Fc-Based Recombinant Henipavirus Vaccines Elicit Broad Neutralizing Antibody Responses in Mice
Abstract
1. Introduction
2. Materials and Methods
2.1. Phylogenetic Analysis of the G Proteins
2.2. Protein Expression and Purification
2.2.1. G Proteins of HNVs
2.2.2. Monoclonal Antibody M102.4
2.2.3. Fc-Fusion Proteins
2.3. Coupling of Purified G Glycoproteins to Microspheres
2.4. Multiplex Microsphere Receptor Binding Assay
2.5. Animal Immunization
2.6. Multiplex Microsphere Receptor Inhibition Assay
2.7. Enzyme-Linked Immunosorbent Assay (ELISA)
2.8. Pseudotyped Virus Packaging
2.9. Pseudovirus Neutralization Assay
2.10. Biolayer Interferometry Assay
3. Results
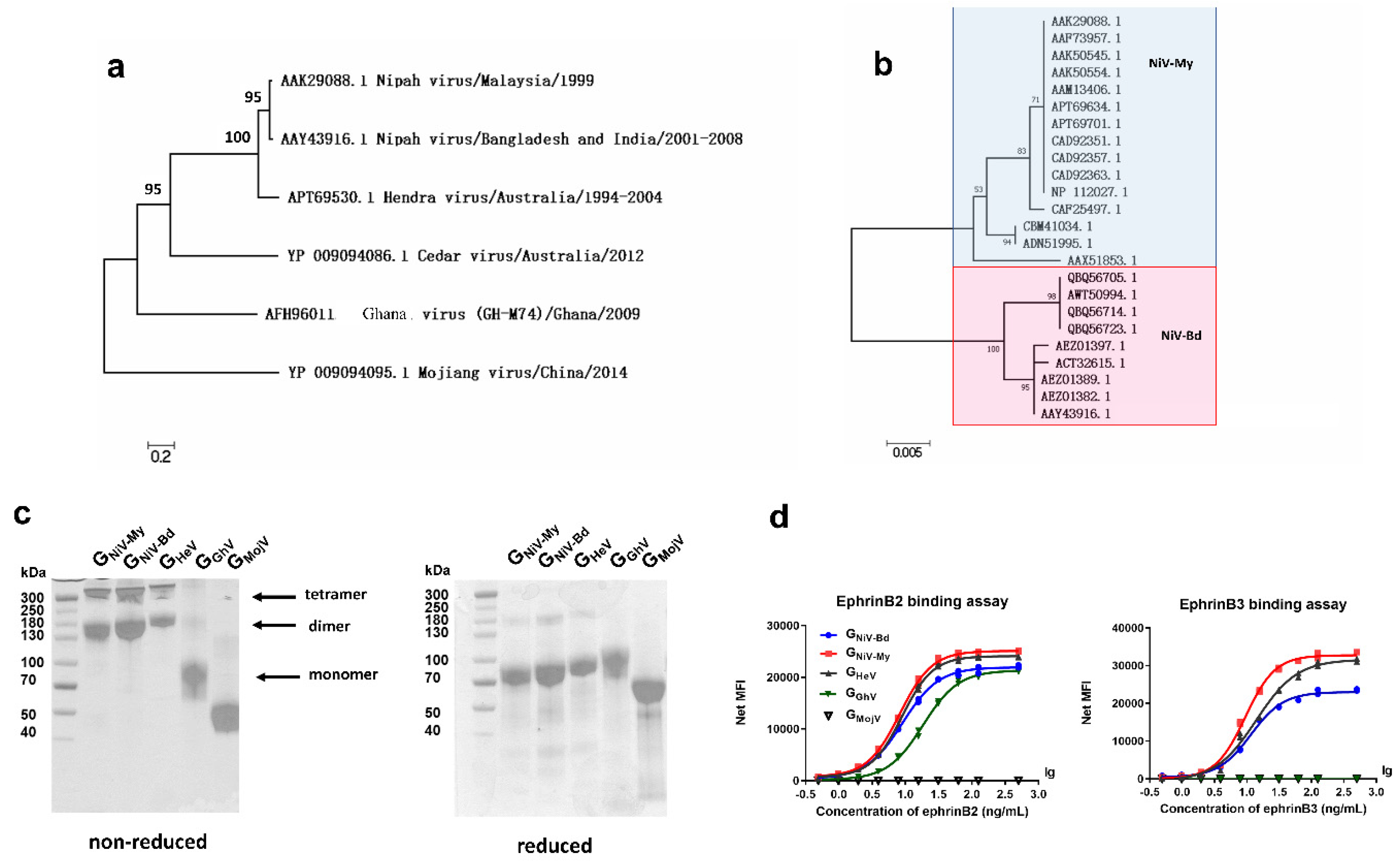
3.1. Phylogenetic Analysis and Expression of G Proteins
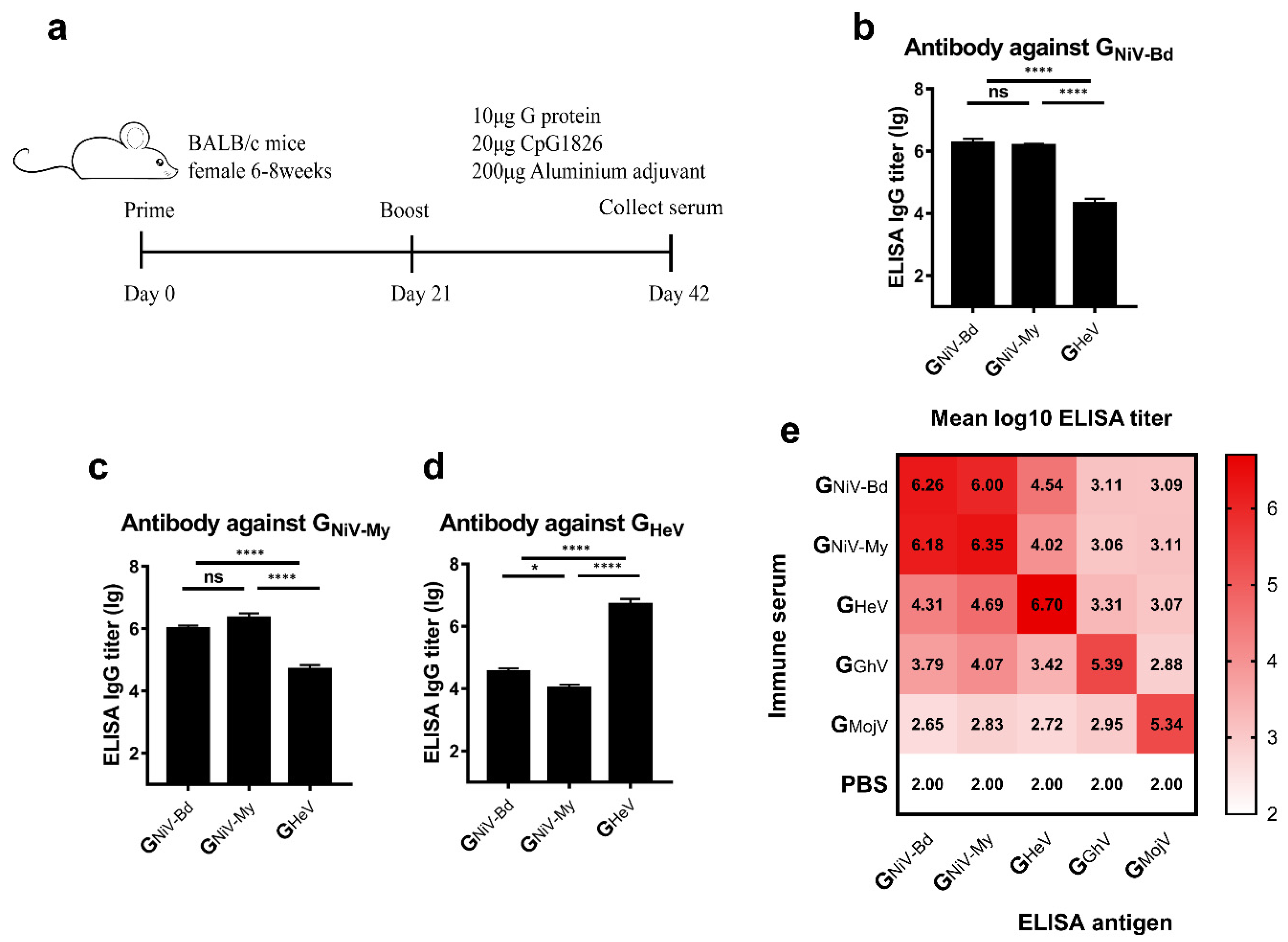
3.2. HNV-G Proteins Elicited Limited a Cross-Reactive Specific Antibody Response
3.3. HNV-G Proteins Elicited a Limited Cross-Reactive Neutralizing Antibody Response
3.3.1. Receptor Competitive Inhibition Assay
3.3.2. Pseudovirus Neutralizing Assay
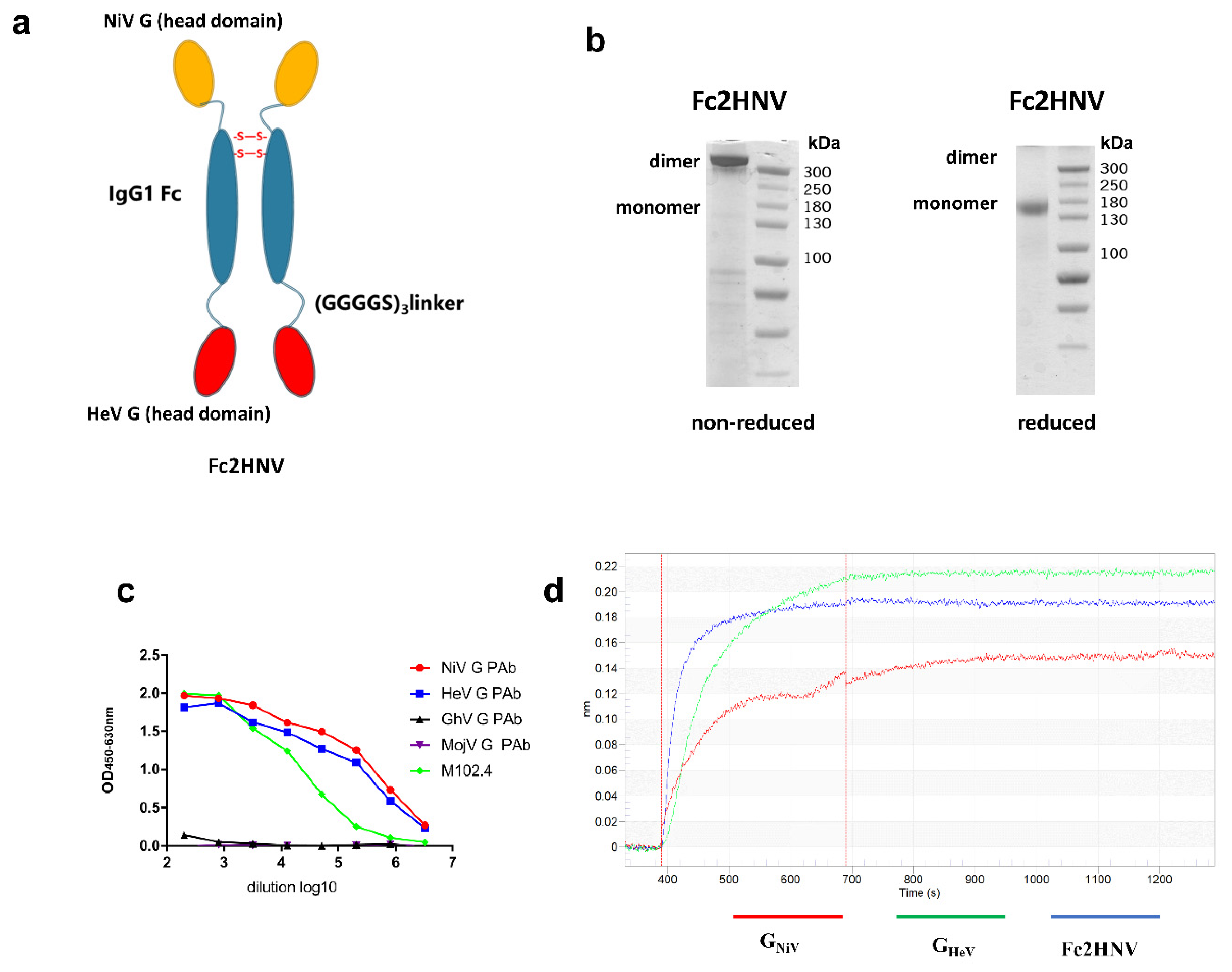
3.4. Construction and Production of Bivalent Fc-Fusion Protein
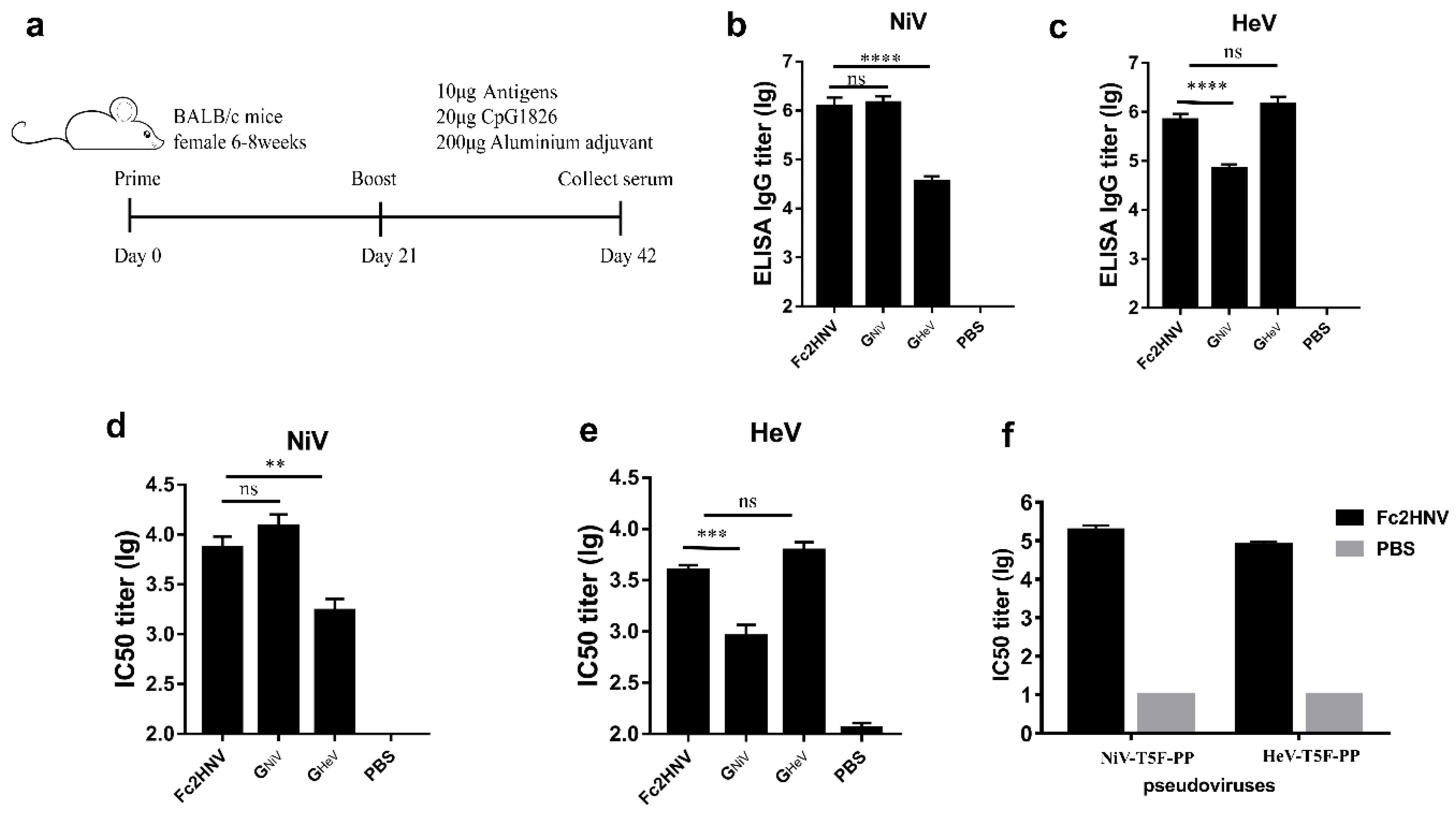
3.5. Bivalent Fc-Fusion Antigens Elicit a Broad Antibody Response against NiV and HeV in Mice
3.6. Tetravalent Fc-Fusion Protein Triggers Broad Antibody Response against HNVs
4. Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Amarasinghe, G.K.; Bao, Y.; Basler, C.F.; Bavari, S.; Beer, M.; Bejerman, N.; Blasdell, K.R.; Bochnowski, A.; Briese, T.; Bukreyev, A.; et al. Taxonomy of the order Mononegavirales: Update 2017. Arch. Virol. 2017, 162, 2493–2504. [Google Scholar] [CrossRef] [PubMed]
- Rahman, S.A.; Hassan, S.S.; Olival, K.J.; Mohamed, M.; Chang, L.Y.; Hassan, L.; Saad, N.M.; Shohaimi, S.A.; Mamat, Z.C.; Naim, M.S.; et al. Characterization of Nipah virus from naturally infected Pteropus vampyrus bats, Malaysia. Emerg. Infect. Dis. 2010, 16, 1990–1993. [Google Scholar] [CrossRef] [PubMed]
- Chua, K.B.; Koh, C.L.; Hooi, P.S.; Wee, K.F.; Khong, J.H.; Chua, B.H.; Chan, Y.P.; Lim, M.E.; Lam, S.K. Isolation of Nipah virus from Malaysian Island flying-foxes. Microbes Infect. 2002, 4, 145–151. [Google Scholar] [CrossRef]
