Jumping Ahead with Sleeping Beauty: Mechanistic Insights into Cut-and-Paste Transposition
Abstract
1. Introduction
2. Structural Features of the Sleeping Beauty Transposon System
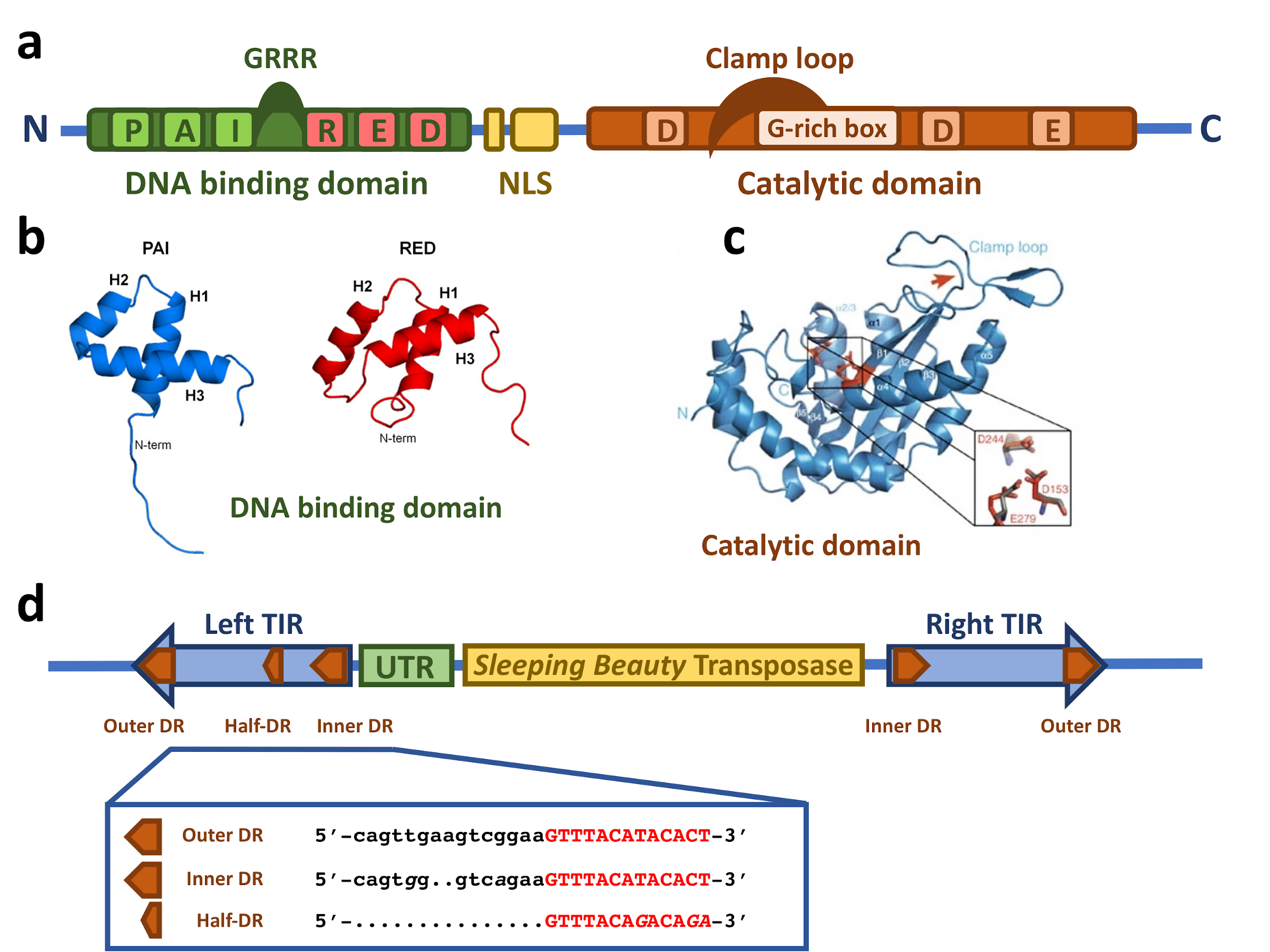
2.1. The Sleeping Beauty Transposase
2.2. The Sleeping Beauty Transposable Element
3. The Mechanism of Sleeping Beauty Transposition
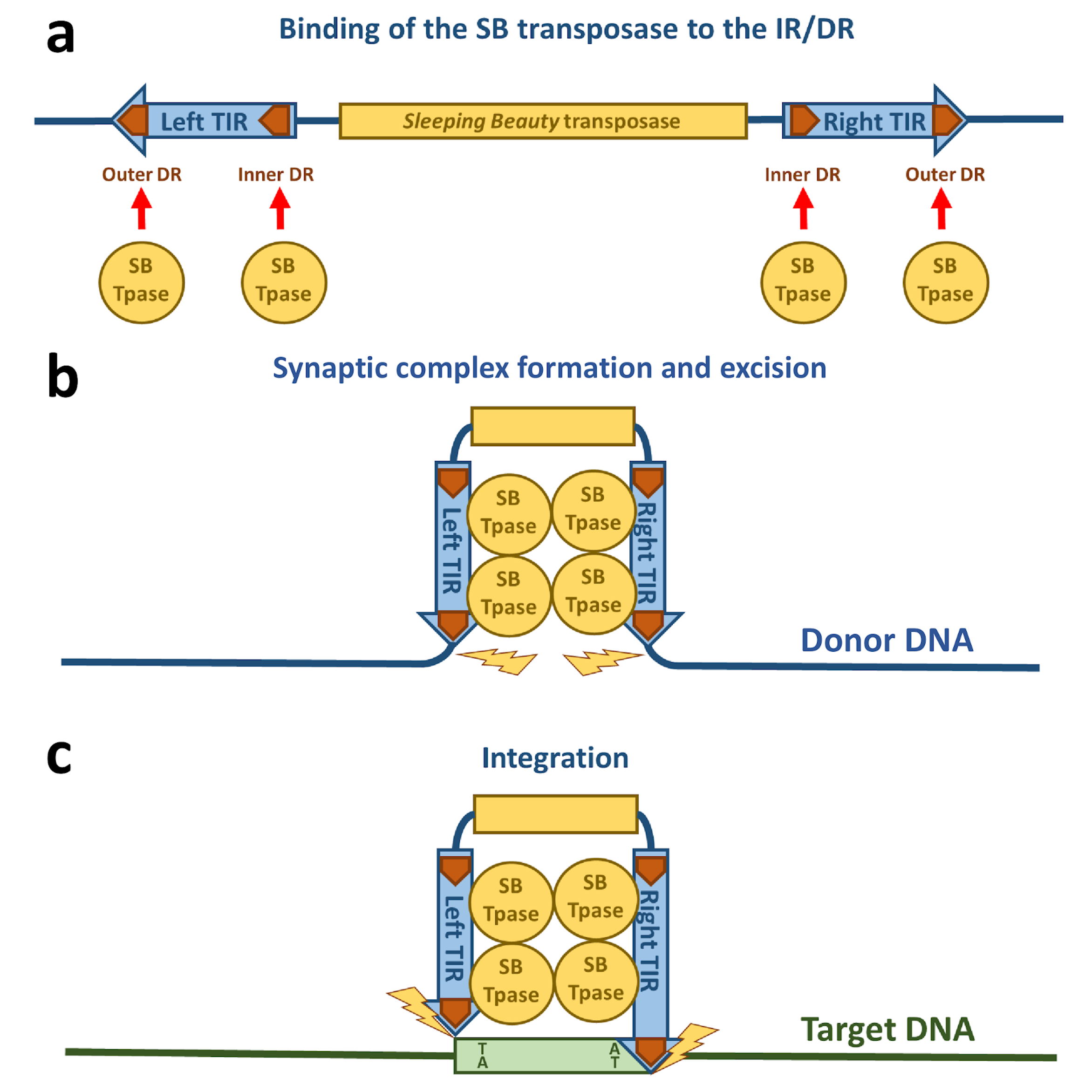
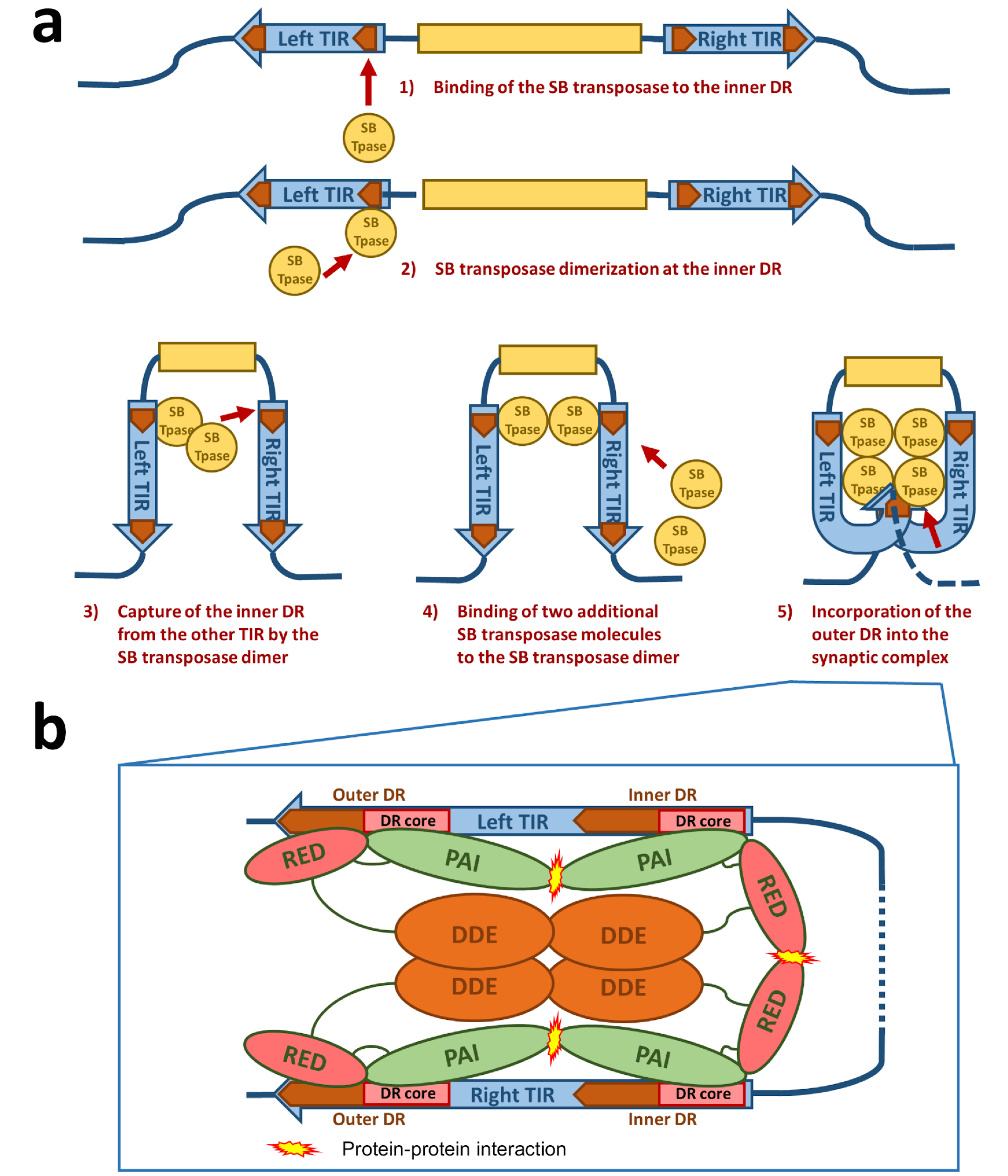
3.1. DNA Binding of the Sleeping Beauty Transposase
3.2. Synaptic Complex Formation
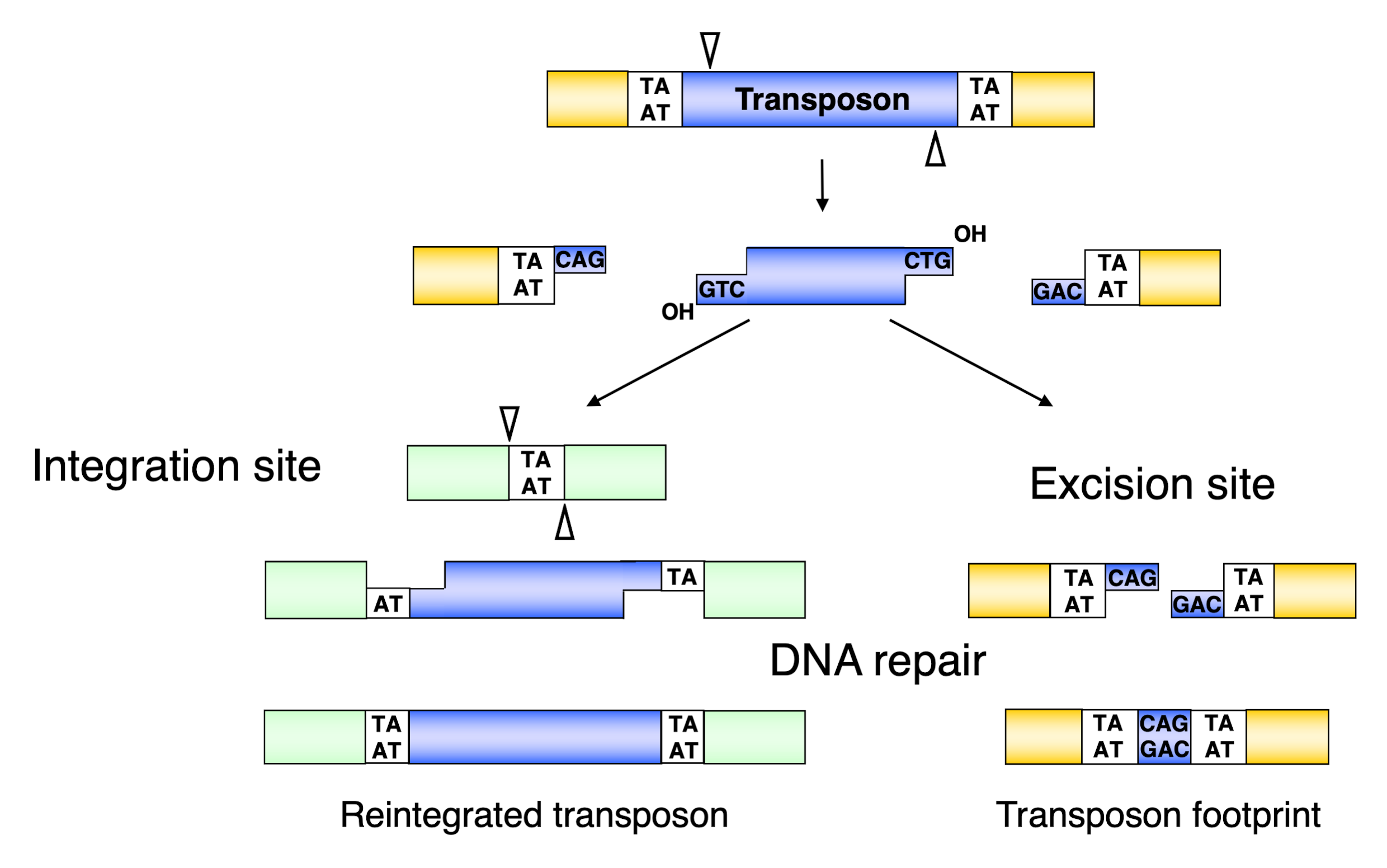
3.3. Excision of the Sleeping Beauty Transposon
3.4. Integration of the Sleeping Beauty Transposon
4. New Sleeping Beauty Variants Offering New Possibilities for Genetic Engineering
4.1. Hyperactive Sleeping Beauty Transposase Variants
4.2. New Vector Platforms for Sleeping Beauty Transposition
4.3. Integration-Deficient Sleeping Beauty Transposase Variant
4.4. Targeted Sleeping Beauty Transposition
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Seelamgari, A.; Maddukuri, A.; Berro, R.; de La Fuente, C.; Kehn, K.; Deng, L.; Dadgar, S.; Bottazzi, M.E.; Ghedin, E.; Pumfery, A.; et al. Role of viral regulatory and accessory proteins in HIV-1 replication. Front. Biosci. 2004, 9, 2388–2413. [Google Scholar] [CrossRef] [PubMed]
- Frankel, A.D.; Young, J.A. HIV-1: Fifteen proteins and an RNA. Annu. Rev. Biochem. 1998, 67, 1–25. [Google Scholar] [CrossRef] [PubMed]
- Bannert, N.; Kurth, R. The evolutionary dynamics of human endogenous retroviral families. Annu. Rev. Genom. Hum. Genet. 2006, 7, 149–173. [Google Scholar] [CrossRef]
- Jasin, M.; Rothstein, R. Repair of strand breaks by homologous recombination. Cold Spring Harb. Perspect. Biol. 2013, 5, a012740. [Google Scholar] [CrossRef] [PubMed]
- McClintock, B. Induction of Instability at Selected Loci in Maize. Genetics 1953, 38, 579–599. [Google Scholar] [PubMed]
- Wicker, T.; Sabot, F.; Hua-Van, A.; Bennetzen, J.L.; Capy, P.; Chalhoub, B.; Flavell, A.; Leroy, P.; Morgante, M.; Panaud, O.; et al. A unified classification system for eukaryotic transposable elements. Nat. Rev. Genet. 2007, 8, 973–982. [Google Scholar] [CrossRef] [PubMed]
- Beck, C.R.; Garcia-Perez, J.L.; Badge, R.M.; Moran, J.V. LINE-1 elements in structural variation and disease. Annu. Rev. Genomics Hum. Genet. 2011, 12, 187–215. [Google Scholar] [CrossRef]
- Deininger, P. Alu elements: Know the SINEs. Genome Biol. 2011, 12, 236. [Google Scholar] [CrossRef]
- Kapitonov, V.V.; Jurka, J. RAG1 core and V(D)J recombination signal sequences were derived from Transib transposons. PLoS Biol. 2005, 3, e181. [Google Scholar] [CrossRef]
- Kapitonov, V.V.; Jurka, J. Rolling-circle transposons in eukaryotes. Proc. Natl. Acad. Sci. USA 2001, 98, 8714–8719. [Google Scholar] [CrossRef]
- Sarkar, A.; Sim, C.; Hong, Y.S.; Hogan, J.R.; Fraser, M.J.; Robertson, H.M.; Collins, F.H. Molecular evolutionary analysis of the widespread piggyBac transposon family and related “domesticated” sequences. Mol. Genet. Genom. 2003, 270, 173–180. [Google Scholar] [CrossRef] [PubMed]
- Jurka, J.; Kapitonov, V.V. PIFs meet Tourists and Harbingers: A superfamily reunion. Proc. Natl. Acad. Sci. USA 2001, 98, 12315–12316. [Google Scholar] [CrossRef] [PubMed]
