Does the Zinc Finger Antiviral Protein (ZAP) Shape the Evolution of Herpesvirus Genomes?
Abstract
1. Introduction
2. Nucleotide and Dinucleotide Bias
3. Zinc Finger Antiviral Protein (ZAP)
4. CpG Profiles of Herpesviruses
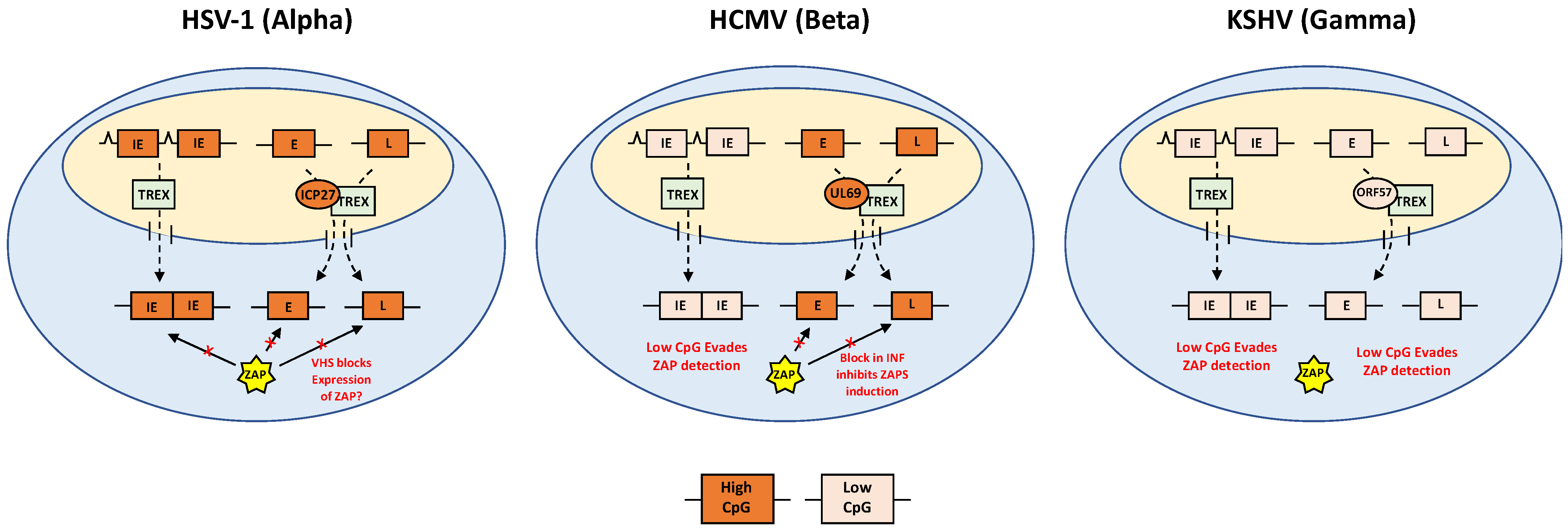
5. Evasion of ZAP by Alpha-Herpesviruses
6. Evasion of ZAP by Beta-Herpesviruses
7. Evasion of ZAP by Gamma-Herpesviruses
8. CpG Dinucleotide Bias, Gene Expression Efficiency and Splicing
9. Enhancement of Unspliced Viral Transcripts by a Family of Herpesvirus RNA Binding Proteins
10. Concluding Remarks
Author Contributions
Funding
Conflicts of Interest
References
- Plotkin, J.B.; Kudla, G. Synonymous but not the same: The causes and consequences of codon bias. Nat. Rev. Genet. 2011, 12, 32–42. [Google Scholar] [CrossRef]
- Gaunt, E.R.; Digard, P. Compositional biases in RNA viruses: Causes, consequences and applications. Wiley Interdiscip. Rev. RNA 2021, e1679. [Google Scholar] [CrossRef]
- Knipe, D.M.; Howley, P. Fields Virology; Wolters Kluwer Health: Philadelphia, PA, USA, 2013. [Google Scholar]
- Gonzalez-Perez, A.C.; Stempel, M.; Chan, B.; Brinkmann, M.M. One Step Ahead: Herpesviruses Light the Way to Understanding Interferon-Stimulated Genes (ISGs). Front. Microbiol. 2020, 11, 124. [Google Scholar] [CrossRef]
- Davison, A.J. Evolution of the herpesviruses. Vet. Microbiol. 2002, 86, 69–88. [Google Scholar] [CrossRef]
- Sausen, D.G.; Reed, K.M.; Bhutta, M.S.; Gallo, E.S.; Borenstein, R. Evasion of the Host Immune Response by Betaherpesviruses. Int. J. Mol. Sci. 2021, 22, 7503. [Google Scholar] [CrossRef] [PubMed]
- Carpentier, K.S.; Geballe, A.P. An Evolutionary View of the Arms Race between Protein Kinase R and Large DNA Viruses. J. Virol. 2016, 90, 3280–3283. [Google Scholar] [CrossRef] [PubMed]
- Child, S.J.; Greninger, A.L.; Geballe, A.P. Rapid adaptation to human protein kinase R by a unique genomic rearrangement in rhesus cytomegalovirus. PLoS Pathog. 2021, 17, e1009088. [Google Scholar] [CrossRef]
- Lander, E.S.; Linton, L.M.; Birren, B.; Nusbaum, C.; Zody, M.C.; Baldwin, J.; Devon, K.; Dewar, K.; Doyle, M.; FitzHugh, W.; et al. Initial sequencing and analysis of the human genome. Nature 2001, 409, 860–921. [Google Scholar] [CrossRef]
- Brown, J.C. High G+C Content of Herpes Simplex Virus DNA: Proposed Role in Protection Against Retrotransposon Insertion. Open Biochem. J. 2007, 1, 33–42. [Google Scholar] [CrossRef]
- Honess, R.W. Herpes simplex and ‘the herpes complex’: Diverse observations and a unifying hypothesis. The eighth Fleming lecture. J. Gen. Virol. 1984, 65, 2077–2107. [Google Scholar] [CrossRef]
- Simmonds, P.; Xia, W.; Baillie, J.K.; McKinnon, K. Modelling mutational and selection pressures on dinucleotides in eukaryotic phyla—Selection against CpG and UpA in cytoplasmically expressed RNA and in RNA viruses. BMC Genom. 2013, 14, 610. [Google Scholar] [CrossRef]
