Oropouche Fever: A Review
Abstract
1. Introduction
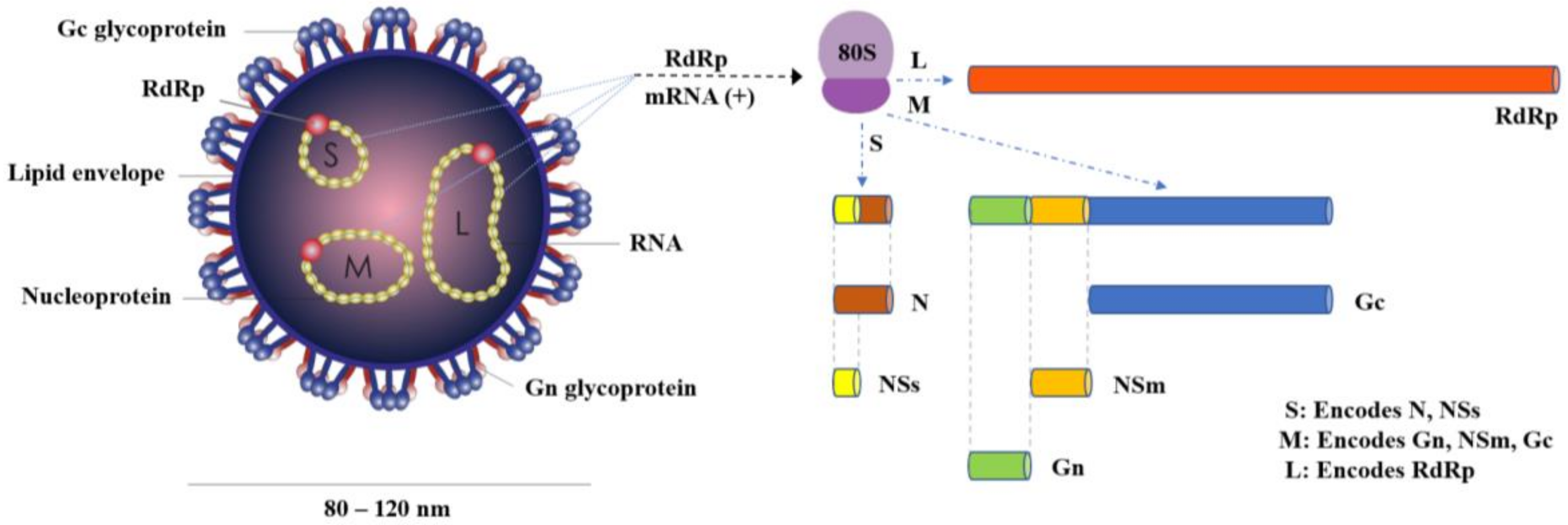
2. The Virus
3. Arthropod Vectors
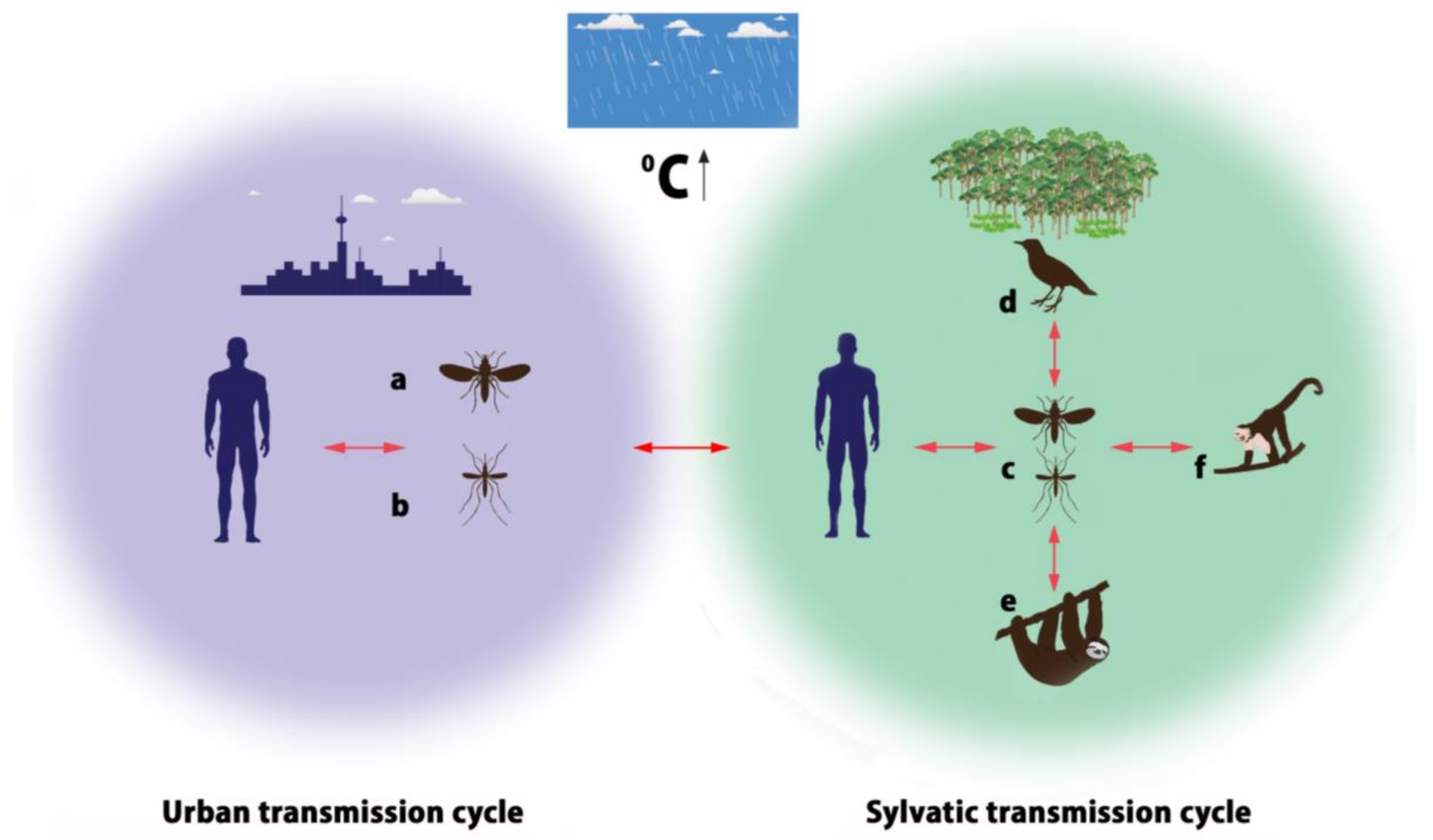
4. Transmission Cycles
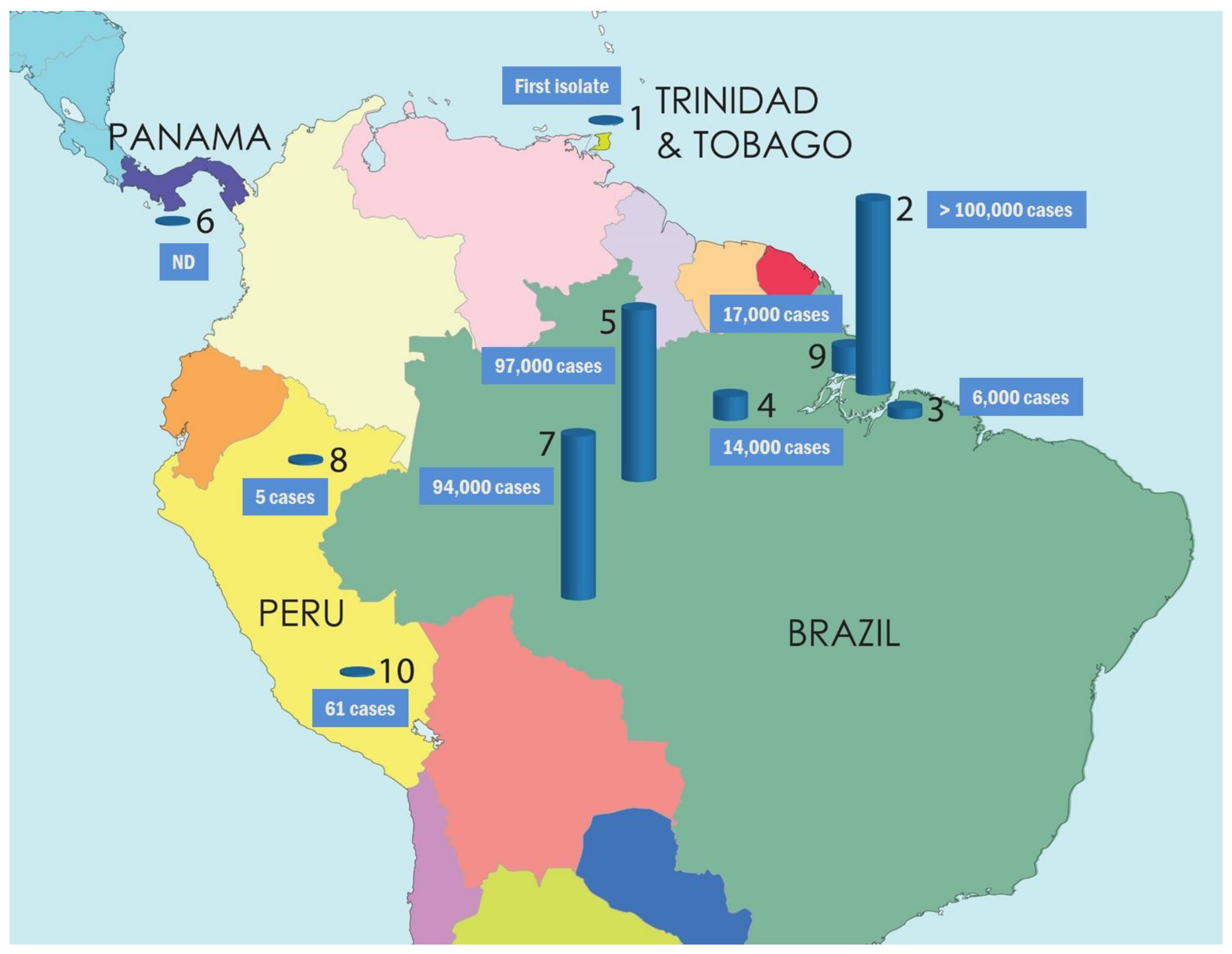
5. Disease Epidemiology
6. Pathogenesis
7. Clinical Manifestations
8. Diagnosis
9. Treatment and Prevention Options
10. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Anderson, C.R.; Spence, L.; Downs, W.G.; Aitken, T.H. Oropouche virus: A new human disease agent from Trinidad, West Indies. Am. J. Trop. Med. Hyg. 1961, 10, 574–578. [Google Scholar] [CrossRef] [PubMed]
- Adams, M.; Lefkowitz, E.; King, A.M.Q.; Harrach, B.; Harrison, R.; Knowles, N.J.; Kropinski, A.M.; Krupovic, M.; Kuhn, J.; Mushegian, A.; et al. Changes to taxonomy and the international code of virus classification and nomenclature ratified by the International Committee on Taxonomy of Viruses. Arch. Virol. 2017, 162, 2505–2538. [Google Scholar] [CrossRef] [PubMed]
- Mourão, M.P.; Bastos, M.S.; Gimaqu, J.B.; Mota, B.R.; Souza, G.; Grimmer, G.H.; Galusso, E.; Arruda, E.; Figueiredo, L.T. Oropouche fever outbreak, Manaus, Brazil, 2007–2008. Emerg. Infect. Dis. 2009, 15, 2063–2064. [Google Scholar] [CrossRef] [PubMed]
- Cardoso, B.F.; Serra, O.P.; Heinen, L.B.; Zuchi, N.; de Souza, V.C.; Naveca, F.G.; dos Santos, M.A.M.; Slhessarenko, R.D. Detection of Oropouche virus segment S in patients and in Culex quinquefasciatus in the state of Mato Grosso, Brazil. Mem. Inst. Oswaldo Cruz 2015, 110, 745–754. [Google Scholar] [CrossRef] [PubMed]
- Travassos da Rosa, J.F.; de Souza, W.M.; Pinheiro, F.P.; Figueiredo, M.L.; Cardoso, J.F.; Acrani, G.O.; Nunes, M.R.T. Oropouche Virus: Clinical, Epidemiological, and Molecular Aspects of a Neglected Orthobunyavirus. Am. J. Trop. Med. Hyg. 2017, 96, 1019–1030. [Google Scholar] [PubMed]
- Tilston-Lunel, N.L.; Hughes, J.; Acrani, G.O.; da Silva, D.E.; Azevedo, R.S.S.; Rodrigues, S.G.; Vasconcelos, P.F.C.; Nunes, M.R.T.; Elliott, R.M. Genetic analysis of members of the species Oropouche virus and identification of a novel M segment sequence. J. Gen. Virol. 2015, 96, 1636–1650. [Google Scholar] [CrossRef] [PubMed]
- Ladner, J.T.; Savji, N.; Lofts, L.; Travassos da Rosa, A.; Wiley, M.R.; Gestole, M.C.; Rosen, G.E.; Guzman, H.; Vasconcelos, P.F.C.; Nunes, M.R.T.; et al. Genomic and phylogenetic characterization of viruses included in the Manzanilla and Oropouche species complexes of the genus Orthobunyavirus, family Bunyaviridae. J. Gen. Virol. 2014, 95, 1055–1066. [Google Scholar] [CrossRef] [PubMed]
