Antiviral Activity of Vitis vinifera Leaf Extract against SARS-CoV-2 and HSV-1
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material
2.2. Polyphenols Extraction
2.3. LC-MS Analysis of Flavonoids
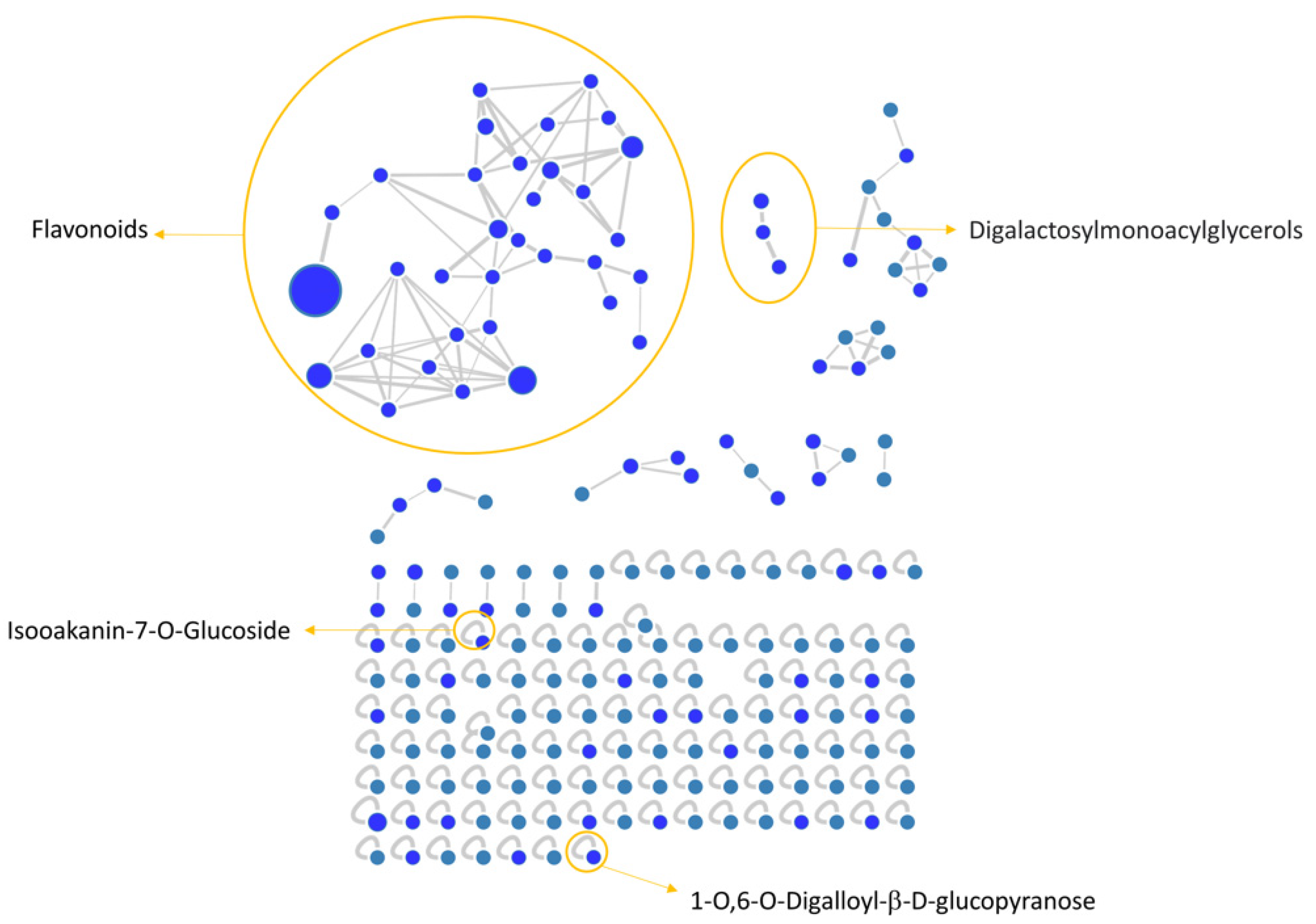
2.4. Molecular Networking and Spectral Library Search
2.5. Cell Lines and Virus
2.6. Cytotoxicity Test
2.7. Antiviral Activity
2.8. Gene Expression
3. Results and Discussion
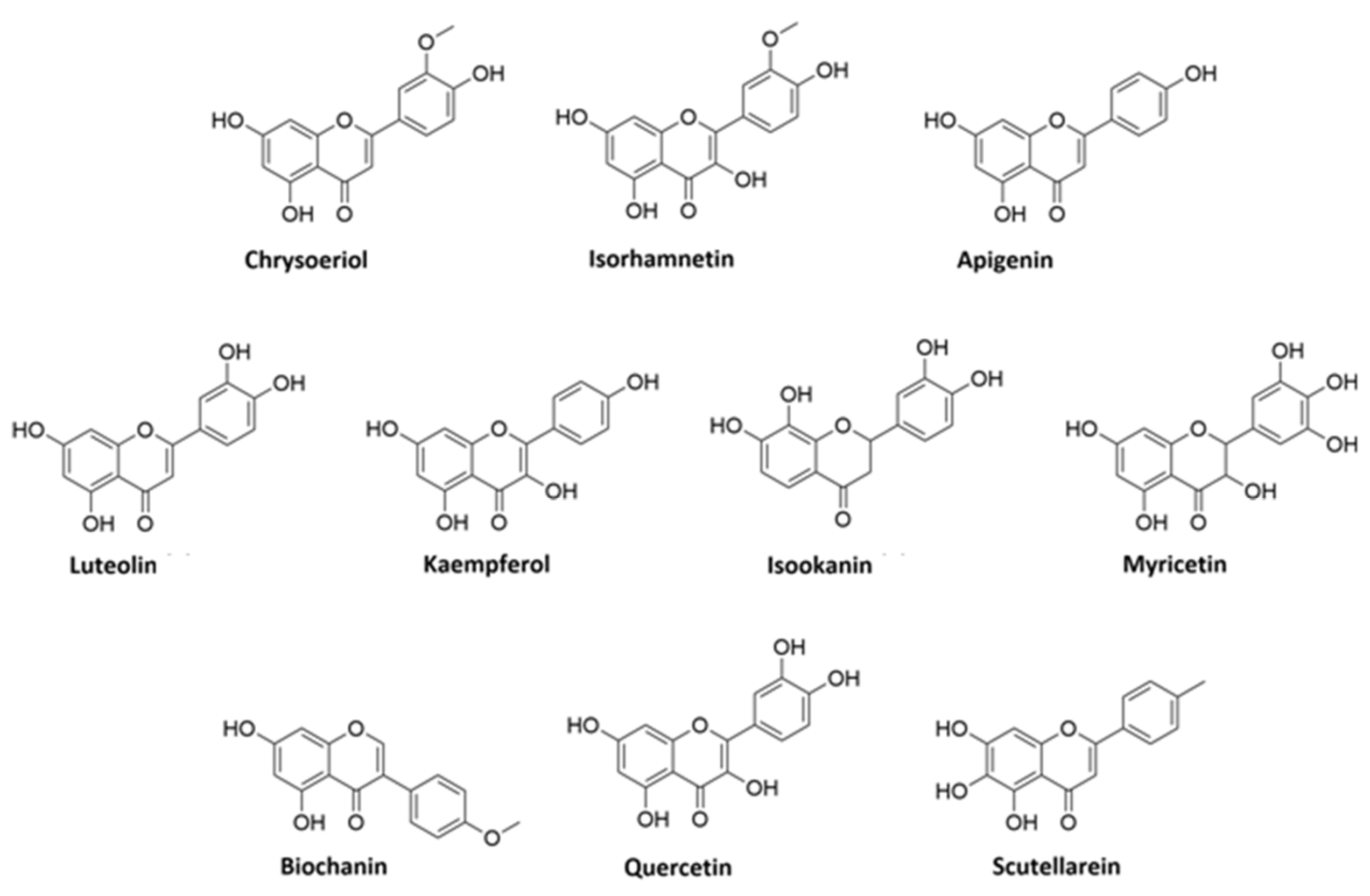
3.1. Flavonoid Composition of V. vinifera Leaves
3.2. Identification of Kaempferol and Luteolin Derivatives
3.3. Identification of Apigenin Derivatives
3.4. Identification of Quercetin Derivatives
3.5. Identification of Isorhamnetin Derivatives
3.6. Other Flavonoids Assignment
3.7. DGMG Assignment
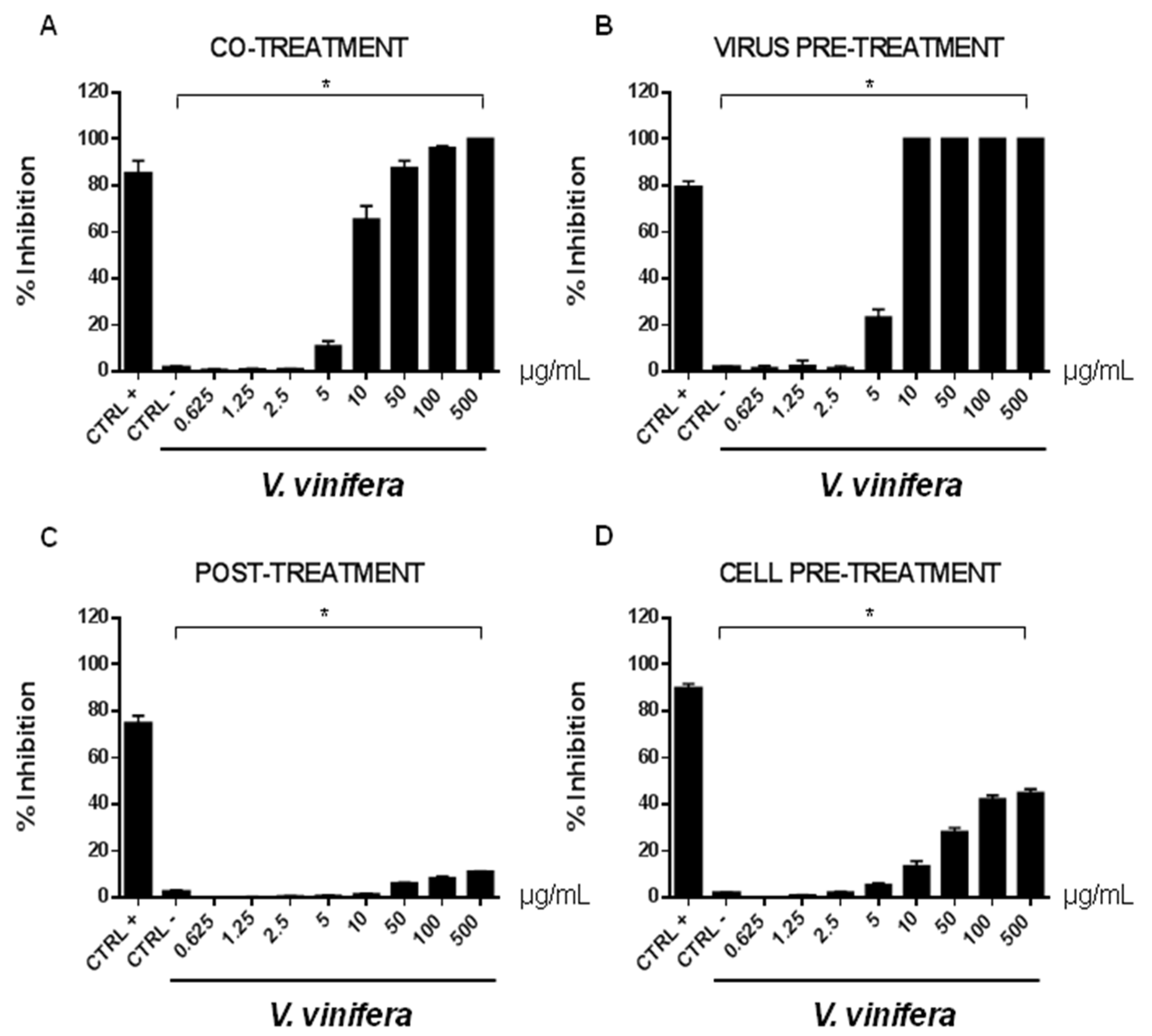
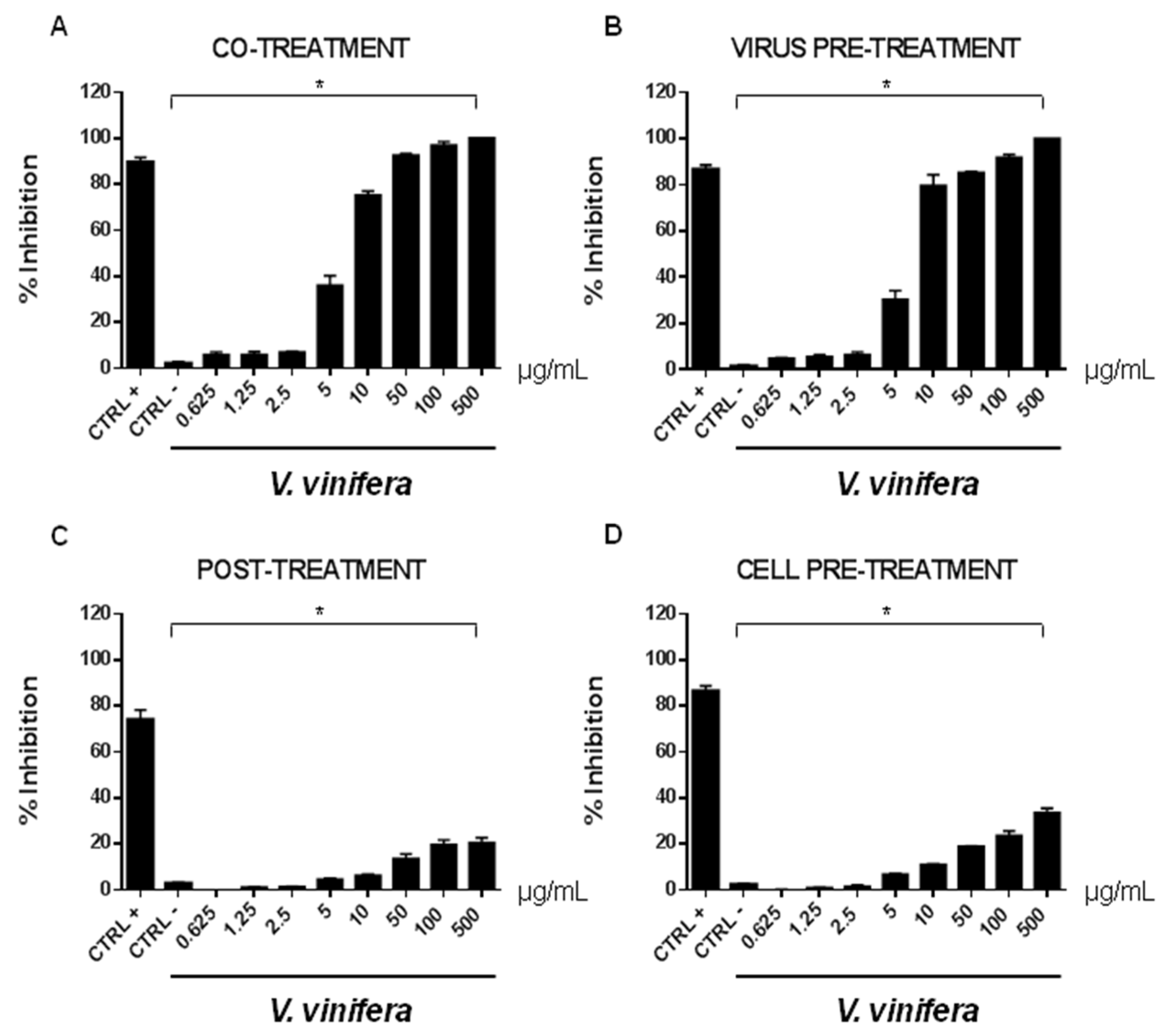
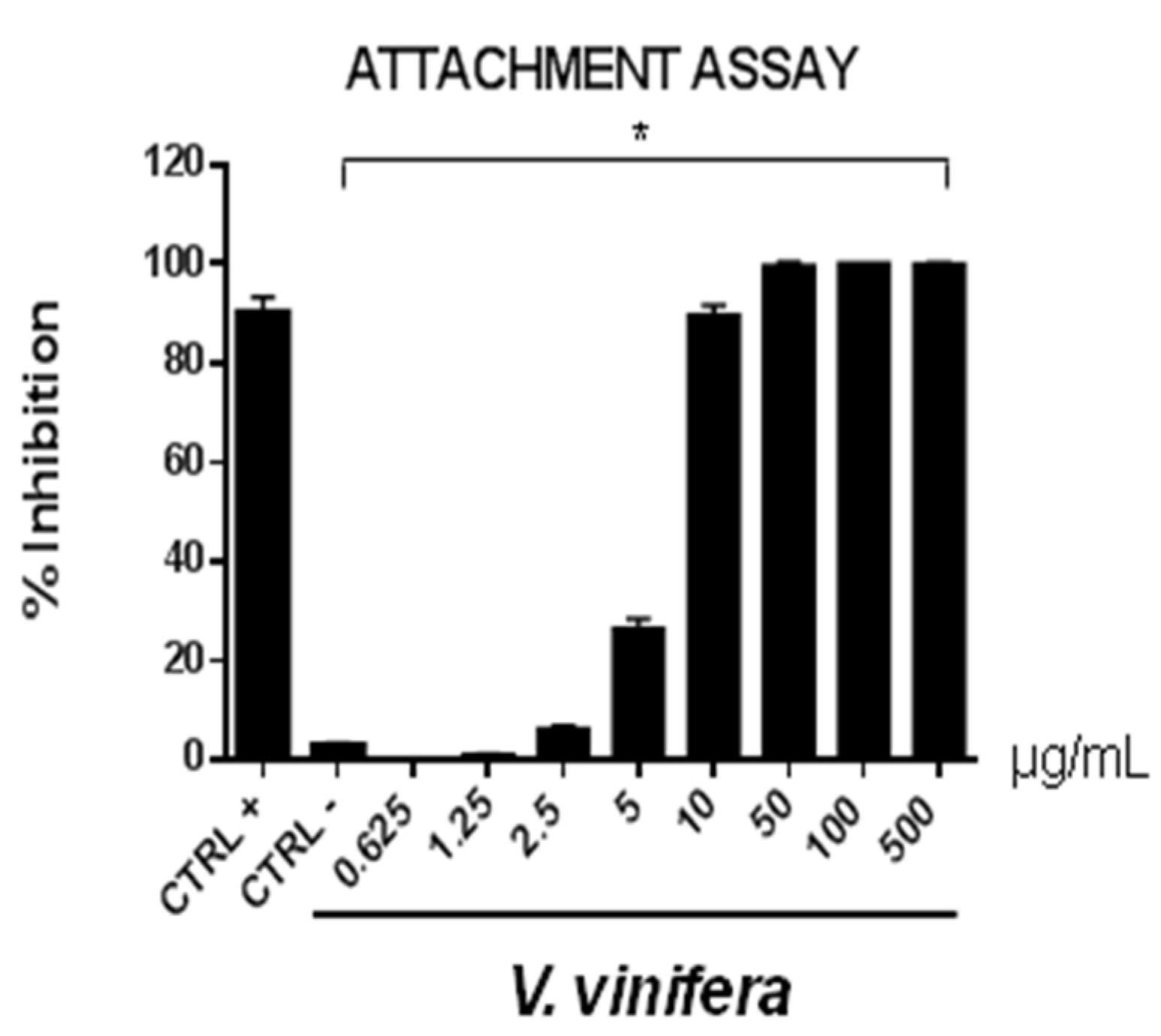
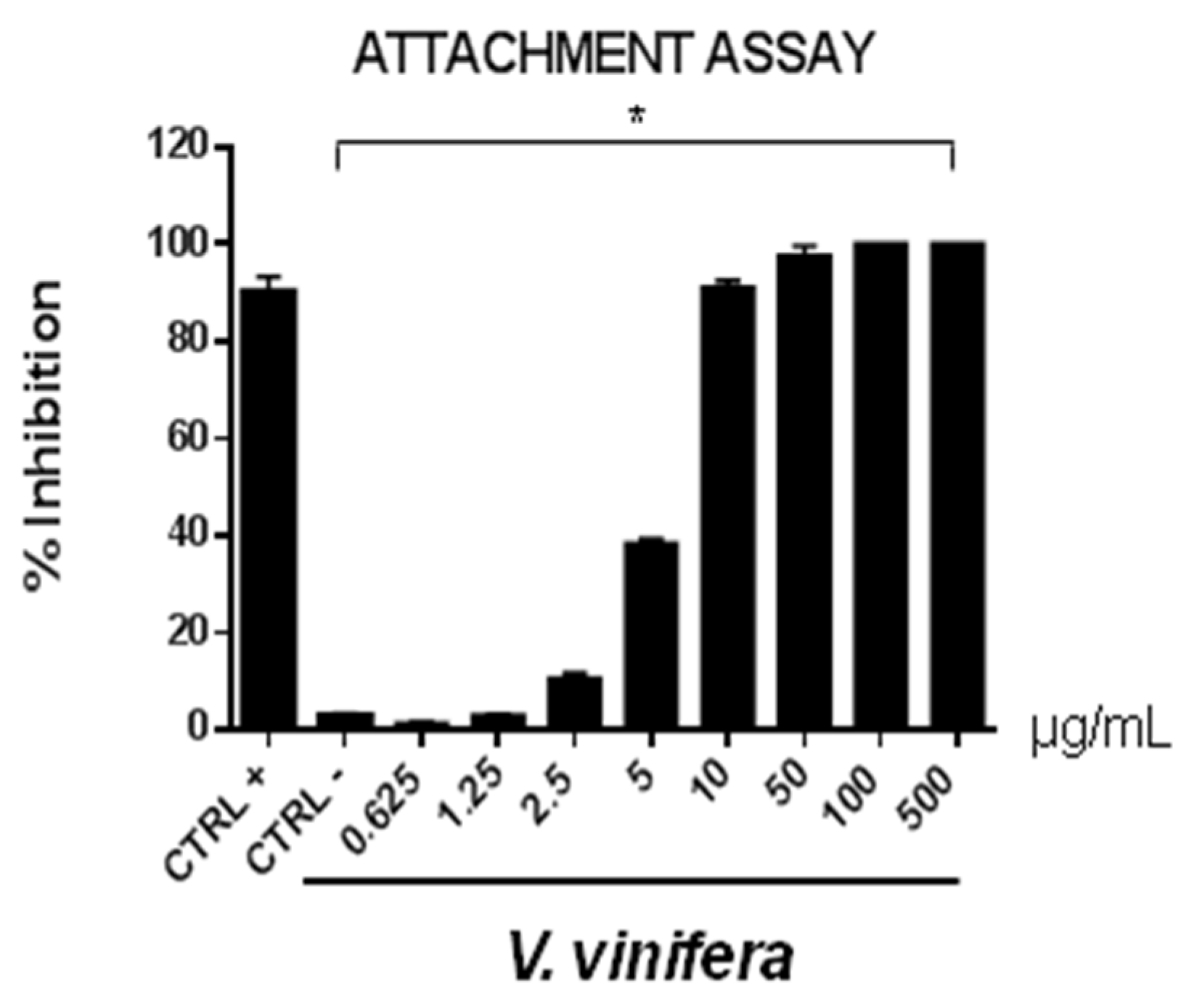
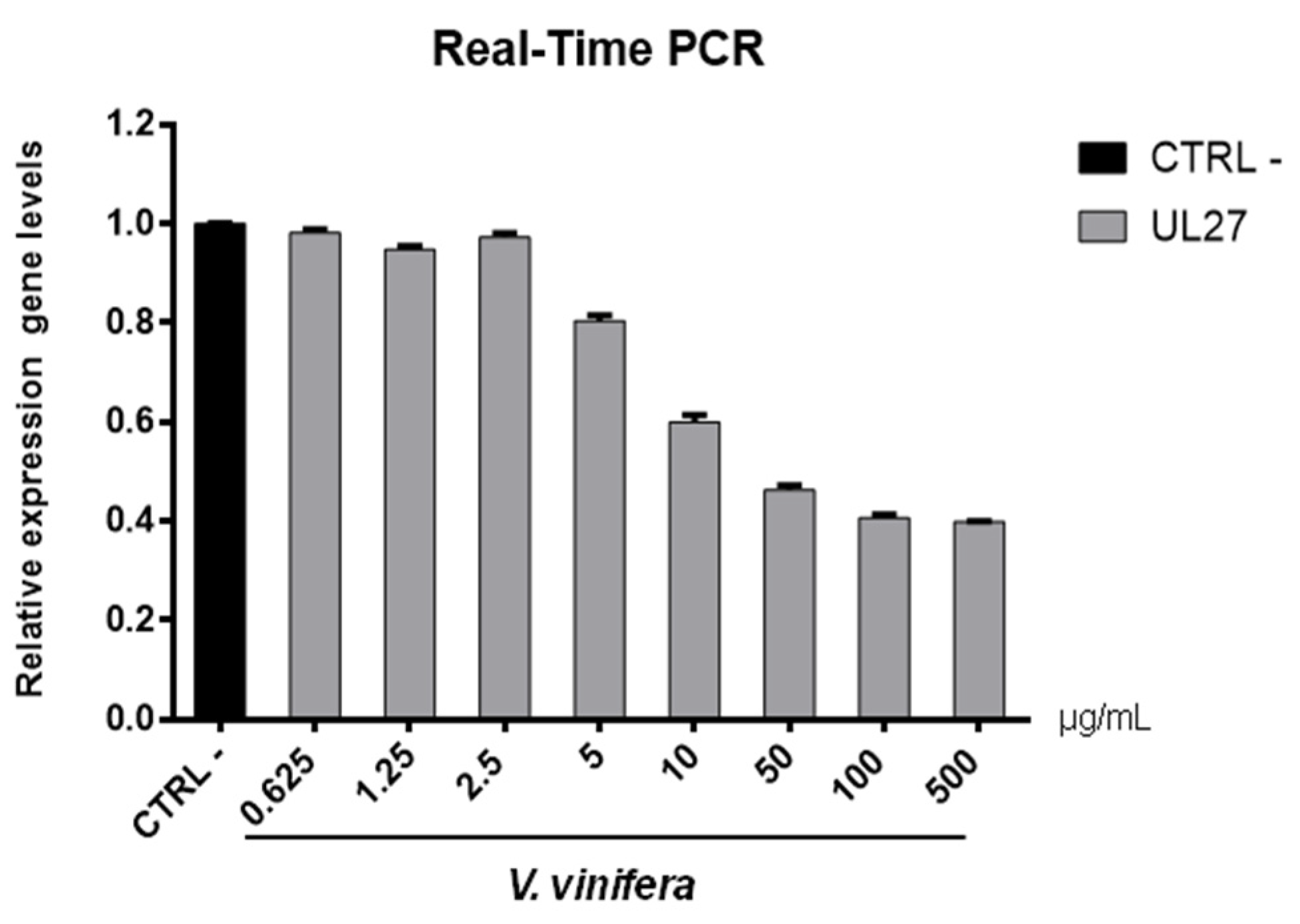
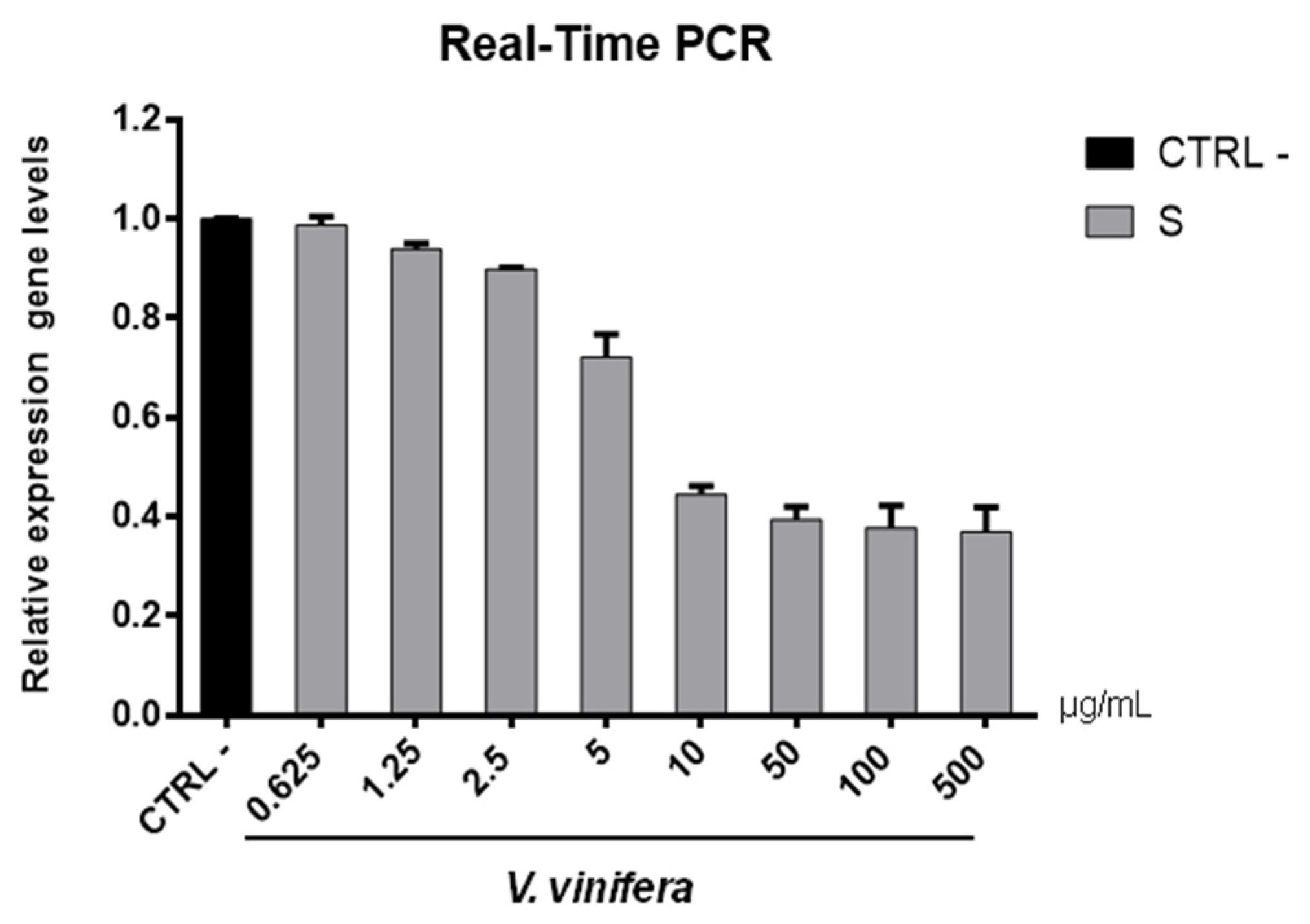
3.8. Antiviral Activity
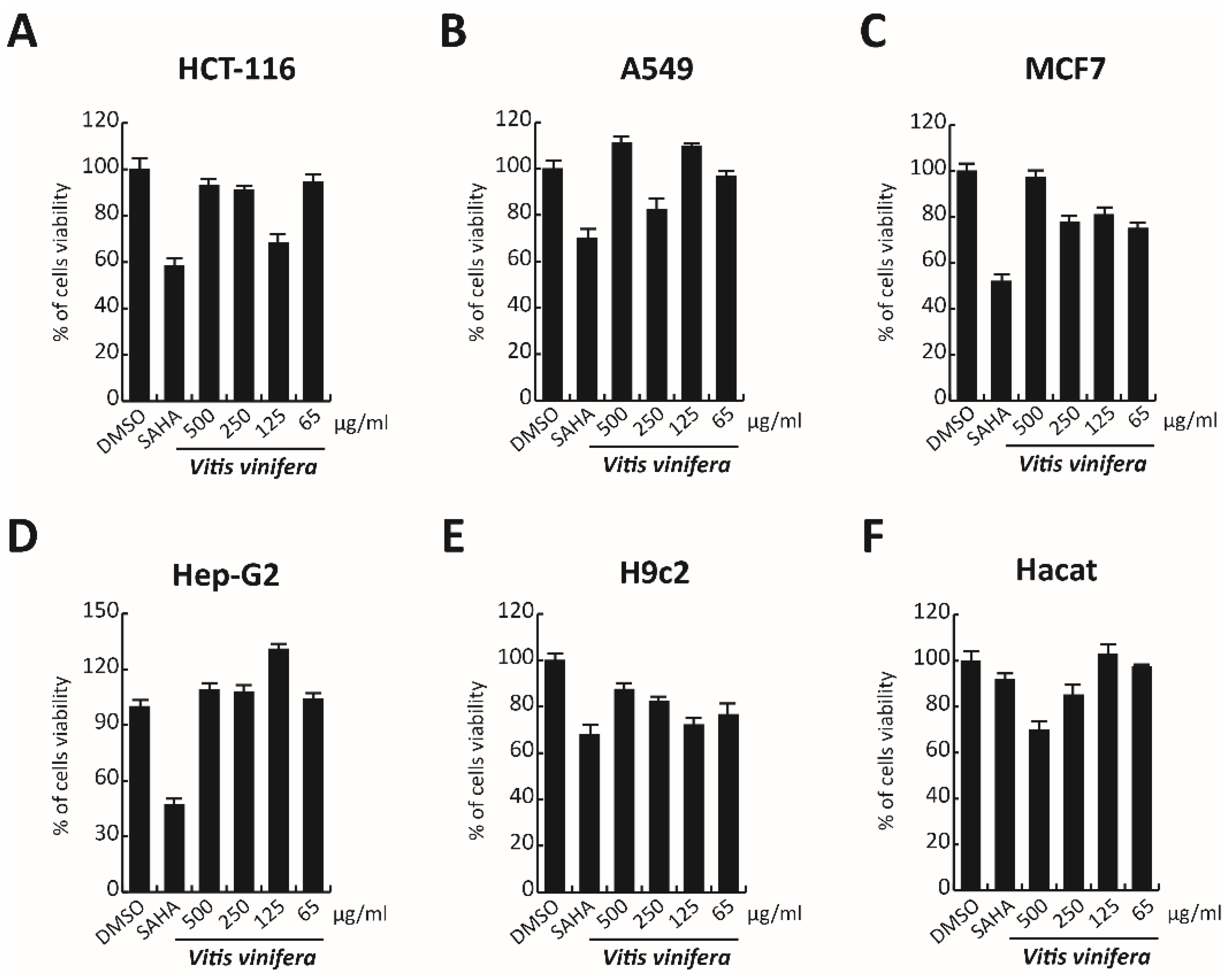
3.9. Cytotoxicity Evaluation
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Rinaldi, A.; Villano, C.; Lanzillo, C.; Tamburrino, A., Jr.; Jourdes, M.; Teissedre, P.L.; Moio, L.; Frusciante, L.; Carputo, D.; Aversano, R. Metabolic and RNA profiling elucidates proanthocyanidins accumulation in Aglianico grape. Food Chem. 2017, 233, 52–59. [Google Scholar] [CrossRef] [PubMed]
- Gratl, V.; Sturm, S.; Zini, E.; Letschka, T.; Stefanini, M.; Vezzulli, S.; Stuppner, H. Comprehensive polyphenolic profiling in promising resistant grapevine hybrids including 17 novel breeds in northern Italy. J. Sci. Food Agric. 2021, 101, 2380–2388. [Google Scholar] [CrossRef]
- Sikuten, I.; Stambuk, P.; Andabaka, Z.; Tomaz, I.; Markovic, Z.; Stupic, D.; Maletic, E.; Kontic, J.K.; Preiner, D. Grapevine as a Rich Source of Polyphenolic Compounds. Molecules 2020, 25, 5604. [Google Scholar] [CrossRef]
- Ju, Y.L.; Yue, X.F.; Min, Z.; Wang, X.H.; Fang, Y.L.; Zhang, J.X. VvNAC17, a novel stress-responsive grapevine (Vitis vinifera L.) NAC transcription factor, increases sensitivity to abscisic acid and enhances salinity, freezing, and drought tolerance in transgenic Arabidopsis. Plant. Physiol. Biochem. 2020, 146, 98–111. [Google Scholar] [CrossRef]
- Batiha, G.E.; Beshbishy, A.M.; Ikram, M.; Mulla, Z.S.; El-Hack, M.E.A.; Taha, A.E.; Algammal, A.M.; Elewa, Y.H.A. The Pharmacological Activity, Biochemical Properties, and Pharmacokinetics of the Major Natural Polyphenolic Flavonoid: Quercetin. Foods 2020, 9, 374. [Google Scholar] [CrossRef]
- Duan, J.; Zhan, J.C.; Wang, G.Z.; Zhao, X.C.; Huang, W.D.; Zhou, G.B. The red wine component ellagic acid induces autophagy and exhibits anti-lung cancer activity in vitro and in vivo. J. Cell Mol. Med. 2019, 23, 143–154. [Google Scholar] [CrossRef]
- Saadaoui, N.; Weslati, A.; Barkaoui, T.; Khemiri, I.; Gadacha, W.; Souli, A.; Mokni, M.; Harbi, M.; Ben-Attia, M. Gastroprotective effect of leaf extract of two varieties grapevine (Vitis vinifera L.) native wild and cultivar grown in North of Tunisia against the oxidative stress induced by ethanol in rats. Biomarkers 2020, 25, 48–61. [Google Scholar] [CrossRef]
