Epidemiological, Molecular, and Clinical Features of Norovirus Infections among Pediatric Patients in Qatar
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection, Characteristic, and Follow-Up
2.2. Processing and Extraction of Viral RNA
2.3. NoV Sequencing and Genotyping
2.4. Vesikari Score System for Severity of NoV Infection
2.5. Statistical Analysis
3. Results
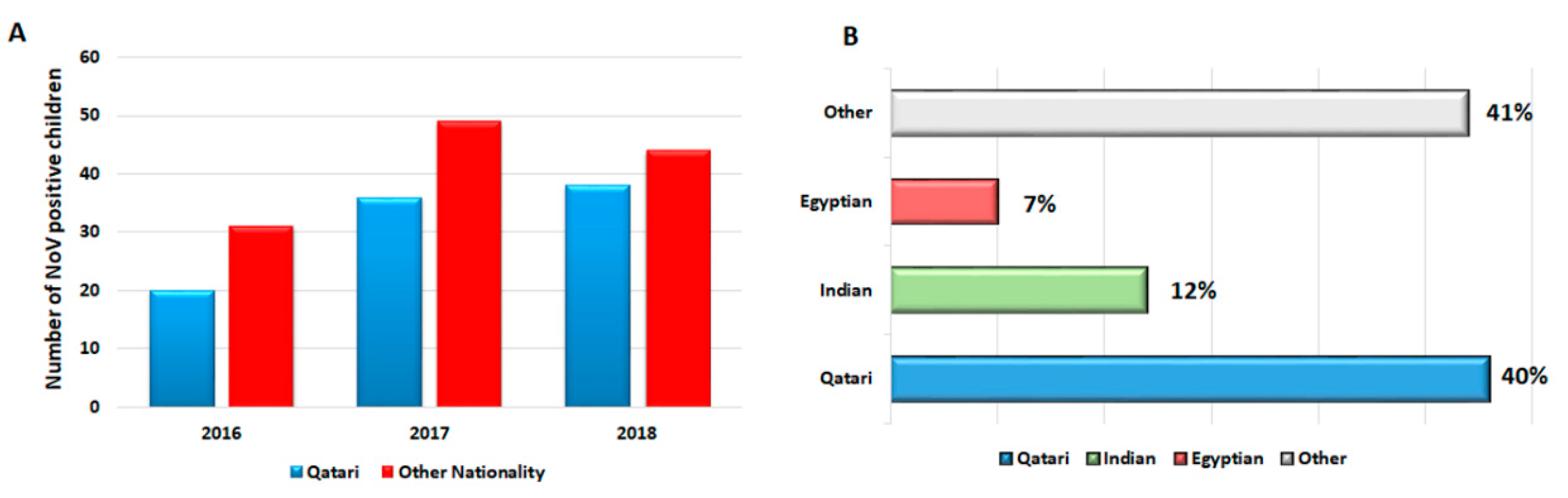
3.1. Demography and Clinical Outcomes of the Study Subjects
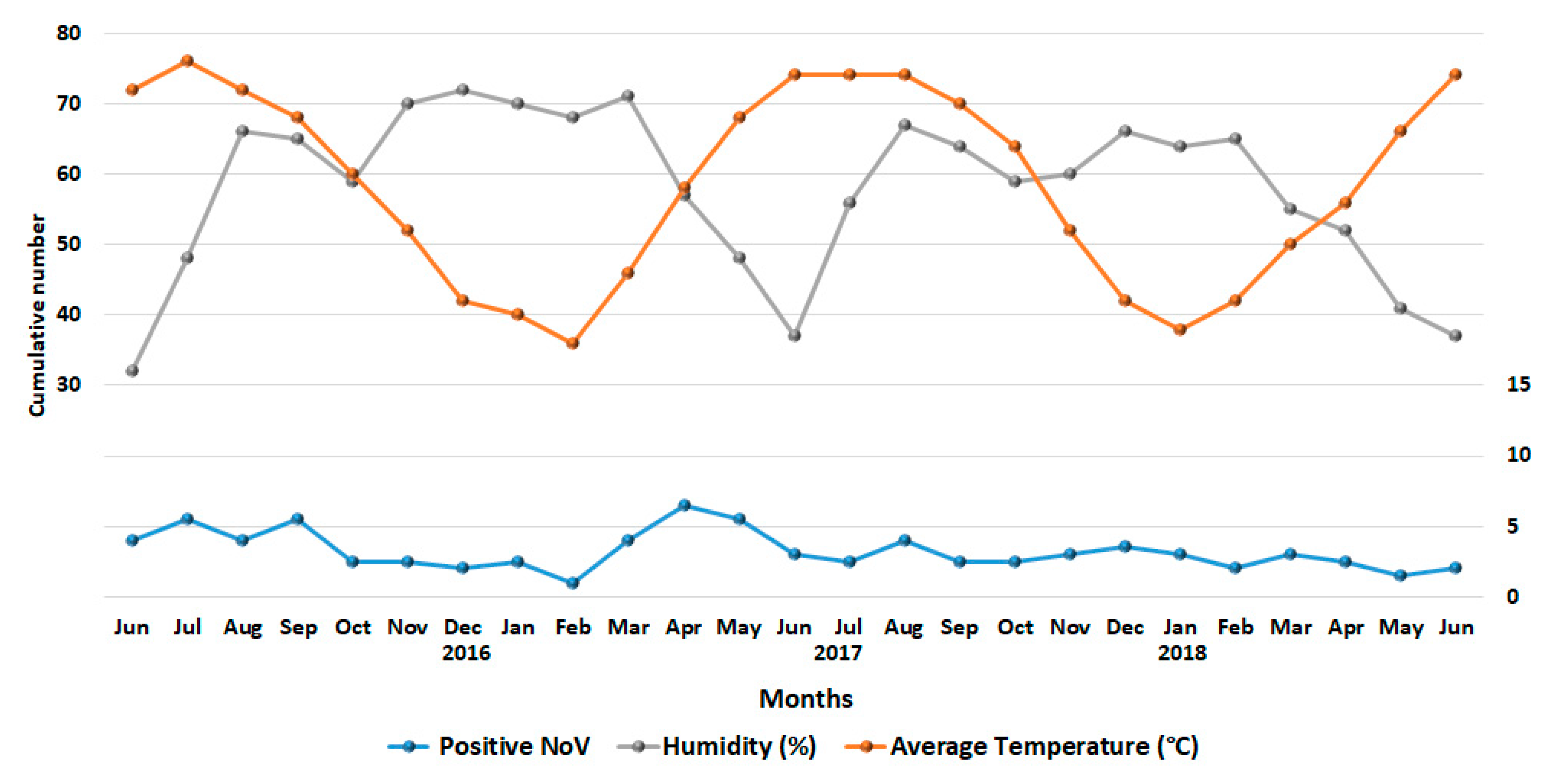
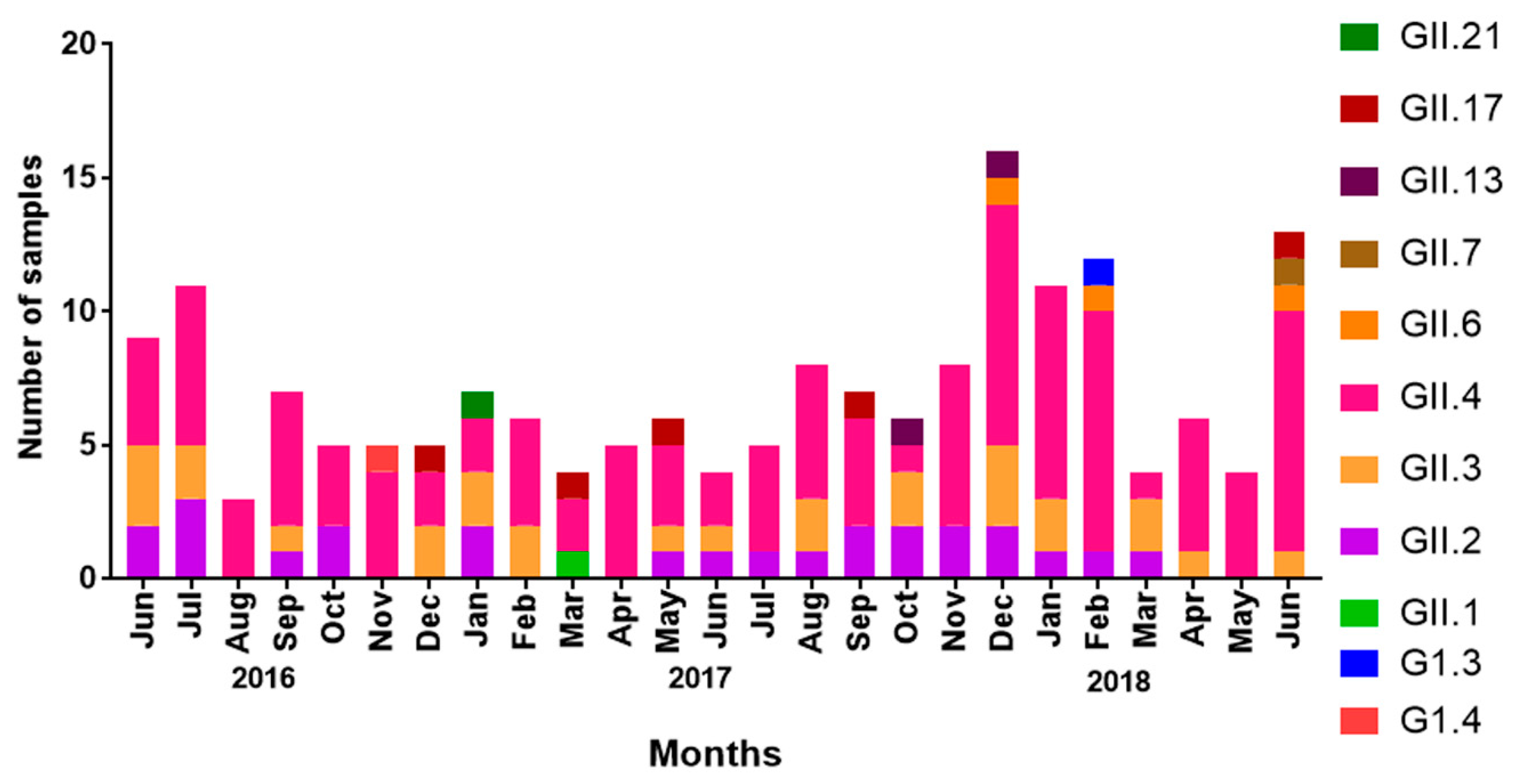
3.2. Seasonality of NoV Infections
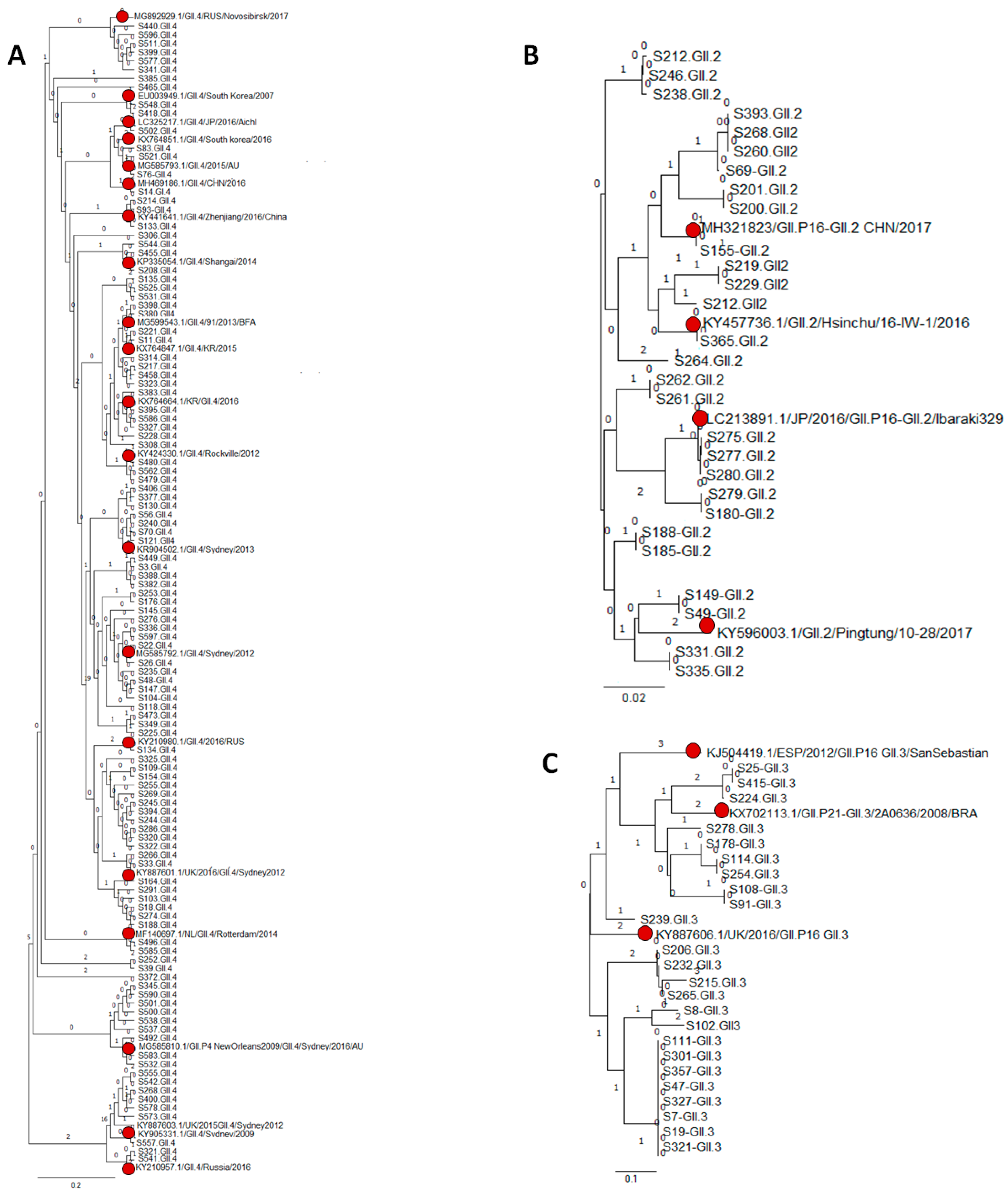
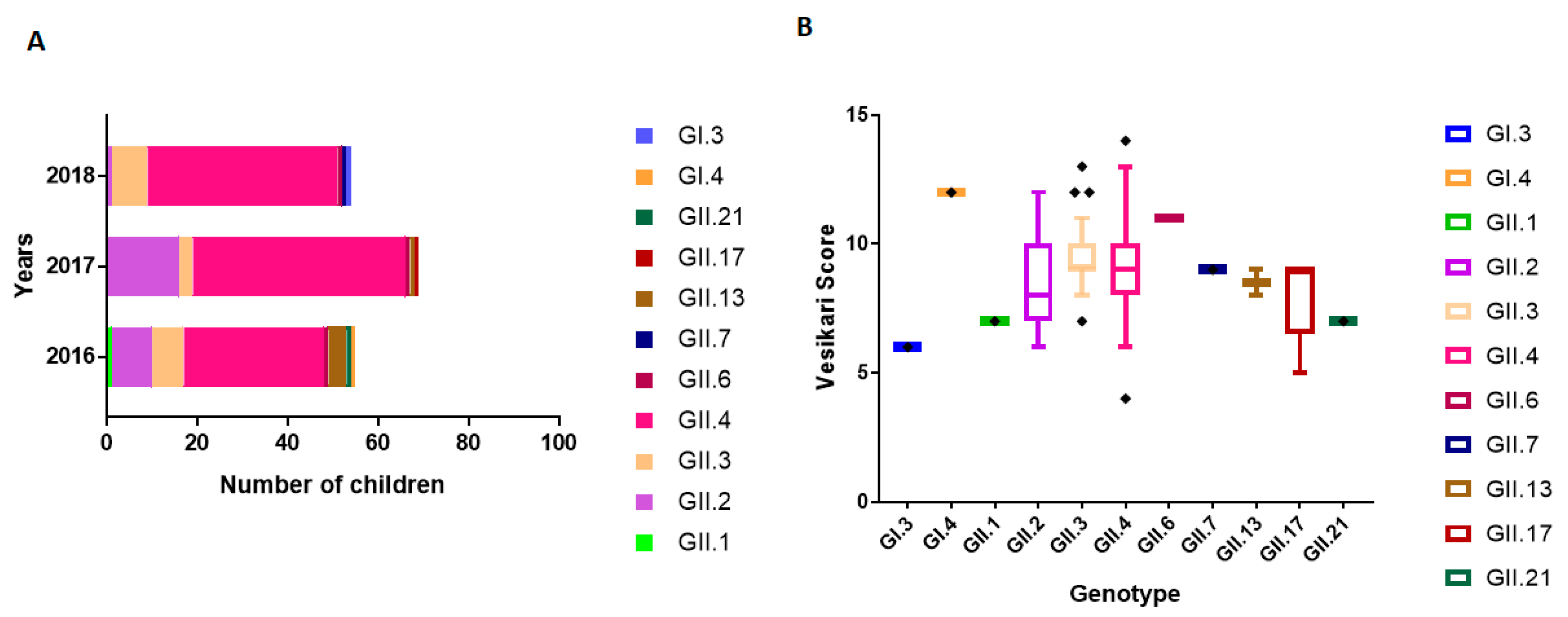
3.3. Genetic Variability of NoV Infections
3.4. NoV Genotypes and Age of Children
3.5. NoV Genotype-Associated Clinical Outcomes
3.6. Seasonal Distribution of NoV Genotypes
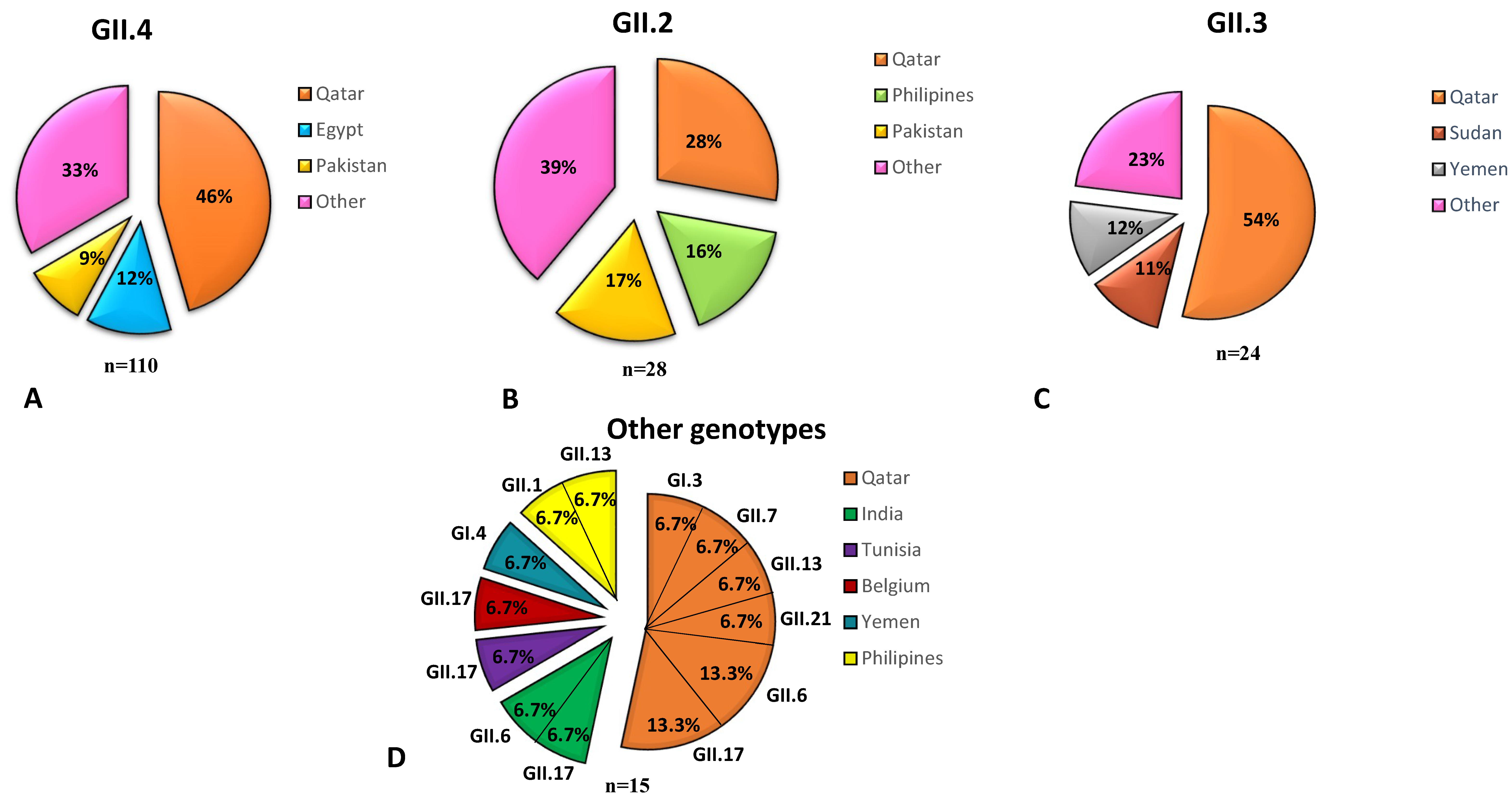
3.7. NoV Genotypes Associated with the Nationality of Children
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- CDC. Norovirus Worldwide. Available online: https://www.cdc.gov/norovirus/worldwide.html (accessed on 1 June 2018).
- Gray, J.J.; Kohli, E.; Ruggeri, F.M.; Vennema, H.; Sanchez-Fauquier, A.; Schreier, E.; Gallimore, C.I.; Iturriza-Gomara, M.; Giraudon, H.; Pothier, P.; et al. European multicenter evaluation of commercial enzyme immunoassays for detecting norovirus antigen in fecal samples. Clin. Vaccine Immunol. 2007, 14, 1349–1355. [Google Scholar] [CrossRef]
- Ettayebi, K.; Crawford, S.E.; Murakami, K.; Broughman, J.R.; Karandikar, U.; Tenge, V.R.; Neill, F.H.; Blutt, S.E.; Zeng, X.L.; Qu, L.; et al. Replication of human noroviruses in stem cell-derived human enteroids. Science 2016, 353, 1387–1393. [Google Scholar] [CrossRef] [PubMed]
