A Case of Phage Therapy against Pandrug-Resistant Achromobacter xylosoxidans in a 12-Year-Old Lung-Transplanted Cystic Fibrosis Patient
Abstract
1. Case Presentation
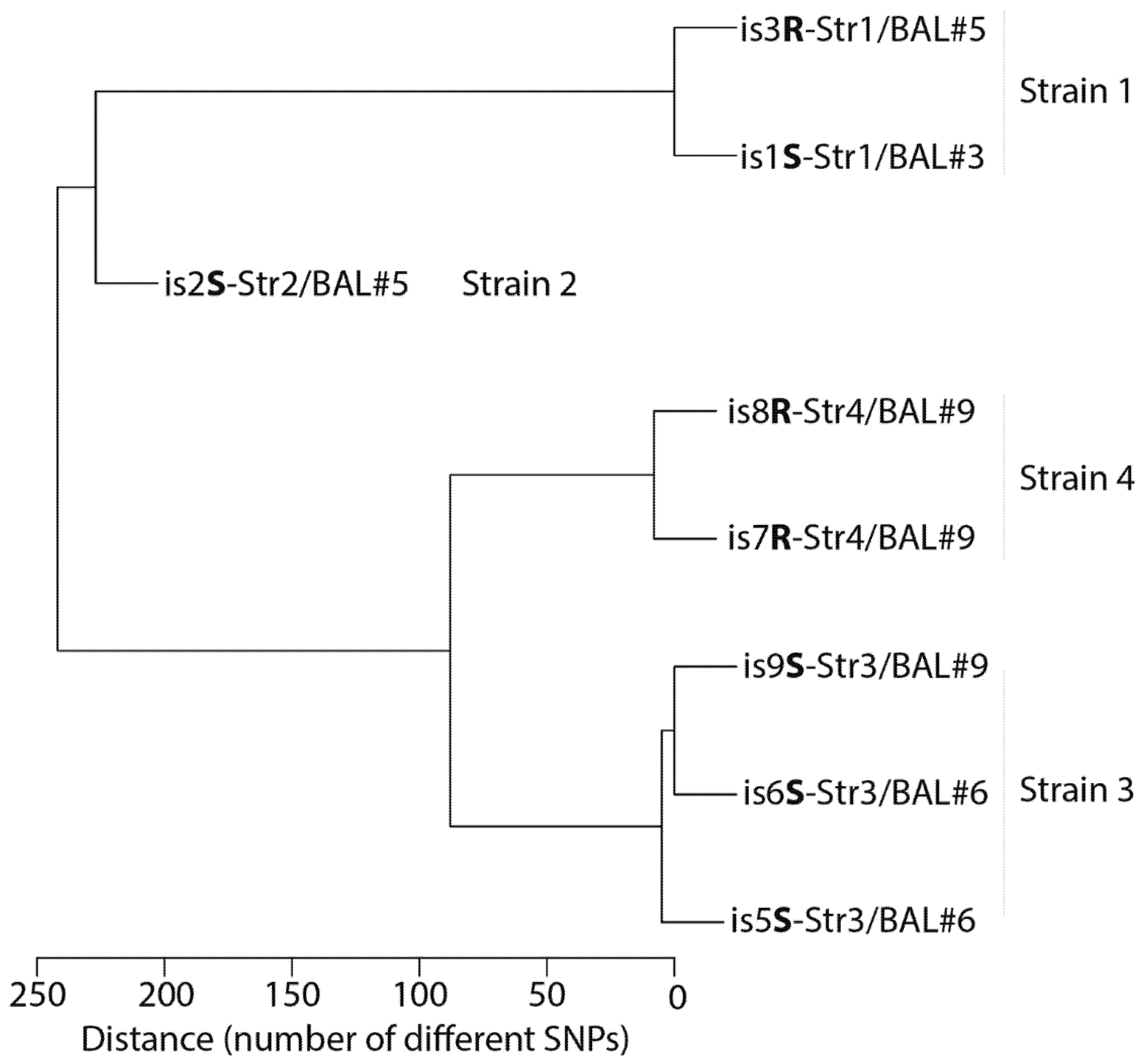
2. Microbiological Analysis of Eight A. xylosoxidans Isolates
3. Discussion and Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Magiorakos, A.P.; Srinivasan, A.; Carey, R.B.; Carmeli, Y.; Falagas, M.E.; Giske, C.G.; Harbarth, S.; Hindler, J.F.; Kahlmeter, G.; Olsson-Liljequist, B.; et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: An international expert proposal for interim standard definitions for acquired resistance. Clin. Microbiol. Infect. 2012, 18, 268–281. [Google Scholar] [CrossRef] [PubMed]
- Dreiseikelmann, B.; Bunk, B.; Sproer, C.; Rohde, M.; Nimtz, M.; Wittmann, J. Characterization and genome comparisons of three Achromobacter phages of the family Siphoviridae. Arch. Virol. 2017, 162, 2191–2201. [Google Scholar] [CrossRef] [PubMed]
- Wittmann, J.; Dreiseikelmann, B.; Rohde, M.; Meier-Kolthoff, J.P.; Bunk, B.; Rohde, C. First genome sequences of Achromobacter phages reveal new members of the N4 family. Virol. J. 2014, 11, 14. [Google Scholar] [CrossRef]
- Merabishvili, M.; Pirnay, J.P.; Verbeken, G.; Chanishvili, N.; Tediashvili, M.; Lashkhi, N.; Glonti, T.; Krylov, V.; Mast, J.; Van Parys, L.; et al. Quality-controlled small-scale production of a well-defined bacteriophage cocktail for use in human clinical trials. PLoS ONE 2009, 4, e4944. [Google Scholar] [CrossRef] [PubMed]
- Cools, P.; Ho, E.; Vranckx, K.; Schelstraete, P.; Wurth, B.; Franckx, H.; Ieven, G.; Van Simaey, L.; Van daele, S.; Verhulst, S.; et al. Epidemic Achromobacter xylosoxidans strain among Belgian cystic fibrosis patients and review of literature. BMC Microbiol. 2016, 16, 122. [Google Scholar] [CrossRef] [PubMed]
- Deschaght, P.; Van Simaey, L.; Decat, E.; Van Mechelen, E.; Brisse, S.; Vaneechoutte, M. Rapid genotyping of Achromobacter xylosoxidans, Acinetobacter baumannii, Klebsiella pneumoniae, Pseudomonas aeruginosa and Stenotrophomonas maltophilia isolates using melting curve analysis of RAPD-generated DNA fragments (McRAPD). Res. Microbiol. 2011, 162, 386–392. [Google Scholar] [CrossRef] [PubMed]
- Hantke, K. Compilation of Escherichia coli K-12 outer membrane phage receptors—Their function and some historical remarks. FEMS Microbiol. Lett. 2020, 367. [Google Scholar] [CrossRef] [PubMed]
- Abedon, S.T. Bacteriophage clinical use as antibacterial “drugs”: Utility and precedent. Microbiol. Spectr. 2017, 5. [Google Scholar] [CrossRef]
