Abstract
Extracellular vesicles (EVs) are cell-derived membranous particles secreted by all cell types (including virus infected and uninfected cells) into the extracellular milieu. EVs carry, protect, and transport a wide array of bioactive cargoes to recipient/target cells. EVs regulate physiological and pathophysiological processes in recipient cells and are important in therapeutics/drug delivery. Despite these great attributes of EVs, an efficient protocol for EV separation from biofluids is lacking. Numerous techniques have been adapted for the separation of EVs with size exclusion chromatography (SEC)-based methods being the most promising. Here, we review the SEC protocols used for EV separation, and discuss opportunities for significant improvements, such as the development of novel particle purification liquid chromatography (PPLC) system capable of tandem purification and characterization of biological and synthetic particles with near-single vesicle resolution. Finally, we identify future perspectives and current issues to make PPLC a tool capable of providing a unified, automated, adaptable, yet simple and affordable particle separation resource.
1. Introduction
EVs are intercellular communication vehicles that are revolutionizing the biomedicine field. However, a few people know that the discovery of EVs can be traced back to the 19th century, although under different nomenclature. In the 1840s, Gulliver’s editorial report (based on the work of Hewson in the 1770s) on milky particles in the blood serum ––that he called “the molecular base of the chyle” and characterized as very small globules of active Brownian movement and of uniform size ranging from ~0.5 to 1 micron––is perhaps the first encounter of EVs in the literature (see [1], pages 7 and 13). In the 1870s, Edmunds [2] described the normal blood serum as filled with a “nebulous haze of points, as is mote-laden air in a sunbeam” when visualized under a microscope. This finding was reproduced by Muller, who coined the particles “Haemokonien" (or blood dust) [3]. However, it was not until the 1930s when Frazer and Stewart reported on different subpopulations of particles in the blood serum, which they described as "large bright particles, small bright particles, and dull particles [4]. The authors also reported particle abundance during fasting and after meals, and estimated particle size to be ~1, ~0.33–0.5, and 0.035 micron, respectively for the above-mentioned particles [5]. In 1946, Chargaff and West [6] isolated the coagulation components of the platelet-free plasma by high-speed centrifugation (31,000× g), but it was not clear whether these structures were the same as Haemokonien. Other names describing blood serum particles such as protein particles, fat particles, fat dust, disintegrated platelets, chylomicrons, or white blood cells were also introduced [7]. In 1967, Wolf introduced the term “platelet-dust” to distinguish the platelet-secreted coagulant-material-carrying particles from chylomicrons. The “platelet-dust” were pelleted by high-speed centrifugation (134,211× g, 3 h) and characterized by electron microscopy and immunological techniques [8]. In the early 1980s, EVs were identified as structures of either membranous or endosomal origins exhibiting enzymatic activities. In the last two decades, EVs started gaining attention after the discovery of their pivotal roles in various physiological and pathological processes, but also in diagnostics and therapeutics (see [9,10,11] for excellent reviews on the subject).
Today, it is established that almost every cell type from the three kingdoms of life––Eukarya, Bacteria, and Archaea––produces EVs. Thus, EV research has become a subject of interest to scientists across numerous disciplines from basic science in chemistry, biology, and virology to applied science in diagnostics, pharmaceutics, and medicine. Such global interest in the EV field prompted a series of organizational actions in the past decade. The international Society for Extracellular Vesicles (ISEV) was created (2012 in Sweden) [12], and the Journal of Extracellular Vesicles (JEV, dedicated for EV research) was launched [13]. National EV societies across the world (United States—ASEMV, Spain—GEIVEX, Austria—ASEV, Germany—GSEV, Netherlands—NLSEV, Japan—JSEV, United Kingdom—UKEV, Taiwan—TSEV, Singapore—SOCRATES, Italy—EVITA, France—FSEV, Belgium—BESEV, and Korea—KSEV) were established, along with some regional networks (Regional Research Network on Extracellular Vesicles and the Gordon Research Conference Extracellular Vesicles biannual meeting). Furthermore, the number of scientific publications have exponentially increased, concomitant with an increase in industrial and governmental investment in EV research, such as the National Institute of Health (NIH) initiation of the Extracellular RNA consortium [14]. These measures were unsurprisingly accompanied with a massive increase in the global EV-based diagnostics and therapeutics market.
However, the progress and innovation in EV research has been largely impeded by challenges with EV separation and characterization [15]. Examples of such challenges include (1) insufficiently pure EV preparations; (2) interference of isolation reagents with downstream analyses; (3) lack of EV subpopulation separation; and (4) lack of scale-up capabilities. These challenges are exacerbated by the lack of automated equipment that readily isolate EVs from a given biofluid, the outcome of which is poor reproducibility of the EV preparations. Numerous NIH request for applications (RFA) have been activated to close this gap, but to date, no standardized EV separation method has been established. In addition to the lack of a standardized EV separation method, debate on whether single-vesicle resolution is needed for advancing the EV field is ongoing. Pro single-vesicle researchers argue that, because of the heterogeneity of EVs, identification of the specific bioactive EV cargo requires a single-vesicle approach. In contrast, researchers from another school of thought argue from a physiological standpoint that embracing rather than ignoring the inherent EV heterogeneity is key to understanding the biological and functional significance of EVs, since that is how EVs naturally occur in vivo. However, whether researchers are pro-single EV or not, it should be noted that before EVs can be efficiently used, the population(s) of interest must be isolated in pure and reproducible ways and retrieved in preparative quantities for downstream use. While searching for a consensus on standardized EV separation [16], the ISEV has published guidelines that urge EV scientists to thoroughly control for their EV content, use a combination of separation and characterization techniques, and report as much experimental details as possible [17,18,19,20,21].
As of 2016, ultracentrifugation was the most used technique for EV isolation, whether alone or in combination with other methods [22], despite its disadvantages [23]. Nevertheless, efforts concomitant with rapid technological advancement recently generated new exciting protocols for EV subpopulation separation (reviewed elsewhere [24]); e.g., asymmetric flow field-flow fractionation (AF4) [25], microfluidic isolation [26], high resolution flow cytometry [27,28,29,30,31], and immunocapture [32], to cite but a few. On the other hand, SEC has also been gaining popularity in the EV field, because of its simplicity, reproducibility, and gentle process allowing for purer and more active EVs, as compared to ultracentrifugation [33]. While EV separation techniques have been extensively discussed in various reviews [24,34] and protocols [35,36], this current review will focus on SEC and its variations as methods of EV separation. We will also discuss potential SEC add-ons that may help achieve EV subpopulation separation with near single-vesicle resolution.
2. History of Size Exclusion Chromatography (SEC) and Commercially Available SEC Beads
The first successful chemical separation by liquid-liquid partition chromatography can be traced back to the 1940s where Martin and Synge used a 1 cm silica gel column to separate amino acids [37,38,39]. For this innovation, the group was awarded the 1952 Nobel prize in chemistry [40]. However, the first size exclusion separation was described by Lindqvist and Storgårds who used a 2 cm × 120 cm starch column to separate cheese extracts into two peaks of peptides and amino-acids, collected in 275 fractions of 1 mL [41]. Soon after, glucan, a bacterial-derived carbohydrate polymer of glucose, was crosslinked with epichlorohydrin to yield crosslinked dextran, a more inert material with extended mechanical properties compared to starch and comparable fractionation capacity [42]. The capability of a dextran column was illustrated by separating 3 mL of fresh serum on a 17.5 cm × 2.5 cm into a large proteinaceous peak and small non-proteinaceous peak. By spiking the serum with a solution of saturated ammonium sulfate, the authors demonstrated the latter to be salts [42]. These dextran-based beads were commercialized at the time by (Pharmacia, Sweden) as Sephadex, and are still the most popular to date (Table 1). Ever since, minimal optimization was noted in which beads were rendered porous and the bead and pore sizes more controlled. Shortly after, Pharmacia commercialized Sepharose (agarose) and Sephacryl (polyacrylamide) beads, with a focus on improving mechanical strength to allow for high flow rates separation in fast purification liquid chromatography (FPLC) applications [43,44,45,46]. Variations of Sepharose were also developed for affinity purification [47,48], permitting fractionation of diverse bioactive molecules such as mammalian tRNA [49], human high density lipoproteins [50], and multiple types of growth factors in bone matrix [51]. Concurrently, Bio-Rad (CA, United States) commercialized Bio-Gel A (agarose) and P (polyacrylamide), originally prepared at the University of Uppsala (Sweden) by Hjertén and Mosbach [52,53] in 1962 for large and small molecules separation, respectively. Similar to Sephadex and Sepharose, the Bio-Gel products were also used in many discoveries [54,55,56,57,58], and are still on the market to date. Moreover, Tosoh, a Japanese chemical company, has recently extended its ToyoPEARL product line, a polymethacrylate-based resin mainly used for polymer purification with organic solvents, to include products for biological samples processing. A non-exhaustive list of commercially available SEC resins is presented in Table 1. Resins with an opportunity to isolate EV subpopulations are those with highest exclusion limit such as BioGel A-50 m, BioGelA-150 m, and Sephadex G-200. However, these products have been discontinued by their respective manufacturers.

Table 1.
Non-exhaustive list of commercially available SEC resins.
3. Current SEC Methods for EV Separation
EVs is an umbrella term used to generically designate exosomes, ectosomes, microvesicles, microparticles, apoptotic bodies, oncosomes, exomeres, migrasomes, and many others [59,60]. EVs are present in cell culture fluids and bodily fluids and secretions, such as blood, saliva, tears, urine, cerebrospinal fluid (CSF), ascites, synovial fluid, amniotic fluid, semen, and breast milk [61,62,63,64]. EV are heterogeneous in structure and size, ranging from 30 nanometers up to 5 microns. This heterogeneity is driven by the type of the producer cell, its homeostatic condition, the type of biofluid, and the EV separation method. From a biophysical standpoint, the size of an EV is a critical parameter that defines the EV surface curvature, and thus dictates the surface cargo. For instance, the number of proteins that can physically fit on a large vesicle surface is orders of magnitude larger than that of a small vesicle (Figure 1). Similarly, large vesicles can carry more soluble cargo (i.e., nucleic acids and soluble proteins) in their intravesicular space, compared to smaller vesicles. In other words, the protein/lipid ratio (or nucleic acids/lipid ratio) is higher in large EVs versus small EVs. and may thus be an indication of EV bioactivity (Figure 1). This analysis is based on an assumption that a given EV population is homogeneous compositionally (i.e., secreted from a specific cellular subset), although in complex biofluids, size does not necessarily correlate with particle density. Nevertheless, the variation in EV size posits that EVs can theoretically be separated using size-guided techniques, such as SEC. In this section, we discussed the studies that used SEC for EV separation, grouped them into biofluid-type subsections, and highlighted the type of chromatographic methods used, in an effort to distill information that may facilitate in designing novel and improved SEC columns capable of effective EV separation. We will maintain the EV nomenclature used by authors in their publication.
