Evaluation of the 50% Infectious Dose of Human Norovirus Cin-2 in Gnotobiotic Pigs: A Comparison of Classical and Contemporary Methods for Endpoint Estimation
Abstract
1. Introduction
2. Materials and Methods
2.1. Virus Inoculum
2.2. Gnotobiotic Pigs and Treatments
2.3. HBGA-Typing of Gn Pigs by Immunofluorescence Assay
2.4. Assessment of Fecal Consistency and Detection of HuNoV Shedding by RT-qPCR
2.5. Statistical Analysis
2.5.1. Analysis of Variance among Challenge Doses
2.5.2. ID50 and DD50 Calculations Using Different Approaches
Reed–Muench, Dragstedt–Behrens and Spearman–Karber Methods
Exponential and Beta-Poisson Dose–Response Models
3. Results
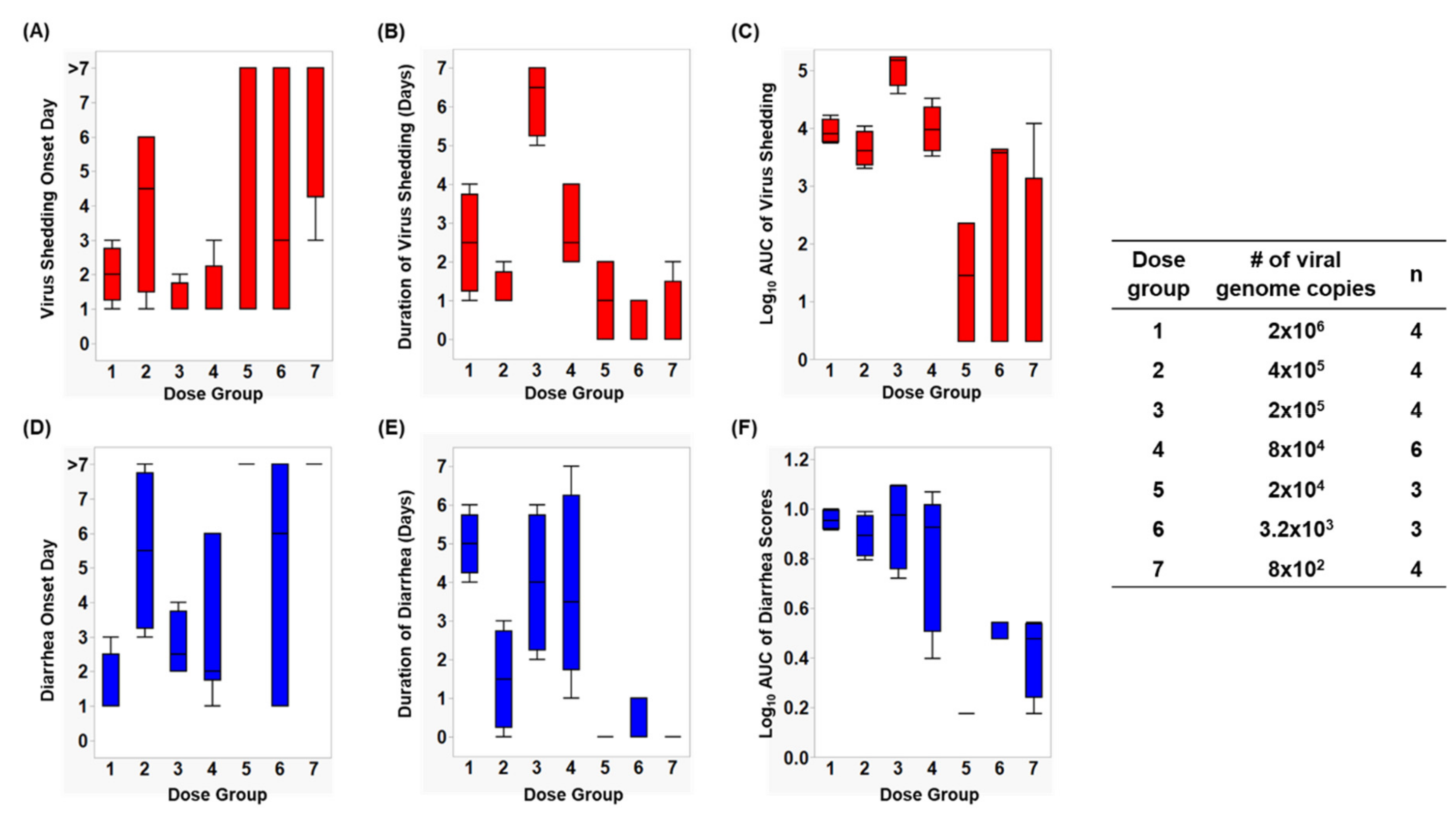
3.1. Assessment of Infection Status in Gn Pigs
3.2. Assessment of Diarrhea Status in Gn Pigs
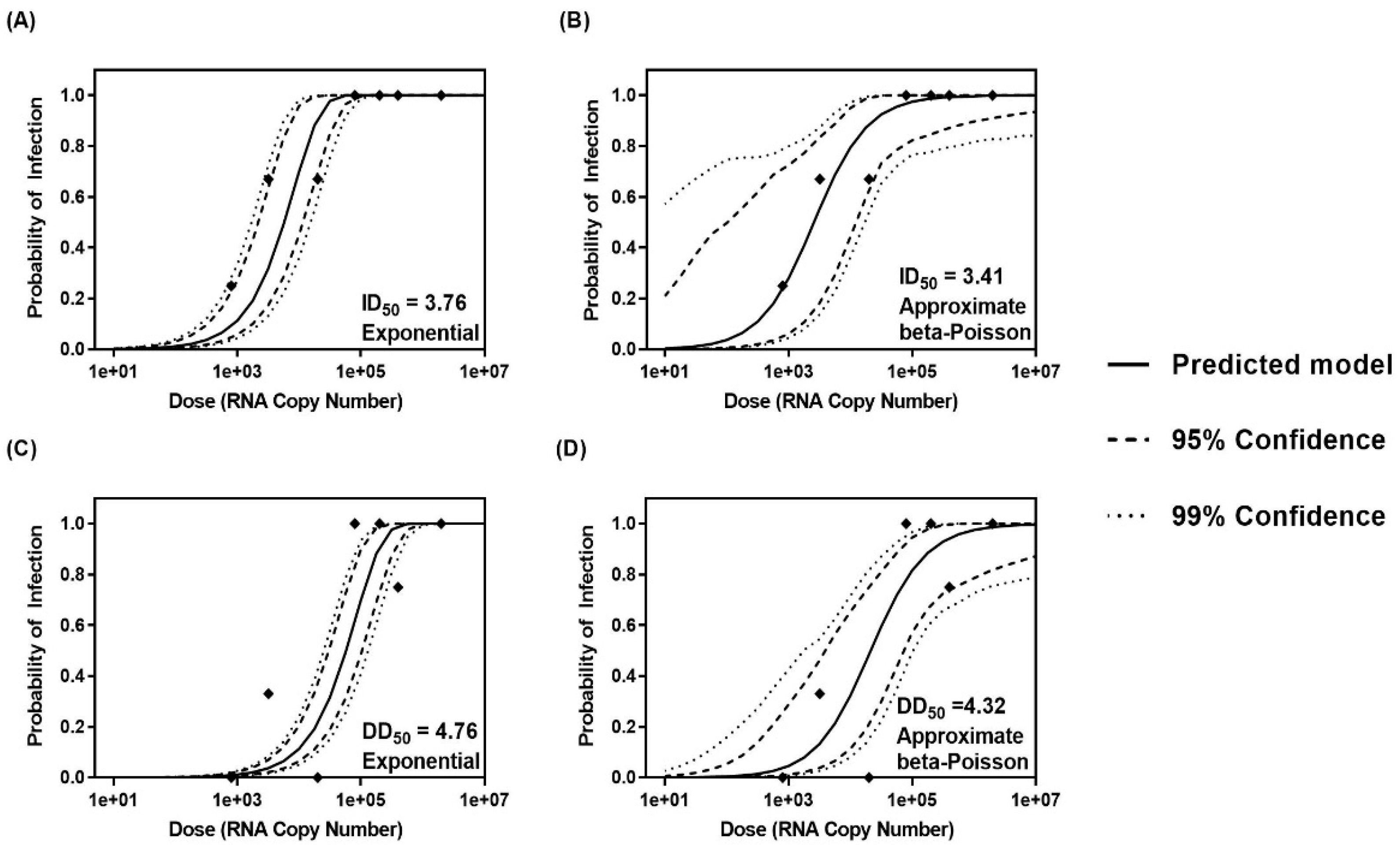
3.3. Determination of ID50 and DD50 Using Various Dose–Response Models
3.4. Comparison of Infectiousness of HuNoV GII.4/2003 Variants Cin-1 and Cin-2 in Humans and Gn Pigs
3.5. Comparison of Methods for Dose–response Analysis of Cin-2
3.6. Determination of an Optimal Challenge Dose
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Payne, D.C.; Vinjé, J.; Szilagyi, P.G.; Edwards, K.M.; Staat, M.A.; Weinberg, G.A.; Hall, C.B.; Chappell, J.; Bernstein, D.I.; Curns, A.T.; et al. Norovirus and Medically Attended Gastroenteritis in U.S. Children. N. Engl. J. Med. 2013, 368, 1121–1130. [Google Scholar] [CrossRef] [PubMed]
- Pringle, K.; Lopman, B.; Vega, E.; Vinje, J.; Parashar, U.D.; Hall, A.J. Noroviruses: Epidemiology, immunity and prospects for prevention. Future Microbiol. 2015, 10, 53–67. [Google Scholar] [CrossRef] [PubMed]
- Rouhani, S.; Peñataro Yori, P.; Paredes Olortegui, M.; Siguas Salas, M.; Rengifo Trigoso, D.; Mondal, D.; Bodhidatta, L.; Platts-Mills, J.; Samie, A.; Kabir, F.; et al. Norovirus Infection and Acquired Immunity in 8 Countries: Results From the MAL-ED Study. Clin. Infect. Dis. Off. Publ. Infect. Dis. Soc. Am. 2016, 62, 1210–1217. [Google Scholar] [CrossRef]
- Kirk, M.D.; Pires, S.M.; Black, R.E.; Caipo, M.; Crump, J.A.; Devleesschauwer, B.; Döpfer, D.; Fazil, A.; Fischer-Walker, C.L.; Hald, T.; et al. World Health Organization Estimates of the Global and Regional Disease Burden of 22 Foodborne Bacterial, Protozoal, and Viral Diseases, 2010: A Data Synthesis. PLoS Med. 2015, 12, e1001921. [Google Scholar] [CrossRef]
- Atmar, R.L.; Baehner, F.; Cramer, J.P.; Lloyd, E.; Sherwood, J.; Borkowski, A.; Mendelman, P.M.; Group, N.O.R.S. Antibody persistence to two Virus-Like Particle norovirus vaccine candidate formulations in healthy adults: One-year follow-up with memory probe vaccination. J. Infect. Dis. 2019. [Google Scholar] [CrossRef]