- Halpin, K.; Young, P.L.; Field, H.E.; Mackenzie, J.S. Isolation of Hendra virus from pteropid bats: A natural reservoir of Hendra virus. J. Gen. Virol. 2000, 81, 1927–1932. [Google Scholar] [CrossRef] [PubMed]
- Reynes, J.M.; Counor, D.; Ong, S.; Faure, C.; Seng, V.; Molia, S.; Walston, J.; Georges-Courbot, M.C.; Deubel, V.; Sarthou, J.L. Nipah virus in Lyle’s flying foxes, Cambodia. Emerg. Infect. Dis. 2005, 11, 1042–1047. [Google Scholar] [CrossRef] [PubMed]
- Gaudino, M.; Aurine, N.; Dumont, C.; Fouret, J.; Ferren, M.; Mathieu, C.; Reynard, O.; Volchkov, V.E.; Legras-Lachuer, C.; Georges-Courbot, M.C.; et al. High Pathogenicity of Nipah Virus from Pteropus lylei Fruit Bats, Cambodia. Emerg. Infect. Dis. 2020, 26, 104–113. [Google Scholar] [CrossRef]
- Sharma, V.; Kaushik, S.; Kumar, R.; Yadav, J.P.; Kaushik, S. Emerging trends of Nipah virus: A review. Rev. Med. Virol. 2018, e2010. [Google Scholar] [CrossRef]
- Mohd Nor, M.N.; Gan, C.H.; Ong, B.L. Nipah virus infection of pigs in peninsular Malaysia. Rev. Sci. Et Tech. 2000, 19, 160–165. [Google Scholar] [CrossRef]
- Arunkumar, G.; Chandni, R.; Mourya, D.T.; Singh, S.K.; Sadanandan, R.; Sudan, P.; Bhargava, B.; Nipah Investigators People and Health Study Group. Outbreak investigation of Nipah Virus Disease in Kerala, India, 2018. J. Infect. Dis. 2018. [Google Scholar] [CrossRef]
- Harcourt, B.H.; Lowe, L.; Tamin, A.; Liu, X.; Bankamp, B.; Bowden, N.; Rollin, P.E.; Comer, J.A.; Ksiazek, T.G.; Hossain, M.J.; et al. Genetic characterization of Nipah virus, Bangladesh, 2004. Emerg. Infect. Dis. 2005, 11, 1594–1597. [Google Scholar] [CrossRef]
- Mire, C.E.; Satterfield, B.A.; Geisbert, J.B.; Agans, K.N.; Borisevich, V.; Yan, L.; Chan, Y.P.; Cross, R.W.; Fenton, K.A.; Broder, C.C.; et al. Pathogenic Differences between Nipah Virus Bangladesh and Malaysia Strains in Primates: Implications for Antibody Therapy. Sci. Rep. 2016, 6, 30916. [Google Scholar] [CrossRef] [PubMed]
- Marsh, G.A.; de Jong, C.; Barr, J.A.; Tachedjian, M.; Smith, C.; Middleton, D.; Yu, M.; Todd, S.; Foord, A.J.; Haring, V.; et al. Cedar virus: A novel Henipavirus isolated from Australian bats. PLoS Pathog. 2012, 8, e1002836. [Google Scholar] [CrossRef]