- Elliott, R.; Blakqori, G. Molecular Biology of Orthobunyaviruses. In Bunyaviridae: Molecular and Cellular Biology; Plyusnin, A., Elliott, R., Eds.; Caister Academic Press: Norfolk, UK, 2011; pp. 1–39. ISBN 9781904455905. [Google Scholar]
- Saeed, M.F.; Wang, H.; Nunes, M.; Vasconcelos, P.F.; Weaver, S.C.; Shope, R.E.; Watts, D.M.; Tesh, R.B.; Barrett, A.D.T. Nucleotide sequences and phylogeny of the nucleocapsid gene of Oropouche virus. J. Gen. Virol. 2000, 81, 743–748. [Google Scholar] [CrossRef] [PubMed]
- Vasconcelos, H.B.; Nunes, M.R.; Casseb, L.M.; Carvalho, V.L.; Pinto da Silva, E.V.; Silva, M.; Casseb, S.M.M.; Vasconcelos, P.F.C. Molecular epidemiology of Oropouche virus, Brazil. Emerg. Infect. Dis. 2011, 17, 800–806. [Google Scholar] [CrossRef] [PubMed]
- Nunes, M.R.; Martins, L.C.; Rodrigues, S.G.; Chiang, J.O.; Azevedo, R.S.; Travassos da Rosa, A.; Vasconcelos, P.F.C. Oropouche virus isolation, southeast Brazil. Emerg. Infect. Dis. 2005, 11, 1610–1613. [Google Scholar] [CrossRef] [PubMed]
- Bernardes-Terzian, A.C.; de-Moraes-Bronzoni, R.V.; Drumond, B.P.; Da Silva-Nunes, M.; Ferreira, M.U.; Speranca, M.A.; Nogueira, M.L. Sporadic Oropouche virus infection, Acre, Brazil. Emerg. Infect. Dis. 2009, 15, 348–350. [Google Scholar] [CrossRef] [PubMed]
- Azevedo, R.S.; Nunes, M.R.; Chiang, J.O.; Bensabath, G.; Vasconcelos, H.B.; Pinto, A.Y.; Martins, L.C.; Monteiro, H.A.; Rodrigues, S.G.; Vasconcelos, P.F.C. Reemergence of Oropouche fever, northern Brazil. Emerg. Infect. Dis. 2007, 13, 912–915. [Google Scholar] [CrossRef] [PubMed]
- Aguilar, P.V.; Barrett, A.D.; Saeed, M.F.; Watts, D.M.; Russell, K.; Guevara, C.; Ampuero, J.; Suarez, L.; Cespedes, M.; Montgomery, J.; et al. Iquitos virus: A novel reassortant Orthobunya virus associated with human illness in Peru. PLoS Negl. Trop. Dis. 2011, 5, e1315. [Google Scholar] [CrossRef] [PubMed]
- Briese, T.; Calisher, C.; Higgs, S. Viruses of the family Bunyaviridae: Are all available isolates reassortants? Virology 2013, 446, 207–216. [Google Scholar] [CrossRef] [PubMed]
- Mellor, P.S.; Boorman, J.; Baylis, M. Culicoides biting midges: Their role as arbovirus vectors. Annu. Rev. Entomol. 2000, 45, 307–340. [Google Scholar] [CrossRef] [PubMed]
- Augot, D.; Mathieu, B.; Hadj-Henni, L.; Barriel, V.; Zapata Mena, S.; Smolis, S.; Slama, D.; Randrianambinintsoa, F.J.; Trueba, G.; Kaltenbach, M.; et al. Molecular phylogeny of 42 species of Culicoides (Diptera, Ceratopogonidae) from three continents. Parasite 2017, 24, 23. [Google Scholar] [CrossRef] [PubMed]
