Transcriptomic Analysis of the Campylobacter jejuni Response to T4-Like Phage NCTC 12673 Infection
Abstract
1. Introduction
2. Materials and Methods
2.1. Bacterial Growth Conditions
2.2. Phage Propagation, Titration and Concentration
2.3. Total RNA Extraction
2.4. RNA Sequencing
2.5. Efficiency of Plating (EOP) Assays
3. Results
3.1. NCTC 12673 Phage Infection Resulted in Clear Differences in Overall C. jejuni Gene Transcription
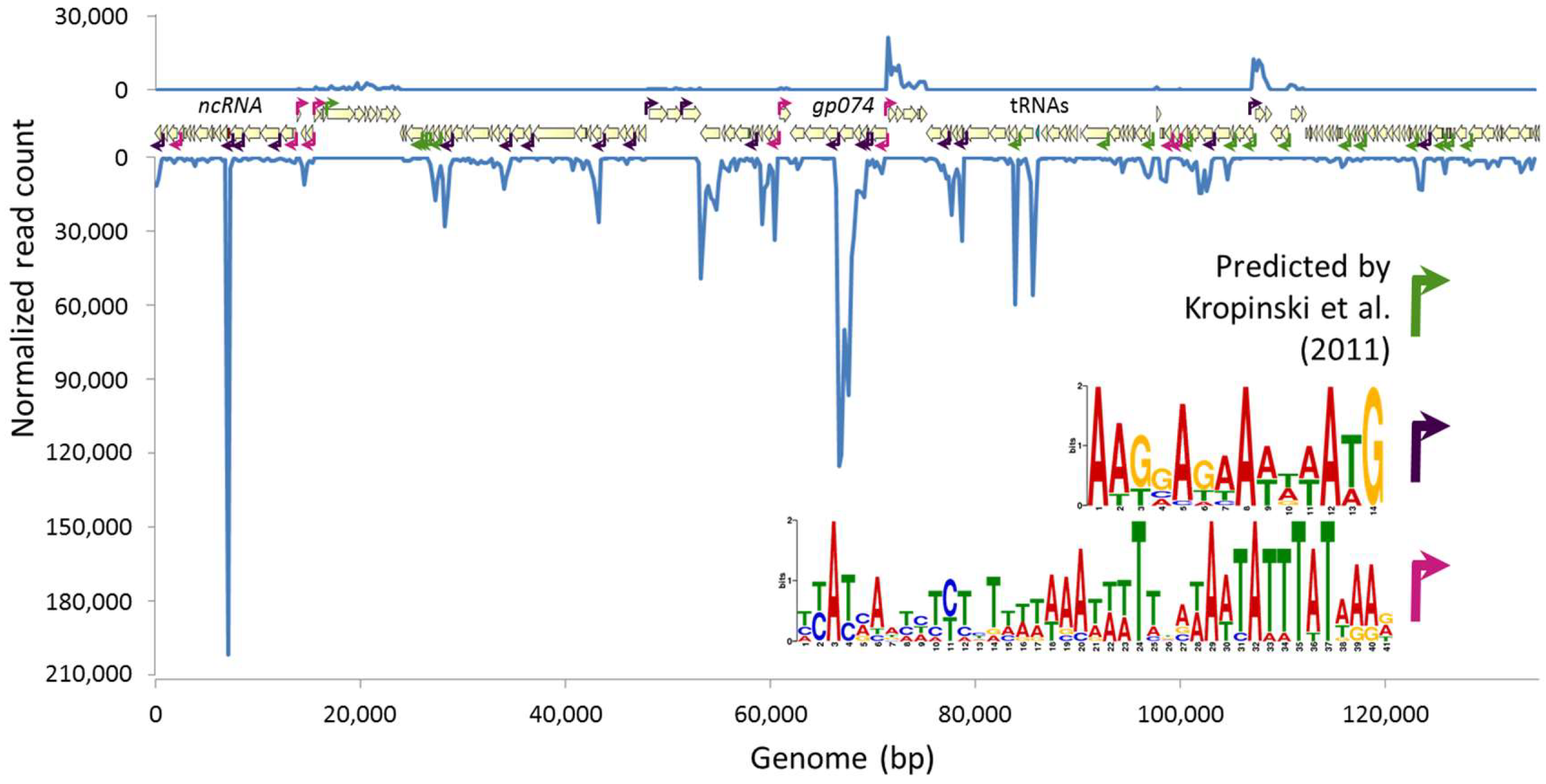
3.2. Phage Transcriptional Analysis Validates in Silico Predictions and Suggests Two New Promoter Motifs
3.3. Phage Infection Induces Widespread Induction of Host Genes and Downregulation of Energy Metabolism Pathways
3.4. Genes Involved in DNA, RNA, Amino Acid, and Protein Synthesis are Upregulated in Infected Cultures
3.5. Canonical Phage Defense Systems are not Differentially Expressed Upon Phage Infection
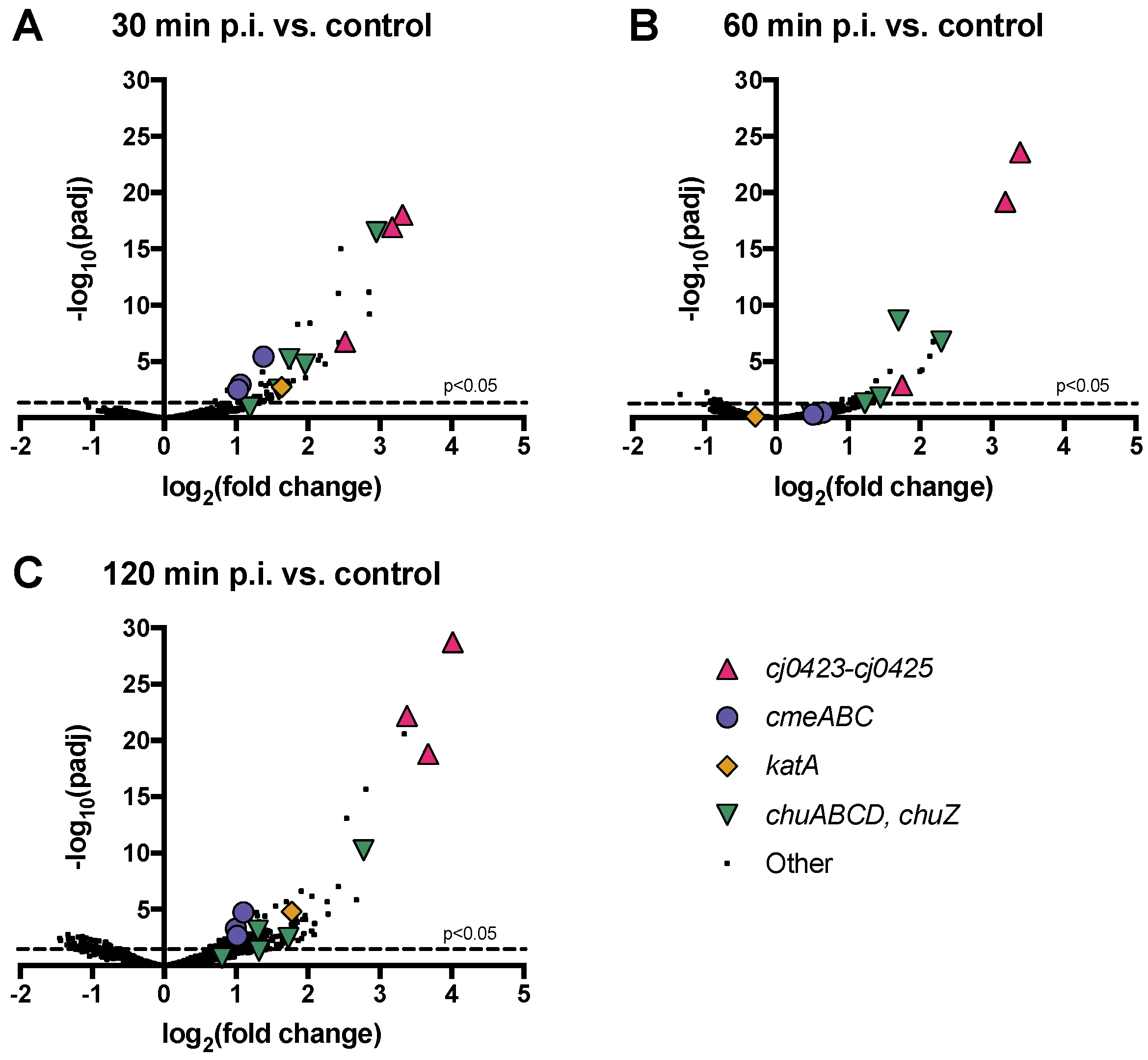
3.6. Host Operons cmeABC, chuABCD and cj0423–cj0425 are Among the Most Highly Upregulated upon Phage Infection
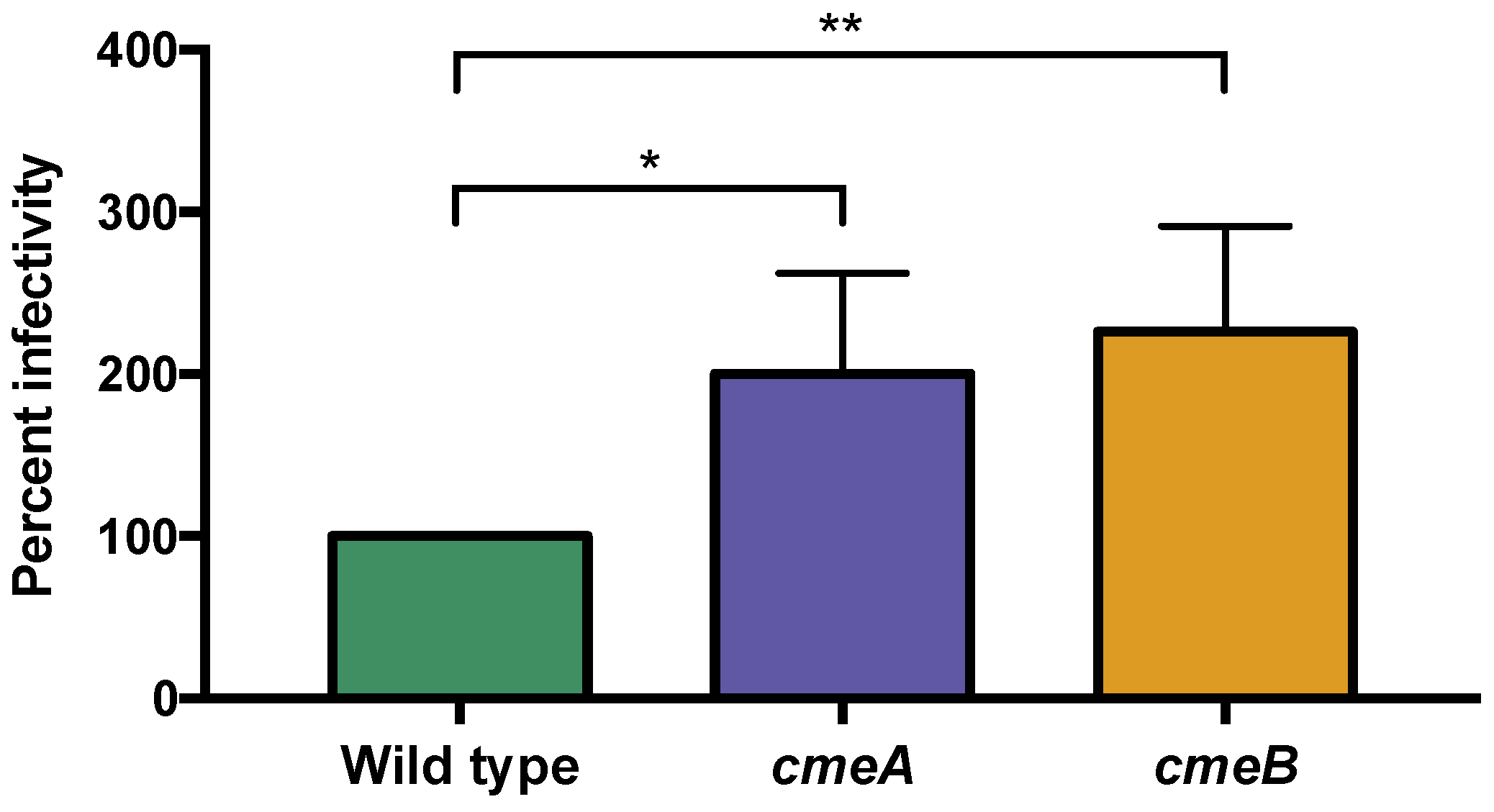
3.6.1. The C. jejuni CmeABC Multi-Drug Efflux Pump May Function in Phage Defense
3.6.2. Oxidative Stress Defense May Affect C. jejuni Interactions with Phage NCTC 12673
3.6.3. cj0423–cj0425 Is the Most Highly Upregulated Host Operon upon Phage Infection
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of Interest
References
- O’Neill, J. Antimicrobial Resistance: Tackling a Crisis for the Health and Wealth of Nations; The Review on Antimicrobial Resistance; UK Department of Health: London, UK, 2014. Available online: http://amr-review.org/sites/default/files/AMR%20Review%20Paper%20-%20Tackling%20a%20crisis%20for%20the%20health%20and%20wealth%20of%20nations_1.pdf (accessed on 23 April 2018).
- Huys, I.; Pirnay, J.P.; Lavigne, R.; Jennes, S.; de Vos, D.; Casteels, M.; Verbeken, G. Paving a regulatory pathway for phage therapy. Europe should muster the resources to financially, technically and legally support the introduction of phage therapy. EMBO Rep. 2013. [Google Scholar] [CrossRef]
- Roach, D.R.; Debarbieux, L. Phage therapy: Awakening a sleeping giant. Emerg. Top. Life Sci. 2017, 1, 93–103. [Google Scholar] [CrossRef]
- Umaraw, P.; Prajapati, A.; Verma, A.K.; Pathak, V.; Singh, V.P. Control of Campylobacter in poultry industry from farm to poultry processing unit: A review. Crit. Rev. Food Sci. Nutr. 2017, 57, 659–665. [Google Scholar] [CrossRef] [PubMed]
- Blasdel, B.; Ceyssens, P.-J.; Lavigne, R. Preparing cDNA Libraries from Lytic Phage-Infected Cells for Whole Transcriptome Analysis by RNA-Seq. In Methods Molecular Biology; Springer: Berlin, Germany, 2018; pp. 185–194. [Google Scholar]
- Leskinen, K.; Blasdel, B.G.; Lavigne, R.; Skurnik, M. RNA-sequencing reveals the progression of Phage-Host interactions between φR1-37 and Yersinia enterocolitica. Viruses 2016, 8, 111. [Google Scholar] [CrossRef] [PubMed]
- Ceyssens, P.-J.; Minakhin, L.; Van den Bossche, A.; Yakunina, M.; Klimuk, E.; Blasdel, B.; de Smet, J.; Noben, J.-P.; Bläsi, U.; Severinov, K.; et al. Development of giant bacteriophage ϕKZ is independent of the host transcription apparatus. J. Virol. 2014, 88, 10501–10510. [Google Scholar] [CrossRef] [PubMed]