- Jones, M.K.; Grau, K.R.; Costantini, V.; Kolawole, A.O.; de Graaf, M.; Freiden, P.; Graves, C.L.; Koopmans, M.; Wallet, S.M.; Tibbetts, S.A.; et al. Human norovirus culture in B cells. Nat. Protoc 2015, 10, 1939–1947. [Google Scholar] [CrossRef]
- Van Dycke, J.; Ny, A.; Conceicao-Neto, N.; Maes, J.; Hosmillo, M.; Cuvry, A.; Goodfellow, I.; de Araujo Nogueira, T.C.; Verbeken, E.; Matthijnssens, J.; et al. A robust human norovirus replication model in zebrafish larvae. BioRxiv 2019. [Google Scholar] [CrossRef]
- Fankhauser, R.L.; Monroe, S.S.; Noel, J.S.; Humphrey, C.D.; Bresee, J.S.; Parashar, U.D.; Ando, T.; Glass, R.I. Epidemiologic and molecular trends of "Norwalk-like viruses" associated with outbreaks of gastroenteritis in the United States. J. Infect. Dis. 2002, 186, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Kumazaki, M.; Usuku, S. Norovirus genotype distribution in outbreaks of acute gastroenteritis among children and older people: An 8-year study. Bmc Infect. Dis. 2016, 16, 643. [Google Scholar] [CrossRef] [PubMed]
- Miyazaki, N.; Taylor, D.W.; Hansman, G.S.; Murata, K. Antigenic and Cryo-Electron Microscopy Structure Analysis of a Chimeric Sapovirus Capsid. J. Virol. 2016, 90, 2664–2675. [Google Scholar] [CrossRef]
- Shanker, S.; Czako, R.; Sankaran, B.; Atmar, R.L.; Estes, M.K.; Prasad, B.V. Structural analysis of determinants of histo-blood group antigen binding specificity in genogroup I noroviruses. J. Virol. 2014, 88, 6168–6180. [Google Scholar] [CrossRef] [PubMed]
