Return of the Coronavirus: 2019-nCoV
Abstract
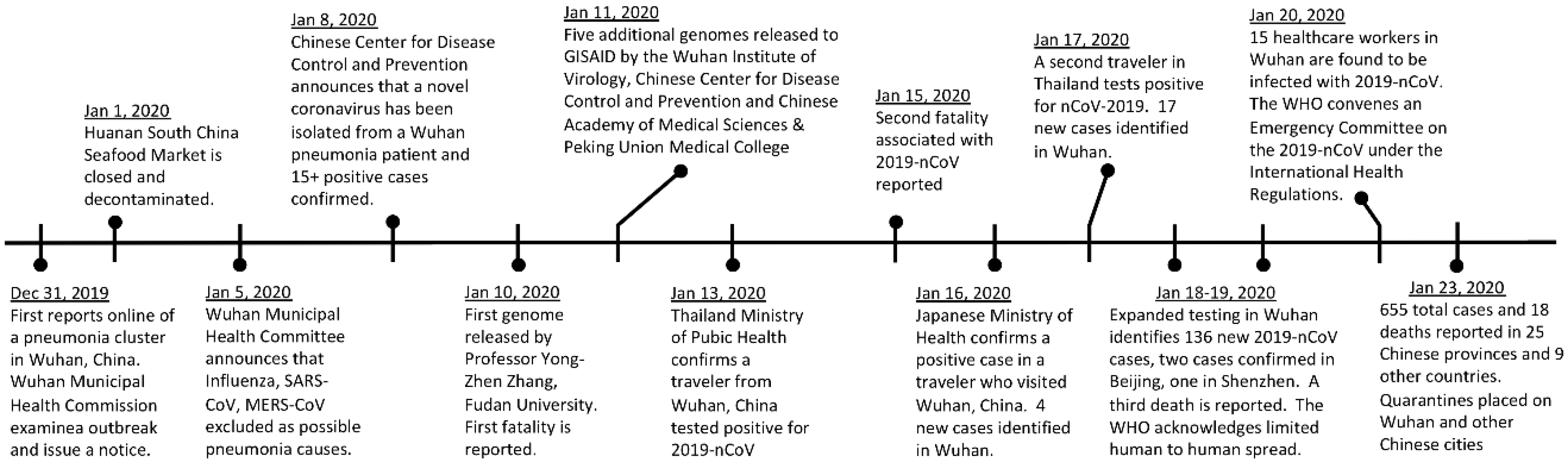
1. Emergence
2. Origins of 2019-nCoV
3. Susceptible Populations
4. Insights from the 2019-nCoV Sequence
5. Achieving Koch Postulates
6. Threat for Spread: Human to Human, Health Care Workers, and Super Spreaders
7. Emerging Diseases in the Age of Social Media
Author Contributions
Funding
Conflicts of Interest
References
- Wuhan Municipal Health Commision. Wuhan Municipal Health and Health Commission’s Briefing on the Current Pneumonia Epidemic Situation in Our City. Available online: http://wjw.wuhan.gov.cn/front/web/showDetail/2019123108989 (accessed on 19 January 2020).
- Kahn, N. New Virus Discovered by Chinese Scientists Investigating Pneumonia Outbreak. Wall Street Journal, 2020. [Google Scholar]
- World Health Organization. WHO Statement Regarding Cluster of Pneumonia Cases in Wuhan, China. Available online: https://www.who.int/china/news/detail/09-01-2020-who-statement-regarding-cluster-of-pneumonia-cases-in-wuhan-china (accessed on 19 January 2020).
- Virological.org. Novel 2019 Coronavirus Genome. Available online: http://virological.org/t/novel-2019-coronavirus-genome/319 (accessed on 19 January 2020).
- GSAID Database. 2020 Coronavirus. Available online: https://www.gisaid.org/CoV2020/ (accessed on 19 January 2020).
- World Health Organization. Novel Coronavirus—Japan (ex-China). Available online: https://www.who.int/csr/don/16-january-2020-novel-coronavirus-japan-ex-china/en/ (accessed on 19 January 2020).
- World Health Organization. Laboratory Testing for 2019 Novel Coronavirus (2019-nCoV) in Suspected Human Cases. Available online: https://www.who.int/health-topics/coronavirus/laboratory-diagnostics-for-novel-coronavirus (accessed on 19 January 2020).
- Imai, N.; Dorigattig, I.; Cori, A.; Riley, S.; Ferguson, N.M. Estimating the Potential Total Number of Novel Coronavirus (2019-nCoV) Cases in Wuhan City, China. Available online: https://www.imperial.ac.uk/mrc-global-infectious-disease-analysis/news--wuhan-coronavirus/ (accessed on 19 January 2020).