- Lee, B.; Pernet, O.; Ahmed, A.A.; Zeltina, A.; Beaty, S.M.; Bowden, T.A. Molecular recognition of human ephrinB2 cell surface receptor by an emergent African henipavirus. Proc. Natl. Acad. Sci. USA 2015, 112, E2156–E2165. [Google Scholar] [CrossRef] [PubMed]
- Drexler, J.F.; Corman, V.M.; Gloza-Rausch, F.; Seebens, A.; Annan, A.; Ipsen, A.; Kruppa, T.; Muller, M.A.; Kalko, E.K.; Adu-Sarkodie, Y.; et al. Henipavirus RNA in African bats. PLoS ONE 2009, 4, e6367. [Google Scholar] [CrossRef] [PubMed]
- Wu, Z.; Yang, L.; Yang, F.; Ren, X.; Jiang, J.; Dong, J.; Sun, L.; Zhu, Y.; Zhou, H.; Jin, Q. Novel Henipa-like virus, Mojiang Paramyxovirus, in rats, China, 2012. Emerg. Infect. Dis. 2014, 20, 1064–1066. [Google Scholar] [CrossRef] [PubMed]
- Mehand, M.S.; Millett, P.; Al-Shorbaji, F.; Roth, C.; Kieny, M.P.; Murgue, B. World Health Organization Methodology to Prioritize Emerging Infectious Diseases in Need of Research and Development. Emerg. Infect. Dis. 2018, 24. [Google Scholar] [CrossRef] [PubMed]
- Thakur, N.; Bailey, D. Advances in diagnostics, vaccines and therapeutics for Nipah virus. Microbes Infect. 2019, 21, 278–286. [Google Scholar] [CrossRef]
- Satterfield, B.A.; Dawes, B.E.; Milligan, G.N. Status of vaccine research and development of vaccines for Nipah virus. Vaccine 2016, 34, 2971–2975. [Google Scholar] [CrossRef]
- Van Doremalen, N.; Lambe, T.; Sebastian, S.; Bushmaker, T.; Fischer, R.; Feldmann, F.; Haddock, E.; Letko, M.; Avanzato, V.A.; Rissanen, I.; et al. A single-dose ChAdOx1-vectored vaccine provides complete protection against Nipah Bangladesh and Malaysia in Syrian golden hamsters. PLoS Negl. Trop. Dis. 2019, 13, e0007462. [Google Scholar] [CrossRef]
- Prescott, J.; DeBuysscher, B.L.; Feldmann, F.; Gardner, D.J.; Haddock, E.; Martellaro, C.; Scott, D.; Feldmann, H. Single-dose live-attenuated vesicular stomatitis virus-based vaccine protects African green monkeys from Nipah virus disease. Vaccine 2015, 33, 2823–2829. [Google Scholar] [CrossRef]
- Weingartl, H.M.; Berhane, Y.; Caswell, J.L.; Loosmore, S.; Audonnet, J.C.; Roth, J.A.; Czub, M. Recombinant nipah virus vaccines protect pigs against challenge. J. Virol. 2006, 80, 7929–7938. [Google Scholar] [CrossRef] [PubMed]
