Detection and Characterization of Homologues of Human Hepatitis Viruses and Pegiviruses in Rodents and Bats in Vietnam
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.2. Nucleic Acid Extraction
2.3. Screening of Hepatitis Viruses and Pegivirus
2.4. Complete Genome Sequencing
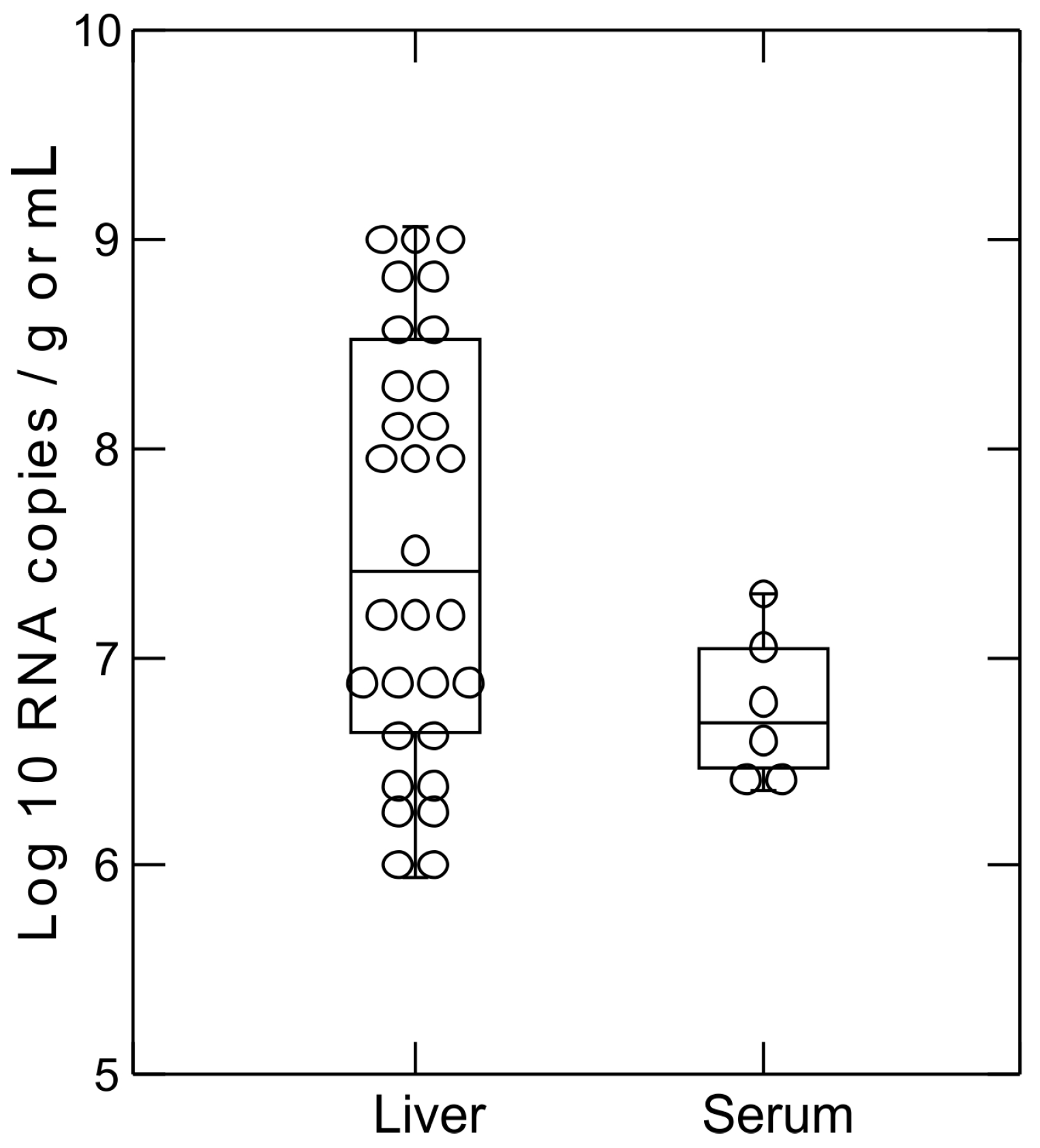
2.5. Hepacivirus RNA Titer Measurement
2.6. Sequence Analysis
3. Results
3.1. Detection of Hepatitis Viruses in Bat and Rodent Liver Samples
3.2. Screening of Rodent Serum Samples for Hepacivirus and Pegivirus
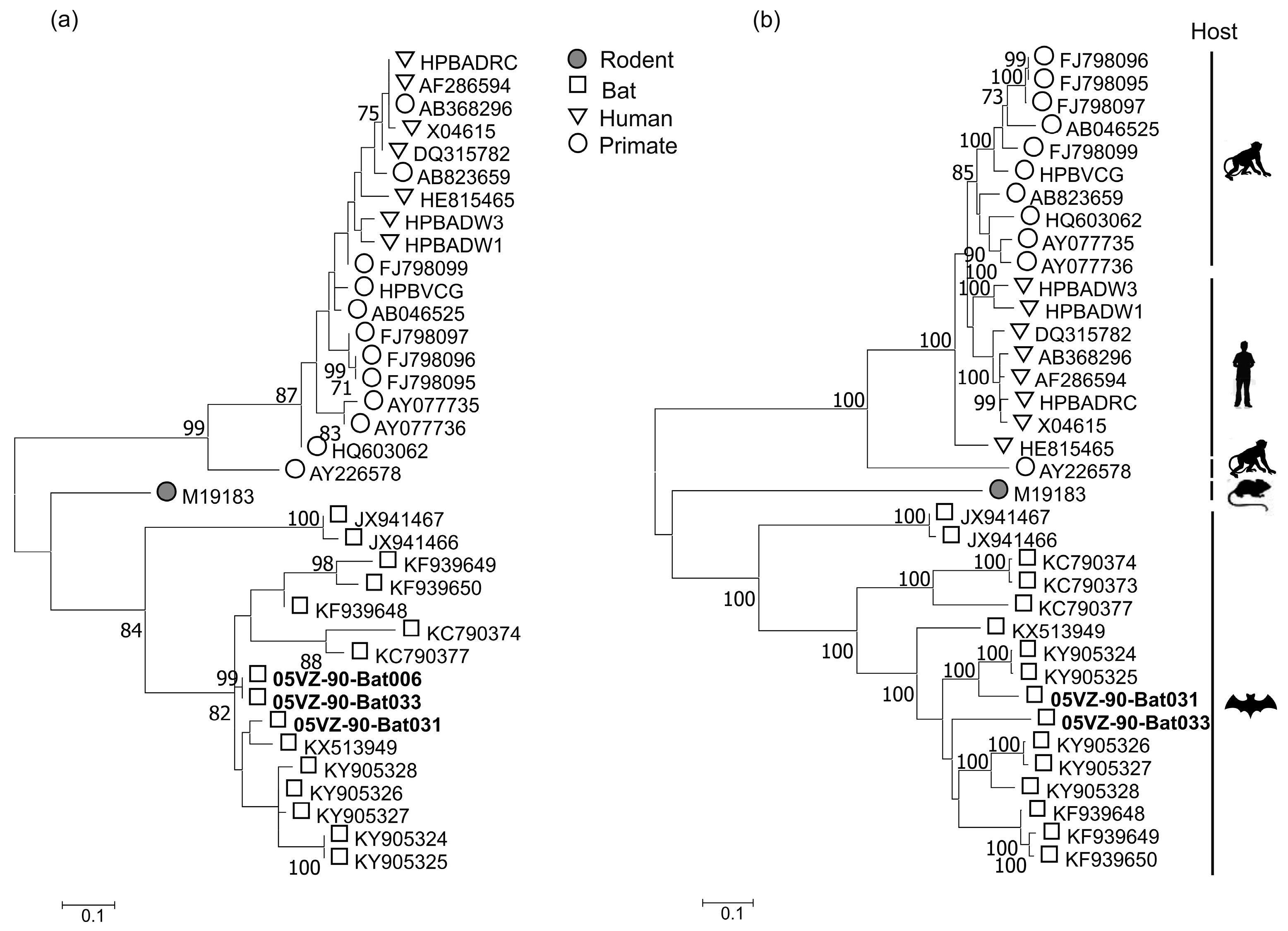
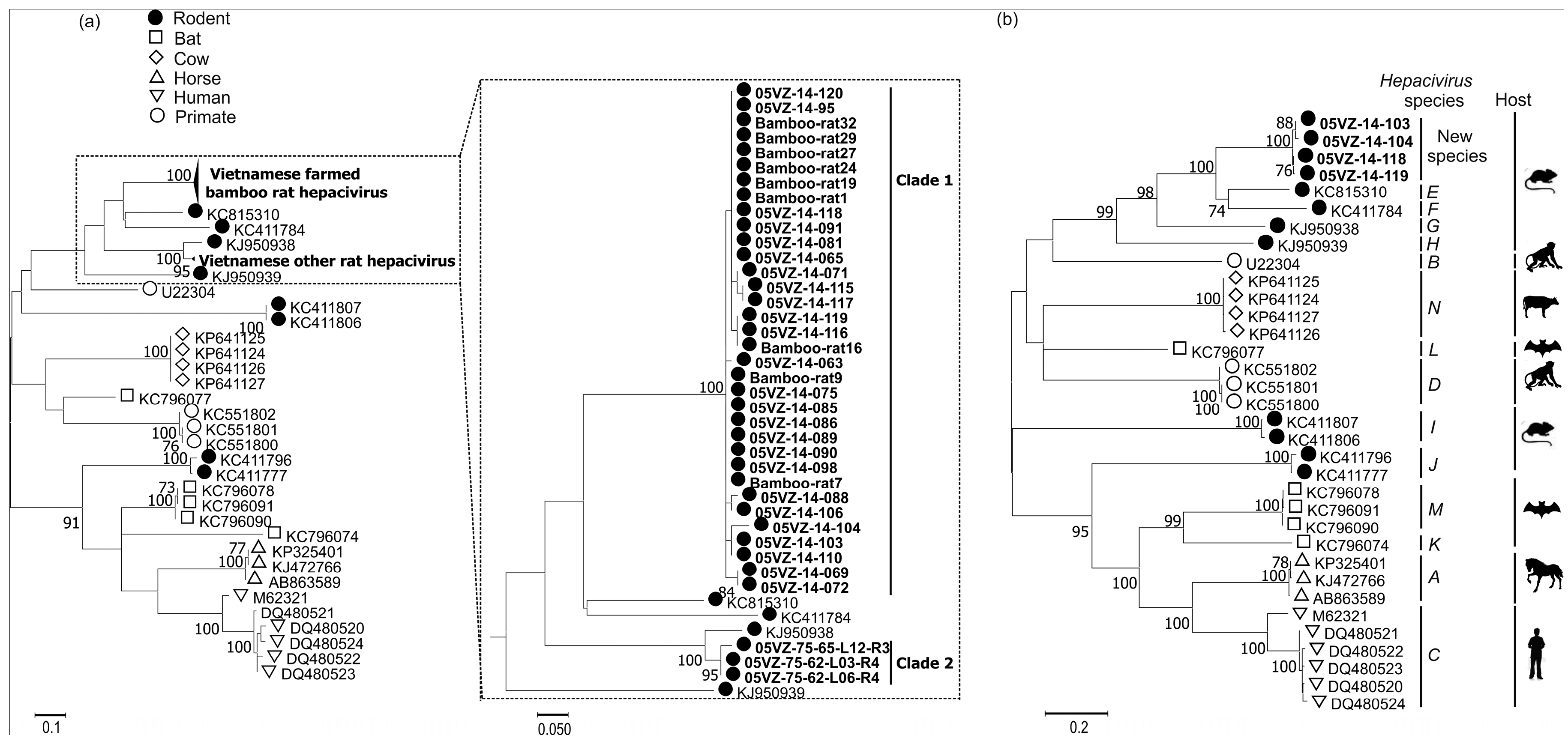
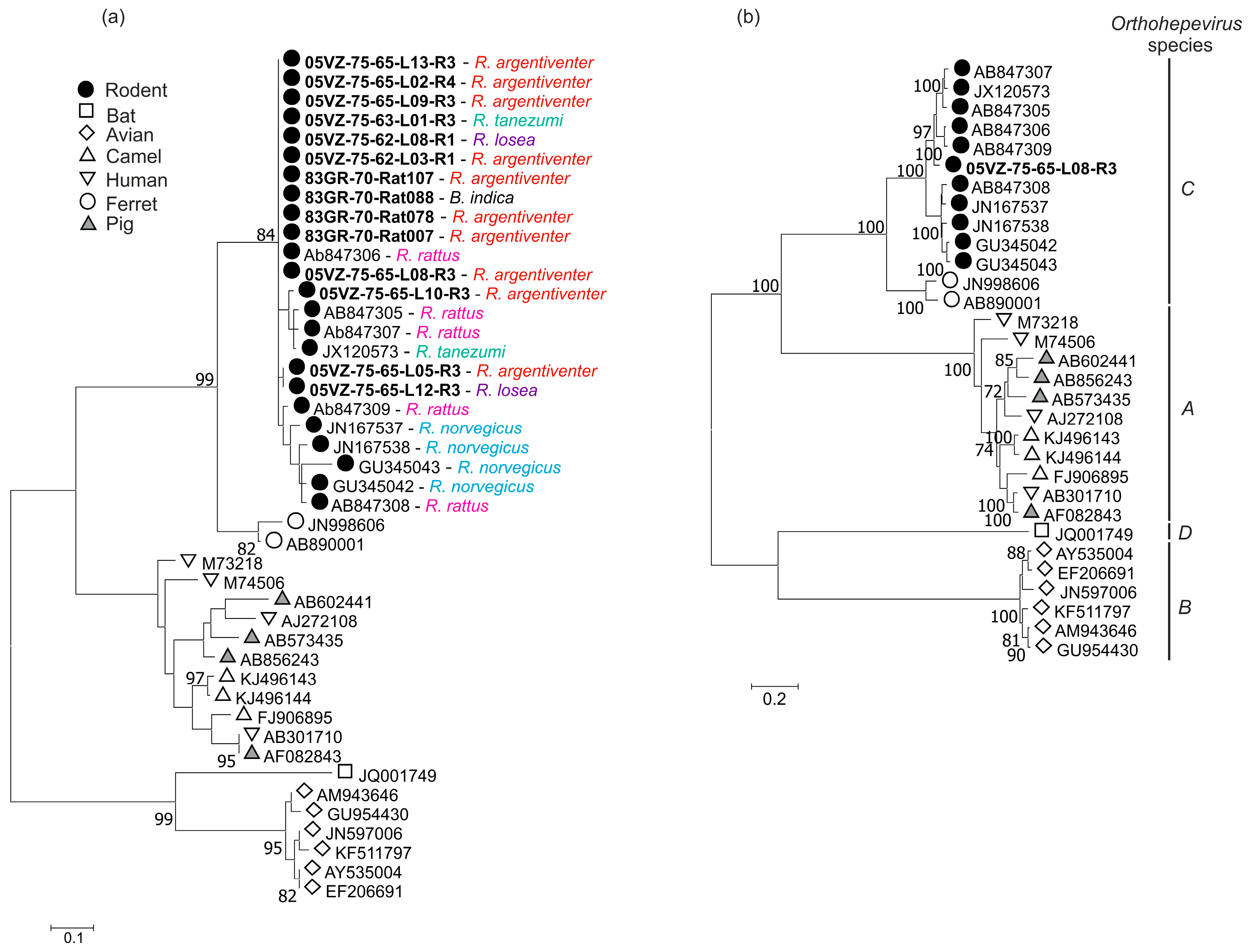
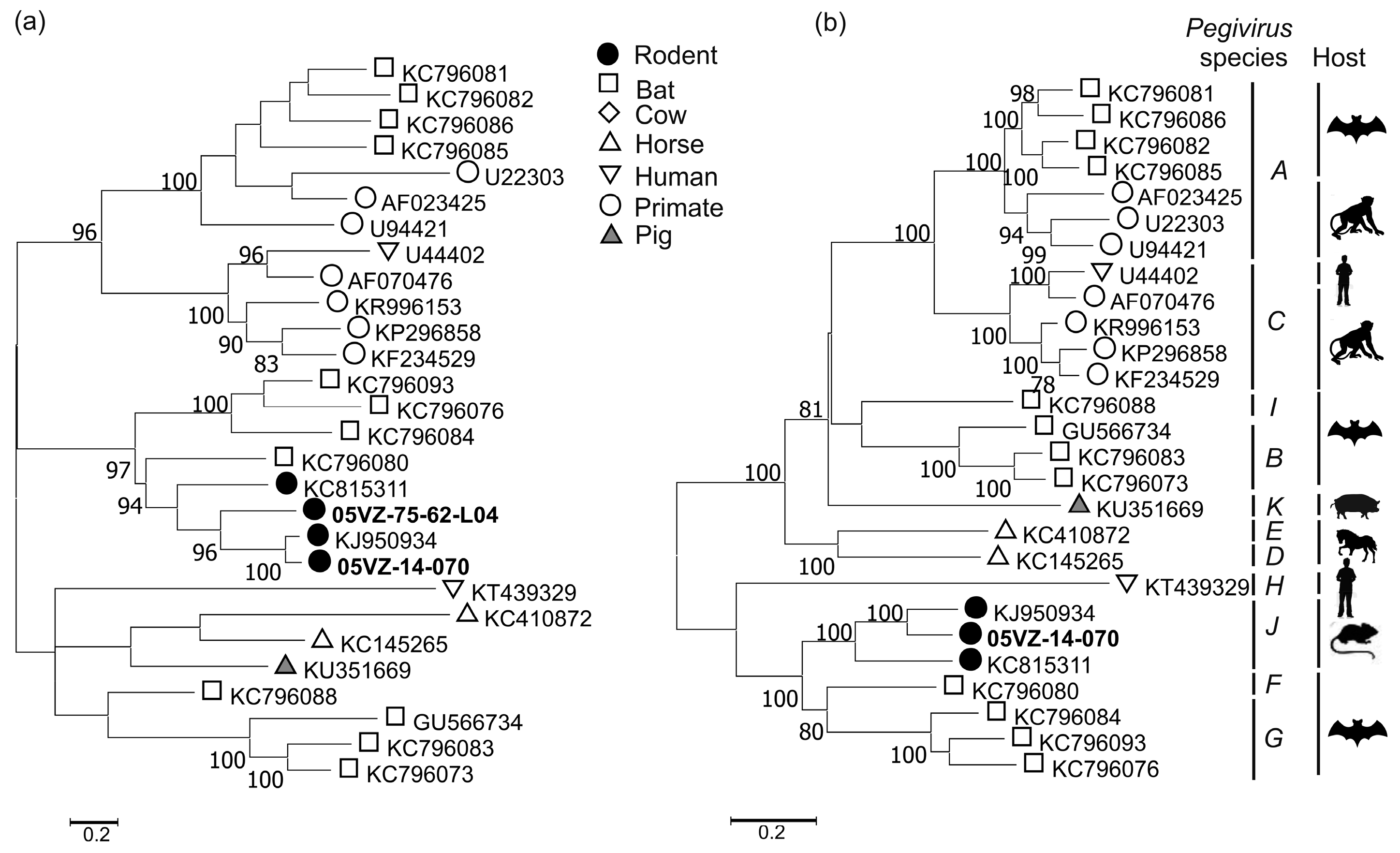
3.3. Sequence and Phylogenetic Analysis
4. Discussion
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Appendix A
References
- Stanaway, J.D.; Flaxman, A.D.; Naghavi, M.; Fitzmaurice, C.; Vos, T.; Abubakar, I.; Abu-Raddad, L.J.; Assadi, R.; Bhala, N.; Cowie, B.; et al. The global burden of viral hepatitis from 1990 to 2013: Findings from the Global Burden of Disease Study 2013. Lancet 2016, 388, 1081–1088. [Google Scholar] [CrossRef]
- Rein, D.B.; Stevens, G.A.; Theaker, J.; Wittenborn, J.S.; Wiersma, S.T. The global burden of hepatitis E virus genotypes 1 and 2 in 2005. Hepatology 2012, 55, 988–997. [Google Scholar] [CrossRef] [PubMed]
- Liang, T.J. Hepatitis B: The Virus and Disease. Hepatology 2009, 49, S13–S21. [Google Scholar] [CrossRef] [PubMed]
- Choo, Q.L.; Richman, K.H.; Han, J.H.; Berger, K.; Lee, C.; Dong, C.; Gallegos, C.; Coit, D.; Medina-Selby, R.; Barr, P.J. Genetic organization and diversity of the hepatitis C virus. Proc. Natl. Acad. Sci. USA 1991, 88, 2451–2455. [Google Scholar] [CrossRef] [PubMed]
