Estimating the Risk of Human Herpesvirus 6 and Cytomegalovirus Transmission to Ugandan Infants from Viral Shedding in Saliva by Household Contacts
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Cohort and Data
2.2. Definitions of Exposure and Transmission Events
2.3. Survival Analysis and Dose-Response Modeling
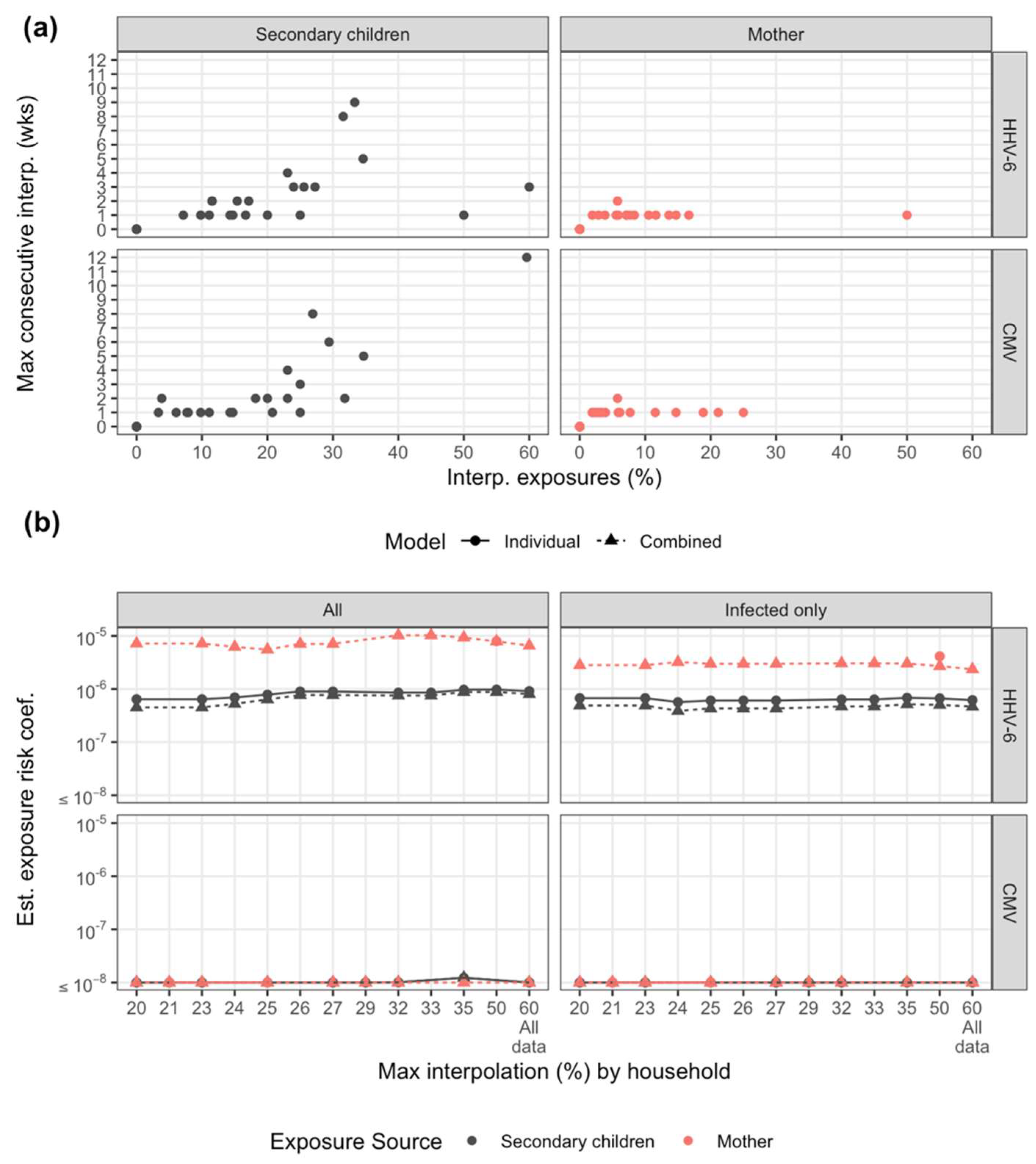
2.4. Sensitivity Analysis
2.5. Software, Data, and Code Availability
3. Results
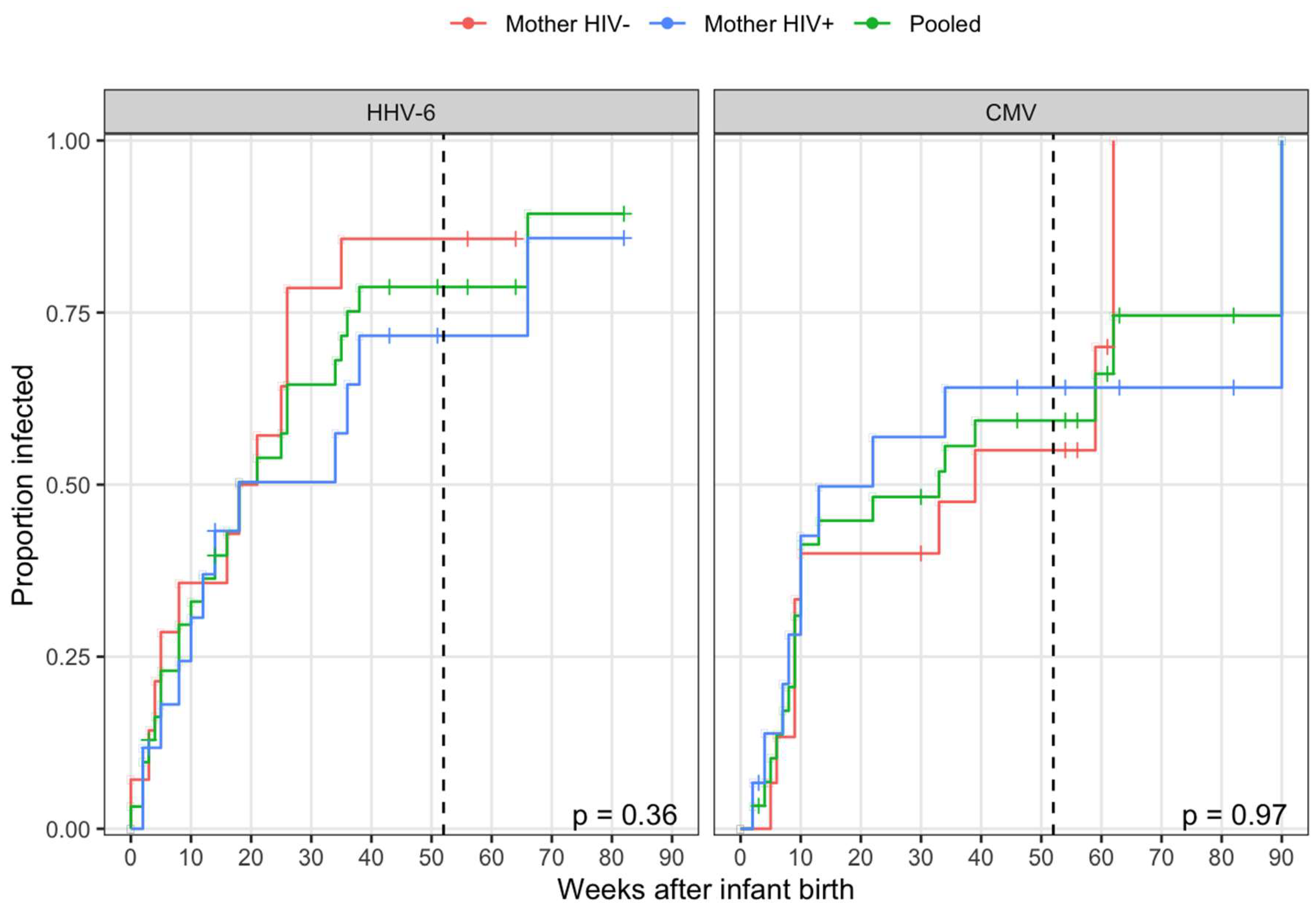
3.1. HHV-6 and CMV Transmission Occurred at High Rate Early During Infancy
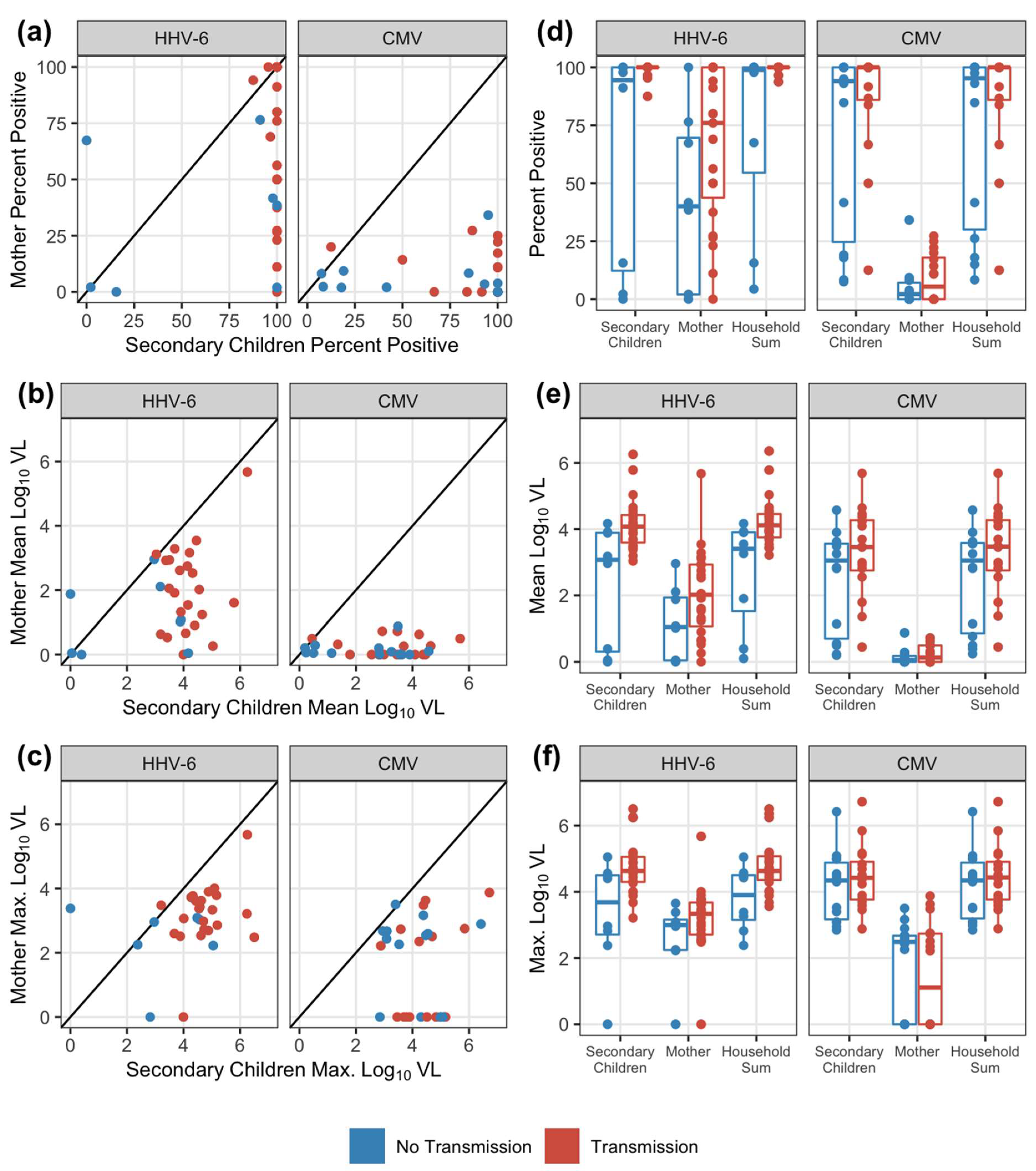
3.2. Secondary Children Shed at Higher Viral Loads than Mothers
3.3. Correlation between household shedding and transmission to infants
3.4. HHV-6, but not CMV, Acquisition in Infants was Associated with Viral Loads of Oral Shedding Exposures
3.5. A Combined Exposure Model for HHV-6 Highlights Differential Risk between Mother and Secondary Children Exposures
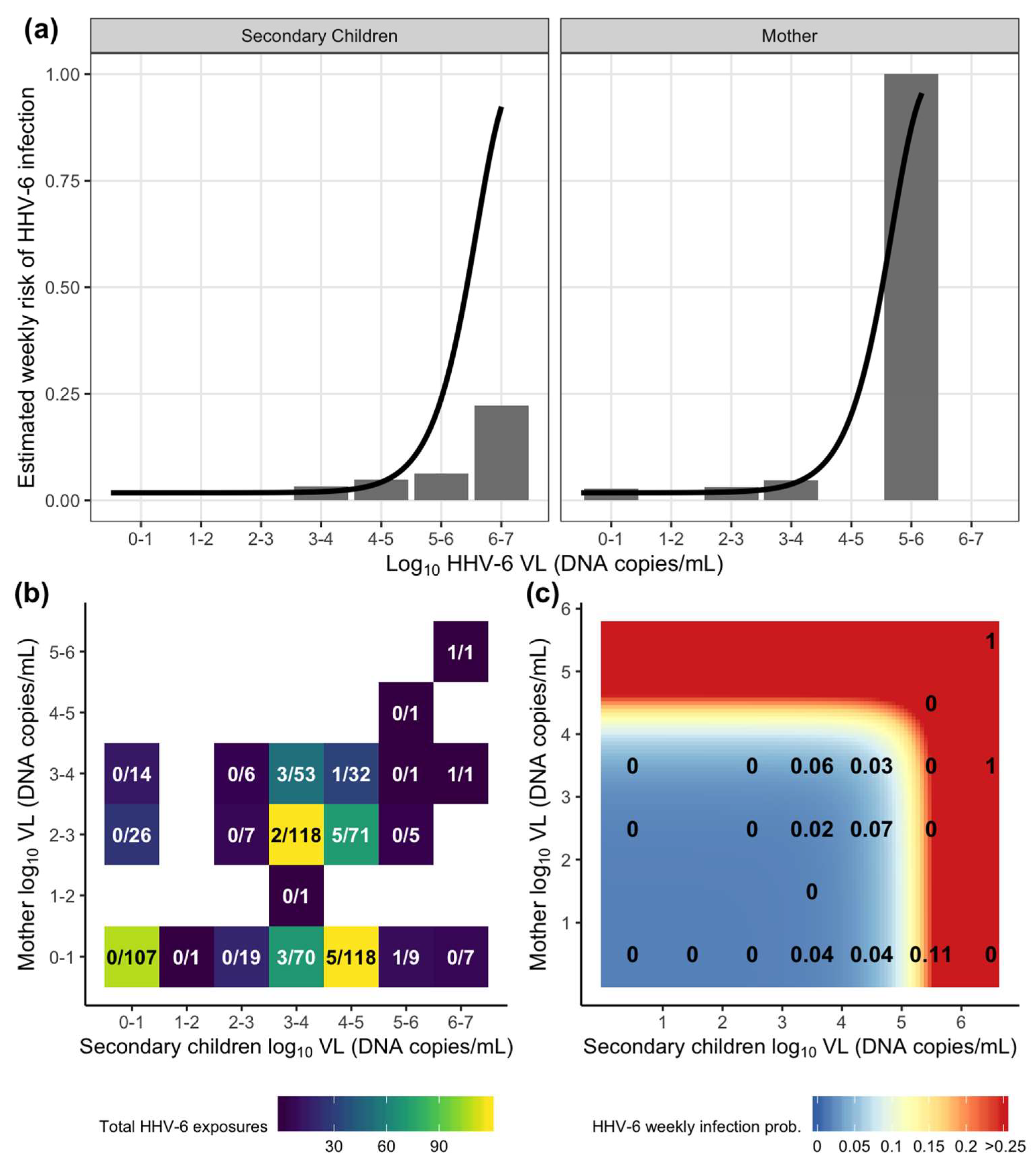
3.6. The Dose–Response Relationship between the Viral Load of HHV-6 oral Shedding Exposure and Weekly Infant Acquisition
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Appendix A
Appendix B

| Virus | Model | Parameter | Estimate | -Log likelihood |
|---|---|---|---|---|
| HHV-6 | Constant risk (null) | b0 | 3.67 × 10−2 | 91.2 |
| Secondary children | b0 | 2.10 × 10−2 | 84.8 | |
| bS | 9.13 × 10−7 | |||
| Mother | b0 | 2.93 × 10−2 | 87.5 | |
| bM | 7.96 × 10−6 | |||
| Household sum | b0 | 2.03 × 10−2 | 84.6 | |
| bHH | 9.12 × 10−7 | |||
| Combined model | b0 | 1.82 × 10−2 | 84.2 | |
| bM | 6.56 × 10−6 | |||
