Genetic Diversity of Enteric Viruses in Children under Five Years Old in Gabon
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.2. Viral RNA Extraction
2.3. PCR Detection and Genotyping
2.4. Nucleotide Sequencing and Phylogenetic Analysis
2.5. Statistical Analysis
3. Results
3.1. Study Population
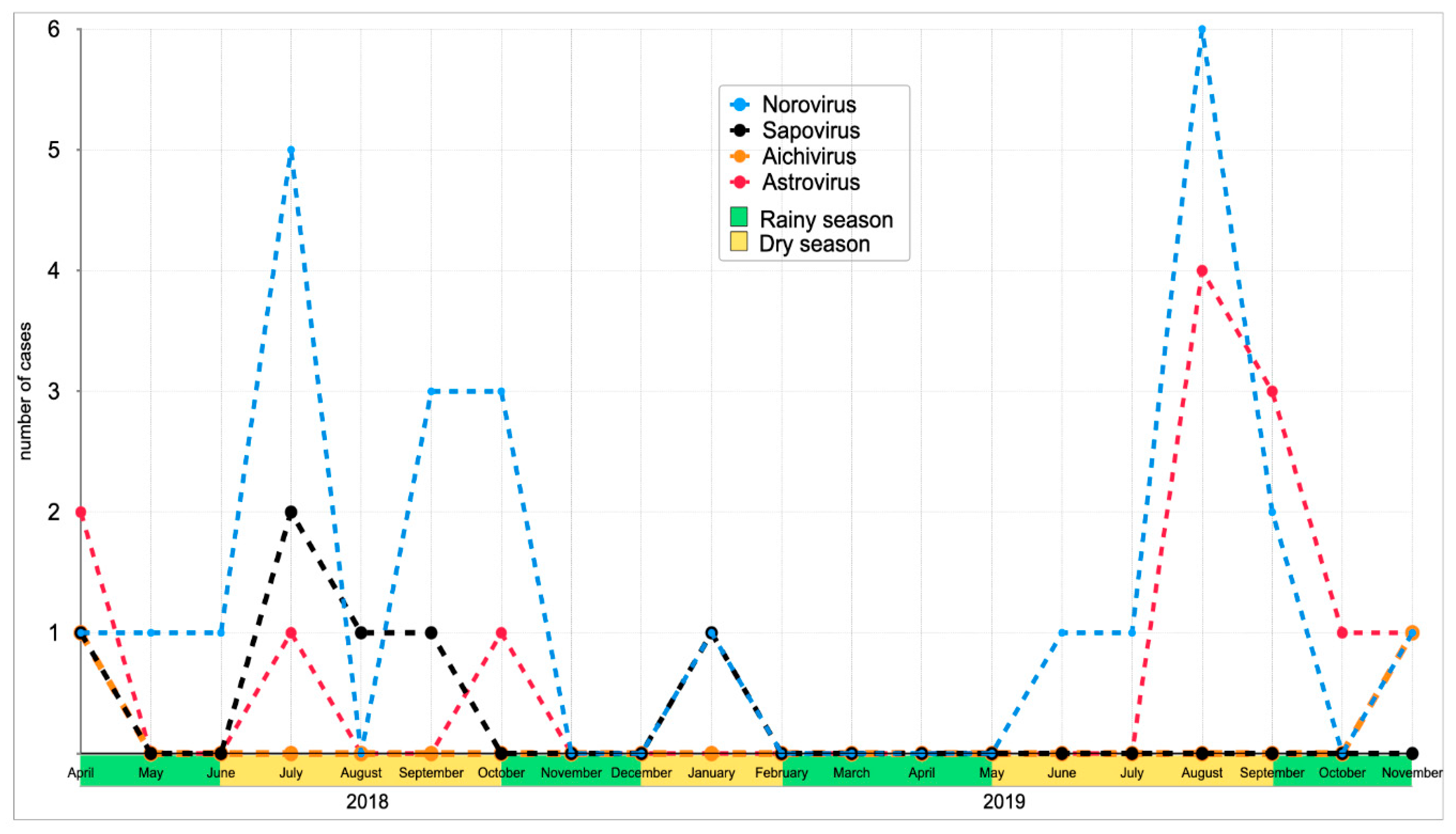
3.2. Detection Rate of Enteric Viruses in the Study Population
3.3. Sequence and Phylogenetic Analyses of Gastroenteritis Viruses
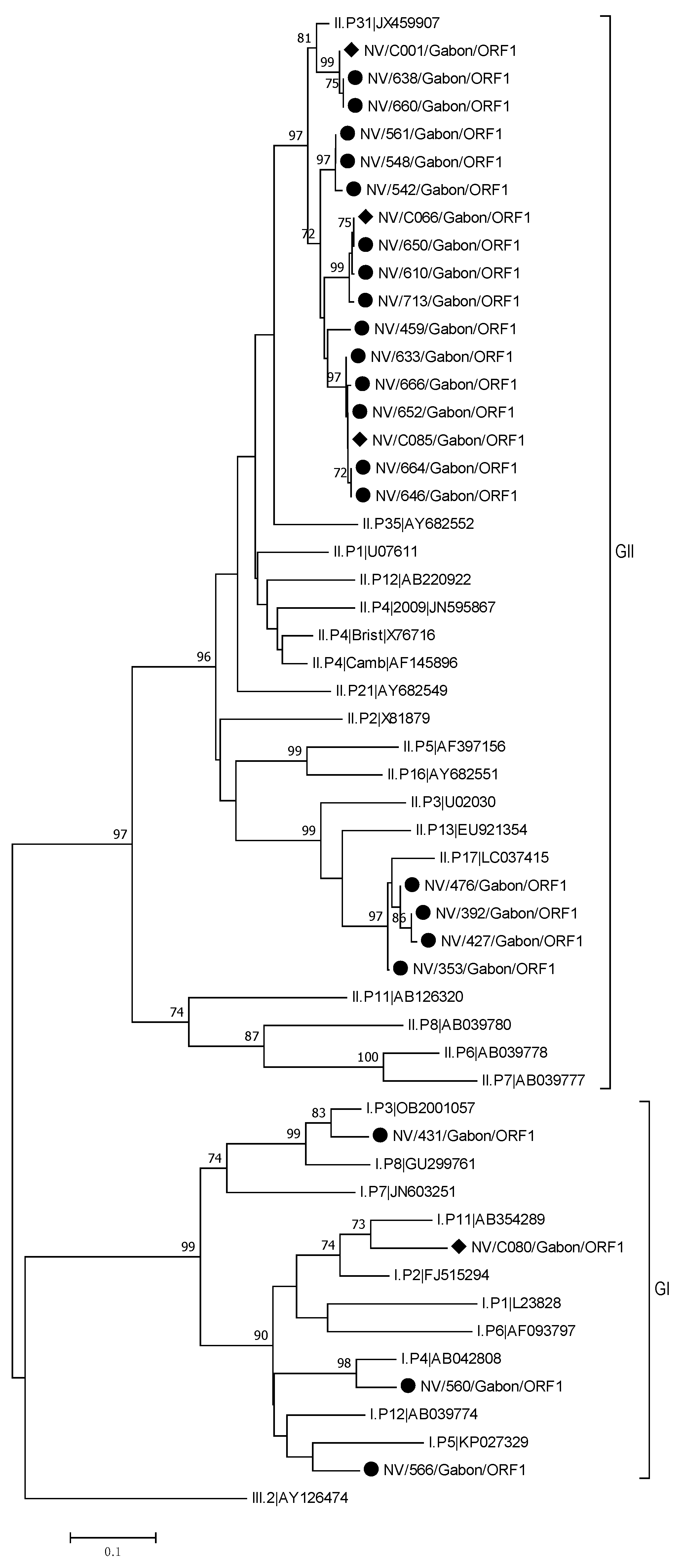
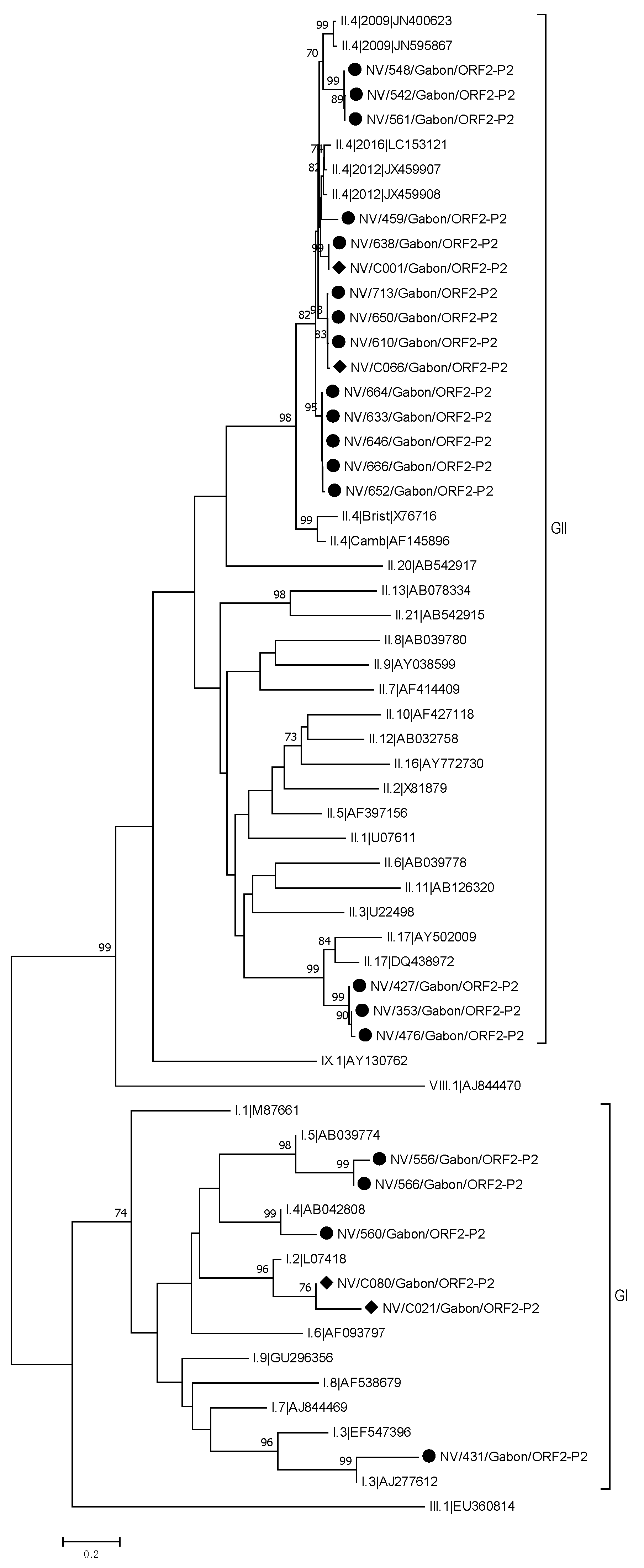
3.3.1. Noroviruses
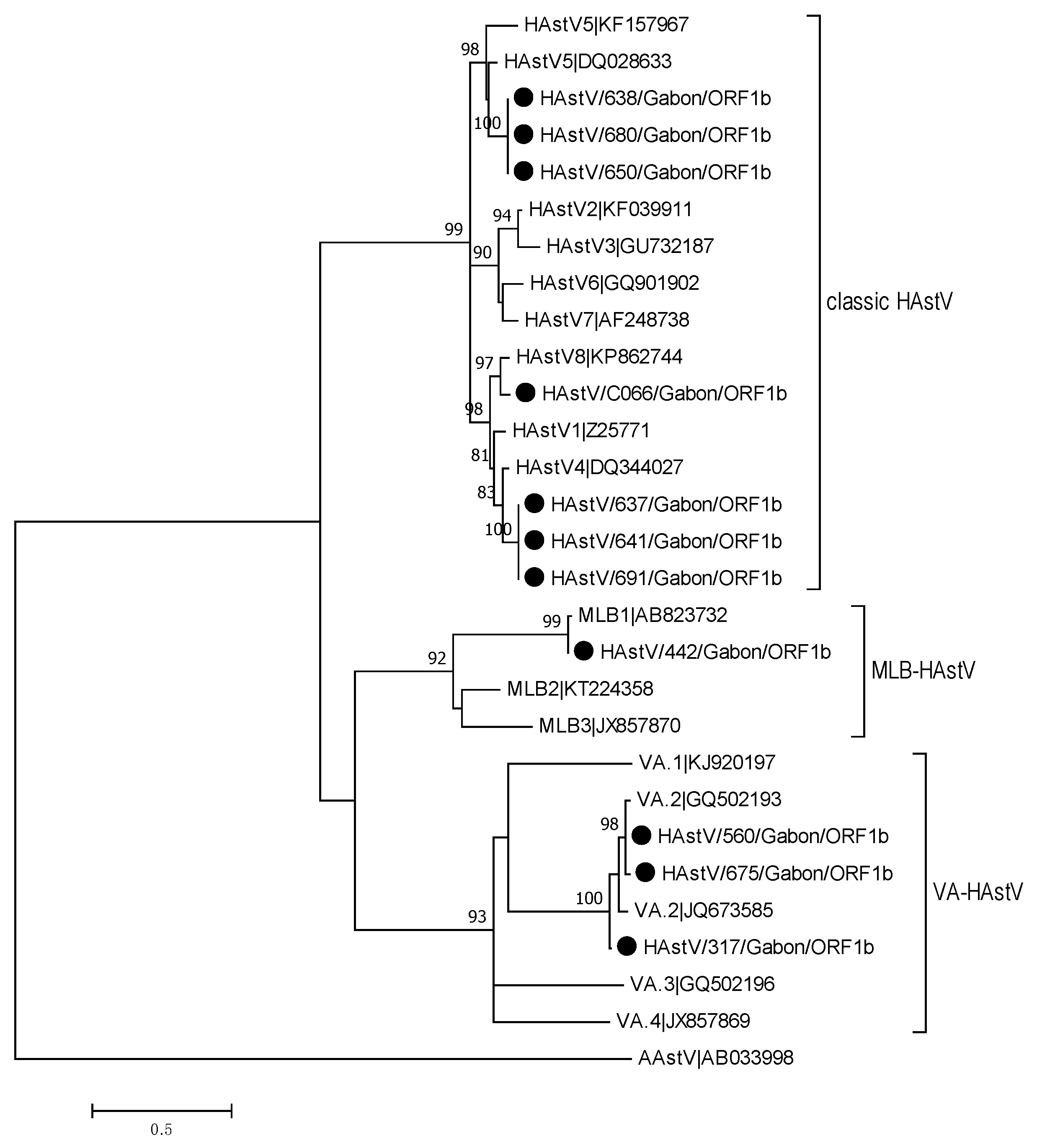
3.3.2. Astroviruses
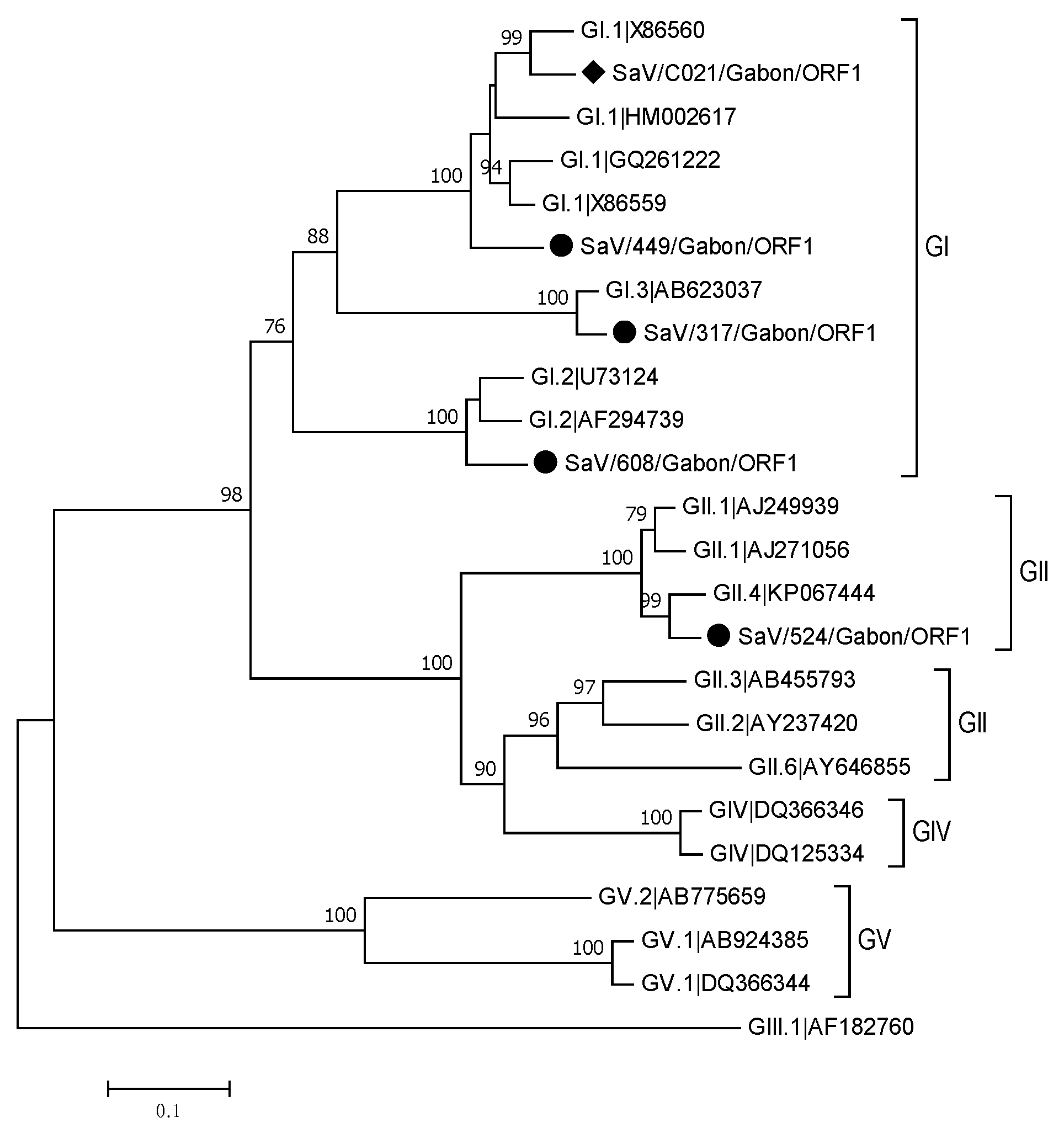
3.3.3. Sapoviruses
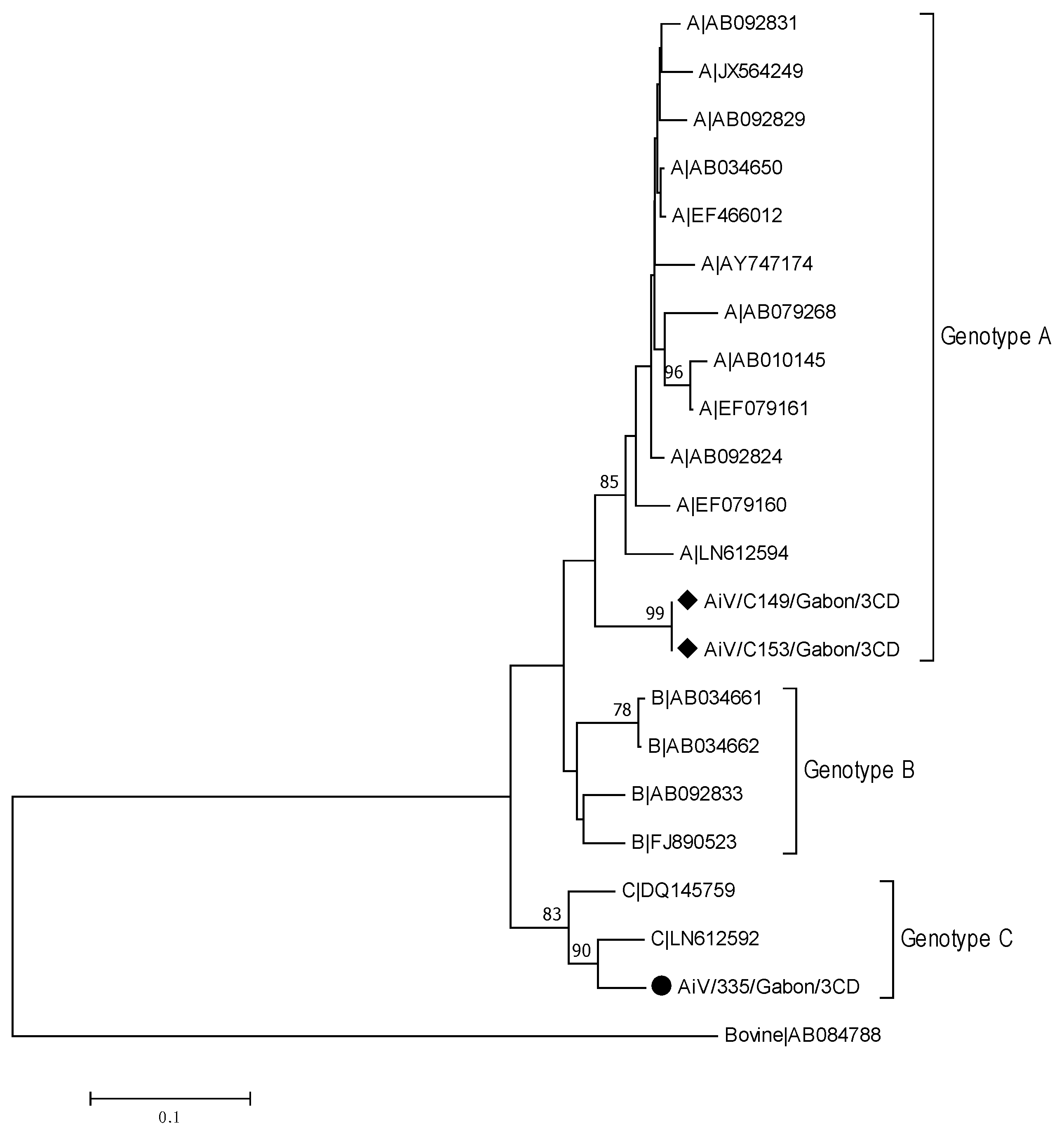
3.3.4. Aichiviruses
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Gautam, R.; Lyde, F.; Esona, M.D.; Quaye, O.; Michael, D.; Viruses, R. Detection of Rotavirus Antigen in Stool Specimens. J. Clin. Virol. 2013, 58, 292–294. [Google Scholar] [CrossRef]
- Black, R.E.; Cousens, S.; Johnson, H.L.; Lawn, J.E.; Rudan, I.; Bassani, D.G.; Jha, P.; Campbell, H.; Walker, C.F.; Cibulskis, R.; et al. Global, regional, and national causes of child mortality in 2008: A systematic analysis. Lancet 2010, 375, 1969–1987. [Google Scholar] [CrossRef]
- Li, V. The Epidemiology of Noroviruses in Ghana: A Case Study of Norovirus Detection. J. Glob. Health 2013, 3, 11–14. [Google Scholar]
- Corcoran, M.S.; Van Well, G.T.J.; Van Loo, I.H.M. Diagnosis of viral gastroenteritis in children: Interpretation of real-time PCR results and relation to clinical symptoms. Eur. J. Clin. Microbiol. Infect. Dis. 2014, 33, 1663–1673. [Google Scholar] [CrossRef]
- Sdiri-Loulizi, K.; Gharbi-Khélifi, H.; De Rougemont, A.; Chouchane, S.; Sakly, N.; Ambert-Balay, K.; Hassine, M.; Guédiche, M.N.; Aouni, M.; Pothier, P. Acute infantile gastroenteritis associated with human enteric viruses in Tunisia. J. Clin. Microbiol. 2008, 46, 1349–1355. [Google Scholar] [CrossRef]
- Ouyang, Y.; Ma, H.; Jin, M.; Wang, X.; Wang, J.; Xu, L.; Lin, S.; Shen, Z.; Chen, Z.; Qiu, Z.; et al. Etiology and epidemiology of viral diarrhea in children under the age of five hospitalized in Tianjin, China. Arch. Virol. 2012, 157, 881–887. [Google Scholar] [CrossRef] [PubMed]
