Atovaquone and Berberine Chloride Reduce SARS-CoV-2 Replication In Vitro
Abstract
:1. Introduction
2. Materials and Methods
2.1. Cells and Viruses
2.2. Plaque Assays
2.3. Compound Treatment
2.4. Cell Cytotoxicity Assays
2.5. Cell Proliferation Experiment
2.6. Human Airway Epithelial (HAE) Cultures and Infections
2.7. Human Colonic Organoid Cultures and Infections
2.8. Berberine Chloride (BBC) Cell and Direct Virus Treatment Assay
2.9. Data Analysis and Statistics
3. Results
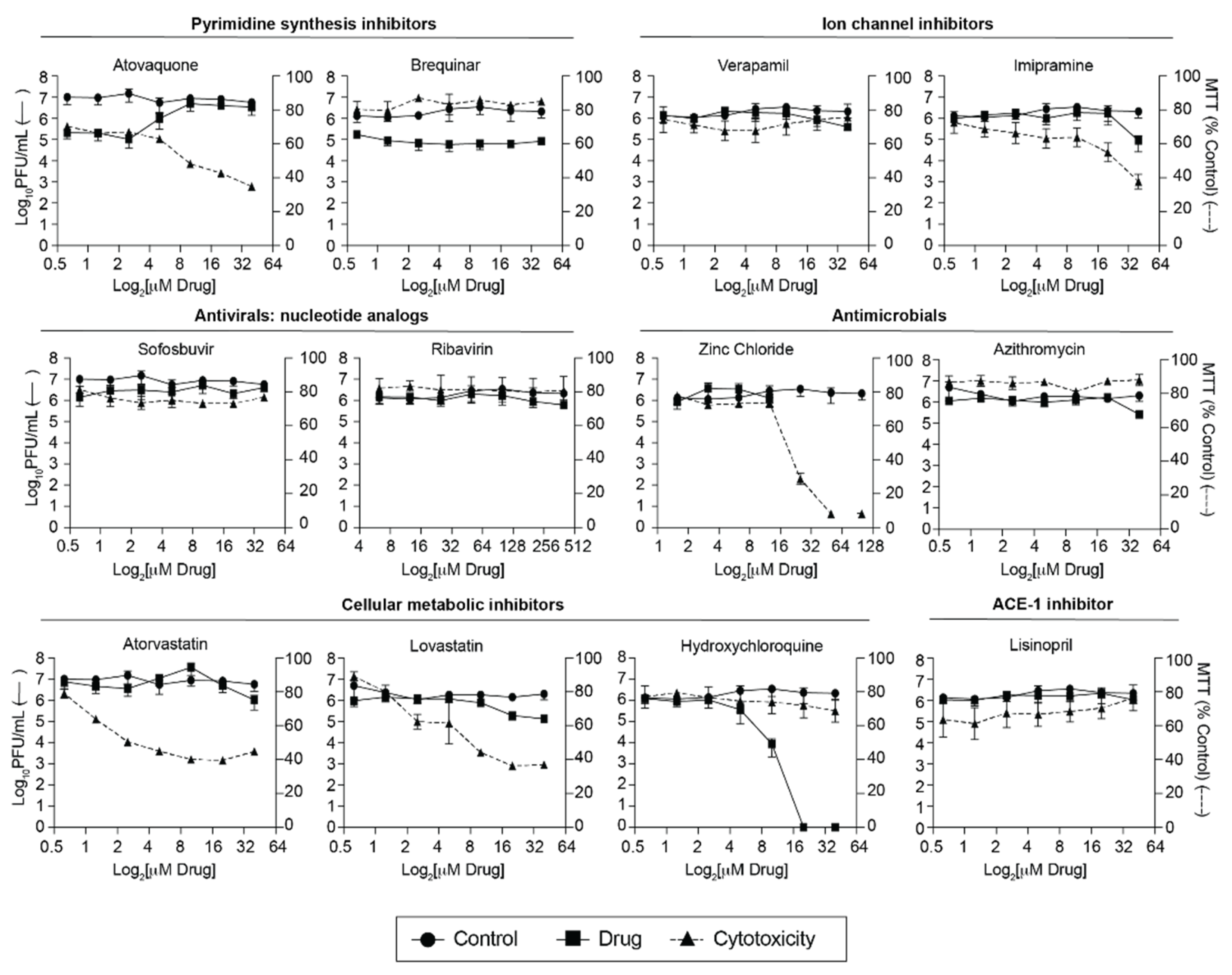
3.1. Screen of FDA-Approved Compounds against SARS-CoV-2
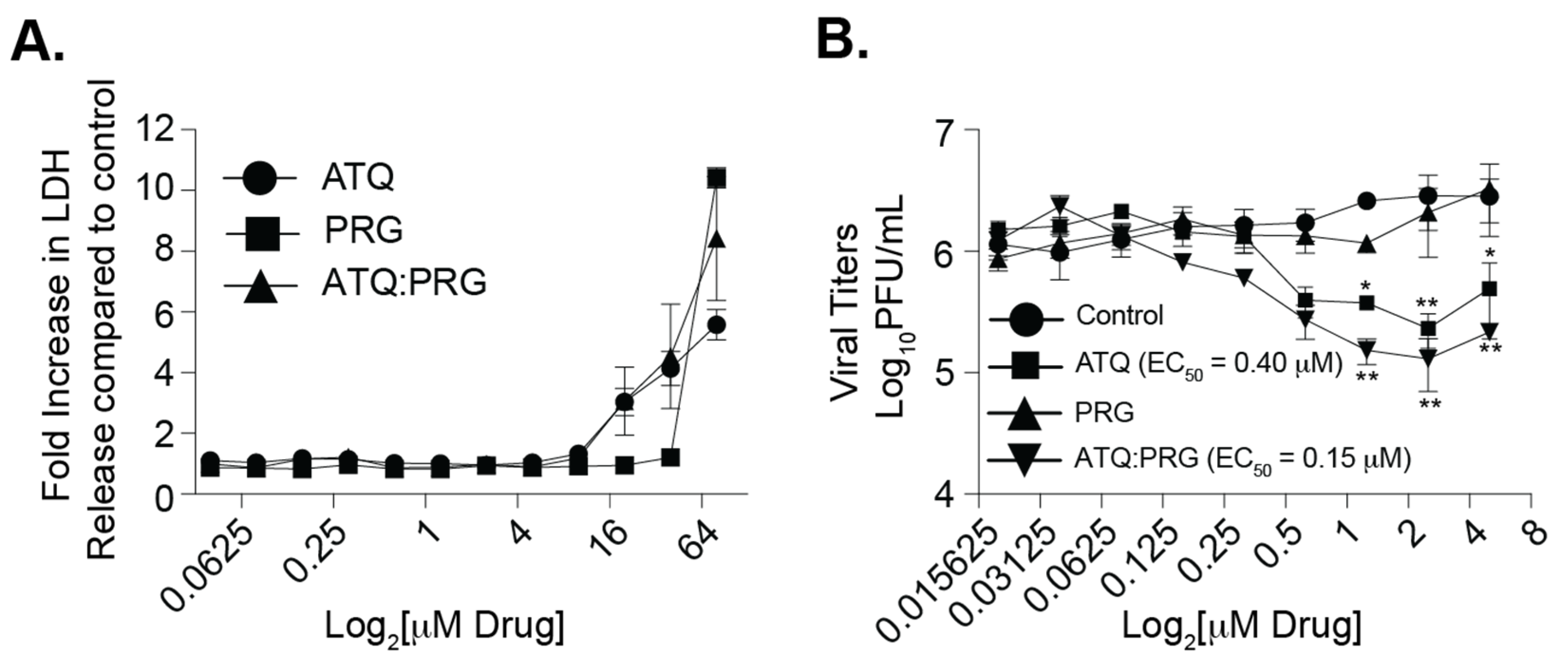
3.2. Atovaquone and Proguanil Hydrochloride Function Together to Reduce SARS-CoV-2 Infectious Particle Production
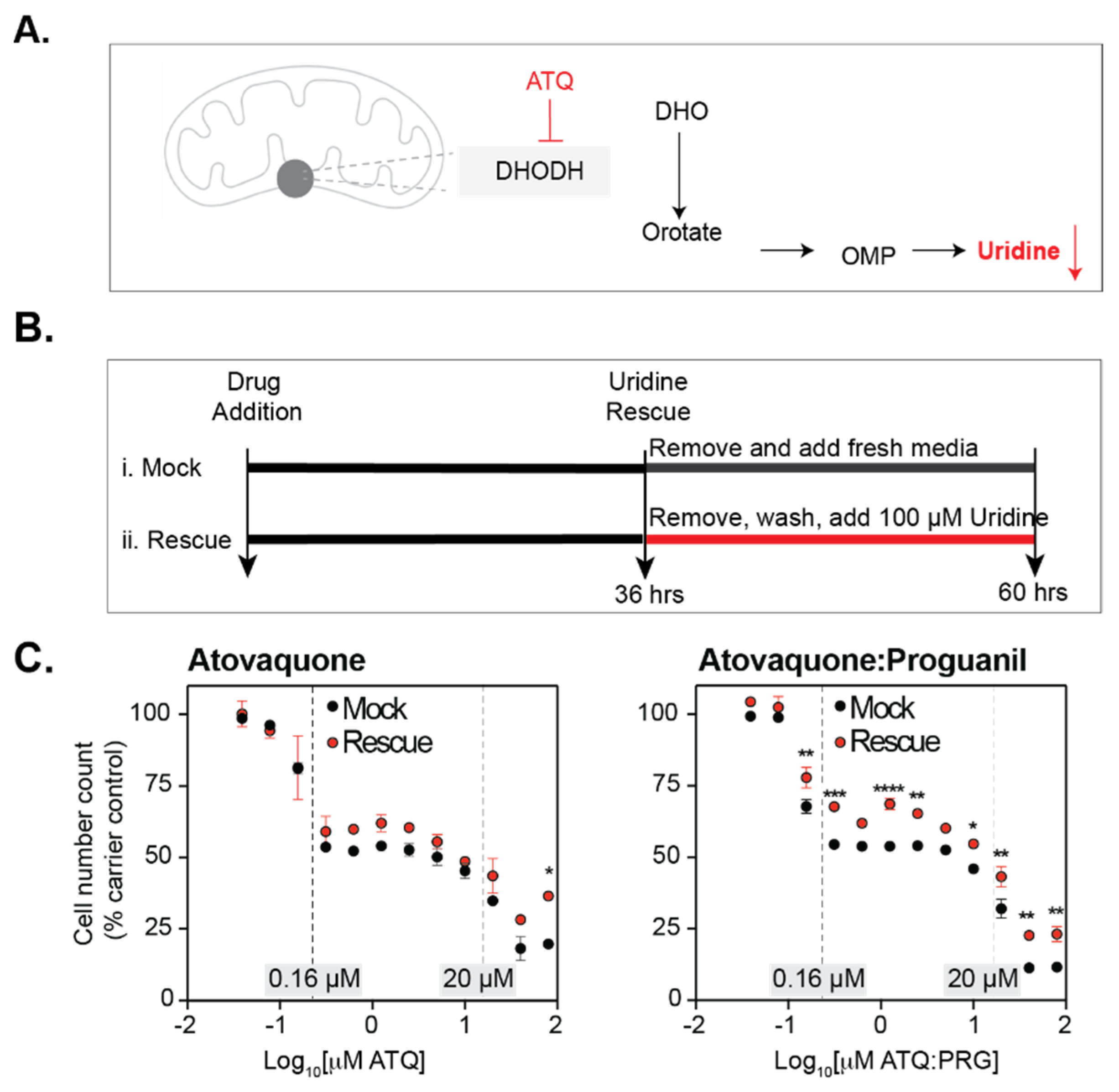
3.3. Atovaquone Affects Cell Proliferation
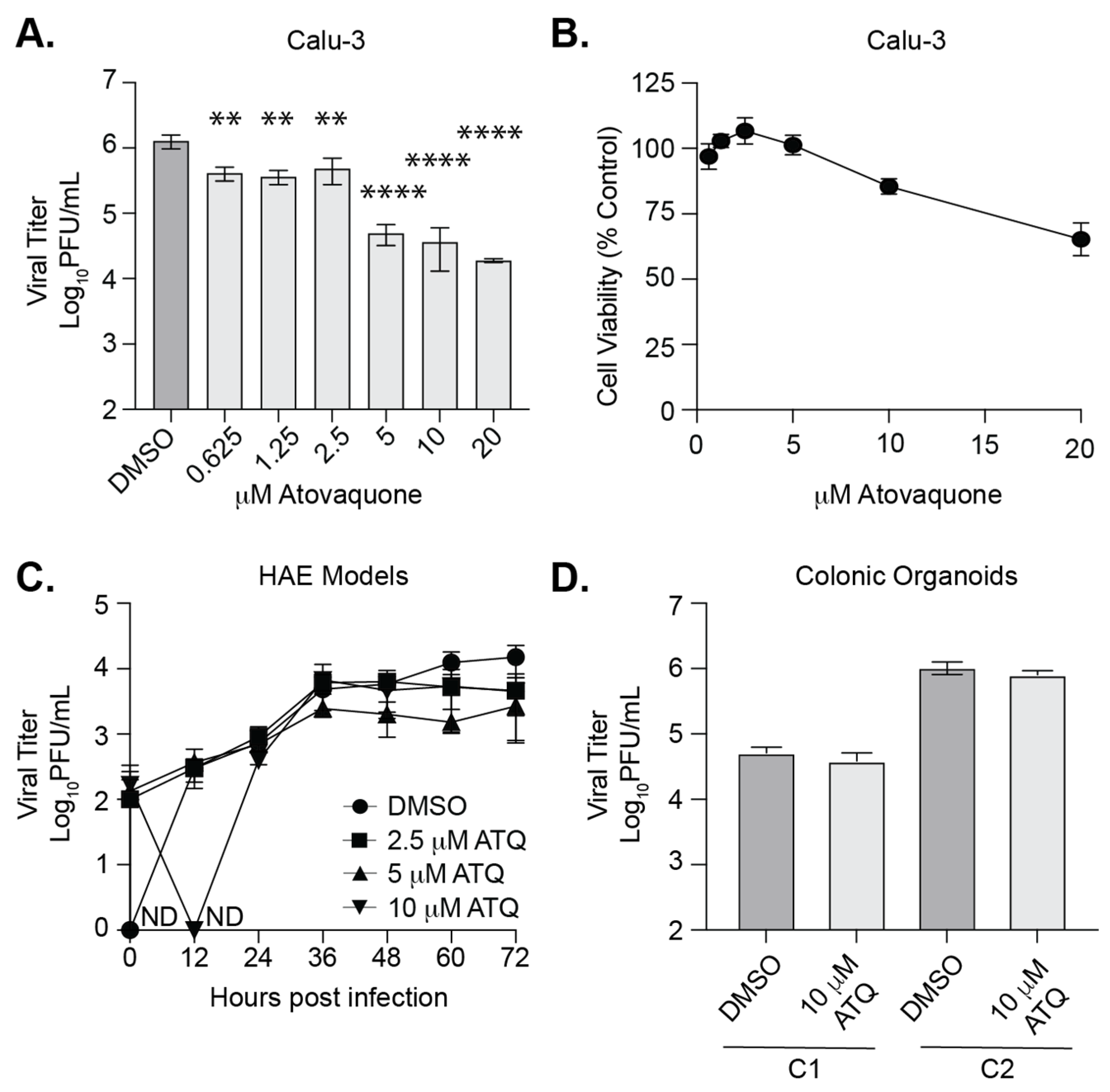
3.4. Atovaquone Reduces SARS-CoV-2 Infection in Calu-3 Cells But Not in Human Airway Epithelial Cultures or Colonic Organoids
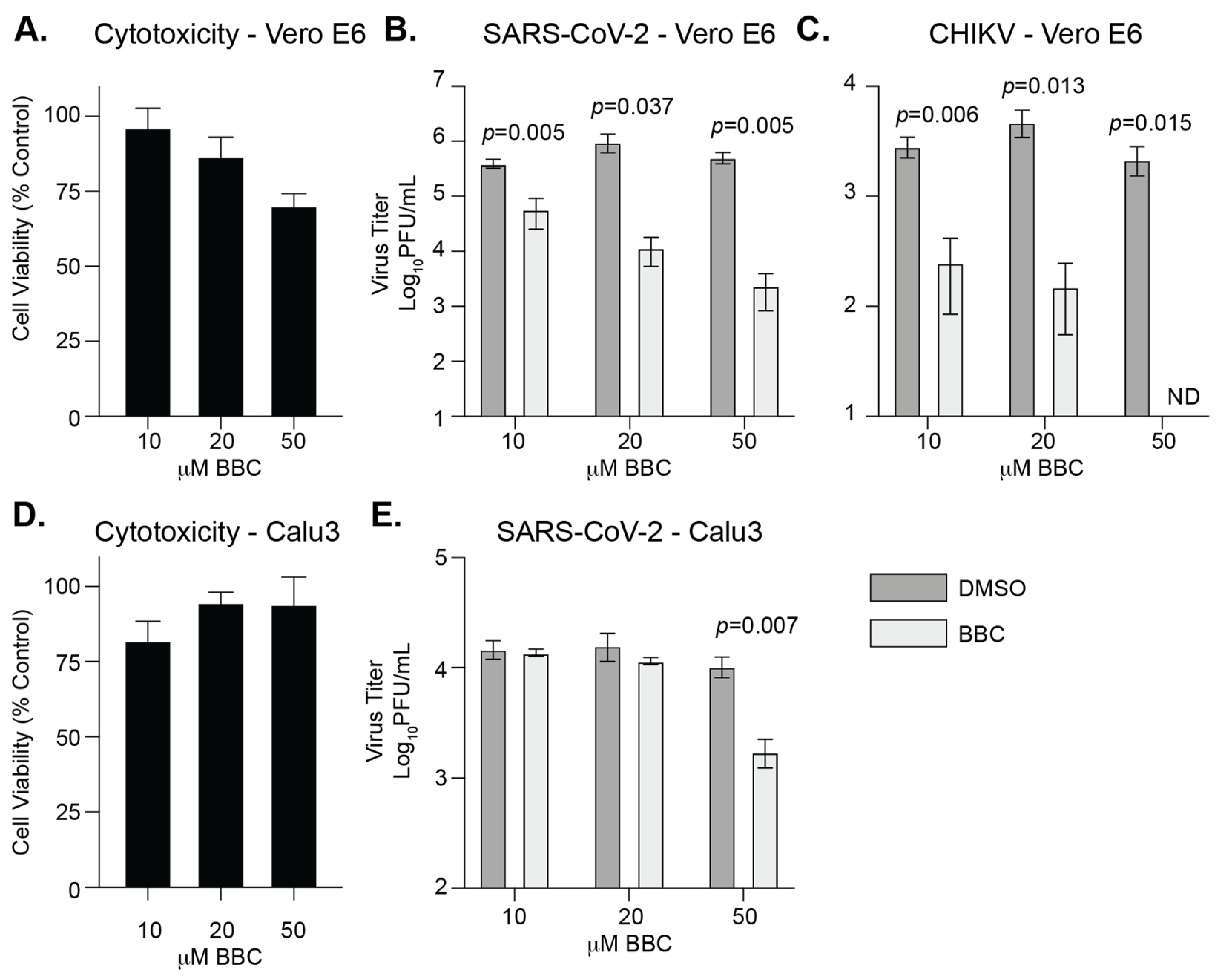
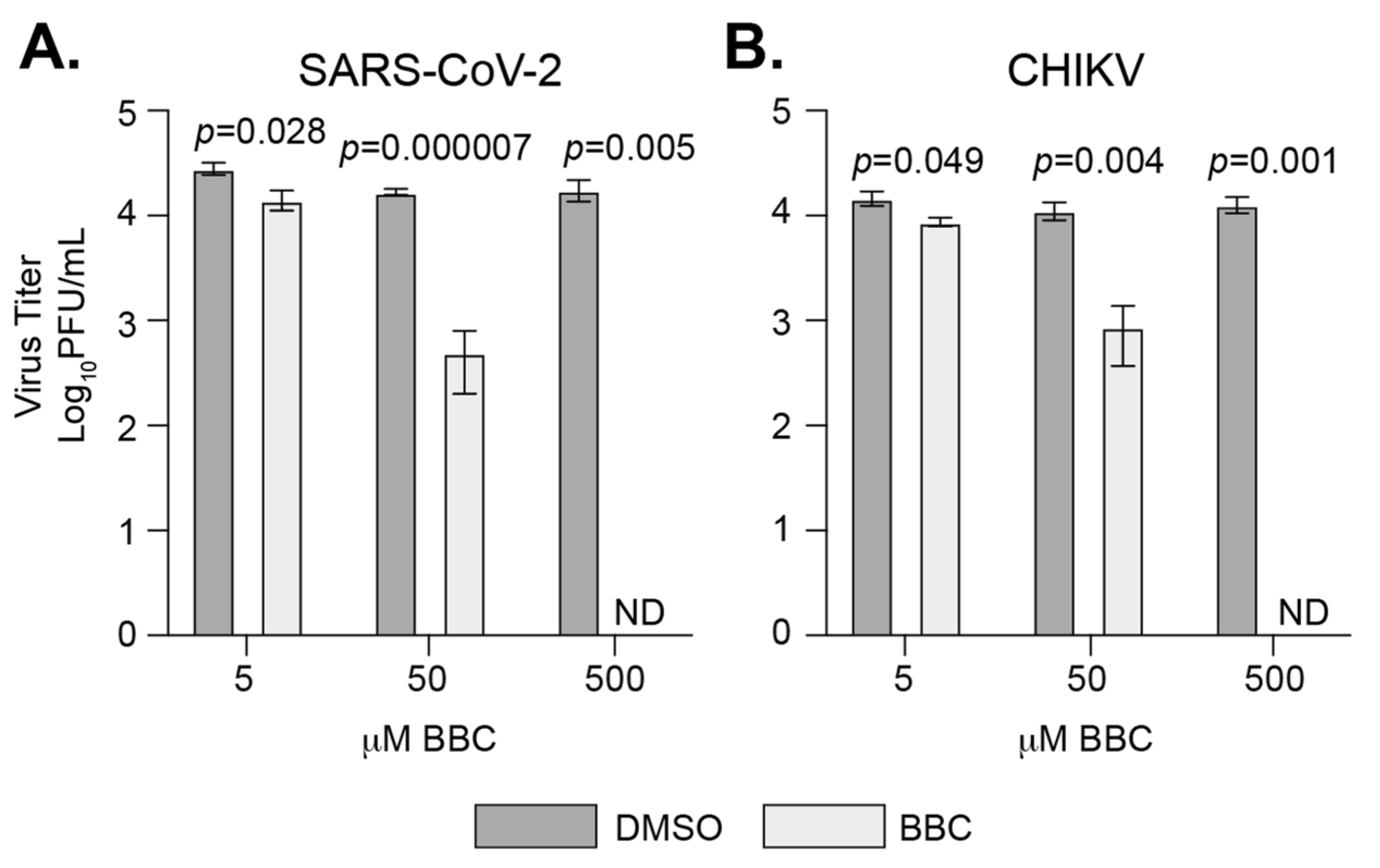
3.5. Berberine Chloride Inhibits SARS-CoV-2 Replication and Infectivity
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Dittmar, M.; Lee, J.S.; Whig, K.; Segrist, E.; Li, M.; Kamalia, B.; Castellana, L.; Ayyanathan, K.; Cardenas-Diaz, F.L.; Morrisey, E.E.; et al. Drug repurposing screens reveal cell-type-specific entry pathways and FDA-approved drugs active against SARS-Cov-2. Cell Rep. 2021, 35, 108959. [Google Scholar] [CrossRef] [PubMed]
- Bakowski, M.A.; Beutler, N.; Wolff, K.C.; Kirkpatrick, M.G.; Chen, E.; Nguyen, T.-T.H.; Riva, L.; Shaabani, N.; Parren, M.; Ricketts, J.; et al. Drug repurposing screens identify chemical entities for the development of COVID-19 interventions. Nat. Commun. 2021, 12, 3309. [Google Scholar] [CrossRef] [PubMed]
- Puhl, A.C.; Fritch, E.J.; Lane, T.R.; Tse, L.V.; Yount, B.L.; Sacramento, C.Q.; Fintelman-Rodrigues, N.; Tavella, T.A.; Costa, F.T.M.; Weston, S.; et al. Repurposing the ebola and marburg virus inhibitors tilorone, quinacrine, and pyronaridine: In vitro activity against SARS-CoV-2 and potential mechanisms. ACS Omega 2021, 6, 7454–7468. [Google Scholar] [CrossRef]
- Sheahan, T.P.; Sims, A.C.; Graham, R.L.; Menachery, V.D.; Gralinski, L.E.; Case, J.B.; Leist, S.R.; Pyrc, K.; Feng, J.Y.; Trantcheva, I.; et al. Broad-spectrum antiviral GS-5734 inhibits both epidemic and zoonotic coronaviruses. Sci. Transl. Med. 2017, 9, 9. [Google Scholar] [CrossRef] [Green Version]