- Sundararajan, A.; Sangster, M.Y.; Frey, S.; Atmar, R.L.; Chen, W.H.; Ferreira, J.; Bargatze, R.; Mendelman, P.M.; Treanor, J.J.; Topham, D.J. Robust mucosal-homing antibody-secreting B cell responses induced by intramuscular administration of adjuvanted bivalent human norovirus-like particle vaccine. Vaccine 2015, 33, 568–576. [Google Scholar] [CrossRef]
- Bernstein, D.I.; Atmar, R.L.; Lyon, G.M.; Treanor, J.J.; Chen, W.H.; Jiang, X.; Vinje, J.; Gregoricus, N.; Frenck, R.W., Jr.; Moe, C.L.; et al. Norovirus vaccine against experimental human GII.4 virus illness: A challenge study in healthy adults. J. Infect. Dis. 2015, 211, 870–878. [Google Scholar] [CrossRef]
- Treanor, J.J.; Atmar, R.L.; Frey, S.E.; Gormley, R.; Chen, W.H.; Ferreira, J.; Goodwin, R.; Borkowski, A.; Clemens, R.; Mendelman, P.M. A novel intramuscular bivalent norovirus virus-like particle vaccine candidate--reactogenicity, safety, and immunogenicity in a phase 1 trial in healthy adults. J. Infect. Dis. 2014, 210, 1763–1771. [Google Scholar] [CrossRef]
- Gerdts, V.; Wilson, H.L.; Meurens, F.; van Drunen Littel-van den Hurk, S.; Wilson, D.; Walker, S.; Wheler, C.; Townsend, H.; Potter, A.A. Large Animal Models for Vaccine Development and Testing. ILAR J. 2015, 56, 53–62. [Google Scholar] [CrossRef]
- Bok, K.; Parra, G.I.; Mitra, T.; Abente, E.; Shaver, C.K.; Boon, D.; Engle, R.; Yu, C.; Kapikian, A.Z.; Sosnovtsev, S.V.; et al. Chimpanzees as an animal model for human norovirus infection and vaccine development. Proc. Natl. Acad. Sci. USA 2011, 108, 325–330. [Google Scholar] [CrossRef]
- Cheetham, S.; Souza, M.; McGregor, R.; Meulia, T.; Wang, Q.; Saif, L.J. Binding Patterns of Human Norovirus-Like Particles to Buccal and Intestinal Tissues of Gnotobiotic Pigs in Relation to A/H Histo-Blood Group Antigen Expression. J. Virol. 2007, 81, 3535–3544. [Google Scholar] [CrossRef]
- Souza, M.; Azevedo, M.S.; Jung, K.; Cheetham, S.; Saif, L.J. Pathogenesis and immune responses in gnotobiotic calves after infection with the genogroup II.4-HS66 strain of human norovirus. J. Virol. 2008, 82, 1777–1786. [Google Scholar] [CrossRef]
- Taube, S.; Kolawole, A.O.; Höhne, M.; Wilkinson, J.E.; Handley, S.A.; Perry, J.W.; Thackray, L.B.; Akkina, R.; Wobus, C.E. A mouse model for human norovirus. MBio 2013, 4, e00450-13. [Google Scholar] [CrossRef]
- Todd, K.V.; Tripp, R.A. Human Norovirus: Experimental Models of Infection. Viruses 2019, 11, 151. [Google Scholar] [CrossRef]
