Timeline of SARS-CoV-2 Spread in Italy: Results from an Independent Serological Retesting
Abstract
:1. Introduction
2. Materials and Methods
3. Results
3.1. Retesting of Lung Cancer Screening Participants
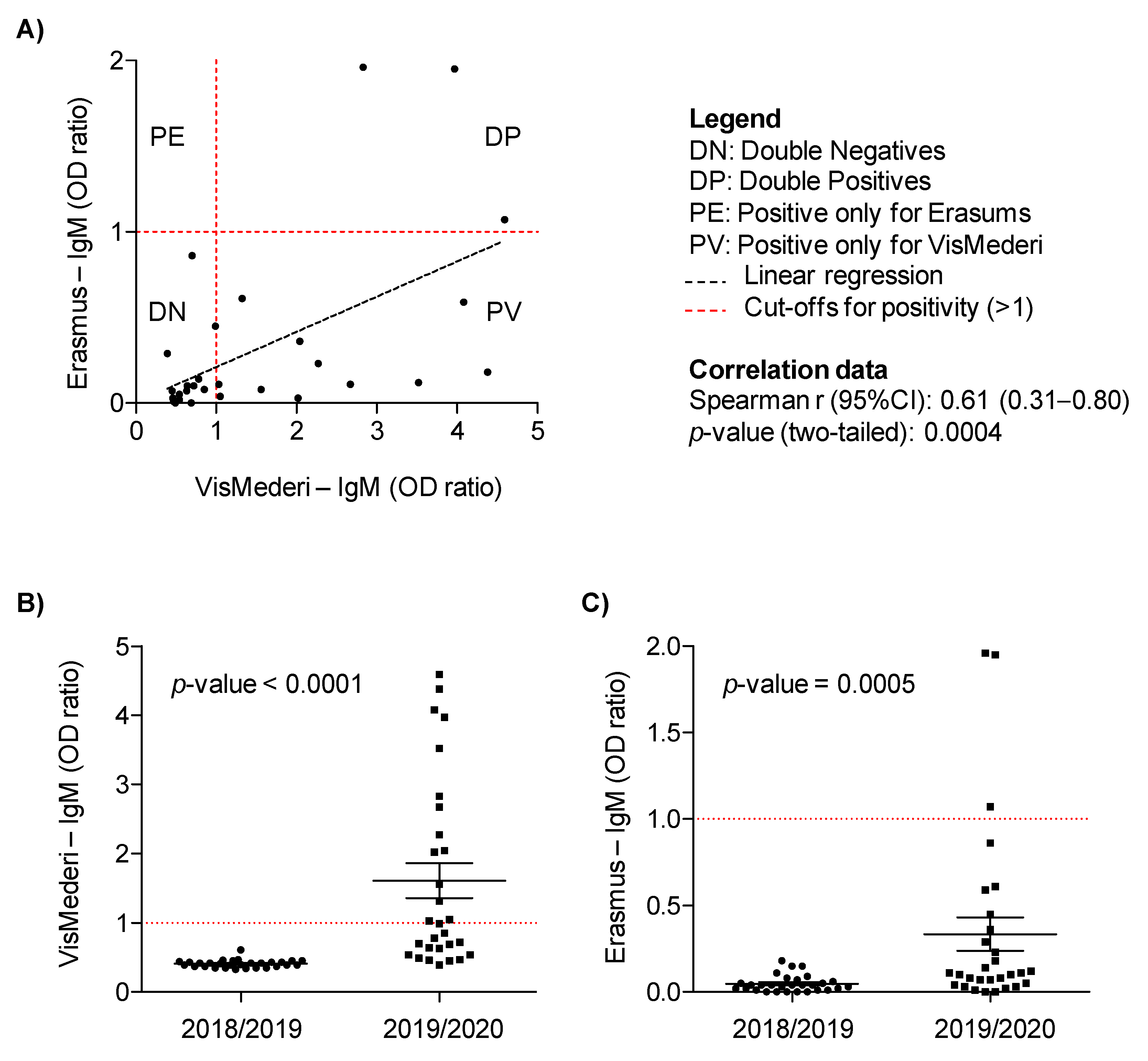
3.2. Analysis of COVID-19 Control Samples
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Mallapaty, S. Where did COVID come from? Five mysteries that remain. Nature 2021, 591, 188–189. [Google Scholar] [CrossRef]
- Wu, J.T.; Leung, K.; Lam, T.T.Y.; Ni, M.Y.; Wong, C.K.H.; Peiris, J.S.M.; Leung, G.M. Nowcasting epidemics of novel pathogens: Lessons from COVID-19. Nat. Med. 2021, 27, 388–395. [Google Scholar] [CrossRef] [PubMed]
- La Rosa, G.; Mancini, P.; Bonanno Ferraro, G.; Veneri, C.; Iaconelli, M.; Bonadonna, L.; Lucentini, L.; Suffredini, E. SARS-CoV-2 has been circulating in northern Italy since December 2019: Evidence from environmental monitoring. Sci. Total Environ. 2021, 750, 141711. [Google Scholar] [CrossRef]
- Chavarria-Miró, G.; Anfruns-Estrada, E.; Martínez-Velázquez, A.; Vázquez-Portero, M.; Guix, S.; Paraira, M.; Galofré, B.; Sánchez, G.; Pintó, R.M.; Bosch, A. Time-evolution of SARS-CoV-2 in wastewater during the first pandemic wave of COVID-19 in the metropolitan area of Barcelona. Appl. Environ. Microbiol. 2021, 87, e02750-20. [Google Scholar] [CrossRef] [PubMed]
- Deslandes, A.; Berti, V.; Tandjaoui-Lambotte, Y.; Alloui, C.; Carbonnelle, E.; Zahar, J.R.; Brichler, S.; Cohen, Y. SARS-CoV-2 was already spreading in France in late December 2019. Int. J. Antimicrob. Agents 2020, 55, 106006. [Google Scholar] [CrossRef]
- Amendola, A.; Bianchi, S.; Gori, M.; Colzani, D.; Canuti, M.; Borghi, E.; Raviglione, M.C.; Zuccotti, G.V.; Tanzi, E. Evidence of SARS-CoV-2 RNA in an oropharyngeal swab specimen, Milan, Italy, early December 2019. Emerg. Infect. Dis. 2021, 27, 648–650. [Google Scholar] [CrossRef] [PubMed]
- Gianotti, R.; Barberis, M.; Fellegara, G.; Galván-Casas, C.; Gianotti, E. COVID-19-related dermatosis in November 2019: Could this case be Italy’s patient zero? Br. J. Dermatol. 2021, 84, 970–971. [Google Scholar] [CrossRef]
- Basavaraju, S.V.; Patton, M.E.; Grimm, K.; Rasheed, M.A.U.; Lester, S.; Mills, L.; Stumpf, M.; Freeman, B.; Tamin, A.; Harcourt, J.; et al. Serologic Testing of US Blood Donations to Identify Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2)–Reactive Antibodies: December 2019–January 2020. Clin. Infect. Dis. 2020, 72, e1004–e1009. [Google Scholar] [CrossRef]