- Mushahwar, I.K. Hepatitis E virus: Molecular virology, clinical features, diagnosis, transmission, epidemiology, and prevention. J. Med. Virol. 2008, 80, 646–658. [Google Scholar] [CrossRef] [PubMed]
- Drexler, J.F.; Geipel, A.; König, A.; Corman, V.M.; van Riel, D.; Leijten, L.M.; Bremer, C.M.; Rasche, A.; Cottontail, V.M.; Maganga, G.D.; et al. Bats carry pathogenic hepadnaviruses antigenically related to hepatitis B virus and capable of infecting human hepatocytes. Proc. Natl. Acad. Sci. USA 2013, 110, 16151–16156. [Google Scholar] [CrossRef] [PubMed]
- Wang, B.; Yang, X.-L.; Li, W.; Zhu, Y.; Ge, X.-Y.; Zhang, L.-B.; Zhang, Y.-Z.; Bock, C.-T.; Shi, Z.-L. Detection and genome characterization of four novel bat hepadnaviruses and a hepevirus in China. Virol. J. 2017, 14, 40. [Google Scholar] [CrossRef] [PubMed]
- He, B.; Fan, Q.; Yang, F.; Hu, T.; Qiu, W.; Feng, Y.; Li, Z.; Li, Y.; Zhang, F.; Guo, H.; et al. Hepatitis Virus in Long-Fingered Bats, Myanmar. Emerg. Infect. Dis. 2013, 19, 638–640. [Google Scholar] [CrossRef] [PubMed]
- Robert, H.P.; Ronald, E.E.; Michael, P.R.; Yamina, K.-L.; Hanh, T.N.; Sugantha, G.; Marisa St, C.; Suzanne, U.E. Hepatitis E Virus in Rats, Los Angeles, California, USA. Emerg. Infect. Dis. 2011, 17, 2216–2222. [Google Scholar]
- Lack, J.B.; Volk, K.; Van Den Bussche, R.A. Hepatitis E Virus Genotype 3 in Wild Rats, United States. Emerg. Infect. Dis. 2012, 18, 1268–1273. [Google Scholar] [CrossRef] [PubMed]
- Drexler, J.F.; Seelen, A.; Corman, V.M.; Fumie Tateno, A.; Cottontail, V.; Melim Zerbinati, R.; Gloza-Rausch, F.; Klose, S.M.; Adu-Sarkodie, Y.; Oppong, S.K.; et al. Bats Worldwide Carry Hepatitis E Virus-Related Viruses That Form a Putative Novel Genus within the Family Hepeviridae. J. Virol. 2012, 86, 9134–9147. [Google Scholar] [CrossRef] [PubMed]