- Pedrera, M.; Macchi, F.; McLean, R.K.; Franceschi, V.; Thakur, N.; Russo, L.; Medfai, L.; Todd, S.; Tchilian, E.Z.; Audonnet, J.C.; et al. Bovine Herpesvirus-4-Vectored Delivery of Nipah Virus Glycoproteins Enhances T Cell Immunogenicity in Pigs. Vaccines (Basel) 2020, 8, 115. [Google Scholar] [CrossRef] [PubMed]
- Mire, C.E.; Geisbert, J.B.; Agans, K.N.; Feng, Y.R.; Fenton, K.A.; Bossart, K.N.; Yan, L.; Chan, Y.P.; Broder, C.C.; Geisbert, T.W. A recombinant Hendra virus G glycoprotein subunit vaccine protects nonhuman primates against Hendra virus challenge. J. Virol. 2014, 88, 4624–4631. [Google Scholar] [CrossRef] [PubMed]
- Pallister, J.A.; Klein, R.; Arkinstall, R.; Haining, J.; Long, F.; White, J.R.; Payne, J.; Feng, Y.R.; Wang, L.F.; Broder, C.C.; et al. Vaccination of ferrets with a recombinant G glycoprotein subunit vaccine provides protection against Nipah virus disease for over 12 months. Virol. J. 2013, 10, 237. [Google Scholar] [CrossRef]
- Pallister, J.; Middleton, D.; Wang, L.F.; Klein, R.; Haining, J.; Robinson, R.; Yamada, M.; White, J.; Payne, J.; Feng, Y.R.; et al. A recombinant Hendra virus G glycoprotein-based subunit vaccine protects ferrets from lethal Hendra virus challenge. Vaccine 2011, 29, 5623–5630. [Google Scholar] [CrossRef]
- McEachern, J.A.; Bingham, J.; Crameri, G.; Green, D.J.; Hancock, T.J.; Middleton, D.; Feng, Y.R.; Broder, C.C.; Wang, L.F.; Bossart, K.N. A recombinant subunit vaccine formulation protects against lethal Nipah virus challenge in cats. Vaccine 2008, 26, 3842–3852. [Google Scholar] [CrossRef]
- Pickering, B.S.; Hardham, J.M.; Smith, G.; Weingartl, E.T.; Dominowski, P.J.; Foss, D.L.; Mwangi, D.; Broder, C.C.; Roth, J.A.; Weingartl, H.M. Protection against henipaviruses in swine requires both, cell-mediated and humoral immune response. Vaccine 2016, 34, 4777–4786. [Google Scholar] [CrossRef]
- Rissanen, I.; Ahmed, A.A.; Azarm, K.; Beaty, S.; Hong, P.; Nambulli, S.; Duprex, W.P.; Lee, B.; Bowden, T.A. Idiosyncratic Mojiang virus attachment glycoprotein directs a host-cell entry pathway distinct from genetically related henipaviruses. Nat. Commun. 2017, 8, 16060. [Google Scholar] [CrossRef]