- Shao, H.; Tu, Z. Expanding the diversity of the IS630-Tc1-mariner superfamily: Discovery of a unique DD37E transposon and reclassification of the DD37D and DD39D transposons. Genetics 2001, 159, 1103–1115. [Google Scholar] [PubMed]
- Ivics, Z.; Hackett, P.B.; Plasterk, R.H.; Izsvák, Z. Molecular Reconstruction of Sleeping Beauty, a Tc1-like Transposon from Fish, and Its Transposition in Human Cells. Cell 1997, 91, 501–510. [Google Scholar] [CrossRef]
- Amberger, M.; Ivics, Z. Latest Advances for the Sleeping Beauty Transposon System: 23 Years of Insomnia but Prettier than Ever: Refinement and Recent Innovations of the Sleeping Beauty Transposon System Enabling Novel, Nonviral Genetic Engineering Applications. Bioessays 2020, 42, e2000136. [Google Scholar] [CrossRef] [PubMed]
- Izsvák, Z.; Khare, D.; Behlke, J.; Heinemann, U.; Plasterk, R.H.; Ivics, Z. Involvement of a bifunctional, paired-like DNA-binding domain and a transpositional enhancer in Sleeping Beauty transposition. J. Biol. Chem. 2002, 277, 34581–34588. [Google Scholar] [CrossRef]
- Pabo, C.O.; Sauer, R.T. Transcription factors: Structural families and principles of DNA recognition. Annu. Rev. Biochem. 1992, 61, 1053–1095. [Google Scholar] [CrossRef]
- Czerny, T.; Schaffner, G.; Busslinger, M. DNA sequence recognition by Pax proteins: Bipartite structure of the paired domain and its binding site. Genes Dev. 1993, 7, 2048–2061. [Google Scholar] [CrossRef]
- Brennan, R.G.; Matthews, B.W. The helix-turn-helix DNA binding motif. J. Biol. Chem. 1989, 264, 1903–1906. [Google Scholar] [CrossRef]
- Aravind, L.; Anantharaman, V.; Balaji, S.; Babu, M.M.; Iyer, L.M. The many faces of the helix-turn-helix domain: Transcription regulation and beyond. FEMS Microbiol. Rev. 2005, 29, 231–262. [Google Scholar] [CrossRef]
- Konnova, T.A.; Singer, C.M.; Nesmelova, I.V. NMR solution structure of the RED subdomain of the Sleeping Beauty transposase. Protein Sci. 2017, 26, 1171–1181. [Google Scholar] [CrossRef] [PubMed]
- Carpentier, C.E.; Schreifels, J.M.; Aronovich, E.L.; Carlson, D.F.; Hackett, P.B.; Nesmelova, I.V. NMR structural analysis of Sleeping Beauty transposase binding to DNA. Protein Sci. 2014, 23, 23–33. [Google Scholar] [CrossRef] [PubMed]
- Voigt, F.; Wiedemann, L.; Zuliani, C.; Querques, I.; Sebe, A.; Mátés, L.; Izsvák, Z.; Ivics, Z.; Barabas, O. Sleeping Beauty transposase structure allows rational design of hyperactive variants for genetic engineering. Nat. Commun. 2016, 7, 11126. [Google Scholar] [CrossRef] [PubMed]
- Hickman, A.B.; Chandler, M.; Dyda, F. Integrating prokaryotes and eukaryotes: DNA transposases in light of structure. Crit. Rev. Biochem. Mol. Biol. 2010, 45, 50–69. [Google Scholar] [CrossRef]
- Rice, P.A.; Baker, T.A. Comparative architecture of transposase and integrase complexes. Nat. Struct Biol. 2001, 8, 302–307. [Google Scholar] [CrossRef]
- Montaño, S.P.; Rice, P.A. Moving DNA around: DNA transposition and retroviral integration. Curr. Opin. Struct. Biol. 2011, 21, 370–378. [Google Scholar] [CrossRef]
- Yang, W.; Lee, J.Y.; Nowotny, M. Making and breaking nucleic acids: Two-Mg2+-ion catalysis and substrate specificity. Mol. Cell 2006, 22, 5–13. [Google Scholar] [CrossRef]
- Cui, Z.; Geurts, A.M.; Liu, G.; Kaufman, C.D.; Hackett, P.B. Structure–Function Analysis of the Inverted Terminal Repeats of the Sleeping Beauty Transposon. J. Mol. Biol. 2002, 318, 1221–1235. [Google Scholar] [CrossRef]
