Codon Usage Bias Analysis of Citrus tristeza virus: Higher Codon Adaptation to Citrus reticulata Host
Abstract
1. Introduction
2. Materials and Methods
2.1. Dataset
2.2. Nucleotide Composition Analysis and Effective Number of Codons (ENc)
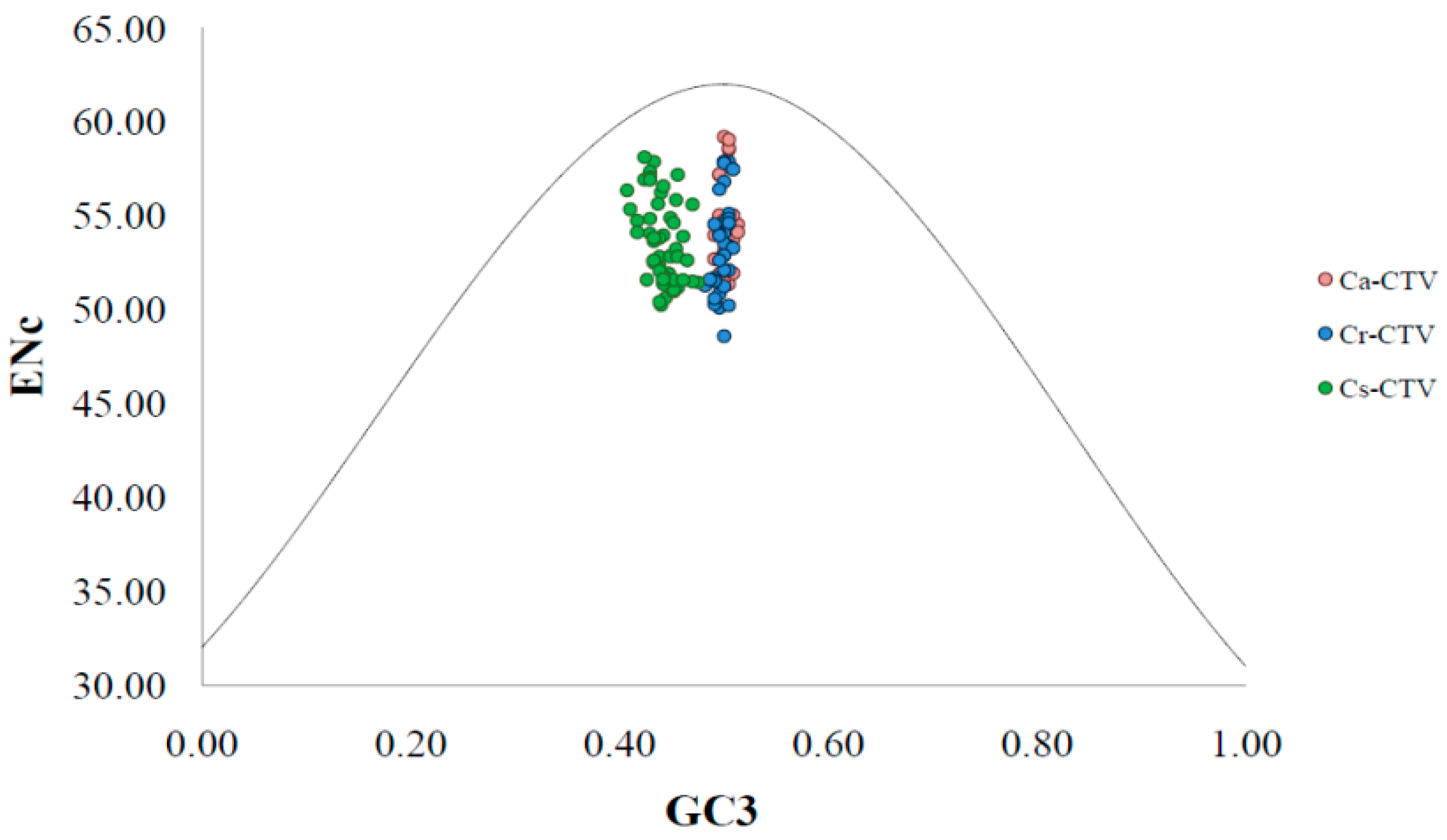
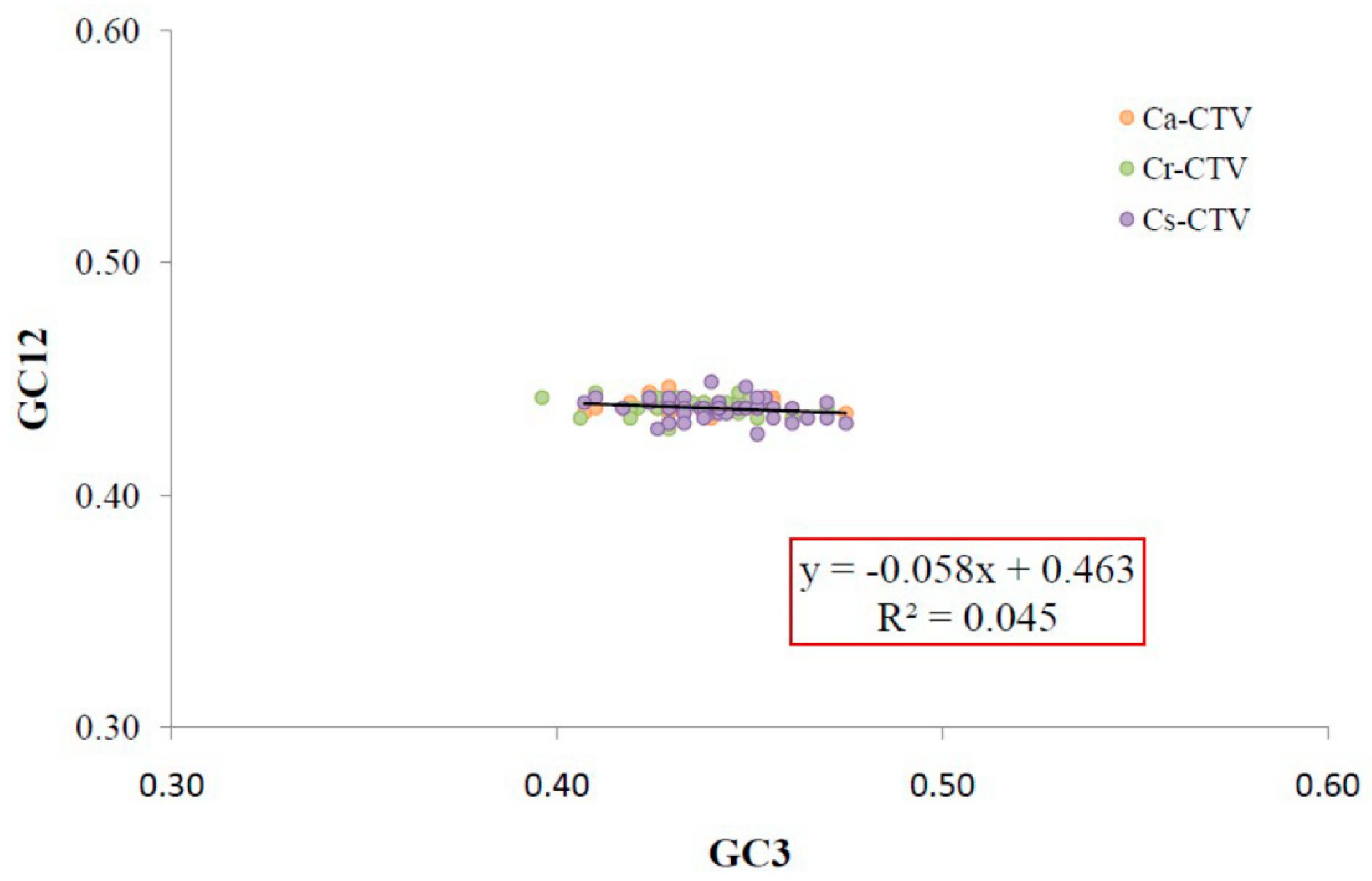
2.3. ENc-GC3 Plot and Neutrality Plot
2.4. Relative Synonymous Codon Usage (RSCU) and Contribution of High-Frequency Codon (CHFC)
2.5. Codon Adaptation Index (CAI)
2.6. Correspondence Analysis (COA)
2.7. Statistical Analysis
3. Results and Discussion
3.1. Preference of G/U-Ended Codon Over A/C-Ended Codon in the AU-Rich CTV CP Gene
3.2. CTVCP Gene Displays Low Codon Usage Bias (CUB) and Higher Genomic Stability
3.3. Natural Selection and Mutation Pressure Both Play Roles in Codon Usage Bias of CTV
3.4. Natural Selection Plays Key Role in Shaping the Codon Usage Bias of CTV
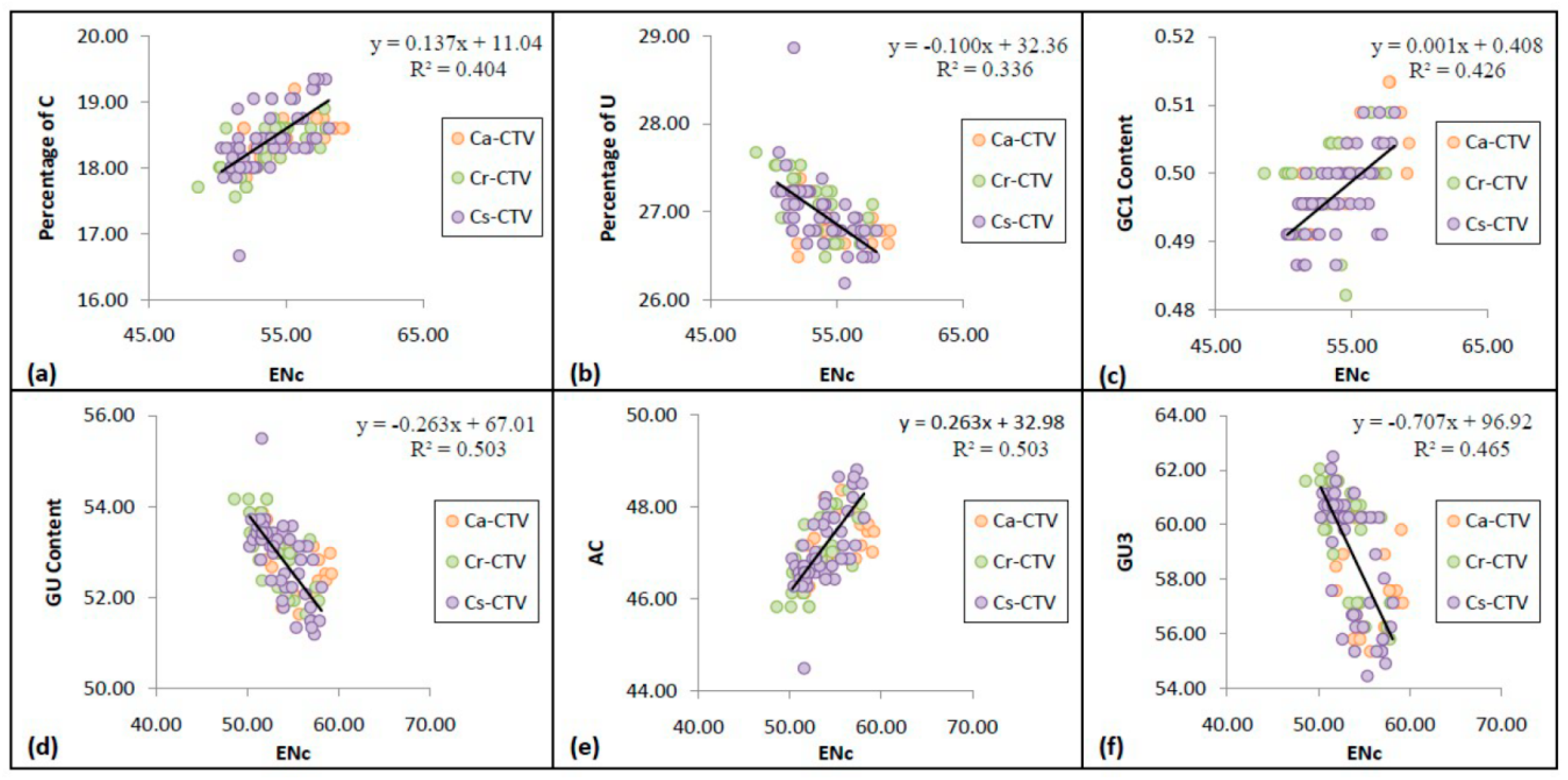
3.5. Codon Usage Bias Has Significant Correlation with the Nucleotide Compositional Constraint in CTV
3.6. Higher Codon Usage Variation in Ca-CTV Subgroup
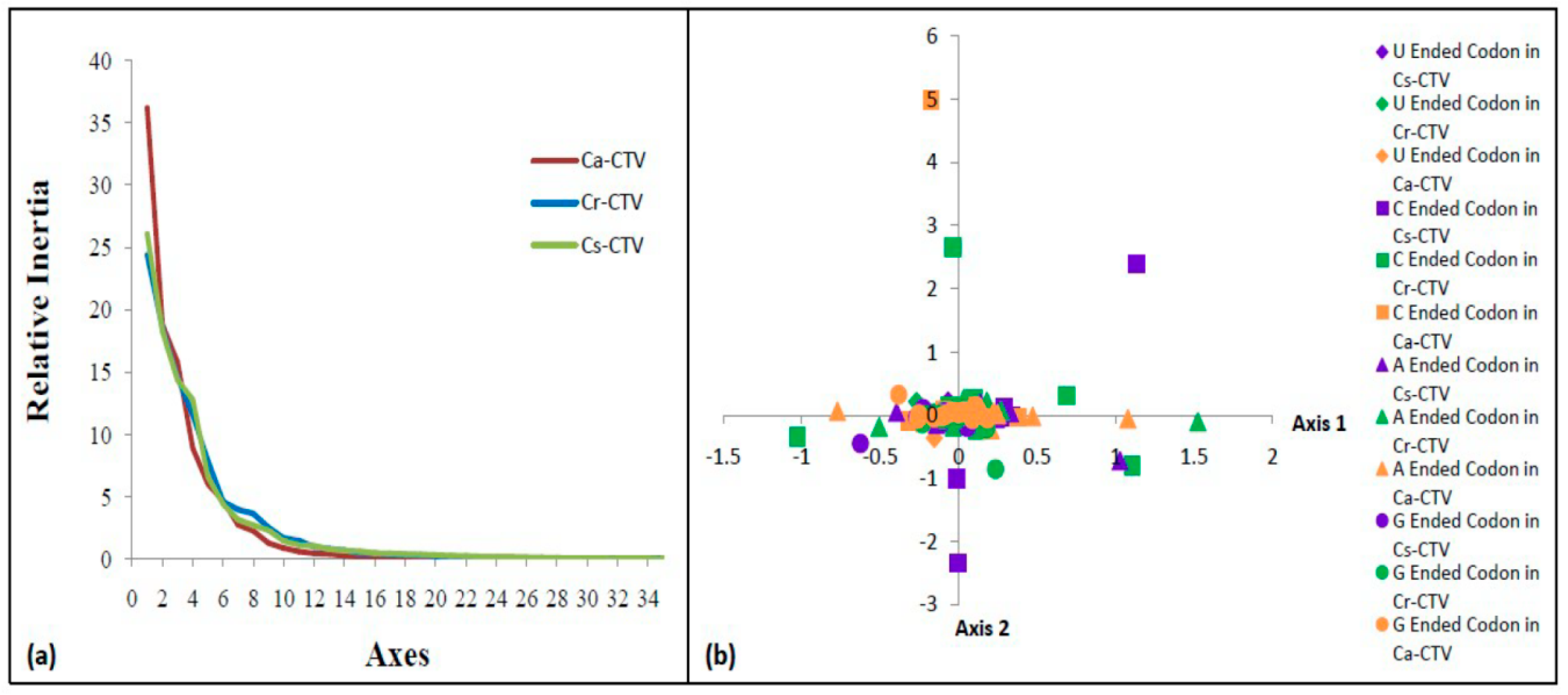
3.7. G/U-Ended Codons Display Higher Influence on Codon Usage of CTV
3.8. The High-Frequency Codons Are Evolutionarily Conserved in CTV
3.9. The Codon Usage Pattern of Cs-CTV Is Different from That of Ca-and Cr-CTV
3.10. CTV CP Gene Exhibits Higher Codon Usage Bias toward U-Ending Codons
3.11. CTV Is Biased toward Its Host Codon Usage Pattern
3.12. CTV Shows Mixture of Coincident and Antagonistic Codon Usage Patterns to Its Respective Host
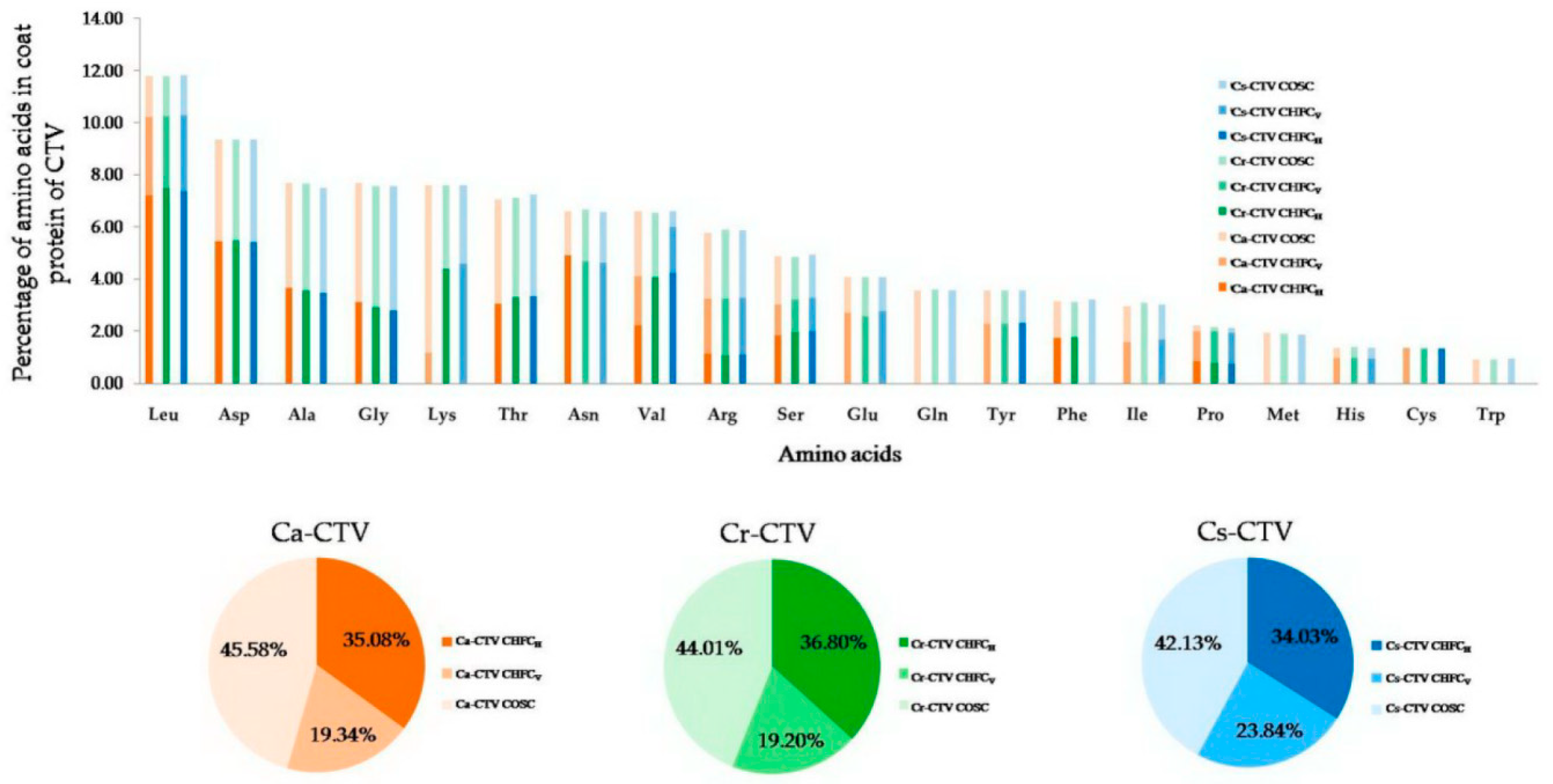
3.13. Host-Preferred High-Frequency Codons Exert Greater Effect on the Codon Usage of CTV
3.14. Varied Degrees of CTV Codon Usage Adaptation to Different Citrus Hosts
3.15. Cr-CTV Isolates Display Higher Codon Usage Adaptation to C. reticulata Host
3.16. The Correlation between Varied Magnitude of High-Frequency Codons and Host–Virus Interactions
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Rocha-Pena, M.A.; Lee, R.F.; Lastra, R.; Niblett, C.L.; Ochoa-Corona, F.M.; Garnsey, S.M.; Yokomi, R.K. Citrustristeza virus and its aphid vector Toxopteracitricida: Threats to citrus production in the Caribbean and Central and North America. Plant Dis. 1995, 79, 437–445. [Google Scholar] [CrossRef]
- Moreno, P.; Ambros, S.; Albiach-Marti, M.R.; Guerri, J.; Pena, L. Citrus tristeza virus: A pathogen that changed the course of the citrus industry. Mol. Plant Pathol. 2008, 9, 251–268. [Google Scholar] [CrossRef]
- Karasev, A.V.; Boyko, V.P.; Gowda, S.; Nikolaeva, O.V.; Hilf, M.E.; Koonin, E.V.; Niblett, C.L.; Cline, K.; Gumpf, D.J.; Lee, R.F.; et al. Complete sequence of the Citrus tristeza virus RNA genome. Virology 1995, 208, 511–520. [Google Scholar] [CrossRef] [PubMed]