- Yuan, L.; Jobst, P.M.; Weiss, M. Gnotobiotic Pigs. In Gnotobiotics; Schoeb, T.R., Eaton, K.A., Eds.; Academic Press: Cambridge, MA, USA, 2017; pp. 349–368. [Google Scholar]
- Lei, S.; Twitchell, E.; Yuan, L. Pathogenesis, Immunity and the Role of Microbiome/Probiotics in Enteric Virus Infections in Humans and Animal Models. In Mechanisms Underlying Host-Microbiome Interactions in Pathophysiology of Human Diseases; Sun, J., Dudeja, P.K., Eds.; Springer: Boston, MA, USA, 2018; pp. 55–78. [Google Scholar]
- Kocher, J.; Bui, T.; Giri-Rachman, E.; Wen, K.; Li, G.; Yang, X.; Liu, F.; Tan, M.; Xia, M.; Zhong, W.; et al. Intranasal P particle vaccine provided partial cross-variant protection against human GII.4 norovirus diarrhea in gnotobiotic pigs. J. Virol. 2014, 88, 9728–9743. [Google Scholar] [CrossRef]
- Bui, T.; Kocher, J.; Li, Y.; Wen, K.; Li, G.; Liu, F.; Yang, X.; LeRoith, T.; Tan, M.; Xia, M.; et al. Median infectious dose of human norovirus GII.4 in gnotobiotic pigs is decreased by simvastatin treatment and increased by age. J. Gen. Virol. 2013, 94, 2005–2016. [Google Scholar] [CrossRef]
- Souza, M.; Costantini, V.; Azevedo, M.S.; Saif, L.J. A human norovirus-like particle vaccine adjuvanted with ISCOM or mLT induces cytokine and antibody responses and protection to the homologous GII.4 human norovirus in a gnotobiotic pig disease model. Vaccine 2007, 25, 8448–8459. [Google Scholar] [CrossRef] [PubMed]
- Cheetham, S.; Souza, M.; Meulia, T.; Grimes, S.; Han, M.G.; Saif, L.J. Pathogenesis of a genogroup II human norovirus in gnotobiotic pigs. J. Virol. 2006, 80, 10372–10381. [Google Scholar] [CrossRef]
- Atmar, R.L.; Bernstein, D.I.; Harro, C.D.; Al-Ibrahim, M.S.; Chen, W.H.; Ferreira, J.; Estes, M.K.; Graham, D.Y.; Opekun, A.R.; Richardson, C.; et al. Norovirus Vaccine against Experimental Human Norwalk Virus Illness. N. Engl. J. Med. 2011, 365, 2178–2187. [Google Scholar] [CrossRef]
- Le Guyader, F.S.; Neill, F.H.; Dubois, E.; Bon, F.; Loisy, F.; Kohli, E.; Pommepuy, M.; Atmar, R.L. A semiquantitative approach to estimate Norwalk-like virus contamination of oysters implicated in an outbreak. Int. J. Food Microbiol. 2003, 87, 107–112. [Google Scholar] [CrossRef]
- Le Guyader, F.S.; Krol, J.; Ambert-Balay, K.; Ruvoen-Clouet, N.; Desaubliaux, B.; Parnaudeau, S.; Le Saux, J.-C.; Ponge, A.; Pothier, P.; Atmar, R.L.; et al. Comprehensive Analysis of a Norovirus-Associated Gastroenteritis Outbreak, from the Environment to the Consumer. J. Clin. Microbiol. 2010, 48, 915–920. [Google Scholar] [CrossRef]