- Morimoto, D.; Kimura, S.; Sako, Y.; Yoshida, T. Transcriptome Analysis of a Bloom-Forming Cyanobacterium Microcystis aeruginosa during Ma-LMM01 Phage Infection. Front. Microbiol. 2018, 9, 2. [Google Scholar] [CrossRef] [PubMed]
- Doron, S.; Fedida, A.; Hernández-Prieto, M.A.; Sabehi, G.; Karunker, I.; Stazic, D.; Feingersch, R.; Steglich, C.; Futschik, M.; Lindell, D.; et al. Transcriptome dynamics of a broad host-range cyanophage and its hosts. ISME J. 2016, 10, 1437–1455. [Google Scholar] [CrossRef] [PubMed]
- Mojardín, L.; Salas, M. Global transcriptional analysis of virus-host interactions between phage φ29 and Bacillus subtilis. J. Virol. 2016, 90, JVI.01245-16. [Google Scholar] [CrossRef] [PubMed]
- Brathwaite, K.J.; Siringan, P.; Connerton, P.L.; Connerton, I.F. Host adaption to the bacteriophage carrier state of Campylobacter jejuni. Res. Microbiol. 2015, 166, 504–515. [Google Scholar] [CrossRef] [PubMed]
- Howard-Varona, C.; Roux, S.; Dore, H.; Solonenko, N.E.; Holmfeldt, K.; Markillie, L.M.; Orr, G.; Sullivan, M.B. Regulation of infection efficiency in a globally abundant marine Bacteriodetes virus. ISME J. 2017, 11, 284–295. [Google Scholar] [CrossRef] [PubMed]
- Chevallereau, A.; Blasdel, B.G.; De Smet, J.; Monot, M.; Zimmermann, M.; Kogadeeva, M.; Sauer, U.; Jorth, P.; Whiteley, M.; Debarbieux, L.; et al. Next-Generation “-omics” Approaches Reveal a Massive Alteration of Host RNA Metabolism during Bacteriophage Infection of Pseudomonas aeruginosa. PLoS Genet. 2016. [Google Scholar] [CrossRef] [PubMed]
- Blasdel, B.G.; Ceyssens, P.-J.; Chevallereau, A.; Debarbieux, L.; Lavigne, R. Comparative transcriptomics reveals a conserved Bacterial Adaptive Phage Response (BAPR) to viral predation. bioRxiv 2018. [Google Scholar] [CrossRef]
- Blasdel, B.G.; Chevallereau, A.; Monot, M.; Lavigne, R.; Debarbieux, L. Comparative transcriptomics analyses reveal the conservation of an ancestral infectious strategy in two bacteriophage genera. ISME J. 2017. [Google Scholar] [CrossRef] [PubMed]
- Lin, X.; Ding, H.; Zeng, Q. Transcriptomic response during phage infection of a marine cyanobacterium under phosphorus-limited conditions. Environ. Microbiol. 2016, 18, 450–460. [Google Scholar] [CrossRef] [PubMed]
- Parkhill, J.; Wren, B.W.; Mungall, K.; Ketley, J.M.; Churcher, C.; Basham, D.; Chillingworth, T.; Davies, R.M.; Feltwell, T.; Holroyd, S.; et al. The genome sequence of the food-borne pathogen Campylobacter jejuni reveals hypervariable sequences. Nature 2000. [Google Scholar] [CrossRef] [PubMed]
- Gundogdu, O.; Bentley, S.D.; Holden, M.T.; Parkhill, J.; Dorrell, N.; Wren, B.W. Re-annotation and re-analysis of the Campylobacter jejuni NCTC11168 genome sequence. BMC Genom. 2007, 8. [Google Scholar] [CrossRef] [PubMed]
- Kaakoush, N.O.; Mitchell, H.M.; Man, S.M. Campylobacter. In Molecular Medical Microbiology; Academic Press: Cambridge, MA, USA, 2015; pp. 1187–1236. ISBN 978-0-12-397169-2. [Google Scholar]
- Wassenaar, T.M. Following an imaginary Campylobacter population from farm to fork and beyond: A bacterial perspective. Lett. Appl. Microbiol. 2011, 53, 253–263. [Google Scholar] [CrossRef] [PubMed]