- Gordon, D.E.; Jang, G.M.; Bouhaddou, M.; Xu, J.; Obernier, K.; White, K.M.; O’Meara, M.J.; Rezelj, V.V.; Guo, J.Z.; Swaney, D.L.; et al. A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature 2020, 583, 459–468. [Google Scholar] [CrossRef] [PubMed]
- Pruijssers, A.J.; George, A.S.; Schäfer, A.; Leist, S.R.; Gralinksi, L.E.; Dinnon, K.H., 3rd; Yount, B.L.; Agostini, M.L.; Stevens, L.J.; Chappell, J.D.; et al. Remdesivir Inhibits SARS-CoV-2 in human lung cells and chimeric SARS-CoV expressing the SARS-CoV-2 RNA polymerase in mice. Cell Rep. 2020, 32, 107940. [Google Scholar] [CrossRef]
- De Vries, M.; Mohamed, A.S.; Prescott, R.A.; Valero-Jimenez, A.M.; Desvignes, L.; O’Connor, R.; Steppan, C.; Devlin, J.C.; Ivanova, E.; Herrera, A.; et al. A comparative analysis of SARS-CoV-2 antivirals characterizes 3CLpro inhibitor PF-00835231 as a potential new treatment for COVID-19. J. Virol. 2021, 95, e01819-20. [Google Scholar] [CrossRef]
- Harcourt, J.; Tamin, A.; Lu, X.; Kamili, S.; Sakthivel, S.K.; Murray, J.; Queen, K.; Tao, Y.; Paden, C.R.; Zhang, J.; et al. Severe acute respiratory syndrome coronavirus 2 from patient with coronavirus disease, United States. Emerg. Infect. Dis. 2020, 26, 1266–1273. [Google Scholar] [CrossRef]
- Xie, X.; Muruato, A.; Lokugamage, K.G.; Narayanan, K.; Zhang, X.; Zou, J.; Liu, J.; Schindewolf, C.; Bopp, N.E.; Aguilar, P.V.; et al. An Infectious cDNA Clone of SARS-CoV-2. Cell Host Microbe 2020, 27, 841–848.e3. [Google Scholar] [CrossRef]
- Coffey, L.L.; Vignuzzi, M. Host alternation of chikungunya virus increases fitness while restricting population diversity and adaptability to novel selective pressures. J. Virol. 2011, 85, 1025–1035. [Google Scholar] [CrossRef] [Green Version]
- Españo, E.; Nam, J.-H.; Song, E.-J.; Song, D.; Lee, C.-K.; Kim, J.-K. Lipophilic statins inhibit Zika virus production in Vero cells. Sci. Rep. 2019, 9, 11461. [Google Scholar] [CrossRef]
- Wani, M.A.; Mukherjee, S.; Mallick, S.; Akbar, I.; Basu, A. Atorvastatin ameliorates viral burden and neural stem/progenitor cell (NSPC) death in an experimental model of Japanese encephalitis. J. Biosci. 2020, 45, 77. [Google Scholar] [CrossRef]