- Noel, J.S.; Fankhauser, R.L.; Ando, T.; Monroe, S.S.; Glass, R.I. Identification of a distinct common strain of "Norwalk-like viruses" having a global distribution. J. Infect. Dis. 1999, 179, 1334–1344. [Google Scholar] [CrossRef]
- Vinje, J.; Green, J.; Lewis, D.C.; Gallimore, C.I.; Brown, D.W.; Koopmans, M.P. Genetic polymorphism across regions of the three open reading frames of "Norwalk-like viruses". Arch. Virol. 2000, 145, 223–241. [Google Scholar] [CrossRef] [PubMed]
- Mattison, K.; Grudeski, E.; Auk, B.; Brassard, J.; Charest, H.; Dust, K.; Gubbay, J.; Hatchette, T.F.; Houde, A.; Jean, J.; et al. Analytical performance of norovirus real-time RT-PCR detection protocols in Canadian laboratories. J. Clin. Virol. 2011, 50, 109–113. [Google Scholar] [CrossRef]
- Mattison, K.; Grudeski, E.; Auk, B.; Charest, H.; Drews, S.J.; Fritzinger, A.; Gregoricus, N.; Hayward, S.; Houde, A.; Lee, B.E.; et al. Multicenter comparison of two norovirus ORF2-based genotyping protocols. J. Clin. Microbiol. 2009, 47, 3927–3932. [Google Scholar] [CrossRef]
- Trujillo, A.A.; McCaustland, K.A.; Zheng, D.P.; Hadley, L.A.; Vaughn, G.; Adams, S.M.; Ando, T.; Glass, R.I.; Monroe, S.S. Use of TaqMan real-time reverse transcription-PCR for rapid detection, quantification, and typing of norovirus. J. Clin. Microbiol. 2006, 44, 1405–1412. [Google Scholar] [CrossRef]
- Zheng, D.P.; Ando, T.; Fankhauser, R.L.; Beard, R.S.; Glass, R.I.; Monroe, S.S. Norovirus classification and proposed strain nomenclature. Virology 2006, 346, 312–323. [Google Scholar] [CrossRef] [PubMed]