- Boudraa, M.; Perrin, P. CpG and TpA frequencies in the plant system. Nucleic Acids Res. 1987, 15, 5729–5737. [Google Scholar] [CrossRef][Green Version]
- Di Giallonardo, F.; Schlub, T.E.; Shi, M.; Holmes, E.C. Dinucleotide Composition in Animal RNA Viruses Is Shaped More by Virus Family than by Host Species. J. Virol. 2017, 91, e02381-16. [Google Scholar] [CrossRef]
- Coulondre, C.; Miller, J.H.; Farabaugh, P.J.; Gilbert, W. Molecular basis of base substitution hotspots in Escherichia coli. Nature 1978, 274, 775–780. [Google Scholar] [CrossRef] [PubMed]
- Gaunt, E.; Wise, H.M.; Zhang, H.; Lee, L.N.; Atkinson, N.J.; Nicol, M.Q.; Highton, A.J.; Klenerman, P.; Beard, P.M.; Dutia, B.M.; et al. Elevation of CpG frequencies in influenza A genome attenuates pathogenicity but enhances host response to infection. eLife 2016, 5, e12735. [Google Scholar] [CrossRef] [PubMed]
- Fros, J.J.; Visser, I.; Tang, B.; Yan, K.; Nakayama, E.; Visser, T.M.; Koenraadt, C.J.M.; van Oers, M.M.; Pijlman, G.P.; Suhrbier, A.; et al. The dinucleotide composition of the Zika virus genome is shaped by conflicting evolutionary pressures in mammalian hosts and mosquito vectors. PLoS Biol. 2021, 19, e3001201. [Google Scholar] [CrossRef] [PubMed]
- Fros, J.J.; Dietrich, I.; Alshaikhahmed, K.; Passchier, T.C.; Evans, D.J.; Simmonds, P. CpG and UpA dinucleotides in both coding and non-coding regions of echovirus 7 inhibit replication initiation post-entry. eLife 2017, 6, e29112. [Google Scholar] [CrossRef]
- Takata, M.A.; Goncalves-Carneiro, D.; Zang, T.M.; Soll, S.J.; York, A.; Blanco-Melo, D.; Bieniasz, P.D. CG dinucleotide suppression enables antiviral defence targeting non-self RNA. Nature 2017, 550, 124–127. [Google Scholar] [CrossRef]
- Ficarelli, M.; Neil, S.J.D.; Swanson, C.M. Targeted Restriction of Viral Gene Expression and Replication by the ZAP Antiviral System. Annu. Rev. Virol. 2021, 8. [Google Scholar] [CrossRef]
- Li, M.M.H.; Aguilar, E.G.; Michailidis, E.; Pabon, J.; Park, P.; Wu, X.; de Jong, Y.P.; Schneider, W.M.; Molina, H.; Rice, C.M.; et al. Characterization of Novel Splice Variants of Zinc Finger Antiviral Protein (ZAP). J. Virol. 2019, 93, e00715–e00719. [Google Scholar] [CrossRef]
- Charron, G.; Li, M.M.; MacDonald, M.R.; Hang, H.C. Prenylome profiling reveals S-farnesylation is crucial for membrane targeting and antiviral activity of ZAP long-isoform. Proc. Natl. Acad. Sci. USA 2013, 110, 11085–11090. [Google Scholar] [CrossRef]
- Guo, X.; Carroll, J.W.; Macdonald, M.R.; Goff, S.P.; Gao, G. The zinc finger antiviral protein directly binds to specific viral mRNAs through the CCCH zinc finger motifs. J. Virol. 2004, 78, 12781–12787. [Google Scholar] [CrossRef]
- Gao, G.; Guo, X.; Goff, S.P. Inhibition of retroviral RNA production by ZAP, a CCCH-type zinc finger protein. Science 2002, 297, 1703–1706. [Google Scholar] [CrossRef] [PubMed]