- Kottkamp, A.; De Jesus, E.; Grande, R.; Brown, J.A.; Jacobs, A.R.; Lim, J.K.; Stapleford, K.A. Atovaquone Inhibits Arbovirus Replication through the Depletion of Intracellular Nucleotides. J. Virol. 2019, 93, 93. [Google Scholar] [CrossRef] [Green Version]
- Retallack, H.; Di Lullo, E.; Arias, C.; Knopp, K.A.; Laurie, M.T.; Sandoval-Espinosa, C.; Mancia Leon, W.R.; Krencik, R.; Ullian, E.M.; Spatazza, J.; et al. Zika virus cell tropism in the developing human brain and inhibition by azithromycin. Proc. Natl. Acad. Sci. USA 2016, 113, 14408–14413. [Google Scholar] [CrossRef] [Green Version]
- Chen, J.; Huang, X.; Tao, C.; Wang, L.; Chen, Z.; Li, X.; Zeng, Q.; Ma, M.; Zhang, R.; Wu, Z. Berberine chloride suppresses non-small cell lung cancer by deregulating Sin3A/TOP2B pathway in vitro and in vivo. Cancer Chemother. Pharmacol. 2020, 86, 151–161. [Google Scholar] [CrossRef] [PubMed]
- Qing, M.; Zou, G.; Wang, Q.-Y.; Xu, H.Y.; Dong, H.; Yuan, Z.; Shi, P.-Y. Characterization of dengue virus resistance to brequinar in cell culture. Antimicrob. Agents Chemother. 2010, 54, 3686–3695. [Google Scholar] [CrossRef] [Green Version]
- Li, S.-F.; Gong, M.-J.; Sun, Y.-F.; Shao, J.-J.; Zhang, Y.-G.; Chang, H.-Y. Antiviral activity of brequinar against foot-and-mouth disease virus infection in vitro and in vivo. Biomed. Pharmacother. 2019, 116, 108982. [Google Scholar] [CrossRef]
- Cao, B.; Parnell, L.A.; Diamond, M.S.; Mysorekar, I.U. Inhibition of autophagy limits vertical transmission of Zika virus in pregnant mice. J. Exp. Med. 2017, 214, 2303–2313. [Google Scholar] [CrossRef] [PubMed]
- Wichit, S.; Hamel, R.; Bernard, E.; Talignani, L.; Diop, F.; Ferraris, P.; Liegeois, F.; Ekchariyawat, P.; Luplertlop, N.; Surasombatpattana, P.; et al. Imipramine inhibits chikungunya virus replication in human skin fibroblasts through interference with intracellular cholesterol trafficking. Sci. Rep. 2017, 7, 3145. [Google Scholar] [CrossRef] [Green Version]
- Blanshard, A.; Hine, P. Atovaquone-proguanil for treating uncomplicated Plasmodium falciparum malaria. Cochrane Database Syst. Rev. 2021, 2021, CD004529. [Google Scholar] [CrossRef]
- Beaucourt, S.; Vignuzzi, M. Ribavirin: A drug active against many viruses with multiple effects on virus replication and propagation. Molecular basis of ribavirin resistance. Curr. Opin. Virol. 2014, 8, 10–15. [Google Scholar] [CrossRef]