- Vega, E.; Barclay, L.; Gregoricus, N.; Williams, K.; Lee, D.; Vinje, J. Novel surveillance network for norovirus gastroenteritis outbreaks, United States. Emerg. Infect. Dis. 2011, 17, 1389–1395. [Google Scholar] [CrossRef] [PubMed]
- Debbink, K.; Lindesmith, L.C.; Donaldson, E.F.; Costantini, V.; Beltramello, M.; Corti, D.; Swanstrom, J.; Lanzavecchia, A.; Vinje, J.; Baric, R.S. Emergence of new pandemic GII.4 Sydney norovirus strain correlates with escape from herd immunity. J. Infect. Dis. 2013, 208, 1877–1887. [Google Scholar] [CrossRef]
- Hasing, M.E.; Lee, B.E.; Preiksaitis, J.K.; Tellier, R.; Honish, L.; Senthilselvan, A.; Pang, X.L. Emergence of a new norovirus GII.4 variant and changes in the historical biennial pattern of norovirus outbreak activity in Alberta, Canada, from 2008 to 2013. J. Clin. Microbiol. 2013, 51, 2204–2211. [Google Scholar]
- Lindesmith, L.C.; Kocher, J.F.; Donaldson, E.F.; Debbink, K.; Mallory, M.L.; Swann, E.W.; Brewer-Jensen, P.D.; Baric, R.S. Emergence of Novel Human Norovirus GII.17 Strains Correlates With Changes in Blockade Antibody Epitopes. J. Infect. Dis. 2017, 216, 1227–1234. [Google Scholar] [CrossRef] [PubMed]
- Bull, R.A.; Eden, J.S.; Rawlinson, W.D.; White, P.A. Rapid evolution of pandemic noroviruses of the GII.4 lineage. Plos Pathog. 2010, 6, e1000831. [Google Scholar] [CrossRef]
- Boon, D.; Mahar, J.E.; Abente, E.J.; Kirkwood, C.D.; Purcell, R.H.; Kapikian, A.Z.; Green, K.Y.; Bok, K. Comparative Evolution of GII.3 and GII.4 Norovirus over a 31-Year Period. J. Virol. 2011, 85, 8656–8666. [Google Scholar]