- Zampara, A.; Sørensen, M.C.H.; Elsser-Gravesen, A.; Brøndsted, L. Significance of phage-host interactions for biocontrol of Campylobacter jejuni in food. Food Control 2017, 73, 1169–1175. [Google Scholar] [CrossRef]
- Fischer, S.; Kittler, S.; Klein, G.; Glünder, G. Impact of a Single Phage and a Phage Cocktail Application in Broilers on Reduction of Campylobacter jejuni and Development of Resistance. PLoS ONE 2013, 8. [Google Scholar] [CrossRef] [PubMed]
- Kittler, S.; Fischer, S.; Abdulmawjood, A.; Glünder, G.; Kleina, G. Effect of bacteriophage application on Campylobacter jejuni loads in commercial broiler flocks. Appl. Environ. Microbiol. 2013, 79, 7525–7533. [Google Scholar] [CrossRef] [PubMed]
- Grajewski, B.A.; Kusek, J.W.; Gelfand, H.M. Development of bacteriophage typing system for Campylobacter jejuni and Campylobacter coli. J. Clin. Microbiol. 1985, 22, 13–18. [Google Scholar] [PubMed]
- Kropinski, A.M.; Arutyunov, D.; Foss, M.; Cunningham, A.; Ding, W.; Singh, A.; Pavlov, A.R.; Henry, M.; Evoy, S.; Kelly, J.; et al. Genome and proteome of Campylobacter jejuni bacteriophage NCTC 12673. Appl. Environ. Microbiol. 2011, 77, 8265–8271. [Google Scholar] [CrossRef] [PubMed]
- Javed, M.A.; Ackermann, H.W.; Azeredo, J.; Carvalho, C.M.; Connerton, I.; Evoy, S.; Hammerl, J.A.; Hertwig, S.; Lavigne, R.; Singh, A.; et al. A suggested classification for two groups of Campylobacter myoviruses. Arch. Virol. 2014, 159, 181–190. [Google Scholar] [CrossRef] [PubMed]
- Sørensen, M.C.H.; Gencay, Y.E.; Birk, T.; Baldvinsson, S.B.; Jäckel, C.; Hammerl, J.A.; Vegge, C.S.; Neve, H.; Brøndsted, L. Primary isolation strain determines both phage type and receptors recognised by Campylobacter jejuni bacteriophages. PLoS ONE 2015, 10. [Google Scholar] [CrossRef] [PubMed]
- Coward, C.; Grant, A.J.; Swift, C.; Philp, J.; Towler, R.; Heydarian, M.; Frost, J.A.; Maskell, D.J. Phase-variable surface structures are required for infection of Campylobacter jejuni by bacteriophages. Appl. Environ. Microbiol. 2006, 72, 4638–4647. [Google Scholar] [CrossRef] [PubMed]
- Javed, M.A.; van Alphen, L.B.; Sacher, J.; Ding, W.; Kelly, J.; Nargang, C.; Smith, D.F.; Cummings, R.D.; Szymanski, C.M. A receptor-binding protein of Campylobacter jejuni bacteriophage NCTC 12673 recognizes flagellin glycosylated with acetamidino-modified pseudaminic acid. Mol. Microbiol. 2015, 95, 101–115. [Google Scholar] [CrossRef] [PubMed]
- Javed, M.A.; Sacher, J.C.; van Alphen, L.B.; Patry, R.T.; Szymanski, C.M. A flagellar glycan-specific protein encoded by Campylobacter phages inhibits host cell growth. Viruses 2015, 7, 6661–6674. [Google Scholar] [CrossRef] [PubMed]
- Sørensen, M.C.H.; van Alphen, L.B.; Harboe, A.; Li, J.; Christensen, B.B.; Szymanski, C.M.; Brøndsted, L. Bacteriophage F336 recognizes the capsular phosphoramidate modification of Campylobacter jejuni NCTC11168. J. Bacteriol. 2011, 193, 6742–6749. [Google Scholar] [CrossRef] [PubMed]
- Frost, J.A.; Kramer, J.M.; Gillanders, S.A. Phage typing of Campylobacter jejuni and Campylobacter coli and its use as an adjunct to serotyping. Epidemiol. Infect. 1999, 123, S095026889900254X. [Google Scholar] [CrossRef]
- Sørensen, M.C.H.; Gencay, Y.E.; Brøndsted, L. Methods for initial characterization of Campylobacter jejuni bacteriophages. Methods Mol. Biol. 2017, 1512, 91–105. [Google Scholar] [PubMed]