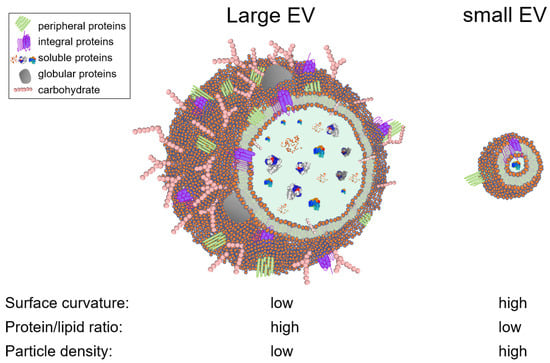
Figure 1.
Schematic depicting the effects of the particle size on the composition and physiochemical properties of EVs. EVs contain other biomolecules that are not represented here, such as nucleic acids, small molecules.
3.1. SEC in Separating Cell Culture Medium
One of the first separations of cell-derived EVs using SEC columns is traced back to a 2012 study [65], where the authors used 16/60 Hiprep Sephacryl S400 HR column to purify exosomes secreted by mouse bone marrow-derived mesenchymal stem cells (MSCs) and MSCs from human umbilical cord Wharton’s jelly. Following separation, the authors showed that the anti-inflammatory activity of MSCs conditioned media (CM) is exosome-associated. Shortly after, two studies reported on the use of Superose 6 column for concentrated CM fractionation [66,67]. In the first study, the authors used a 50 cm column to separate astrocytes CM into four fractions, three of which were EVs [66]. In the second, a preparative Superose 6 column was used to isolate EVs from MCF-7, MDA-MB-231, L3.6pL, and U937 cells [67]. Interestingly, the representative separation profile that was shown [67], was very different from that of astrocytes [66], with only a small EV peak being detected in the void volume. Subsequently, Nordin et al. [68] reported higher yield and preservation of physical and functional properties of EVs isolated from conditioned stem cell media by ultra-filtration (UF)-SEC (S-400 Sephacryl column) compared to ultracentrifugation. Willms et al. [69] combined SEC on a S-1000 Sephacryl column to a 110,000 g ultracentrifugation pellet from 250 mL CM of B16F10 cells to show two compositionally different EV populations, large exosomes enriched for ACTN4 and CCNY and small exosomes enriched for EPHA2, although a complete exosome characterization that includes tetraspanins’ markers was not reported. It was also unclear what column was used because, in the Section 3, a HiPrep Sephacryl S-400 HR 16/60 column was reported, instead of a Sephacryl S-1000 [69]. In a separate study, SEC using a Sephacryl S-400 column outperformed ultracentrifugation when comparing the function of EVs isolated from the CM of 400 million CPC cells [70], but neither the EV chromatogram nor the SEC experimental details were reported [70]. Nonetheless, the same conclusion was reached by another group who cultured BEAS-2B cells and fractionated their CM on a 10 mL Sepharose CL-4B column after an ultrafiltration step (10 KDa), as opposed to the ultracentrifugation method (117,734× g for 2.5 h) [71]. Another size exclusion based technique that involves tangential flow filtration coupled with bind-elute SEC (TFF/BE-SEC) was developed and validated for the CM from two mouse cell lines, neuroblastoma N2a and myoblast C2C12 [72]. Today, pre-packed columns that can be used with or without FPLC with straightforward protocols are commercially available [73], although various cell culture conditions and parameters still need to be empirically determined.
3.2. Blood
One of the first successful EV separations from blood using SEC was reported in 2003, when Taylor et al. [74] isolated membrane fragments (MFs) from the sera of women with stage III ovarian cancer using a 1.5 cm × 4.5 cm bioGel A50m column. The authors found that compared to a healthy group, sera from ovarian cancer women contained significantly elevated levels of MFs. Fas Ligand (FasL), a type-II transmembrane protein that belongs to the tumor necrosis factor family, was associated with the tumor-derived MFs and was responsible for apoptosis of T lymphocytes, through loss of ζ-chain expression [74]. Taylor et al. also used Sepharose 2B column (1 cm × 35 cm) to isolate FasL-positive microvesicles that induced apoptosis of activated T lymphocytes from sera of oral cancer patients [75]. Later, the authors described a protocol for EV separation from blood plasma (as well as ascites) on a 2% agarose-based gel column (2.5 cm × 16 cm) for proteomics analysis and RNA profiling [76]. Based on their cumulative evidence in using SEC for isolating macromolecules, the authors concluded that use of precipitation reagents, such as ExoQuick, yielded increased quantity and quality of exosomal proteins and RNA, followed by SEC, then immunocapture; and the least effective method was ultracentrifugation [76]. However, as the science and technology of EV separation evolves, the precipitation method has been found to co-isolate EVs with other contaminants, and as such, is considered the least EV-specific method [77,78,79,80,81,82].
Arroyo et al. used Sephacryl S-500 to fractionate human plasma and showed that the majority of circulating miRNAs are associated with non-vesicular Ago-2 complexes, whereas only a few miRNA were EV-associated [83]. For instance, microRNAs miR-16 and miR-92a were mostly EV-free, whereas circulating let-7a was enriched in the EV fractions. On the other hand, by separating EVs from the High Density Lipoprotein (HDL) fractions using Superose 6 or Sephacryl S-200 FPLC columns, Vickers et al. showed that plasma miRNA are mostly HDL-associated [84]. In contrast, there exist studies demonstrating that extracellular RNAs are carried in EVs from human [85,86] and mouse [87] blood plasma. However, these studies did not use SEC for EV separation, making it difficult to compare.
Aside from commercially available SEC columns, the use of in-house columns has also been reported. Böing et al. [88] packed Sepharose CL-2B resin in a 10 mL plastic syringe stuffed with nylon stocking to create a 1.6 cm × 6.2 cm column. The column was used to demonstrate a single-step protocol for vesicles separation from human body fluids [88]. Soon after, this SEC method of EV separation from plasma was compared to that using a commercially available spin column (Exo-SpinTM) and, after detailed mass spectrometry analysis, the early SEC fractions, but not Exo-Spin derived EVs, were found to contain known EV markers [89]. This observation contradicts a concurrent report that concluded that Exo-Spin columns are excellent 10-min EV separation protocol from complex biofluid such as plasma [90]. The differences in these observations may be due to variations in donor EV cargo [62]. In their study, Baranyai et al. [91] used three different SEC columns (Sephacryl S-400, Sepharose 4B, Sepharose CL-2B) and compared the efficiency of blood plasma EVs separation to multiple rounds of ultracentrifugation. The authors showed that Sephacryl S-400 and Sepharose 4B, but not Sepharose CL-2B, efficiently separated EVs from albumin. It should be noted that EV mass ranges from mega to gigadaltons [92], and thus, several orders of magnitude larger than albumin which is 66.4 kDa. Thus, a mere separation between EV markers and albumin likely hides contaminants from other plasma particles with in-between molecular weight, such as HDL, LDL, and other immunoglobulins complexes. Nevertheless, Hong et al. [93] reported separation of intact and bioactive EVs from plasma of healthy controls and cancer patients using a mini-SEC 1.5 cm × 12 cm Sepharose 2B packed column. Subsequently, Gamez Valero et al. [94] compared this SEC method to the PEG precipitation method as well as the total protein organic solvent precipitation method from plasma samples to find that SEC outperformed the latter two methods, as evidenced by nanoparticle tracking analysis (NTA), flow cytometry, cryo-TEM, and immunoblot data. However, what is often ignored is that blood plasma is a complex fluid that contains multiple bionanoparticles of similar sizes and similar densities. Hence, it is likely very difficult to develop a one-step protocol that can efficiently separate all bionanoparticles from plasma [95], although this is still a matter of debate [96,97]. This limitation is highlighted in studies where plasma EVs were not separated from lipoproteins for proteomics studies, until a two-step protocol (density-gradient ultracentrifugation followed by SEC of the high-density fraction) was performed [98], this is also visible where studies that only used commercially available SEC columns co-purified non-EV particles with the EV preparations [99,100]. In any case, what is needed and not systematically performed is to test SEC fractions for other bionanoparticle markers, such as ApoB100, ApoA1, and Ago-2 proteins, in addition to tetraspanins and albumin. Another important control that is often overlooked is the inclusion of the non-EV fractions as a negative control for NTA, TEM, and immunoblot analyses.
3.3. Semen
Prostasomes are a class of EVs secreted by the prostate gland into the seminal plasma. The term “prostasomes” was coined in 1985 by Gunnar Ronquist (for more details about the history and function of prostasomes, see [101,102,103]). The chromatographic fractionation of seminal plasma is also decades old. Peeker et al. [104] fractionated clarified seminal plasma using a 1.6 cm × 60 cm Superdex 200 column, although their focus was not on prostasomes. On the other hand, Saez et al. [105] demonstrated an anti-oxidant property of human prostasomes that they purified using a combination of ultracentrifugation and Sephadex G-200 chromatography. More recently, Jones et al. [106] used SEC on a 35 mL Sephacryl S-300 column as an orthogonal method to demonstrate the presence of galectin-3, an immunomodulatory galactose-specific lectin, in human prostasomes. Two distinct prostasomes populations were isolated from seminal fluids from healthy vasectomized men using a combination of SEC on 70 cm × 2.6 cm Sephacryl S-1000 and density gradient ultracentrifugation [107,108]. The density-gradient ultracentrifugation step clearly separated a high-density luminal GLIPR2-rich population from a low-density membranous ANXA1-rich population [107]. Both populations exhibited atypical high levels of sphingomyelin, cholesterol, and glycosphingolipids and low levels of phosphatidylcholine, and differed from each other in the relative levels [108]. However, the chromatogram of the separation was not shown and it appears that the density gradient step is what allowed separation of seminal plasma into two different sizes of seminal EV (SEV)-populations, whereas SEC was used for washing EVs [108]. In their study, Lyu et al., used a 22 cm × 1 cm Sephadex G-50 column to isolate SEVs from HIV-infected and uninfected men who use or did not use illicit substances [109]. The separation profiles showed two peaks with the void peak containing the SEVs, as demonstrated by NTA, TEM, western blot, and turbidity data [109]. The group investigated the effects of HIV-1 infection and illicit drug use on the structure and function of SEVs, and concluded that HIV-SEV, Drug-SEV, and HIV-Drug-SEV may mediate pathological processes by differentially inducing monocytes’ actin-cytoskeletal re-organization, adhesion, ECM-modifying metalloprotease secretion, and chemotactic migration toward HIV secretome [109]. Lessons from study [109] led the group to develop a gradient SEC (gSEC) column in which all available Sephadex beads (Table 1) were layered in a 100 × 1 cm column [110]. The gSEC column yielded four distinct peaks, the first two of which correspond to two (large and small) EV populations, the third peak was enriched in membraneless condensates, comprising small RNAs and a unique protein signature, but clearly lacking a membrane whereas the fourth contained small molecules and salts [110]. Thus far, seminal plasma is one of the highest EV-containing body fluids, with diverse cargos and functions [62,111,112,113,114].