| bS | 8.03 × 10−7 | |||
| CMV | Constant risk (null) | b0 | 1.98 × 10−2 | 74.1 |
| Secondary children | b0 | 1.94 × 10−2 | ||
| bS | 0 | |||
| Mother | b0 | 1.94 × 10−2 | ||
| bM | 0 | |||
| Household sum | b0 | 1.94 × 10−2 | ||
| bHH | 0 | |||
| Combined model | b0 | 1.94 × 10−2 | ||
| bM | 0 | |||
| bS | 0 |
References
- Condon, L.M.; Cederberg, L.E.; Rabinovitch, M.D.; Liebo, R.V.; Go, J.C.; Delaney, A.S.; Schmeling, D.O.; Thomas, W.; Balfour, H.H., Jr. Age-specific prevalence of Epstein-Barr virus infection among Minnesota children: Effects of race/ethnicity and family environment. Clin Infect Dis. 2014, 59, 501–508. [Google Scholar] [CrossRef] [PubMed]
- Kusuhara, K.; Takabayashi, A.; Ueda, K.; Hidaka, Y.; Minamishima, I.; Take, H.; Fujioka, K.; Imai, S.; Osato, T. Breast milk is not a significant source for early Epstein-Barr virus or human herpesvirus 6 infection in infants: A seroepidemiologic study in 2 endemic areas of human T-cell lymphotropic virus type I in Japan. Microbiol. Immunol. 1997, 41, 309–312. [Google Scholar] [CrossRef] [PubMed]
- Zerr, D.M.; Meier, A.S.; Selke, S.S.; Frenkel, L.M.; Huang, M.L.; Wald, A.; Rhoads, M.P.; Nguy, L.; Bornemann, R.; Morrow, R.A.; et al. A population-based study of primary human herpesvirus 6 infection. N. Engl. J. Med. 2005, 352, 768–776. [Google Scholar] [CrossRef] [PubMed]
- Eimer, W.A.; Vijaya Kumar, D.K.; Navalpur Shanmugam, N.K.; Rodriguez, A.S.; Mitchell, T.; Washicosky, K.J.; Gyorgy, B.; Breakefield, X.O.; Tanzi, R.E.; Moir, R.D. Alzheimer’s Disease-Associated beta-Amyloid Is Rapidly Seeded by Herpesviridae to Protect against Brain Infection. Neuron 2018, 100, 1527–1532. [Google Scholar] [CrossRef] [PubMed]
