Abstract
Paulownia witches’ broom (PaWB) disease caused by phytoplasmas is a fatal disease that leads to considerable economic losses. Long non-coding RNAs (lncRNAs) have been demonstrated to play critical regulatory roles in posttranscriptional and transcriptional regulation. However, lncRNAs and their functional roles remain poorly characterized in Paulownia. To identify lncRNAs and investigate their roles in the response to PaWB phytoplasmas, RNA sequencing was performed for healthy Paulownia tomentosa, PaWB-infected P. tomentosa, and for healthy and PaWB-infected P. tomentosa treated with 100 mg L−1 rifampicin. A total of 28,614 unique mRNAs and 3693 potential lncRNAs were identified. Comparisons between lncRNAs and coding genes indicated that lncRNAs tended to have shorter transcripts and fewer exon numbers, and displayed significant expression specificity. Based on our comparison scheme, 1063 PaWB-related mRNAs and 110 PaWB-related lncRNAs were identified; among them, 12 PaWB-related candidate target genes that were regulated by nine PaWB-related lncRNAs were characterized. This study provides the first catalog of lncRNAs expressed in Paulownia and gives a revealing insight into the molecular mechanism responsible for PaWB.
1. Introduction
Phytoplasmas are minute bacteria without cell walls and are the causal agents of witches’ broom disease in Paulownia. Witches’ broom disease occurs in many countries and is the biggest threat to Paulownia production, causing serious economic losses [1,2]. In Paulownia, witches’ broom disease has been found in saplings and big trees, and leads to malformations of branch, leaf, stem, and flower, such as witches’ broom, yellowing, phloem necrosis, and phyllody, respectively [3,4].
Paulownia trees are native to China and have been introduced into many countries [5]. The superior traits of Paulownia, such as distinguished adaptive capacity to poor habitat, extremely fast growth, and high-quality timber, make it a popular tree species with important economic and ecological values [6]. Accordingly, Paulownia witches’ broom (PaWB) disease has been studied by biologists for many years. Vitamin C plays an important role in the resistance to PaWB [7], and plant hormones are known to be related to the morphogenesis of PaWB [8]. The Sec protein translocation system [9], elongation factor EF-Tu [10], and two plasmids [11] of the PaWB phytoplasma have been analyzed. Transgenic technology has been used in the breeding of PaWB-resistant Paulownia [12]. Rifampicin treatment can make PaWB-infected Paulownia recover to normal morphology [13]. Furthermore, the rapid development of ‘omics’ has allowed for the investigation of the transcriptomes [3,14,15,16,17,18], microRNAs (miRNAs), degradomes [19,20,21,22], and proteomes [4,23] of Paulownia species, and the results have revealed changes after PaWB infection at transcriptional, post-transcriptional, and translational levels.
Long non-coding RNAs (lncRNAs) are generally defined as non-coding transcripts that are more than 200 nucleotides in length [24]. Large numbers of lncRNAs have been identified in Arabidopsis thaliana [25,26,27], rice [28,29], wheat [30,31], maize [32], cotton [33,34], Brassica napus [35], and Populus [36,37], and have been found to play important roles in growth, development, differentiation, and stress responses. LncRNAs involved in the responses of plants to biotic stresses have also been identified. For example, in wheat, four lncRNAs involved in the response to stripe rust pathogen infection have been identified [31]; in Arabidopsis, lncRNAs responsive to Fusarium oxysporum infection were revealed and played an important role in antifungal immunity [38]; and in tomato, lncRNAs have been identified as endogenous target mimics for miRNAs in response to Tomato yellow leaf curl virus infection [39], and a lncRNA (lncRNA16397) that conveys resistance to Phytophthora infestans by co-expressing glutaredoxin has also been detected [40]. However, in studies of PaWB, lncRNAs that play important roles in the Paulownia–phytoplasma interaction have not yet been identified.
In this study, we used next-generation deep-sequencing technologies to identify PaWB-related genes and lncRNAs among healthy Paulownia tomentosa (PT), PaWB-infected P. tomentosa (PTI), and PT and PTI treated with 100 mg L−1 rifampicin (PT-100 and PTI-100, respectively). Our results may help to clarify the molecular mechanisms of PaWB, and accelerate the process of preventing the disastrous effects of PaWB.