3.4. Urine
Urinary extracellular vesicles (uEV) have been proposed as source of biomarkers for kidney disorders, such as diabetic nephropathy, glomerular disease, and bladder cancer [115,116]. The clinical relevance of urine is not new and urinary proteins have been chromatographically fractionated for decades [117]. However, pathology-associated differences in uEV were not known until the study by Rood et al. [118]. The team used ultracentrifugation followed by SEC on a BioSep-SEC-S4000 column from Phenomenex (Torrance, CA, USA) to profile uEVs. The authors noted differences in chromatograms of urine from patients with membranous nephropathy or with focal segmental glomerulosclerosis compared to that of a healthy patient. Noteworthy is that the authors’ protocol involved multiple ultracentrifugation steps and protein reduction by DTT to remove soluble Tamm–Horsfall protein prior to SEC [118]. Interestingly, Kaddour et al. [110] obtained better resolved chromatogram of healthy urine using ultrafiltration followed by chromatography on a 100 cm × 1 cm Sephadex gSEC column, without ultracentrifugation or protein reduction. On the other hand, Lozano-Ramos [119] used a 10 mL Sepharose-CL-2B packed column to show clear EV separation from bulk urine proteins, as evidenced by NTA, flow-cytometry, cryo-TEM, immunoblot, and silver stain. The same group then performed in a separate study focusing on small RNA-Seq and proteomics analyses of uEVs from living and deceased kidney donors, but found only minor differences in the EV profiles that could not discriminate between the two groups [120]. Thus, questions on whether uEVs can be used as biomarkers for health and diseases remains largely open.
3.5. Other Bodily Fluids
SEC has been used to isolate EVs from other bodily fluids, including saliva, tears, milk, CSF, peritoneal dialysis efflux, and synovial fluid. Ogawa et al. [121] used 1.5 cm × 50 cm Sepharose CL-4B column to isolate saliva EVs containing active dipeptidyl peptidase IV (DPP IV) and galectin-3 and immunoglobulin A. Soon after, the same group used Sephacryl S-500 packed column (1.5 cm × 50 cm) to identify two saliva-derived EV populations [122]. Aqrawi et al. [123] used commercially available (Izon Science Ltd., New Zealand) qEV column to isolate EVs from saliva and tears. Sephacryl S-500 was also successfully used to fractionate milk fluff and serum into 4 and 5 peaks, respectively; allowing human and bovine milk EVs (MEVs) separation in a reproducible manner [124]. In their study, Kaddour et al. [110] used gSEC Sephadex column to obtain a 4-peak profile from cow milk, identifying a decreasing intensity in the EV peak in the order fat-free <2% fat < whole milk. Clarified CSF from young (<2 years) and old (>70 years) patients was purified on a S400-HR Sephacryl spin column and the filtrates were concentrated by the precipitation method (ExoQuick or total exosome isolation reagent) [125]. Although this method, has not been proven efficient for EV separation from any other biofluid, much less for CSF, the study identified over 50 differentially present miRNA between the two groups [125]. Sepharose CL-2B column was used post ultrafiltration as a concentrating step to isolate EVs from peritoneal dialysis (PD) efflux. A subsequent proteomics analysis revealed a good hierarchical clustering of samples from long term (>18 month) vs short-term (<10 month) PD patients [126]. Finally, separation of human synovial fluid (SF) EVs devoid of serum albumin and apolipoprotein A-I, was achieved on a HiPrep 26/60 Sephacryl S-500 HR column, outperforming ultracentrifugation and sucrose density-gradient ultracentrifugation [127]. Taken together, these studies demonstrate the superiority and usefulness of the SEC technique for separation of EVs from any biofluid type.
4. Discussion of the SEC Techniques Used for EV Separation
Particle size has been the key physical feature for EV definition; thus, a size-guided separation of EVs is unavoidable, potential crossover between subtypes notwithstanding. Yet, current size-exclusion columns, as reported in the section above, have been mostly used for bulk EV purification, with differing success. While our intent here was not to conduct a systematic review, it becomes evident from the 35 revisited studies that SEC is far from being standardized in the field, even within studies from the same group (Table 2). Furthermore, the use of automated systems such as HPLC and FPLC in the separation protocols are reported as automation is critical to overall performance and reproducibility, compared to the gravity-driven separations (Table 2). However, it becomes now evident that for heterogeneous bionanoparticle separation from complex biofluids, heterogeneous columns are needed, not only for separating EVs from non-EV components, but also for efficient one-step subpopulation separation. In this regard, gSEC columns coupled with a Particle Purification Liquid Chromatography (PPLC) system that can characterize EVs in real-time during separation would be valuable. The proof-of-concept of such approach was recently demonstrated by Kaddour et al., [110]. With empirical attention to separation parameters such as column length and width, sample volume, flow rate, column pressure, detectors parameters, and volume of collected fractions (see section below), it is possible to envisage a one-step protocol that achieves EV subpopulation separation with remarkable resolution.

Table 2.
A non-exhaustive list of studies that used SEC for EV separation.
5. Future Directions: Towards a Particle Purification Liquid Chromatography System
Despite the unanimous agreement in the field that SEC is a superior particle separation method [33,128], the method is still in need of empirical standardization with respect to the columns, the separation conditions, and the choice of the chromatographic system. We will discuss these parameters in this section. We will also highlight the benefits of a new EV purification system such as PPLC and how such a system may alleviate EVs separation challenges (Figure 2, Table 3).
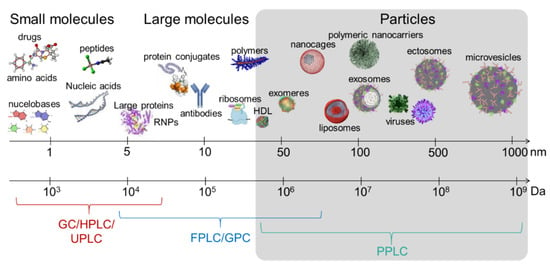
Figure 2.
Schematic depicting the limits of the commercially available chromatographic systems in EV separation. GC, gas chromatography; HPLC, high performance liquid chromatography; UPLC, ultra-high performance liquid chromatography; FPLC, fast purification liquid chromatography; GPC, gel permeation chromatography. PPLC, particle purification liquid chromatography, which is currently under development (Kaddour et al.,), is tailored for mega- to gigadalton materials.

Table 3.
Summary of EV separation challenges and possible solutions using a PPLC system.
5.1. Column Chemistry
In general, the column chemistry is of utmost importance in chromatographic separation. We have seen in Section 2 that resin manufacturing has been focused on bead homogeneity and decreased bead size, for increased reproducibility and accommodation of higher flow rates and faster runs. We have also seen a trend of resins with high exclusion limit discontinued from manufacturing. However, for the sought goals of EV subpopulation separation, both practices are amiss. Indeed, as noted previously EVs are in the mega- to gigadalton range [92], thus one would need beads with large exclusion limits (i.e, >500 MDa) and a gradient of different pore sizes [110] for a more resolved EV separation. An additional approach that may add significant resolution to EV separation is to employ columns of hybrid chemistries. In this regard, Van Deun et al. [129] showed that a dual-mode chromatography (DMC), employing ion exchange chromatography (IEC) and SEC column, can help isolate purer plasma EVs compared to SEC alone. While the proof of concept was validated, some basic but critical chromatographic principles were overlooked. For instance, the authors omitted an important control in which the IEC-bound fraction would be eluted and tested. It is possible that differently charged EV particles were eliminated as suggested by the decreased CD63 levels in DMC compared to SEC alone. In addition, the SEC resin was layered atop the IEC resin; but in this design, elution of the IEC-bound fraction would have not benefited from the presence of the SEC resin. In other words, the IEC resin played the role of a trap instead of a second dimension of separation. In contrast, if the IEC portion of the column were atop, the SEC portion of the column could have thus fractionated both IEC-unbound and IEC-bound material and operated as a desalting step for the bound-elution, resulting in a one-step 2D separation profile. Nevertheless, the results demonstrated a promising proof-of-concept that needs improvement. Another example of hybrid chromatography proof-of-concept is illustrated in an old study that used a PBS-washed 104k pellet of human seminal plasma in a three-step chromatographic purification: carboxymethyl cellulose (a cation exchange), Concanavalin A (a carbohydrate-binding protein), and 1.5 cm × 30 cm Sepharose CL-6B (size exclusion) to isolate and characterize an anti-fertility factor [130]. Although each of the separation steps was used independently and the resulting peaks were re-concentrated by ultracentrifugation before being applied to the next column, the successful separations support the concept of hybrid chromatography. In summary, available SEC columns and resins on the market do not provide enough resolution for EV subpopulation separation. However, there exist numerous opportunities for optimization, such as development of hybrid columns, that may significantly expand the dimensions for complex biofluid separation.
5.2. Online Characterization
Although SEC is often thought as a purification technique, it arguably offers a wide range of analytical potential [131] that is not well appreciated in the field. For instance, the molecular mass of a solute can mathematically be inferred, but also solute interactions and solute self-association can be characterized in terms of reaction stoichiometry and equilibrium constant [131]. These are valuable biochemical information. Furthermore, UV-Vis spectra can be employed to calculate biophysical information such as particle size and concentration. Strikingly however, the EV field has to date almost exclusively relied on analytical techniques that utilize light scattering detectors: DLS, NTA, and flow cytometry, known to have detection limits on the particle size, concentration determination, and labeling, respectively. There is paucity of studies in the literature that applied turbidity-based modeling for size determination of liposomes [132] and protocells [133]. However, Kaddour et al., [110] was the first to adapt Lorenz-Mie solution for EV size and concentration determination, taking advantage of HoloPy, a publicly available python package for digital holograms and light scattering analyses [134]. Moreover, SEC studies that used HPLC and FPLC systems often monitored elution profiles at wavelengths (220, 230, 260, or 280 nm) that confound EVs with non-EVs materials, such as extracellular condensates and other cell-free nucleic acids, proteins, and small molecules [110]. Since EVs are membranous and scatter light in the visible range, it is thus more relevant to monitor EVs in the turbidity range (400–600 nm). As other biological molecules may also exhibit absorbance around 400 nm, turbidity ratios R1 = A400/A600 and R2 = A600/A650 were defined to precisely distinguish between EVs and non-EVs during SEC separation [110].