- Parashar, U.D.; Hummelman, E.G.; Bresee, J.S.; Miller, M.A.; Glass, R.I. Global illness and deaths caused by rotavirus disease in children. Emerg. Infect. Dis. 2003, 9, 565–572. [Google Scholar] [CrossRef] [PubMed]
- Tate, J.E.; Burton, A.H.; Boschi-Pinto, C.; Parashar, U.D. Global, Regional, and National Estimates of Rotavirus Mortality in Children <5 Years of Age, 2000–2013. Clin. Infect. Dis. 2016, 62 (Suppl. 2), S96–S105. [Google Scholar] [CrossRef]
- Arnold, M.M. Rotavirus Vaccines: Why Continued Investment in Research Is Necessary. Curr. Clin. Microbiol. Rep. 2018, 5, 73–81. [Google Scholar] [CrossRef]
- da Silva Poló, T.; Peiró, J.R.; Mendes, L.C.N.; Ludwig, L.F.; De Oliveira-Filho, E.F.; Bucardo, F.; Huynen, P.; Melin, P.; Thiry, E.; Mauroy, A. Human norovirus infection in Latin America. J. Clin. Virol. 2016, 78, 111–119. [Google Scholar] [CrossRef]
- Bartsch, S.M.; Lopman, B.A.; Ozawa, S.; Hall, A.J.; Lee, B.Y. Global economic burden of norovirus gastroenteritis. PLoS ONE 2016, 11, e0151219. [Google Scholar] [CrossRef]
- Carmona-Vicente, N.; Vila-Vicent, S.; Allen, D.; Gozalbo-Rovira, R.; Iturriza-Gómara, M.; Buesa, J.; Rodríguez-Díaz, J. Characterization of a Novel Conformational GII.4 Norovirus Epitope: Implications for Norovirus-Host Interactions. J. Virol. 2016, 90, 7703–7714. [Google Scholar]
- Robilotti, E.; Deresinski, S.; Pinsky, B.A. Norovirus. Clin. Microbiol. Rev. 2015, 28, 134–164. [Google Scholar] [CrossRef] [PubMed]
- Chhabra, P.; de Graaf, M.; Parra, G.I.; Chan, M.C.-W.; Green, K.; Martella, V.; Wang, Q.; White, P.A.; Katayama, K.; Vennema, H.; et al. Erratum: Updated classification of norovirus genogroups and genotypes (Journal of General Virology (2020) 100 (1393–1406) doi:10.1099/jgv.0.001318). J. Gen. Virol. 2020, 101, 893. [Google Scholar] [CrossRef]
- Tohma, K.; Lepore, C.J.; Gao, Y.; Ford-Siltz, L.A.; Parra, G.I. Population Genomics of GII.4 Noroviruses Reveal Complex Diversification and New Antigenic Sites Involved in the Emergence of Pandemic Strains. mBio 2019, 10, e02202-19. [Google Scholar] [CrossRef] [PubMed]