- Lowther, J.A.; Gustar, N.E.; Hartnell, R.E.; Lees, D.N. Comparison of norovirus RNA levels in outbreak-related oysters with background environmental levels. J. Food Prot. 2012, 75, 389–393. [Google Scholar] [CrossRef]
- Teunis, P.F.; Moe, C.L.; Liu, P.; Miller, S.E.; Lindesmith, L.; Baric, R.S.; Le Pendu, J.; Calderon, R.L. Norwalk virus: How infectious is it? J. Med. Virol. 2008, 80, 1468–1476. [Google Scholar] [CrossRef]
- Atmar, R.L.; Opekun, A.R.; Gilger, M.A.; Estes, M.K.; Crawford, S.E.; Neill, F.H.; Ramani, S.; Hill, H.; Ferreira, J.; Graham, D.Y. Determination of the 50% human infectious dose for Norwalk virus. J. Infect. Dis. 2014, 209, 1016–1022. [Google Scholar] [CrossRef]
- Haas, C.N.; Rose, J.B.; Gerba, C.P. Quantitative Microbial Risk Assessment; Wiley: Hoboken, NJ, USA, 2014. [Google Scholar]
- World Health Organization. Quantitative Microbial Risk Assessment: Application for Water Safety Management; World Health Organization: Geneva, Switzerland, 2016. [Google Scholar]
- Schmidt, P.J. Norovirus Dose-Response: Are Currently Available Data Informative Enough to Determine How Susceptible Humans Are to Infection from a Single Virus? Risk Anal. Off. Publ. Soc. Risk Anal. 2015, 35, 1364–1383. [Google Scholar] [CrossRef]
- Messner, M.J.; Berger, P.; Nappier, S.P. Fractional poisson--a simple dose-response model for human norovirus. Risk Anal. Off. Publ. Soc. Risk Anal. 2014, 34, 1820–1829. [Google Scholar] [CrossRef]
- Villabruna, N.; Koopmans, M.P.G.; de Graaf, M. Animals as Reservoir for Human Norovirus. Viruses 2019, 11, 478. [Google Scholar] [CrossRef]
- Frenck, R.; Bernstein, D.I.; Xia, M.; Huang, P.; Zhong, W.; Parker, S.; Dickey, M.; McNeal, M.; Jiang, X. Predicting susceptibility to norovirus GII.4 by use of a challenge model involving humans. J. Infect. Dis. 2012, 206, 1386–1393. [Google Scholar] [CrossRef]
- Lei, S.; Ryu, J.; Wen, K.; Twitchell, E.; Bui, T.; Ramesh, A.; Weiss, M.; Li, G.; Samuel, H.; Clark-Deener, S.; et al. Increased and prolonged human norovirus infection in RAG2/IL2RG deficient gnotobiotic pigs with severe combined immunodeficiency. Sci. Rep. 2016, 6, 25222. [Google Scholar] [CrossRef]
- Lei, S.; Samuel, H.; Twitchell, E.; Bui, T.; Ramesh, A.; Wen, K.; Weiss, M.; Li, G.; Yang, X.; Jiang, X.; et al. Enterobacter cloacae inhibits human norovirus infectivity in gnotobiotic pigs. Sci. Rep. 2016, 6, 25017. [Google Scholar] [CrossRef]
- Lei, S.; Ramesh, A.; Twitchell, E.; Wen, K.; Bui, T.; Weiss, M.; Yang, X.; Kocher, J.; Li, G.; Giri-Rachman, E.; et al. High Protective Efficacy of Probiotics and Rice Bran against Human Norovirus Infection and Diarrhea in Gnotobiotic Pigs. Front. Microbiol. 2016, 7, 1699. [Google Scholar] [CrossRef]