- Readhead, B.; Haure-Mirande, J.V.; Funk, C.C.; Richards, M.A.; Shannon, P.; Haroutunian, V.; Sano, M.; Liang, W.S.; Beckmann, N.D.; Price, N.D.; et al. Multiscale Analysis of Independent Alzheimer’s Cohorts Finds Disruption of Molecular, Genetic, and Clinical Networks by Human Herpesvirus. Neuron 2018, 99, 64–82 e67. [Google Scholar] [CrossRef] [PubMed]
- Zerr, D.M. Human herpesvirus 6 and central nervous system disease in hematopoietic cell transplantation. J. Clin. Virol. 2006, 37, S52–S56. [Google Scholar] [CrossRef]
- Agut, H.; Bonnafous, P.; Gautheret-Dejean, A. Laboratory and clinical aspects of human herpesvirus 6 infections. Clin. Microbiol. Rev. 2015, 28, 313–335. [Google Scholar] [CrossRef]
- Kenneson, A.; Cannon, M.J. Review and meta-analysis of the epidemiology of congenital cytomegalovirus (CMV) infection. Rev. Med. Virol. 2007, 17, 253–276. [Google Scholar] [CrossRef]
- Gantt, S.; Orem, J.; Krantz, E.M.; Morrow, R.A.; Selke, S.; Huang, M.L.; Schiffer, J.T.; Jerome, K.R.; Nakaganda, A.; Wald, A.; et al. Prospective Characterization of the Risk Factors for Transmission and Symptoms of Primary Human Herpesvirus Infections Among Ugandan Infants. J. Infect. Dis 2016, 214, 36–44. [Google Scholar] [CrossRef]
- Pass, R.F. Epidemiology and transmission of cytomegalovirus. J. Infect. Dis 1985, 152, 243–248. [Google Scholar] [CrossRef]
- Boeckh, M.; Geballe, A.P. Cytomegalovirus: Pathogen, paradigm, and puzzle. J. Clin. Invest. 2011, 121, 1673–1680. [Google Scholar] [CrossRef] [PubMed]