- Chanishvili, N. Bacteriophages as therapeutic and prophylactic means: Summary of the Soviet and post Soviet experiences. Curr. Drug Deliv. 2016, 13, 309–323. [Google Scholar] [CrossRef]
- Jault, P.; Leclerc, T.; Jennes, S.; Pirnay, J.P.; Que, Y.A.; Resch, G.; Rousseau, A.F.; Ravat, F.; Carsin, H.; Le Floch, R.; et al. Efficacy and tolerability of a cocktail of bacteriophages to treat burn wounds infected by Pseudomonas aeruginosa (PhagoBurn): A randomised, controlled, double-blind phase 1/2 trial. Lancet Infect. Dis. 2019, 19, 35–45. [Google Scholar] [CrossRef]
- Leitner, L.; Sybesma, W.; Chanishvili, N.; Goderdzishvili, M.; Chkhotua, A.; Ujmajuridze, A.; Schneider, M.P.; Sartori, A.; Mehnert, U.; Bachmann, L.M.; et al. Bacteriophages for treating urinary tract infections in patients undergoing transurethral resection of the prostate: A randomized, placebo-controlled, double-blind clinical trial. BMC Urol. 2017, 17, 90. [Google Scholar] [CrossRef]
- Wright, A.; Hawkins, C.H.; Anggård, E.E.; Harper, D.R. A controlled clinical trial of a therapeutic bacteriophage preparation in chronic otitis due to antibiotic-resistant Pseudomonas aeruginosa; A preliminary report of efficacy. Clin. Otolaryngol. 2009, 34, 349–357. [Google Scholar] [CrossRef] [PubMed]
- Pirnay, J.P.; Kutter, E. Bacteriophages: It’s a medicine, Jim, but not as we know it. Lancet Infect. Dis. 2020. [Google Scholar] [CrossRef]
- Ferry, T.; Boucher, F.; Fevre, C.; Perpoint, T.; Chateau, J.; Petitjean, C.; Josse, J.; Chidiac, C.; L’Hostis, G.; Leboucher, G.; et al. Innovations for the treatment of a complex bone and joint infection due to XDR Pseudomonas aeruginosa including local application of a selected cocktail of bacteriophages. J. Antimicrob. Chemother. 2018, 73, 2901–2903. [Google Scholar] [CrossRef] [PubMed]
- Ferry, T.; Leboucher, G.; Fevre, C.; Herry, Y.; Conrad, A.; Josse, J.; Batailler, C.; Chidiac, C.; Medina, M.; Lustig, S.; et al. Salvage debridement, antibiotics and implant retention (“DAIR”) with local injection of a selected cocktail of bacteriophages: Is it an option for an elderly patient with relapsing Staphylococcus aureus prosthetic-joint infection? Open Forum Infect. Dis. 2018, 5, ofy269. [Google Scholar] [CrossRef] [PubMed]
- Jennes, S.; Merabishvili, M.; Soentjens, P.; Pang, K.W.; Rose, T.; Keersebilck, E.; Soete, O.; Francois, P.M.; Teodorescu, S.; Verween, G.; et al. Use of bacteriophages in the treatment of colistin-only-sensitive Pseudomonas aeruginosa septicaemia in a patient with acute kidney injury—A case report. Crit. Care 2017, 21, 129. [Google Scholar] [CrossRef] [PubMed]