- Satyanayanana, T.; Gowda, S.; Mawassi, M.; Albiach-Marti, M.R.; Ayllon, M.A.; Robertson, C.; Garnsey, S.M.; Dawson, W.O. Closterovirus encoded HSP70 homolog and p61 in addition to both coat proteins function inefficient virion assembly. Virology 2000, 278, 253–265. [Google Scholar] [CrossRef]
- Ahlawat, Y.S. Viruses, greening bacterium and viroids associated with citrus (Citrus species) decline in India. Indian J. Agric. Sci. 1997, 67, 51–57. [Google Scholar]
- Biswas, K.K. Molecular diagnosis of Citrus tristeza virus in mandarin (Citrus reticulata) orchards of hills of West Bengal. Indian J. Virol. 2008, 19, 26–31. [Google Scholar]
- Biswas, K.K. Molecular characterization of Citrus tristeza virus isolates from the Northeastern Himalayan region of India. Arch. Virol. 2010, 155, 959–963. [Google Scholar] [CrossRef] [PubMed]
- Sharma, S.K.; Tarafdar, A.; Khatun, D.; Sumita, K.; Biswas, K.K. Intra-farm diversity and evidence of genetic recombination of Citrus tristeza virus isolates in Delhi region of India. J. Plant Biochem. Biotech. 2012, 21, 38–43. [Google Scholar] [CrossRef]
- Biswas, K.K.; Tarafdar, A.; Diwedi, S.; Lee, R.F. Distribution, genetic diversity and recombination analysis of Citrus tristeza virus of India. Virus Genes 2012, 45, 139–148. [Google Scholar]
- Singh, J.K.; Tarafdar, A.; Sharma, S.K.; Biswas, K.K. Evidence of recombinant Citrus tristeza virus isolate occurring in acid lime cv. Pant Lemon orchard in Uttarakhand Terai Region of Northern Himalaya in India. Indian J. Virol. 2013, 24, 35–41. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Tarafdar, A.; Godara, S.; Dwivedi, S.; Jayakumar, B.K.; Biswas, K.K. Characterization of Citrus tristeza virus and determination of genetic variability in North-east and South India. Indian Phytopath. 2013, 66, 302–307. [Google Scholar]
- Palchoudhury, S.; Sharma, S.K.; Biswas, M.K.; Biswas, K.K. Diversified Citrus tristeza virus causing decline disease in Mandarin orange in Manipur state of NEH region in India. J. Mycol. Plant Pathol. 2015, 45, 317–323. [Google Scholar]
- Roy, A.; Brlansky, R.H. Genome analysis of an orange stem pitting Citrus tristeza virus isolate reveals a novel recombinant genotype. Virus Res. 2010, 151, 118–130. [Google Scholar] [CrossRef]
- Melzer, M.J.; Borth, W.B.; Sether, D.M.; Ferreira, S.; Gonsalves, D.; Hu, J.S. Genetic diversity and evidence for recent modular recombination in Hawaiian Citrus tristeza virus. Virus Genes 2010, 40, 111–118. [Google Scholar] [CrossRef]