- Mentec, H.; Leport, C.; Leport, J.; Marche, C.; Harzic, M.; Vilde, J.L. Cytomegalovirus colitis in HIV-1-infected patients: A prospective research in 55 patients. AIDS 1994, 8, 461–467. [Google Scholar] [CrossRef]
- Maclean, H.; Dhillon, B. Cytomegalovirus retinitis: Diagnosis and treatment. Int J. STD AIDS 1993, 4, 322–325. [Google Scholar] [CrossRef]
- Adachi, K.; Xu, J.; Ank, B.; Watts, D.H.; Camarca, M.; Mofenson, L.M.; Pilotto, J.H.; Joao, E.; Gray, G.; Theron, G.; et al. Congenital Cytomegalovirus and HIV Perinatal Transmission. Pediatr Infect. Dis. J. 2018, 37, 1016–1021. [Google Scholar] [CrossRef] [PubMed]
- Looney, R.J.; Falsey, A.; Campbell, D.; Torres, A.; Kolassa, J.; Brower, C.; McCann, R.; Menegus, M.; McCormick, K.; Frampton, M.; et al. Role of cytomegalovirus in the T cell changes seen in elderly individuals. Clin. Immunol. 1999, 90, 213–219. [Google Scholar] [CrossRef]
- Karrer, U.; Sierro, S.; Wagner, M.; Oxenius, A.; Hengel, H.; Koszinowski, U.H.; Phillips, R.E.; Klenerman, P. Memory inflation: Continuous accumulation of antiviral CD8+ T cells over time. J. Immunol. 2003, 170, 2022–2029. [Google Scholar] [CrossRef] [PubMed]
- Weltevrede, M.; Eilers, R.; de Melker, H.E.; van Baarle, D. Cytomegalovirus persistence and T-cell immunosenescence in people aged fifty and older: A systematic review. Exp. Gerontol. 2016, 77, 87–95. [Google Scholar] [CrossRef] [PubMed]
- Simanek, A.M.; Dowd, J.B.; Pawelec, G.; Melzer, D.; Dutta, A.; Aiello, A.E. Seropositivity to cytomegalovirus, inflammation, all-cause and cardiovascular disease-related mortality in the United States. PLoS ONE 2011, 6, e16103. [Google Scholar] [CrossRef] [PubMed]