- Guo, X.; Ma, J.; Sun, J.; Gao, G. The zinc-finger antiviral protein recruits the RNA processing exosome to degrade the target mRNA. Proc. Natl. Acad. Sci. USA 2007, 104, 151–156. [Google Scholar] [CrossRef]
- Zhu, Y.; Chen, G.; Lv, F.; Wang, X.; Ji, X.; Xu, Y.; Sun, J.; Wu, L.; Zheng, Y.T.; Gao, G. Zinc-finger antiviral protein inhibits HIV-1 infection by selectively targeting multiply spliced viral mRNAs for degradation. Proc. Natl. Acad. Sci. USA 2011, 108, 15834–15839. [Google Scholar] [CrossRef]
- Muller, S.; Moller, P.; Bick, M.J.; Wurr, S.; Becker, S.; Gunther, S.; Kummerer, B.M. Inhibition of filovirus replication by the zinc finger antiviral protein. J. Virol. 2007, 81, 2391–2400. [Google Scholar] [CrossRef]
- Liu, C.H.; Zhou, L.; Chen, G.; Krug, R.M. Battle between influenza A virus and a newly identified antiviral activity of the PARP-containing ZAPL protein. Proc. Natl. Acad. Sci. USA 2015, 112, 14048–14053. [Google Scholar] [CrossRef]
- Chiu, H.P.; Chiu, H.; Yang, C.F.; Lee, Y.L.; Chiu, F.L.; Kuo, H.C.; Lin, R.J.; Lin, Y.L. Inhibition of Japanese encephalitis virus infection by the host zinc-finger antiviral protein. PLoS Pathog. 2018, 14, e1007166. [Google Scholar] [CrossRef]
- Mao, R.; Nie, H.; Cai, D.; Zhang, J.; Liu, H.; Yan, R.; Cuconati, A.; Block, T.M.; Guo, J.T.; Guo, H. Inhibition of hepatitis B virus replication by the host zinc finger antiviral protein. PLoS Pathog. 2013, 9, e1003494. [Google Scholar] [CrossRef]
- Hayakawa, S.; Shiratori, S.; Yamato, H.; Kameyama, T.; Kitatsuji, C.; Kashigi, F.; Goto, S.; Kameoka, S.; Fujikura, D.; Yamada, T.; et al. ZAPS is a potent stimulator of signaling mediated by the RNA helicase RIG-I during antiviral responses. Nat. Immunol. 2011, 12, 37–44. [Google Scholar] [CrossRef]
- Li, M.; Yan, K.; Wei, L.; Yang, J.; Lu, C.; Xiong, F.; Zheng, C.; Xu, W. Zinc finger antiviral protein inhibits coxsackievirus B3 virus replication and protects against viral myocarditis. Antiviral. Res. 2015, 123, 50–61. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez-Perez, A.C.; Stempel, M.; Wyler, E.; Urban, C.; Piras, A.; Hennig, T.; Ganskih, S.; Wei, Y.; Heim, A.; Landthaler, M.; et al. The Zinc Finger Antiviral Protein ZAP Restricts Human Cytomegalovirus and Selectively Binds and Destabilizes Viral UL4/UL5 Transcripts. mBio 2021, 12, e02683-20. [Google Scholar] [CrossRef]
- Lin, Y.T.; Chiweshe, S.; McCormick, D.; Raper, A.; Wickenhagen, A.; DeFillipis, V.; Gaunt, E.; Simmonds, P.; Wilson, S.J.; Grey, F. Human cytomegalovirus evades ZAP detection by suppressing CpG dinucleotides in the major immediate early 1 gene. PLoS Pathog. 2020, 16, e1008844. [Google Scholar] [CrossRef]
- Peng, C.; Wyatt, L.S.; Glushakow-Smith, S.G.; Lal-Nag, M.; Weisberg, A.S.; Moss, B. Zinc-finger antiviral protein (ZAP) is a restriction factor for replication of modified vaccinia virus Ankara (MVA) in human cells. PLoS Pathog. 2020, 16, e1008845. [Google Scholar] [CrossRef]