- Bonaparte, M.I.; Dimitrov, A.S.; Bossart, K.N.; Crameri, G.; Mungall, B.A.; Bishop, K.A.; Choudhry, V.; Dimitrov, D.S.; Wang, L.F.; Eaton, B.T.; et al. Ephrin-B2 ligand is a functional receptor for Hendra virus and Nipah virus. Proc. Natl. Acad. Sci. USA 2005, 102, 10652–10657. [Google Scholar] [CrossRef]
- Negrete, O.A.; Wolf, M.C.; Aguilar, H.C.; Enterlein, S.; Wang, W.; Muhlberger, E.; Su, S.V.; Bertolotti-Ciarlet, A.; Flick, R.; Lee, B. Two key residues in ephrinB3 are critical for its use as an alternative receptor for Nipah virus. PLoS Pathog. 2006, 2, e7. [Google Scholar] [CrossRef]
- Xu, K.; Rajashankar, K.R.; Chan, Y.P.; Himanen, J.P.; Broder, C.C.; Nikolov, D.B. Host cell recognition by the henipaviruses: Crystal structures of the Nipah G attachment glycoprotein and its complex with ephrin-B3. Proc. Natl. Acad. Sci. USA 2008, 105, 9953–9958. [Google Scholar] [CrossRef] [PubMed]
- Bowden, T.A.; Aricescu, A.R.; Gilbert, R.J.; Grimes, J.M.; Jones, E.Y.; Stuart, D.I. Structural basis of Nipah and Hendra virus attachment to their cell-surface receptor ephrin-B2. Nat. Struct. Mol. Biol. 2008, 15, 567–572. [Google Scholar] [CrossRef]
- Czajkowsky, D.M.; Hu, J.; Shao, Z.; Pleass, R.J. Fc-fusion proteins: New developments and future perspectives. Embo Mol. Med. 2012, 4, 1015–1028. [Google Scholar] [CrossRef] [PubMed]
- Sockolosky, J.T.; Szoka, F.C. The neonatal Fc receptor, FcRn, as a target for drug delivery and therapy. Adv. Drug Deliv. Rev. 2015, 91, 109–124. [Google Scholar] [CrossRef] [PubMed]
- Zhao, B.; Zhang, X.; Krummenacher, C.; Song, S.; Gao, L.; Zhang, H.; Xu, M.; Feng, L.; Feng, Q.; Zeng, M.; et al. Immunization With Fc-Based Recombinant Epstein-Barr Virus gp350 Elicits Potent Neutralizing Humoral Immune Response in a BALB/c Mice Model. Front. Immunol. 2018, 9, 932. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhou, Z.; Zhu, S.L.; Zu, X.; Wang, Z.; Zhang, L.K.; Wang, W.; Xiao, G. A novel RSV F-Fc fusion protein vaccine reduces lung injury induced by respiratory syncytial virus infection. Antivir. Res. 2019, 165, 11–22. [Google Scholar] [CrossRef]