- Kageyama, T.; Kojima, S.; Shinohara, M.; Uchida, K.; Fukushi, S.; Hoshino, F.B.; Takeda, N.; Katayama, K. Broadly reactive and highly sensitive assay for Norwalk-like viruses based on real-time quantitative reverse transcription-PCR. J. Clin. Microbiol. 2003, 41, 1548–1557. [Google Scholar] [CrossRef]
- Reed, L.J.; Muench, H. A Simple Method of Estimating Fifty Percent Endpoints. Am. J. Epidemiol. 1938, 27, 493–497. [Google Scholar] [CrossRef]
- Dragstedt, C.A.; Lang, V.F. Respiratory Stimulants in Acute Cocaine Poisoning in Rabbits. J. Pharmacol. Exp. Ther. 1928, 32, 215. [Google Scholar]
- James, G.S. The Behrens-Fisher Distribution and Weighted Means. J. R. Stat. Soc. Ser. B (Methodol.) 1959, 21, 73–90. [Google Scholar] [CrossRef]
- Siev, D. Non-Parametric Estimation of Median Effective Dose. Available online: https://www.aphis.usda.gov/animal_health/vet_biologics/publications/STATWI0001.pdf (accessed on 9 May 2019).
- Han, N.; Chen, Z.; Wan, H.; Huang, G.; Li, J.; Jin, B.R. A rapid method for titration of ascovirus infectivity. J. Virol. Methods 2018, 255, 101–106. [Google Scholar] [CrossRef]
- Ramakrishnan, M.A. Determination of 50% endpoint titer using a simple formula. World J. Virol. 2016, 5, 85–86. [Google Scholar] [CrossRef]
- Sudeep, A.B.; Vyas, P.B.; Parashar, D.; Shil, P. Differential susceptibility & replication potential of Vero E6, BHK-21, RD, A-549, C6/36 cells & Aedes aegypti mosquitoes to three strains of chikungunya virus. Indian J. Med. Res. 2019, 149, 771–777. [Google Scholar] [CrossRef]
- Spackman, E.; Malladi, S.; Ssematimba, A.; Stephens, C.B. Assessment of replicate numbers for titrating avian influenza virus using dose-response models. J. Vet. Diagn. Investig. 2019, 31, 616–619. [Google Scholar] [CrossRef]
- Elikaei, A.; Hosseini, S.M.; Sharifi, Z. Inactivation of model viruses and bacteria in human fresh frozen plasma using riboflavin and long wave ultraviolet rays. Iran. J. Microbiol. 2017, 9, 50–54. [Google Scholar]
- Zhao, F.R.; Xie, Y.L.; Liu, Z.Z.; Shao, J.J.; Li, S.F.; Zhang, Y.G.; Chang, H.Y. Lithium chloride inhibits early stages of foot-and-mouth disease virus (FMDV) replication in vitro. J. Med. Virol. 2017, 89, 2041–2046. [Google Scholar] [CrossRef]
- USDA. Statistical Methods (STATWIs). Available online: https://www.aphis.usda.gov/aphis/ourfocus/animalhealth/veterinary-biologics/biologics-regulations-and-guidance/ct_vb_statwi (accessed on 23 June 2020).