- Purse, B.V.; Carpenter, S.; Venter, G.J.; Bellis, G.; Mullens, B.A. Bionomics of temperate and tropical Culicoides midges: Knowledge gaps and consequences for transmission of Culicoides-borne viruses. Annu. Rev. Entomol. 2015, 60, 373–392. [Google Scholar] [CrossRef] [PubMed]
- Aybar, C.A.; Juri, M.J.; de Grosso, M.S.; Spinell, G.R. New records of Culicoides species (Diptera: Ceratopogonidae) for Bolivia. J. Am. Mosq. Control Assoc. 2011, 27, 306–307. [Google Scholar] [CrossRef] [PubMed]
- Lassen, S.; Nielsen, S.A.; Kristensen, M.L. Identity and diversity of blood meal hosts of biting midges (Diptera: Ceratopogonidae: Culicoides Latreille) in Denmark. Parasites Vectors 2012, 5, 143. [Google Scholar] [CrossRef] [PubMed]
- Schnettler, E.; Ratinier, M.; Watson, M.; Shaw, A.E.; Mc Farlane, M.; Varela, M.; Elliott, R.; Palmarini, M.; Kohl, A. RNA interference targets arbovirus replication in Culicoides cells. J. Virol. 2013, 87, 2441–2454. [Google Scholar] [CrossRef] [PubMed]
- Augot, D.; Hadj-Henni, L.; Strutz, S.E.; Slama, D.; Millot, C.; Depaquit, J.; Millot, J.M. Association between host species choice and morphological characters of main sensory structures of Culicoides in the Palaeartic region. PeerJ 2017, 5, e3478. [Google Scholar] [CrossRef] [PubMed]
- Temmam, S.; Monteil-Bouchard, S.; Robert, C.; Baudoin, J.P.; Sambou, M.; Aubadie-Ladrix, M.; Labas, N.; Raoult, D.; Meddianikov, O.; Desnues, C. Characterization of Viral Communities of Biting Midges and Identification of Novel Thogoto virus Species and Rhabdovirus Genus. Viruses 2016, 8, 77. [Google Scholar] [CrossRef] [PubMed]
- Wirth, W.; Felippe-Bauer, M.L. The neotropical biting midges related to Culicoides paraensis (Diptera: Ceratopogonidae). Mem. Inst. Oswaldo Cruz 1989, 84, 551–565. [Google Scholar] [CrossRef]
- Aybar, C.A.; Juri, M.J.; De Grosso, M.S.; Spinelli, G.R. Species diversity and seasonal abundance of Culicoides biting midges in northwestern Argentina. Med. Vet. Entomol. 2010, 24, 95–98. [Google Scholar] [CrossRef] [PubMed]
- Aybar, C.A.; Juri, M.J.; Santana, M.; de Grosso, M.S.; Spinelli, G. The spatio-temporal distribution patterns of biting midges of the genus Culicoides in Salta province, Argentina. J. Insect Sci. 2012, 12, 145. [Google Scholar] [CrossRef] [PubMed]
- Carpenter, S.; Groschup, M.H.; Garros, C.; Felippe-Bauer, M.L.; Purse, B.V. Culicoides biting midges, arboviruses and public health in Europe. Antivir. Res. 2013, 100, 102–113. [Google Scholar] [CrossRef] [PubMed]