- Chan-It, W.; Thongprachum, A.; Khamrin, P.; Kobayashi, M.; Okitsu, S.; Mizuguchi, M.; Ushijima, H. Emergence of a new norovirus GII.6 variant in Japan, 2008-2009. J. Med. Virol. 2012, 84, 1089–1096. [Google Scholar]
- Kreidieh, K.; Charide, R.; Dbaibo, G.; Melhem, N.M. The epidemiology of Norovirus in the Middle East and North Africa (MENA) region: A systematic review. Virol. J. 2017, 14, 220. [Google Scholar] [CrossRef]
- Rohayem, J. Norovirus seasonality and the potential impact of climate change. Clin. Microbiol. Infect. 2009, 15, 524–527. [Google Scholar] [CrossRef]
- Goodman, A. The Development of the Qatar Healthcare System: A Review of the Literature. Int. J. Clin. Med. 2015, 6, 177. [Google Scholar] [CrossRef]
- Al-Thani, A.; Baris, M.; Al-Lawati, N.; Al-Dhahry, S. Characterising the aetiology of severe acute gastroenteritis among patients visiting a hospital in Qatar using real-time polymerase chain reaction. BMC Infect. Dis. 2013, 13, 329. [Google Scholar] [CrossRef] [PubMed]
- Kojima, S.; Kageyama, T.; Fukushi, S.; Hoshino, F.B.; Shinohara, M.; Uchida, K.; Natori, K.; Takeda, N.; Katayama, K. Genogroup-specific PCR primers for detection of Norwalk-like viruses. J. Virol. Methods 2002, 100, 107–114. [Google Scholar] [CrossRef]
- Chen, Y.; Li, Z.; Han, D.; Cui, D.; Chen, X.; Zheng, S.; Yu, F.; Liu, J.; Lai, S.; Yan, Y.; et al. Viral agents associated with acute diarrhea among outpatient children in southeastern China. Pediatr. Infect. Dis. J. 2013, 32, e285–e290. [Google Scholar] [CrossRef] [PubMed]
- Available online: http://www.fhcrc.org/science/labs/hahn/methods/mol_bio_meth/Big%20Dye%20Protocol.pdf (accessed on 29 April 2019).