- Weir, M.H.; Mitchell, J.; Flynn, W.; Pope, J.M. Development of a microbial dose response visualization and modelling application for QMRA modelers and educators. Environ. Model. Softw. 2017, 88, 74–83. [Google Scholar] [CrossRef]
- Spouge, J.L. Infectious Dose or Dilution (ID50) Server. Available online: https://www.ncbi.nlm.nih.gov/CBBresearch/Spouge/html_ncbi/html/id50/id50.cgi (accessed on 29 March 2019).
- Spouge, J.L. Statistical analysis of sparse infection data and its implications for retroviral treatment trials in primates. Proc. Natl. Acad. Sci. USA 1992, 89, 7581–7585. [Google Scholar] [CrossRef]
- Schmidt, P.J.; Pintar, K.D.M.; Fazil, A.M.; Topp, E. Harnessing the Theoretical Foundations of the Exponential and Beta-Poisson Dose-Response Models to Quantify Parameter Uncertainty Using Markov Chain Monte Carlo. Risk Anal. 2013, 33, 1677–1693. [Google Scholar] [CrossRef]
- Green, K. Caliciviridae: The Noroviruses. In Fields Virology, 6th ed.; Knipe, D.M., Howley, P.M., Eds.; Wolters Kluwer Health/Lippincott Williams & Wilkins: Philadelphia, PA, USA, 2013; Volume 1, pp. 582–608. [Google Scholar]
- Kapikian, A.Z.; Wyatt, R.G.; Dolin, R.; Thornhill, T.S.; Kalica, A.R.; Chanock, R.M. Visualization by immune electron microscopy of a 27-nm particle associated with acute infectious nonbacterial gastroenteritis. J. Virol. 1972, 10, 1075–1081. [Google Scholar] [CrossRef]
- Otto, P.H.; Clarke, I.N.; Lambden, P.R.; Salim, O.; Reetz, J.; Liebler-Tenorio, E.M. Infection of Calves with Bovine Norovirus GIII.1 Strain Jena Virus: An Experimental Model To Study the Pathogenesis of Norovirus Infection. J. Virol. 2011, 85, 12013. [Google Scholar] [CrossRef]
- Santiana, M.; Ghosh, S.; Ho, B.A.; Rajasekaran, V.; Du, W.-L.; Mutsafi, Y.; De Jésus-Diaz, D.A.; Sosnovtsev, S.V.; Levenson, E.A.; Parra, G.I.; et al. Vesicle-Cloaked Virus Clusters Are Optimal Units for Inter-organismal Viral Transmission. Cell Host Microbe 2018, 24, 208–220.e208. [Google Scholar] [CrossRef]
- Schmidt, P.J.; Emelko, M.B.; Thompson, M.E. Recognizing Structural Nonidentifiability: When Experiments Do Not Provide Information About Important Parameters and Misleading Models Can Still Have Great Fit. Risk Anal. 2020, 40, 352–369. [Google Scholar] [CrossRef]
- Tu, E.T.-V.; Bull, R.A.; Greening, G.E.; Hewitt, J.; Lyon, M.J.; Marshall, J.A.; McIver, C.J.; Rawlinson, W.D.; White, P.A. Epidemics of Gastroenteritis during 2006 Were Associated with the Spread of Norovirus GII.4 Variants 2006a and 2006b. Clin. Infect. Dis. 2008, 46, 413–420. [Google Scholar] [CrossRef]
- Chen, S.Y.; Feng, Y.; Chao, H.C.; Lai, M.W.; Huang, W.L.; Lin, C.Y.; Tsai, C.N.; Chen, C.L.; Chiu, C.H. Emergence in Taiwan of novel norovirus GII.4 variants causing acute gastroenteritis and intestinal haemorrhage in children. J. Med. Microbiol. 2015, 64, 544–550. [Google Scholar] [CrossRef]
- Kumazaki, M.; Usuku, S. Genetic Analysis of Norovirus GII.4 Variant Strains Detected in Outbreaks of Gastroenteritis in Yokohama, Japan, from the 2006-2007 to the 2013-2014 Seasons. PLoS ONE 2015, 10, e0142568. [Google Scholar] [CrossRef]