- Johnson, C.; Hargest, V.; Cortez, V.; Meliopoulos, V.A.; Schultz-Cherry, S. Astrovirus pathogenesis. Viruses 2017, 9, 22. [Google Scholar] [CrossRef]
- Bosch, A.; Pintó, R.M.; Guix, S. Human Astroviruses. Clin. Microbiol. Rev. 2014, 27, 1048–1074. [Google Scholar] [CrossRef]
- Vu, D.-L.; Bosch, A.; Pintó, R.M.; Guix, S. Epidemiology of classic and novel human astrovirus: Gastroenteritis and beyond. Viruses 2017, 9, 33. [Google Scholar] [CrossRef]
- Magwalivha, M.; Kabue, J.-P.; Traore, A.N.; Potgieter, N. Prevalence of Human Sapovirus in Low and Middle Income Countries. Adv. Virol. 2018, 2018, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Oka, T.; Wang, Q.; Katayama, K.; Saif, L.J. Comprehensive review of human sapoviruses. Clin. Microbiol. Rev. 2015, 28, 32–53. [Google Scholar] [CrossRef]
- Ambert-Balay, K.; Lorrot, M.; Bon, F.; Giraudon, H.; Kaplon, J.; Wolfer, M.; Lebon, P.; Gendrel, D.; Pothier, P. Prevalence and genetic diversity of Aichi virus strains in stool samples from community and hospitalized patients. J. Clin. Microbiol. 2008, 46, 1252–1258. [Google Scholar] [CrossRef] [PubMed]
- Taghinejad, M.; Ghaderi, M.; Mousavi-Nasab, S.D. Aichivirus with acute gastroenteritis in Iran. Pediatric Infect. Dis. J. 2020, 39, 576–579. [Google Scholar] [CrossRef] [PubMed]
- Kotloff, K.L. The Burden and Etiology of Diarrheal Illness in Developing Countries. Pediatric Clin. N. Am. 2017, 64, 799–814. [Google Scholar] [CrossRef]
- Mans, J.; Armah, G.E.; Steele, A.D.; Taylor, M.B. Norovirus epidemiology in Africa: A review. PLoS ONE 2016, 11, e0146280. [Google Scholar] [CrossRef]
- Lekana-Douki, S.E.; Kombila-Koumavor, C.; Nkoghe, D.; Drosten, C.; Drexler, J.F.; Leroy, E.M. Molecular epidemiology of enteric viruses and genotyping of rotavirus A, adenovirus and astrovirus among children under 5 years old in Gabon. Int. J. Infect. Dis. 2015, 34, 90–95. [Google Scholar] [CrossRef]
- Niendorf, S.; Faber, M.; Tröger, A.; Hackler, J.; Jacobsen, S. Diversity of noroviruses throughout outbreaks in Germany 2018. Viruses 2020, 12, 1157. [Google Scholar] [CrossRef] [PubMed]