- Palyada, K.; Threadgill, D.; Stintzi, A. Iron acquisition and regulation in Campylobacter jejuni. J. Bacteriol. 2004, 186, 4714–4729. [Google Scholar] [CrossRef] [PubMed]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15. [Google Scholar] [CrossRef] [PubMed]
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 2013, 29, 15–21. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M.; Furumichi, M.; Tanabe, M.; Sato, Y.; Morishima, K. KEGG: New perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 2017, 45, D353–D361. [Google Scholar] [CrossRef] [PubMed]
- Luo, W.; Friedman, M.S.; Shedden, K.; Hankenson, K.D.; Woolf, P.J. GAGE: Generally applicable gene set enrichment for pathway analysis. BMC Bioinform. 2009, 10. [Google Scholar] [CrossRef] [PubMed]
- Bailey, T.L.; Elkan, C. Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc. Int. Conf. Intell. Syst. Mol. Biol. 1994, 2, 28–36. [Google Scholar] [PubMed]
- Grant, C.E.; Bailey, T.L.; Noble, W.S. FIMO: scanning for occurrences of a given motif. Bioinformatics 2011, 27, 1017–1018. [Google Scholar] [CrossRef] [PubMed]
- Palyada, K.; Sun, Y.Q.; Flint, A.; Butcher, J.; Naikare, H.; Stintzi, A. Characterization of the oxidative stress stimulon and PerR regulon of Campylobacter jejuni. BMC Genom. 2009, 10, 481. [Google Scholar] [CrossRef] [PubMed]
- Patry, R.T.; Stahl, M.; Perez-Munoz, M.E.; Nothaft, H.; Wenzel, C.Q.; Sacher, J.C.; Coros, C.; Walter, J.; Vallance, B.A.; Szymanski, C.M. Bacterial warfare: growth inhibition of ganglioside-mimicking gut bacteria by AB5 toxins. Nat. Microbiol. 2018, in press. [Google Scholar]
- Akiba, M.; Lin, J.; Barton, Y.-W.; Zhang, Q. Interaction of CmeABC and CmeDEF in conferring antimicrobial resistance and maintaining cell viability in Campylobacter jejuni. J. Antimicrob. Chemother. 2006, 57, 52–60. [Google Scholar] [CrossRef] [PubMed]
- Loc Carrillo, C.; Atterbury, R.J.; El-Shibiny, A.; Connerton, P.L.; Dillon, E.; Scott, A.; Connerton, I.F. Bacteriophage therapy to reduce Campylobacter jejuni colonization of broiler chickens. Appl. Environ. Microbiol. 2005, 71, 6554–6563. [Google Scholar] [CrossRef] [PubMed]
- Erez, Z.; Steinberger-Levy, I.; Shamir, M.; Doron, S.; Stokar-Avihail, A.; Peleg, Y.; Melamed, S.; Leavitt, A.; Savidor, A.; Albeck, S.; et al. Communication between viruses guides lysis-lysogeny decisions. Nature 2017, 541, 488–493. [Google Scholar] [CrossRef] [PubMed]
- Abedon, S.T. Commentary: Communication between Viruses Guides Lysis-Lysogeny Decisions. Front. Microbiol. 2017, 8, 983. [Google Scholar] [CrossRef] [PubMed]
- Miller, E.S.; Kutter, E.; Mosig, G.; Arisaka, F.; Kunisawa, T.; Rüger, W. Bacteriophage T4 genome. Microbiol. Mol. Biol. Rev. 2003, 67, 86–156. [Google Scholar] [CrossRef] [PubMed]
- De Smet, J.; Zimmermann, M.; Kogadeeva, M.; Ceyssens, P.J.; Vermaelen, W.; Blasdel, B.; Bin Jang, H.; Sauer, U.; Lavigne, R. High coverage metabolomics analysis reveals phage-specific alterations to Pseudomonas aeruginosa physiology during infection. ISME J. 2016. [Google Scholar] [CrossRef] [PubMed]
- Ankrah, N.Y.D.; May, A.L.; Middleton, J.L.; Jones, D.R.; Hadden, M.K.; Gooding, J.R.; LeCleir, G.R.; Wilhelm, S.W.; Campagna, S.R.; Buchan, A. Phage infection of an environmentally relevant marine bacterium alters host metabolism and lysate composition. ISME J. 2014, 8, 1089–1100. [Google Scholar] [CrossRef] [PubMed]