- Petrovic Fabijan, A.; Lin, R.C.Y.; Ho, J.; Maddocks, S.; Ben Zakour, N.L.; Iredell, J.R.; Team, W.B.T. Safety of bacteriophage therapy in severe Staphylococcus aureus infection. Nat. Microbiol. 2020, 5, 465–472. [Google Scholar] [CrossRef]
- Sulakvelidze, A.; Alavidze, Z.; Morris, J.G., Jr. Bacteriophage therapy. Antimicrob. Agents Chemother. 2001, 45, 649–659. [Google Scholar] [CrossRef]
- Trend, S.; Fonceca, A.M.; Ditcham, W.G.; Kicic, A.; Cf, A. The potential of phage therapy in cystic fibrosis: Essential human-bacterial-phage interactions and delivery considerations for use in Pseudomonas aeruginosa-infected airways. J. Cyst. Fibros. 2017, 16, 663–670. [Google Scholar] [CrossRef]
- Law, N.; Logan, C.; Yung, G.; Furr, C.L.; Lehman, S.M.; Morales, S.; Rosas, F.; Gaidamaka, A.; Bilinsky, I.; Grint, P.; et al. Successful adjunctive use of bacteriophage therapy for treatment of multidrug-resistant Pseudomonas aeruginosa infection in a cystic fibrosis patient. Infection 2019, 47, 665–668. [Google Scholar] [CrossRef]
- Hoyle, N.; Zhvaniya, P.; Balarjishvili, N.; Bolkvadze, D.; Nadareishvili, L.; Nizharadze, D.; Wittmann, J.; Rohde, C.; Kutateladze, M. Phage therapy against Achromobacter xylosoxidans lung infection in a patient with cystic fibrosis: A case report. Res. Microbiol. 2018, 169, 540–542. [Google Scholar] [CrossRef]
- Carrigy, N.B.; Chang, R.Y.; Leung, S.S.Y.; Harrison, M.; Petrova, Z.; Pope, W.H.; Hatfull, G.F.; Britton, W.J.; Chan, H.K.; Sauvageau, D.; et al. Anti-tuberculosis bacteriophage D29 delivery with a vibrating mesh nebulizer, jet nebulizer, and soft mist inhaler. Pharm. Res. 2017, 34, 2084–2096. [Google Scholar] [CrossRef]
- Rohde, C.; Resch, G.; Pirnay, J.P.; Blasdel, B.G.; Debarbieux, L.; Gelman, D.; Górski, A.; Hazan, R.; Huys, I.; Kakabadze, E.; et al. Expert Opinion on Three Phage Therapy Related Topics: Bacterial Phage Resistance, Phage Training and Prophages in Bacterial Production Strains. Viruses 2018, 10, 178. [Google Scholar] [CrossRef]
- Hughes, D.; Andersson, D.I. Environmental and genetic modulation of the phenotypic expression of antibiotic resistance. FEMS Microbiol. Rev. 2017, 41, 374–391. [Google Scholar] [CrossRef] [PubMed]
- Urmi, U.L.; Nahar, S.; Rana, M.; Sultana, F.; Jahan, N.; Hossain, B.; Alam, M.S.; Mosaddek, A.S.M.; McKimm, J.; Rahman, N.A.A.; et al. Genotypic to phenotypic resistance discrepancies identified involving β-lactamase genes, blaKPC, blaIMP, blaNDM-1, and blaVIM in uropathogenic Klebsiella pneumoniae. Infect. Drug Resist. 2020, 13, 2863–2875. [Google Scholar] [CrossRef]
- Bull, J.J.; Vegge, C.S.; Schmerer, M.; Chaudhry, W.N.; Levin, B.R. Phenotypic resistance and the dynamics of bacterial escape from phage control. PLoS ONE 2014, 9, e94690. [Google Scholar] [CrossRef] [PubMed]
