The Anticoagulant Nafamostat Potently Inhibits SARS-CoV-2 S Protein-Mediated Fusion in a Cell Fusion Assay System and Viral Infection In Vitro in a Cell-Type-Dependent Manner
Abstract
1. Introduction
2. Materials and Methods
2.1. Protease Inhibitors and Anti-Coagulants
2.2. Cell Lines and Transient Transfection
2.3. Construction of Expression Vectors
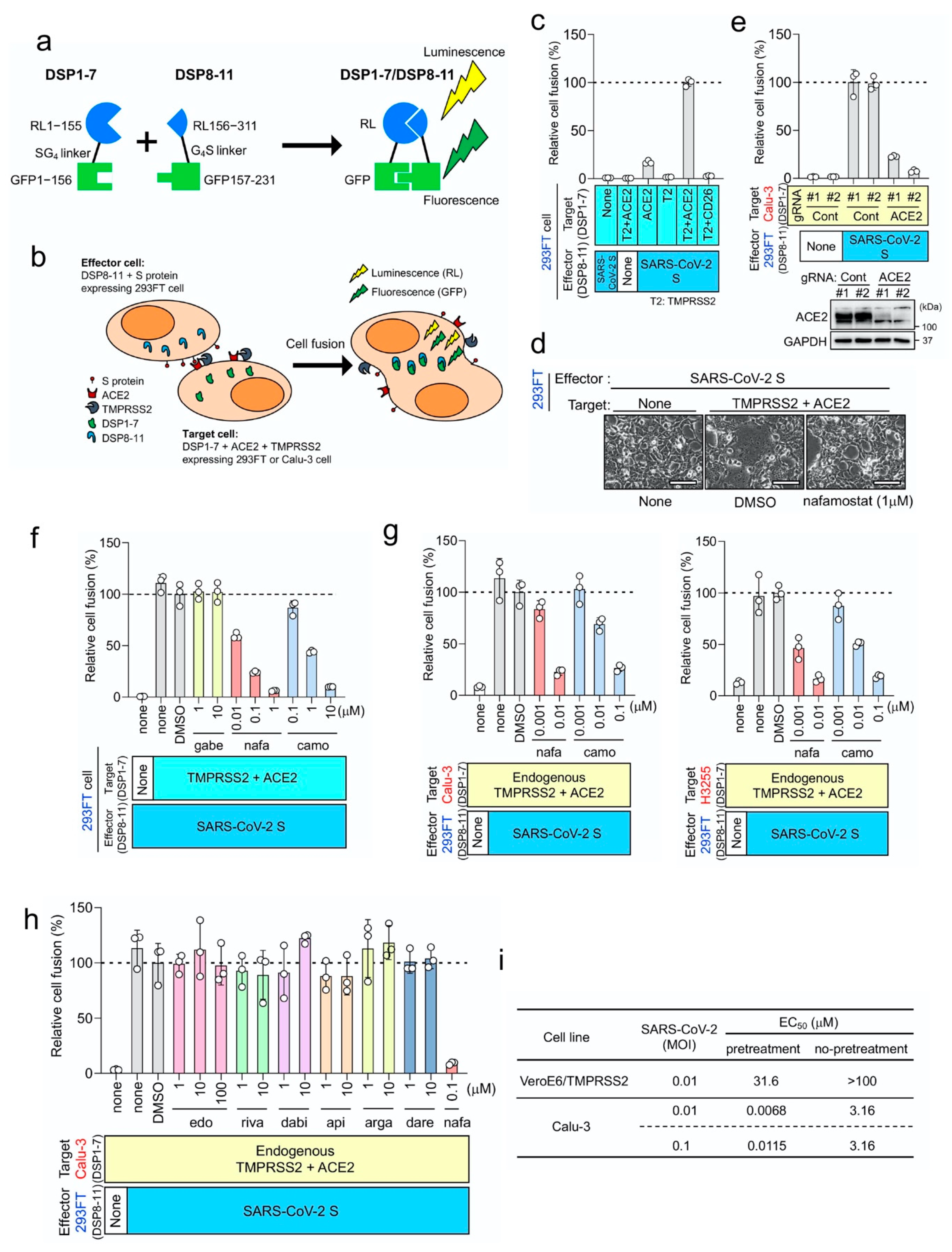
2.4. DSP Assay to Monitor Membrane Fusion
2.5. Western Blotting
2.6. Isolation of SARS-CoV-2
2.7. SARS-CoV-2 Infection Assay
3. Results
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Zhou, P.; Yang, X.L.; Wang, X.G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.R.; Zhu, Y.; Li, B.; Huang, C.L.; et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 2020, 579, 270–273. [Google Scholar] [CrossRef] [PubMed]
- Shen, L.W.; Mao, H.J.; Wu, Y.L.; Tanaka, Y.; Zhang, W. TMPRSS2: A potential target for treatment of influenza virus and coronavirus infections. Biochimie 2017, 142, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Shang, J.; Wan, Y.; Luo, C.; Ye, G.; Geng, Q.; Auerbach, A.; Li, F. Cell entry mechanisms of SARS-CoV-2. Proc. Natl. Acad. Sci. USA 2020, 117, 11727–11734. [Google Scholar] [CrossRef] [PubMed]
- Zhou, L.; Niu, Z.; Jiang, X.; Zhang, Z.; Zheng, Y.; Wang, Z.; Zhu, Y.; Wang, X.; Sun, Q. Systemic Analysis of Tissue Cells Potentially Vulnerable to SARS-CoV-2 Infection by the Protein-Proofed Single-Cell RNA Profiling of ACE2, TMPRSS2 and Furin Proteases. 2020. Available online: https://www.biorxiv.org/content/10.1101/2020.04.06.028522v1 (accessed on 7 June 2020).