- Guccione, E.; del Rocio Leon-Kempis, M.; Pearson, B.M.; Hitchin, E.; Mulholland, F.; van Diemen, P.M.; Stevens, M.P.; Kelly, D.J. Amino acid-dependent growth of Campylobacter jejuni : key roles for aspartase (AspA) under microaerobic and oxygen-limited conditions and identification of AspB (Cj0762), essential for growth on glutamate. Mol. Microbiol. 2008, 69, 77–93. [Google Scholar] [CrossRef] [PubMed]
- Sacher, J.C. Campylobacter Phage Propagating Strain NCTC 12661 Propagates Phage NCTC 12673 More Efficiently Than Does Strain NCTC 11168; Department of Biological Sciences, University of Alberta: Edmonton, Alberta, 2017. [Google Scholar]
- Anjum, A.; Brathwaite, K.J.; Aidley, J.; Connerton, P.L.; Cummings, N.J.; Parkhill, J.; Connerton, I.; Bayliss, C.D. Phase variation of a Type IIG restriction-modification enzyme alters site-specific methylation patterns and gene expression in Campylobacter jejuni strain NCTC11168. Nucleic Acids Res. 2016, 44, 4581–4594. [Google Scholar] [CrossRef] [PubMed]
- Hooton, S.P.T.; Connerton, I.F. Campylobacter jejuni acquire new host-derived CRISPR spacers when in association with bacteriophages harboring a CRISPR-like Cas4 protein. Front. Microbiol. 2015, 6, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Gardner, S.P.; Olson, J.W. Barriers to Horizontal Gene Transfer in Campylobacter jejuni. Adv. Appl. Microbiol. 2012, 79, 19–42. [Google Scholar] [CrossRef] [PubMed]
- Louwen, R.; van Baarlen, P. Are bacteriophage defence and virulence two sides of the same coin in Campylobacter jejuni? Biochem. Soc. Trans. 2013, 41, 1475–1481. [Google Scholar] [CrossRef] [PubMed]
- McNally, D.J.; Lamoureux, M.P.; Karlyshev, A.V.; Fiori, L.M.; Li, J.; Thacker, G.; Coleman, R.A.; Khieu, N.H.; Wren, B.W.; Brisson, J.R.; et al. Commonality and biosynthesis of the O-methyl phosphoramidate capsule modification in Campylobacter jejuni. J. Biol. Chem. 2007, 282, 28566–28576. [Google Scholar] [CrossRef] [PubMed]
- Ridley, K.A.; Rock, J.D.; Li, Y.; Ketley, J.M. Heme utilization in Campylobacter jejuni. J. Bacteriol. 2006, 188, 7862–7875. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.; Michel, L.O.; Zhang, Q. CmeABC functions as a multidrug efflux system in Campylobacter jejuni. Antimicrob. Agents Chemother. 2002, 46, 2124–2131. [Google Scholar] [CrossRef] [PubMed]
- Pumbwe, L.; Piddock, L.J.V. Identification and molecular characterisation of CmeB, a Campylobacter jejuni multidrug efflux pump. FEMS Microbiol. Lett. 2002, 206, 185–189. [Google Scholar] [CrossRef] [PubMed]
- De Smet, J.; Hendrix, H.; Blasdel, B.G.; Danis-Wlodarczyk, K.; Lavigne, R. Pseudomonas predators: Understanding and exploiting phage-host interactions. Nat. Rev. Microbiol. 2017, 15, 517–530. [Google Scholar] [CrossRef] [PubMed]
- Butcher, J.; Stintzi, A. The Transcriptional Landscape of Campylobacter jejuni under Iron Replete and Iron Limited Growth Conditions. PLoS ONE 2013, 8, e79475. [Google Scholar] [CrossRef] [PubMed]
- Grant, K.A.; Park, S.F. Molecular characterization of katA from Campylobacter jejuni and generation of a catalase-deficient mutant of Campylobacter coli by interspecific allelic exchange. Microbiology 1995, 141, 1369–1376. [Google Scholar] [CrossRef] [PubMed]