- Jacobsen, S.; Höhne, M.; Marques, A.M.; Beslmüller, K.; Bock, C.-T.; Niendorf, S. Co-circulation of classic and novel astrovirus strains in patients with acute gastroenteritis in Germany. J. Infect. 2018, 76, 457–464. [Google Scholar] [CrossRef] [PubMed]
- Höhne, M.; Niendorf, S.; Marques, A.M.; Bock, C.-T. Use of sequence analysis of the P2 domain for characterization of norovirus strains causing a large multistate outbreak of norovirus gastroenteritis in Germany 2012. Int. J. Med. Microbiol. 2015, 305, 612–618. [Google Scholar] [CrossRef]
- Oh, D.-Y.; Silva, P.A.; Hauroeder, B.; Diedrich, S.; Cardoso, D.D.P.; Schreier, E. Molecular characterization of the first Aichi viruses isolated in Europe and in South America. Arch. Virol. 2006, 151, 1199–1206. [Google Scholar] [CrossRef]
- Japhet, M.O.; Famurewa, O.; Adesina, O.; Opaleye, O.; Wang, B.; Höhne, M.; Bock, C.; Marques, A.M.; Niendorf, S. Viral gastroenteritis among children of 0–5 years in Nigeria: Characterization of the first Nigerian aichivirus, recombinant noroviruses and detection of a zoonotic astrovirus. J. Clin. Virol. 2019, 111, 4–11. [Google Scholar] [CrossRef]
- Shen, X.-X.; Qiu, F.-Z.; Li, G.-X.; Zhao, M.-C.; Wang, J.; Chen, C.; Zhao, L.; Qi, J.-J.; Liu, H.; Zhang, Y.; et al. A case control study on the prevalence of enterovirus in children samples and its association with diarrhea. Arch. Virol. 2018, 164, 63–68. [Google Scholar] [CrossRef] [PubMed]
- Han, T.-H.; Park, S.H.; Hwang, E.-S.; Reuter, G.; Chung, J.-Y. Detection of Aichi virus in South Korea. Arch. Virol. 2014, 159, 1835–1839. [Google Scholar] [CrossRef] [PubMed]
- Ayukekbong, J.; Lindh, M.; Nenonen, N.; Tah, F.; Nkuo-Akenji, T.; Bergström, T. Enteric viruses in healthy children in cameroon: Viral load and genotyping of norovirus strains. J. Med. Virol. 2011, 83, 2135–2142. [Google Scholar] [CrossRef] [PubMed]
- Arowolo, K.O.; Ayolabi, C.I.; Lapinski, B.; Santos, J.S.; Raboni, S.M. Epidemiology of enteric viruses in children with gastroenteritis in Ogun State, Nigeria. J. Med. Virol. 2019, 91, 1022–1029. [Google Scholar] [CrossRef] [PubMed]