- Fujita, K.; Aoki, Y.; Suzuki, S. Antidiabetic effects of novel cell culture established from grapevine, Vitis vinifera cv. Koshu. Cytotechnology 2018, 70, 993–999. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.Y.; Hong, M.H.; Yoon, J.J.; Kim, D.S.; Na, S.W.; Jang, Y.J.; Lee, Y.J.; Kang, D.G.; Lee, H.S. Protective Effect of Vitis labrusca Leaves Extract on Cardiovascular Dysfunction through HMGB1-TLR4-NFkappaB Signaling in Spontaneously Hypertensive Rats. Nutrients 2020, 12, 10. [Google Scholar]
- Li, L.; Zhang, M.; Zhang, S.; Cui, Y.; Sun, B. Preparation and Antioxidant Activity of Ethyl-Linked Anthocyanin-Flavanol Pigments from Model Wine Solutions. Molecules 2018, 23, 1066. [Google Scholar] [CrossRef]
- Squillaci, G.; Zannella, C.; Carbone, V.; Minasi, P.; Folliero, V.; Stelitano, D.; Cara, F.L.; Galdiero, M.; Franci, G.; Morana, A. Grape Canes from Typical Cultivars of Campania (Southern Italy) as a Source of High-Value Bioactive Compounds: Phenolic Profile, Antioxidant and Antimicrobial Activities. Molecules 2021, 26, 2746. [Google Scholar] [CrossRef]
- Aliano-Gonzalez, M.J.; Richard, T.; Cantos-Villar, E. Grapevine Cane Extracts: Raw Plant Material, Extraction Methods, Quantification, and Applications. Biomolecules 2020, 10, 1195. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, M.C.; Ribeiro-Vidal, H.; Esteban-Fernandez, A.; Bartolome, B.; Figuero, E.; Moreno-Arribas, M.V.; Sanz, M.; Herrera, D. Antimicrobial activity of red wine and oenological extracts against periodontal pathogens in a validated oral biofilm model. BMC Complement. Altern Med. 2019, 19, 145. [Google Scholar] [CrossRef]
- Lee, J.W.; Kim, Y.I.; Im, C.N.; Kim, S.W.; Kim, S.J.; Min, S.; Joo, Y.H.; Yim, S.V.; Chung, N. Grape Seed Proanthocyanidin Inhibits Mucin Synthesis and Viral Replication by Suppression of AP-1 and NF-kappaB via p38 MAPKs/JNK Signaling Pathways in Respiratory Syncytial Virus-Infected A549 Cells. J. Agric. Food Chem. 2017, 65, 4472–4483. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.Y.; Li, W.G.; Wu, Y.J.; Zheng, T.Z.; Li, W.; Qu, S.Y.; Liu, N.F. Proanthocyanidin from grape seeds potentiates anti-tumor activity of doxorubicin via immunomodulatory mechanism. Int. Immunopharmacol. 2005, 5, 1247–1257. [Google Scholar] [CrossRef]
- Lin, Y.S.; Chen, S.F.; Liu, C.L.; Nieh, S. The chemoadjuvant potential of grape seed procyanidins on p53-related cell death in oral cancer cells. J. Oral Pathol. Med. 2012, 41, 322–331. [Google Scholar] [CrossRef] [PubMed]
- Zykova, T.A.; Zhu, F.; Zhai, X.; Ma, W.Y.; Ermakova, S.P.; Lee, K.W.; Bode, A.M.; Dong, Z. Resveratrol directly targets COX-2 to inhibit carcinogenesis. Mol. Carcinog. 2008, 47, 797–805. [Google Scholar] [CrossRef] [PubMed]
- Sun, Q.; Prasad, R.; Rosenthal, E.; Katiyar, S.K. Grape seed proanthocyanidins inhibit the invasive potential of head and neck cutaneous squamous cell carcinoma cells by targeting EGFR expression and epithelial-to-mesenchymal transition. BMC Complement. Altern. Med. 2011, 11, 134. [Google Scholar] [CrossRef] [PubMed]