- Liu, G.; Aronovich, E.L.; Cui, Z.; Whitley, C.B.; Hackett, P.B. Excision of Sleeping Beauty transposons: Parameters and applications to gene therapy. J. Gene Med. 2004, 6, 574–583. [Google Scholar] [CrossRef]
- Zayed, H.; Izsvák, Z.; Walisko, O.; Ivics, Z. Development of hyperactive sleeping beauty transposon vectors by mutational analysis. Mol. Ther. 2004, 9, 292–304. [Google Scholar] [CrossRef]
- Geurts, A.M.; Yang, Y.; Clark, K.J.; Liu, G.; Cui, Z.; Dupuy, A.J.; Bell, J.B.; Largaespada, D.A.; Hackett, P.B. Gene transfer into genomes of human cells by the sleeping beauty transposon system. Mol. Ther. 2003, 8, 108–117. [Google Scholar] [CrossRef]
- Walisko, O.; Schorn, A.; Rolfs, F.; Devaraj, A.; Miskey, C.; Izsvák, Z.; Ivics, Z. Transcriptional activities of the Sleeping Beauty transposon and shielding its genetic cargo with insulators. Mol. Ther. 2008, 16, 359–369. [Google Scholar] [CrossRef] [PubMed]
- Moldt, B.; Yant, S.R.; Andersen, P.R.; Kay, M.A.; Mikkelsen, J.G. Cis-acting gene regulatory activities in the terminal regions of sleeping beauty DNA transposon-based vectors. Hum. Gene Ther. 2007, 18, 1193–1204. [Google Scholar] [CrossRef] [PubMed]
- Izsvák, Z.; Ivics, Z.; Plasterk, R.H. Sleeping Beauty, a wide host-range transposon vector for genetic transformation in vertebrates. J. Mol. Biol. 2000, 302, 93–102. [Google Scholar] [CrossRef] [PubMed]
- Zayed, H.; Izsvák, Z.; Khare, D.; Heinemann, U.; Ivics, Z. The DNA-bending protein HMGB1 is a cellular cofactor of Sleeping Beauty transposition. Nucleic Acids Res. 2003, 31, 2313–2322. [Google Scholar] [CrossRef]
- Wang, Y.; Pryputniewicz-Dobrinska, D.; Nagy, E.É.; Kaufman, C.D.; Singh, M.; Yant, S.; Wang, J.; Dalda, A.; Kay, M.A.; Ivics, Z.; et al. Regulated complex assembly safeguards the fidelity of Sleeping Beauty transposition. Nucleic Acids Res. 2017, 45, 311–326. [Google Scholar] [CrossRef]
- Richardson, J.M.; Colloms, S.D.; Finnegan, D.J.; Walkinshaw, M.D. Molecular architecture of the Mos1 paired-end complex: The structural basis of DNA transposition in a eukaryote. Cell 2009, 138, 1096–1108. [Google Scholar] [CrossRef]
- Watkins, S.; van Pouderoyen, G.; Sixma, T.K. Structural analysis of the bipartite DNA-binding domain of Tc3 transposase bound to transposon DNA. Nucleic Acids Res. 2004, 32, 4306–4312. [Google Scholar] [CrossRef]
- Bouuaert, C.C.; Liu, D.; Chalmers, R. A simple topological filter in a eukaryotic transposon as a mechanism to suppress genome instability. Mol. Cell. Biol. 2011, 31, 317–327. [Google Scholar] [CrossRef]
- Wang, Y.; Wang, J.; Devaraj, A.; Singh, M.; Jimenez Orgaz, A.; Chen, J.-X.; Selbach, M.; Ivics, Z.; Izsvák, Z. Suicidal autointegration of sleeping beauty and piggyBac transposons in eukaryotic cells. PLoS Genet. 2014, 10, e1004103. [Google Scholar] [CrossRef]
- West, R.B.; Lieber, M.R. The RAG-HMG1 complex enforces the 12/23 rule of V(D)J recombination specifically at the double-hairpin formation step. Mol. Cell. Biol. 1998, 18, 6408–6415. [Google Scholar] [CrossRef][Green Version]
- Van Gent, D.C.; Hiom, K.; Paull, T.T.; Gellert, M. Stimulation of V(D)J cleavage by high mobility group proteins. EMBO J. 1997, 16, 2665–2670. [Google Scholar] [CrossRef]