- Odon, V.; Fros, J.J.; Goonawardane, N.; Dietrich, I.; Ibrahim, A.; Alshaikhahmed, K.; Nguyen, D.; Simmonds, P. The role of ZAP and OAS3/RNAseL pathways in the attenuation of an RNA virus with elevated frequencies of CpG and UpA dinucleotides. Nucleic Acids Res. 2019, 47, 8061–8083. [Google Scholar] [CrossRef]
- Luo, X.; Wang, X.; Gao, Y.; Zhu, J.; Liu, S.; Gao, G.; Gao, P. Molecular Mechanism of RNA Recognition by Zinc-Finger Antiviral Protein. Cell Rep. 2020, 30, 46–52. [Google Scholar] [CrossRef]
- Kmiec, D.; Nchioua, R.; Sherrill-Mix, S.; Sturzel, C.M.; Heusinger, E.; Braun, E.; Gondim, M.V.P.; Hotter, D.; Sparrer, K.M.J.; Hahn, B.H.; et al. CpG Frequency in the 5’ Third of the env Gene Determines Sensitivity of Primary HIV-1 Strains to the Zinc-Finger Antiviral Protein. mBio 2020, 11, e02903–e02919. [Google Scholar] [CrossRef] [PubMed]
- Ficarelli, M.; Antzin-Anduetza, I.; Hugh-White, R.; Firth, A.E.; Sertkaya, H.; Wilson, H.; Neil, S.J.D.; Schulz, R.; Swanson, C.M. CpG Dinucleotides Inhibit HIV-1 Replication through Zinc Finger Antiviral Protein (ZAP)-Dependent and -Independent Mechanisms. J. Virol. 2020, 94, e01337-19. [Google Scholar] [CrossRef] [PubMed]
- Meyerson, N.R.; Zhou, L.; Guo, Y.R.; Zhao, C.; Tao, Y.J.; Krug, R.M.; Sawyer, S.L. Nuclear TRIM25 Specifically Targets Influenza Virus Ribonucleoproteins to Block the Onset of RNA Chain Elongation. Cell Host Microbe 2017, 22, 627–638. [Google Scholar] [CrossRef]
- Zheng, X.; Wang, X.; Tu, F.; Wang, Q.; Fan, Z.; Gao, G. TRIM25 Is Required for the Antiviral Activity of Zinc Finger Antiviral Protein. J. Virol. 2017, 91, e00088-17. [Google Scholar] [CrossRef]
- Li, M.M.; Lau, Z.; Cheung, P.; Aguilar, E.G.; Schneider, W.M.; Bozzacco, L.; Molina, H.; Buehler, E.; Takaoka, A.; Rice, C.M.; et al. TRIM25 Enhances the Antiviral Action of Zinc-Finger Antiviral Protein (ZAP). PLoS Pathog. 2017, 13, e1006145. [Google Scholar] [CrossRef] [PubMed]
- Ficarelli, M.; Wilson, H.; Pedro Galao, R.; Mazzon, M.; Antzin-Anduetza, I.; Marsh, M.; Neil, S.J.; Swanson, C.M. KHNYN is essential for the zinc finger antiviral protein (ZAP) to restrict HIV-1 containing clustered CpG dinucleotides. eLife 2019, 8, e46767. [Google Scholar] [CrossRef] [PubMed]
- Fields, B.N.; Knipe, D.M.; Chanock, R.M.; Hirsch, M.S.; Melnick, J.L.; Monath, T.P.; Roizman, B. Fields Virology; Wolters Kluwer Health/Lippincott Williams & Wilkins: Philadelphia, PA, USA, 2007. [Google Scholar]
- Jha, H.C.; Banerjee, S.; Robertson, E.S. The Role of Gammaherpesviruses in Cancer Pathogenesis. Pathogens 2016, 5, 18. [Google Scholar] [CrossRef]
- Wilson, J.B.; Bell, J.L.; Levine, A.J. Expression of Epstein-Barr virus nuclear antigen-1 induces B cell neoplasia in transgenic mice. EMBO J. 1996, 15, 3117–3126. [Google Scholar] [CrossRef] [PubMed]