- Schlachterman, A.; Valle, F.; Wall, K.M.; Azios, N.G.; Castillo, L.; Morell, L.; Washington, A.V.; Cubano, L.A.; Dharmawardhane, S.F. Combined resveratrol, quercetin, and catechin treatment reduces breast tumor growth in a nude mouse model. Transl. Oncol. 2008, 1, 19–27. [Google Scholar] [CrossRef] [PubMed]
- Cai, W.; Chen, G.; Luo, Q.; Liu, J.; Guo, X.; Zhang, T.; Ma, F.; Yuan, L.; Li, B.; Cai, J. PMP22 Regulates Self-Renewal and Chemoresistance of Gastric Cancer Cells. Mol. Cancer Ther. 2017, 16, 1187–1198. [Google Scholar] [CrossRef]
- Borai, I.H.; Ezz, M.K.; Rizk, M.Z.; Aly, H.F.; El-Sherbiny, M.; Matloub, A.A.; Fouad, G.I. Therapeutic impact of grape leaves polyphenols on certain biochemical and neurological markers in AlCl3-induced Alzheimer’s disease. Biomed. Pharmacother. 2017, 93, 837–851. [Google Scholar] [CrossRef]
- Di Meo, F.; Aversano, R.; Diretto, G.; Demurtas, O.C.; Villano, C.; Cozzolino, S.; Filosa, S.; Carputo, D.; Crispi, S. Anti-cancer activity of grape seed semi-polar extracts in human mesothelioma cell lines. J. Funct. Foods 2019, 61, 103515. [Google Scholar] [CrossRef]
- Radulescu, C.; Buruleanu, L.C.; Nicolescu, C.M.; Olteanu, R.L.; Bumbac, M.; Holban, G.C.; Simal-Gandara, J. Phytochemical Profiles, Antioxidant and Antibacterial Activities of Grape (Vitis vinifera L.) Seeds and Skin from Organic and Conventional Vineyards. Plants 2020, 9, 1470. [Google Scholar] [CrossRef] [PubMed]
- Al-Mousawi, A.H.; Al-Kaabi, S.J.; Albaghdadi, A.J.H.; Almulla, A.F.; Raheem, A.; Algon, A.A.A. Effect of Black Grape Seed Extract (Vitis vinifera) on Biofilm Formation of Methicillin-Resistant Staphylococcus aureus and Staphylococcus haemolyticus. Curr Microbiol 2020, 77, 238–245. [Google Scholar] [CrossRef] [PubMed]
- Pavic, V.; Kujundzic, T.; Kopic, M.; Jukic, V.; Braun, U.; Schwander, F.; Drenjancevic, M. Effects of Defoliation on Phenolic Concentrations, Antioxidant and Antibacterial Activity of Grape Skin Extracts of the Varieties Blaufrankisch and Merlot (Vitis vinifera L.). Molecules 2019, 24, 2444. [Google Scholar] [CrossRef]
- Olejar, K.J.; Ricci, A.; Swift, S.; Zujovic, Z.; Gordon, K.C.; Fedrizzi, B.; Versari, A.; Kilmartin, P.A. Characterization of an Antioxidant and Antimicrobial Extract from Cool Climate, White Grape Marc. Antioxidants 2019, 8, 232. [Google Scholar] [CrossRef]
- Santella, B.; Folliero, V.; Pirofalo, G.M.; Serretiello, E.; Zannella, C.; Moccia, G.; Santoro, E.; Sanna, G.; Motta, O.; De Caro, F.; et al. Sepsis-A Retrospective Cohort Study of Bloodstream Infections. Antibiotics 2020, 9, 851. [Google Scholar] [CrossRef]
- Zannella, C.; Shinde, S.; Vitiello, M.; Falanga, A.; Galdiero, E.; Fahmi, A.; Santella, B.; Nucci, L.; Gasparro, R.; Galdiero, M.; et al. Antibacterial Activity of Indolicidin-Coated Silver Nanoparticles in Oral Disease. Appl. Sci. 2020, 10, 1837. [Google Scholar] [CrossRef]