- Hoch, A.L.; Roberts, R.; Pinheiro, F.D. Breeding sites of Culicoides paraensis and options for control by environmental management. Bull. Pan Am. Health Organ. 1986, 20, 284–293. [Google Scholar] [PubMed]
- Tesh, R.B. The emerging epidemiology of Venezuelan hemorrhagic fever and Oropouche fever in tropical South America. Ann. N. Y. Acad. Sci. 1994, 740, 129–137. [Google Scholar] [CrossRef] [PubMed]
- Mercer, D.R.; Spinelli, G.R.; Watts, D.M.; Tesh, R.B. Biting rates and developmental substrates for biting midges (Diptera: Ceratopogonidae) in Iquitos, Peru. J. Med. Entomol. 2003, 40, 807–812. [Google Scholar] [CrossRef] [PubMed]
- Roberts, D.R.; Hoch, A.L.; Dixon, K.E.; Llewellyn, C.H. Oropouche virus. III. Entomological observations from three epidemics in Pará, Brazil, 1975. Am. J. Trop. Med. Hyg. 1981, 30, 165–171. [Google Scholar] [CrossRef] [PubMed]
- Pinheiro, F.; Travassos da Rosa, A.; Travassos da Rosa, J.; Ishak, R.; Freitas, R.B.; Gomes, L.M.C.; LeDuc, J.W.; Oliva, O.F.P. Oropouche Virus I. A Review of Clinical, Epidemiological, and Ecological Findings. Am. J. Trop. Med. Hyg. 1981, 30, 149–160. [Google Scholar] [CrossRef] [PubMed]
- De Souza Luna, L.K.; Rodrigues, A.H.; Santos, R.I.; Sesti-Costa, R.; Criado, M.F.; Martins, R.B.; Silva, M.L.; Delcaro, L.S.; Proenca-Modena, J.L.; Figueiredo, L.T.M.; et al. Oropouche virus is detected in peripheral blood leukocytes from patients. J. Med. Virol. 2017, 89, 1108–1111. [Google Scholar] [CrossRef] [PubMed]
- Cardoso, J.C.; Almeida, M.A.B.; dos Santos, E.; da Fonseca, D.F.; Sallum, M.A.M.; Noll, C.; Monteiro, A.O.; Cruz, A.C.R.; Carvalho, V.L.; Pinto, E.V.; et al. Yellow fever virus in Haemagogus leucocelaenus and Aedes serratus mosquitoes, Southern Brazil, 2008. Emerg. Infect. Dis. 2010, 16, 1918–1924. [Google Scholar] [CrossRef] [PubMed]
- Alencar, J.; Pacheco, J.B.; Correa, F.F.; dos Santos Silva, J.; Guimaraes, A.E. New report on the bionomics of Coquillettidia venezuelensis in temporary breeding sites (Diptera: Culicidae). Rev. Soc. Bras. Med. Trop. 2011, 44, 247–248. [Google Scholar] [CrossRef] [PubMed]
- Guo, X.X.; Li, C.X.; Deng, Y.Q.; Xing, D.; Liu, Q.M.; Wu, Q.; Sun, A.J.; Dong, Y.D.; Cao, W.C.; Qin, C.F.; et al. Culex pipiens quinquefasciatus: A potential vector to transmit Zika virus. Emerg. Microbe Inf. 2016, 5, e102. [Google Scholar] [CrossRef] [PubMed]
- Bhattacharya, S.; Basu, P. The southern house mosquito, Culex quinquefasciatus: Profile of a smart vector. J. Entomol. Zool Stud. 2016, 4, 73–81. [Google Scholar]