- Biswas, K.K.; Tarafdar, A.; Sharma, S.K. Complete genome of mandarin decline Citrus tristeza virus of Northeastern Himalayan hill region of India: Comparative analyses determine recombinant. Arch. Virol. 2012, 157, 579–583. [Google Scholar] [CrossRef]
- Harper, S.J. Citrus tristeza virus: Evolution of complex and varied genotypic groups. Front. Microbiol. 2013, 4, 1–18. [Google Scholar] [CrossRef]
- Martin, S.; Sambade, A.; Rubio, L.; Vives, M.C.; Moya, P.; Guerri, J.; Elena, S.F.; Moreno, P. Contribution of recombination and selection to molecular evolution of Citrus tristeza virus. J. Gen. Virol. 2009, 90, 1527–1538. [Google Scholar] [CrossRef]
- Ikemura, T. Codon usage and tRNA content in unicellular and multicellular organisms. Mol. Biol. Evol. 1985, 2, 13–34. [Google Scholar]
- Sharp, P.M.; Averof, M.; Lloyd, A.T.; Matassi, G.; Peden, J.F. DNA sequence evolution: The sounds of silence. Phil. Trans. R. Soc. B. 1995, 349, 241–247. [Google Scholar]
- Shabalina, S.A.; Spridonov, N.A.; Kashina, A. Sounds of silence: Synonymous nucleotides as a key to biological regulation and complexity. Nucleic Acids Res. 2013, 41, 2073–2094. [Google Scholar] [CrossRef]
- Angellotti, M.C.; Bhuiyan, S.B.; Chen, G.; Wan, X.F. CodonO: Codon usage bias analysis within and across genomes. Nucleic Acids Res. 2007, 35, 132–136. [Google Scholar] [CrossRef]
- Butt, A.M.; Nasrullah, I.; Tong, Y. Genome-wide analysis of codon usage and influencing factors in chikungunya viruses. PLoS ONE 2014, 9. [Google Scholar] [CrossRef]
- Adams, M.J.; Antoniw, J.F. Codon usage bias amongst plant viruses. Arch. Virol. 2003, 149, 113–135. [Google Scholar]
- Xu, X.Z.; Liu, Q.P.; Fan, L.J.; Cui, X.F.; Zhou, X.P. Analysis of synonymous codon usage and evolution of begomoviruses. J. Zhejiang Univ. Sci. B 2008, 9, 667–674. [Google Scholar] [CrossRef]
- Cheng, X.F.; Wu, X.Y.; Wang, H.Z.; Sun, Y.Q.; Qian, Y.S.; Luo, L. High codon adaptation in Citrus tristeza virus to its citrus host. Virol. J. 2012, 9, 113. [Google Scholar] [CrossRef]
- Chakraborty, P.; Das, S.; Saha, B.; Sarkar, P.; Karmakar, A.; Saha, A.; Saha, D.; Saha, A. Phylogeny and synonymous codon usage pattern of Papaya ringspot virus coat protein gene in the sub-Himalayan region of north-east India. Can. J. Microbiol. 2015, 61, 555–564. [Google Scholar] [CrossRef]
- Sharp, P.M.; Li, W.H. Codon usage in regulatory genes in Escherichia coli does not reflect selection for ‘rare’ codons. Nucleic Acids Res. 1986, 14, 7737–7749. [Google Scholar] [CrossRef]
- Bulmer, M. The selection-mutation-drift theory of synonymous codon usage. Genetics 1991, 12, 897–907. [Google Scholar]
- Hershberg, R.; Petrov, D.A. Selection on codon bias. Annu. Rev. Genet. 2008, 42, 287–299. [Google Scholar] [CrossRef]