- Agrawal, A.; Eastman, Q.M.; Schatz, D.G. Transposition mediated by RAG1 and RAG2 and its implications for the evolution of the immune system. Nature 1998, 394, 744–751. [Google Scholar] [CrossRef]
- Jones, J.M.; Gellert, M. Ordered assembly of the V(D)J synaptic complex ensures accurate recombination. EMBO J. 2002, 21, 4162–4171. [Google Scholar] [CrossRef] [PubMed]
- Mizuuchi, K. Polynucleotidyl transfer reactions in transpositional DNA recombination. J. Biol. Chem. 1992, 267, 21273–21276. [Google Scholar] [CrossRef]
- Craig, N.L. Unity in transposition reactions. Science 1995, 270, 253–254. [Google Scholar] [CrossRef] [PubMed]
- Hickman, A.B.; Dyda, F. Mechanisms of DNA Transposition. Microbiol. Spectr. 2015, 3, 531–553. [Google Scholar] [CrossRef]
- Hencken, C.G.; Li, X.; Craig, N.L. Functional characterization of an active Rag-like transposase. Nat. Struct. Mol. Biol. 2012, 19, 834–836. [Google Scholar] [CrossRef] [PubMed]
- Zhou, L.; Mitra, R.; Atkinson, P.W.; Hickman, A.B.; Dyda, F.; Craig, N.L. Transposition of hAT elements links transposable elements and V(D)J recombination. Nature 2004, 432, 995–1001. [Google Scholar] [CrossRef]
- Mitra, R.; Fain-Thornton, J.; Craig, N.L. piggyBac can bypass DNA synthesis during cut and paste transposition. EMBO J. 2008, 27, 1097–1109. [Google Scholar] [CrossRef]
- Bischerour, J.; Chalmers, R. Base flipping in tn10 transposition: An active flip and capture mechanism. PLoS ONE 2009, 4, e6201. [Google Scholar] [CrossRef] [PubMed]
- Bischerour, J.; Lu, C.; Roth, D.B.; Chalmers, R. Base flipping in V(D)J recombination: Insights into the mechanism of hairpin formation, the 12/23 rule, and the coordination of double-strand breaks. Mol. Cell. Biol. 2009, 29, 5889–5899. [Google Scholar] [CrossRef] [PubMed]
- Richardson, J.M.; Dawson, A.; O’Hagan, N.; Taylor, P.; Finnegan, D.J.; Walkinshaw, M.D. Mechanism of Mos1 transposition: Insights from structural analysis. EMBO J. 2006, 25, 1324–1334. [Google Scholar] [CrossRef] [PubMed]
- Izsvák, Z.; Stüwe, E.E.; Fiedler, D.; Katzer, A.; Jeggo, P.A.; Ivics, Z. Healing the Wounds Inflicted by Sleeping Beauty Transposition by Double-Strand Break Repair in Mammalian Somatic Cells. Mol. Cell 2004, 13, 279–290. [Google Scholar] [CrossRef]
- Claeys Bouuaert, C.; Chalmers, R. A single active site in the mariner transposase cleaves DNA strands of opposite polarity. Nucleic Acids Res. 2017, 45, 11467–11478. [Google Scholar] [CrossRef]
- Dawson, A.; Finnegan, D.J. Excision of the Drosophila Mariner Transposon Mos1. Mol. Cell 2003, 11, 225–235. [Google Scholar] [CrossRef]
- Miskey, C.; Papp, B.; Mátés, L.; Sinzelle, L.; Keller, H.; Izsvák, Z.; Ivics, Z. The ancient mariner sails again: Transposition of the human Hsmar1 element by a reconstructed transposase and activities of the SETMAR protein on transposon ends. Mol. Cell. Biol. 2007, 27, 4589–4600. [Google Scholar] [CrossRef]
- Lampe, D.J.; Churchill, M.E.; Robertson, H.M. A purified mariner transposase is sufficient to mediate transposition in vitro. EMBO J. 1996, 15, 5470–5479. [Google Scholar] [CrossRef]
- Lohe, A.R.; Timmons, C.; Beerman, I.; Lozovskaya, E.R.; Hartl, D.L. Self-inflicted wounds, template-directed gap repair and a recombination hotspot. Effects of the mariner transposase. Genetics 2000, 154, 647–656. [Google Scholar]