- Honess, R.W.; Gompels, U.A.; Barrell, B.G.; Craxton, M.; Cameron, K.R.; Staden, R.; Chang, Y.N.; Hayward, G.S. Deviations from expected frequencies of CpG dinucleotides in herpesvirus DNAs may be diagnostic of differences in the states of their latent genomes. J. Gen. Virol. 1989, 70, 837–855. [Google Scholar] [CrossRef]
- Su, C.; Zhang, J.; Zheng, C. Herpes simplex virus 1 UL41 protein abrogates the antiviral activity of hZAP by degrading its mRNA. Virol. J. 2015, 12, 203. [Google Scholar] [CrossRef]
- Shu, M.; Taddeo, B.; Zhang, W.; Roizman, B. Selective degradation of mRNAs by the HSV host shutoff RNase is regulated by the UL47 tegument protein. Proc. Natl. Acad. Sci. USA 2013, 110, E1669–E1675. [Google Scholar] [CrossRef]
- Weekes, M.P.; Tomasec, P.; Huttlin, E.L.; Fielding, C.A.; Nusinow, D.; Stanton, R.J.; Wang, E.C.Y.; Aicheler, R.; Murrell, I.; Wilkinson, G.W.G.; et al. Quantitative temporal viromics: An approach to investigate host-pathogen interaction. Cell 2014, 157, 1460–1472. [Google Scholar] [CrossRef]
- Collins-McMillen, D.; Kamil, J.; Moorman, N.; Goodrum, F. Control of Immediate Early Gene Expression for Human Cytomegalovirus Reactivation. Front. Cell. Infect. Microbiol. 2020, 10, 476. [Google Scholar] [CrossRef]
- Murray, M.J.; Peters, N.E.; Reeves, M.B. Navigating the Host Cell Response during Entry into Sites of Latent Cytomegalovirus Infection. Pathogens 2018, 7, 30. [Google Scholar] [CrossRef]
- Xuan, Y.; Liu, L.; Shen, S.; Deng, H.; Gao, G. Zinc finger antiviral protein inhibits murine gammaherpesvirus 68 M2 expression and regulates viral latency in cultured cells. J. Virol. 2012, 86, 12431–12434. [Google Scholar] [CrossRef] [PubMed]
- Xuan, Y.; Gong, D.; Qi, J.; Han, C.; Deng, H.; Gao, G. ZAP inhibits murine gammaherpesvirus 68 ORF64 expression and is antagonized by RTA. J. Virol. 2013, 87, 2735–2743. [Google Scholar] [CrossRef] [PubMed]
- Luo, M.J.; Reed, R. Splicing is required for rapid and efficient mRNA export in metazoans. Proc. Natl. Acad. Sci. USA 1999, 96, 14937–14942. [Google Scholar] [CrossRef] [PubMed]
- Luo, M.L.; Zhou, Z.; Magni, K.; Christoforides, C.; Rappsilber, J.; Mann, M.; Reed, R. Pre-mRNA splicing and mRNA export linked by direct interactions between UAP56 and Aly. Nature 2001, 413, 644–647. [Google Scholar] [CrossRef] [PubMed]
- Mordstein, C.; Savisaar, R.; Young, R.S.; Bazile, J.; Talmane, L.; Luft, J.; Liss, M.; Taylor, M.S.; Hurst, L.D.; Kudla, G. Codon Usage and Splicing Jointly Influence mRNA Localization. Cell Syst. 2020, 10, 351–362. [Google Scholar] [CrossRef]
- Minarovits, J. Epigenotypes of latent herpesvirus genomes. Curr. Top. Microbiol. Immunol. 2006, 310, 61–80. [Google Scholar] [CrossRef]
- Gunther, T.; Grundhoff, A. The epigenetic landscape of latent Kaposi sarcoma-associated herpesvirus genomes. PLoS Pathog. 2010, 6, e1000935. [Google Scholar] [CrossRef]
- Niller, H.H.; Tarnai, Z.; Decsi, G.; Zsedenyi, A.; Banati, F.; Minarovits, J. Role of epigenetics in EBV regulation and pathogenesis. Future Microbiol. 2014, 9, 747–756. [Google Scholar] [CrossRef]