- Wuhan Municipal Health Commision. Wuhan Municipal Commission of Health and Health on Pneumonia of New Coronavirus Infection. Available online: http://wjw.wuhan.gov.cn/front/web/showDetail/2020012009077 (accessed on 20 January 2020).
- Stein, R.A. Super-spreaders in infectious diseases. Int. J. Infect. Dis. 2011, 15, e510–e513. [Google Scholar] [CrossRef]
- The Straits Times. Wuhan Virus: China Locks down Huanggang, Shuts Down Railway Station in Ezhou after Wuhan Lockdown. Available online: https://www.straitstimes.com/asia/east-asia/china-locks-down-two-more-cities-huanggang-and-ezhou-after-wuhan (accessed on 23 January 2020).
- Ji, W.; Wang, W.; Zhao, X.; Zai, J.; Li, X. Homologous recombination within the spike glycoprotein of the newly identified coronavirus may boost cross-species transmission from snake to human. J. Med. Virol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Robertson, D.; Jiang, X. nCoV’s Relationship to Bat Coronaviruses and Recombination Signals No Snakes. Available online: http://virological.org/t/ncovs-relationship-to-bat-coronaviruses-recombination-signals-no-snakes/331 (accessed on 23 January 2020).
- Kan, B.; Wang, M.; Jing, H.; Xu, H.; Jiang, X.; Yan, M.; Liang, W.; Zheng, H.; Wan, K.; Liu, Q.; et al. Molecular evolution analysis and geographic investigation of severe acute respiratory syndrome coronavirus-like virus in palm civets at an animal market and on farms. J. Virol. 2005, 79, 11892–11900. [Google Scholar] [CrossRef] [PubMed]
- Lau, S.K.; Woo, P.C.; Li, K.S.; Huang, Y.; Tsoi, H.W.; Wong, B.H.; Wong, S.S.; Leung, S.Y.; Chan, K.H.; Yuen, K.Y. Severe acute respiratory syndrome coronavirus-like virus in Chinese horseshoe bats. Proc. Natl. Acad. Sci. USA 2005, 102, 14040–14045. [Google Scholar] [CrossRef] [PubMed]
- Menachery, V.D.; Yount, B.L., Jr.; Debbink, K.; Agnihothram, S.; Gralinski, L.E.; Plante, J.A.; Graham, R.L.; Scobey, T.; Ge, X.Y.; Donaldson, E.F.; et al. A SARS-like cluster of circulating bat coronaviruses shows potential for human emergence. Nat. Med. 2015, 21, 1508–1513. [Google Scholar] [CrossRef]
- Menachery, V.D.; Yount, B.L., Jr.; Sims, A.C.; Debbink, K.; Agnihothram, S.S.; Gralinski, L.E.; Graham, R.L.; Scobey, T.; Plante, J.A.; Royal, S.R.; et al. SARS-like WIV1-CoV poised for human emergence. Proc. Natl. Acad. Sci. USA 2016, 113, 3048–3053. [Google Scholar] [CrossRef]
- Wang, N.; Li, S.Y.; Yang, X.L.; Huang, H.M.; Zhang, Y.J.; Guo, H.; Luo, C.M.; Miller, M.; Zhu, G.; Chmura, A.A.; et al. Serological Evidence of Bat SARS-Related Coronavirus Infection in Humans, China. Virol. Sin 2018, 33, 104–107. [Google Scholar] [CrossRef]
- Anthony, S.J.; Gilardi, K.; Menachery, V.D.; Goldstein, T.; Ssebide, B.; Mbabazi, R.; Navarrete-Macias, I.; Liang, E.; Wells, H.; Hicks, A.; et al. Further Evidence for Bats as the Evolutionary Source of Middle East Respiratory Syndrome Coronavirus. mBio 2017, 8. [Google Scholar] [CrossRef]
- Quan, P.L.; Firth, C.; Street, C.; Henriquez, J.A.; Petrosov, A.; Tashmukhamedova, A.; Hutchison, S.K.; Egholm, M.; Osinubi, M.O.; Niezgoda, M.; et al. Identification of a severe acute respiratory syndrome coronavirus-like virus in a leaf-nosed bat in Nigeria. mBio 2010, 1. [Google Scholar] [CrossRef]
- Azhar, E.I.; El-Kafrawy, S.A.; Farraj, S.A.; Hassan, A.M.; Al-Saeed, M.S.; Hashem, A.M.; Madani, T.A. Evidence for camel-to-human transmission of MERS coronavirus. N. Engl. J. Med. 2014, 370, 2499–2505. [Google Scholar] [CrossRef]
- Fehr, A.R.; Channappanavar, R.; Perlman, S. Middle East Respiratory Syndrome: Emergence of a Pathogenic Human Coronavirus. Annu. Rev. Med. 2017, 68, 387–399. [Google Scholar] [CrossRef] [PubMed]
