The Cajal Body in Plant-Virus Interactions
Abstract
1. Cajal Bodies in Plants
2. The Emerging Role of the Cajal Body in Plant-Virus Interactions
3. Conclusions and Outlook
Funding
Acknowledgments
Conflicts of Interest
References
- Cajal, S.R. Un sencillo metodo de coloracion seletiva del reticulo protoplasmatico y sus efectos en los diversos organos nerviosos de vertebrados e invertebrados. Trab. Lab. Investig. Biol. Univ. Madrid 1903, 2, 129–221. [Google Scholar]
- Love, A.J.; Yu, C.; Petukhova, N.V.; Kalinina, N.O.; Chen, J.; Taliansky, M.E. Cajal bodies and their role in plant stress and disease responses. RNA Biol. 2017, 14, 779–790. [Google Scholar] [CrossRef] [PubMed]
- Machyna, M.; Neugebauer, K.M.; Stanek, D. Coilin: The first 25 years. RNA Biol. 2015, 12, 590–596. [Google Scholar] [CrossRef] [PubMed]
- Collier, S.; Pendle, A.; Boudonck, K.; van Rij, T.; Dolan, L.; Shaw, P. A distant coilin homologue is required for the formation of cajal bodies in Arabidopsis. Mol. Biol. Cell 2006, 17, 2942–2951. [Google Scholar] [CrossRef] [PubMed]
- Shaw, J.; Love, A.J.; Makarova, S.S.; Kalinina, N.O.; Harrison, B.D.; Taliansky, M.E. Coilin, the signature protein of Cajal bodies, differentially modulates the interactions of plants with viruses in widely different taxa. Nucleus 2014, 5, 85–94. [Google Scholar] [CrossRef]
- Ogg, S.C.; Lamond, A.I. Cajal bodies and coilin--moving towards function. J. Cell Biol. 2002, 159, 17–21. [Google Scholar] [CrossRef]
- Barneche, F.S.F.; Echeverrıa, M. Fibrillarin genes encode both a conserved nucleolar protein and a novel small nucleolar RNA involved in ribosomal RNA methylation in Arabidopsis thaliana. J. Biol. Chem. 2000, 275, 27212–27220. [Google Scholar]
- Matera, A.G.; Shpargel, K.B. Pumping RNA: Nuclear bodybuilding along the RNP pipeline. Curr. Opin. Cell Biol. 2006, 18, 317–324. [Google Scholar] [CrossRef]
- Stanek, D.; Neugebauer, K.M. The Cajal body: A meeting place for spliceosomal snRNPs in the nuclear maze. Chromosoma 2006, 115, 343–354. [Google Scholar] [CrossRef]
- Gall, J.G. Cajal bodies: The first 100 years. Annu. Rev. Cell Dev. Biol. 2000, 16, 273–300. [Google Scholar] [CrossRef]
- Bassett, C.L. Cajal Bodies and Plant RNA Metabolism. Crit. Rev. Plant Sci. 2012, 31, 258–270. [Google Scholar] [CrossRef]
- Cioce, M.; Lamond, A.I. Cajal bodies: A long history of discovery. Annu. Rev. Cell Dev. Biol. 2005, 21, 105–131. [Google Scholar] [CrossRef] [PubMed]