The concept of online EV characterization during SEC is not limited to UV-Vis. Other sophisticated detectors can also be employed as long as they are non-invasive. In fact, DLS has been used for EV monitoring to precisely identify the EV-containing fractions when the EVs were overwhelmed by the presence of free-proteins in the UV range [135]. Fluorescence has also been employed for EV analysis during SEC separations and from which fluctuation correlation spectroscopy (FCS) analysis was employed to derive particle size [136]. However, DLS is beneficial only when EV are sufficiently diluted, unlike UV-Vis that can withstand relatively high EV concentrations. Additionally, DLS does not differentiate between EVs and possibly high scattering non-EVs particles such as HDL, LDL, and VLDL. On the other hand, fluorescence detection has great potential because, in addition to its capability to orthogonally determine vesicle size, antibody fluorescence can be used to determine levels of EV-associated tissue-specific markers in body fluids [136]. For instance, with a six-channel detector, it is possible to multiplex EV-positive, EV-negative, and tissue- or disease-specific markers, and obtain real-time information during the separation. This can be achieved in a one-step protocol, with minimal hands-on time, and without the need for a flow cytometer.
5.3. Fraction Collection
One of the fundamental limitations in the EV field arises from the lack of a high-resolution and reproducible EV sub-fractionation, exacerbating the EV inherent heterogeneity problem. What is also lacking is the retrieval of EV sub-fractions for subsequent functional studies, such as the determination of the protein and nucleic acid levels, as well as the deciphering of molecular and phenotypic effects of EVs on the target cells. Newly developed single-vesicle techniques (microfluidic devices, high-resolution confocal imaging, and nano-flow cytometry [137]) offer unique opportunities to tackle the formidable EV heterogeneity challenge, but leave the EV-retrieval problem unaddressed. The drawback of these techniques is that they require very diluted input material and/or are destructive in nature. In contrast, PPLC columns as described above, when coupled with a fast and small-volume fraction collector, have the potential of achieving EV sub-fractionation in near-native conditions and preparative amounts. Indeed, PPLC uses a multi-96 well plate fraction collector with as little as 20 microliters per fraction [110], whereas the majority of currently available FPLC and HPLC fraction collectors are designed for relatively large volume. Thus, the combination of high-resolution columns and small volume fraction collector of PPLC may allow retrieval of intact EV subpopulations in a convenient format for downstream compositional or functional studies.
5.4. Additional Factors for Consideration
5.4.1. Pre and Post Column Concentrating Steps and Mobile Phase
Some of the samples such as cell culture supernatants and urine may require a pre-concentrating step before EV separation. Furthermore, depending on the downstream applications, purified EVs may also require re-concentration. The most used concentration method is ultrafiltration using membrane centrifugal devices ranging from 3 to 100 kDa. However, particles may irreversibly bind to the membrane filter leading to significant loss in yield. To overcome this hurdle, membrane pre-saturation with a 1% BSA solution and membrane wash with PBS post-concentrating are often recommended. However, when using a hybrid column (see Section 5.1 above), a pre-concentrating step of dilute input samples may not be needed anymore, because the ion exchange portion of the column can act as a pre-concentrating step. By contrast, hybrid IEX/SEC columns require longer runs with additional mobile phase solutions for a gradient- or step-elution of the IEX-bound material. Regarding the post-concentration step, an inexpensive and efficient alternative to ultrafiltration is concentration by reduced-pressure evaporation or by freeze-drying. In this case, the amount of salt in the sample should be noted and ideally, the mobile phase solution adapted (i.e., 0.1X PBS mobile phase in a 10X re-concentration protocol). In other cases, for instance when preparing samples for TEM analysis, 10 mM Tris buffer is more suitable than 1X PBS. However, changing buffers or lowering their concentration in mobile phase may allow non-specific binding between the sample proteins and the stationary phase. Thus, these separation parameters should be empirically determined and optimized per sample type.
5.4.2. Scalability
As tissue-specific EVs circulating in some body fluids may be rare events compared to the other circulating materials, sample scale-up becomes unavoidable to obtaining preparative amounts of the specific EV subtype, needed for downstream applications. By contrast, other samples such as CSF and mosquito EVs [138] are very limited requiring rather a down-scale to accommodate low-volume input. Fortunately, PPLC, in particular, and chromatography, in general, are versatile. Indeed, chromatography separation utilizes various column sizes ranging from as little as a hundred microliters bead volume to over a hundred liters in industrial settings. Within a specific setting, chromatography systems are also versatile, by accommodating easily interchangeable tubing with various inner diameters. Finally, the PPLC fraction collection (see Section 5.3 above) can also be easily adapted to collect fractions of as little as 20 microliters up to 5 mm. The PPLC adaptability to the sample type and the needed scale, within the limits of the detectors at the given settings, is invaluable because it will permit freedom in experimental design for both analytical and preparative purposes.
5.4.3. Biohazard Containment
There is need for systems that account for biohazard management, where separate columns/piping are used for separating infectious fluids and non-infectious fluids. For example, PPLC can be used for separating viruses from other EVs. Since PPLC is a closed system, it can be operated in a BSL1 laboratory. However, infectious fluids should be processed in a Class II BSC and in the BSL2+ laboratory, as adequate environment for handling fluids containing infectious materials, such as HIV. Furthermore, sodium azide (0.01–0.05%) may be added to the mobile phase to prevent bacterial contamination, especially when handling patient samples. However, if isolated EVs are meant for functional cell-culture based studies, sodium azide should be removed from the system.
6. Conclusions
In this review we have methodically provided summaries of primary data as evidence that SEC, and chromatography in general, can be leveraged to develop a multi-parameter resource for the purification, online characterization, and retrieval of preparative quantities of near-native, pure, and functional EV subpopulations from diverse biofluids. However, in spite of its long existence, affordability, and widespread availability, SEC is still in its infancy in the EV field. Hence, resources are needed for researchers and innovators to develop and standardize SEC-based EV separation protocols for each biofluid. Based on the EV separation techniques that are currently available, along with the potential of chromatography and recent improvements, such as PPLC, we envisage that the development of hybrid columns on a PPLC platform will be a compelling unifier for EV subpopulation separation and characterization. Such a hybrid system can also be applied to other biological vesicles such as viruses, or synthetic nanoparticles, including liposomes, polymers, and nanocages.
7. Patents
Stony Brook University filed a US patent application (PCT US2020/030914) covering the PPLC platform, with HK and CMO as inventors. Issues related to intellectual properties will be managed by the Office of Technology Licensing and Industry Relations (OTLIR) at Stony Brook University, according to existing and standard policies.
Author Contributions
Conceptualization, H.K. and C.M.O.; writing—original draft, H.K.; writing—review and editing, H.K., M.T. and C.M.O.; funding acquisition, C.M.O. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the National Institute on Drug Abuse (NIDA), grants number DA042348, DA050169, and DA053643 to C.M.O.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Acknowledgments
The authors would like to apologize to colleagues whose work were inadvertently not cited. The authors thank Nadia Shouman from the Okeoma Laboratory for assembling Table 2, as well as the other members for helpful discussions. Our gratitude goes to Bryson Okeoma from Stony Brook University for his critical reading of the manuscript.