- Ferreira, A.C.; Zaverucha-Do-Valle, C.; Reis, P.A.; Barbosa-Lima, G.; Vieira, Y.R.; Mattos, M.; Silva, P.D.P.; Sacramento, C.; de Castro Faria Neto, H.C.; Campanati, L.; et al. Sofosbuvir protects Zika virus-infected mice from mortality, preventing short- and long-term sequelae. Sci. Rep. 2017, 7, 9409. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- De Freitas, C.S.; Higa, L.M.; Sacramento, C.Q.; Ferreira, A.C.; Reis, P.A.; Delvecchio, R.; Monteiro, F.L.; Barbosa-Lima, G.; Westgarth, H.J.; Vieira, Y.R.; et al. Yellow fever virus is susceptible to sofosbuvir both in vitro and in vivo. PLoS Negl. Trop. Dis. 2019, 13, e0007072. [Google Scholar] [CrossRef] [Green Version]
- Nugent, K.M.; Shanley, J.D. Verapamil inhibits influenza A virus replication. Arch. Virol. 1984, 81, 163–170. [Google Scholar] [CrossRef] [PubMed]
- Choi, E.-K.; Lee, H.-H.; Kang, M.-S.; Kim, B.-G.; Lim, H.-S.; Kim, S.-M.; Kang, I.-C. Potentiation of bacterial killing activity of zinc chloride by pyrrolidine dithiocarbamate. J. Microbiol. 2010, 48, 40–43. [Google Scholar] [CrossRef]
- Neil, J.A.; Matsuzawa-Ishimoto, Y.; Kernbauer-Hölzl, E.; Schuster, S.L.; Sota, S.; Venzon, M.; Dallari, S.; Neto, A.G.; Hine, A.; Hudesman, D.; et al. IFN-I and IL-22 mediate protective effects of intestinal viral infection. Nat. Microbiol. 2019, 4, 1737–1749. [Google Scholar] [CrossRef] [PubMed]
- Matsuzawa-Ishimoto, Y.; Hine, A.; Shono, Y.; Rudensky, E.; Lazrak, A.; Yeung, F.; Neil, J.A.; Yao, X.; Chen, Y.-H.; Heaney, T.; et al. An intestinal organoid–based platform that recreates susceptibility to T-cell–mediated tissue injury. Blood 2020, 135, 2388–2401. [Google Scholar] [CrossRef]
- Wan, J.J.; Brown, R.S.; Kielian, M. Berberine chloride is an alphavirus inhibitor that targets nucleocapsid assembly. mBio 2020, 11, e01382-20. [Google Scholar] [CrossRef]
- Saib, A.; Amara, W.; Wang, P.; Cattan, S.; Dellal, A.; Regaieg, K.; Nahon, S.; Nallet, O.; Nguyen, L.S. Lack of efficacy of hydroxychloroquine and azithromycin in patients hospitalized for COVID-19 pneumonia: A retrospective study. PLoS ONE 2021, 16, e0252388. [Google Scholar] [CrossRef]
- Ulrich, R.J.; Troxel, A.B.; Carmody, E.; Eapen, J.; Bäcker, M.; Dehovitz, J.A.; Prasad, P.J.; Li, Y.; Delgado, C.; Jrada, M.; et al. Treating Covid-19 With Hydroxychloroquine (TEACH): A multicenter, double-blind, randomized controlled trial in hospitalized patients. Open Forum Infect. Dis. 2020, 7, ofaa446. [Google Scholar] [CrossRef]
