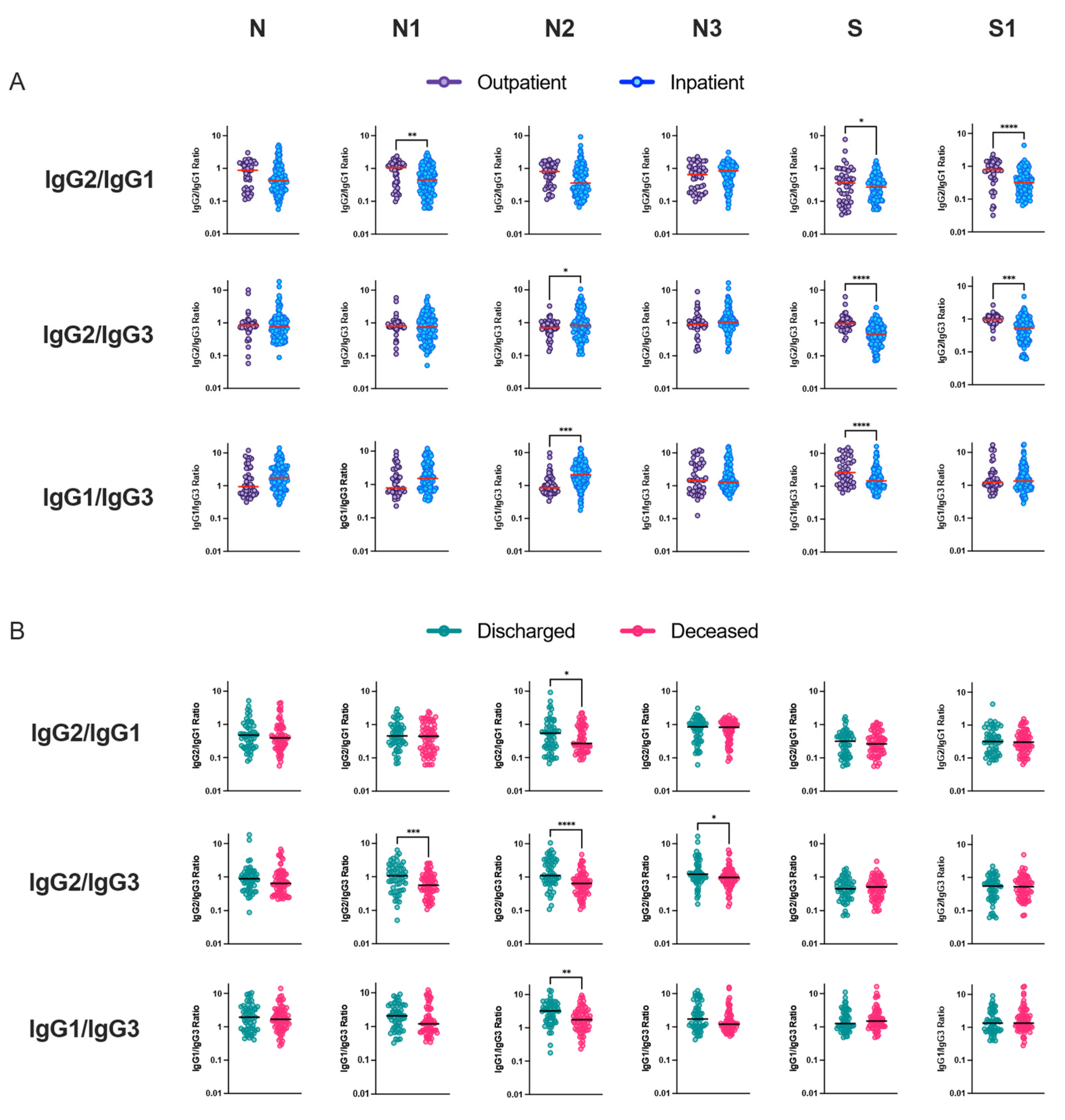
3.4. Relationship between IgG Subclass Signature and COVID-19 Outcome
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Brizzi, A.; Whittaker, C.; Servo, L.M.S.; Hawryluk, I.; Prete, C.A.; de Souza, W.M.; Aguiar, R.S.; Araujo, L.J.T.; Bastos, L.S.; Blenkinsop, A.; et al. Spatial and temporal fluctuations in COVID-19 fatality rates in Brazilian hospitals. Nat. Med. 2022, 28, 1476–1485. [Google Scholar] [CrossRef] [PubMed]
- Earle, K.A.; Ambrosino, D.M.; Fiore-Gartland, A.; Goldblatt, D.; Gilbert, P.B.; Siber, G.R.; Dull, P.; Plotkin, S.A. Evidence for antibody as a protective correlate for COVID-19 vaccines. Vaccine 2021, 39, 4423–4428. [Google Scholar] [CrossRef] [PubMed]
- Wu, F.; Zhao, S.; Yu, B.; Chen, Y.-M.; Wang, W.; Song, Z.-G.; Hu, Y.; Tao, Z.-W.; Tian, J.-H.; Pei, Y.-Y.; et al. A new coronavirus associated with human respiratory disease in China. Nature 2020, 579, 265–269. [Google Scholar] [CrossRef] [PubMed]
- Lu, R.; Zhao, X.; Li, J.; Niu, P.; Yang, B.; Wu, H.; Wang, W.; Song, H.; Huang, B.; Zhu, N.; et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: Implications for virus origins and receptor binding. Lancet 2020, 395, 565–574. [Google Scholar] [CrossRef]
- V’Kovski, P.; Kratzel, A.; Steiner, S.; Stalder, H.; Thiel, V. Coronavirus biology and replication: Implications for SARS-CoV-2. Nat. Rev. Microbiol. 2021, 19, 155–170. [Google Scholar] [CrossRef]
- Bojkova, D.; Klann, K.; Koch, B.; Widera, M.; Krause, D.; Ciesek, S.; Cinatl, J.; Münch, C. Proteomics of SARS-CoV-2-infected host cells reveals therapy targets. Nature 2020, 583, 469–472. [Google Scholar] [CrossRef]
- Jayaram, J.; Youn, S.; Collisson, E.W. The virion N protein of infectious bronchitis virus is more phosphorylated than the N protein from infected cell lysates. Virology 2005, 339, 127–135. [Google Scholar] [CrossRef]
- Poland, G.A.; Ovsyannikova, I.G.; Kennedy, R.B. SARS-CoV-2 immunity: Review and applications to phase 3 vaccine candidates. Lancet 2020, 396, 1595–1606. [Google Scholar] [CrossRef]
- Deeks, J.J.; Dinnes, J.; Takwoingi, Y.; Davenport, C.; Spijker, R.; Taylor-Phillips, S.; Adriano, A.; Beese, S.; Dretzke, J.; di Ruffano, L.F.; et al. Antibody tests for identification of current and past infection with SARS-CoV-2. Cochrane Database Syst. Rev. 2020, 6, CD013652. [Google Scholar] [CrossRef]
- Fazolo, T.; Lima, K.; Fontoura, J.C.; de Souza, P.O.; Hilario, G.; Zorzetto, R.; Júnior, L.R.; Pscheidt, V.M.; Neto, J.d.C.F.; Haubert, A.F.; et al. Pediatric COVID-19 patients in South Brazil show abundant viral mRNA and strong specific anti-viral responses. Nat. Commun. 2021, 12, 6844. [Google Scholar] [CrossRef]