- Konduru, K.; Shurtleff, A.C.; Bradfute, S.B.; Nakamura, S.; Bavari, S.; Kaplan, G. Ebolavirus Glycoprotein Fc Fusion Protein Protects Guinea Pigs against Lethal Challenge. PLoS ONE 2016, 11, e0162446. [Google Scholar] [CrossRef]
- Lu, L.; Palaniyandi, S.; Zeng, R.; Bai, Y.; Liu, X.; Wang, Y.; Pauza, C.D.; Roopenian, D.C.; Zhu, X. A neonatal Fc receptor-targeted mucosal vaccine strategy effectively induces HIV-1 antigen-specific immunity to genital infection. J. Virol. 2011, 85, 10542–10553. [Google Scholar] [CrossRef]
- Ridgway, J.B.; Presta, L.G.; Carter, P. ‘Knobs-into-holes’ engineering of antibody CH3 domains for heavy chain heterodimerization. Protein Eng. 1996, 9, 617–621. [Google Scholar] [CrossRef]
- Jones, D.T.; Taylor, W.R.; Thornton, J.M. The rapid generation of mutation data matrices from protein sequences. Comput. Appl. Biosci. Cabios 1992, 8, 275–282. [Google Scholar] [CrossRef]
- Felsenstein, J. Confidence Limits on Phylogenies: An Approach Using the Bootstrap. Evolution 1985, 39, 783–791. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Bossart, K.N.; Crameri, G.; Dimitrov, A.S.; Mungall, B.A.; Feng, Y.R.; Patch, J.R.; Choudhary, A.; Wang, L.F.; Eaton, B.T.; Broder, C.C. Receptor binding, fusion inhibition, and induction of cross-reactive neutralizing antibodies by a soluble G glycoprotein of Hendra virus. J. Virol. 2005, 79, 6690–6702. [Google Scholar] [CrossRef] [PubMed]
- Geisbert, T.W.; Mire, C.E.; Geisbert, J.B.; Chan, Y.P.; Agans, K.N.; Feldmann, F.; Fenton, K.A.; Zhu, Z.; Dimitrov, D.S.; Scott, D.P.; et al. Therapeutic treatment of Nipah virus infection in nonhuman primates with a neutralizing human monoclonal antibody. Sci. Transl. Med. 2014, 6, 242ra282. [Google Scholar] [CrossRef] [PubMed]
- Xu, K.; Rockx, B.; Xie, Y.; DeBuysscher, B.L.; Fusco, D.L.; Zhu, Z.; Chan, Y.P.; Xu, Y.; Luu, T.; Cer, R.Z.; et al. Crystal structure of the Hendra virus attachment G glycoprotein bound to a potent cross-reactive neutralizing human monoclonal antibody. PLoS Pathog. 2013, 9, e1003684. [Google Scholar] [CrossRef] [PubMed]