- Hung, I.F.N.; Cheng, V.C.C.; Wu, A.K.L.; Tang, B.S.F.; Chan, K.H.; Chu, C.M.; Wong, M.M.L.; Hui, W.T.; Poon, L.L.M.; Tse, D.M.W.; et al. Viral Loads in Clinical Specimens and SARS Manifestations. Emerg. Infect. Dis. 2004, 10, 1550–1557. [Google Scholar] [CrossRef] [PubMed]
- Assiri, A.; Al-Tawfiq, J.A.; Al-Rabeeah, A.A.; Al-Rabiah, F.A.; Al-Hajjar, S.; Al-Barrak, A.; Flemban, H.; Al-Nassir, W.N.; Balkhy, H.H.; Al-Hakeem, R.F.; et al. Epidemiological, demographic, and clinical characteristics of 47 cases of Middle East respiratory syndrome coronavirus disease from Saudi Arabia: A descriptive study. Lancet Infect. Dis. 2013, 13, 752–761. [Google Scholar] [CrossRef]
- Rahman, A.; Sarkar, A. Risk Factors for Fatal Middle East Respiratory Syndrome Coronavirus Infections in Saudi Arabia: Analysis of the WHO Line List, 2013–2018. Am. J. Public Health 2019, 109, 1288–1293. [Google Scholar] [CrossRef]
- CIDRAP. SARS Severe Acute Respiratory Syndrome. Available online: http://www.cidrap.umn.edu/infectious-disease-topics/sars (accessed on 20 January 2020).
- CIDRAP. MERS-CoV. Available online: http://www.cidrap.umn.edu/infectious-disease-topics/mers-cov (accessed on 20 January 2020).
- Rambaut, A. Preliminary Phylogenetic Analysis of 11 nCoV2019 Genomes, 2020-01-19. Available online: http://virological.org/t/preliminary-phylogenetic-analysis-of-11-ncov2019-genomes-2020-01-19/329 (accessed on 20 January 2020).
- Bedford, T.; Neher, R. Genomic Epidemiology of Novel Coronavirus (nCoV) Using Data Generated by Fudan University, China CDC, Chinese Academy of Medical Sciences, Chinese Academy of Sciences and the Thai National Institute of Health Shared via GISAID. Available online: https://nextstrain.org/ncov (accessed on 20 January 2020).
- Shi, Z.-L.; Zhou, P.; Yang, X.-L.; Wang, X.-G.; Hu, B.; Zhang, L.; Zhang, W.; Si, H.-R.; Zhu, Y.; Li, B.; et al. Discovery of a novel coronavirus associated with the recent pneumonia outbreak in humans and its potential bat origin. bioRxiv 2020. [Google Scholar] [CrossRef]
- Menachery, V.D.; Graham, R.L.; Baric, R.S. Jumping species-a mechanism for coronavirus persistence and survival. Curr. Opin. Virol. 2017, 23, 1–7. [Google Scholar] [CrossRef]
- Becker, M.M.; Graham, R.L.; Donaldson, E.F.; Rockx, B.; Sims, A.C.; Sheahan, T.; Pickles, R.J.; Corti, D.; Johnston, R.E.; Baric, R.S.; et al. Synthetic recombinant bat SARS-like coronavirus is infectious in cultured cells and in mice. Proc. Natl. Acad. Sci. USA 2008, 105, 19944–19949. [Google Scholar] [CrossRef]
- Li, W.; Sui, J.; Huang, I.C.; Kuhn, J.H.; Radoshitzky, S.R.; Marasco, W.A.; Choe, H.; Farzan, M. The S proteins of human coronavirus NL63 and severe acute respiratory syndrome coronavirus bind overlapping regions of ACE2. Virology 2007, 367, 367–374. [Google Scholar] [CrossRef]
- Zeng, L.P.; Gao, Y.T.; Ge, X.Y.; Zhang, Q.; Peng, C.; Yang, X.L.; Tan, B.; Chen, J.; Chmura, A.A.; Daszak, P.; et al. Bat Severe Acute Respiratory Syndrome-Like Coronavirus WIV1 Encodes an Extra Accessory Protein, ORFX, Involved in Modulation of the Host Immune Response. J. Virol. 2016, 90, 6573–6582. [Google Scholar] [CrossRef]
- Xu, X.; Chen, P.; Wang, J.; Feng, J.; Zhou, H.; Li, X.; Zhong, W.; Hao, P. Evolution of the novel coronavirus from the ongoing Wuhan outbreak and modeling of its spike protein for risk of human transmission. Sci. China Life Sci. 2020. [Google Scholar] [CrossRef]
- Letko, M.C.; Munster, V. Functional assessment of cell entry and receptor usage for lineage B β-coronaviruses, including 2019-nCoV. bioRxiv 2020. [Google Scholar] [CrossRef]