- Kroneman, A.; Vennema, H.; Deforche, K.; v d Avoort, H.; Penaranda, S.; Oberste, M.S.; Vinje, J.; Koopmans, M. An automated genotyping tool for enteroviruses and noroviruses. J. Clin. Virol. 2011, 51, 121–125. [Google Scholar] [CrossRef]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef] [PubMed]
- Vesikari, T.; Rautanen, T.; Varis, T.; Beards, G.M.; Kapikian, A.Z. Rhesus Rotavirus candidate vaccine. Clinical trial in children vaccinated between 2 and 5 months of age. Am. J. Dis. Child. 1990, 144, 285–289. [Google Scholar] [CrossRef]
- Mabasa, V.V.; Meno, K.D.; Taylor, M.B.; Mans, J. Environmental Surveillance for Noroviruses in Selected South African Wastewaters 2015-2016: Emergence of the Novel GII.17. Food Environ. Virol. 2018, 10, 16–28. [Google Scholar] [CrossRef]
- Kabue, J.P.; Meader, E.; Hunter, P.R.; Potgieter, N. Genetic characterisation of Norovirus strains in outpatient children from rural communities of Vhembe district/South Africa, 2014-2015. J. Clin. Virol. 2017, 94, 100–106. [Google Scholar] [CrossRef]
- Abugalia, M.; Cuevas, L.; Kirby, A.; Dove, W.; Nakagomi, O.; Nakagomi, T.; Kara, M.; Gweder, R.; Smeo, M.; Cunliffe, N. Clinical features and molecular epidemiology of rotavirus and norovirus infections in Libyan children. J. Med. Virol. 2011, 83, 1849–1856. [Google Scholar] [CrossRef]
- Rahouma, A.; Klena, J.D.; Krema, Z.; Abobker, A.A.; Treesh, K.; Franka, E.; Abusnena, O.; Shaheen, H.I.; El Mohammady, H.; Abudher, A.; et al. Enteric pathogens associated with childhood diarrhea in Tripoli-Libya. Am. J. Trop Med. Hyg 2011, 84, 886–891. [Google Scholar] [CrossRef]
- Muhsen, K.; Kassem, E.; Rubinstein, U.; Schachter, Y.; Kremer, A.; Goren, S.; Zilberstein, I.; Ephros, M.; Cohen, D.; Shulman, L.M. Incidence and characteristics of sporadic norovirus gastroenteritis associated with hospitalization of children less than 5 years of age in Israel. Pediatr. Infect. Dis. J. 2013, 32, 688–690. [Google Scholar] [CrossRef] [PubMed]
- Leshem, E.; Givon-Lavi, N.; Vinje, J.; Gregoricus, N.; Parashar, U.; Dagan, R. Differences in Norovirus-Associated Hospital Visits Between Jewish and Bedouin Children in Southern Israel. Pediatr. Infect. Dis. J. 2015, 34, 1036–1038. [Google Scholar] [CrossRef] [PubMed]
- Kaplan, N.M.; Kirby, A.; Abd-Eldayem, S.A.; Dove, W.; Nakagomi, T.; Nakagomi, O.; Cunliffe, N.A. Detection and molecular characterisation of rotavirus and norovirus infections in Jordanian children with acute gastroenteritis. Arch. Virol. 2011, 156, 1477–1480. [Google Scholar] [CrossRef] [PubMed]
- El-Mohammady, H.; Mansour, A.; Shaheen, H.I.; Henien, N.H.; Motawea, M.S.; Raafat, I.; Moustafa, M.; Adib-Messih, I.A.; Sebeny, P.J.; Young, S.Y.; et al. Increase in the detection rate of viral and parasitic enteric pathogens among Egyptian children with acute diarrhea. J. Infect. Dev. Ctries 2012, 6, 774–781. [Google Scholar] [CrossRef] [PubMed]
- Sdiri-Loulizi, K.; Hassine, M.; Gharbi-Khelifi, H.; Aouni, Z.; Chouchane, S.; Sakly, N.; Neji-Guediche, M.; Pothier, P.; Ambert-Balay, K.; Aouni, M. Molecular detection of genogroup I sapovirus in Tunisian children suffering from acute gastroenteritis. Virus Genes 2011, 43, 6–12. [Google Scholar] [CrossRef]
- Ben Salem-Ben Nejma, I.; Hassine Zaafrane, M.; Hassine, F.; Sdiri-Loulizi, K.; Ben Said, M.; Aouni, M.; Mzoughi, R. Etiology of Acute Diarrhea in Tunisian Children with Emphasis on Diarrheagenic Escherichia coli: Prevalence and Identification of E. coli Virulence Markers. Iran. J. Public Health 2014, 43, 947–960. [Google Scholar] [PubMed]
- Najafi, A.; Najafi, S.; Vahdat, K.; Kargar, M.; Javdani, N. Importance of viral pathogens in children with acute gastroenteritis in the south of Iran. Ann. Saudi Med. 2013, 33, 124–129. [Google Scholar] [CrossRef] [PubMed]
- El Qazoui, M.; Oumzil, H.; Baassi, L.; El Omari, N.; Sadki, K.; Amzazi, S.; Benhafid, M.; El Aouad, R. Rotavirus and norovirus infections among acute gastroenteritis children in Morocco. BMC Infect. Dis. 2014, 14, 1471–2334. [Google Scholar] [CrossRef] [PubMed]
- Melhem, N.M.; Zaraket, H.; Kreidieh, K.; Ali, Z.; Hammadi, M.; Ghanem, S.; Hajar, F.; Haidar, A.; Inati, A.; Rajab, M.; et al. Clinical and epidemiological characteristics of norovirus gastroenteritis among hospitalized children in Lebanon. World J. Gastroenterol. 2016, 22, 10557–10565. [Google Scholar] [CrossRef] [PubMed]
- Bicer, S.; Col, D.; Erdag, G.C.; Giray, T.; Gurol, Y.; Yilmaz, G.; Vitrinel, A.; Ozelgun, B. A retrospective analysis of acute gastroenteritis agents in children admitted to a university hospital pediatric emergency unit. Jundishapur J. Microbiol. 2014, 7, 1. [Google Scholar]
- Hoa Tran, T.N.; Trainor, E.; Nakagomi, T.; Cunliffe, N.A.; Nakagomi, O. Molecular epidemiology of noroviruses associated with acute sporadic gastroenteritis in children: Global distribution of genogroups, genotypes and GII.4 variants. J. Clin. Virol. 2013, 56, 185–193. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, S.M.; Lopman, B.A.; Levy, K. A systematic review and meta-analysis of the global seasonality of norovirus. PLoS ONE 2013, 8, e75922. [Google Scholar] [CrossRef] [PubMed]
- Oldak, E.; Sulik, A.; Rozkiewicz, D.; Liwoch-Nienartowicz, N. Norovirus infections in children under 5 years of age hospitalized due to the acute viral gastroenteritis in northeastern Poland. Eur. J. Clin. Microbiol. Infect. Dis. 2012, 31, 417–422. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Hullegie, S.; Bruijning-Verhagen, P.; Uiterwaal, C.S.P.M.; van der Ent, C.K.; Smit, H.A.; de Hoog, M.L.A. First-year Daycare and Incidence of Acute Gastroenteritis. Pediatrics 2016, 137, e20153356. [Google Scholar] [CrossRef] [PubMed]