- Engels, W.R.; Johnson-Schlitz, D.M.; Eggleston, W.B.; Sved, J. High-frequency P element loss in Drosophila is homolog dependent. Cell 1990, 62, 515–525. [Google Scholar] [CrossRef]
- Luo, G.; Ivics, Z.; Izsvák, Z.; Bradley, A. Chromosomal transposition of a Tc1/mariner-like element in mouse embryonic stem cells. Proc. Natl. Acad. Sci. USA 1998, 95, 10769–10773. [Google Scholar] [CrossRef] [PubMed]
- Daniel, R.; Katz, R.A.; Skalka, A.M. A role for DNA-PK in retroviral DNA integration. Science 1999, 284, 644–647. [Google Scholar] [CrossRef] [PubMed]
- Jackson, S.P.; Jeggo, P.A. DNA double-strand break repair and V(D)J recombination: Involvement of DNA-PK. Trends Biochem. Sci. 1995, 20, 412–415. [Google Scholar] [CrossRef]
- Miskey, C.; Izsvák, Z.; Kawakami, K.; Ivics, Z. DNA transposons in vertebrate functional genomics. Cell. Mol. Life Sci. 2005, 62, 629–641. [Google Scholar] [CrossRef] [PubMed]
- Yoder, J.A.; Walsh, C.P.; Bestor, T.H. Cytosine methylation and the ecology of intragenomic parasites. Trends Genet. 1997, 13, 335–340. [Google Scholar] [CrossRef]
- Yusa, K.; Takeda, J.; Horie, K. Enhancement of Sleeping Beauty transposition by CpG methylation: Possible role of heterochromatin formation. Mol. Cell. Biol. 2004, 24, 4004–4018. [Google Scholar] [CrossRef]
- Jursch, T.; Miskey, C.; Izsvák, Z.; Ivics, Z. Regulation of DNA transposition by CpG methylation and chromatin structure in human cells. Mob. DNA 2013, 4, 15. [Google Scholar] [CrossRef]
- Li, X.; Ewis, H.; Hice, R.H.; Malani, N.; Parker, N.; Zhou, L.; Feschotte, C.; Bushman, F.D.; Atkinson, P.W.; Craig, N.L. A resurrected mammalian hAT transposable element and a closely related insect element are highly active in human cell culture. Proc. Natl. Acad. Sci. USA 2013, 110, E478–E487. [Google Scholar] [CrossRef]
- Liu, G.; Geurts, A.M.; Yae, K.; Srinivasan, A.R.; Fahrenkrug, S.C.; Largaespada, D.A.; Takeda, J.; Horie, K.; Olson, W.K.; Hackett, P.B. Target-site preferences of Sleeping Beauty transposons. J. Mol. Biol. 2005, 346, 161–173. [Google Scholar] [CrossRef]
- Voigt, K.; Gogol-Döring, A.; Miskey, C.; Chen, W.; Cathomen, T.; Izsvák, Z.; Ivics, Z. Retargeting sleeping beauty transposon insertions by engineered zinc finger DNA-binding domains. Mol. Ther. 2012, 20, 1852–1862. [Google Scholar] [CrossRef]
- Moldt, B.; Miskey, C.; Staunstrup, N.H.; Gogol-Döring, A.; Bak, R.O.; Sharma, N.; Mátés, L.; Izsvák, Z.; Chen, W.; Ivics, Z.; et al. Comparative genomic integration profiling of Sleeping Beauty transposons mobilized with high efficacy from integrase-defective lentiviral vectors in primary human cells. Mol. Ther. 2011, 19, 1499–1510. [Google Scholar] [CrossRef] [PubMed]
- de Jong, J.; Akhtar, W.; Badhai, J.; Rust, A.G.; Rad, R.; Hilkens, J.; Berns, A.; van Lohuizen, M.; Wessels, L.F.A.; de Ridder, J. Chromatin landscapes of retroviral and transposon integration profiles. PLoS Genet. 2014, 10, e1004250. [Google Scholar] [CrossRef] [PubMed]
- Montaño, S.P.; Pigli, Y.Z.; Rice, P.A. The μ transpososome structure sheds light on DDE recombinase evolution. Nature 2012, 491, 413–417. [Google Scholar] [CrossRef] [PubMed]
- Morris, E.R.; Grey, H.; McKenzie, G.; Jones, A.C.; Richardson, J.M. A bend, flip and trap mechanism for transposon integration. Elife 2016, 5. [Google Scholar] [CrossRef] [PubMed]