- Andelkovic, M.; Radovanovic, B.; Andelkovic, A.M.; Radovanovic, V. Phenolic Compounds and Bioactivity of Healthy and Infected Grapevine Leaf Extracts from Red Varieties Merlot and Vranac (Vitis vinifera L.). Plant Foods Hum. Nutr. 2015, 70, 317–323. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez Vaquero, M.J.; Alberto, M.R.; Manca de Nadra, M.C. Influence of phenolic compounds from wines on the growth of Listeria monocytogenes. Food Control. 2007, 18, 587–593. [Google Scholar] [CrossRef]
- Papadopoulou, C.; Soulti, K.; Roussis, I. Potential Antimicrobial Activity of Red and White Wine Phenolic Extracts against Strains of Staphylococcus aureus, Escherichia coli and Candida albicans. Food Technol. Biotechnol. 2005, 43, 41–46. [Google Scholar]
- Lin, S.C.; Ho, C.T.; Chuo, W.H.; Li, S.; Wang, T.T.; Lin, C.C. Effective inhibition of MERS-CoV infection by resveratrol. BMC Infect. Dis. 2017, 17, 144. [Google Scholar] [CrossRef]
- Yang, T.; Li, S.; Zhang, X.; Pang, X.; Lin, Q.; Cao, J. Resveratrol, sirtuins, and viruses. Rev. Med. Virol. 2015, 25, 431–445. [Google Scholar] [CrossRef]
- Friedman, M. Antibacterial, antiviral, and antifungal properties of wines and winery byproducts in relation to their flavonoid content. J. Agric. Food Chem. 2014, 62, 6025–6042. [Google Scholar] [CrossRef]
- Vazquez-Calvo, A.; Jimenez de Oya, N.; Martin-Acebes, M.A.; Garcia-Moruno, E.; Saiz, J.C. Antiviral Properties of the Natural Polyphenols Delphinidin and Epigallocatechin Gallate against the Flaviviruses West Nile Virus, Zika Virus, and Dengue Virus. Front. Microbiol. 2017, 8, 1314. [Google Scholar] [CrossRef]
- Madeddu, S.; Marongiu, A.; Sanna, G.; Zannella, C.; Falconieri, D.; Porcedda, S.; Manzin, A.; Piras, A. Bovine Viral Diarrhea Virus (BVDV): A Preliminary Study on Antiviral Properties of Some Aromatic and Medicinal Plants. Pathogens 2021, 10, 403. [Google Scholar] [CrossRef] [PubMed]
- Matias, A.A.; Serra, A.T.; Silva, A.C.; Perdigao, R.; Ferreira, T.B.; Marcelino, I.; Silva, S.; Coelho, A.V.; Alves, P.M.; Duarte, C.M. Portuguese winemaking residues as a potential source of natural anti-adenoviral agents. Int. J. Food Sci. Nutr. 2010, 61, 357–368. [Google Scholar] [CrossRef]
- Sharaf, M.; El-Deeb, N.; Eladawi, H. The Potentiality of Grape Seed Extract as a Novel Anti-hepatitis C virus Agent. J. Med. Sci. 2012, 12, 107. [Google Scholar] [CrossRef][Green Version]
- Su, X.; D’Souza, D.H. Grape seed extract for control of human enteric viruses. Appl. Environ. Microbiol. 2011, 77, 3982–3987. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Massive Proportion of World’s Population Are Living with Herpes Infection; World Health Organization: Geneva, Switzerland, 2020. [Google Scholar]
- World Health Organization. Globally, an Estimated Two-Thirds of the Population Under 50 are Infected with Herpes Simplex Virus Type 1; World Health Organization: Geneva, Switzerland, 2015. [Google Scholar]
- Stelitano, D.; Franci, G.; Chianese, A.; Galdiero, S.; Morelli, G.; Galdiero, M. HSV membrane glycoproteins, their function in viral entry and their use in vaccine studies. In Amino Acids, Peptides and Proteins: Volume 43; The Royal Society of Chemistry: London, UK, 2019; Volume 43, pp. 14–43. [Google Scholar]