- Carmo-Fonseca, M. New clues to the function of the Cajal body. EMBO Rep. 2002, 3, 726–727. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Ohtani, M. Plant snRNP Biogenesis: A Perspective from the Nucleolus and Cajal Bodies. Front. Plant Sci. 2017, 8, 2184. [Google Scholar] [CrossRef] [PubMed]
- He, X.-J.; Hsu, Y.-F.; Zhu, S.; Wierzbicki, A.T.; Pontes, O.; Pikaard, C.S.; Liu, H.L.; Wang, C.S.; Jin, H.; Zhu, J.K. An effector of RNA-directed DNA methylation in Arabidopsis is an ARGONAUTE 4- and RNA-binding protein. Cell 2009, 137, 498–508. [Google Scholar] [CrossRef] [PubMed]
- Gao, Z.; Liu, H.L.; Daxinger, L.; Pontes, O.; He, X.; Qian, W.; Lin, H.; Xie, M.; Lorkovic, Z.J.; Zhang, S.; et al. An RNA polymerase II- and AGO4-associated protein acts in RNA-directed DNA methylation. Nature 2010, 465, 106–109. [Google Scholar] [CrossRef] [PubMed]
- Li, C.F.; Pontes, O.; El-Shami, M.; Henderson, I.R.; Bernatavichute, Y.V.; Chan, S.W.; Lagrange, T.; Pikaard, C.S.; Jacobsen, S.E. An ARGONAUTE4-containing nuclear processing center colocalized with Cajal bodies in Arabidopsis thaliana. Cell 2006, 126, 93–106. [Google Scholar] [CrossRef]
- Li, C.F.; Henderson, I.R.; Song, L.; Fedoroff, N.; Lagrange, T.; Jacobsen, S.E. Dynamic regulation of ARGONAUTE4 within multiple nuclear bodies in Arabidopsis thaliana. PLoS Genet. 2008, 4, e27. [Google Scholar] [CrossRef]
- Kalinina, N.O.; Makarova, S.; Makhotenko, A.; Love, A.J.; Taliansky, M. The Multiple Functions of the Nucleolus in Plant Development, Disease and Stress Responses. Front. Plant Sci. 2018, 9, 132. [Google Scholar] [CrossRef]
- Boulon, S.; Westman, B.J.; Hutten, S.; Boisvert, F.M.; Lamond, A.I. The nucleolus under stress. Mol. Cell 2010, 40, 216–227. [Google Scholar] [CrossRef]
- Ruiz-Ruiz, S.; Soler, N.; Sanchez-Navarro, J.; Fagoaga, C.; Lopez, C.; Navarro, L.; Moreno, P.; Pena, L.; Flores, R. Citrus tristeza virus p23: Determinants for nucleolar localization and their influence on suppression of RNA silencing and pathogenesis. Mol. Plant Microbe Interact. 2013, 26, 306–318. [Google Scholar] [CrossRef] [PubMed]
- Rajamaki, M.L.; Valkonen, J.P. Control of nuclear and nucleolar localization of nuclear inclusion protein a of picorna-like Potato virus a in Nicotiana species. Plant Cell 2009, 21, 2485–2502. [Google Scholar] [CrossRef] [PubMed]