- Bossart, K.N.; Geisbert, T.W.; Feldmann, H.; Zhu, Z.; Feldmann, F.; Geisbert, J.B.; Yan, L.; Feng, Y.R.; Brining, D.; Scott, D.; et al. A neutralizing human monoclonal antibody protects african green monkeys from hendra virus challenge. Sci. Transl. Med. 2011, 3, 105ra103. [Google Scholar] [CrossRef]
- Bossart, K.N.; Zhu, Z.; Middleton, D.; Klippel, J.; Crameri, G.; Bingham, J.; McEachern, J.A.; Green, D.; Hancock, T.J.; Chan, Y.P.; et al. A neutralizing human monoclonal antibody protects against lethal disease in a new ferret model of acute nipah virus infection. Plos Pathog. 2009, 5, e1000642. [Google Scholar] [CrossRef]
- Palomares, K.; Vigant, F.; Van Handel, B.; Pernet, O.; Chikere, K.; Hong, P.; Sherman, S.P.; Patterson, M.; An, D.S.; Lowry, W.E.; et al. Nipah virus envelope-pseudotyped lentiviruses efficiently target ephrinB2-positive stem cell populations in vitro and bypass the liver sink when administered in vivo. J. Virol. 2013, 87, 2094–2108. [Google Scholar] [CrossRef]
- Connor, R.I.; Chen, B.K.; Choe, S.; Landau, N.R. Vpr is required for efficient replication of human immunodeficiency virus type-1 in mononuclear phagocytes. Virology 1995, 206, 935–944. [Google Scholar] [CrossRef]
- Bossart, K.N.; McEachern, J.A.; Hickey, A.C.; Choudhry, V.; Dimitrov, D.S.; Eaton, B.T.; Wang, L.F. Neutralization assays for differential henipavirus serology using Bio-Plex protein array systems. J. Virol. Methods 2007, 142, 29–40. [Google Scholar] [CrossRef]
- Khetawat, D.; Broder, C.C. A functional henipavirus envelope glycoprotein pseudotyped lentivirus assay system. Virol. J. 2010, 7, 312. [Google Scholar] [CrossRef] [PubMed]
- Bossart, K.N.; Rockx, B.; Feldmann, F.; Brining, D.; Scott, D.; LaCasse, R.; Geisbert, J.B.; Feng, Y.R.; Chan, Y.P.; Hickey, A.C.; et al. A Hendra virus G glycoprotein subunit vaccine protects African green monkeys from Nipah virus challenge. Sci. Transl. Med. 2012, 4, 146ra107. [Google Scholar] [CrossRef] [PubMed]
- Hayman, D.T.; Suu-Ire, R.; Breed, A.C.; McEachern, J.A.; Wang, L.; Wood, J.L.; Cunningham, A.A. Evidence of henipavirus infection in West African fruit bats. PLoS ONE 2008, 3, e2739. [Google Scholar] [CrossRef] [PubMed]