- Carrat, F.; Figoni, J.; Henny, J.; Desenclos, J.C.; Kab, S.; de Lamballerie, X.; Zins, M. Evidence of early circulation of SARS-CoV-2 in France: Findings from the population-based “CONSTANCES” cohort. Eur. J. Epidemiol. 2021, 36, 219–222. [Google Scholar] [CrossRef]
- Apolone, G.; Montomoli, E.; Manenti, A.; Boeri, M.; Sabia, F.; Hyseni, I.; Mazzini, L.; Martinuzzi, D.; Cantone, L.; Milanese, G.; et al. Unexpected detection of SARS-CoV-2 antibodies in the prepandemic period in Italy. Tumori J. 2020, 107, 446–451. [Google Scholar] [CrossRef]
- Andreano, E.; Nicastri, E.; Paciello, I.; Pileri, P.; Manganaro, N.; Piccini, G.; Manenti, A.; Pantano, E.; Kabanova, A.; Troisi, M.; et al. Extremely potent human monoclonal antibodies from COVID-19 convalescent patients. Cell 2021, 184, 1821–1835. [Google Scholar] [CrossRef]
- Mazzini, L.; Martinuzzi, D.; Hyseni, I.; Benincasa, L.; Molesti, E.; Casa, E.; Lapini, G.; Piu, P.; Trombetta, C.M.; Marchi, S.; et al. Comparative analyses of SARS-CoV-2 binding (IgG, IgM, IgA) and neutralizing antibodies from human serum samples. J. Immunol. Methods 2021, 489, 112937. [Google Scholar] [CrossRef] [PubMed]
- GeurtsvanKessel, C.H.; Okba, N.M.A.; Igloi, Z.; Bogers, S.; Embregts, C.W.E.; Laksono, B.M.; Leijten, L.; Rokx, C.; Rijnders, B.; Rahamat-Langendoen, J.; et al. An evaluation of COVID-19 serological assays informs future diagnostics and exposure assessment. Nat. Commun. 2020, 11, 3436. [Google Scholar] [CrossRef]
- Van Tol, S.; Mögling, R.; Li, W.; Godeke, G.J.; Swart, A.; Bergmans, B.; Brandenburg, A.; Kremer, K.; Murk, J.L.; van Beek, J.; et al. Accurate serology for SARS-CoV-2 and common human coronaviruses using a multiplex approach. Emerg. Microbes Infect. 2020, 9, 1965–1973. [Google Scholar] [CrossRef]
- Milani, G.P.; Dioni, L.; Favero, C.; Cantone, L.; Macchi, C.; Delbue, S.; Bonzini, M.; Montomoli, E.; Bollati, V.; Albetti, B.; et al. Serological follow-up of SARS-CoV-2 asymptomatic subjects. Sci. Rep. 2020, 10, 1–7. [Google Scholar] [CrossRef]
- Piccoli, L.; Park, Y.J.; Tortorici, M.A.; Czudnochowski, N.; Walls, A.C.; Beltramello, M.; Silacci-Fregni, C.; Pinto, D.; Rosen, L.E.; Bowen, J.E.; et al. Mapping Neutralizing and Immunodominant Sites on the SARS-CoV-2 Spike Receptor-Binding Domain by Structure-Guided High-Resolution Serology. Cell 2020, 183, 1024–1042. [Google Scholar] [CrossRef] [PubMed]
- Ding, S.; Laumaea, A.; Benlarbi, M.; Beaudoin-Bussières, G.; Gasser, R.; Medjahed, H.; Pancera, M.; Stamatatos, L.; McGuire, A.; Bazin, R.; et al. Antibody Binding to SARS-CoV-2 S Glycoprotein Correlates with but Does Not Predict Neutralization. Viruses 2020, 12, 1214. [Google Scholar] [CrossRef]