- Zheng, L.; He, J.; Ding, Z.; Zhang, C.; Meng, R. Identification of Functional Domain(s) of Fibrillarin Interacted with p2 of Rice stripe virus. Can. J. Infect. Dis. Med. Microbiol. 2018, 2018. [Google Scholar] [CrossRef] [PubMed]
- Shaw, J.; Yu, C.; Makhotenko, A.V.; Makarova, S.S.; Love, A.J.; Kalinina, N.O.; MacFarlane, S.; Chen, J.; Taliansky, M.E. Interaction of a plant virus protein with the signature Cajal body protein coilin facilitates salicylic acid-mediated plant defence responses. New Phytol. 2019, 224, 439–453. [Google Scholar] [CrossRef]
- Kim, S.H.; Macfarlane, S.; Kalinina, N.O.; Rakitina, D.V.; Ryabov, E.V.; Gillespie, T.; Haupt, S.; Brown, J.W.; Taliansky, M. Interaction of a plant virus-encoded protein with the major nucleolar protein fibrillarin is required for systemic virus infection. Proc. Natl. Acad. Sci. USA 2007, 104, 11115–11120. [Google Scholar] [CrossRef]
- Semashko, M.A.; Rakitina, D.V.; Gonzalez, I.; Canto, T.; Kalinina, N.O.; Taliansky, M.E. Movement protein of hordeivirus interacts in vitro and in vivo with coilin, a major structural protein of Cajal bodies. Dokl. Biochem. Biophys. 2012, 442, 57–60. [Google Scholar] [CrossRef]
- Wang, L.; Ding, Y.; He, L.; Zhang, G.; Zhu, J.-K.; Lozano-Duran, R. A virus-encoded protein suppresses methylation of the viral genome in the Cajal body through its interaction with AGO4. bioRxiv 2020. [Google Scholar] [CrossRef]
- Xu, M.; Mazur, M.J.; Tao, X.; Kormelink, R. Cellular RNA Hubs: Friends and Foes of Plant Viruses. Mol. Plant Microbe Interact. 2020, 33, 40–54. [Google Scholar] [CrossRef]
- Guo, T.W.; Vimalesvaran, D.; Thompson, J.R.; Perry, K.L.; Krenz, B. Subcellular localization of grapevine red blotch-associated virus ORFs V2 and V3. Virus Genes 2015, 51, 156–158. [Google Scholar] [CrossRef]
- Ryabov, E.V.; Robinson, D.J.; Taliansky, M. Umbravirus-encoded proteins both stabilize heterologous viral RNA and mediate its systemic movement in some plant species. Virology 2001, 288, 391–400. [Google Scholar] [CrossRef]
- Ryabov, E.V.; Robinson, D.J.; Taliansky, M.E. A plant virus-encoded protein facilitates long-distance movement of heterologous viral RNA. Proc. Natl. Acad. Sci. USA 1999, 96, 1212–1217. [Google Scholar] [CrossRef] [PubMed]
- Ryabov, E.V.; Kim, S.H.; Taliansky, M. Identification of a nuclear localization signal and nuclear export signal of the umbraviral long-distance RNA movement protein. J. Gen. Virol. 2004, 85, 1329–1333. [Google Scholar] [CrossRef] [PubMed]