- Brodin, P.; Jojic, V.; Gao, T.; Bhattacharya, S.; Angel, C.J.; Furman, D.; Shen-Orr, S.; Dekker, C.L.; Swan, G.E.; Butte, A.J.; et al. Variation in the human immune system is largely driven by non-heritable influences. Cell 2015, 160, 37–47. [Google Scholar] [CrossRef] [PubMed]
- Amin, M.M.; Bialek, S.R.; Dollard, S.C.; Wang, C. Urinary Cytomegalovirus Shedding in the United States: The National Health and Nutrition Examination Surveys, 1999-2004. Clin. Infect. Dis. 2018, 67, 587–592. [Google Scholar] [CrossRef] [PubMed]
- Prendergast, A.J.; Goga, A.E.; Waitt, C.; Gessain, A.; Taylor, G.P.; Rollins, N.; Abrams, E.J.; Lyall, E.H.; de Perre, P.V. Transmission of CMV, HTLV-1, and HIV through breastmilk. Lancet Child. Adolesc Health 2019, 3, 264–273. [Google Scholar] [CrossRef]
- Hamprecht, K.; Maschmann, J.; Vochem, M.; Dietz, K.; Speer, C.P.; Jahn, G. Epidemiology of transmission of cytomegalovirus from mother to preterm infant by breastfeeding. Lancet 2001, 357, 513–518. [Google Scholar] [CrossRef]
- Stagno, S.; Reynolds, D.W.; Pass, R.F.; Alford, C.A. Breast milk and the risk of cytomegalovirus infection. N. Engl J. Med. 1980, 302, 1073–1076. [Google Scholar] [CrossRef] [PubMed]
- Emery, V.C.; Clark, D.A. HHV-6A, 6B, and 7: Persistence in the population, epidemiology and transmission. In Human Herpesviruses: Biology, Therapy, and Immunoprophylaxis; Arvin, A., Campadelli-Fiume, G., Mocarski, E., Moore, P.S., Roizman, B., Whitley, R., Yamanishi, K., Eds.; Cambridge University Press: Cambridge, UK, 2007. [Google Scholar]