- Criscuolo, E.; Diotti, R.A.; Strollo, M.; Rolla, S.; Ambrosi, A.; Locatelli, M.; Burioni, R.; Mancini, N.; Clementi, M.; Clementi, N. Weak correlation between antibody titers and neutralizing activity in sera from SARS-CoV-2 infected subjects. J. Med. Virol. 2021, 93, 2160–2167. [Google Scholar] [CrossRef]
- Alteri, C.; Cento, V.; Piralla, A.; Costabile, V.; Tallarita, M.; Colagrossi, L.; Renica, S.; Giardina, F.; Novazzi, F.; Gaiarsa, S.; et al. Genomic epidemiology of SARS-CoV-2 reveals multiple lineages and early spread of SARS-CoV-2 infections in Lombardy, Italy. Nat. Commun. 2021, 12, 434. [Google Scholar] [CrossRef] [PubMed]
- Lai, A.; Bergna, A.; Acciarri, C.; Galli, M.; Zehender, G. Early phylogenetic estimate of the effective reproduction number of SARS-CoV-2. J. Med. Virol. 2020, 92, 675–679. [Google Scholar] [CrossRef] [Green Version]
- Giardina, F.; Galli, C.; Pellegrinelli, L.; Paolucci, S.; Pariani, E.; Piralla, A.; Baldanti, F. No evidence of SARS-CoV-2 circulation in the framework of influenza surveillance between October 2019 and February 2020 in Lombardy, Italy. Travel Med. Infect. Dis. 2021, 40, 102002. [Google Scholar] [CrossRef]
- Percivalle, E.; Cambiè, G.; Cassaniti, I.; Nepita, E.V.; Maserati, R.; Ferrari, A.; Di Martino, R.; Isernia, P.; Mojoli, F.; Bruno, R.; et al. Prevalence of SARS-CoV-2 specific neutralising antibodies in blood donors from the Lodi Red Zone in Lombardy, Italy, as at 06 April 2020. Eurosurveillance 2020, 25, 2001031. [Google Scholar] [CrossRef]
- Amendola, A.; Canuti, M.; Bianchi, S.; Kumar, S.; Fappani, C.; Gori, M.; Colzani, D.; Pond, S.L.; Miura, S.; Baggieri, M.; et al. Molecular Evidence for SARS-CoV-2 in Samples Collected from Patients With Morbilliform Eruptions Since Late Summer 2019 in Lombardy, Northern Italy. SSRN Electron. J. 2021, 27p. [Google Scholar] [CrossRef]
- Cereda, D.; Manica, M.; Tirani, M.; Rovida, F.; Demicheli, V.; Ajelli, M.; Poletti, P.; Trentini, F.; Guzzetta, G.; Marziano, V.; et al. The early phase of the COVID-19 epidemic in Lombardy, Italy. Epidemics 2021, 37, 100528. [Google Scholar] [CrossRef]
- Chiara, M.; Horner, D.S.; Gissi, C.; Pesole, G. Comparative Genomics Reveals Early Emergence and Biased Spatiotemporal Distribution of SARS-CoV-2. Mol. Biol. Evol. 2021, 38, 2547–2565. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Tao, Q.; Weaver, S.; Sanderford, M.; Caraballo-Ortiz, M.A.; Sharma, S.; Pond, S.L.K.; Miura, S. An Evolutionary Portrait of the Progenitor SARS-CoV-2 and Its Dominant Offshoots in COVID-19 Pandemic. Mol. Biol. Evol. 2021, 38, 3046–3059. [Google Scholar] [CrossRef] [PubMed]