- Kovacs, J. Efficacy of atovaquone in treatment of toxoplasmosis in patients with AIDS. The NIAID-clinical center intramural AIDS program. Lancet 1992, 340, 637–638. [Google Scholar] [CrossRef]
- Krause, P.J.; Lepore, T.; Sikand, V.K.; Gadbaw, J.; Burke, G.; Telford, S.R., 3rd; Brassard, P.; Pearl, D.; Azlanzadeh, J.; Christianson, D.; et al. Atovaquone and azithromycin for the treatment of babesiosis. N. Engl. J. Med. 2000, 343, 1454–1458. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Srivastava, I.K.; Vaidya, A.B. A Mechanism for the synergistic antimalarial action of atovaquone and proguanil. Antimicrob. Agents Chemother. 1999, 43, 1334–1339. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nixon, G.L.; Pidathala, C.; Shone, A.E.; Antoine, T.; Fisher, N.; O’Neill, P.M.; Ward, S.A.; Biagini, G.A. Targeting the mitochondrial electron transport chain of Plasmodium falciparum: New strategies towards the development of improved antimalarials for the elimination era. Future Med. Chem. 2013, 5, 1573–1591. [Google Scholar] [CrossRef] [PubMed]
- Knecht, W.; Henseling, J.; Löffler, M. Kinetics of inhibition of human and rat dihydroorotate dehydrogenase by atovaquone, lawsone derivatives, brequinar sodium and polyporic acid. Chem. Interact. 2000, 124, 61–76. [Google Scholar] [CrossRef]
- Ashton, T.M.; Fokas, E.; Kunz-Schughart, L.A.; Folkes, L.K.; Anbalagan, S.; Huether, M.; Kelly, C.J.; Pirovano, G.; Buffa, F.M.; Hammond, E.M.; et al. The anti-malarial atovaquone increases radiosensitivity by alleviating tumour hypoxia. Nat. Commun. 2016, 7, 12308. [Google Scholar] [CrossRef] [Green Version]
- Messina, E.; Gazzaniga, P.; Micheli, V.; Guaglianone, M.R.; Barbato, S.; Morrone, S.; Frati, L.; Aglianò, A.M.; Giacomello, A. Guanine nucleotide depletion triggers cell cycle arrest and apoptosis in human neuroblastoma cell lines. Int. J. Cancer 2004, 108, 812–817. [Google Scholar] [CrossRef]
- Cohen, M.B.; Sadee, W. Contributions of the depletions of guanine and adenine nucleotides to the toxicity of purine starvation in the mouse T lymphoma cell line. Cancer Res. 1983, 43, 1587–1591. [Google Scholar]
- Zang, R.; Gomez Castro, M.F.; McCune, B.T.; Zeng, Q.; Rothlauf, P.W.; Sonnek, N.M.; Liu, Z.; Brulois, K.F.; Wang, X.; Greenberg, H.B.; et al. TMPRSS2 and TMPRSS4 promote SARS-CoV-2 infection of human small intestinal enterocytes. Sci. Immunol. 2020, 5, eabc3582. [Google Scholar] [CrossRef]