- Sharp, P.M.; Cowe, E. Synonymous codon usage in Saccharomyces cerevisiae. Yeast 1991, 7, 657–678. [Google Scholar] [CrossRef]
- Zhou, J.; Liu, W.J.; Peng, S.W.; Sun, X.Y.; Frazer, I. Papillomavirus capsid protein expression level depends on the match between codon usage and tRNA availability. J. Virol. 1999, 73, 4972–4982. [Google Scholar]
- Jenkins, G.M.; Holmes, E.C. The extent of codon usage bias in human RNA viruses and its evolutionary origin. Virus Res. 2003, 92, 1–7. [Google Scholar] [CrossRef]
- Tsai, C.T.; Lin, C.H.; Chang, C.Y. Analysis of codon usage bias and base compositional constraints in iridovirus genomes. Virus Res. 2007, 126, 196–206. [Google Scholar] [CrossRef]
- Bahir, I.; Fromer, M.; Prat, Y.; Linial, M. Viral adaptation to host: A proteome-based analysis of codon usage and amino acid preferences. Mol. Syst. Biol. 2009, 5, 311. [Google Scholar] [CrossRef]
- Biswas, K.K.; Tarafdar, A.; Sharma, S.K.; Singh, J.K.; Dwivedi, S.; Biswas, K.; Jayakumar, B.K. Current status of Citrus tristeza virus incidence and its spatial distribution in citrus growing geographical zones of India. Indian J. Agric. Sci. 2014, 84, 8–13. [Google Scholar]
- Athey, J.; Alexaki, A.; Osipova, E.; Rostovtsev, A.; Santana-Quintero, L.V.; Katneni, U.; Simonyan, V.; Kimchi-Sarfaty, C. A new and updated resource for codon usage tables. BMC Bioinform. 2017, 18, 391. [Google Scholar] [CrossRef]
- Peden, J.F. Analysis of Codon Usage. Master’s Thesis, Nottingham University, Nottingham, UK, 1999. [Google Scholar]
- Puigbo, P.; Bravo, I.G.; Garcia-Vallve, S. CAIcal: A combined set of tools to assess codon usage adaptation. Biol. Direct. 2008, 3, 38. [Google Scholar] [CrossRef]
- Wright, F. The ‘effective number of codons’ used in a gene. Gene 1990, 87, 23–29. [Google Scholar] [CrossRef]
- Sueoka, N. Directional mutation pressure and neutral molecular evolution. Proc. Natl. Acad. Sci. USA 1988, 85, 2653–2657. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis Version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef]
- Greenacre, M. Theory and Applications of Correspondence Analysis; Academic Press: London, UK, 1984. [Google Scholar]
- Kumar, N.; Bera, B.C.; Greenbaum, B.D.; Bhatia, S.; Sood, R.; Selvaraj, P.; Anand, T.; Tripathi, B.N.; Virmani, N. Revelation of influencing factors in overall codon usage bias of Equine influenza viruses. PLoS ONE 2016, 11, e0154376. [Google Scholar] [CrossRef]
- Butt, A.M.; Nasrullah, I.; Qamar, R.; Tong, Y. Evolution of codon usage in Zika virus genomes is host and vector specific. Emerg. Microbes Infect. 2016, 5, e107. [Google Scholar] [CrossRef]