- Peel, A.J.; Baker, K.S.; Crameri, G.; Barr, J.A.; Hayman, D.T.; Wright, E.; Broder, C.C.; Fernandez-Loras, A.; Fooks, A.R.; Wang, L.F.; et al. Henipavirus neutralising antibodies in an isolated island population of African fruit bats. PLoS ONE 2012, 7, e30346. [Google Scholar] [CrossRef]
- Pernet, O.; Schneider, B.S.; Beaty, S.M.; LeBreton, M.; Yun, T.E.; Park, A.; Zachariah, T.T.; Bowden, T.A.; Hitchens, P.; Ramirez, C.M.; et al. Evidence for henipavirus spillover into human populations in Africa. Nat. Commun. 2014, 5, 5342. [Google Scholar] [CrossRef]
- Peel, A.J.; Sargan, D.R.; Baker, K.S.; Hayman, D.T.S.; Barr, J.A.; Crameri, G.; Suu-Ire, R.; Broder, C.C.; Lembo, T.; Wang, L.F.; et al. Continent-wide panmixia of an African fruit bat facilitates transmission of potentially zoonotic viruses. Nat. Commun. 2013, 4, 2770. [Google Scholar] [CrossRef]
- Liu, H.; Saxena, A.; Sidhu, S.S.; Wu, D. Fc Engineering for Developing Therapeutic Bispecific Antibodies and Novel Scaffolds. Front. Immunol. 2017, 8, 38. [Google Scholar] [CrossRef]
- Ye, L.; Zeng, R.; Bai, Y.; Roopenian, D.C.; Zhu, X. Efficient mucosal vaccination mediated by the neonatal Fc receptor. Nat. Biotechnol. 2011, 29, 158–163. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, Y.; Li, R.; Wang, M.; Liu, Y.; Yin, Y.; Zai, X.; Song, X.; Chen, Y.; Xu, J.; Chen, W. Fc-Based Recombinant Henipavirus Vaccines Elicit Broad Neutralizing Antibody Responses in Mice. Viruses 2020, 12, 480. https://doi.org/10.3390/v12040480
Li Y, Li R, Wang M, Liu Y, Yin Y, Zai X, Song X, Chen Y, Xu J, Chen W. Fc-Based Recombinant Henipavirus Vaccines Elicit Broad Neutralizing Antibody Responses in Mice. Viruses. 2020; 12(4):480. https://doi.org/10.3390/v12040480
Chicago/Turabian StyleLi, Yaohui, Ruihua Li, Meirong Wang, Yujiao Liu, Ying Yin, Xiaodong Zai, Xiaohong Song, Yi Chen, Junjie Xu, and Wei Chen. 2020. "Fc-Based Recombinant Henipavirus Vaccines Elicit Broad Neutralizing Antibody Responses in Mice" Viruses 12, no. 4: 480. https://doi.org/10.3390/v12040480
APA StyleLi, Y., Li, R., Wang, M., Liu, Y., Yin, Y., Zai, X., Song, X., Chen, Y., Xu, J., & Chen, W. (2020). Fc-Based Recombinant Henipavirus Vaccines Elicit Broad Neutralizing Antibody Responses in Mice. Viruses, 12(4), 480. https://doi.org/10.3390/v12040480