Conflicts of Interest
The authors declare no conflict of interest, beyond the patent application related to this work. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
References
- Gage, S.H.; Fish, P.A. Fat digestion, absorption, and assimilation in man and animals as determined by the dark-field microscope, and a fat-soluble dye. Am. J. Anat. 1924, 34, 1–85. [Google Scholar] [CrossRef]
- Edmunds, J. Note on a New Paraboloid Illuminator for use beneath the Microscope Stage. Also Note on the Resolution of Podura Scale by means of the New Paraboloid. Mon. Microsc. J. 1877, 18, 78–85. [Google Scholar] [CrossRef][Green Version]
- Müller, F. Über einen bisher nicht beobachteten Formbestandteil des Blutes. Cent. f. Pathol. Anat 1896, 7, 529–539. [Google Scholar]
- Frazer, A.C.; Stewart, H.C. Ultramicroscopic particles in normal human blood. J. Physiol. 1937, 90, 18–30. [Google Scholar] [CrossRef] [PubMed]
- Elkes, J.J.; Frazer, A.C.; Stewart, H.C. The composition of particles seen in normal human blood under dark-ground illumination. J. Physiol. 1939, 95, 68–82. [Google Scholar] [CrossRef] [PubMed]
- Chargaff, E.; West, R. The biological significance of the thromboplastic protein of blood. J. Biol. Chem. 1946, 166, 189–197. [Google Scholar] [CrossRef]
- Frailing, J.M.; Owen, C.I. Chylomicrons of Blood: A Resumé of Literature and a Recommended Counting Procedure. Am. J. Clin. Pathol. 1951, 21, 508–514. [Google Scholar] [CrossRef]
- Wolf, P. The Nature and Significance of Platelet Products in Human Plasma. Br. J. Haematol. 1967, 13, 269–288. [Google Scholar] [CrossRef]
- Abels, E.R.; Breakefield, X.O. Introduction to Extracellular Vesicles: Biogenesis, RNA Cargo Selection, Content, Release, and Uptake. Cell. Mol. Neurobiol. 2016, 36, 301–312. [Google Scholar] [CrossRef]
- Kalra, H.; Drummen, G.P.; Mathivanan, S. Focus on extracellular vesicles: Introducing the next small big thing. Int. J. Mol. Sci. 2016, 17, 170. [Google Scholar] [CrossRef]
- Raposo, G.; Stoorvogel, W. Extracellular vesicles: Exosomes, microvesicles, and friends. J. Cell Biol. 2013, 200, 373–383. [Google Scholar] [CrossRef] [PubMed]
- Araldi, E.; Krämer-Albers, E.-M.; Hoen, E.N.-t.; Peinado, H.; Psonka-Antonczyk, K.M.; Rao, P.; van Niel, G.; Yáñez-Mó, M.; Nazarenko, I. International Society for Extracellular Vesicles: First annual meeting, April 17–21, 2012: ISEV-2012. J. Extracell. Vesicles 2012, 1, 19995. [Google Scholar] [CrossRef]
- Lötvall, J.; Rajendran, L.; Gho, Y.-S.; Thery, C.; Wauben, M.; Raposo, G.; Sjöstrand, M.; Taylor, D.; Telemo, E.; Breakefield, X.O. The launch of Journal of Extracellular Vesicles (JEV), the official journal of the International Society for Extracellular Vesicles-about microvesicles, exosomes, ectosomes and other extracellular vesicles. J. Extracell. Vesicles 2012, 1. [Google Scholar] [CrossRef] [PubMed]
- Ainsztein, A.M.; Brooks, P.J.; Dugan, V.G.; Ganguly, A.; Guo, M.; Howcroft, T.K.; Kelley, C.A.; Kuo, L.S.; Labosky, P.A.; Lenzi, R. The NIH extracellular RNA communication consortium. J. Extracell. Vesicles 2015, 4, 27493. [Google Scholar] [CrossRef]
- Laurent, L.C.; Abdel-Mageed, A.B.; Adelson, P.D.; Arango, J.; Balaj, L.; Breakefield, X.; Carlson, E.; Carter, B.S.; Majem, B.; Chen, C.C.; et al. Meeting report: Discussions and preliminary findings on extracellular RNA measurement methods from laboratories in the NIH Extracellular RNA Communication Consortium. J. Extracell. Vesicles 2015, 4, 26533. [Google Scholar] [CrossRef]
- Witwer, K.W.; Buzás, E.I.; Bemis, L.T.; Bora, A.; Lässer, C.; Lötvall, J.; Nolte-‘t Hoen, E.N.; Piper, M.G.; Sivaraman, S.; Skog, J.; et al. Standardization of sample collection, isolation and analysis methods in extracellular vesicle research. J. Extracell. Vesicles 2013, 2, 20360. [Google Scholar] [CrossRef] [PubMed]
- Hill, A.F.; Pegtel, D.M.; Lambertz, U.; Leonardi, T.; O’driscoll, L.; Pluchino, S.; Ter-Ovanesyan, D.; Nolte-‘t Hoen, E.N. ISEV position paper: Extracellular vesicle RNA analysis and bioinformatics. J. Extracell. Vesicles 2013, 2, 22859. [Google Scholar] [CrossRef]
- Lener, T.; Gimona, M.; Aigner, L.; Börger, V.; Buzas, E.; Camussi, G.; Chaput, N.; Chatterjee, D.; Court, F.A.; Portillo, H.A.d.; et al. Applying extracellular vesicles based therapeutics in clinical trials–an ISEV position paper. J. Extracell. Vesicles 2015, 4, 30087. [Google Scholar] [CrossRef]
- Mateescu, B.; Kowal, E.J.; van Balkom, B.W.; Bartel, S.; Bhattacharyya, S.N.; Buzás, E.I.; Buck, A.H.; de Candia, P.; Chow, F.W.; Das, S. Obstacles and opportunities in the functional analysis of extracellular vesicle RNA–an ISEV position paper. J. Extracell. Vesicles 2017, 6, 1286095. [Google Scholar] [CrossRef]
- Théry, C.; Witwer, K.W.; Aikawa, E.; Alcaraz, M.J.; Anderson, J.D.; Andriantsitohaina, R.; Antoniou, A.; Arab, T.; Archer, F.; Atkin-Smith, G.K. Minimal information for studies of extracellular vesicles 2018 (MISEV2018): A position statement of the International Society for Extracellular Vesicles and update of the MISEV2014 guidelines. J. Extracell. Vesicles 2018, 7, 1535750. [Google Scholar] [CrossRef]
- Coumans, F.A.; Brisson, A.R.; Buzas, E.I.; Dignat-George, F.; Drees, E.E.; El-Andaloussi, S.; Emanueli, C.; Gasecka, A.; Hendrix, A.; Hill, A.F. Methodological guidelines to study extracellular vesicles. Circ. Res. 2017, 120, 1632–1648. [Google Scholar] [CrossRef] [PubMed]
- Gardiner, C.; Vizio, D.D.; Sahoo, S.; Théry, C.; Witwer, K.W.; Wauben, M.; Hill, A.F. Techniques used for the isolation and characterization of extracellular vesicles: Results of a worldwide survey. J. Extracell. Vesicles 2016, 5, 32945. [Google Scholar] [CrossRef] [PubMed]
- Linares, R.; Tan, S.; Gounou, C.; Arraud, N.; Brisson, A.R. High-speed centrifugation induces aggregation of extracellular vesicles. J. Extracell. Vesicles 2015, 4, 29509. [Google Scholar] [CrossRef] [PubMed]
- Shao, H.; Im, H.; Castro, C.M.; Breakefield, X.; Weissleder, R.; Lee, H. New Technologies for Analysis of Extracellular Vesicles. Chem. Rev. 2018, 118, 1917–1950. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Freitas, D.; Kim, H.S.; Fabijanic, K.; Li, Z.; Chen, H.; Mark, M.T.; Molina, H.; Martin, A.B.; Bojmar, L. Identification of distinct nanoparticles and subsets of extracellular vesicles by asymmetric flow field-flow fractionation. Nat. Cell Biol. 2018, 20, 332–343. [Google Scholar] [CrossRef] [PubMed]
- Shao, H.; Chung, J.; Balaj, L.; Charest, A.; Bigner, D.D.; Carter, B.S.; Hochberg, F.H.; Breakefield, X.O.; Weissleder, R.; Lee, H. Protein typing of circulating microvesicles allows real-time monitoring of glioblastoma therapy. Nat. Med. 2012, 18, 1835. [Google Scholar] [CrossRef]
- Arkesteijn, G.; Nauwelaers, F.; van den Engh, G.; Wauben, M. Prerequisites for the analysis and sorting of extracellular vesicle subpopulations by high-resolution flow cytometry. Cytometry. Part A J. Int. Soc. Anal. Cytol. 2015, 89, 135–147. [Google Scholar]
- Kormelink, T.G.; Arkesteijn, G.J.; Nauwelaers, F.A.; van den Engh, G.; Nolte-’t Hoen, E.N.; Wauben, M.H. Prerequisites for the analysis and sorting of extracellular vesicle subpopulations by high-resolution flow cytometry. Cytom. Part A 2016, 89, 135–147. [Google Scholar] [CrossRef]
- Tian, Y.; Ma, L.; Gong, M.; Su, G.; Zhu, S.; Zhang, W.; Wang, S.; Li, Z.; Chen, C.; Li, L. Protein profiling and sizing of extracellular vesicles from colorectal cancer patients via flow cytometry. ACS Nano 2018, 12, 671–680. [Google Scholar] [CrossRef]
- Choi, D.; Montermini, L.; Jeong, H.; Sharma, S.; Meehan, B.; Rak, J. Mapping subpopulations of cancer cell-derived extracellular vesicles and particles by nano-flow cytometry. ACS Nano 2019, 13, 10499–10511. [Google Scholar] [CrossRef]
- van der Pol, E.; de Rond, L.; Coumans, F.A.; Gool, E.L.; Böing, A.N.; Sturk, A.; Nieuwland, R.; van Leeuwen, T.G. Absolute sizing and label-free identification of extracellular vesicles by flow cytometry. Nanomed. Nanotechnol. Biol. Med. 2018, 14, 801–810. [Google Scholar] [CrossRef]
- Clayton, A.; Court, J.; Navabi, H.; Adams, M.; Mason, M.D.; Hobot, J.A.; Newman, G.R.; Jasani, B. Analysis of antigen presenting cell derived exosomes, based on immuno-magnetic isolation and flow cytometry. J. Immunol. Methods 2001, 247, 163–174. [Google Scholar] [CrossRef]
- Monguió-Tortajada, M.; Gálvez-Montón, C.; Bayes-Genis, A.; Roura, S.; Borràs, F.E. Extracellular vesicle isolation methods: Rising impact of size-exclusion chromatography. Cell. Mol. Life Sci. 2019, 76, 2369–2382. [Google Scholar] [CrossRef]
- Gholizadeh, S.; Draz, M.S.; Zarghooni, M.; Sanati-Nezhad, A.; Ghavami, S.; Shafiee, H.; Akbari, M. Microfluidic approaches for isolation, detection, and characterization of extracellular vesicles: Current status and future directions. Biosens. Bioelectron. 2017, 91, 588–605. [Google Scholar] [CrossRef]