- Dixon, K.E.; Travassos da Rosa, A.P.; Travassos da Rosa, J.F.; Llewellyn, C.H. Oropouche virus. II. Epidemiological observations during an epidemic in Santarém, Pará, Brazil in 1975. Am. J. Trop. Med. Hyg. 1981, 30, 161–164. [Google Scholar] [CrossRef] [PubMed]
- Mourão, M.P.; Bastos, M.S.; Figueiredo, R.M.; Gimaque, J.B.; Alves, V.C.; Saraiva, M.; Figueiredo, M.; Ramasawmy, R.; Nogueira, M.L.; Figueiredo, L.T.M. Arboviral diseases in the Western Brazilian Amazon: A perspective and analysis from a tertiary health & research center in Manaus, State of Amazonas. Rev. Soc. Bras. Med. Trop. 2015, 48, 20–26. [Google Scholar] [PubMed]
- Pinheiro, F.P.; Rocha, A.G.; Freitas, R.B. Meningitis associated with Oropouche virus infection. Rev. Inst. Med. Trop. Sao Paulo 1982, 24, 246–251. [Google Scholar] [PubMed]
- Pinheiro, F.P.; Travassos da Rosa, A.P.; Travassos da Rosa, J.F.; Bensabath, G. An outbreak of Oropouche virus disease in the vicinity of Santarem, Para, Brazil. Tropenmed. Parasitol. 1976, 27, 213–223. [Google Scholar] [PubMed]
- Batista, P.M.; Andreotti, R.; Almeida, P.S.; Marques, A.C.; Rodrigues, S.G.; Chiang, J.O.; Vasconcelos, P.F.C. Detection of arboviruses of public health interest in free-living New World primates (Sapajus spp.; Alouatta caraya) captured in Mato Grosso do Sul, Brazil. Rev. Soc. Bras. Med. Trop. 2013, 46, 684–690. [Google Scholar] [CrossRef] [PubMed]
- Romero-Alvarez, D.; Escobar, L.E. Vegetation loss and the 2016 Oropouche fever outbreak in Peru. Mem. Inst. Oswaldo Cruz 2017, 112, 292–298. [Google Scholar] [CrossRef] [PubMed]
- Vasconcelos, P.F.; Calisher, C.H. Emergence of Human Arboviral Diseases in the Americas, 2000–2016. Vector Borne Zoonotic Dis. 2016, 16, 295–301. [Google Scholar] [CrossRef] [PubMed]
- Forshey, B.M.; Guevara, C.; Laguna-Torres, V.A.; Cespedes, M.; Vargas, J.; Gianella, A.; Vallejo, E.; Madrid, C.; Aguayo, N.; Gotuzzo, E.; et al. Arboviral etiologies of acute febrile illnesses in Western South America, 2000–2007. PLoS Negl. Trop. Dis. 2010, 4, e787. [Google Scholar] [CrossRef] [PubMed]
- Proenca-Modena, J.L.; Sesti-Costa, R.; Pinto, A.K.; Richner, J.M.; Lazear, H.M.; Lucas, T.; Hyde, J.; Diamond, M.S. Oropouche virus infection and pathogenesis are restricted by MAVS, IRF-3, IRF-7, and type I interferon signaling pathways in nonmyeloidcells. J. Virol. 2015, 89, 4720–4737. [Google Scholar] [CrossRef] [PubMed]
- Gibrail, M.M.; Fiaccadori, F.S.; Souza, M.; Almeida, T.N.; Chiang, J.O.; Martins, L.C.; Ferreira, M.S.; de Paula Cardoso, D.D. Detection of antibodies to Oropouche virus in non-human primates in Goiânia City, Goiás. Rev. Soc. Bras. Med. Trop. 2016, 49, 357–360. [Google Scholar] [CrossRef] [PubMed]