- World Health Organization. WHO Coronavirus (COVID-19) Dashboard; World Health Organization: Geneva, Switzerland, 2021. [Google Scholar]
- Martines, R.B.; Ritter, J.M.; Matkovic, E.; Gary, J.; Bollweg, B.C.; Bullock, H.; Goldsmith, C.S.; Silva-Flannery, L.; Seixas, J.N.; Reagan-Steiner, S.; et al. Pathology and Pathogenesis of SARS-CoV-2 Associated with Fatal Coronavirus Disease, United States. Emerg. Infect. Dis. 2020, 26, 2005–2015. [Google Scholar] [CrossRef]
- Docimo, T.; Francese, G.; Ruggiero, A.; Batelli, G.; De Palma, M.; Bassolino, L.; Toppino, L.; Rotino, G.L.; Mennella, G.; Tucci, M. Phenylpropanoids Accumulation in Eggplant Fruit: Characterization of Biosynthetic Genes and Regulation by a MYB Transcription Factor. Front. Plant. Sci. 2015, 6, 1233. [Google Scholar] [CrossRef]
- Pandey, P.; Khan, F.; Rana, A.K.; Srivastava, Y. A drug repurposing approach towards elucidating the potential of flavonoids as COVID-19 spike protein inhibitors. Biointerface Res. Appl. Chem. 2021, 11, 8482–8501. [Google Scholar]
- Sukovic, D.; Knezevic, B.; Gasic, U.; Sredojevic, M.; Ciric, I.; Todic, S.; Mutic, J.; Tesic, Z. Phenolic Profiles of Leaves, Grapes and Wine of Grapevine Variety Vranac (Vitis vinifera L.) from Montenegro. Foods 2020, 9, 138. [Google Scholar] [CrossRef]
- Mouffouk, C.; Mouffouk, S.; Mouffouk, S.; Hambaba, L.; Haba, H. Flavonols as potential antiviral drugs targeting SARS-CoV-2 proteases (3CL(pro) and PL(pro)), spike protein, RNA-dependent RNA polymerase (RdRp) and angiotensin-converting enzyme II receptor (ACE2). Eur. J. Pharmacol. 2021, 891, 173759. [Google Scholar] [CrossRef] [PubMed]
- Yao, X.; Ye, F.; Zhang, M.; Cui, C.; Huang, B.; Niu, P.; Liu, X.; Zhao, L.; Dong, E.; Song, C.; et al. In Vitro Antiviral Activity and Projection of Optimized Dosing Design of Hydroxychloroquine for the Treatment of Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2). Clin. Infect. Dis. 2020, 71, 732–739. [Google Scholar] [CrossRef] [PubMed]
- Singh, M.; Zannella, C.; Folliero, V.; Di Girolamo, R.; Bajardi, F.; Chianese, A.; Altucci, L.; Damasco, A.; Del Sorbo, M.R.; Imperatore, C.; et al. Combating Actions of Green 2D-Materials on Gram Positive and Negative Bacteria and Enveloped Viruses. Front. Bioeng. Biotechnol. 2020, 8, 569967. [Google Scholar] [CrossRef] [PubMed]
- Ferhi, S.; Santaniello, S.; Zerizer, S.; Cruciani, S.; Fadda, A.; Sanna, D.; Dore, A.; Maioli, M.; D’Hallewin, G. Total Phenols from Grape Leaves Counteract Cell Proliferation and Modulate Apoptosis-Related Gene Expression in MCF-7 and HepG2 Human Cancer Cell Lines. Molecules 2019, 24, 612. [Google Scholar] [CrossRef]
- Hassan, W.H.B.; Abdelaziz, S.; Al Yousef, H.M. Chemical Composition and Biological Activities of the Aqueous Fraction of Parkinsonea aculeata L. Growing in Saudi Arabia. Arab. J. Chem. 2019, 12, 377–387. [Google Scholar] [CrossRef]
- Pluskal, T.; Castillo, S.; Villar-Briones, A.; Oresic, M. MZmine 2: Modular framework for processing, visualizing, and analyzing mass spectrometry-based molecular profile data. BMC Bioinform. 2010, 11, 395. [Google Scholar] [CrossRef]