2. Materials and Methods
2.1. Plant Materials
All the biological materials used in this study were obtained from the Institute of Paulownia, Henan Agricultural University, China. The following four groups of P. tomentosa seedlings were set up: PT, PTI, PT-100, and PTI-100. The cultivation and rifampin treatment procedures were as described by Fan et al. [13]. The terminal buds from three individual plants were combined to form one biological replicate, and at least three biological replicates were used for each treatment.
2.2. Library Construction and Sequencing
Total RNA was extracted from the terminal buds of P. tomentosa using TRIzol reagent (Invitrogen, Carlsbad, CA, USA) and treated with RNase-free DNase (Promega, Madison, WI, USA) according to the manufacturer’s instructions. The quality and quantity of total RNA were measured using an Agilent 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA). Total RNA was used to construct RNA sequencing (RNA-seq) libraries using a TruSeq Stranded Total RNA preparation kit with RiboZero Plant (Illumina Inc., San Diego, CA, USA) according to the manufacturer’s instructions. The libraries were sequenced on an Illumina HiSeq 2000 platform by the Beijing Genomics Institute, Shenzhen, Guang dong, China.
2.3. Genome Mapping and Transcript Assembly
Reads with more than 10% N (unable to determine base information), with adapter sequence, or of low quality were removed from the raw reads to obtain clean reads. To remove ribosomal RNA reads, the clean reads were mapped to the SILVA rRNA database (http://www.arb-silva.de/) using the short read alignment software SOAP2 (BGI, Shenzhen, China). Finally, the clean reads were mapped to the Paulownia genome sequence (http://paulownia.genomics.cn) using TopHat2 [41]. Cufflinks 2.0.0 was used to assemble the mapped reads [42]. Transcripts that were no less than 200 nucleotides in length were selected for further analysis.
2.4. LncRNAs Identification
Based on the assembled transcripts annotated to the Paulownia genome sequence, transcripts encoding known proteins were identified and the unknown transcripts (<200 bp) were excluded. Non-coding transcripts were identified based on a coding potential score of less than 0, as calculated by the Coding Potential Calculator (CPC; http://cpc.cbi.pku.edu.cn). The candidate lncRNAs were classified into several categories, according to their genomic locations and previous descriptions of Sun et al. [43].
2.5. Genomic Characterization and Function Prediction of lncRNAs
We determined the distribution of the identified lncRNA on the Paulownia chromosomes according to the description of Li et al. [44]. To explore lncRNA conservation, all the lncRNA sequences identified in this study were aligned against the CANTATAdb (http://yeti.amu.edu.pl/CANTATA) using BLASTN (E-value < 1×10−5). To identify lncRNAs that may act as miRNA precursors, we aligned the lncRNA sequences to the miRNA precursor sequences in miRBase (Release 21, http://www.mirbase.org/). The secondary structures of the lncRNA and miRNA precursor sequences were predicted using RNAfold in the Vienna RNA package (http://rna.tbi.univie.ac.at/). To explore the homology of the lncRNAs based on their consensus RNA secondary structures, we classified the predicted lncRNAs into different non-coding RNA families using INFERNAL [45].
We used an algorithm to search for potential cis target genes within a 10-kb region upstream and downstream of the identified lncRNAs. Potential trans target genes were detected based on sequence complementarity and RNA duplex energy prediction to assess the impact of lncRNA binding on complete mRNAs. BLAST was used to determine sequence complementarity (identify ≥ 95%, E-value < 0.05), and RNAplex was used to calculated the energy of potential mRNA–lncRNA interactions. The RNAplex parameters were set as e < −30. The criteria used for the prediction of potential trans targets are described in previous studies [36,46,47].
2.6. Identification of PaWB-Related Genes and lncRNAs
All the sequencing data were uploaded to SRA database with accession number SRP117715. Expression levels of genes and lncRNAs were measured as fragments per kilobase of exon model per million mapped reads (FPKM). Differentially expressed genes (DEGs) and differentially expressed lncRNAs (DELs) were identified according to the methods described in previous studies [36,40].