- Sungnak, W.; Huang, N.; Bécavin, C.; Berg, M.; Queen, R.; Litvinukova, M.; Talavera-López, C.; Maatz, H.; Reichart, D.; Sampaziotis, F.; et al. SARS-CoV-2 entry factors are highly expressed in nasal epithelial cells together with innate immune genes. Nat. Med. 2020, 26, 681–687. [Google Scholar] [CrossRef] [PubMed]
- Ziegler, C.G.K.; Allon, S.J.; Nyquist, S.K.; Mbano, I.M.; Miao, V.N.; Tzouanas, C.N.; Cao, Y.; Yousif, A.S.; Bals, J.; Hauser, B.M.; et al. SARS-CoV-2 Receptor ACE2 Is an Interferon-Stimulated Gene in Human Airway Epithelial Cells and Is Detected in Specific Cell Subsets across Tissues. Cell 2020, 181, 1016–1035. [Google Scholar] [CrossRef] [PubMed]
- Iwata-Yoshikawa, N.; Okamura, T.; Shimizu, Y.; Hasegawa, H.; Takeda, M.; Nagata, N. TMPRSS2 Contributes to Virus Spread and Immunopathology in the Airways of Murine Models after Coronavirus Infection. J. Virol. 2019, 93. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, M.; Matsuyama, S.; Li, X.; Takeda, M.; Kawaguchi, Y.; Inoue, J.I.; Matsuda, Z. Identification of Nafamostat as a Potent Inhibitor of Middle East Respiratory Syndrome Coronavirus S Protein-Mediated Membrane Fusion Using the Split-Protein-Based Cell-Cell Fusion Assay. Antimicrob. Agents Chemother. 2016, 60, 6532–6539. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Li, X.; Nakane, S.; Liu, S.; Ishikawa, H.; Iwamoto, A.; Matsuda, Z. Co-expression of foreign proteins tethered to HIV-1 envelope glycoprotein on the cell surface by introducing an intervening second membrane-spanning domain. PLoS ONE 2014, 9, e96790. [Google Scholar] [CrossRef] [PubMed]
- Kitamura, T.; Koshino, Y.; Shibata, F.; Oki, T.; Nakajima, H.; Nosaka, T.; Kumagai, H. Retrovirus-mediated gene transfer and expression cloning: Powerful tools in functional genomics. Exp. Hematol. 2003, 31, 1007–1014. [Google Scholar] [CrossRef]
- Yamamoto, M.; Abe, C.; Wakinaga, S.; Sakane, K.; Yumiketa, Y.; Taguchi, Y.; Matsumura, T.; Ishikawa, K.; Fujimoto, J.; Semba, K.; et al. TRAF6 maintains mammary stem cells and promotes pregnancy-induced mammary epithelial cell expansion. Commun. Biol. 2019, 2, 292. [Google Scholar] [CrossRef] [PubMed]
- Matsuyama, S.; Nao, N.; Shirato, K.; Kawase, M.; Saito, S.; Takayama, I.; Nagata, N.; Sekizuka, T.; Katoh, H.; Kato, F.; et al. Enhanced isolation of SARS-CoV-2 by TMPRSS2-expressing cells. Proc. Natl. Acad. Sci. USA 2020, 117, 7001–7003. [Google Scholar] [CrossRef] [PubMed]
- Ramakrishnan, M.A. Determination of 50% endpoint titer using a simple formula. World J. Virol. 2016, 5, 85–86. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Krüger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.H.; Nitsche, A.; et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell 2020, 181, 271–280. [Google Scholar] [CrossRef] [PubMed]
- Rensi, S.; Altman, R.B.; Liu, T.; Lo, Y.C.; McInnes, G.; Derry, A.; Keys, A. Homology Modeling of TMPRSS2 Yields Candidate Drugs That May Inhibit Entry of SARS-CoV-2 into Human Cells. ChemRxiv 2020. [Google Scholar] [CrossRef]
- Kim, J.; Zhang, J.; Cha, Y.; Kolitz, S.; Funt, J.; Chong, R.E.; Barrett, S.; Zeskind, B.; Kusko, R.; Kaufman, H. Advanced Bioinformatics Rapidly Identifies Existing Therapeutics for Patients with Coronavirus Disease—2019 (COVID-19). ChemRxiv 2020. [Google Scholar] [CrossRef]
- Matsuyama, S.; Kawase, M.; Nao, N.; Shirato, K.; Ujike, M.; Kamitani, W.; Shimojima, M.; Fukushi, S. The inhaled corticosteroid ciclesonide blocks coronavirus RNA replication by targeting viral NSP15. BioRxiv 2020. [Google Scholar] [CrossRef]
- Kawase, M.; Shirato, K.; Van der Hoek, L.; Taguchi, F.; Matsuyama, S. Simultaneous treatment of human bronchial epithelial cells with serine and cysteine protease inhibitors prevents severe acute respiratory syndrome coronavirus entry. J. Virol. 2012, 86, 6537–6545. [Google Scholar] [CrossRef] [PubMed]
- Kyoto Encyclopedia of Genes and Genomes (KEGG) MEDICUS. Available online: https://www.kegg.jp/medicus-bin/japic_med?japic_code=00048710 (accessed on 7 June 2020). (In Japanese).