- Kubat, N.J.; Tran, R.K.; McAnany, P.; Bloom, D.C. Specific histone tail modification and not DNA methylation is a determinant of herpes simplex virus type 1 latent gene expression. J. Virol. 2004, 78, 1139–1149. [Google Scholar] [CrossRef]
- Brooks, A.R.; Harkins, R.N.; Wang, P.; Qian, H.S.; Liu, P.; Rubanyi, G.M. Transcriptional silencing is associated with extensive methylation of the CMV promoter following adenoviral gene delivery to muscle. J. Gene. Med. 2004, 6, 395–404. [Google Scholar] [CrossRef] [PubMed]
- Pereira-Montecinos, C.; Valiente-Echeverria, F.; Soto-Rifo, R. Epitranscriptomic regulation of viral replication. Biochim. Biophys. Acta (BBA)-Gene Regul. Mech. 2017, 1860, 460–471. [Google Scholar] [CrossRef]
- Yang, X.; Yang, Y.; Sun, B.F.; Chen, Y.S.; Xu, J.W.; Lai, W.Y.; Li, A.; Wang, X.; Bhattarai, D.P.; Xiao, W.; et al. 5-methylcytosine promotes mRNA export-NSUN2 as the methyltransferase and ALYREF as an m(5)C reader. Cell Res. 2017, 27, 606–625. [Google Scholar] [CrossRef]
- Groenke, N.; Trimpert, J.; Merz, S.; Conradie, A.M.; Wyler, E.; Zhang, H.; Hazapis, O.G.; Rausch, S.; Landthaler, M.; Osterrieder, N.; et al. Mechanism of Virus Attenuation by Codon Pair Deoptimization. Cell Rep. 2020, 31, 107586. [Google Scholar] [CrossRef]
- Roychoudhury, S.; Mukherjee, D. A detailed comparative analysis on the overall codon usage pattern in herpesviruses. Virus Res. 2010, 148, 31–43. [Google Scholar] [CrossRef]
- Fu, M. Codon usage bias in herpesvirus. Arch. Virol. 2010, 155, 391–396. [Google Scholar] [CrossRef] [PubMed]
- Mordstein, C.; Cano, L.; Morales, A.C.; Young, B.; Ho, A.T.; Rice, A.M.; Liss, M.; Hurst, L.D.; Kudla, G. Transcription, mRNA export and immune evasion shape the codon usage of viruses. Genome Biol. Evol. 2021, 12. [Google Scholar] [CrossRef]
- Shi, Y. Mechanistic insights into precursor messenger RNA splicing by the spliceosome. Nat. Rev. Mol. Cell. Biol. 2017, 18, 655–670. [Google Scholar] [CrossRef] [PubMed]
- Kohler, A.; Hurt, E. Exporting RNA from the nucleus to the cytoplasm. Nat. Rev. Mol. Cell. Biol. 2007, 8, 761–773. [Google Scholar] [CrossRef]
- Rondon, A.G.; Jimeno, S.; Aguilera, A. The interface between transcription and mRNP export: From THO to THSC/TREX-2. Biochim. Biophys. Acta 2010, 1799, 533–538. [Google Scholar] [CrossRef]
- Strasser, K.; Masuda, S.; Mason, P.; Pfannstiel, J.; Oppizzi, M.; Rodriguez-Navarro, S.; Rondon, A.G.; Aguilera, A.; Struhl, K.; Reed, R.; et al. TREX is a conserved complex coupling transcription with messenger RNA export. Nature 2002, 417, 304–308. [Google Scholar] [CrossRef]
- Hautbergue, G.M.; Hung, M.L.; Golovanov, A.P.; Lian, L.Y.; Wilson, S.A. Mutually exclusive interactions drive handover of mRNA from export adaptors to TAP. Proc. Natl. Acad. Sci. USA 2008, 105, 5154–5159. [Google Scholar] [CrossRef]
- Sandri-Goldin, R.M. The many roles of the highly interactive HSV protein ICP27, a key regulator of infection. Future Microbiol. 2011, 6, 1261–1277. [Google Scholar] [CrossRef]