- Capparelli, R.; Nocerino, N.; Iannaccone, M.; Ercolini, D.; Parlato, M.; Chiara, M.; Iannelli, D. Bacteriophage therapy of Salmonella enterica: A fresh appraisal of bacteriophage therapy. J. Infect. Dis. 2010, 201, 52–61. [Google Scholar] [CrossRef]
- Hung, C.H.; Kuo, C.F.; Wang, C.H.; Wu, C.M.; Tsao, N. Experimental phage therapy in treating Klebsiella pneumoniae-mediated liver abscesses and bacteremia in mice. Antimicrob. Agents Chemother. 2011, 55, 1358–1365. [Google Scholar] [CrossRef] [PubMed]
- Pouillot, F.; Chomton, M.; Blois, H.; Courroux, C.; Noelig, J.; Bidet, P.; Bingen, E.; Bonacorsi, S. Efficacy of bacteriophage therapy in experimental sepsis and meningitis caused by a clone O25b:H4-ST131 Escherichia coli strain producing CTX-M-15. Antimicrob. Agents Chemother. 2012, 56, 3568–3575. [Google Scholar] [CrossRef] [PubMed]
| Date | 19 May 2017 | 23 May 2017 | 6 June 2017 | 12 June 2017 | 25 July 2017 | 5 September 2017 | 8 September 2017 | 15 September 2017 | 12 October 2017 | 27 October 2017 | 23 January 2018 | 8 February 2018 | 5 March 2018 | 18 May 2018 | 19 February 2019 | 19 June 2019 | 27 August 2019 | 30 April 2020 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Phage therapy | ROUND 1 | ROUND 2 | ||||||||||||||||
| Tigecycline | from 29 June to 31 July | |||||||||||||||||
| Imipenem | From 31 July 2017, to 16 February 2018 | |||||||||||||||||
| Number of days, relative to the first round of phage therapy | −112 | −108 | −94 | −88 | −45 | −3 | +7 | +37 | +49 | +137 | +153 | +178 | +252 | +529 | +649 | +718 | +963 | |
| BAL | ||||||||||||||||||
| BAL Number * | #1 | #2 | #3 | #4 | #5 | #6 | #7 | #8 | #9 | #10 | #11 | #12 | ||||||
| Leukocytes (/mm3) | >1000 | 800 | >1000 | 500 | >1000 | 60 | >1000 | 810 | >1000 | NA | NA | NA | ||||||
| PMN cells (%) | 90 | 88 | 90 | 91 | 43 | 30 | 80 | 69 | 82 | NA | NA | NA | ||||||
| Quantification of A. x culture (CFU/mL) | 104 | 103 | 104 | 103 | 104 | 103 | 104 | 106 | 105 | 104 | 0 | 0 | ||||||
| Other bacteria (CFU/mL) | None | None | None | None | None | None | None | None | None | None | 104 S. a | 106 S. a | ||||||
| Sputum | ||||||||||||||||||
| Leucocytes (/field) | >25 | >25 | >25 | >25 | NA | NA | NA | |||||||||||
| Quantification of A. x culture (CFU/mL) | 105 | 105 | 105 | 106 | 103 | 103 | 103 | |||||||||||
| Other bacteria (CFU/mL) | None | None | None | None | 108 M. c | None | 106 S. a | |||||||||||
| Complete Name | Isolate | Strain | Source of Sampling | Date of Sampling | Whole Genome Sequence * |
|---|---|---|---|---|---|