- Borba, M.G.S.; Val, F.F.A.; Sampaio, V.S.; Alexandre, M.A.A.; Melo, G.C.; Brito, M.; Mourao, M.P.G.; Brito-Sousa, J.D.; Baia-da-Silva, D.; Guerra, M.V.F.; et al. Effect of High vs Low Doses of Chloroquine Diphosphate as Adjunctive Therapy for Patients Hospitalized With Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) Infection: A Randomized Clinical Trial. JAMA Netw. Open 2020, 3, e208857. [Google Scholar] [CrossRef] [PubMed]
- Lalwani, P.; Salgado, B.B.; Pereira Filho, I.V.; Silva, D.S.S.d.; Morais, T.B.d.N.d.; Jordão, M.F.; Barbosa, A.R.C.; Cordeiro, I.B.; Neto, J.N.d.S.; Assunção, E.N.d.; et al. SARS-CoV-2 Seroprevalence and Associated Factors in Manaus, Brazil: Baseline Results from the DETECTCoV-19 Cohort Study. Int. J. Infect. Dis. 2021, 110, 141–150. [Google Scholar] [CrossRef] [PubMed]
- Alvim, R.G.F.; Lima, T.M.; Rodrigues, D.A.S.; Marsili, F.F.; Bozza, V.B.T.; Higa, L.M.; Monteiro, F.L.; Leitao, I.C.; Carvalho, R.S.; Galliez, R.M.; et al. From a recombinant key antigen to an accurate, affordable serological test: Lessons learnt from COVID-19 for future pandemics. Biochem. Eng. J. 2022, 186, 108537. [Google Scholar] [CrossRef] [PubMed]
- Souza, W.M.; Amorim, M.R.; Sesti-Costa, R.; Coimbra, L.D.; Brunetti, N.S.; Toledo-Teixeira, D.A.; de Souza, G.F.; Muraro, S.P.; Parise, P.L.; Barbosa, P.P.; et al. Neutralisation of SARS-CoV-2 lineage P.1 by antibodies elicited through natural SARS-CoV-2 infection or vaccination with an inactivated SARS-CoV-2 vaccine: An immunological study. Lancet Microbe 2021, 2, e527–e535. [Google Scholar] [CrossRef]
- Long, Q.-X.; Liu, B.-Z.; Deng, H.-J.; Wu, G.-C.; Deng, K.; Chen, Y.-K.; Liao, P.; Qiu, J.-F.; Lin, Y.; Cai, X.-F.; et al. Antibody responses to SARS-CoV-2 in patients with COVID-19. Nat. Med. 2020, 26, 845–848. [Google Scholar] [CrossRef]
- Guthmiller, J.J.; Stovicek, O.; Wang, J.; Changrob, S.; Li, L.; Halfmann, P.; Zheng, N.-Y.; Utset, H.; Stamper, C.T.; Dugan, H.L.; et al. SARS-CoV-2 Infection Severity Is Linked to Superior Humoral Immunity against the Spike. mBio 2021, 12, e02940-20. [Google Scholar] [CrossRef]
- Röltgen, K.; Powell, A.E.; Wirz, O.F.; Stevens, B.A.; Hogan, C.A.; Najeeb, J.; Hunter, M.; Wang, H.; Sahoo, M.K.; Huang, C.; et al. Defining the features and duration of antibody responses to SARS-CoV-2 infection associated with disease severity and outcome. Sci. Immunol. 2020, 5, abe0240. [Google Scholar] [CrossRef]