- Passos, D.O.; Li, M.; Yang, R.; Rebensburg, S.V.; Ghirlando, R.; Jeon, Y.; Shkriabai, N.; Kvaratskhelia, M.; Craigie, R.; Lyumkis, D. Cryo-EM structures and atomic model of the HIV-1 strand transfer complex intasome. Science 2017, 355, 89–92. [Google Scholar] [CrossRef]
- Yin, Z.; Shi, K.; Banerjee, S.; Pandey, K.K.; Bera, S.; Grandgenett, D.P.; Aihara, H. Crystal structure of the Rous sarcoma virus intasome. Nature 2016, 530, 362–366. [Google Scholar] [CrossRef]
- Maertens, G.N.; Hare, S.; Cherepanov, P. The mechanism of retroviral integration from X-ray structures of its key intermediates. Nature 2010, 468, 326–329. [Google Scholar] [CrossRef]
- Maskell, D.P.; Renault, L.; Serrao, E.; Lesbats, P.; Matadeen, R.; Hare, S.; Lindemann, D.; Engelman, A.N.; Costa, A.; Cherepanov, P. Structural basis for retroviral integration into nucleosomes. Nature 2015, 523, 366–369. [Google Scholar] [CrossRef]
- Yanagihara, K.; Mizuuchi, K. Mismatch-targeted transposition of Mu: A new strategy to map genetic polymorphism. Proc. Natl. Acad. Sci. USA 2002, 99, 11317–11321. [Google Scholar] [CrossRef]
- Fuller, J.R.; Rice, P.A. Target DNA bending by the Mu transpososome promotes careful transposition and prevents its reversal. Elife 2017, 6. [Google Scholar] [CrossRef]
- Kuduvalli, P.N.; Rao, J.E.; Craig, N.L. Target DNA structure plays a critical role in Tn7 transposition. EMBO J. 2001, 20, 924–932. [Google Scholar] [CrossRef] [PubMed]
- Pribil, P.A.; Haniford, D.B. Target DNA Bending is an Important Specificity Determinant in Target Site Selection in Tn10 Transposition. J. Mol. Biol. 2003, 330, 247–259. [Google Scholar] [CrossRef]
- Benjamin, H.W.; Kleckner, N. Intramolecular transposition by Tn10. Cell 1989, 59, 373–383. [Google Scholar] [CrossRef]
- Maxwell, A.; Craigie, R.; Mizuuchi, K. B protein of bacteriophage mu is an ATPase that preferentially stimulates intermolecular DNA strand transfer. Proc. Natl. Acad. Sci. USA 1987, 84, 699–703. [Google Scholar] [CrossRef] [PubMed]
- Karsi, A.; Moav, B.; Hackett, P.; Liu, Z. Effects of insert size on transposition efficiency of the sleeping beauty transposon in mouse cells. Mar. Biotechnol. 2001, 3, 241–245. [Google Scholar] [CrossRef] [PubMed]
- Lampe, D.J.; Grant, T.E.; Robertson, H.M. Factors affecting transposition of the Himar1 mariner transposon in vitro. Genetics 1998, 149, 179–187. [Google Scholar]
- Fischer, S.E.; van Luenen, H.G.; Plasterk, R.H. Cis requirements for transposition of Tc1-like transposons in C. elegans. Mol. Genet. Genom. 1999, 262, 268–274. [Google Scholar] [CrossRef]
- Mansharamani, M.; Graham, D.R.M.; Monie, D.; Lee, K.K.; Hildreth, J.E.K.; Siliciano, R.F.; Wilson, K.L. Barrier-to-autointegration factor BAF binds p55 Gag and matrix and is a host component of human immunodeficiency virus type 1 virions. J. Virol. 2003, 77, 13084–13092. [Google Scholar] [CrossRef]
- Suzuki, Y.; Craigie, R. Regulatory mechanisms by which barrier-to-autointegration factor blocks autointegration and stimulates intermolecular integration of Moloney murine leukemia virus preintegration complexes. J. Virol. 2002, 76, 12376–12380. [Google Scholar] [CrossRef]
- Lee, M.S.; Craigie, R. A previously unidentified host protein protects retroviral DNA from autointegration. Proc. Natl. Acad. Sci. USA 1998, 95, 1528–1533. [Google Scholar] [CrossRef]