- Wang, M.; Carver, J.J.; Phelan, V.V.; Sanchez, L.M.; Garg, N.; Peng, Y.; Nguyen, D.D.; Watrous, J.; Kapono, C.A.; Luzzatto-Knaan, T.; et al. Sharing and community curation of mass spectrometry data with Global Natural Products Social Molecular Networking. Nat. Biotechnol. 2016, 34, 828–837. [Google Scholar] [CrossRef]
- Vitale, G.A.; Sciarretta, M.; Cassiano, C.; Buonocore, C.; Festa, C.; Mazzella, V.; Nunez Pons, L.; D’Auria, M.V.; de Pascale, D. Molecular Network and Culture Media Variation Reveal a Complex Metabolic Profile in Pantoea cf. eucrina D2 Associated with an Acidified Marine Sponge. Int. J. Mol. Sci. 2020, 21, 6307. [Google Scholar] [CrossRef]
- Brito, A.; Ramirez, J.E.; Areche, C.; Sepulveda, B.; Simirgiotis, M.J. HPLC-UV-MS profiles of phenolic compounds and antioxidant activity of fruits from three citrus species consumed in Northern Chile. Molecules 2014, 19, 17400–17421. [Google Scholar] [CrossRef]
- Geng, P.; Sun, J.; Zhang, M.; Li, X.; Harnly, J.M.; Chen, P. Comprehensive characterization of C-glycosyl flavones in wheat (Triticum aestivum L.) germ using UPLC-PDA-ESI/HRMS(n) and mass defect filtering. J. Mass Spectrom. 2016, 51, 914–930. [Google Scholar] [CrossRef]
- Elsadig Karar, M.; Kuhnert, N. UPLC-ESI-Q-TOF-MS/MS Characterization of Phenolics from Crataegus monogyna and Crataegus laevigata (Hawthorn) Leaves, Fruits and their Herbal Derived Drops (Crataegutt Tropfen). J. Chem. Biol. Ther. 2016, 1, 102. [Google Scholar] [CrossRef]
- Spinola, V.; Llorent-Martinez, E.J.; Castilho, P.C. Antioxidant polyphenols of Madeira sorrel (Rumex maderensis): How do they survive to in vitro simulated gastrointestinal digestion? Food Chem. 2018, 259, 105–112. [Google Scholar] [CrossRef] [PubMed]
- Dueñas, M.; Mingo-Chornet, H.; Pérez-Alonso, J.; Di Paola, R.; Gonzalez-paramas, A.M.; Santos Buelga, C. Preparation of quercetin glucuronides and characterization by HPLC–DAD–ESI/MS. Eur. Food Res. Technol. 2008, 227, 1069–1076. [Google Scholar] [CrossRef]
- Chen, G.; Li, X.; Saleri, F.; Guo, M. Analysis of Flavonoids in Rhamnus davurica and Its Antiproliferative Activities. Molecules 2016, 21, 1275. [Google Scholar] [CrossRef]
- Roriz, C.L.; Barros, L.; Carvalho, A.M.; Santos-Buelga, C.; Ferreira, I.C. Scientific validation of synergistic antioxidant effects in commercialised mixtures of Cymbopogon citratus and Pterospartum tridentatum or Gomphrena globosa for infusions preparation. Food Chem. 2015, 185, 16–24. [Google Scholar] [CrossRef][Green Version]
- Marczak, Ł.; Znajdek-Awiżeń, P.; Bylka, W. The Use of Mass Spectrometric Techniques to Differentiate Isobaric and Isomeric Flavonoid Conjugates from Axyris amaranthoides. Molecules 2016, 21, 1229. [Google Scholar] [CrossRef]
- World Health Organization. Coronavirus Disease (COVID-19) Pandemic; World Health Organization: Geneva, Switzerland, 2020. [Google Scholar]
- Chianese, A.; Santella, B.; Ambrosino, A.; Stelitano, D.; Rinaldi, L.; Galdiero, M.; Zannella, C.; Franci, G. Oncolytic Viruses in Combination Therapeutic Approaches with Epigenetic Modulators: Past, Present, and Future Perspectives. Cancers 2021, 13, 2761. [Google Scholar] [CrossRef]