- Okba, N.M.; Raj, V.S.; Widjaja, I.; GeurtsvanKessel, C.H.; de Bruin, E.; Chandler, F.D.; Park, W.B.; Kim, N.-J.; Farag, E.A.; Al-Hajri, M.; et al. Sensitive and Specific Detection of Low-Level Antibody Responses in Mild Middle East Respiratory Syndrome Coronavirus Infections. Emerg. Infect. Dis. 2019, 25, 1868–1877. [Google Scholar] [CrossRef]
- Alshukairi, A.N.; Khalid, I.; Ahmed, W.A.; Dada, A.M.; Bayumi, D.T.; Malic, L.S.; Althawadi, S.; Ignacio, K.; Alsalmi, H.S.; Al-Abdely, H.M.; et al. Antibody Response and Disease Severity in Healthcare Worker MERS Survivors. Emerg. Infect. Dis. 2016, 22, 1113–1115. [Google Scholar] [CrossRef]
- Drosten, C.; Meyer, B.; Müller, M.A.; Corman, V.M.; Al-Masri, M.; Hossain, R.; Madani, H.; Sieberg, A.; Bosch, B.J.; Lattwein, E.; et al. Transmission of MERS-Coronavirus in Household Contacts. N. Engl. J. Med. 2014, 371, 828–835. [Google Scholar] [CrossRef]
- Nilsson, A.C.; Holm, D.K.; Justesen, U.S.; Gorm-Jensen, T.; Andersen, N.S.; Øvrehus, A.; Johansen, I.S.; Michelsen, J.; Sprogøe, U.; Lillevang, S.T. Comparison of six commercially available SARS-CoV-2 antibody assays—Choice of assay depends on intended use. Int. J. Infect. Dis. 2020, 103, 381–388. [Google Scholar] [CrossRef] [PubMed]
- Anna, F.; Goyard, S.; Lalanne, A.I.; Nevo, F.; Gransagne, M.; Souque, P.; Louis, D.; Gillon, V.; Turbiez, I.; Bidard, F.C.; et al. High seroprevalence but short-lived immune response to SARS-CoV-2 infection in Paris. Eur. J. Immunol. 2021, 51, 180–190. [Google Scholar] [CrossRef] [PubMed]
- Moncunill, G.; Mayor, A.; Santano, R.; Jiménez, A.; Vidal, M.; Tortajada, M.; Sanz, S.; Méndez, S.; Llupià, A.; Aguilar, R.; et al. SARS-CoV-2 Seroprevalence and Antibody Kinetics Among Health Care Workers in a Spanish Hospital After 3 Months of Follow-up. J. Infect. Dis. 2020, 223, 62–71. [Google Scholar] [CrossRef] [PubMed]
- Dobaño, C.; Santano, R.; Jiménez, A.; Vidal, M.; Chi, J.; Melero, N.R.; Popovic, M.; López-Aladid, R.; Fernández-Barat, L.; Tortajada, M.; et al. Immunogenicity and crossreactivity of antibodies to the nucleocapsid protein of SARS-CoV-2: Utility and limitations in seroprevalence and immunity studies. Transl. Res. 2021, 232, 60–74. [Google Scholar] [CrossRef]
- Atyeo, C.; Fischinger, S.; Zohar, T.; Slein, M.D.; Burke, J.; Loos, C.; McCulloch, D.J.; Newman, K.L.; Wolf, C.; Yu, J.; et al. Distinct Early Serological Signatures Track with SARS-CoV-2 Survival. Immunity 2020, 53, 524–532.e4. [Google Scholar] [CrossRef]
- Iles, J.K.; Zmuidinaite, R.; Sadee, C.; Gardiner, A.; Lacey, J.; Harding, S.; Wallis, G.; Patel, R.; Roblett, D.; Heeney, J.; et al. Determination of IgG1 and IgG3 SARS-CoV-2 Spike Protein and Nucleocapsid Binding—Who Is Binding Who and Why? Int. J. Mol. Sci. 2022, 23, 6050. [Google Scholar] [CrossRef]