- Isakbaeva, E.T.; Widdowson, M.-A.; Beard, R.S.; Bulens, S.N.; Mullins, J.; Monroe, S.S.; Bresee, J.; Sassano, P.; Cramer, E.H.; Glass, R.I. Norovirus transmission on cruise ship. Emerg. Infect. Dis. 2005, 11, 154–158. [Google Scholar] [CrossRef]
- Bull, R.A.; Tu, E.T.V.; McIver, C.J.; Rawlinson, W.D.; White, P.A. Emergence of a New Norovirus Genotype II.4 Variant Associated with Global Outbreaks of Gastroenteritis. J. Clin. Microbiol. 2006, 44, 327–333. [Google Scholar] [CrossRef]
- Johnston, C.P.; Qiu, H.; Ticehurst, J.R.; Dickson, C.; Rosenbaum, P.; Lawson, P.; Stokes, A.B.; Lowenstein, C.J.; Kaminsky, M.; Cosgrove, S.E.; et al. Outbreak Management and Implications of a Nosocomial Norovirus Outbreak. Clin. Infect. Dis. 2007, 45, 534–540. [Google Scholar] [CrossRef]
- Lei, S.; Twitchell, E.L.; Ramesh, A.K.; Bui, T.; Majette, E.; Tin, C.M.; Avery, R.; Arango-Argoty, G.; Zhang, L.; Becker-Dreps, S.; et al. Enhanced GII.4 human norovirus infection in gnotobiotic pigs transplanted with a human gut microbiota. J. Gen. Virol. 2019. [Google Scholar] [CrossRef]
- Armitage, P.; Allen, I. Methods of estimating the LD 50 in quantal response data. J. Hyg. (Lond.) 1950, 48, 298–322. [Google Scholar] [CrossRef]
- Grimes, S. A Basic Laboratory Manual for the Small-Scale Production and Testing of I-2 Newcastle Disease Vaccine. In FAO Corporate Document Repository; FAO, Ed.; RAP Publications: Rome, Italy, 2002. [Google Scholar]
- Miller, R.G. Nonparametric Estimators of the Mean Tolerance in Bioassay. Biometrika 1973, 60, 535–542. [Google Scholar] [CrossRef]
- Miller, J.; Ulrich, R. On the analysis of psychometric functions: The Spearman-Kärber method. Percept. Psychophys. 2001, 63, 1399–1420. [Google Scholar] [CrossRef]
- Haas, C.N. Conditional dose-response relationships for microorganisms: Development and application. Risk Anal. Off. Publ. Soc. Risk Anal. 2002, 22, 455–463. [Google Scholar] [CrossRef]
- Roederer, M. Parsimonious Determination of the Optimal Infectious Dose of a Pathogen for Nonhuman Primate Models. PLoS Pathog. 2015, 11, e1005100. [Google Scholar] [CrossRef]
| Dose Group | # of Viral Genome Copies | n | Virus Shedding | Diarrhea | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| (%) a | Mean Duration Days (SEM) c–e | AUC d,f | Mean Onset Day (SEM) c,d | (%) b | Mean Duration Days (SEM) c–e | AUC d,f | Mean Onset Day (SEM) c,d | |||
| 1 | 2 × 106 | 4 | 4 (100%) | 2.5 (0.6) BC | 9506 B | 2 (0.4) B | 4 (100%) | 5.0 (0.3) A | 9.06 A | 1.5 (0.5) D |
| 2 | 4 × 105 | 4 | 4 (100%) | 1.3 (0.3) CD | 5232 B | 4 (1.2) AB | 3 (75%) | 1.3 (0.9) ABC | 7.04 A | 5.5 (1.2) ABC |
| 3 | 2 × 105 | 4 | 4 (100%) | 6.3 (0.5) A | 126774 A | 1.3 (0.3) B | 4 (100%) | 4.0 (0.9) AB | 9.31 A | 2.8 (0.5) CD |
| 4 | 8 × 104 | 6 | 6 (100%) | 2.8 (0.5) B | 13495 B | 1.5 (0.3) B | 6 (100%) | 3.8 (1) AB | 7.46 A | 3.2 (0.9) CD |
| 5 | 2 × 104 | 3 | 2 (67%) | 1 (0.6) B | 93 B | 3.3 (2.3) AB | 0 (0%) | 0.0 (0) ABC | 1.50 B | 6.3 (1.7) AB |
| 6 | 3.2 × 103 | 3 | 2 (67%) | 1.3 (0.9) BCD | 2667 B | 4 (2.1) AB | 1 (33%) | 1.0 (0) BC | 3.17 AB | 5 (2.1) BC |
| 7 | 8 × 102 | 4 | 1 (25%) | 0.5 (0.5) D | 2972 B | 6.8 (1.3) A | 0 (0%) | 0.0 (0) C | 2.75 AB | 8 (0) A |
| Method | Log10ID50 | ID50 | Log10DD50 | DD50 |