- Lee, M.S.; Craigie, R. Protection of retroviral DNA from autointegration: Involvement of a cellular factor. Proc. Natl. Acad. Sci. USA 1994, 91, 9823–9827. [Google Scholar] [CrossRef] [PubMed]
- Tower, J.; Karpen, G.H.; Craig, N.; Spradling, A.C. Preferential transposition of Drosophila P elements to nearby chromosomal sites. Genetics 1993, 133, 347–359. [Google Scholar]
- Ruf, S.; Symmons, O.; Uslu, V.V.; Dolle, D.; Hot, C.; Ettwiller, L.; Spitz, F. Large-scale analysis of the regulatory architecture of the mouse genome with a transposon-associated sensor. Nat. Genet. 2011, 43, 379–386. [Google Scholar] [CrossRef]
- Dupuy, A.J.; Fritz, S.; Largaespada, D.A. Transposition and gene disruption in the male germline of the mouse. Genesis 2001, 30, 82–88. [Google Scholar] [CrossRef] [PubMed]
- Kokubu, C.; Horie, K.; Abe, K.; Ikeda, R.; Mizuno, S.; Uno, Y.; Ogiwara, S.; Ohtsuka, M.; Isotani, A.; Okabe, M.; et al. A transposon-based chromosomal engineering method to survey a large cis-regulatory landscape in mice. Nat. Genet. 2009, 41, 946–952. [Google Scholar] [CrossRef] [PubMed]
- Liang, Q.; Kong, J.; Stalker, J.; Bradley, A. Chromosomal mobilization and reintegration of Sleeping Beauty and PiggyBac transposons. Genesis 2009, 47, 404–408. [Google Scholar] [CrossRef] [PubMed]
- Horie, K.; Yusa, K.; Yae, K.; Odajima, J.; Fischer, S.E.J.; Keng, V.W.; Hayakawa, T.; Mizuno, S.; Kondoh, G.; Ijiri, T.; et al. Characterization of Sleeping Beauty transposition and its application to genetic screening in mice. Mol. Cell. Biol. 2003, 23, 9189–9207. [Google Scholar] [CrossRef]
- Carlson, C.M.; Dupuy, A.J.; Fritz, S.; Roberg-Perez, K.J.; Fletcher, C.F.; Largaespada, D.A. Transposon mutagenesis of the mouse germline. Genetics 2003, 165, 243–256. [Google Scholar]
- Fischer, S.E.; Wienholds, E.; Plasterk, R.H. Regulated transposition of a fish transposon in the mouse germ line. Proc. Natl. Acad. Sci. USA 2001, 98, 6759–6764. [Google Scholar] [CrossRef]
- Yergeau, D.A.; Kelley, C.M.; Kuliyev, E.; Zhu, H.; Johnson Hamlet, M.R.; Sater, A.K.; Wells, D.E.; Mead, P.E. Remobilization of Sleeping Beauty transposons in the germline of Xenopus tropicalis. Mob. DNA 2011, 2, 15. [Google Scholar] [CrossRef]
- Keng, V.W.; Yae, K.; Hayakawa, T.; Mizuno, S.; Uno, Y.; Yusa, K.; Kokubu, C.; Kinoshita, T.; Akagi, K.; Jenkins, N.A.; et al. Region-specific saturation germline mutagenesis in mice using the Sleeping Beauty transposon system. Nat. Methods 2005, 2, 763–769. [Google Scholar] [CrossRef] [PubMed]
- Knapp, S.; Larondelle, Y.; Rossberg, M.; Furtek, D.; Theres, K. Transgenic tomato lines containing Ds elements at defined genomic positions as tools for targeted transposon tagging. Mol. Genet. Genom. 1994, 243, 666–673. [Google Scholar] [CrossRef] [PubMed]
- Mátés, L.; Chuah, M.K.L.; Belay, E.; Jerchow, B.; Manoj, N.; Acosta-Sanchez, A.; Grzela, D.P.; Schmitt, A.; Becker, K.; Matrai, J.; et al. Molecular evolution of a novel hyperactive Sleeping Beauty transposase enables robust stable gene transfer in vertebrates. Nat. Genet. 2009, 41, 753–761. [Google Scholar] [CrossRef] [PubMed]
- Rostovskaya, M.; Fu, J.; Obst, M.; Baer, I.; Weidlich, S.; Wang, H.; Smith, A.J.H.; Anastassiadis, K.; Stewart, A.F. Transposon-mediated BAC transgenesis in human ES cells. Nucleic Acids Res. 2012, 40, e150. [Google Scholar] [CrossRef] [PubMed]