- Dutra, H.L.; Caragata, E.P.; Moreira, L.A. The re-emerging arboviral threat: Hidden, enemies: The emergence of obscure arboviral diseases, and the potential use of Wolbachia in their control. Bioessays 2017, 39, 1600175. [Google Scholar] [CrossRef] [PubMed]
- Figueiredo, L.T.M. Emerging arboviruses in Brazil. Rev. Soc. Braz. Med. Trop. 2007, 40, 224–229. [Google Scholar] [CrossRef]
- Tilston-Lunel, N.L.; Acrani, G.O.; Randall, R.E.; Elliott, R.M. Generation of Recombinant Oropouche Viruses Lacking the Nonstructural Protein NSm or NSs. J. Virol. 2016, 90, 2616–2627. [Google Scholar] [CrossRef] [PubMed]
- Acrani, G.O.; Tilston-Lunel, N.L.; Spiegel, M.; Weidmann, M.; Dilcher, M.; da Silva, D.E.A.; Nunes, M.R.T.; Elliott, R.M. Establishment of a minigenome system for Oropouche virus reveals the S genome segment to be significantly longer than reported previously. J. Gen. Virol. 2015, 96, 513–523. [Google Scholar] [CrossRef] [PubMed]
- Culquichicón, C.; Cardona-Ospina, J.A.; Patiño-Barbosa, A.M.; Rodriguez-Morales, A.J. Bibliometric analysis of Oropouche research: Impact on the surveillance of emerging arboviruses in Latin America. F1000 Res. 2017, 6, 194. [Google Scholar] [CrossRef] [PubMed]
- Pinheiro, F.P.; Travassos da Rosa, A.P.; Gomes, M.L.; LeDuc, J.W.; Hoch, A.L. Transmission of Oropouche virus from man to hamster by the midge Culicoides paraensis. Science 1982, 215, 1251–1253. [Google Scholar] [CrossRef] [PubMed]
- Vasconcelos, H.B.; Azevedo, R.S.; Casseb, S.M.; Nunes-Neto, J.P.; Chiang, J.O.; Cantuaria, P.C.; Segura, M.N.O.; Martins, L.C.; Monteiro, H.A.O.; Rodrigues, S.G.; et al. Oropouche fever epidemic in Northern Brazil: Epidemiology and molecular characterization of isolates. J. Clin. Virol. 2009, 44, 129–133. [Google Scholar] [CrossRef] [PubMed]
- Vasconcelos, P.F.; Travassos Da Rosa, J.F.; Guerreiro, S.C.; Dégallier, N.; Travassos da Rosa, E.S.; Travassos da Rosa, A.P. 1st register of an epidemic caused by Oropouche virus in the states of Maranhão and Goiás, Brazil. Rev. Inst. Med. Trop. Sao Paulo 1989, 31, 271–278. [Google Scholar] [CrossRef] [PubMed]
- Rosa, A.P.; Rodrigues, S.G.; Nunes, M.R.; Magalhães, M.T. Outbreak of oropouche virus fever in Serra Pelada, municipality of Curionópolis, Pará, 1994. Rev. Soc. Bras. Med. Trop 1996, 29, 537–541. [Google Scholar] [CrossRef] [PubMed]
- Encyclopaedia Britannica. Available online: https://www.britannica.com/place/Brazil (accessed on 27 March 2018).