- Corey, L.; Wald, A.; Patel, R.; Sacks, S.L.; Tyring, S.K.; Warren, T.; Douglas, J.M., Jr.; Paavonen, J.; Morrow, R.A.; Beutner, K.R.; et al. Once-daily valacyclovir to reduce the risk of transmission of genital herpes. N. Engl. J. Med. 2004, 350, 11–20. [Google Scholar] [CrossRef]
- Schiffer, J.T.; Mayer, B.T.; Fong, Y.; Swan, D.A.; Wald, A. Herpes simplex virus-2 transmission probability estimates based on quantity of viral shedding. J. R Soc. Interface 2014, 11, 20140160. [Google Scholar] [CrossRef]
- Quinn, T.C.; Wawer, M.J.; Sewankambo, N.; Serwadda, D.; Li, C.; Wabwire-Mangen, F.; Meehan, M.O.; Lutalo, T.; Gray, R.H. Viral load and heterosexual transmission of human immunodeficiency virus type 1. Rakai Project Study Group. N. Engl. J. Med. 2000, 342, 921–929. [Google Scholar] [CrossRef]
- Baeten, J.M.; Kahle, E.; Lingappa, J.R.; Coombs, R.W.; Delany-Moretlwe, S.; Nakku-Joloba, E.; Mugo, N.R.; Wald, A.; Corey, L.; Donnell, D.; et al. Genital HIV-1 RNA predicts risk of heterosexual HIV-1 transmission. Sci. Transl. Med. 2011, 3, 77ra29. [Google Scholar] [CrossRef]
- Johnston, C.; Orem, J.; Okuku, F.; Kalinaki, M.; Saracino, M.; Katongole-Mbidde, E.; Sande, M.; Ronald, A.; McAdam, K.; Huang, M.L.; et al. Impact of HIV infection and Kaposi sarcoma on human herpesvirus-8 mucosal replication and dissemination in Uganda. PLoS ONE 2009, 4, e4222. [Google Scholar] [CrossRef]
- Boeckh, M.; Huang, M.; Ferrenberg, J.; Stevens-Ayers, T.; Stensland, L.; Nichols, W.G.; Corey, L. Optimization of quantitative detection of cytomegalovirus DNA in plasma by real-time PCR. J. Clin. Microbiol. 2004, 42, 1142–1148. [Google Scholar] [CrossRef]