- Leblanc, D.; Inglis, G.D.; Boras, V.F.; Brassard, J.; Houde, A. The prevalence of enteric RNA viruses in stools from diarrheic and non-diarrheic people in southwestern Alberta, Canada. Arch. Virol. 2017, 162, 117–128. [Google Scholar] [CrossRef]
- Cannon, J.L.; Lopman, B.A.; Payne, D.C.; Vinjé, J. Birth Cohort Studies Assessing Norovirus Infection and Immunity in Young Children: A Review. Clin. Infect. Dis. 2019, 69, 357–365. [Google Scholar] [CrossRef]
- Louya, V.M.; Vouvoungui, C.; Koukouikila-Koussounda, F.; Veas, F.; Kobawila, S.C.; Ntoumi, F.; Mikounou, V.; Charles, K.S. Molecular characterization of norovirus infection responsible for acute diarrhea in Congolese hospitalized children under five years old in Brazzaville, Republic of Congo. Int. J. Infect. Dis. 2019, 88, 41–48. [Google Scholar] [CrossRef]
- Wu, C.Y.; Chi, H.; Liu, C.-C.; Huang, Y.-C.; Huang, Y.-C.; Lin, H.-C.; Ho, Y.-H.; Huang, L.-M.; Huang, C.-Y.; Shih, S.-M.; et al. Clinical characteristics and risk factors for children with norovirus gastroenteritis in Taiwan. J. Microbiol. Immunol. Infect. 2020. [Google Scholar] [CrossRef]
- Ouedraogo, N.; Ngangas, S.M.T.; Bonkoungou, I.J.O.; Tiendrebeogo, A.B.; Traore, K.A.; Sanou, I.; Traore, A.S.; Barro, N. Temporal distribution of gastroenteritis viruses in Ouagadougou, Burkina Faso: Seasonality of rotavirus. BMC Public Health 2017, 17, 1–8. [Google Scholar] [CrossRef]
- Ouédraogo, N.; Kaplon, J.; Bonkoungou, I.J.O.; Traoré, A.S.; Pothier, P.; Barro, N.; Balay, K.A. Prevalence and genetic diversity of enteric viruses in children with diarrhea in Ouagadougou, Burkina Faso. PLoS ONE 2016, 11, e0153652. [Google Scholar] [CrossRef]
- Bucardo, F.; Reyes, Y.; Svensson, L.; Nordgren, J. Predominance of norovirus and sapovirus in nicaragua after implementation of universal rotavirus vaccination. PLoS ONE 2014, 9, e98201. [Google Scholar] [CrossRef]
- El Qazoui, M.; Oumzil, H.; Baassi, L.; El Omari, N.; Sadki, K.; Amzazi, S.; Benhafid, M.; El Aouad, R. Rotavirus and Norovirus infections among acute gastroenteritis children in Morocco. BMC Infect. Dis. 2014, 14, 300. [Google Scholar] [CrossRef] [PubMed]