- Ryabov, E.V.; Oparka, K.J.; Santa Cruz, S.; Robinson, D.J.; Taliansky, M.E. Intracellular location of two groundnut rosette umbravirus proteins delivered by PVX and TMV vectors. Virology 1998, 242, 303–313. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Kim, S.H.; Ryabov, E.V.; Kalinina, N.O.; Rakitina, D.V.; Gillespie, T.; MacFarlane, S.; Haupt, S.; Brown, J.W.; Taliansky, M. Cajal bodies and the nucleolus are required for a plant virus systemic infection. EMBO J. 2007, 26, 2169–2179. [Google Scholar] [CrossRef]
- Li, Z.; Zhang, Y.; Jiang, Z.; Jin, X.; Zhang, K.; Wang, X.; Han, C.; Yu, J.; Li, D. Hijacking of the nucleolar protein fibrillarin by TGB1 is required for cell-to-cell movement of Barley stripe mosaic virus. Mol. Plant Pathol. 2018, 19, 1222–1237. [Google Scholar] [CrossRef]
- Wang, M.B.; Masuta, C.; Smith, N.A.; Shimura, H. RNA silencing and plant viral diseases. Mol. Plant Microbe Interact. 2012, 25, 1275–1285. [Google Scholar] [CrossRef]
- Matzke, M.A.; Kanno, T.; Matzke, A.J.M. RNA-Directed DNA Methylation: The Evolution of a Complex Epigenetic Pathway in Flowering Plants. Annu. Rev. Plant Biol. 2015, 66, 243–267. [Google Scholar] [CrossRef]
- Pontes, O.; Li, C.F.; Costa Nunes, P.; Haag, J.; Ream, T.; Vitins, A.; Jacobsen, S.E.; Pikaard, C.S. The Arabidopsis chromatin-modifying nuclear siRNA pathway involves a nucleolar RNA processing center. Cell 2006, 126, 79–92. [Google Scholar] [CrossRef]
- Wierzbicki, A.T.; Haag, J.R.; Pikaard, C.S. Noncoding Transcription by RNA Polymerase Pol IVb/Pol V Mediates Transcriptional Silencing of Overlapping and Adjacent Genes. Cell 2008, 135, 635–648. [Google Scholar] [CrossRef]
- Wang, B.; Li, F.; Huang, C.; Yang, X.; Qian, Y.; Xie, Y.; Zhou, X. V2 of tomato yellow leaf curl virus can suppress methylation-mediated transcriptional gene silencing in plants. J. Gen. Virol. 2014, 95, 225–230. [Google Scholar] [CrossRef]
- Rothenstein, D.; Krenz, B.; Selchow, O.; Jeske, H. Tissue and cell tropism of Indian cassava mosaic virus (ICMV) and its AV2 (precoat) gene product. Virology 2007, 359, 137–145. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Wu, Y.; Gong, Q.; Ismayil, A.; Yuan, Y.; Lian, B.; Jia, Q.; Han, M.; Deng, H.; Hong, Y.; et al. Geminiviral V2 Protein Suppresses Transcriptional Gene Silencing through Interaction with AGO4. J. Virol. 2019, 93. [Google Scholar] [CrossRef]