- Debing, Y.; Emerson, S.U.; Purcell, R.H.; Neyts, J.; Dallmeier, K. Complete Genome Sequence of a Rat Hepatitis E Virus Strain Isolated in the United States. Genome Announc. 2014, 2. [Google Scholar] [CrossRef] [PubMed]
- Kanai, Y.; Miyasaka, S.; Uyama, S.; Kawami, S.; Kato-Mori, Y.; Tsujikawa, M.; Yunoki, M.; Nishiyama, S.; Ikuta, K.; Hagiwara, K. Hepatitis E virus in Norway rats (Rattus norvegicus) captured around a pig farm. BMC Res. Notes 2012, 5, 4. [Google Scholar] [CrossRef] [PubMed]
- Johne, R.; Heckel, G.; Plenge-Bonig, A.; Kindler, E.; Maresch, C.; Reetz, J.; Schielke, A.; Ulrich, R.G. Novel hepatitis e virus genotype in norway rats, Germany. Emerg. Infect. Dis. 2010, 16, 1452–1455. [Google Scholar] [CrossRef] [PubMed]
- Johne, R.; Plenge-Bonig, A.; Hess, M.; Ulrich, R.G.; Reetz, J.; Schielke, A. Detection of a novel hepatitis E-like virus in faeces of wild rats using a nested broad-spectrum RT-PCR. J. Gen. Virol. 2009, 91, 750–758. [Google Scholar] [CrossRef] [PubMed]
- Li, T.-C.; Ami, Y.; Suzaki, Y.; Yasuda, S.P.; Yoshimatsu, K.; Arikawa, J.; Takeda, N.; Takaji, W. Characterization of Full Genome of Rat Hepatitis E Virus Strain from Vietnam. Emerg. Infect. Dis. 2013, 19, 115–118. [Google Scholar] [CrossRef] [PubMed]
- Kapoor, A.; Simmonds, P.; Scheel, T.K.H.; Hjelle, B.; Cullen, J.M.; Burbelo, P.D.; Chauhan, L.V.; Duraisamy, R.; Sanchez Leon, M.; Jain, K.; et al. Identification of Rodent Homologs of Hepatitis C Virus and Pegiviruses. mBio 2013, 4. [Google Scholar] [CrossRef] [PubMed]
- Firth, C.; Bhat, M.; Firth, M.A.; Williams, S.H.; Frye, M.J.; Simmonds, P.; Conte, J.M.; Ng, J.; Garcia, J.; Bhuva, N.P.; et al. Detection of Zoonotic Pathogens and Characterization of Novel Viruses Carried by Commensal Rattus norvegicus in New York City. mBio 2014, 5, e01933-14. [Google Scholar] [CrossRef] [PubMed]