- Falanga, A.; Del Genio, V.; Kaufman, E.A.; Zannella, C.; Franci, G.; Weck, M.; Galdiero, S. Engineering of Janus-Like Dendrimers with Peptides Derived from Glycoproteins of Herpes Simplex Virus Type 1: Toward a Versatile and Novel Antiviral Platform. Int. J. Mol. Sci. 2021, 22, 6488. [Google Scholar] [CrossRef]
- Monda, V.; Valenzano, A.; Monda, M. Modifications of Activity of Autonomic Nervous System, and Resting Energy Expenditure in Women Using Hormone-Replacement Therapy. Biol. Med. 2016, 8, 1. [Google Scholar]
- Schiattarella, A.; Riemma, G.; La Verde, M.; Franci, G.; Chianese, A.; Fasulo, D.; Fichera, M.; Gallo, P.; De Franciscis, P. Polycystic ovary syndrome and probiotics: A natural approach to an inflammatory disease. Curr. Womens Health Rev. 2021, 17, 14–20. [Google Scholar] [CrossRef]
- Russo, M.; Moccia, S.; Spagnuolo, C.; Tedesco, I.; Russo, G.L. Roles of flavonoids against coronavirus infection. Chem. Biol. Interact. 2020, 328, 109211. [Google Scholar] [CrossRef]
- Urmenyi, F.G.; Saraiva, G.D.; Casanova, L.M.; Matos, A.D.; de Magalhaes Camargo, L.M.; Romanos, M.T.; Costa, S.S. Anti-HSV-1 and HSV-2 Flavonoids and a New Kaempferol Triglycoside from the Medicinal Plant Kalanchoe daigremontiana. Chem. Biodivers. 2016, 13, 1707–1714. [Google Scholar] [CrossRef]
- Abdallah, H.M.; El-Halawany, A.M.; Sirwi, A.; El-Araby, A.M.; Mohamed, G.A.; Ibrahim, S.R.M.; Koshak, A.E.; Asfour, H.Z.; Awan, Z.A. Repurposing of Some Natural Product Isolates as SARS-COV-2 Main Protease Inhibitors via In Vitro Cell Free and Cell-Based Antiviral Assessments and Molecular Modeling Approaches. Pharmaceuticals 2021, 14, 213. [Google Scholar] [CrossRef]
- Conceicao, C.; Thakur, N.; Human, S.; Kelly, J.T.; Logan, L.; Bialy, D.; Bhat, S.; Stevenson-Leggett, P.; Zagrajek, A.K.; Hollinghurst, P.; et al. The SARS-CoV-2 Spike protein has a broad tropism for mammalian ACE2 proteins. PLoS Biol. 2020, 18, e3001016. [Google Scholar] [CrossRef]
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Kruger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.H.; Nitsche, A.; et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell 2020, 181, 271–280.e8. [Google Scholar] [CrossRef]
- Zhao, P.; Praissman, J.L.; Grant, O.C.; Cai, Y.; Xiao, T.; Rosenbalm, K.E.; Aoki, K.; Kellman, B.P.; Bridger, R.; Barouch, D.H.; et al. Virus-Receptor Interactions of Glycosylated SARS-CoV-2 Spike and Human ACE2 Receptor. Cell Host Microbe 2020, 28, 586–601.e6. [Google Scholar] [CrossRef]
- Jain, A.S.; Sushma, P.; Dharmashekar, C.; Beelagi, M.S.; Prasad, S.K.; Shivamallu, C.; Prasad, A.; Syed, A.; Marraiki, N.; Prasad, K.S. In silico evaluation of flavonoids as effective antiviral agents on the spike glycoprotein of SARS-CoV-2. Saudi J. Biol. Sci. 2021, 28, 1040–1051. [Google Scholar] [CrossRef] [PubMed]
- Taban, A.K.; Süntar, İ. An Overview on Flavonoids as Potential Antiviral Strategies against Coronavirus Infections. Gazi Med. J. 2020. [Google Scholar] [CrossRef]
- Effect of Quercetin on Prophylaxis and Treatment of COVID-19. Available online: https://clinicaltrials.gov/ct2/show/NCT04377789 (accessed on 28 June 2021).