- Lamers, M.M.; Beumer, J.; van der Vaart, J.; Knoops, K.; Puschhof, J.; Breugem, T.I.; Ravelli, R.B.G.; van Schayck, J.P.; Mykytyn, A.Z.; Duimel, H.Q.; et al. SARS-CoV-2 productively infects human gut enterocytes. Science 2020, 369, 50–54. [Google Scholar] [CrossRef]
- Dickson, I. Organoids demonstrate gut infection by SARS-CoV-2. Nat. Rev. Gastroenterol. Hepatol. 2020, 17, 383. [Google Scholar] [CrossRef]
- Luganini, A.; Mercorelli, B.; Messa, L.; Palù, G.; Gribaudo, G.; Loregian, A. The isoquinoline alkaloid berberine inhibits human cytomegalovirus replication by interfering with the viral Immediate Early-2 (IE2) protein transactivating activity. Antivir. Res. 2019, 164, 52–60. [Google Scholar] [CrossRef]
- Hayashi, K.; Minoda, K.; Nagaoka, Y.; Hayashi, T.; Uesato, S. Antiviral activity of berberine and related compounds against human cytomegalovirus. Bioorg. Med. Chem. Lett. 2007, 17, 1562–1564. [Google Scholar] [CrossRef] [PubMed]
- Wojtyczka, R.D.; Dziedzic, A.; Kępa, M.; Kubina, R.; Kabała-Dzik, A.; Mularz, T.; Idzik, D. berberine enhances the antibacterial activity of selected antibiotics against coagulase-negative staphylococcus strains in vitro. Molecules 2014, 19, 6583–6596. [Google Scholar] [CrossRef]
- Dziedzic, A.; Wojtyczka, R.D.; Kubina, R. Inhibition of oral streptococci growth induced by the complementary action of berberine chloride and antibacterial compounds. Molecules 2015, 20, 13705–13724. [Google Scholar] [CrossRef] [PubMed]
- Varghese, F.S.; Thaa, B.; Amrun, S.N.; Simarmata, D.; Rausalu, K.; Nyman, T.; Merits, A.; McInerney, G.M.; Ng, L.F.P.; Ahola, T. The antiviral alkaloid berberine reduces chikungunya virus-induced mitogen-activated protein kinase signaling. J. Virol. 2016, 90, 9743–9757. [Google Scholar] [CrossRef] [Green Version]
- Pickard, A.; Calverley, B.C.; Chang, J.; Garva, R.; Gago, S.; Lu, Y.; Kadler, K.E. Discovery of re-purposed drugs that slow SARS-CoV-2 replication in human cells. PLoS Pathog. 2021, 17, e1009840. [Google Scholar] [CrossRef]
- Carter-Timofte, M.E.; Arulanandam, R.; Kurmasheva, N.; Fu, K.; Laroche, G.; Taha, Z.; van der Horst, D.; Cassin, L.; van der Sluis, R.M.; Palermo, E.; et al. Antiviral potential of the antimicrobial drug atovaquone against SARS-CoV-2 and emerging variants of concern. ACS Infect. Dis. 2021, 7, 3034–3051. [Google Scholar] [CrossRef]