- Quan, P.-L.; Firth, C.; Conte, J.M.; Williams, S.H.; Zambrana-Torrelio, C.M.; Anthony, S.J.; Ellison, J.A.; Gilbert, A.T.; Kuzmin, I.V.; Niezgoda, M.; et al. Bats are a major natural reservoir for hepaciviruses and pegiviruses. Proc. Natl. Acad. Sci. USA 2013, 110, 8194–8199. [Google Scholar] [CrossRef] [PubMed]
- Drexler, J.F.; Corman, V.M.; Müller, M.A.; Lukashev, A.N.; Gmyl, A.; Coutard, B.; Adam, A.; Ritz, D.; Leijten, L.M.; van Riel, D.; et al. Evidence for Novel Hepaciviruses in Rodents. PLoS Pathog. 2013, 9, e1003438. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, C.V.; Carrique-Mas, J.; Vo Be, H.; An, N.N.; Tue, N.T.; Anh, N.L.; Anh, P.H.; Phuc, N.T.; Baker, S.; Voutilainen, L.; et al. Rodents and Risk in the Mekong Delta of Vietnam: Seroprevalence of Selected Zoonotic Viruses in Rodents and Humans. Vector-Borne Zoonotic Dis. 2015, 15, 65–72. [Google Scholar]
- Khiem, N.T.; Cuong, L.Q.; Chien, H.V. Market study of meat from field rats in the Mekong Delta. In Rats, Mice and People: Rodent Biology and Management; Singleton, G.R., Hinds, L.A., Krebs, C.J., Spratt, D.M., Eds.; ACIAR Monograph, Australian Centre for Internationa Agricultural Research: Canberra, Australia, 2003; pp. 543–547. [Google Scholar]
- Meerburg, B.G.; Singleton, G.R.; Kijlstra, A. Rodent-borne diseases and their risks for public health. Crit. Rev. Microbiol. 2009, 35, 221–270. [Google Scholar] [CrossRef] [PubMed]
- Berto, A.; Anh, P.H.; Carrique-Mas, J.J.; Simmonds, P.; Van Cuong, N.; Tue, N.T.; Van Dung, N.; Woolhouse, M.E.; Smith, I.; Marsh, G.A.; et al. Detection of potentially novel paramyxovirus and coronavirus viral RNA in bats and rats in the Mekong Delta region of southern Viet Nam. Zoonoses Public Health 2018, 65, 30–42. [Google Scholar] [CrossRef] [PubMed]