- Théry, C.; Amigorena, S.; Raposo, G.; Clayton, A. Isolation and Characterization of Exosomes from Cell Culture Supernatants and Biological Fluids. Curr. Protoc. Cell Biol. 2006, 30, 3–22. [Google Scholar] [CrossRef]
- Zhang, H.; Lyden, D. Asymmetric-flow field-flow fractionation technology for exomere and small extracellular vesicle separation and characterization. Nat. Protoc. 2019, 14, 1027–1053. [Google Scholar] [CrossRef] [PubMed]
- Gordon, A.H.; Martin, A.J.; Synge, R.L. Partition chromatography in the study of protein constituents. Biochem. J. 1943, 37, 79–86. [Google Scholar] [CrossRef] [PubMed]
- Gordon, A.; Martin, A.; Synge, R. Technical notes on the partition chromatography of acetamino-acids with silica gel. Biochem. J. 1944, 38, 65–68. [Google Scholar] [CrossRef] [PubMed]
- Martin, A.; Synge, R. Separation of the higher monoamino-acids by counter-current liquid-liquid extraction: The amino-acid composition of wool. Biochem. J. 1941, 35, 91–121. [Google Scholar] [CrossRef]
- Ettre, L.S. The birth of partition chromatography. LCGC N. Am. 2001, 19, 506–512. [Google Scholar]
- Lindqvist, B.; StorgÅRds, T. Molecular-sieving Properties of Starch. Nature 1955, 175, 511–512. [Google Scholar] [CrossRef]
- Porath, J.; Flodin, P.E.R. Gel Filtration: A Method for Desalting and Group Separation. Nature 1959, 183, 1657–1659. [Google Scholar] [CrossRef]
- Belew, M.; Porath, J.; Fohlman, J.; Janson, J.-C. Adsorption phenomena on sephacryl® S-200 superfine. J. Chromatogr. A 1978, 147, 205–212. [Google Scholar] [CrossRef]
- Morgan-Capner, P.; Davies, E.; Pattison, J. Rubella-specific IgM detection using Sephacryl S-300 gel filtration. J. Clin. Pathol. 1980, 33, 1082–1085. [Google Scholar] [CrossRef]
- Morgan, G.; Ramsden, D.B. Comparison of the gel chromatographic properties of Sephacryl S-200 Superfine and Sephadex G-150. J. Chromatogr. A 1978, 161, 319–323. [Google Scholar] [CrossRef]
- Iverius, P.-H. Coupling of glycosaminoglycans to agarose beads (Sepharose 4B). Biochem. J. 1971, 124, 677–683. [Google Scholar] [CrossRef]
- Thompson, S.T.; Cass, K.H.; Stellwagen, E. Blue dextran-sepharose: An affinity column for the dinucleotide fold in proteins. Proc. Natl. Acad. Sci. USA 1975, 72, 669–672. [Google Scholar] [CrossRef] [PubMed]
- Ogata, S.-I.; Muramatsu, T.; Kobata, A. Fractionation of Glycopeptides by Affinity Column Chromatography on Concanavalin A-Sepharose1. J. Biochem. 1975, 78, 687–696. [Google Scholar] [CrossRef]
- Okada, N.; Shindo-Okada, N.; Nishimura, S. Isolation of mammalian tRNAAsp and tRNATyr by lectin-Sepharose affinity column chromatography. Nucleic Acids Res. 1977, 4, 415–423. [Google Scholar] [CrossRef]
- Weisgraber, K.H.; Mahley, R.W. Subfractionation of human high density lipoproteins by heparin-Sepharose affinity chromatography. J. Lipid Res. 1980, 21, 316–325. [Google Scholar] [CrossRef]
- Hauschka, P.; Mavrakos, A.; Iafrati, M.; Doleman, S.; Klagsbrun, M. Growth factors in bone matrix. Isolation of multiple types by affinity chromatography on heparin-Sepharose. J. Biol. Chem. 1986, 261, 12665–12674. [Google Scholar] [CrossRef]
- Hjertén, S. A new method for preparation of agarose for gel electrophoresis. Biochim. Biophys. Acta 1962, 62, 445–449. [Google Scholar] [CrossRef]
- Hjertén, S.; Mosbach, R. “Molecular-sieve” chromatography of proteins on columns of cross-linked polyacrylamide. Anal. Biochem. 1962, 3, 109–118. [Google Scholar] [CrossRef]
- Draus, F. Separation of a cat submaxillary mucoid on Biogel. Arch. Oral Biol. 1965, 10, 719–721. [Google Scholar] [CrossRef]
- Jos, J.; Charbonnier, L.; Mosse, J.; Olives, J.; De Tand, M.-F.; Rey, J. The toxic fraction of gliadin digests in coeliac disease. Isolation by chromatography on Biogel P-10. Clin. Chim. Acta 1982, 119, 263–274. [Google Scholar] [CrossRef]
- Van der Westhuyzen, D.; Böhm, E.; Von Holt, C. Fractionation of chicken erythrocyte whole histone into the six main components by gel exclusion chromatography. Biochim. Et Biophys. Acta (BBA)-Protein Struct. 1974, 359, 341–345. [Google Scholar] [CrossRef]
- Huang, H.-C.; Cole, R.D. Fractionation of chromatin fragments on columns of Biogel A50-m at different salt concentrations. FEBS Lett. 1985, 183, 270–274. [Google Scholar] [CrossRef]
- Takemoto, L.J.; Azari, P. Isolation and characterization of covalently linked, high molecular weight proteins from human cataractous lens. Exp. Eye Res. 1977, 24, 63–70. [Google Scholar] [CrossRef]
- Meehan, B.; Rak, J.; Di Vizio, D. Oncosomes–large and small: What are they, where they came from? J. Extracell. Vesicles 2016, 5, 33109. [Google Scholar] [CrossRef] [PubMed]
- Witwer, K.W.; Théry, C. Extracellular vesicles or exosomes? On primacy, precision, and popularity influencing a choice of nomenclature. J. Extracell. Vesicles 2019, 8, 1648167. [Google Scholar] [CrossRef]
- Foster, B.; Balassa, T.; Benen, T.; Dominovic, M.; Elmadjian, G.; Florova, V.; Fransolet, M.; Kestlerova, A.; Kmiecik, G.; Kostadinova, I. Extracellular vesicles in blood, milk and body fluids of the female and male urogenital tract and with special regard to reproduction. Crit. Rev. Clin. Lab. Sci. 2016, 53, 379–395. [Google Scholar] [CrossRef]
- Kaddour, H.; Lyu, Y.; Welch, J.L.; Paromov, V.; Mandape, S.N.; Sakhare, S.S.; Pandhare, J.; Stapleton, J.T.; Pratap, S.; Dash, C. Proteomics profiling of autologous blood and semen exosomes from HIV-infected and uninfected individuals reveals compositional and functional variabilities. Mol. Cell. Proteom. 2020, 19, 78–100. [Google Scholar] [CrossRef]
- Dixon, C.L.; Sheller-Miller, S.; Saade, G.R.; Fortunato, S.J.; Lai, A.; Palma, C.; Guanzon, D.; Salomon, C.; Menon, R. Amniotic Fluid Exosome Proteomic Profile Exhibits Unique Pathways of Term and Preterm Labor. Endocrinology 2018, 159, 2229–2240. [Google Scholar] [CrossRef] [PubMed]
- Kalluri, R.; LeBleu, V.S. The biology, function, and biomedical applications of exosomes. Science 2020, 367. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.; Mitsialis, S.A.; Aslam, M.; Vitali Sally, H.; Vergadi, E.; Konstantinou, G.; Sdrimas, K.; Fernandez-Gonzalez, A.; Kourembanas, S. Exosomes Mediate the Cytoprotective Action of Mesenchymal Stromal Cells on Hypoxia-Induced Pulmonary Hypertension. Circulation 2012, 126, 2601–2611. [Google Scholar] [CrossRef] [PubMed]
- Hajj, G.N.M.; Arantes, C.P.; Dias, M.V.S.; Roffé, M.; Costa-Silva, B.; Lopes, M.H.; Porto-Carreiro, I.; Rabachini, T.; Lima, F.R.; Beraldo, F.H.; et al. The unconventional secretion of stress-inducible protein 1 by a heterogeneous population of extracellular vesicles. Cell. Mol. Life Sci. 2013, 70, 3211–3227. [Google Scholar] [CrossRef]
- Redzic, J.S.; Kendrick, A.A.; Bahmed, K.; Dahl, K.D.; Pearson, C.G.; Robinson, W.A.; Robinson, S.E.; Graner, M.W.; Eisenmesser, E.Z. Extracellular Vesicles Secreted from Cancer Cell Lines Stimulate Secretion of MMP-9, IL-6, TGF-β1 and EMMPRIN. PLoS ONE 2013, 8, e71225. [Google Scholar] [CrossRef]
- Nordin, J.Z.; Lee, Y.; Vader, P.; Mäger, I.; Johansson, H.J.; Heusermann, W.; Wiklander, O.P.B.; Hällbrink, M.; Seow, Y.; Bultema, J.J.; et al. Ultrafiltration with size-exclusion liquid chromatography for high yield isolation of extracellular vesicles preserving intact biophysical and functional properties. Nanomed. Nanotechnol. Biol. Med. 2015, 11, 879–883. [Google Scholar] [CrossRef]
- Willms, E.; Johansson, H.J.; Mäger, I.; Lee, Y.; Blomberg, K.E.M.; Sadik, M.; Alaarg, A.; Smith, C.I.E.; Lehtiö, J.; El Andaloussi, S.; et al. Cells release subpopulations of exosomes with distinct molecular and biological properties. Sci. Rep. 2016, 6, 22519. [Google Scholar] [CrossRef]
- Mol, E.A.; Goumans, M.-J.; Doevendans, P.A.; Sluijter, J.P.G.; Vader, P. Higher functionality of extracellular vesicles isolated using size-exclusion chromatography compared to ultracentrifugation. Nanomed. Nanotechnol. Biol. Med. 2017, 13, 2061–2065. [Google Scholar] [CrossRef]
- Benedikter, B.J.; Bouwman, F.G.; Vajen, T.; Heinzmann, A.C.A.; Grauls, G.; Mariman, E.C.; Wouters, E.F.M.; Savelkoul, P.H.; Lopez-Iglesias, C.; Koenen, R.R.; et al. Ultrafiltration combined with size exclusion chromatography efficiently isolates extracellular vesicles from cell culture media for compositional and functional studies. Sci. Rep. 2017, 7, 15297. [Google Scholar] [CrossRef] [PubMed]
- Corso, G.; Mäger, I.; Lee, Y.; Görgens, A.; Bultema, J.; Giebel, B.; Wood, M.J.A.; Nordin, J.Z.; Andaloussi, S.E.L. Reproducible and scalable purification of extracellular vesicles using combined bind-elute and size exclusion chromatography. Sci. Rep. 2017, 7, 11561. [Google Scholar] [CrossRef] [PubMed]