- Calistri, A.; Luganini, A.; Mognetti, B.; Elder, E.; Sibille, G.; Conciatori, V.; Del Vecchio, C.; Sainas, S.; Boschi, D.; Montserrat, N.; et al. The New Generation hDHODH Inhibitor MEDS433 Hinders the in vitro replication of SARS-CoV-2 and other human coronaviruses. Microorganisms 2021, 9, 1731. [Google Scholar] [CrossRef]
- Ko, M.; Chang, S.; Byun, S.; Ianevski, A.; Choi, I.; D’Orengiani, A.-L.P.H.D.; Ravlo, E.; Wang, W.; Bjørås, M.; Kainov, D.; et al. Screening of FDA-Approved Drugs Using a MERS-CoV Clinical isolate from south korea identifies potential therapeutic options for COVID-19. Viruses 2021, 13, 651. [Google Scholar] [CrossRef]
- Luban, J.; Sattler, R.A.; Mühlberger, E.; Graci, J.D.; Cao, L.; Weetall, M.; Trotta, C.; Colacino, J.M.; Bavari, S.; Strambio-De-Castillia, C.; et al. The DHODH inhibitor PTC299 arrests SARS-CoV-2 replication and suppresses induction of inflammatory cytokines. Virus Res. 2021, 292, 198246. [Google Scholar] [CrossRef]
- Bonavia, A.; Franti, M.; Keaney, E.P.; Kuhen, K.; Seepersaud, M.; Radetich, B.; Shao, J.; Honda, A.; Dewhurst, J.; Balabanis, K.; et al. Identification of broad-spectrum antiviral compounds and assessment of the druggability of their target for efficacy against respiratory syncytial virus (RSV). Proc. Natl. Acad. Sci. USA 2011, 108, 6739–6744. [Google Scholar] [CrossRef] [Green Version]
- Yang, C.-W.; Peng, T.-T.; Hsu, H.-Y.; Lee, Y.-Z.; Wu, S.-H.; Lin, W.-H.; Ke, Y.-Y.; Hsu, T.-A.; Yeh, T.-K.; Huang, W.-Z.; et al. Repurposing old drugs as antiviral agents for coronaviruses. Biomed. J. 2020, 43, 368–374. [Google Scholar] [CrossRef] [PubMed]
- Coelho, A.R.; Oliveira, P.J. Dihydroorotate dehydrogenase inhibitors in SARS-CoV-2 infection. Eur. J. Clin. Investig. 2020, 50, e13366. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Li, K.; Maskey, A.R.; Huang, W.; Toutov, A.A.; Yang, N.; Srivastava, K.; Geliebter, J.; Tiwari, R.; Miao, M.; et al. A small molecule compound berberine as an orally active therapeutic candidate against COVID-19 and SARS: A computational and mechanistic study. FASEB J. 2021, 35, e21360. [Google Scholar] [CrossRef] [PubMed]
- Varghese, F.; van Woudenbergh, E.; Overheul, G.; Eleveld, M.; Kurver, L.; van Heerbeek, N.; van Laarhoven, A.; Miesen, P.; Hartog, G.D.; de Jonge, M.; et al. Berberine and Obatoclax Inhibit SARS-Cov-2 Replication in Primary Human Nasal Epithelial Cells In Vitro. Viruses 2021, 13, 282. [Google Scholar] [CrossRef]