- Rabaa, M.; Tue, N.; Phuc, T.; Carrique-Mas, J.; Saylors, K.; Cotten, M.; Bryant, J.; Nghia, H.; Cuong, N.; Pham, H.; et al. The Vietnam Initiative on Zoonotic Infections (VIZIONS): A Strategic Approach to Studying Emerging Zoonotic Infectious Diseases. EcoHealth 2015, 12, 726–735. [Google Scholar] [CrossRef] [PubMed]
- Anh, P.H.; Van Cuong, N.; Son, N.T.; Tue, N.T.; Kosoy, M.; Woolhouse, M.E.J.; Baker, S.; Bryant, J.E.; Thwaites, G.; Carrique-Mas, J.J.; et al. Diversity of Bartonella spp. in Bats, Southern Vietnam. Emerg. Infect. Dis. 2015, 21, 1266–1267. [Google Scholar] [CrossRef] [PubMed]
- Loan, H.K.; Cuong, N.V.; Takhampunya, R.; Klangthong, K.; Osikowicz, L.; Kiet, B.T.; Campbell, J.; Bryant, J.; Promstaporn, S.; Kosoy, M.; et al. Bartonella Species and Trombiculid Mites of Rats from the Mekong Delta of Vietnam. Vector Borne Zoonotic Dis. 2015, 15, 40–47. [Google Scholar] [CrossRef] [PubMed]
- Loan, H.K.; Van Cuong, N.; Takhampunya, R.; Kiet, B.T.; Campbell, J.; Them, L.N.; Bryant, J.E.; Tippayachai, B.; Van Hoang, N.; Morand, S.; et al. How Important Are Rats As Vectors of Leptospirosis in the Mekong Delta of Vietnam? Vector Borne Zoonotic Dis. 2015, 15, 56–64. [Google Scholar] [CrossRef] [PubMed]
- Francis, C.M.; Barrett, P. A Guide to the Mammals of Southeast Asia; Princeton University Press: Princeton, NJ, USA, 2008. [Google Scholar]
- Van Dung, N.; Anh, P.H.; Van Cuong, N.; Hoa, N.T.; Carrique-Mas, J.; Hien, V.B.; Sharp, C.; Rabaa, M.; Berto, A.; Campbell, J.; et al. Large-scale screening and characterization of enteroviruses and kobuviruses infecting pigs in Vietnam. J. Gen. Virol. 2016, 97, 378–388. [Google Scholar] [CrossRef] [PubMed]
- Simmonds, P. SSE: A nucleotide and amino acid sequence analysis platform. BMC Res. Notes 2012, 5, 50. [Google Scholar] [CrossRef] [PubMed]