- Wang, M.; Cao, R.; Zhang, L.; Yang, X.; Liu, J.; Xu, M.; Shi, Z.; Hu, Z.; Zhong, W.; Xiao, G. Remdesivir and chloroquine effectively inhibit the recently emerged novel coronavirus (2019-nCoV) in vitro. Cell Res. 2020, 30, 269–271. [Google Scholar] [CrossRef] [PubMed]
- Esumi, Y.; Takaichi, M.; Washino, T.; Jin, Y.; Kakurai, Y. Pharmacokinetic studies of FUT-175 (Nafamostat mesilate) (6)-blood level profiles, distribution, metabolism and excretion in rats after constant-rate infusion. Clin. Rep. 1984, 15, 4050–4066. [Google Scholar]
- Hoffmann, M.; Schroeder, S.; Kleine-Weber, H.; Müller, M.A.; Drosten, C.; Pöhlmann, S. Nafamostat mesylate blocks activation of SARS-CoV-2: New treatment option for COVID-19. Antimicrob. Agents Chemother. 2020. [Google Scholar] [CrossRef] [PubMed]
- Tang, N.; Bai, H.; Chen, X.; Gong, J.; Li, D.; Sun, Z. Anticoagulant treatment is associated with decreased mortality in severe coronavirus disease 2019 patients with coagulopathy. J. Thromb. Haemost. 2020. [Google Scholar] [CrossRef] [PubMed]
- Asakura, H.; Ogawa, H. Potential of Heparin and Nafamostat Combination Therapy for COVID-19. J. Thromb. Haemost. 2020. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.L.; Wang, S.D.; Zeng, Z.Y.; Lin, K.J.; Kao, S.T.; Tani, T.; Yu, C.K.; Wang, J.Y. Serine protease inhibitors nafamostat mesilate and gabexate mesilate attenuate allergen-induced airway inflammation and eosinophilia in a murine model of asthma. J. Allergy Clin. Immunol. 2006, 118, 105–112. [Google Scholar] [CrossRef] [PubMed]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yamamoto, M.; Kiso, M.; Sakai-Tagawa, Y.; Iwatsuki-Horimoto, K.; Imai, M.; Takeda, M.; Kinoshita, N.; Ohmagari, N.; Gohda, J.; Semba, K.; et al. The Anticoagulant Nafamostat Potently Inhibits SARS-CoV-2 S Protein-Mediated Fusion in a Cell Fusion Assay System and Viral Infection In Vitro in a Cell-Type-Dependent Manner. Viruses 2020, 12, 629. https://doi.org/10.3390/v12060629
Yamamoto M, Kiso M, Sakai-Tagawa Y, Iwatsuki-Horimoto K, Imai M, Takeda M, Kinoshita N, Ohmagari N, Gohda J, Semba K, et al. The Anticoagulant Nafamostat Potently Inhibits SARS-CoV-2 S Protein-Mediated Fusion in a Cell Fusion Assay System and Viral Infection In Vitro in a Cell-Type-Dependent Manner. Viruses. 2020; 12(6):629. https://doi.org/10.3390/v12060629
Chicago/Turabian StyleYamamoto, Mizuki, Maki Kiso, Yuko Sakai-Tagawa, Kiyoko Iwatsuki-Horimoto, Masaki Imai, Makoto Takeda, Noriko Kinoshita, Norio Ohmagari, Jin Gohda, Kentaro Semba, and et al. 2020. "The Anticoagulant Nafamostat Potently Inhibits SARS-CoV-2 S Protein-Mediated Fusion in a Cell Fusion Assay System and Viral Infection In Vitro in a Cell-Type-Dependent Manner" Viruses 12, no. 6: 629. https://doi.org/10.3390/v12060629
APA StyleYamamoto, M., Kiso, M., Sakai-Tagawa, Y., Iwatsuki-Horimoto, K., Imai, M., Takeda, M., Kinoshita, N., Ohmagari, N., Gohda, J., Semba, K., Matsuda, Z., Kawaguchi, Y., Kawaoka, Y., & Inoue, J.-i. (2020). The Anticoagulant Nafamostat Potently Inhibits SARS-CoV-2 S Protein-Mediated Fusion in a Cell Fusion Assay System and Viral Infection In Vitro in a Cell-Type-Dependent Manner. Viruses, 12(6), 629. https://doi.org/10.3390/v12060629