- Chavez, R.; Colan, E.; Phillips, I. Fiebre de Oropuche en Iquitos: Reporte preliminar de 5 casos. Rev. Farmacol. Terap. 1992, 2, 12–14. [Google Scholar]
- Watts, D.M.; Phillips, I.; Callahan, J.D.; Griebenow, W.; Hyams, K.C.; Hayes, C.G. Oropouche virus transmission in the Amazon River basin of Peru. Am. J. Trop. Med. Hyg. 1997, 56, 148–152. [Google Scholar] [CrossRef] [PubMed]
- Baisley, K.J.; Watts, D.M.; Munstermann, L.E.; Wilson, M.L. Epidemiology of endemic Oropouche virus transmission in upper Amazonian Peru. Am. J. Trop. Med. Hyg. 1998, 59, 710–716. [Google Scholar] [CrossRef] [PubMed]
- Navarro, J.C.; Giambalvo, D.; Hernandez, R.; Auguste, A.J.; Tesh, R.B.; Weaver, S.C.; Montanez, H.; Liria, J.; Lima, A.; Travassos da Rosa, J.F.; et al. Isolation of Madre de Dios virus (Orthobunyavirus; Bunyaviridae), an Oropouche virus species Reassortment, from a monkey in Venezuela. Am. J. Trop. Med. Hyg. 2016, 95, 328–338. [Google Scholar] [CrossRef] [PubMed]
- Manock, S.R.; Jacobsen, K.H.; de Bravo, N.B.; Russell, K.L.; Negrete, M.; Olson, J.G.; Sanchez, J.L.; Blair, P.J.; Smalligan, R.D.; Quist, B.K.; et al. Etiology of acute undifferentiated febrile illness in the Amazon basin of Ecuador. Am. J. Trop. Med. Hyg. 2009, 81, 146–151. [Google Scholar] [PubMed]
- Romero-Alvarez, D.; Escobar, L.E. Oropouche fever, an emergent disease from the Americas. Microbes Infect. 2017, 20, 135–146. [Google Scholar] [CrossRef] [PubMed]
- Rodrigues, A.H.; Santos, R.I.; Arisi, G.M.; Bernardes, E.S.; Silva, M.L.; Rossi, M.A.; Lopes, M.B.S.; Arruda, E. Oropouche virus experimental infection in the golden hamster (Mesocrisetus auratus). Virus Res. 2011, 155, 35–41. [Google Scholar] [CrossRef] [PubMed]
- Santos, R.I.; Almeida, M.F.; Paula, F.E.; Rodrigues, A.H.; Saranzo, A.M.; Paula, A.E.; Silva, M.C.; Correa, V.M.A.; Acrani, G.O.; Neder, L.; et al. Experimental infection of suckling mice by subcutaneous inoculation with Oropouche virus. Virus Res. 2012, 170, 25–33. [Google Scholar] [CrossRef] [PubMed]
- Araujo, R.; Dias, L.B.; Araujo, M.T.; Pinheiro, F.; Oliva, O.F. Ultrastructure changes in the hamster liver after experimental inoculation with Oropouche arbovirus. Rev. Inst. Med. Trop. Sao Paulo 1978, 20, 45–54. [Google Scholar] [PubMed]
- Pulzova, L.; Bhide, M.; Andrej, K. Pathogen translocation across the blood-brain barrier. Pathog. Dis. 2009, 57, 203–213. [Google Scholar] [CrossRef] [PubMed]
- Proenca-Modena, J.L.; Hyde, J.L.; Sesti-Costa, R.; Lucas, T.; Pinto, A.K.; Richner, J.M.; Gorman, M.J.; Lazear, H.M.; Diamond, M.S. Interferon-Regulatory Factor 5-Dependent Signaling Restricts Orthobunyavirus Dissemination to the Central Nervous System. J. Virol. 2016, 90, 189–205. [Google Scholar] [CrossRef] [PubMed]
- Santos, R.I.; Bueno-Júnior, L.S.; Ruggiero, R.N.; Almeida, M.F.; Silva, M.L.; Paula, F.E.; Correa, V.M.A.; Arruda, E. Spread of Oropouche virus into the central nervous system in mouse. Viruses 2014, 6, 3827–3836. [Google Scholar] [CrossRef] [PubMed]
- Acrani, G.O.; Gomes, R.; Proença-Módena, J.L.; da Silva, A.; Carminati, P.O.; Silva, M.L.; Santos, R.I.M.; Arruda, E. Apoptosis induced by Oropouche virus infection in HeLa cells is dependent on virus protein expression. Virus Res. 2010, 149, 56–63. [Google Scholar] [CrossRef] [PubMed]
- Bastos, M.S.; Figueiredo, L.T.; Naveca, F.G.; Monte, R.L.; Lessa, N.; Figueiredo, R.M.P.; Gimaque, J.B.; Joao, G.P.; Ramasawmy, R.; Mourao, M.P.G. Short report: Identification of Oropouche Orthobunyavirus in the cerebrospinal fluid of three patients in the Amazonas Brazil. Am. J. Trop. Med. Hyg. 2012, 86, 732–735. [Google Scholar] [CrossRef] [PubMed]