- Lobb, R.; Möller, A. Size exclusion chromatography: A simple and reliable method for exosome purification. In Extracellular Vesicles; Springer: Berlin/Heidelberg, Germany, 2017; pp. 105–110. [Google Scholar]
- Taylor, D.D.; Gerçel-Taylor, Ç.; Lyons, K.S.; Stanson, J.; Whiteside, T.L. T-Cell Apoptosis and Suppression of T-Cell Receptor/CD3-ζ by Fas Ligand-Containing Membrane Vesicles Shed from Ovarian Tumors. Clin. Cancer Res. 2003, 9, 5113–5119. [Google Scholar] [PubMed]
- Kim, J.W.; Wieckowski, E.; Taylor, D.D.; Reichert, T.E.; Watkins, S.; Whiteside, T.L. Fas ligand–positive membranous vesicles isolated from sera of patients with oral cancer induce apoptosis of activated T lymphocytes. Clin. Cancer Res. 2005, 11, 1010–1020. [Google Scholar] [PubMed]
- Taylor, D.D.; Zacharias, W.; Gercel-Taylor, C. Exosome Isolation for Proteomic Analyses and RNA Profiling. In Serum/Plasma Proteomics: Methods and Protocols; Simpson, R.J., Greening, D.W., Eds.; Humana Press: Totowa, NJ, USA, 2011; pp. 235–246. [Google Scholar]
- Shu, S.L.; Yang, Y.; Allen, C.L.; Hurley, E.; Tung, K.H.; Minderman, H.; Wu, Y.; Ernstoff, M.S. Purity and yield of melanoma exosomes are dependent on isolation method. J. Extracell. Vesicles 2020, 9, 1692401. [Google Scholar] [CrossRef]
- Li, M.; Lou, D.; Chen, J.; Shi, K.; Wang, Y.; Zhu, Q.; Liu, F.; Zhang, Y. Deep dive on the proteome of salivary extracellular vesicles: Comparison between ultracentrifugation and polymer-based precipitation isolation. Anal. Bioanal. Chem. 2021, 413, 365–375. [Google Scholar] [CrossRef]
- Van Deun, J.; Mestdagh, P.; Sormunen, R.; Cocquyt, V.; Vermaelen, K.; Vandesompele, J.; Bracke, M.; De Wever, O.; Hendrix, A. The impact of disparate isolation methods for extracellular vesicles on downstream RNA profiling. J. Extracell. Vesicles 2014, 3, 24858. [Google Scholar] [CrossRef]
- Karttunen, J.; Heiskanen, M.; Navarro-Ferrandis, V.; Das Gupta, S.; Lipponen, A.; Puhakka, N.; Rilla, K.; Koistinen, A.; Pitkänen, A. Precipitation-based extracellular vesicle isolation from rat plasma co-precipitate vesicle-free microRNAs. J. Extracell. Vesicles 2019, 8, 1555410. [Google Scholar] [CrossRef]
- Wang, T.; Anderson, K.W.; Turko, I.V. Assessment of Extracellular Vesicles Purity Using Proteomic Standards. Anal. Chem. 2017, 89, 11070–11075. [Google Scholar] [CrossRef]
- DDong, L.; Zieren, R.C.; Horie, K.; Kim, C.; Mallick, E.; Jing, Y.; Feng, M.; Kuczler, M.D.; Green, J.; Amend, S.R.; et al. Comprehensive evaluation of methods for small extracellular vesicles separation from human plasma, urine and cell culture medium. J. Extracell. Vesicles 2020, 10, e12044. [Google Scholar] [CrossRef] [PubMed]
- Arroyo, J.D.; Chevillet, J.R.; Kroh, E.M.; Ruf, I.K.; Pritchard, C.C.; Gibson, D.F.; Mitchell, P.S.; Bennett, C.F.; Pogosova-Agadjanyan, E.L.; Stirewalt, D.L.; et al. Argonaute2 complexes carry a population of circulating microRNAs independent of vesicles in human plasma. Proc. Natl. Acad. Sci. USA 2011, 108, 5003. [Google Scholar] [CrossRef]
- Vickers, K.C.; Palmisano, B.T.; Shoucri, B.M.; Shamburek, R.D.; Remaley, A.T. MicroRNAs are transported in plasma and delivered to recipient cells by high-density lipoproteins. Nat. Cell Biol. 2011, 13, 423–433. [Google Scholar] [CrossRef] [PubMed]
- Zeöld, A.; Sándor, G.O.; Kiss, A.; Soós, A.Á.; Tölgyes, T.; Bursics, A.; Szűcs, Á.; Harsányi, L.; Kittel, Á.; Gézsi, A.; et al. Shared extracellular vesicle miRNA profiles of matched ductal pancreatic adenocarcinoma organoids and blood plasma samples show the power of organoid technology. Cell. Mol. Life Sci. 2021, 78, 3005–3020. [Google Scholar] [CrossRef] [PubMed]
- Just, J.; Yan, Y.; Farup, J.; Sieljacks, P.; Sloth, M.; Venø, M.; Gu, T.; de Paoli, F.V.; Nyengaard, J.R.; Bæk, R.; et al. Blood flow-restricted resistance exercise alters the surface profile, miRNA cargo and functional impact of circulating extracellular vesicles. Sci. Rep. 2020, 10, 5835. [Google Scholar] [CrossRef] [PubMed]
- Cheng, L.; Quek, C.; Li, X.; Bellingham, S.A.; Ellett, L.J.; Shambrook, M.; Zafar, S.; Zerr, I.; Lawson, V.A.; Hill, A.F. Distribution of microRNA profiles in pre-clinical and clinical forms of murine and human prion disease. Commun. Biol. 2021, 4, 411. [Google Scholar] [CrossRef] [PubMed]
- Böing, A.N.; van der Pol, E.; Grootemaat, A.E.; Coumans, F.A.W.; Sturk, A.; Nieuwland, R. Single-step isolation of extracellular vesicles by size-exclusion chromatography. J. Extracell. Vesicles 2014, 3, 23430. [Google Scholar] [CrossRef]
- de Menezes-Neto, A.; Sáez, M.J.F.; Lozano-Ramos, I.; Segui-Barber, J.; Martin-Jaular, L.; Ullate, J.M.E.; Fernandez-Becerra, C.; Borrás, F.E.; Del Portillo, H.A. Size-exclusion chromatography as a stand-alone methodology identifies novel markers in mass spectrometry analyses of plasma-derived vesicles from healthy individuals. J. Extracell. Vesicles 2015, 4, 27378. [Google Scholar] [CrossRef]
- Welton, J.L.; Webber, J.P.; Botos, L.-A.; Jones, M.; Clayton, A. Ready-made chromatography columns for extracellular vesicle isolation from plasma. J. Extracell. Vesicles 2015, 4, 27269. [Google Scholar] [CrossRef]
- Baranyai, T.; Herczeg, K.; Onódi, Z.; Voszka, I.; Módos, K.; Marton, N.; Nagy, G.; Mäger, I.; Wood, M.J.; El Andaloussi, S. Isolation of exosomes from blood plasma: Qualitative and quantitative comparison of ultracentrifugation and size exclusion chromatography methods. PLoS ONE 2015, 10, e0145686. [Google Scholar] [CrossRef]
- Brown, B.A.; Zeng, X.; Todd, A.R.; Barnes, L.F.; Winstone, J.M.A.; Trinidad, J.C.; Novotny, M.V.; Jarrold, M.F.; Clemmer, D.E. Charge Detection Mass Spectrometry Measurements of Exosomes and other Extracellular Particles Enriched from Bovine Milk. Anal. Chem. 2020, 92, 3285–3292. [Google Scholar] [CrossRef]
- Hong, C.-S.; Funk, S.; Muller, L.; Boyiadzis, M.; Whiteside, T.L. Isolation of biologically active and morphologically intact exosomes from plasma of patients with cancer. J. Extracell. Vesicles 2016, 5, 29289. [Google Scholar] [CrossRef]
- Gámez-Valero, A.; Monguió-Tortajada, M.; Carreras-Planella, L.; Franquesa, M.l.; Beyer, K.; Borràs, F.E. Size-Exclusion Chromatography-based isolation minimally alters Extracellular Vesicles’ characteristics compared to precipitating agents. Sci. Rep. 2016, 6, 33641. [Google Scholar] [CrossRef] [PubMed]
- Simonsen, J.B. What are we looking at? Extracellular vesicles, lipoproteins, or both? Circ. Res. 2017, 121, 920–922. [Google Scholar] [CrossRef] [PubMed]
- Sluijter, J.P.G.; Davidson, S.M.; Boulanger, C.M.; Buzas, E.I.; De Kleijn, D.P.V.; Engel, F.B.; Giricz, Z.; Hausenloy, D.J.; Kishore, R.; Lecour, S. Extracellular vesicles in diagnostics and therapy of the ischaemic heart: Position Paper from the Working Group on Cellular Biology of the Heart of the European Society of Cardiology. Cardiovasc. Res. 2018, 114, 19–34. [Google Scholar] [CrossRef]
- Ridger, V.C.; Boulanger, C.M.; Angelillo-Scherrer, A.; Badimon, L.; Blanc-Brude, O.; Bochaton-Piallat, M.-L.; Boilard, E.; Buzas, E.I.; Caporali, A.; Dignat-George, F. Microvesicles in vascular homeostasis and diseases. Position paper of the European Society of Cardiology (ESC) working group on atherosclerosis and vascular biology. Thromb. Haemost. 2017, 117, 1296–1316. [Google Scholar] [PubMed]
- Karimi, N.; Cvjetkovic, A.; Jang, S.C.; Crescitelli, R.; Hosseinpour Feizi, M.A.; Nieuwland, R.; Lötvall, J.; Lässer, C. Detailed analysis of the plasma extracellular vesicle proteome after separation from lipoproteins. Cell. Mol. Life Sci. 2018, 75, 2873–2886. [Google Scholar] [CrossRef]
- Buschmann, D.; Kirchner, B.; Hermann, S.; Märte, M.; Wurmser, C.; Brandes, F.; Kotschote, S.; Bonin, M.; Steinlein, O.K.; Pfaffl, M.W.; et al. Evaluation of serum extracellular vesicle isolation methods for profiling miRNAs by next-generation sequencing. J. Extracell. Vesicles 2018, 7, 1481321. [Google Scholar] [CrossRef]
- Takov, K.; Yellon, D.M.; Davidson, S.M. Comparison of small extracellular vesicles isolated from plasma by ultracentrifugation or size-exclusion chromatography: Yield, purity and functional potential. J. Extracell. Vesicles 2019, 8, 1560809. [Google Scholar] [CrossRef]
- Ronquist, G. Prostasomes: Their Characterisation: Implications for Human Reproduction. In The Male Role in Pregnancy Loss and Embryo Implantation Failure; Bronson, R., Ed.; Springer International Publishing: Cham, Switzerland, 2015; pp. 191–209. [Google Scholar]
- Ronquist, G. Prostasomes are mediators of intercellular communication: From basic research to clinical implications. J. Intern. Med. 2012, 271, 400–413. [Google Scholar] [CrossRef]
- Ronquist, G.; Brody, I. The prostasome: Its secretion and function in man. Biochim. Biophys. Acta 1985, 822, 203–218. [Google Scholar] [CrossRef]