- Cristina, J.; Moreno, P.; Moratorio, G.; Musto, H. Genome-wide analysis of codon usage bias in Ebolavirus. Virus Res. 2014, 196, 87–93. [Google Scholar] [CrossRef]
- Hu, J.S.; Wang, Q.Q.; Zhang, J.; Chen, H.T.; Xu, Z.W.; Ling, Z.; Ding, Y.Z.; Ma, L.N.; Xu, K.; Gu, Y.X. The characteristic of codon usage pattern and its evolution of hepatitis C virus. Infect. Genet. Evol. 2011, 11, 2098–2102. [Google Scholar] [CrossRef]
- Elena, S.F.; Sanjuán, R. Adaptive value of high mutation rates of RNA viruses: Separating causes from consequences. J. Virol. 2005, 79, 11555–11558. [Google Scholar] [CrossRef]
- Hemert, F.; van der Kuyl, A.C.; Berkhout, B. Impact of the biased nucleotide composition of viral RNA genomes on RNA structure and codon usage. J. Gen. Virol. 2016, 97, 2608–2619. [Google Scholar] [CrossRef]
- Cheng, X.; Virk, N.; Chen, W.; Ji, S.; Ji, S.; Sun, Y.; Wu, X. CpG usage in RNA viruses: Data and hypotheses. PLoS ONE 2013, 8, e0074109. [Google Scholar] [CrossRef]
- Ahmad, T.; Sablok, G.; Tatarinova, T.V.; Xu, Q.; Deng, X.X.; Guo, W.W. Evaluation of codon biology in Citrus and Poncirustrifoliata based on genomic features and frame corrected expressed sequence tags. DNA Res. 2013, 20, 135–150. [Google Scholar] [CrossRef]
- Sanchez, G.; Bosch, A.; Pinto, R.M. Genome variability and capsid structural constraints of hepatitis a virus. J. Virol. 2003, 77, 452–459. [Google Scholar] [CrossRef]
- Mueller, S.; Papamichail, D.; Coleman, J.R.; Skiena, S.; Wimmer, E. Reduction of the rate of poliovirus protein synthesis through large-scale codon deoptimization causes attenuation of viral virulence by lowering specific infectivity. J. Virol. 2006, 80, 9687–9696. [Google Scholar] [CrossRef]
- Wu, G.A.; Prochnik, S.; Jenkins, J.; Salse, J.; Hellsten, U.; Florent, M.; Perrier, X.; Ruiz, M.; Scalabrin, S.; Terol, J.F. Sequencing of diverse mandarin, pummelo and orange genomes reveals complex history of admixture during citrus domestication. Nat. Biotechnol. 2014, 32, 656–662. [Google Scholar] [CrossRef]
- Silva, G.; Marques, N.; Nolasco, G. The evolutionary rate of Citrus tristeza virus ranks among the rates of the slowest RNA viruses. J. Gen. Virol. 2012, 93, 419–429. [Google Scholar] [CrossRef]
- Dawson, W.O.; Garnsey, S.M.; Tatineni, S.; Folimonova, S.Y.; Harper, S.J.; Gowda, S. Citrustristeza virus-host interactions. Front Microbiol. 2013, 4, 1–10. [Google Scholar] [CrossRef]
| Nt Composition | Ca-CTV | Cr-CTV | Cs-CTV | Overall CTV |
|---|---|---|---|---|
| A% | 28.78 (0.10) | 28.72 (0.26) | 28.71 (0.48) | 28.73 (0.05) |
| C% | 18.41 (0.36) | 18.30 (0.51) | 18.42 (0.66) | 18.38 (0.04) |
| U% | 26.91 (0.09) | 27.06 (0.29) | 26.98 (0.51) | 26.99 (0.03) |
| G% | 25.90 (0.15) | 25.92 (0.33) | 25.88 (0.53) | 25.90 (0.05) |