- Petti, S. Undetected and relatively sustained SARS-CoV-2 circulation worldwide during the year 2019. Clin. Infect. Dis. 2021, ciab727. [Google Scholar] [CrossRef]
- WHO-Convened Global Study of Origins of SARS-CoV-2: China Part. 2021. Available online: https://www.who.int/publications/i/item/who-convened-global-study-of-origins-of-sars-cov-2-china-part (accessed on 23 December 2021).
| Date of Collection | Italian Region | IgG | IgM | Neutralization Assay | |||
|---|---|---|---|---|---|---|---|
| VisMederi | Erasmus | VisMederi | Erasmus | MN VisMederi | PRNT Erasmus | ||
| 23 July 2019 | N/A | Neg | Neg | Neg | Neg | N/A | N/A |
| 24 July 2019 | N/A | Neg | Neg | Neg | Neg | N/A | N/A |
| 3 September 2019 | Lombardy | Neg | Neg | Neg | Neg | N/A | N/A |
| 3 September 2019 | Veneto | Neg | Neg | Pos (RBD) | Neg | Neg | N/A |
| 5 September 2019 | Liguria | Neg | Neg | Borderline | Neg | Neg | N/A |
| 10 September 2019 | N/A | Neg | Neg | Neg | Neg | N/A | N/A |
| 12 September 2019 | Lombardy | Neg | Neg | Neg | Neg | N/A | N/A |
| 17 September 2019 | N/A | Neg | Neg | Neg | Neg | N/A | N/A |
| 2 October 2019 | Piedmont | Pos (RBD) | Neg | Neg | Neg | Neg | Neg |
| 7 October 2019 | Valle’d’Aosta | Neg | Neg | Pos (RBD) | Neg | Pos | N/A |
| 7 October 2019 | Liguria | Pos (RBD) | Neg | Neg | Neg | Neg | Neg |
| 7 October 2019 | Lombardy | Neg | Neg | Pos (RBD) | Neg | Pos | N/A |
| 8 October 2019 | Lombardy | Neg | Neg | Pos (RBD) | Neg | Pos | N/A |
| 10 October 2019 | Lombardy | Pos (RBD) | Neg | Neg | Neg | Neg | Neg |
| 10 October 2019 | Lombardy | Neg | Neg | Pos (RBD) | Pos (RBD) | Neg | Neg |
| 15 October 2019 | Lombardy | Neg | Neg | Pos (RBD) | Neg | Neg | N/A |
| 15 October 2019 | N/A | Neg | Neg | Neg | Neg | N/A | Neg |
| 21 October 2019 | Tuscany | Neg | Neg | Pos (RBD) | Neg | Pos | N/A |
| 22 October 2019 | Lombardy | Neg | Neg | Neg | Neg | N/A | N/A |
| 7 November 2019 | Lombardy | Neg | Neg | Borderline | Neg | Pos | N/A |
| 11 November 2019 | Lombardy | Neg | Pos (S1) | Pos (RBD) | Borderline | Neg | Neg |
| 26 November 2019 | Lazio | Pos (RBD) | Neg | Neg | Neg | Neg | Neg |
| 13 December 2019 | Lombardy | Pos (RBD) | Neg | Borderline | Neg | Neg | Neg |
| 17 December 2019 | Campania | Pos (RBD) | Pos (S1) | Pos (RBD) | Neg | Neg | Neg |
| 18 December 2019 | Liguria | Neg | Neg | Pos (RBD) | Neg | Neg | N/A |
| 28 January 2020 | Lombardy | Pos (RBD) | Pos (S1,NP) | Neg | Neg | Neg | Neg |