- Day, W.A.; Sajecki, J.L.; Pitts, T.M.; Joens, L.A.; Joens, L.A. Role of catalase in Campylobacter jejuni intracellular survival. Infect. Immun. 2000, 68, 6337–6345. [Google Scholar] [CrossRef] [PubMed]
- Flint, A.; Sun, Y.Q.; Stintzi, A. Cj1386 is an ankyrin-containing protein involved in HEME trafficking to catalase in Campylobacter jejuni. J. Bacteriol. 2012, 194, 334–345. [Google Scholar] [CrossRef] [PubMed]
- Holberger, L.E.; Garza-Sánchez, F.; Lamoureux, J.; Low, D.A.; Hayes, C.S. A novel family of toxin/antitoxin proteins in Bacillus species. FEBS Lett. 2012, 586, 132–136. [Google Scholar] [CrossRef] [PubMed]
- Fernández, L.; Rodríguez, A.; García, P. Phage or foe: an insight into the impact of viral predation on microbial communities. ISME J. 2018. [Google Scholar] [CrossRef] [PubMed]
- Fernández, L.; González, S.; Campelo, A.B.; Martínez, B.; Rodríguez, A.; García, P. Low-level predation by lytic phage phiIPLA-RODI promotes biofilm formation and triggers the stringent response in Staphylococcus aureus. Sci. Rep. 2017, 7, 40965. [Google Scholar] [CrossRef] [PubMed]
- Moreau, P.; Diggle, S.P.; Friman, V.-P. Bacterial cell-to-cell signaling promotes the evolution of resistance to parasitic bacteriophages. Ecol. Evol. 2017, 7, 1936–1941. [Google Scholar] [CrossRef] [PubMed]
- Stahl, M.; Butcher, J.; Stintzi, A. Nutrient acquisition and metabolism by Campylobacter jejuni. Front. Cell. Infect. Microbiol. 2012, 2, 5. [Google Scholar] [CrossRef] [PubMed]
- Gots, J.S.; Hunt, G.R. Amino acid requirements for the maturation of bacteriophage in lysogenic Escherichia coli. J. Bacteriol. 1953, 66, 353–361. [Google Scholar] [PubMed]
- Bryan, D.; El-Shibiny, A.; Hobbs, Z.; Porter, J.; Kutter, E.M. Bacteriophage T4 infection of stationary phase E. coli: Life after log from a phage perspective. Front. Microbiol. 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- Dugar, G.; Herbig, A.; Förstner, K.U.; Heidrich, N.; Reinhardt, R.; Nieselt, K.; Sharma, C.M. High-Resolution Transcriptome Maps Reveal Strain-Specific Regulatory Features of Multiple Campylobacter jejuni Isolates. PLoS Genet. 2013, 9, e1003495. [Google Scholar] [CrossRef] [PubMed]
- Louwen, R.; Horst-Kreft, D.; de Boer, A.G.; Van Der Graaf, L.; De Knegt, G.; Hamersma, M.; Heikema, A.P.; Timms, A.R.; Jacobs, B.C.; Wagenaar, J.A.; et al. A novel link between Campylobacter jejuni bacteriophage defence, virulence and Guillain-Barre syndrome. Eur. J. Clin. Microbiol. Infect. Dis. 2013, 32, 207–226. [Google Scholar] [CrossRef] [PubMed]
- Dugar, G.; Leenay, R.T.; Eisenbart, S.K.; Bischler, T.; Aul, B.U.; Beisel, C.L.; Sharma, C.M. CRISPR RNA-Dependent Binding and Cleavage of Endogenous RNAs by the Campylobacter jejuni Cas9. Mol. Cell 2018, 69, 893–905. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Fang, L.; Tan, S.; Yu, M.; Li, X.; He, S.; Wei, Y.; Li, G.; Jiang, J.; Wu, M. Type I CRISPR-Cas targets endogenous genes and regulates virulence to evade mammalian host immunity. Cell Res. 2016, 26, 1273–1287. [Google Scholar] [CrossRef] [PubMed]
- Strutt, S.C.; Torrez, R.M.; Kaya, E.; Negrete, O.A.; Doudna, J.A. RNA-dependent RNA targeting by CRISPR-Cas9. eLife 2018, 7. [Google Scholar] [CrossRef] [PubMed]