- Cone, R.W.; Huang, M.L.; Ashley, R.; Corey, L. Human herpesvirus 6 DNA in peripheral blood cells and saliva from immunocompetent individuals. J. Clin. Microbiol 1993, 31, 1262–1267. [Google Scholar] [CrossRef]
- Zerr, D.M.; Gupta, D.; Huang, M.L.; Carter, R.; Corey, L. Effect of antivirals on human herpesvirus 6 replication in hematopoietic stem cell transplant recipients. Clin. Infect. Dis 2002, 34, 309–317. [Google Scholar] [CrossRef] [PubMed]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2019. [Google Scholar]
- Wickham, H.; Averick, M.; Bryan, J.; Chang, W.; McGowan, L.D.A.; François, R.; Grolemund, G.; Hayes, A.; Henry, L.; Hester, J.; et al. Welcome to the tidyverse. J. Open Source Softw. 2019, 4, 1686. [Google Scholar] [CrossRef]
- Therneau, T.M. A Package for Survival Analysis in S, version 2.38; 2015; Available online: http://cran.r-project.org/package=survival (accessed on 31 January 2020).
- Hothorn, T.; Hornik, K.; van de Wiel, M.A.; Zeileis, A. Implementing a Class of Permutation Tests: The coin Package. J. Stat. Softw. 2008, 28. [Google Scholar] [CrossRef]
- Hothorn, T.; Hornik, K.; van de Wiel, M.A.; Zeileis, A. A Lego System for Conditional Inference. Am. Stat. 2006, 60, 257–263. [Google Scholar] [CrossRef]
- Nocedal, J.; Wright, S.J. Numerical Optimization, 2nd ed.; Springer: New York, NY, USA, 2006. [Google Scholar]
- Nelder, J.A.; Mead, R. A Simplex Method for Function Minimization. Comput. J. 1965, 7, 308–313. [Google Scholar] [CrossRef]
- Carnell, R. lhs: Latin Hypercube Samples, version 1.0.1; 2019; Available online: https://github.com/bertcarnell/lhs (accessed on 6 January 2020).