- Smith, D.B.; Becher, P.; Bukh, J.; Gould, E.A.; Meyers, G.; Monath, T.; Muerhoff, A.S.; Pletnev, A.; Rico-Hesse, R.; Stapleton, J.T.; et al. Proposed update to the taxonomy of the genera Hepacivirus and Pegivirus within the Flaviviridae family. J. Gen. Virol. 2016, 97, 2894–2907. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Smith, D.B.; Simmonds, P.; International Committee on the Taxonomy of Viruses Hepeviridae Study Group; Jameel, S.; Emerson, S.U.; Harrison, T.J.; Meng, X.-J.; Okamoto, H.; Van der Poel, W.H.M.; Purdy, M.A. Consensus proposals for classification of the family Hepeviridae. J. Gen. Virol. 2014, 95, 2223–2232. [Google Scholar] [CrossRef] [PubMed]
- Coltman, D.W.; Pilkington, J.G.; Smith, J.A.; Pemberton, J.M. Parasite-Mediated Selection against Inbred Soay Sheep in a Free-Living, Island Population. Evolution 1999, 53, 1259–1267. [Google Scholar] [CrossRef] [PubMed]
- Acevedo-Whitehouse, K.; Gulland, F.; Greig, D.; Amos, W. Disease susceptibility in California sea lions. Nature 2003, 422, 35. [Google Scholar] [CrossRef] [PubMed]
- Lyons, E.J.; Frodsham, A.J.; Zhang, L.; Hill, A.V.S.; Amos, W. Consanguinity and susceptibility to infectious diseases in humans. Biol. Lett. 2009, 5, 574–576. [Google Scholar] [CrossRef] [PubMed]
- Pybus, O.G.; Thézé, J. Hepacivirus cross-species transmission and the origins of the hepatitis C virus. Curr. Opin. Virol. 2016, 16, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Lyons, S.; Kapoor, A.; Schneider, B.S.; Wolfe, N.D.; Culshaw, G.; Corcoran, B.; Durham, A.E.; Burden, F.; McGorum, B.C.; Simmonds, P. Viraemic frequencies and seroprevalence of non-primate hepacivirus and equine pegiviruses in horses and other mammalian species. J. Gen. Virol. 2014, 95, 1701–1711. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Pybus, O.G.; Gray, R.R. Virology: The virus whose family expanded. Nature 2013, 498, 310–311. [Google Scholar] [CrossRef] [PubMed]