- Lun, J.H.; Hewitt, J.; Yan, G.J.H.; Enosi Tuipulotu, D.; Rawlinson, W.D.; White, P.A. Recombinant GII.P16/GII.4 Sydney 2012 Was the Dominant Norovirus Identified in Australia and New Zealand in 2017. Viruses 2018, 10, 548. [Google Scholar]
- Centers for Disease Control and Prevention (CDC). Emergence of new norovirus strain GII.4 Sydney-United States, 2012. Mmwr Morb. Mortal. Wkly. Rep. 2013, 62, 55. [Google Scholar]
- Eden, J.-S.; Hewitt, J.; Lim, K.L.; Boni, M.F.; Merif, J.; Greening, G.; Ratcliff, R.M.; Holmes, E.C.; Tanaka, M.M.; Rawlinson, W.D.; et al. The emergence and evolution of the novel epidemic norovirus GII.4 variant Sydney 2012. Virology 2014, 450–451, 106–113. [Google Scholar]
- Bennett, S.; MacLean, A.; Miller, R.S.; Aitken, C.; Gunson, R.N. Increased norovirus activity in Scotland in 2012 is associated with the emergence of a new norovirus GII.4 variant. Euro. Surveill. 2013, 18. [Google Scholar]
- Matsushima, Y.; Shimizu, T.; Ishikawa, M.; Komane, A.; Okabe, N.; Ryo, A.; Kimura, H.; Katayama, K.; Shimizu, H. Complete Genome Sequence of a Recombinant GII.P16-GII.4 Norovirus Detected in Kawasaki City, Japan, in 2016. Genome Announc 2016, 4, e01099-16. [Google Scholar]
- Donaldson, E.F.; Lindesmith, L.C.; Lobue, A.D.; Baric, R.S. Viral shape-shifting: Norovirus evasion of the human immune system. Nat. Rev. Microbiol. 2010, 8, 231–241. [Google Scholar] [CrossRef] [PubMed]
- Han, J.; Wu, X.; Chen, L.; Fu, Y.; Xu, D.; Zhang, P.; Ji, L. Emergence of norovirus GII.P16-GII.2 strains in patients with acute gastroenteritis in Huzhou, China, 2016-2017. BMC Infect. Dis. 2018, 18, 342. [Google Scholar]
- Bidalot, M.; Thery, L.; Kaplon, J.; De Rougemont, A.; Ambert-Balay, K. Emergence of new recombinant noroviruses GII.p16-GII.4 and GII.p16-GII.2, France, winter 2016 to 2017. Euro. Surveill. 2017, 22. [Google Scholar] [CrossRef]
- Ao, Y.; Cong, X.; Jin, M.; Sun, X.; Wei, X.; Wang, J.; Zhang, Q.; Song, J.; Yu, J.; Cui, J.; et al. Genetic Analysis of Reemerging GII.P16-GII.2 Noroviruses in 2016-2017 in China. J. Infect. Dis. 2018, 218, 133–143. [Google Scholar]
- Ao, Y.; Wang, J.; Ling, H.; He, Y.; Dong, X.; Wang, X.; Peng, J.; Zhang, H.; Jin, M.; Duan, Z. Norovirus GII.P16/GII.2-Associated Gastroenteritis, China, 2016. Emerg. Infect. Dis. 2017, 23, 1172–1175. [Google Scholar]
- Tohma, K.; Lepore, C.J.; Ford-Siltz, L.A.; Parra, G.I. Phylogenetic Analyses Suggest that Factors Other Than the Capsid Protein Play a Role in the Epidemic Potential of GII.2 Norovirus. mSphere 2017, 2, e00187-00117. [Google Scholar] [CrossRef] [PubMed]
- Ayouni, S.; Estienney, M.; Sdiri-Loulizi, K.; Ambert-Balay, K.; de Rougemont, A.; Aho, S.; Hammami, S.; Aouni, M.; Guediche, M.N.; Pothier, P.; et al. Relationship between GII.3 norovirus infections and blood group antigens in young children in Tunisia. Clin Microbiol Infect. 2015, 21, e871–e878. [Google Scholar] [CrossRef]
- Timurkan, M.O.; Aydin, H.; Aktas, O. Frequency and molecular characterization of human norovirus in Erzurum, Turkey. Turk. J. Med. Sci. 2017, 47, 960–966. [Google Scholar] [CrossRef]
- Lopman, B.; Vennema, H.; Kohli, E.; Pothier, P.; Sanchez, A.; Negredo, A.; Buesa, J.; Schreier, E.; Reacher, M.; Brown, D.; et al. Increase in viral gastroenteritis outbreaks in Europe and epidemic spread of new norovirus variant. Lancet 2004, 363, 682–688. [Google Scholar] [CrossRef]
- Kamel, A.H.; Ali, M.A.; El-Nady, H.G.; de Rougemont, A.; Pothier, P.; Belliot, G. Predominance and circulation of enteric viruses in the region of Greater Cairo, Egypt. J. Clin. Microbiol. 2009, 47, 1037–1045. [Google Scholar] [CrossRef]
- Available online: https://www.worldtravelguide.net/guides/middle-east/qatar/weather-climate-geography/ (accessed on 29 April 2019).
- Hassine-Zaafrane, M.; Sdiri-Loulizi, K.; Kaplon, J.; Salem, I.B.; Pothier, P.; Aouni, M.; Ambert-Balay, K. Prevalence and genetic diversity of norovirus infection in Tunisian children (2007-2010). J. Med. Virol. 2013, 85, 1100–1110. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.Y.; Feng, Y.; Chao, H.C.; Lai, M.W.; Huang, W.L.; Lin, C.Y.; Tsai, C.N.; Chen, C.L.; Chiu, C.H. Emergence in Taiwan of novel norovirus GII.4 variants causing acute gastroenteritis and intestinal haemorrhage in children. J. Med. Microbiol. 2015, 64, 544–550. [Google Scholar] [CrossRef] [PubMed]
- Huhti, L.; Szakal, E.D.; Puustinen, L.; Salminen, M.; Huhtala, H.; Valve, O.; Blazevic, V.; Vesikari, T. Norovirus GII-4 causes a more severe gastroenteritis than other noroviruses in young children. J. Infect. Dis. 2011, 203, 1442–1444. [Google Scholar] [CrossRef] [PubMed]
- Desai, R.; Hembree, C.D.; Handel, A.; Matthews, J.E.; Dickey, B.W.; McDonald, S.; Hall, A.J.; Parashar, U.D.; Leon, J.S.; Lopman, B. Severe outcomes are associated with genogroup 2 genotype 4 norovirus outbreaks: A systematic literature review. Clin. Infect. Dis. 2012, 55, 189–193. [Google Scholar] [CrossRef] [PubMed]
- Siebenga, J.J.; Beersma, M.F.; Vennema, H.; van Biezen, P.; Hartwig, N.J.; Koopmans, M. High prevalence of prolonged norovirus shedding and illness among hospitalized patients: A model for in vivo molecular evolution. J. Infect. Dis. 2008, 198, 994–1001. [Google Scholar] [CrossRef] [PubMed]
- Lindesmith, L.C.; Donaldson, E.F.; Lobue, A.D.; Cannon, J.L.; Zheng, D.P.; Vinje, J.; Baric, R.S. Mechanisms of GII.4 norovirus persistence in human populations. PLoS Med. 2008, 5, e31. [Google Scholar] [CrossRef] [PubMed]
- Parra, G.I.; Squires, R.B.; Karangwa, C.K.; Johnson, J.A.; Lepore, C.J.; Sosnovtsev, S.V.; Green, K.Y. Static and Evolving Norovirus Genotypes: Implications for Epidemiology and Immunity. PLoS Pathog. 2017, 13, e1006136. [Google Scholar] [CrossRef]
- Available online: https://www.cdc.gov/norovirus/downloads/global-burden-report.pdf (accessed on 29 April 2019).