| is1S-Str1/BAL#3 | is1S | Str1 | BAL#3 | 25 July 2017 | ERS5044236-UGAX1 |
| is2S-Str2/BAL#5 | is2S | Str2 | BAL#5 | 15 September 2017 | ERS5044237-UGAX2 |
| is3R-Str1/BAL#5 | is3R | Str1 | BAL#5 | 15 September 2017 | ERS5044238-UGAX3 |
| is5S-Str3/BAL#6 | is5S | Str3 | BAL#6 | 12 October 2017 | ERS5044239-UGAX5 |
| is6S-Str3/BAL#6 | is6S | Str3 | BAL#6 | 12 October 2017 | ERS5044240-UGAX6 |
| is7R-Str4/BAL#9 | is7R | Str4 | BAL#9 | 8 February 2018 | ERS5044241-UGAX7 |
| is8R-Str4/BAL#9 | is8R | Str4 | BAL#9 | 8 February 2018 | ERS5044242-UGAX8 |
| is9S-Str3/BAL#9 | is9S | Str3 | BAL#9 | 8 February 2018 | ERS5044243-UGAX9 |
| Bronchoalveolar Lavage | BAL#5 | BAL#6 | BAL#9 | |
|---|---|---|---|---|
| 15 September 2017 | 12 October 2017 | 8 February 2018 | ||
| qPCR for Strain | Isolates | |||
| Str1 | is1S, is3R | N | NS | P (34.7) |
| Str2 | is2S | P (36.6) | N | N |
| Str3 | is5S, is6S, is9S | P (32.7) | P (28.9) | P (27.7) |
| Str4 | is7R, is8R | NS | NS | P (35.2) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lebeaux, D.; Merabishvili, M.; Caudron, E.; Lannoy, D.; Van Simaey, L.; Duyvejonck, H.; Guillemain, R.; Thumerelle, C.; Podglajen, I.; Compain, F.; et al. A Case of Phage Therapy against Pandrug-Resistant Achromobacter xylosoxidans in a 12-Year-Old Lung-Transplanted Cystic Fibrosis Patient. Viruses 2021, 13, 60. https://doi.org/10.3390/v13010060
Lebeaux D, Merabishvili M, Caudron E, Lannoy D, Van Simaey L, Duyvejonck H, Guillemain R, Thumerelle C, Podglajen I, Compain F, et al. A Case of Phage Therapy against Pandrug-Resistant Achromobacter xylosoxidans in a 12-Year-Old Lung-Transplanted Cystic Fibrosis Patient. Viruses. 2021; 13(1):60. https://doi.org/10.3390/v13010060
Chicago/Turabian StyleLebeaux, David, Maia Merabishvili, Eric Caudron, Damien Lannoy, Leen Van Simaey, Hans Duyvejonck, Romain Guillemain, Caroline Thumerelle, Isabelle Podglajen, Fabrice Compain, and et al. 2021. "A Case of Phage Therapy against Pandrug-Resistant Achromobacter xylosoxidans in a 12-Year-Old Lung-Transplanted Cystic Fibrosis Patient" Viruses 13, no. 1: 60. https://doi.org/10.3390/v13010060
APA StyleLebeaux, D., Merabishvili, M., Caudron, E., Lannoy, D., Van Simaey, L., Duyvejonck, H., Guillemain, R., Thumerelle, C., Podglajen, I., Compain, F., Kassis, N., Mainardi, J.-L., Wittmann, J., Rohde, C., Pirnay, J.-P., Dufour, N., Vermeulen, S., Gansemans, Y., Van Nieuwerburgh, F., & Vaneechoutte, M. (2021). A Case of Phage Therapy against Pandrug-Resistant Achromobacter xylosoxidans in a 12-Year-Old Lung-Transplanted Cystic Fibrosis Patient. Viruses, 13(1), 60. https://doi.org/10.3390/v13010060