- Ramsay, J.D.; Evanoff, R.; Wilkinson, T.E.; Divers, T.J.; Knowles, D.P.; Mealey, R.H. Experimental transmission of equine hepacivirus in horses as a model for hepatitis C virus. Hepatology 2015, 61, 1533–1546. [Google Scholar] [CrossRef] [PubMed]
- Gather, T.; Walter, S.; Todt, D.; Pfaender, S.; Brown, R.J.P.; Postel, A.; Becher, P.; Moritz, A.; Hansmann, F.; Baumgaertner, W.; et al. Vertical transmission of hepatitis C virus-like non-primate hepacivirus in horses. J. Gen. Virol. 2016, 97, 2540–2551. [Google Scholar] [CrossRef] [PubMed]
- Billerbeck, E.; Wolfisberg, R.; Fahnøe, U.; Xiao, J.W.; Quirk, C.; Luna, J.M.; Cullen, J.M.; Hartlage, A.S.; Chiriboga, L.; Ghoshal, K.; et al. Mouse models of acute and chronic hepacivirus infection. Science 2017, 357, 204–208. [Google Scholar] [CrossRef] [PubMed]

| Species | Tested | Positive (%) | ||
|---|---|---|---|---|
| Hepacivirus | HEV | HBV | ||
| Bandicota indica | 38 | 0 | 1 (2.6) | 0 |
| Rattus argentiventer | 275 | 2 (0.7) | 10 (3.6) | 0 |
| Rattus losea | 19 | 1 (5.3) | 2 (10.5) | 0 |
| Rattus norvegicus | 39 | 0 | 0 | 0 |
| Rattus tanezumi | 17 | 0 | 1 (5.9) | 0 |
| Rhizomys pruinosus | 82 | 35 (42.7) | 0 | 0 |
| Total | 470 | 38 (8.1) | 14 (3) | 0 |
| Virus | Sequence | Compared Region | Highest Nucleotide Identity (%) | Highest Amino Acid Identity (%) | Closest Match |
|---|---|---|---|---|---|
| Rodent hepacivirus | 05VZ-14-103 | Complete cds | 59.7 | 63.3 | KC815310 |
| 05VZ-14-104 | Complete cds | 59.7 | 63.2 | KC815310 | |
| 05VZ-14-118 | Complete cds | 59.5 | 63.3 | KC815310 | |
| 05VZ-14-119 | Complete cds | 59.5 | 63.3 | KC815310 | |
| Rodent HEV | 05VZ-75-65-L08-R3 | ORF1 + ORF2 | 81.5 | 93.2 | JX120573 |