- Rivers, T.M. Viruses and Koch’s Postulates. J. Bacteriol. 1937, 33, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Roberts, A.; Deming, D.; Paddock, C.D.; Cheng, A.; Yount, B.; Vogel, L.; Herman, B.D.; Sheahan, T.; Heise, M.; Genrich, G.L.; et al. A mouse-adapted SARS-coronavirus causes disease and mortality in BALB/c mice. PLoS Pathog. 2007, 3, e5. [Google Scholar] [CrossRef] [PubMed]
- McAuliffe, J.; Vogel, L.; Roberts, A.; Fahle, G.; Fischer, S.; Shieh, W.J.; Butler, E.; Zaki, S.; St Claire, M.; Murphy, B.; et al. Replication of SARS coronavirus administered into the respiratory tract of African Green, rhesus and cynomolgus monkeys. Virology 2004, 330, 8–15. [Google Scholar] [CrossRef] [PubMed]
- Kuiken, T.; Fouchier, R.A.; Schutten, M.; Rimmelzwaan, G.F.; van Amerongen, G.; van Riel, D.; Laman, J.D.; de Jong, T.; van Doornum, G.; Lim, W.; et al. Newly discovered coronavirus as the primary cause of severe acute respiratory syndrome. Lancet (Lond. Engl.) 2003, 362, 263–270. [Google Scholar] [CrossRef]
- Van Doremalen, N.; Miazgowicz, K.L.; Milne-Price, S.; Bushmaker, T.; Robertson, S.; Scott, D.; Kinne, J.; McLellan, J.S.; Zhu, J.; Munster, V.J. Host species restriction of Middle East respiratory syndrome coronavirus through its receptor, dipeptidyl peptidase 4. J. Virol. 2014, 88, 9220–9232. [Google Scholar] [CrossRef]
- Cockrell, A.S.; Peck, K.M.; Yount, B.L.; Agnihothram, S.S.; Scobey, T.; Curnes, N.R.; Baric, R.S.; Heise, M.T. Mouse dipeptidyl peptidase 4 is not a functional receptor for Middle East respiratory syndrome coronavirus infection. J. Virol. 2014, 88, 5195–5199. [Google Scholar] [CrossRef]
- De Wit, E.; Rasmussen, A.L.; Falzarano, D.; Bushmaker, T.; Feldmann, F.; Brining, D.L.; Fischer, E.R.; Martellaro, C.; Okumura, A.; Chang, J.; et al. Middle East respiratory syndrome coronavirus (MERS-CoV) causes transient lower respiratory tract infection in rhesus macaques. Proc. Natl. Acad. Sci. USA 2013, 110, 16598–16603. [Google Scholar] [CrossRef]
- Wen, X. 15 Medical Staff in Wuhan Confirmed New Coronavirus Pneumonia, and Another Suspected. Available online: http://china.qianlong.com/2020/0121/3600877.shtml (accessed on 21 January 2020).
- Al-Tawfiq, J.A.; Gautret, P. Asymptomatic Middle East Respiratory Syndrome Coronavirus (MERS-CoV) infection: Extent and implications for infection control: A systematic review. Travel Med. Infect. Dis. 2019, 27, 27–32. [Google Scholar] [CrossRef]
- Wong, G.; Liu, W.; Liu, Y.; Zhou, B.; Bi, Y.; Gao, G.F. MERS, SARS, and Ebola: The Role of Super-Spreaders in Infectious Disease. Cell Host Microbe 2015, 18, 398–401. [Google Scholar] [CrossRef]
- Park, D.; Huh, H.J.; Kim, Y.J.; Son, D.S.; Jeon, H.J.; Im, E.H.; Kim, J.W.; Lee, N.Y.; Kang, E.S.; Kang, C.I.; et al. Analysis of intrapatient heterogeneity uncovers the microevolution of Middle East respiratory syndrome coronavirus. Cold Spring Harb. Mol. Case Stud. 2016, 2, a001214. [Google Scholar] [CrossRef] [PubMed]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gralinski, L.E.; Menachery, V.D. Return of the Coronavirus: 2019-nCoV. Viruses 2020, 12, 135. https://doi.org/10.3390/v12020135
Gralinski LE, Menachery VD. Return of the Coronavirus: 2019-nCoV. Viruses. 2020; 12(2):135. https://doi.org/10.3390/v12020135
Chicago/Turabian StyleGralinski, Lisa E., and Vineet D. Menachery. 2020. "Return of the Coronavirus: 2019-nCoV" Viruses 12, no. 2: 135. https://doi.org/10.3390/v12020135
APA StyleGralinski, L. E., & Menachery, V. D. (2020). Return of the Coronavirus: 2019-nCoV. Viruses, 12(2), 135. https://doi.org/10.3390/v12020135