- Blischak, J.; Carbonetto, P.; Stephens, M. Creating and sharing reproducible research code the workflowr way [version 1; peer review: 3 approved]. F1000Research 2019, 8. [Google Scholar] [CrossRef]
- Rhoads, M.P.; Magaret, A.S.; Zerr, D.M. Family saliva sharing behaviors and age of human herpesvirus-6B infection. J. Infect. 2007, 54, 623–626. [Google Scholar] [CrossRef]
- Murata, H.; Nii, R.; Ito, M.; Ihara, T.; Komada, Y. Quantitative detection of HCMV-DNA in saliva from infants and breast milk on real-time polymerase chain reaction. Pediatr. Int. 2009, 51, 530–534. [Google Scholar] [CrossRef]
- Slyker, J.; Farquhar, C.; Atkinson, C.; Asbjornsdottir, K.; Roxby, A.; Drake, A.; Kiarie, J.; Wald, A.; Boeckh, M.; Richardson, B.; et al. Compartmentalized cytomegalovirus replication and transmission in the setting of maternal HIV-1 infection. Clin. Infect. Dis. 2014, 58, 564–572. [Google Scholar] [CrossRef]
- Boucoiran, I.; Mayer, B.T.; Krantz, E.M.; Marchant, A.; Pati, S.; Boppana, S.; Wald, A.; Corey, L.; Casper, C.; Schiffer, J.T.; et al. Nonprimary Maternal Cytomegalovirus Infection After Viral Shedding in Infants. Pediatr Infect. Dis. J. 2018, 37, 627–631. [Google Scholar] [CrossRef]
- Mayer, B.T.; Koopman, J.S.; Ionides, E.L.; Pujol, J.M.; Eisenberg, J.N. A dynamic dose-response model to account for exposure patterns in risk assessment: A case study in inhalation anthrax. J. R Soc. Interface 2011, 8, 506–517. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Stocki, R. A method to improve design reliability using optimal Latin hypercube sampling. Comput. Assist. Mech. Eng. Sci. 2005, 12, 393–411. [Google Scholar]
| No Exposure | Mean Exposure | Maximum Exposure | |||||
|---|---|---|---|---|---|---|---|
| Virus | Exposure Source | Weekly Risk (%) | Log10 DNA copies/mL | Risk (%) | Log10 DNA Copies/mL | Risk (%) | p-Value |
| HHV-6 | Null | 3.60 | |||||
| Secondary children | 2.07 | 3.98 | 2.92 | 6.50 | 94.66 | <0.001 | |
| Mother | 2.88 | 2.82 | 3.39 | 5.68 | 97.75 | 0.007 | |
| Household sum | 2.01 | 3.94 | 2.79 | 6.50 | 94.63 | <0.001 | |
| CMV | Null | 1.96 | |||||
| Secondary children | 1.92 | 3.72 | 1.92 | 6.72 | 1.92 | 0.939 | |
| Mother | 1.92 | 2.56 | 1.92 | 3.88 | 1.92 | 0.939 | |
| Household sum | 1.92 | 3.77 | 1.92 | 6.72 | 1.92 | 0.939 | |
| No Exposure | Secondary Children Exposure 1 | Maternal Exposure 1 | ||||
|---|---|---|---|---|---|---|
| Virus | Weekly Risk (%) | Mean Exposure Risk (%) | Maximum Exposure Risk (%) | Mean Exposure Risk (%) | Maximum Exposure Risk (%) | p-Value |
| HHV-6 | 1.80 | 2.55 | 92.39 | 2.22 | 95.58 | <0.001 |
| CMV | 1.92 | 1.92 | 1.92 | 1.92 | 1.92 | 0.997 |
| HHV-6 Infectious Dose (ID, log10 DNA copies/mL) | ||||
|---|---|---|---|---|
| Model | Exposure Source | ID25 | ID50 | ID75 |
| Individual | Secondary Children | 5.47 | 5.87 | 6.17 |
| Mother | 4.51 | 4.92 | 5.23 | |
| Combined | Secondary Children | 5.53 | 5.92 | 6.23 |
| Mother | 4.61 | 5.01 | 5.32 | |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mayer, B.T.; Krantz, E.M.; Wald, A.; Corey, L.; Casper, C.; Gantt, S.; Schiffer, J.T. Estimating the Risk of Human Herpesvirus 6 and Cytomegalovirus Transmission to Ugandan Infants from Viral Shedding in Saliva by Household Contacts. Viruses 2020, 12, 171. https://doi.org/10.3390/v12020171
Mayer BT, Krantz EM, Wald A, Corey L, Casper C, Gantt S, Schiffer JT. Estimating the Risk of Human Herpesvirus 6 and Cytomegalovirus Transmission to Ugandan Infants from Viral Shedding in Saliva by Household Contacts. Viruses. 2020; 12(2):171. https://doi.org/10.3390/v12020171
Chicago/Turabian StyleMayer, Bryan T., Elizabeth M. Krantz, Anna Wald, Lawrence Corey, Corey Casper, Soren Gantt, and Joshua T. Schiffer. 2020. "Estimating the Risk of Human Herpesvirus 6 and Cytomegalovirus Transmission to Ugandan Infants from Viral Shedding in Saliva by Household Contacts" Viruses 12, no. 2: 171. https://doi.org/10.3390/v12020171
APA StyleMayer, B. T., Krantz, E. M., Wald, A., Corey, L., Casper, C., Gantt, S., & Schiffer, J. T. (2020). Estimating the Risk of Human Herpesvirus 6 and Cytomegalovirus Transmission to Ugandan Infants from Viral Shedding in Saliva by Household Contacts. Viruses, 12(2), 171. https://doi.org/10.3390/v12020171