- Kiulia, N.M.; Mans, J.; Mwenda, J.M.; Taylor, M.B. Norovirus GII.17 Predominates in Selected Surface Water Sources in Kenya. Food Environ. Virol. 2014, 6, 221–231. [Google Scholar] [CrossRef] [PubMed]
- Kuang, X.; Teng, Z.; Zhang, X. Genotypic prevalence of norovirus GII in gastroenteritis outpatients in Shanghai from 2016 to 2018. Gut Pathog. 2019, 11, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Guarines, K.M.; Mendes, R.P.G.; de Magalhães, J.J.F.; Pena, L. Norovirus-associated gastroenteritis, Pernambuco, Northeast Brazil, 2014–2017. J. Med. Virol. 2020, 92, 1093–1101. [Google Scholar] [CrossRef]
- Hasing, M.E.; Lee, B.E.; Qiu, Y.; Xia, M.; Pabbaraju, K.; Wong, A.; Tipples, G.; Jiang, X.; Pang, X.L. Changes in norovirus genotype diversity in gastroenteritis outbreaks in Alberta, Canada: 2012–2018. BMC Infect. Dis. 2019, 19, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Hernandez, J.M.; Silva, L.D.; Junior, E.C.S.; Cardoso, J.F.; Reymão, T.K.A.; Portela, A.C.R.; De Lima, C.P.S.; Teixeira, D.M.; Lucena, M.S.S.; Nunes, M.R.T.; et al. Evolutionary and Molecular Analysis of Complete Genome Sequences of Norovirus From Brazil: Emerging Recombinant Strain GII.P16/GII.4. Front. Microbiol. 2020, 11, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Becker-Dreps, S.; Bucardo, F.; Vinjé, J. Sapovirus: An important cause of acute gastroenteritis in children. Lancet Child Adolesc. Health 2019, 3, 758–759. [Google Scholar] [CrossRef]
- Kaikkonen, S.; Räsänen, S.; Rämet, M.; Vesikari, T. Aichi virus infection in children with acute gastroenteritis in Finland. Epidemiol. Infect. 2010, 138, 1166–1171. [Google Scholar] [CrossRef]
- Nielsen, A.C.Y.; Gyhrs, M.L.; Nielsen, L.P.; Pedersen, C.; Böttiger, B. Gastroenteritis and the novel picornaviruses aichi virus, cosavirus, saffold virus, and salivirus in young children. J. Clin. Virol. 2013, 57, 239–242. [Google Scholar] [CrossRef]
- Jonsson, N.; Wahlström, K.; Svensson, L.; Serrander, L.; Lindberg, A.M. Aichi virus infection in elderly people in Sweden. Arch. Virol. 2012, 157, 1365–1369. [Google Scholar] [CrossRef] [PubMed]
- Rivadulla, E.; Romalde, J.L. Correction to: A Comprehensive Review on Human Aichi Virus. Virol. Sin. 2020, 12250, 15782. [Google Scholar]