| GC% | 44.31 (0.53) | 44.23 (0.18) | 44.30 (0.20) | 44.28 (0.05) |
| AU% | 55.69 (0.91) | 55.77 (0.47) | 55.70 (0.07) | 55.72 (0.05) |
| A3% | 21.09 (0.40) | 20.64 (0.53) | 20.69 (0.65) | 20.77 (0.14) |
| C3% | 20.26 (0.33) | 19.92 (0.49) | 20.40 (0.64) | 20.22 (0.09) |
| U3% | 33.84 (0.24) | 34.26 (0.17) | 33.86 (0.40) | 33.98 (0.09) |
| G3% | 24.82 (0.34) | 25.18 (0.43) | 25.06 (0.59) | 25.04 (0.15) |
| GC3% | 45.07 (0.61) | 45.10 (0.32) | 45.45 (0.25) | 45.25 (0.13) |
| GA3% | 45.91 (0.58) | 45.82 (0.23) | 45.75 (0.19) | 45.81 (0.06) |
| GU3% | 58.65 (1.08) | 59.43 (0.65) | 58.91(0.32) | 59.01 (0.20) |
| AU3% | 54.93 (0.92) | 54.90 (0.51) | 54.55(0.19) | 54.75 (0.13) |
| CU3% | 54.09 (0.86) | 54.18 (0.44) | 54.25 (0.08) | 54.19 (0.06) |
| GC1 | 0.50 (0.00) | 0.50 (0.00) | 0.50 (0.00) | 0.50 (0.00) |
| GC2 | 0.38 (0.00) | 0.38 (0.00) | 0.38 (0.00) | 0.38 (0.00) |
| GC3 | 0.44 (0.00) | 0.44 (0.00) | 0.44 (0.00) | 0.44 (0.00) |
| GC12 | 0.44 (0.00) | 0.44 (0.01) | 0.44 (0.01) | 0.44(0.01) |
| CBI | 0.05 (0.01) | 0.04 (0.00) | 0.04 (0.00) | 0.05 (0.00) |
| Fop | 0.45 (0.00) | 0.45 (0.00) | 0.45 (0.00) | 0.45 (0.00) |
| ENc | 54.63 (0.46) | 53.51 (0.39) | 53.74 (0.30) | 53.88 (0.22) |
| L_sym | 216.69 (0.08) | 216.76 (0.08) | 216.78 (0.07) | 216.75 (0.04) |
| L_aa | 223.00 (0.00) | 223.00 (0.00) | 223.00 (0.00) | 223.00 (0.00) |
| Gravy | −0.48 (0.00) | −0.49 (0.00) | −0.48 (0.00) | −0.48 (0.00) |
| Aromo | 0.08 (0.00) | 0.08 (0.00) | 0.08 (0.00) | 0.08 (0.00) |
| CAI | CBI | Fop | ENc | GC1 | GC2 | GC12 | GC3 | A | C | U | G | GU | AC | GU3 | AU3 | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| CAI | ||||||||||||||||
| CBI | 0.820 ** | |||||||||||||||
| Fop | 0.826 ** | 0.998 ** | ||||||||||||||
| ENc | 0.487 ** | 0.454 ** | 0.435 ** | |||||||||||||
| GC1 | 0.387 ** | 0.247 ** | 0.233 ** | 0.610 ** | ||||||||||||
| GC2 | −0.278 ** | −0.178 * | −0.172 | 0.047 | 0.056 | |||||||||||
| GC12 | 0.151 | 0.096 | 0.088 | 0.512 ** | 0.827 ** | 0.608 ** | ||||||||||
| GC3 | −0.563 ** | −0.412 ** | −0.402 ** | −0.373 ** | −0.329 ** | 0.093 | −0.210 * | |||||||||
| A | 0.733 ** | 0.704 ** | 0.696 ** | 0.500 ** | 0.191 * | −0.268 ** | 0.001 | −0.795 ** | ||||||||
| C | 0.425 ** | 0.665 ** | 0.653 ** | 0.614 ** | 0.382 ** | 0.090 | 0.355 ** | 0.005 | 0.388 ** | −0.845 ** | ||||||
| U | −0.393 ** | −0.506 ** | −0.504 ** | −0.595 ** | −0.426 ** | −0.175 | −0.438 ** | −0.087 | −0.358 ** | −0.537 ** | 0.313 ** | |||||
| G | −0.704 ** | −0.778 ** | −0.764 ** | −0.514 ** | −0.182 * | 0.273 ** | 0.009 | 0.741 ** | −0.931 ** | −0.788 ** | 0.686 ** | 0.906 ** | ||||
| GU | −0.715 ** | −0.822 ** | −0.810 ** | −0.659 ** | −0.329 ** | 0.131 | −0.188 * | 0.529 ** | −0.874 ** | 0.788 ** | −0.686 ** | −0.906 ** | −0.471 ** | |||