| Bat HBV | Bat031 | P gene | 89 | 87 | KY905324 |
| S gene | 91 | 94 | KY905324 | ||
| X gene | 92.4 | 86 | KY905324 | ||
| C gene | 90 | 96.3 | KY905324 | ||
| Bat033 | P gene | 82.5 | 80 | KY905328 | |
| S gene | 87 | 84 | KY905328 | ||
| X gene | 90 | 80 | KY905324 | ||
| C gene | 89 | 97.2 | KY905327 | ||
| Rodent pegivirus | 05VZ-14-070 | Complete cds | 65.2 | 65.5 | KJ950934 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Van Nguyen, D.; Van Nguyen, C.; Bonsall, D.; Ngo, T.T.; Carrique-Mas, J.; Pham, A.H.; Bryant, J.E.; Thwaites, G.; Baker, S.; Woolhouse, M.; et al. Detection and Characterization of Homologues of Human Hepatitis Viruses and Pegiviruses in Rodents and Bats in Vietnam. Viruses 2018, 10, 102. https://doi.org/10.3390/v10030102
Van Nguyen D, Van Nguyen C, Bonsall D, Ngo TT, Carrique-Mas J, Pham AH, Bryant JE, Thwaites G, Baker S, Woolhouse M, et al. Detection and Characterization of Homologues of Human Hepatitis Viruses and Pegiviruses in Rodents and Bats in Vietnam. Viruses. 2018; 10(3):102. https://doi.org/10.3390/v10030102
Chicago/Turabian StyleVan Nguyen, Dung, Cuong Van Nguyen, David Bonsall, Tue Tri Ngo, Juan Carrique-Mas, Anh Hong Pham, Juliet E. Bryant, Guy Thwaites, Stephen Baker, Mark Woolhouse, and et al. 2018. "Detection and Characterization of Homologues of Human Hepatitis Viruses and Pegiviruses in Rodents and Bats in Vietnam" Viruses 10, no. 3: 102. https://doi.org/10.3390/v10030102
APA StyleVan Nguyen, D., Van Nguyen, C., Bonsall, D., Ngo, T. T., Carrique-Mas, J., Pham, A. H., Bryant, J. E., Thwaites, G., Baker, S., Woolhouse, M., & Simmonds, P. (2018). Detection and Characterization of Homologues of Human Hepatitis Viruses and Pegiviruses in Rodents and Bats in Vietnam. Viruses, 10(3), 102. https://doi.org/10.3390/v10030102