| Virus Name | Family | Genus | Genome | Virus-Encoded Protein | Localization | Known Cajal Body Interactor | Described Function | Ref. |
|---|---|---|---|---|---|---|---|---|
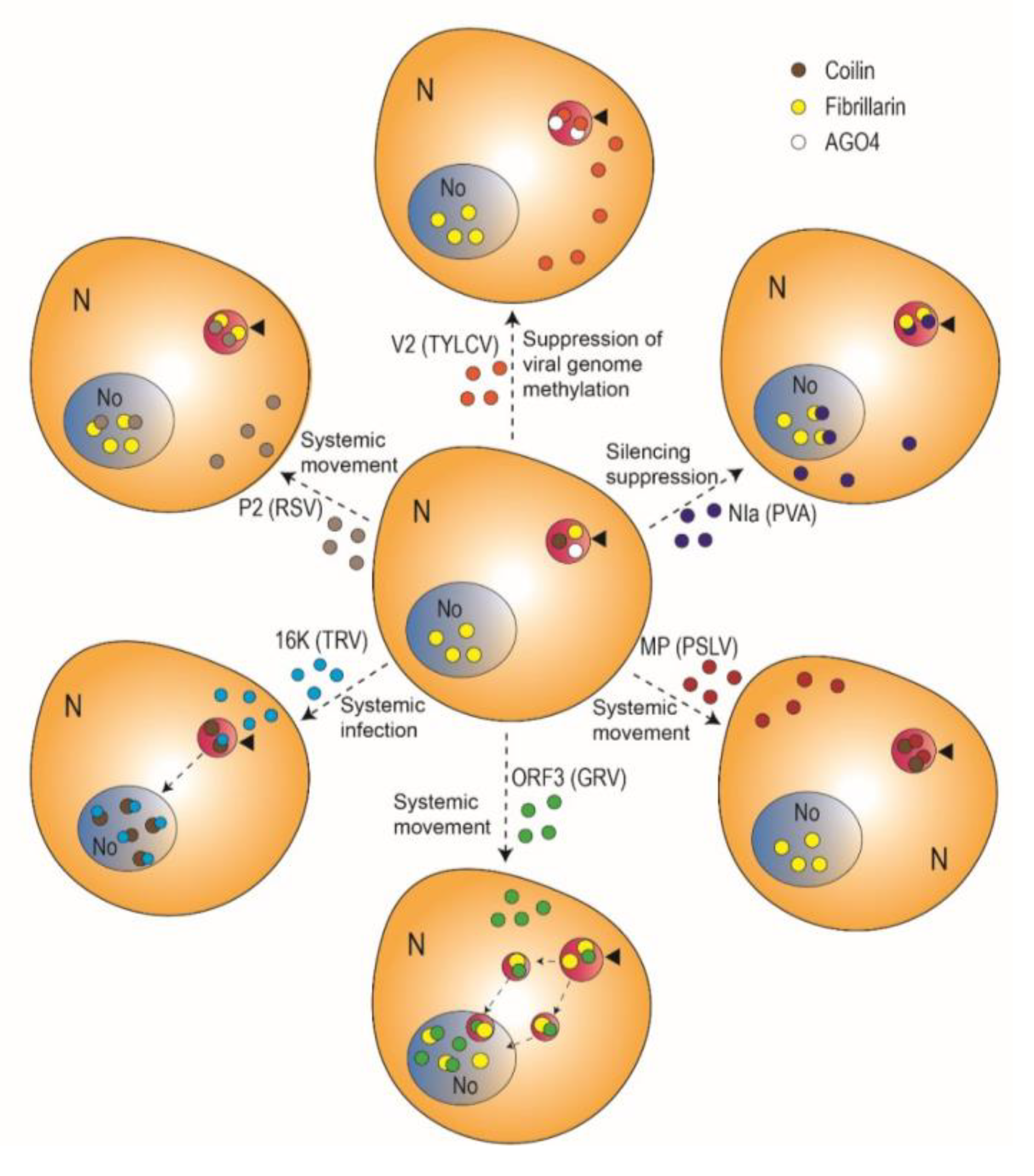
| Rice stripe virus (RSV) | Phenuiviridae | Tenuivirus | (−)ssRNA | P2 | Nucleus, cytoplasm, Cajal bodies, nucleolus | Fibrillarin | Systemic movement | [23] |
| Potato virus A (PVA) | Potyviridae | Potyvirus | (+)ssRNA | Nuclear inclusion protein a (NIa) | Nucleus, nucleolus, Cajal bodies | Fibrillarin | Suppression of RNA silencing | [22] |
| Groundnut rosette virus (GRV) | Tombusviridae | Umbravirus | (+)ssRNA | Open reading frame 3 (ORF3) protein | Cajal bodies, nucleolus, Cajal body (CB)-like structures | Fibrillarin | Systemic movement | [25] |
| Poa semilatent virus (PSLV) | Virgaviridae | Hordeovirus | (+)ssRNA | Movement protein (MP; TGBp1) | Nucleolus, Cajal bodies, inclusions in the nucleoplasm | Coilin | Cell-to-cell and systemic movement | [26] |
| Tobacco rattle virus (TRV) | Virgaviridae | Tobravirus | (−)ssRNA | 16K | Nucleus, nucleolus, Cajal bodies | Coilin | Systemic infection | [24] |
| Tomato yellow leaf curl virus (TYLCV) | Geminiviridae | Begomovirus | ssDNA | V2 | Cajal bodies, nucleoplasm, cytoplasm | AGO4 | Suppression of viral DNA methylation | [27] |
| Grapevine red blotch-associated virus (GRBaV) | Geminiviridae | Grablovirus | ssDNA | V2 | Nucleoplasm, Cajal bodies, cytoplasm, nucleolus | Fibrillarin | Unknown | [29] |
| Citrus tristeza virus (CTV) | Closteroviridae | Closterovirus | (+)ssRNA | P23 | Nucleolus, Cajal bodies, plasmodesma | Unknown | Suppression of RNA silencing | [21] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ding, Y.; Lozano-Durán, R. The Cajal Body in Plant-Virus Interactions. Viruses 2020, 12, 250. https://doi.org/10.3390/v12020250
Ding Y, Lozano-Durán R. The Cajal Body in Plant-Virus Interactions. Viruses. 2020; 12(2):250. https://doi.org/10.3390/v12020250
Chicago/Turabian StyleDing, Yi, and Rosa Lozano-Durán. 2020. "The Cajal Body in Plant-Virus Interactions" Viruses 12, no. 2: 250. https://doi.org/10.3390/v12020250
APA StyleDing, Y., & Lozano-Durán, R. (2020). The Cajal Body in Plant-Virus Interactions. Viruses, 12(2), 250. https://doi.org/10.3390/v12020250