|---|---|---|---|---|
| Reed-Muench | ||||
| Hand calculation “skrmdb” R script | 3.40 | 2.51 × 103 | 4.58 | 3.80 × 104 |
| Dragstedt-Behrens “skrmdb” R script | 3.39 | 2.45 × 103 | 4.58 | 3.80 × 104 |
| Spearman-Karber Hand calculation “skrmdb” R script online calculator | 3.52 | 3.31 × 103 | 4.49 | 3.09 × 104 |
| Logistic Regression online calculator | 3.40 | 2.51 × 103 | 4.34 | 2.18 × 104 |
| Exponentiala R script | 3.76 | 5.75 × 103 | 4.76 | 5.75 × 104 |
| Approximate Beta-Poissona | ||||
| R script | 3.41 | 2.57 × 103 | 4.33 | 2.13 × 104 |
| Host | Age | Challenge Dose | n | Virus Shedding (%) | Mean Duration Days (Range) c | Peak Virus Shedding Day (PID) |
|---|---|---|---|---|---|---|
| Human a | 19–48 years | 5 × 104 | 23 | 70 | 5.2 (2–30) | 3 |
| Human b | 18–50 years | 4.4 × 103 | 34 | 76.5 | - | - |
| Human b | 18–50 years | 4.4 × 103 | 48 | 62.5 | - | - |
| Gn pig | 33–34 days | 2 × 104 | 3 | 67 | 1.0 (1–2) | 2 |
| Gn pig | 33–34 days | 8 × 104 | 6 | 100 | 2.8 (2–4) | 3 |
| Gn pig | 33–34 days | 2 × 105 | 4 | 100 | 6.3 (5–7) | 4 |
| Exponential Model | Approximate Beta Poisson Model | ||||||||
|---|---|---|---|---|---|---|---|---|---|
| r | AIC a | Chi-Squared p-Value | Minimized Deviance | α | N50 | AIC a | Chi-Squared p-Value | Minimized Deviance | |
| ID50 | 1.20 × 10−4 | 10.71 | 0.71 | 3.74 | 0.998 | 2572 | 10.68 | 0.89 | 1.71 |
| DD50 | 1.20 × 10−5 | 21.46 | 0.0 b | 16.09 | 0.928 | 21,340 | 17.92 | 0.06 | 10.5 |
| Norovirus Variant | Optimal Dose (Viral Genome Copies) | ID50 Dose | Method of ID50 Determination | |
|---|---|---|---|---|
| GII.4/2003 Cin-2 | 2.0 × 105 | 2.57 × 103 | Approximate Beta-Poisson | This study |
| GII.4/2006b | 6.43 × 105 | 6.43 × 104 | Reed-Muench | Bui et al. 2013 [18]; Kocher et al. 2014 [17]; Lei et al. 2016 [16] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ramesh, A.K.; Parreño, V.; Schmidt, P.J.; Lei, S.; Zhong, W.; Jiang, X.; Emelko, M.B.; Yuan, L. Evaluation of the 50% Infectious Dose of Human Norovirus Cin-2 in Gnotobiotic Pigs: A Comparison of Classical and Contemporary Methods for Endpoint Estimation. Viruses 2020, 12, 955. https://doi.org/10.3390/v12090955
Ramesh AK, Parreño V, Schmidt PJ, Lei S, Zhong W, Jiang X, Emelko MB, Yuan L. Evaluation of the 50% Infectious Dose of Human Norovirus Cin-2 in Gnotobiotic Pigs: A Comparison of Classical and Contemporary Methods for Endpoint Estimation. Viruses. 2020; 12(9):955. https://doi.org/10.3390/v12090955
Chicago/Turabian StyleRamesh, Ashwin K., Viviana Parreño, Philip J. Schmidt, Shaohua Lei, Weiming Zhong, Xi Jiang, Monica B. Emelko, and Lijuan Yuan. 2020. "Evaluation of the 50% Infectious Dose of Human Norovirus Cin-2 in Gnotobiotic Pigs: A Comparison of Classical and Contemporary Methods for Endpoint Estimation" Viruses 12, no. 9: 955. https://doi.org/10.3390/v12090955
APA StyleRamesh, A. K., Parreño, V., Schmidt, P. J., Lei, S., Zhong, W., Jiang, X., Emelko, M. B., & Yuan, L. (2020). Evaluation of the 50% Infectious Dose of Human Norovirus Cin-2 in Gnotobiotic Pigs: A Comparison of Classical and Contemporary Methods for Endpoint Estimation. Viruses, 12(9), 955. https://doi.org/10.3390/v12090955