- Widdowson, M.-A.; Sulka, A.; Bulens, S.N.; Beard, R.S.; Chaves, S.S.; Hammond, R.; Salehi, E.D.; Swanson, E.; Totaro, J.; Woron, R.; et al. Norovirus and foodborne disease, United States, 1991–2000. Emerg. Infect. Dis. 2005, 11, 95–102. [Google Scholar] [CrossRef] [PubMed]
| Virus | Diarrhea (n = 177) | No Diarrhea (n = 67) | p-Value | Total (n = 244) |
|---|---|---|---|---|
| NoV | 26 (14.7%) | 6 (9.0%) | 0.2764 | 32 (13.1%) |
| NoV GI | 6 (3.4%) | 3 (4.5%) | 1.0000 | 9 (3.7%) |
| NoV GII | 20 (11.3%) | 3 (4.5%) | 0.1046 | 23 (9.4%) |
| AstV | 13(7.3%) | 3 (4.5%) | 0.5371 | 16 (6.6%) |
| SaV | 6 (3.4%) | 1 (1.5%) | 0.6212 | 7 (2.9%) |
| AiV | 2 (1.1%) | 4 (6.0%) | 0.1184 | 6 (2.5%) |
| Diarrhea Cases | ||||||
| NoV n (%) | NoV GI n (%) | NoV GII n (%) | AstV n (%) | SaV n (%) | AiV n (%) | |
| Age group (Months) | ||||||
| 0–6 (n = 47) | 11 (23.4) | 1 (2.1) | 10 (21.3) | 4 (8.5) | 0 (0) | 0 (0) |
| 7–12 (n = 47) | 8 (17.0) | 3 (6.4) | 5 (10.6) | 6 (12.8%) | 1 (2.1) | 2 (4.3) |
| 13–18 (n = 45) | 5 (11.1) | 2 (4.4) | 3 (6.7) | 2 (4.4) | 2 (4.4) | 0 (0) |
| 19–24 (n = 16) | 2 (12.5) | 0 (0) | 2 (12.5) | 1 (6.3) | 3 (18.8) | 0 (0) |
| 25–59 (n = 22) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) |
| Gender | ||||||
| F (n = 76) | 13 (17.1) | 2 (2.6) | 11 (14.5) | 9 (11.8) | 4 (5.3) | 1 (1.3) |
| M (n = 101) | 13 (12.9) | 4 (4.0) | 9 (8.9) | 4 (4.0) | 2 (2.0) | 1 (1.0) |
| Residence | ||||||
| Surrounding villages (n = 53) | 10 (18.9) | 3 (5.7) | 7 (13.2) | 3 (5.7) | 4 (7.5) | 1 (1.9) |
| Lambaréné (n = 124) | 16 (12.9) | 3 (2.4) | 13 (10.5) | 10 (8.1) | 2 (1.6) | 1 (0.8) |
| Healthy children | ||||||
| NoV n (%) | NoV GI n (%) | NoV GII n (%) | AstV n (%) | SaV n (%) | AiV n (%) | |
| Age group (Months) | ||||||
| 0–6 (n = 7) | 1 (14.3) | 0 (0) | 1 (14.3) | 0 (0) | 0 (0) | 0 (0) |
| 7–12 (n = 8) | 0 (0) | 0 (0) | 0 (0) | 1 (12.5) | 0 (0) | 0 (0) |
| 13–18 (n = 4) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) |
| 19–24 (n = 16) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 0 (0) | 1 (6.3) |
| 25–59 (n = 32) | 5 (15.6) | 3 (9.4) | 2 (6.3) | 2 (6.3) | 1 (3.1) | 3 (9.4) |
| Gender | ||||||