- Sørensen, M.C.H.; van Alphen, L.B.; Fodor, C.; Crowley, S.M.; Christensen, B.B.; Szymanski, C.M.; Brøndsted, L. Phase Variable Expression of Capsular Polysaccharide Modifications Allows Campylobacter jejuni to Avoid Bacteriophage Infection in Chickens. Front. Cell. Infect. Microbiol. 2012, 2. [Google Scholar] [CrossRef] [PubMed]
- Aidley, J.; Holst Sørensen, M.C.; Bayliss, C.D.; Brøndsted, L. Phage exposure causes dynamic shifts in the expression states of specific phase-variable genes of Campylobacter jejuni. Microbiology 2017, 163, 911–919. [Google Scholar] [CrossRef] [PubMed]
- Gencay, Y.E.; Sørensen, M.C.H.; Wenzel, C.Q.; Szymanski, C.M.; Brøndsted, L. Phase Variable Expression of a Single Phage Receptor in Campylobacter jejuni NCTC12662 Influences Sensitivity Toward Several Diverse CPS-Dependent Phages. Front. Microbiol. 2018, 9, 82. [Google Scholar] [CrossRef] [PubMed]
- Hyman, P.; Abedon, S.T. Bacteriophage host range and bacterial resistance. Adv. Appl. Microbiol. 2010, 70, 217–248. [Google Scholar] [PubMed]
- Chan, B.K.; Sistrom, M.; Wertz, J.E.; Kortright, K.E.; Narayan, D.; Turner, P.E. Phage selection restores antibiotic sensitivity in MDR Pseudomonas aeruginosa. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Kim, Y.; Ma, Q.; Hong, S.H.; Pokusaeva, K.; Sturino, J.M.; Wood, T.K. Cryptic prophages help bacteria cope with adverse environments. Nat. Commun. 2010, 1, 147. [Google Scholar] [CrossRef] [PubMed]
- Schuch, R.; Fischetti, V.A. Detailed genomic analysis of the Wbeta and gamma phages infecting Bacillus anthracis: implications for evolution of environmental fitness and antibiotic resistance. J. Bacteriol. 2006. [Google Scholar] [CrossRef] [PubMed]
- Lu, M.J.; Henning, U. The immunity (IMM) gene of Escherichia coli bacteriophage T4. J. Virol. 1989, 63, 3472–3478. [Google Scholar] [PubMed]
- Labrie, S.J.; Samson, J.E.; Moineau, S. Bacteriophage resistance mechanisms. Nat. Rev. Microbiol. 2010, 8, 317–327. [Google Scholar] [CrossRef] [PubMed]
- Fineran, P.C.; Blower, T.R.; Foulds, I.J.; Humphreys, D.P.; Lilley, K.S.; Salmond, G.P.C. The phage abortive infection system, ToxIN, functions as a protein-RNA toxin-antitoxin pair. Proc. Natl. Acad. Sci. USA 2009, 106, 894–899. [Google Scholar] [CrossRef] [PubMed]
- Kaakoush, N.O.; Miller, W.G.; De Reuse, H.; Mendz, G.L. Oxygen requirement and tolerance of Campylobacter jejuni. Res. Microbiol. 2007, 158, 644–650. [Google Scholar] [CrossRef] [PubMed]
- Woodall, C.A.; Jones, M.A.; Barrow, P.A.; Hinds, J.; Marsden, G.L.; Kelly, D.J.; Dorrell, N.; Wren, B.W.; Maskell, D.J. Campylobacter jejuni gene expression in the chick cecum: Evidence for adaptation to a low-oxygen environment. Infect. Immun. 2005, 73, 5278–5285. [Google Scholar] [CrossRef] [PubMed]
- Gaynor, E.C.; Cawthraw, S.; Manning, G.; MacKichan, J.K.; Falkow, S.; Newell, D.G. The genome-sequenced variant of Campylobacter jejuni NCTC 11168 and the original clonal clinical isolate differ markedly in colonization, gene expression, and virulence-associated phenotypes. J. Bacteriol. 2004, 186, 503–517. [Google Scholar] [CrossRef] [PubMed]