- Todd E, G.J. Viruses of foodborne origin: A review. Dovepress 2014, 7, 25–45. [Google Scholar] [CrossRef]
- Available online: http://stat.wto.org/CountryProfile/WSDBCountryPFView.aspx?Language=F&Country=QA (accessed on 29 April 2019).
| Pediatric Profile | NoV-Positive Cases No. (%) | NoV-Negative Cases No. (%) | p-Value |
|---|---|---|---|
| No. of Pediatric Patients | 177 (29.5) | 423 (70.5) | - |
| Gender | |||
| Female | 77 (43.5) | 235 (55.5) | 0.0070 |
| Male | 100 (56.5) | 188 (44.5) | |
| Age (years) | |||
| <1 year | 73 (41.2) | 97 (23) | <0.0001 |
| 1–3 years | 88 (49.7) | 209 (49.4) | |
| >3 years | 16 (9.1) | 117 (27.6) | |
| NoV-Positive No. (%) | GI No. (%) | GII No. (%) | |
|---|---|---|---|
| Fever | |||
| Yes | 70 (39.5) | 0 (0) | 70 (40) |
| No | 107(60.5) | 2(100) | 105(60) |
| Max reached fever (°C) | 40.6 | 38.6 | 39.9 |
| Vesikari Score | 7 ≤ scores ≤ 10 | score < 7 | score > 10 |
| Vomiting | |||
| Yes | 165 (93.2) | 2(100.0) | 163(93.1) |
| Frequency (per day) | 2–11 | 2–6 | 3–11 |
| Duration (days) | 1–7 | 3–4 | 1–7 |
| No | 12(6.8) | 0 (0.0) | 12 (6.9) |
| Vesikari Score * | score > 10 | score < 7 | score > 10 |
| Diarrhea | |||
| Yes | 176 (99.4) | 2(100.0) | 174 (99.4) |
| Frequency (per day) | 2–10 | 2–5 | 3–10 |
| Duration (days) | 1–5 | 3–4 | 2–5 |
| No Diarrhea | 1 (0.6) | 0 (0.0) | 1 (0.6) |
| Vesikari Score * | score > 10 | (score < 7) | (score > 10) |
| Dehydration | |||
| Severe | 8 (4.5) | 0 (0.0) | 8 (4.6) |
| Moderate | 76 (43) | 0 (0.0) | 76(43.4) |
| Mild | 89 (50.2) | 2(100.0) | 87 (49.7) |
| No dehydration | 4(2.3) | 0 (0.0) | 4 (2.3) |
| Vesikari Score * | 7 ≤ scores ≤ 10 | (score < 7) | (score > 10) |
| Mean Duration of Hospitalization (days) | 1–7 | 2–3 | 1–7 |
| Treatment ** | 24 (13.6) | 0 (0.0) | 24 (13.6) |
| Fully recovered | 22 (91.7) | 0 (0.0) | 22 (91.7) |
| Partially recovered | 2 (8.3) | 0 (0.0) | 2 (8.3) |
| Worsening | 0 (0.0) | 0 (0.0) | 0 (0.0) |
| Recovered but became sick again | 0 (0.0) | 0 (0.0) | 0 (0.0) |
| No treatment | 153 (86.4) | 2(100.0) | 151 (85.3) |
| Fully recovered | 129 (84.3) | 2(100.0) | 127 (82.3) |
| Partial recovered | 24 (15.7) | 0 (0.0) | 24 (15.7) |
| Worsening | 0 (0.0) | 0 (0.0) | 0 (0.0) |
| Recovered but became sick again | 0 (0.0) | 0 (0.0) | 0 (0.0) |
| Genotype | Total Number | <1 Year | 1–3 Years | <1 Year |
|---|---|---|---|---|
| Genotype I | ||||
| GI.4 | 1 (0.56%) | 1 | 0 | 0 |
| GI.3 | 1 (0.56%) | 0 | 1 | 0 |
| Genotype II | ||||
| GII.4 | 110 (62.2%) | 45 | 60 | 5 |
| All non-GII.4 GII | 65 (36.7%) | 27 | 27 | 11 |
| GII.1 | 1 (0.56%) | 1 | 0 | 0 |
| GII.3 | 28 (15.8%) | 5 | 19 | 4 |
| GII.2 | 24 (13.5%) | 18 | 4 | 2 |
| GII.6 | 3 (1.69%) | 1 | 1 | 1 |
| GII.7 | 1 (0.56%) | 0 | 1 | 0 |
| GII.13 | 2 (1.1%) | 1 | 0 | 1 |
| GII.17 | 5 (2.8%) | 1 | 2 | 2 |
| GII.21 | 1 (0.56%) | 1 | 0 | 0 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mathew, S.; Alansari, K.; K. Smatti, M.; Zaraket, H.; Al Thani, A.A.; Yassine, H.M. Epidemiological, Molecular, and Clinical Features of Norovirus Infections among Pediatric Patients in Qatar. Viruses 2019, 11, 400. https://doi.org/10.3390/v11050400
Mathew S, Alansari K, K. Smatti M, Zaraket H, Al Thani AA, Yassine HM. Epidemiological, Molecular, and Clinical Features of Norovirus Infections among Pediatric Patients in Qatar. Viruses. 2019; 11(5):400. https://doi.org/10.3390/v11050400
Chicago/Turabian StyleMathew, Shilu, Khalid Alansari, Maria K. Smatti, Hassan Zaraket, Asmaa A. Al Thani, and Hadi M. Yassine. 2019. "Epidemiological, Molecular, and Clinical Features of Norovirus Infections among Pediatric Patients in Qatar" Viruses 11, no. 5: 400. https://doi.org/10.3390/v11050400
APA StyleMathew, S., Alansari, K., K. Smatti, M., Zaraket, H., Al Thani, A. A., & Yassine, H. M. (2019). Epidemiological, Molecular, and Clinical Features of Norovirus Infections among Pediatric Patients in Qatar. Viruses, 11(5), 400. https://doi.org/10.3390/v11050400