- Bastos, M.S.; Lessa, N.; Naveca, F.G.; Monte, R.L.; Braga, W.S.; Figueiredo, L.T.M.; Ramasawmy, R.; Mourao, M.P. Detection of Herpesvirus, Enterovirus, and Arbovirus infection in patients with suspected central nervous system viral infection in the Western Brazilian Amazon. J. Med. Virol. 2014, 86, 1522–1527. [Google Scholar] [CrossRef] [PubMed]
- Moreli, M.L.; Aquino, V.H.; Cruz, A.C.; Figueiredo, L.T. Diagnosis of Oropouche virus infection by RT-nested-PCR. J. Med. Virol. 2002, 66, 139–142. [Google Scholar] [CrossRef] [PubMed]
- Saeed, M.F.; Nunes, M.; Vasconcelos, P.F.; Travassos Da Rosa, A.P.; Watts, D.M.; Russell, K.; Shope, R.E.; Tesh, R.B.; Barrett, A.D.T. Diagnosis of Oropouche virus infection using a recombinant nucleocapsid protein-based enzyme immunoassay. J. Clin. Microbiol. 2001, 39, 2445–2452. [Google Scholar] [CrossRef] [PubMed]
- Weidmann, M.; Rudaz, V.; Nunes, M.R.; Vasconcelos, P.F.; Hufert, F.T. Rapid detection of human pathogenic orthobunyaviruses. J. Clin. Microbiol. 2003, 41, 3299–3305. [Google Scholar] [CrossRef] [PubMed]
- Hang, J.; Forshey, B.M.; Kochel, T.J.; Li, T.; Solorzano, V.F.; Halsey, E.S.; Kuschner, R.A. Random amplification and pyrosequencing for identification of novel viral genome sequences. J. Biomol. Tech. 2012, 23, 4–10. [Google Scholar] [CrossRef] [PubMed]
- Naveca, F.G.; Nascimento, V.A.D.; Souza, V.C.; Nunes, B.T.D.; Rodrigues, D.S.G.; Vasconcelos, P.F.C. Multiplexed reverse transcription real-time polymerase chain reaction for simultaneous detection of Mayaro, Oropouche, and Oropouche-like viruses. Mem. Inst. Oswaldo Cruz 2017, 112, 510–513. [Google Scholar] [CrossRef] [PubMed]
- Livonesi, M.C.; De Sousa, R.L.; Badra, S.J.; Figueiredo, L.T. In vitro and in vivo studies of ribavirin action on Brazilian Orthobunyavirus. Am. J. Trop. Med. Hyg. 2006, 75, 1011–1016. [Google Scholar] [PubMed]
- Livonesi, M.C.; de Sousa, R.L.; Badra, S.J.; Figueiredo, L.T. In vitro and in vivo studies of the Interferon-alpha action on distinct Orthobunyavirus. Antivir. Res. 2007, 75, 121–128. [Google Scholar] [CrossRef] [PubMed]
- González, M.; Alarcón-Elbal, P.M.; Valle-Mora, J. Goldarazena, A. Comparison of different light sources for trapping Culicoides biting midges, mosquitoes and other dipterans. Vet. Parasitol. 2016, 226, 44–49. [Google Scholar] [CrossRef] [PubMed]
- Robin, M.; Page, P.; Archer, D.; Baylis, M. African horse sickness: The potential for an outbreak in disease-free regions and current disease control and elimination techniques. Equine Vet. J. 2016, 48, 659–669. [Google Scholar] [CrossRef] [PubMed]
- Benelli, G.; Buttazzoni, L.; Canale, A.; D’Andrea, A.; Serrone, P.; Delrio, G.; Foxi, C.; Mariani, S.; Savini, G.; Vadivalagan, C. Bluetongue outbreaks: Looking for effective control strategies against Culicoides vectors. Res. Vet. Sci. 2017, 115, 263–270. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sakkas, H.; Bozidis, P.; Franks, A.; Papadopoulou, C. Oropouche Fever: A Review. Viruses 2018, 10, 175. https://doi.org/10.3390/v10040175
Sakkas H, Bozidis P, Franks A, Papadopoulou C. Oropouche Fever: A Review. Viruses. 2018; 10(4):175. https://doi.org/10.3390/v10040175
Chicago/Turabian StyleSakkas, Hercules, Petros Bozidis, Ashley Franks, and Chrissanthy Papadopoulou. 2018. "Oropouche Fever: A Review" Viruses 10, no. 4: 175. https://doi.org/10.3390/v10040175
APA StyleSakkas, H., Bozidis, P., Franks, A., & Papadopoulou, C. (2018). Oropouche Fever: A Review. Viruses, 10(4), 175. https://doi.org/10.3390/v10040175