- Ofir, G.; Melamed, S.; Sberro, H.; Mukamel, Z.; Silverman, S.; Yaakov, G.; Doron, S.; Sorek, R. DISARM is a widespread bacterial defence system with broad anti-phage activities. Nat. Microbiol. 2018, 3, 90–98. [Google Scholar] [CrossRef] [PubMed]

| Name | Source | Reference |
|---|---|---|
| C. jejuni NCTC 11168 MP21 | Chicken | [17,18,31] |
| Phage NCTC 12673 | Chicken | [24,25] |
| C. jejuni NCTC 11168 ∆katA | C. jejuni NCTC 11168 | [41] |
| C. jejuni NCTC 11168 ∆ahpC | C. jejuni NCTC 11168 | [41] |
| C. jejuni NCTC 11168 ∆sodB | C. jejuni NCTC 11168 | [41] |
| C. jejuni NCTC 11168 ∆cmeA | C. jejuni NCTC 11168 | Patry et al. submitted [42] |
| C. jejuni NCTC 11168 ∆cmeB | C. jejuni NCTC 11168 | [43] |
| Fold Change (P-Value) a | ||||
|---|---|---|---|---|
| Gene | Annotation | 30 min (p) | 60 min (p) | 120 min (p) |
| Multi-drug efflux pump CmeABC | ||||
| cmeA | Periplasmic fusion protein CmeA (multidrug efflux system CmeABC) | 2.6 (<0.01) | NS | 1.99 (<0.01) |
| cmeB | Inner membrane efflux transporter CmeB (multidrug efflux system CmeABC) | 2.09 (<0.01) | NS | 2.14 (<0.01) |
| cmeC | Outer membrane channel protein CmeC (multidrug efflux system CmeABC) | 2.04 (<0.01) | NS | 2.02 (<0.01) |
| Oxidative stress and iron metabolism | ||||
| katA | Catalase | 3.1 (<0.01) | NS | 3.42 (<0.01) |
| chuA | Haemin uptake system outer membrane receptor | 7.73 (<0.01) | 4.9 (<0.01) | 6.84 (<0.01) |
| chuB | Putative haemin uptake system permease protein | 3.02 (<0.01) | NS | 1.8 (0.02) |
| chuC | Putative haemin uptake system ATP-binding protein | NS | NS | 2.49 (0.05) |
| chuD | Putative haemin uptake system periplasmic haemin-binding protein | 3.9 (<0.01) | 2.73 (0.02) | 3.31 (<0.01) |
| chuZb | Iron-responsive cellular heme oxygenase | 3.34 (<0.01) | 3.25 (<0.01) | 2.47 (<0.01) |
| Uncharacterized operon cj0423–cj0425 | ||||
| cj0423 | Putative integral membrane protein | 5.73 (<0.01) | 3.36 (<0.01) | 16.14 (<0.01) |
| cj0424 | Putative acidic periplasmic protein | 9.95 (<0.01) | 10.51 (<0.01) | 12.75 (<0.01) |
| cj0425 | Putative periplasmic protein | 9.01 (<0.01) | 9.1 (<0.01) | 10.39 (<0.01) |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sacher, J.C.; Flint, A.; Butcher, J.; Blasdel, B.; Reynolds, H.M.; Lavigne, R.; Stintzi, A.; Szymanski, C.M. Transcriptomic Analysis of the Campylobacter jejuni Response to T4-Like Phage NCTC 12673 Infection. Viruses 2018, 10, 332. https://doi.org/10.3390/v10060332
Sacher JC, Flint A, Butcher J, Blasdel B, Reynolds HM, Lavigne R, Stintzi A, Szymanski CM. Transcriptomic Analysis of the Campylobacter jejuni Response to T4-Like Phage NCTC 12673 Infection. Viruses. 2018; 10(6):332. https://doi.org/10.3390/v10060332
Chicago/Turabian StyleSacher, Jessica C., Annika Flint, James Butcher, Bob Blasdel, Hayley M. Reynolds, Rob Lavigne, Alain Stintzi, and Christine M. Szymanski. 2018. "Transcriptomic Analysis of the Campylobacter jejuni Response to T4-Like Phage NCTC 12673 Infection" Viruses 10, no. 6: 332. https://doi.org/10.3390/v10060332
APA StyleSacher, J. C., Flint, A., Butcher, J., Blasdel, B., Reynolds, H. M., Lavigne, R., Stintzi, A., & Szymanski, C. M. (2018). Transcriptomic Analysis of the Campylobacter jejuni Response to T4-Like Phage NCTC 12673 Infection. Viruses, 10(6), 332. https://doi.org/10.3390/v10060332