| AC | 0.715 ** | 0.822 ** | 0.810 ** | 0.659 ** | 0.329 ** | −0.131 | 0.188 * | −0.529 ** | 0.874 ** | −0.725 ** | 0.605 ** | 0.873 ** | 0.938 ** | −0.938 ** | ||
| GU3 | −0.749 ** | −0.836 ** | −0.821 ** | −0.646 ** | −0.282 ** | 0.118 | −0.158 | 0.521 ** | −0.831 ** | 0.000 | 0.097 | −0.748 ** | −0.530 ** | 0.530 ** | −1.000 ** | |
| AU3 | 0.568 ** | 0.431 ** | 0.420 ** | 0.370 ** | 0.325 ** | −0.090 | 0.208 * | −0.996 ** | 0.793 ** | 0.000 | −0.097 | 0.748 ** | 0.530 ** | −0.530 ** | 0.526 ** | −1.000 ** |
| AA | Codon | Ca-CTV | Cr-CTV | Cs-CTV | C. aurantifolia (Ca) | C. reticulata (Cr) | C. sinensis (Cs) |
|---|---|---|---|---|---|---|---|
| Phe | UUU | 1.11 | 1.14 | 1.01 | 1.20 | 1.08 | 1.24 |
| UUC | 0.89 | 0.86 | 0.99 | 0.80 | 0.92 | 0.76 | |
| Leu | UUA | 1.54 | 1.40 | 1.47 | 0.95 | 0.88 | 0.87 |
| UUG | 2.54 | 2.70 | 2.62 | 1.49 | 1.52 | 1.56 | |
| CUU | 1.13 | 1.12 | 1.13 | 1.40 | 1.28 | 1.48 | |
| CUC | 0.01 | 0.01 | 0.01 | 1.16 | 0.91 | 0.71 | |
| CUA | 0.24 | 0.26 | 0.31 | 0.48 | 0.43 | 0.57 | |
| CUG | 0.54 | 0.50 | 0.46 | 0.52 | 0.98 | 0.82 | |
| Ile | AUU | 0.91 | 0.81 | 0.80 | 1.39 | 1.48 | 1.51 |
| AUC | 0.50 | 0.59 | 0.55 | 0.93 | 0.90 | 0.71 | |
| AUA | 1.59 | 1.6 | 1.65 | 0.68 | 0.63 | 0.77 | |
| Val | GUU | 1.34 | 1.37 | 1.35 | 1.57 | 1.34 | 1.65 |
| GUC | 1.00 | 1.01 | 1.07 | 0.90 | 0.84 | 0.63 | |
| GUA | 0.51 | 0.48 | 0.37 | 0.55 | 0.54 | 0.66 | |
| GUG | 1.15 | 1.13 | 1.21 | 0.98 | 1.29 | 1.06 | |
| Pro | CCU | 1.49 | 1.44 | 1.38 | 1.84 | 1.54 | 1.50 |
| CCC | 0.03 | 0.04 | 0.06 | 0.66 | 0.69 | 0.58 | |
| CCA | 0.37 | 0.22 | 0.28 | 1.18 | 1.29 | 1.46 | |
| CCG | 2.11 | 2.30 | 2.28 | 0.32 | 0.49 | 0.45 | |
| Thr | ACU | 1.72 | 1.86 | 1.84 | 1.20 | 1.49 | 1.48 |
| ACC | 0.89 | 0.78 | 0.77 | 0.99 | 0.84 | 0.75 | |
| ACA | 0.66 | 0.64 | 0.65 | 1.34 | 1.21 | 1.36 | |
| ACG | 0.73 | 0.72 | 0.73 | 0.47 | 0.45 | 0.42 | |
| Ala | GCU | 1.90 | 1.86 | 1.85 | 1.88 | 1.41 | 1.65 |
| GCC | 0.76 | 0.82 | 0.87 | 1.08 | 1.01 | 0.69 | |
| GCA | 0.88 | 0.84 | 0.83 | 0.78 | 1.13 | 1.34 | |
| GCG | 0.47 | 0.48 | 0.44 | 0.25 | 0.46 | 0.32 | |
| Tyr | UAU | 1.29 | 1.28 | 1.29 | 0.94 | 1.03 | 1.23 |
| UAC | 0.71 | 0.72 | 0.71 | 1.06 | 0.97 | 0.77 | |
| Ser | UCU | 2.27 | 2.42 | 2.42 | 1.68 | 1.41 | 1.56 |
| UCC | 0.71 | 0.56 | 0.53 | 0.96 | 0.84 | 0.72 | |
| UCA | 0.75 | 0.84 | 0.81 | 1.20 | 1.33 | 1.45 | |
| UCG | 0.38 | 0.32 | 0.30 | 0.48 | 0.69 | 0.42 | |
| AGU | 1.45 | 1.57 | 1.53 | 0.80 | 0.87 | 1.07 | |
| AGC | 0.44 | 0.29 | 0.41 | 0.89 | 0.85 | 0.77 | |
| Arg | AGA | 1.18 | 1.09 | 1.12 | 2.38 | 1.84 | 1.98 |
| AGG | 0.69 | 0.75 | 0.76 | 0.83 | 1.29 | 1.49 | |