| Compound | Source—Catalog Number | Concentrations Tested | Reference |
|---|---|---|---|
| Atorvastatin | TCI America—A24761G | 0.625 μM–40 μM | [11,12] |
| Atovaquone | Tocris Bioscience—6358 | 0.039 μM–80 μM | [13] |
| Azithromycin | TCI America—A20761G | 0.625 μM–40 μM | [14] |
| Berberine chloride | Sigma-Aldrich—B3251 | 10 μM–500 μM | [15] |
| Brequinar | Sigma-Aldrich—SML0113 | 0.039 μM–80 μM | [16,17] |
| Hydroxychloroquine | Santa Cruz—SC-215157 | 0.625 μM–40 μM | [18] |
| Imipramine | TCI America—I09711G | 0.625 μM–40 μM | [19] |
| Lovastatin | TCI America—L02145G | 0.625 μM–40 μM | [11] |
| Proguanil hydrochloride | ACROS—461430250 | 0.039 μM–80 μM | [20] |
| Ribavirin | Sigma-Aldrich—R9644 | 6.25 μM–400 μM | [21] |
| Sofosbuvir | Santa Cruz SC—482362 | 0.625 μM–40 μM | [22,23] |
| Verapamil | ACROS Organics—329330010 | 0.625 μM–40 μM | [24] |
| Zinc chloride | Fisher—Z33-100 | 1.56 μM–100 μM | [25] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rodriguez-Rodriguez, B.A.; Noval, M.G.; Kaczmarek, M.E.; Jang, K.K.; Thannickal, S.A.; Cifuentes Kottkamp, A.; Brown, R.S.; Kielian, M.; Cadwell, K.; Stapleford, K.A. Atovaquone and Berberine Chloride Reduce SARS-CoV-2 Replication In Vitro. Viruses 2021, 13, 2437. https://doi.org/10.3390/v13122437
Rodriguez-Rodriguez BA, Noval MG, Kaczmarek ME, Jang KK, Thannickal SA, Cifuentes Kottkamp A, Brown RS, Kielian M, Cadwell K, Stapleford KA. Atovaquone and Berberine Chloride Reduce SARS-CoV-2 Replication In Vitro. Viruses. 2021; 13(12):2437. https://doi.org/10.3390/v13122437
Chicago/Turabian StyleRodriguez-Rodriguez, Bruno A., Maria G. Noval, Maria E. Kaczmarek, Kyung Ku Jang, Sara A. Thannickal, Angelica Cifuentes Kottkamp, Rebecca S. Brown, Margaret Kielian, Ken Cadwell, and Kenneth A. Stapleford. 2021. "Atovaquone and Berberine Chloride Reduce SARS-CoV-2 Replication In Vitro" Viruses 13, no. 12: 2437. https://doi.org/10.3390/v13122437
APA StyleRodriguez-Rodriguez, B. A., Noval, M. G., Kaczmarek, M. E., Jang, K. K., Thannickal, S. A., Cifuentes Kottkamp, A., Brown, R. S., Kielian, M., Cadwell, K., & Stapleford, K. A. (2021). Atovaquone and Berberine Chloride Reduce SARS-CoV-2 Replication In Vitro. Viruses, 13(12), 2437. https://doi.org/10.3390/v13122437