- Yates, J.L.; Ehrbar, D.J.; Hunt, D.T.; Girardin, R.C.; Dupuis, A.P.; Payne, A.F.; Sowizral, M.; Varney, S.; Kulas, K.E.; Demarest, V.L.; et al. Serological analysis reveals an imbalanced IgG subclass composition associated with COVID-19 disease severity. Cell Rep. Med. 2021, 2, 100329. [Google Scholar] [CrossRef]
- Liu, Z.; Long, W.; Tu, M.; Chen, S.; Huang, Y.; Wang, S.; Zhou, W.; Chen, D.; Zhou, L.; Wang, M.; et al. Lymphocyte subset (CD4+, CD8+) counts reflect the severity of infection and predict the clinical outcomes in patients with COVID-19. J. Infect. 2020, 81, 318–356. [Google Scholar] [CrossRef]
- Huang, W.; Berube, J.; McNamara, M.; Saksena, S.; Hartman, M.; Arshad, T.; Bornheimer, S.J.; O’Gorman, M. Lymphocyte Subset Counts in COVID-19 Patients: A Meta-Analysis. Cytom. Part A 2020, 97, 772–776. [Google Scholar] [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Salgado, B.B.; Jordão, M.F.; de Morais, T.B.d.N.; da Silva, D.S.S.; Pereira Filho, I.V.; Salgado Sobrinho, W.B.; Carvalho, N.O.; dos Santos, R.O.; Forato, J.; Barbosa, P.P.; et al. Antigen-Specific Antibody Signature Is Associated with COVID-19 Outcome. Viruses 2023, 15, 1018. https://doi.org/10.3390/v15041018
Salgado BB, Jordão MF, de Morais TBdN, da Silva DSS, Pereira Filho IV, Salgado Sobrinho WB, Carvalho NO, dos Santos RO, Forato J, Barbosa PP, et al. Antigen-Specific Antibody Signature Is Associated with COVID-19 Outcome. Viruses. 2023; 15(4):1018. https://doi.org/10.3390/v15041018
Chicago/Turabian StyleSalgado, Bárbara Batista, Maele Ferreira Jordão, Thiago Barros do Nascimento de Morais, Danielle Severino Sena da Silva, Ivanildo Vieira Pereira Filho, Wlademir Braga Salgado Sobrinho, Nani Oliveira Carvalho, Rafaella Oliveira dos Santos, Julia Forato, Priscilla Paschoal Barbosa, and et al. 2023. "Antigen-Specific Antibody Signature Is Associated with COVID-19 Outcome" Viruses 15, no. 4: 1018. https://doi.org/10.3390/v15041018
APA StyleSalgado, B. B., Jordão, M. F., de Morais, T. B. d. N., da Silva, D. S. S., Pereira Filho, I. V., Salgado Sobrinho, W. B., Carvalho, N. O., dos Santos, R. O., Forato, J., Barbosa, P. P., Toledo-Teixeira, D. A., Pinto, K. R., Correia, I. S., Cordeiro, I. B., Souza Neto, J. N. d., Assunção, E. N. d., Val, F. F. A., Melo, G. C., Sampaio, V. d. S., ... Lalwani, P. (2023). Antigen-Specific Antibody Signature Is Associated with COVID-19 Outcome. Viruses, 15(4), 1018. https://doi.org/10.3390/v15041018