| 5 February 2020 | Lazio | Neg | Neg | Pos (RBD) | Pos (RBD) | Pos | Pos |
| 12 February 2020 | Lombardy | Neg | Neg | Pos (RBD) | Neg | Neg | Neg |
| 17 February 2020 | Piedmont | Neg | Neg | Pos (RBD) | Neg | Neg | N/A |
| Sample | Center of Collection | IgG | IgM | Neutralization | |||
|---|---|---|---|---|---|---|---|
| VisMederi | Erasmus | VisMederi | Erasmus | MN VisMederi | PRNT Erasmus | ||
| Negative control | NIBSC | Neg | Neg | Neg | Neg | Neg | N/A |
| Negative control | NIBSC | Neg | Neg | Neg | Neg | Neg | N/A |
| Convalescent symptomatic | NIBSC | Pos (RBD) | Pos (S1, ecto, NP) | Neg | N/A | Pos | Pos |
| Convalescent symptomatic | BioIVT | Pos (RBD) | Pos (S1, ecto, NP) | Pos (RBD) | Pos (RBD) | Pos | Pos |
| Convalescent symptomatic | BioIVT | Pos (RBD) | Pos (S1, ecto, NP) | Pos (RBD) | Neg | Pos | Pos |
| Convalescent asymptomatic | UNISI | Pos (RBD) | Neg | Neg | Neg | Neg | N/A |
| Convalescent asymptomatic | UNIMI | Pos (RBD) | Neg | Neg | N/A | Neg | Neg |
| Convalescent asymptomatic | UNIMI | Pos (RBD) | Neg | Neg | Neg | Neg | N/A |
| Convalescent asymptomatic | UNIMI | Pos (RBD) | Neg | Neg | Neg | Neg | N/A |
| Convalescent asymptomatic | UNIMI | Pos (RBD) | Neg | Neg | Neg | Neg | N/A |
| Convalescent asymptomatic | UNIMI | Pos (RBD) | Neg | Neg | N/A | Neg | Neg |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Montomoli, E.; Apolone, G.; Manenti, A.; Boeri, M.; Suatoni, P.; Sabia, F.; Marchianò, A.; Bollati, V.; Pastorino, U.; Sozzi, G. Timeline of SARS-CoV-2 Spread in Italy: Results from an Independent Serological Retesting. Viruses 2022, 14, 61. https://doi.org/10.3390/v14010061
Montomoli E, Apolone G, Manenti A, Boeri M, Suatoni P, Sabia F, Marchianò A, Bollati V, Pastorino U, Sozzi G. Timeline of SARS-CoV-2 Spread in Italy: Results from an Independent Serological Retesting. Viruses. 2022; 14(1):61. https://doi.org/10.3390/v14010061
Chicago/Turabian StyleMontomoli, Emanuele, Giovanni Apolone, Alessandro Manenti, Mattia Boeri, Paola Suatoni, Federica Sabia, Alfonso Marchianò, Valentina Bollati, Ugo Pastorino, and Gabriella Sozzi. 2022. "Timeline of SARS-CoV-2 Spread in Italy: Results from an Independent Serological Retesting" Viruses 14, no. 1: 61. https://doi.org/10.3390/v14010061
APA StyleMontomoli, E., Apolone, G., Manenti, A., Boeri, M., Suatoni, P., Sabia, F., Marchianò, A., Bollati, V., Pastorino, U., & Sozzi, G. (2022). Timeline of SARS-CoV-2 Spread in Italy: Results from an Independent Serological Retesting. Viruses, 14(1), 61. https://doi.org/10.3390/v14010061