- Darquet, A.M.; Cameron, B.; Wils, P.; Scherman, D.; Crouzet, J. A new DNA vehicle for nonviral gene delivery: Supercoiled minicircle. Gene Ther. 1997, 4, 1341–1349. [Google Scholar] [CrossRef] [PubMed]
- Holstein, M.; Mesa-Nuñez, C.; Miskey, C.; Almarza, E.; Poletti, V.; Schmeer, M.; Grueso, E.; Ordóñez Flores, J.C.; Kobelt, D.; Walther, W.; et al. Efficient Non-viral Gene Delivery into Human Hematopoietic Stem Cells by Minicircle Sleeping Beauty Transposon Vectors. Mol. Ther. 2018, 26, 1137–1153. [Google Scholar] [CrossRef]
- Monjezi, R.; Miskey, C.; Gogishvili, T.; Schleef, M.; Schmeer, M.; Einsele, H.; Ivics, Z.; Hudecek, M. Enhanced CAR T-cell engineering using non-viral Sleeping Beauty transposition from minicircle vectors. Leukemia 2017, 31, 186–194. [Google Scholar] [CrossRef] [PubMed]
- Kesselring, L.; Miskey, C.; Zuliani, C.; Querques, I.; Kapitonov, V.; Laukó, A.; Fehér, A.; Palazzo, A.; Diem, T.; Lustig, J.; et al. A single amino acid switch converts the Sleeping Beauty transposase into an efficient unidirectional excisionase with utility in stem cell reprogramming. Nucleic Acids Res. 2020, 48, 316–331. [Google Scholar] [CrossRef]
- Kebriaei, P.; Izsvák, Z.; Narayanavari, S.A.; Singh, H.; Ivics, Z. Gene Therapy with the Sleeping Beauty Transposon System. Trends Genet. 2017, 33, 852–870. [Google Scholar] [CrossRef]
- Kettlun, C.; Galvan, D.L.; George, A.L.; Kaja, A.; Wilson, M.H. Manipulating piggyBac transposon chromosomal integration site selection in human cells. Mol. Ther. 2011, 19, 1636–1644. [Google Scholar] [CrossRef]
- Yant, S.R.; Huang, Y.; Akache, B.; Kay, M.A. Site-directed transposon integration in human cells. Nucleic Acids Res. 2007, 35, e50. [Google Scholar] [CrossRef] [PubMed]
- Klompe, S.E.; Vo, P.L.H.; Halpin-Healy, T.S.; Sternberg, S.H. Transposon-encoded CRISPR-Cas systems direct RNA-guided DNA integration. Nature 2019, 571, 219–225. [Google Scholar] [CrossRef] [PubMed]
- Strecker, J.; Ladha, A.; Gardner, Z.; Schmid-Burgk, J.L.; Makarova, K.S.; Koonin, E.V.; Zhang, F. RNA-guided DNA insertion with CRISPR-associated transposases. Science 2019, 365, 48–53. [Google Scholar] [CrossRef] [PubMed]
- Jinek, M.; Chylinski, K.; Fonfara, I.; Hauer, M.; Doudna, J.A.; Charpentier, E. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 2012, 337, 816–821. [Google Scholar] [CrossRef]
- Kovač, A.; Miskey, C.; Menzel, M.; Grueso, E.; Gogol-Döring, A.; Ivics, Z. RNA-guided retargeting of Sleeping Beauty transposition in human cells. ELife 2020, 9. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ochmann, M.T.; Ivics, Z. Jumping Ahead with Sleeping Beauty: Mechanistic Insights into Cut-and-Paste Transposition. Viruses 2021, 13, 76. https://doi.org/10.3390/v13010076
Ochmann MT, Ivics Z. Jumping Ahead with Sleeping Beauty: Mechanistic Insights into Cut-and-Paste Transposition. Viruses. 2021; 13(1):76. https://doi.org/10.3390/v13010076
Chicago/Turabian StyleOchmann, Matthias T., and Zoltán Ivics. 2021. "Jumping Ahead with Sleeping Beauty: Mechanistic Insights into Cut-and-Paste Transposition" Viruses 13, no. 1: 76. https://doi.org/10.3390/v13010076
APA StyleOchmann, M. T., & Ivics, Z. (2021). Jumping Ahead with Sleeping Beauty: Mechanistic Insights into Cut-and-Paste Transposition. Viruses, 13(1), 76. https://doi.org/10.3390/v13010076