- Jackson, B.R.; Noerenberg, M.; Whitehouse, A. The Kaposi’s Sarcoma-Associated Herpesvirus ORF57 Protein and Its Multiple Roles in mRNA Biogenesis. Front. Microbiol. 2012, 3, 59. [Google Scholar] [CrossRef]
- Lischka, P.; Toth, Z.; Thomas, M.; Mueller, R.; Stamminger, T. The UL69 transactivator protein of human cytomegalovirus interacts with DEXD/H-Box RNA helicase UAP56 to promote cytoplasmic accumulation of unspliced RNA. Mol. Cell. Biol. 2006, 26, 1631–1643. [Google Scholar] [CrossRef]
- Sei, E.; Conrad, N.K. Delineation of a core RNA element required for Kaposi’s sarcoma-associated herpesvirus ORF57 binding and activity. Virology 2011, 419, 107–116. [Google Scholar] [CrossRef]
- Mears, W.E.; Rice, S.A. The RGG box motif of the herpes simplex virus ICP27 protein mediates an RNA-binding activity and determines in vivo methylation. J. Virol. 1996, 70, 7445–7453. [Google Scholar] [CrossRef] [PubMed]
- Corbin-Lickfett, K.A.; Chen, I.H.; Cocco, M.J.; Sandri-Goldin, R.M. The HSV-1 ICP27 RGG box specifically binds flexible, GC-rich sequences but not G-quartet structures. Nucleic Acids Res. 2009, 37, 7290–7301. [Google Scholar] [CrossRef]
- Tang, S.; Patel, A.; Krause, P.R. Herpes simplex virus ICP27 regulates alternative pre-mRNA polyadenylation and splicing in a sequence-dependent manner. Proc. Natl. Acad. Sci. USA 2016, 113, 12256–12261. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Hennig, T.; Whisnant, A.W.; Erhard, F.; Prusty, B.K.; Friedel, C.C.; Forouzmand, E.; Hu, W.; Erber, L.; Chen, Y.; et al. Herpes simplex virus blocks host transcription termination via the bimodal activities of ICP27. Nat. Commun. 2020, 11, 293. [Google Scholar] [CrossRef] [PubMed]


Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lin, Y.-T.; Chau, L.-F.; Coutts, H.; Mahmoudi, M.; Drampa, V.; Lee, C.-H.; Brown, A.; Hughes, D.J.; Grey, F. Does the Zinc Finger Antiviral Protein (ZAP) Shape the Evolution of Herpesvirus Genomes? Viruses 2021, 13, 1857. https://doi.org/10.3390/v13091857
Lin Y-T, Chau L-F, Coutts H, Mahmoudi M, Drampa V, Lee C-H, Brown A, Hughes DJ, Grey F. Does the Zinc Finger Antiviral Protein (ZAP) Shape the Evolution of Herpesvirus Genomes? Viruses. 2021; 13(9):1857. https://doi.org/10.3390/v13091857
Chicago/Turabian StyleLin, Yao-Tang, Long-Fung Chau, Hannah Coutts, Matin Mahmoudi, Vayalena Drampa, Chen-Hsuin Lee, Alex Brown, David J. Hughes, and Finn Grey. 2021. "Does the Zinc Finger Antiviral Protein (ZAP) Shape the Evolution of Herpesvirus Genomes?" Viruses 13, no. 9: 1857. https://doi.org/10.3390/v13091857
APA StyleLin, Y.-T., Chau, L.-F., Coutts, H., Mahmoudi, M., Drampa, V., Lee, C.-H., Brown, A., Hughes, D. J., & Grey, F. (2021). Does the Zinc Finger Antiviral Protein (ZAP) Shape the Evolution of Herpesvirus Genomes? Viruses, 13(9), 1857. https://doi.org/10.3390/v13091857