| CGU | 2.18 | 2.21 | 2.21 | 0.89 | 0.91 | 0.79 | |
| CGC | 0.55 | 0.49 | 0.53 | 0.49 | 0.70 | 0.48 | |
| CGA | 0.95 | 1.01 | 0.92 | 0.89 | 0.69 | 0.70 | |
| CGG | 0.45 | 0.44 | 0.47 | 0.52 | 0.56 | 0.55 | |
| Cys | UGU | 2.00 | 1.95 | 1.95 | 0.97 | 0.76 | 1.13 |
| UGC | 0.00 | 0.05 | 0.05 | 1.03 | 1.24 | 0.87 | |
| His | CAU | 0.59 | 0.62 | 0.61 | 1.15 | 1.05 | 1.32 |
| CAC | 1.41 | 1.38 | 1.39 | 0.85 | 0.95 | 0.68 | |
| Gln | CAA | 1.01 | 1.04 | 1.02 | 1.02 | 1.00 | 1.10 |
| CAG | 0.99 | 0.96 | 0.98 | 0.98 | 1.00 | 0.90 | |
| Asn | AAU | 0.52 | 0.59 | 0.59 | 0.83 | 1.11 | 1.30 |
| AAC | 1.48 | 1.41 | 1.41 | 1.17 | 0.89 | 0.70 | |
| Lys | AAA | 0.84 | 0.84 | 0.79 | 1.05 | 0.93 | 0.99 |
| AAG | 1.16 | 1.16 | 1.21 | 0.95 | 1.07 | 1.01 | |
| Asp | GAU | 1.16 | 1.17 | 1.16 | 1.29 | 1.33 | 1.42 |
| GAC | 0.84 | 0.83 | 0.84 | 0.71 | 0.67 | 0.58 | |
| Glu | GAA | 1.32 | 1.26 | 1.35 | 1.04 | 0.99 | 1.10 |
| GAG | 0.68 | 0.74 | 0.65 | 0.96 | 1.01 | 0.90 | |
| Gly | GGU | 1.62 | 1.54 | 1.48 | 1.26 | 1.09 | 1.20 |
| GGC | 0.65 | 0.73 | 0.75 | 1.11 | 0.98 | 0.73 | |
| GGA | 0.76 | 0.74 | 0.77 | 1.13 | 1.21 | 1.29 | |
| GGG | 0.97 | 1.00 | 0.99 | 0.50 | 0.71 | 0.77 | |
| Trp | UGG | 1.00 | 1.00 | 1.00 | 1.00 | 1.00 | 1.00 |
| Met | AUG | 1.00 | 1.00 | 1.00 | 1.00 | 1.00 | 1.00 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Biswas, K.K.; Palchoudhury, S.; Chakraborty, P.; Bhattacharyya, U.K.; Ghosh, D.K.; Debnath, P.; Ramadugu, C.; Keremane, M.L.; Khetarpal, R.K.; Lee, R.F. Codon Usage Bias Analysis of Citrus tristeza virus: Higher Codon Adaptation to Citrus reticulata Host. Viruses 2019, 11, 331. https://doi.org/10.3390/v11040331
Biswas KK, Palchoudhury S, Chakraborty P, Bhattacharyya UK, Ghosh DK, Debnath P, Ramadugu C, Keremane ML, Khetarpal RK, Lee RF. Codon Usage Bias Analysis of Citrus tristeza virus: Higher Codon Adaptation to Citrus reticulata Host. Viruses. 2019; 11(4):331. https://doi.org/10.3390/v11040331
Chicago/Turabian StyleBiswas, Kajal Kumar, Supratik Palchoudhury, Prosenjit Chakraborty, Utpal K. Bhattacharyya, Dilip K. Ghosh, Palash Debnath, Chandrika Ramadugu, Manjunath L. Keremane, Ravi K. Khetarpal, and Richard F. Lee. 2019. "Codon Usage Bias Analysis of Citrus tristeza virus: Higher Codon Adaptation to Citrus reticulata Host" Viruses 11, no. 4: 331. https://doi.org/10.3390/v11040331
APA StyleBiswas, K. K., Palchoudhury, S., Chakraborty, P., Bhattacharyya, U. K., Ghosh, D. K., Debnath, P., Ramadugu, C., Keremane, M. L., Khetarpal, R. K., & Lee, R. F. (2019). Codon Usage Bias Analysis of Citrus tristeza virus: Higher Codon Adaptation to Citrus reticulata Host. Viruses, 11(4), 331. https://doi.org/10.3390/v11040331