| F (n = 26) | 1 (3.8) | 1 (3.8) | 0 (0) | 1 (3.8) | 0 (0) | 2 (7.7) |
| M (n = 41) | 5 (12.2) | 2 (4.9) | 3 (7.3) | 2 (4.9) | 1 (2.4) | 2 (4.9) |
| Residence | ||||||
| Surrounding villages (n = 39) | 3 (7.7) | 0 (0) | 3 (7.7) | 2 (5.1) | 0 (0) | 3 (7.7) |
| Lambaréné (n = 28) | 3 (10.7) | 3 (10.7) | 0 (0) | 1 (3.6) | 1 (3.6) | 1 (3.6) |
| Polymerase (RdRp) Genotype | Capsid (P2 Domain) Genotype | Number (%) |
|---|---|---|
| Genogroup I | ||
| GI.P3 | GI.3 | 1 (3.2) |
| GI.5 | 1 (3.2) | |
| GI.P4 | GI.4 | 1 (3.2) |
| GI.P5 | GI.5 | 1 (3.2) |
| GI.3 | 2 (6.5) | |
| GI. | 1 (3.2) | |
| GI.2 | 1 (3.2) | |
| GI.P11 | GI.2 | 1 (3.2) |
| Genogroup II | ||
| GII.P17 | GII.17 | 3 (9.7) |
| GII.P17 | GII. | 1 (3.2) |
| GII.P31 | GII.4 Syd | 14 (45.2) |
| GII.6 | 1 (3.2) | |
| GII.P31 | GII.4 NO | 3 (9.7) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Manouana, G.P.; Nguema-Moure, P.A.; Mbong Ngwese, M.; Bock, C.-T.; Kremsner, P.G.; Borrmann, S.; Eibach, D.; Mordmüller, B.; Velavan, T.P.; Niendorf, S.; et al. Genetic Diversity of Enteric Viruses in Children under Five Years Old in Gabon. Viruses 2021, 13, 545. https://doi.org/10.3390/v13040545
Manouana GP, Nguema-Moure PA, Mbong Ngwese M, Bock C-T, Kremsner PG, Borrmann S, Eibach D, Mordmüller B, Velavan TP, Niendorf S, et al. Genetic Diversity of Enteric Viruses in Children under Five Years Old in Gabon. Viruses. 2021; 13(4):545. https://doi.org/10.3390/v13040545
Chicago/Turabian StyleManouana, Gédéon Prince, Paul Alvyn Nguema-Moure, Mirabeau Mbong Ngwese, C.-Thomas Bock, Peter G. Kremsner, Steffen Borrmann, Daniel Eibach, Benjamin Mordmüller, Thirumalaisamy P. Velavan, Sandra Niendorf, and et al. 2021. "Genetic Diversity of Enteric Viruses in Children under Five Years Old in Gabon" Viruses 13, no. 4: 545. https://doi.org/10.3390/v13040545
APA StyleManouana, G. P., Nguema-Moure, P. A., Mbong Ngwese, M., Bock, C.-T., Kremsner, P. G., Borrmann, S., Eibach, D., Mordmüller, B., Velavan, T. P., Niendorf, S., & Adegnika, A. A. (2021). Genetic Diversity of Enteric Viruses in Children under Five Years Old in Gabon. Viruses, 13(4), 545. https://doi.org/10.3390/v13040545