- Peeker, R.; Abramsson, L.; Marklund, S.L. Superoxide dismutase isoenzymes in human seminal plasma and spermatozoa. Mol. Hum. Reprod. 1997, 3, 1061–1066. [Google Scholar] [CrossRef]
- Saez, F.; Motta, C.; Boucher, D.; Grizard, G. Antioxidant capacity of prostasomes in human semen. Mol. Hum. Reprod. 1998, 4, 667–672. [Google Scholar] [CrossRef]
- Jones, J.L.; Saraswati, S.; Block, A.S.; Lichti, C.F.; Mahadevan, M.; Diekman, A.B. Galectin-3 is associated with prostasomes in human semen. Glycoconj. J. 2010, 27, 227–236. [Google Scholar] [CrossRef]
- Aalberts, M.; van Dissel-Emiliani, F.M.F.; van Adrichem, N.P.H.; van Wijnen, M.; Wauben, M.H.M.; Stout, T.A.E.; Stoorvogel, W. Identification of Distinct Populations of Prostasomes That Differentially Express Prostate Stem Cell Antigen, Annexin A1, and GLIPR2 in Humans1. Biol. Reprod. 2012, 86, 82. [Google Scholar] [CrossRef] [PubMed]
- Brouwers, J.F.; Aalberts, M.; Jansen, J.W.A.; van Niel, G.; Wauben, M.H.; Stout, T.A.E.; Helms, J.B.; Stoorvogel, W. Distinct lipid compositions of two types of human prostasomes. Proteomics 2013, 13, 1660–1666. [Google Scholar] [CrossRef] [PubMed]
- Lyu, Y.; Kaddour, H.; Kopcho, S.; Panzner, T.D.; Shouman, N.; Kim, E.-Y.; Martinson, J.; McKay, H.; Martinez-Maza, O.; Margolick, J.B.; et al. Human Immunodeficiency Virus (HIV) Infection and Use of Illicit Substances Promote Secretion of Semen Exosomes that Enhance Monocyte Adhesion and Induce Actin Reorganization and Chemotactic Migration. Cells 2019, 8, 1027. [Google Scholar] [CrossRef]
- Kaddour, H.; Lyu, Y.; Shouman, N.; Mohan, M.; Okeoma, C.M. Development of Novel High-Resolution Size-Guided Turbidimetry-Enabled Particle Purification Liquid Chromatography (PPLC): Extracellular Vesicles and Membraneless Condensates in Focus. Int. J. Mol. Sci. 2020, 21, 5361. [Google Scholar] [CrossRef] [PubMed]
- Welch, J.L.; Kaddour, H.; Winchester, L.; Fletcher, C.V.; Stapleton, J.T.; Okeoma, C.M. Semen Extracellular Vesicles From HIV-1–Infected Individuals Inhibit HIV-1 Replication In Vitro, and Extracellular Vesicles Carry Antiretroviral Drugs In Vivo. JAIDS J. Acquir. Immune Defic. Syndr. 2020, 83, 90–98. [Google Scholar] [CrossRef] [PubMed]
- Kaddour, H.; Panzner, T.D.; Welch, J.L.; Shouman, N.; Mohan, M.; Stapleton, J.T.; Okeoma, C.M. Electrostatic Surface Properties of Blood and Semen Extracellular Vesicles: Implications of Sialylation and HIV-Induced Changes on EV Internalization. Viruses 2020, 12, 1117. [Google Scholar] [CrossRef]
- Welch, J.L.; Stapleton, J.T.; Okeoma, C.M. Vehicles of intercellular communication: Exosomes and HIV-1. J. Gen. Virol. 2019, 100, 350–366. [Google Scholar] [CrossRef]
- Welch, J.L.; Kaddour, H.; Schlievert, P.M.; Stapleton, J.T.; Okeoma, C.M. Semen exosomes promote transcriptional silencing of HIV-1 by disrupting NF-κB/Sp1/Tat circuitry. J. Virol. 2018, 92, e00731-18. [Google Scholar] [CrossRef] [PubMed]
- Sun, I.O.; Lerman, L.O. Urinary Extracellular Vesicles as Biomarkers of Kidney Disease: From Diagnostics to Therapeutics. Diagnostics 2020, 10, 311. [Google Scholar] [CrossRef] [PubMed]
- Erdbrügger, U.; Le, T.H. Extracellular vesicles in renal diseases: More than novel biomarkers? J. Am. Soc. Nephrol. 2016, 27, 12–26. [Google Scholar] [CrossRef] [PubMed]
- Lindblom, H.; Söderberg, L.; Cooper, E.H.; Turner, R. Urinary protein isolation by high-performance ion exchange chromatography. J. Chromatogr. A 1983, 266, 187–196. [Google Scholar] [CrossRef]
- Rood, I.M.; Deegens, J.K.; Merchant, M.L.; Tamboer, W.P.; Wilkey, D.W.; Wetzels, J.F.; Klein, J.B. Comparison of three methods for isolation of urinary microvesicles to identify biomarkers of nephrotic syndrome. Kidney Int. 2010, 78, 810–816. [Google Scholar] [CrossRef]
- Lozano-Ramos, I.; Bancu, I.; Oliveira-Tercero, A.; Armengol, M.P.; Menezes-Neto, A.; Portillo, H.A.D.; Lauzurica-Valdemoros, R.; Borràs, F.E. Size-exclusion chromatography-based enrichment of extracellular vesicles from urine samples. J. Extracell. Vesicles 2015, 4, 27369. [Google Scholar] [CrossRef]
- Lozano-Ramos, S.I.; Bancu, I.; Carreras-Planella, L.; Monguió-Tortajada, M.; Cañas, L.; Juega, J.; Bonet, J.; Armengol, M.P.; Lauzurica, R.; Borràs, F.E. Molecular profile of urine extracellular vesicles from normo-functional kidneys reveal minimal differences between living and deceased donors. BMC Nephrol. 2018, 19, 189. [Google Scholar] [CrossRef]
- Ogawa, Y.; Kanai-Azuma, M.; Akimoto, Y.; Kawakami, H.; Yanoshita, R. Exosome-like vesicles with dipeptidyl peptidase IV in human saliva. Biol. Pharm. Bull. 2008, 31, 1059–1062. [Google Scholar] [CrossRef]
- Ogawa, Y.; Miura, Y.; Harazono, A.; Kanai-Azuma, M.; Akimoto, Y.; Kawakami, H.; Yamaguchi, T.; Toda, T.; Endo, T.; Tsubuki, M. Proteomic analysis of two types of exosomes in human whole saliva. Biol. Pharm. Bull. 2011, 34, 13–23. [Google Scholar] [CrossRef]
- Aqrawi, L.A.; Galtung, H.K.; Vestad, B.; Øvstebø, R.; Thiede, B.; Rusthen, S.; Young, A.; Guerreiro, E.M.; Utheim, T.P.; Chen, X. Identification of potential saliva and tear biomarkers in primary Sjögren’s syndrome, utilising the extraction of extracellular vesicles and proteomics analysis. Arthritis Res. Ther. 2017, 19, 14. [Google Scholar] [CrossRef]
- Blans, K.; Hansen, M.S.; Sørensen, L.V.; Hvam, M.L.; Howard, K.A.; Möller, A.; Wiking, L.; Larsen, L.B.; Rasmussen, J.T. Pellet-free isolation of human and bovine milk extracellular vesicles by size-exclusion chromatography. J. Extracell. Vesicles 2017, 6, 1294340. [Google Scholar] [CrossRef] [PubMed]
- Tietje, A.; Maron, K.N.; Wei, Y.; Feliciano, D.M. Cerebrospinal Fluid Extracellular Vesicles Undergo Age Dependent Declines and Contain Known and Novel Non-coding RNAs. PLoS ONE 2014, 9, e113116. [Google Scholar] [CrossRef]
- Carreras-Planella, L.; Soler-Majoral, J.; Rubio-Esteve, C.; Lozano-Ramos, S.I.; Franquesa, M.; Bonet, J.; Troya-Saborido, M.I.; Borràs, F.E. Characterization and proteomic profile of extracellular vesicles from peritoneal dialysis efflux. PLoS ONE 2017, 12, e0176987. [Google Scholar] [CrossRef] [PubMed]
- Foers, A.D.; Chatfield, S.; Dagley, L.F.; Scicluna, B.J.; Webb, A.I.; Cheng, L.; Hill, A.F.; Wicks, I.P.; Pang, K.C. Enrichment of extracellular vesicles from human synovial fluid using size exclusion chromatography. J. Extracell. Vesicles 2018, 7, 1490145. [Google Scholar] [CrossRef] [PubMed]
- Sidhom, K.; Obi, P.O.; Saleem, A. A Review of Exosomal Isolation Methods: Is Size Exclusion Chromatography the Best Option? Int. J. Mol. Sci. 2020, 21, 6466. [Google Scholar] [CrossRef]
- Van Deun, J.; Jo, A.; Li, H.; Lin, H.Y.; Weissleder, R.; Im, H.; Lee, H. Integrated Dual-Mode Chromatography to Enrich Extracellular Vesicles from Plasma. Adv. Biosyst. 2020, 1900310. [Google Scholar] [CrossRef]
- Audhya, T.; Reddy, J.; Zaneveld, L.J.D. Purification and Partial Chemical Characterization of a Glycoprotein with Antifertility Activity from Human Seminal Plasma1. Biol. Reprod. 1987, 36, 511–521. [Google Scholar] [CrossRef]
- Winzor, D.J. Analytical exclusion chromatography. J. Biochem. Biophys. Methods 2003, 56, 15–52. [Google Scholar] [CrossRef]
- Elsayed, M.M.; Cevc, G. Turbidity spectroscopy for characterization of submicroscopic drug carriers, such as nanoparticles and lipid vesicles: Size determination. Pharm. Res. 2011, 28, 2204–2222. [Google Scholar] [CrossRef]
- Wang, A.; Miller, C.C.; Szostak, J.W. Core-Shell Modeling of Light Scattering by Vesicles: Effect of Size, Contents, and Lamellarity. Biophys. J. 2019, 116, 659–669. [Google Scholar] [CrossRef]
- Wang, A.; Garmann, R.F.; Manoharan, V.N. Tracking E. coli runs and tumbles with scattering solutions and digital holographic microscopy. Opt. Express 2016, 24, 23719–23725. [Google Scholar] [CrossRef]
- Varga, Z.; Yuana, Y.; Grootemaat, A.E.; Van der Pol, E.; Gollwitzer, C.; Krumrey, M.; Nieuwland, R. Towards traceable size determination of extracellular vesicles. J. Extracell. Vesicles 2014, 3, 23298. [Google Scholar] [CrossRef] [PubMed]
- Wyss, R.; Grasso, L.; Wolf, C.; Grosse, W.; Demurtas, D.; Vogel, H. Molecular and dimensional profiling of highly purified extracellular vesicles by fluorescence fluctuation spectroscopy. Anal. Chem. 2014, 86, 7229–7233. [Google Scholar] [CrossRef] [PubMed]
- Tian, Y.; Gong, M.; Hu, Y.; Liu, H.; Zhang, W.; Zhang, M.; Hu, X.; Aubert, D.; Zhu, S.; Wu, L.; et al. Quality and efficiency assessment of six extracellular vesicle isolation methods by nano-flow cytometry. J. Extracell. Vesicles 2020, 9, 1697028. [Google Scholar] [CrossRef] [PubMed]
- Sultana, H.; Sultana, H.; Neelakanta, G.; Neelakanta, G.; Sultana, H.; Sultana, H.; Neelakanta, G.; Neelakanta, G. Arthropod exosomes as bubbles with message(s) to transmit vector-borne diseases. Curr. Opin. Insect Sci. 2020, 40, 39–47. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).