- Grace Nirmala, J.; Evangeline Celsia, S.; Swaminathan, A.; Narendhirakannan, R.T.; Chatterjee, S. Cytotoxicity and apoptotic cell death induced by Vitis vinifera peel and seed extracts in A431 skin cancer cells. Cytotechnology 2018, 70, 537–554. [Google Scholar] [CrossRef] [PubMed]
- Abed, A.; Harb, J.; Khasib, S.; Saad, B. In vitro assessment of cytotoxic, antioxidant and antimicrobial activities of leaves from two grape varieties collected from arid and temperate regions in Palestine. QSci. Connect 2015, 2015, 4. [Google Scholar]
| Gene | Sequence |
|---|---|
| HSV-1 UL27 forward | GCCTTCTTCGCCTTTCGC |
| HSV-1 UL27 reverse | CGCTCGTGCCCTTCTTCTT |
| SARS-CoV-2 S forward | AGGTTGATCACAGGCAGACT |
| SARS-CoV-2 S reverse | GCTGACTGAGGGAAGGAC |
| GAPDH forward | CCTTTCATTGAGCTCCAT |
| GAPDH reverse | CGTACATGGGAGCGTC |
| n. | Precursor Mass [M − H]− | Rt (min) | Putative Compound | Key Fragments | Relative Abundance |
|---|---|---|---|---|---|
| Polyphenols | |||||
| 1 | 482.96 | 5.07 | 1-O,6-O-Digalloyl-β-D-glucopyranose | 483; 301; 312; 271; 211 | Traces |
| 2 | 447.22 | 6.34 | Kaempferol-8-C-glucoside | 357; 327; 297; 285 | Traces |
| 3 | 789.63 | 6.63 | N.I. | 639; 613; 581; 477; 465; 463 | Traces |
| 4 | 447.16 | 7.60 | Luteolin-8-C-glucoside (Orientin) | 357; 339; 327; 299; 298; 297; 285 | Traces |
| 5 | 478.94 | 8.39 | Myricetin-3-O-beta-D-galactopyranoside | 479; 317; 316; 271; 270 | 0.1% |
| 6 | 447.03 | 8.47 | Luteolin-6-C-glucoside (Isoorientin) | 357; 339; 327; 299; 298; 297; 285 | 16.0% |
| 7 | 561.36 | 8.73 | N.I. | 449; 447; 357; 327; 301; 297 | Traces |
| 8 | 447.05 | 8.88 | Luteolin-6-C-glucoside (Isoorientin) | 357; 327; 299; 297; 285; | 4.9% |
| 9 | 447.00 | 9.16 | Luteolin-4-O-glucoside | 447; 327; 285; 284 | Traces |
| 10 | 431.03 | 9.45 | Apigenin-8-C-glucoside (Vitexin) | 377; 353; 341; 323; 311; 283; 282; 268 | 0.7% |
| 11 | 431.06 | 9.85 | Apigenin-6-C-glucoside (Isovitexin) | 341; 323; 311; 293; 283; 281; 269 | 13.2% |
| 12 | 499.07 | 10.02 | N.I. | 499; 323; 301; 300 | 0.1% |
| 13 | 477.03 | 10.10 | Quercetin-O-glucuronide | 477; 301; | 44.2% |
| 14 | 463.07 | 10.30 | Quercetin-3-O-glucoside (Isoquercitrin) | 463; 301; 300; 271; 255 | 3.3% |
| 15 | 447.06 | 10.62 | Biochanin A-7-glucoside (Astragaloside) | 447; 327; 285; 284 | 4.9% |
| 16 | 461.06 | 10.81 | Chrysoeriol 8-C-glucoside (Scoparin) | 371; 341; 326; 313; 299; 298 | Traces |
| 17 | 477.09 | 10.83 | Quercetin 3-O-glucuronide | 477; 302; 301; 214 | Traces |
| 18 | 447.00 | 10.95 | Luteolin-7-O-glucoside (Cynaroside) | 447; 327; 285; 284 | Traces |
| 19 | 505.57 | 11.04 | Quercetin-3-O-glucose-6″-acetate | 505; 301; 300; 271; 255 | Traces |
| 20 | 447.09 | 11.27 | Kaempferol-7-O-glucoside | 447; 357; 327; 285; 284; 255; 256; 227 | 0.3% |
| 21 | 505.37 | 11.29 | Quercetin-O-Acetyl hexoside | 302; 301; 300; 271; 255 | Traces |
| 22 | 462.98 | 11.31 | Quercetin-3-O-galactoside (Hyperoside) | 463; 301; 300; 271 | Traces |
| 23 | 593.15 | 11.54 | Kaempferol-3-O-(6-O-rhamnosyl-galactoside) | 285; 284; 257; 255 | Traces |
| 24 | 483.20 | 11.63 | Luteolin-C-hexoside | 483; 271 | Traces |
| 25 | 461.05 | 11.86 | Kaempferol 3-O-glucuronide | 285; 257; 228 | 0.6% |
| 26 | 447.08 | 11.94 | Kaempferol-3-O-glucoside | 447; 327; 284; 285; 255; 256; 227 | 2.4% |
| 27 | 517.06 | 12.35 | N.I. | 517; 355; 341 | Traces |
| 28 | 491.09 | 12.46 | Isorhamnetin-3(7)-O-glucuronopyranoside | 491; 315; 300 | 8.8% |
| 29 | 431.01 | 12.95 | Apigenin-7-O-glucoside | 431; 283 | Traces |
| 31 | 463.11 | 13.45 | Quercetin-7-O-glucoside | 300; 301 | Traces |
| 32 | 475.06 | 14.34 | 6-O-Methylscutellarin | 285; 284; 255; 256 | 0.3% |
| 33 | 639.15 | 15.22 | Quercetin-O-glucuronide-O-hexoside | 463; 301 | Traces |
| 34 | 477.06 | 16.24 | Isorhamnetin-O-glucoside | 477; 315; 314 | Traces |
| 35 | 461.37 | 18.96 | 4-(3,4-Dihydroxyphenyl)-5-beta-D-glucopyranosyloxy-7-methoxycoumarin | 299;284 | Traces |
| 36 | 449.26 | 19.90 | Isookanin-7-O-glucoside | 449; 431; 287; 269 | Traces |
| Digalactosylmonoacylglycerols | |||||
| 37 | 675.35 | 24.81 | DGMG 18:3 | 415; 397; 277; 235 | 97.1% |
| 38 | 699.34 | 25.54 | DGMG 20:5 | 653; 415; 397; 323; 255; 235 | Traces |
| 39 | 653.81 | 25.55 | DMGM 16:2 | 653; 415; 397; 277; 255; 235 | 2.9% |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zannella, C.; Giugliano, R.; Chianese, A.; Buonocore, C.; Vitale, G.A.; Sanna, G.; Sarno, F.; Manzin, A.; Nebbioso, A.; Termolino, P.; et al. Antiviral Activity of Vitis vinifera Leaf Extract against SARS-CoV-2 and HSV-1. Viruses 2021, 13, 1263. https://doi.org/10.3390/v13071263
Zannella C, Giugliano R, Chianese A, Buonocore C, Vitale GA, Sanna G, Sarno F, Manzin A, Nebbioso A, Termolino P, et al. Antiviral Activity of Vitis vinifera Leaf Extract against SARS-CoV-2 and HSV-1. Viruses. 2021; 13(7):1263. https://doi.org/10.3390/v13071263
Chicago/Turabian StyleZannella, Carla, Rosa Giugliano, Annalisa Chianese, Carmine Buonocore, Giovanni Andrea Vitale, Giuseppina Sanna, Federica Sarno, Aldo Manzin, Angela Nebbioso, Pasquale Termolino, and et al. 2021. "Antiviral Activity of Vitis vinifera Leaf Extract against SARS-CoV-2 and HSV-1" Viruses 13, no. 7: 1263. https://doi.org/10.3390/v13071263
APA StyleZannella, C., Giugliano, R., Chianese, A., Buonocore, C., Vitale, G. A., Sanna, G., Sarno, F., Manzin, A., Nebbioso, A., Termolino, P., Altucci, L., Galdiero, M., de Pascale, D., & Franci, G. (2021). Antiviral Activity of Vitis vinifera Leaf Extract against SARS-CoV-2 and HSV-1. Viruses, 13(7), 1263. https://doi.org/10.3390/v13071263