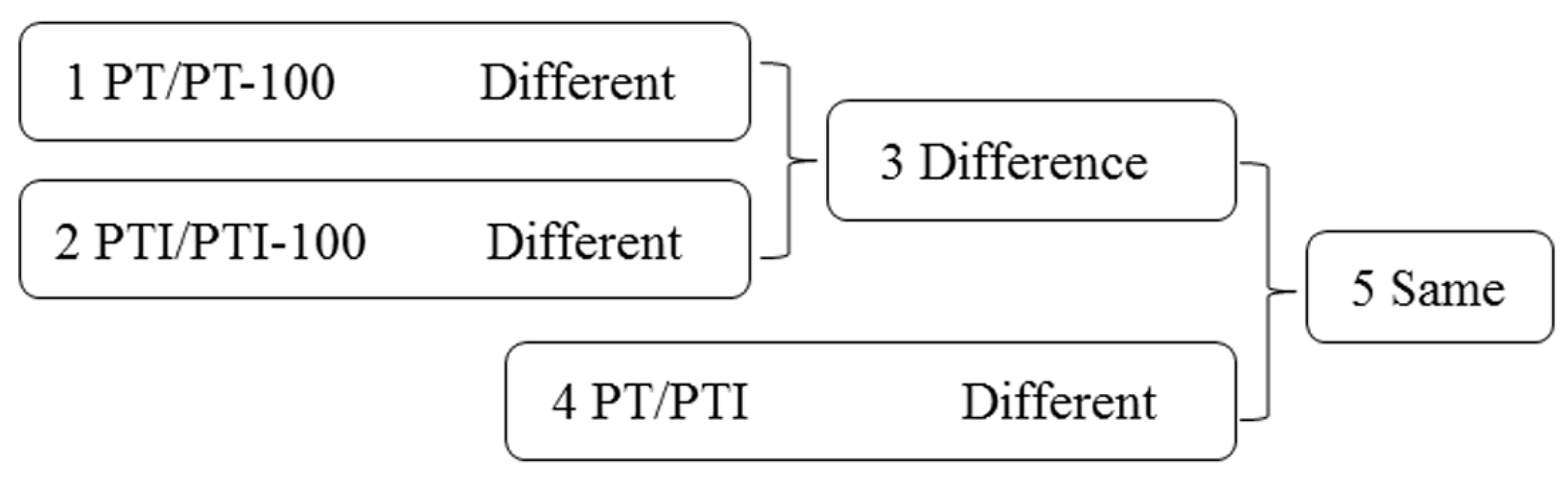
To identify genes and lncRNAs related to PaWB, we made comparisons among the four libraries (Figure 1). The following comparison schemes were considered: (1) DEGs and DELs in PT/PT-100 may be related to the influence of rifampicin; (2) DEGs and DELs in PTI/PTI-100 may be related to the influence of rifampicin and PaWB; (3) Differences between comparisons 1 and 2 may exclude DEGs and DELs related to the influence of rifampicin; (4) DEGs and DELs in PT/PTI may be related to PaWB; (5) The common DEGs and DELs between comparisons 3 and 4 may be related directly to PaWB.
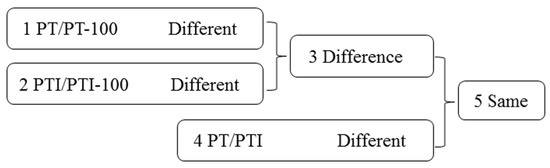
Figure 1.
Comparison schemes of the four samples. PT represents the healthy wild-type sample of P. tomentosa, PTI represents the sample of phytoplasma infected PT. PT-100 represents the sample of 100 mg L−1 rifampicin treated PT, PTI-100 represents the sample of 100 mg L−1 rifampicin treated PTI. Different represents differentially expressed genes/long non-coding RNAs (lncRNAs). Difference represents the union of two groups, except their intersection. Same represents the intersection of two groups.
2.7. Nested-PCR and Quantitative RT-PCR
We applied nested-PCR to detect phytoplasma in seedlings according to the method described by Fan et al. [13]. Total RNA was reverse transcribed into cDNA and used to measure the expression of lncRNAs by quantitative real-time PCR (qRT-PCR). The qRT-PCRs were performed on a DNA Engine Opticon machine (MJ Research, Waltham, MA, USA) using a LightCycler FastStar DNA master SYBR Green I kit (Roche Diagnostics, Mannheim, Germany). The primers were designed using Primer Express 3.0 (Applied Biosystems, Stockholm, Sweden) and the specificity of primer pairs was checked by sequencing the PCR products. All qRT-PCR amplifications were carried out in triplicate, with the standard reaction program. The specificity of the amplified fragments was checked using the generated melting curve. The generated real-time data were analyzed using the Opticon Monitor Analysis Software 3.1 (Bio-Rad, Hercules, CA, USA) tool and standardized to the levels of 18S RNA using the 2−ΔΔCt method [48]. The primers used for the qRT-PCRs are listed in Table S1.
3. Results
3.1. Changes in Morphology of the Four Samples
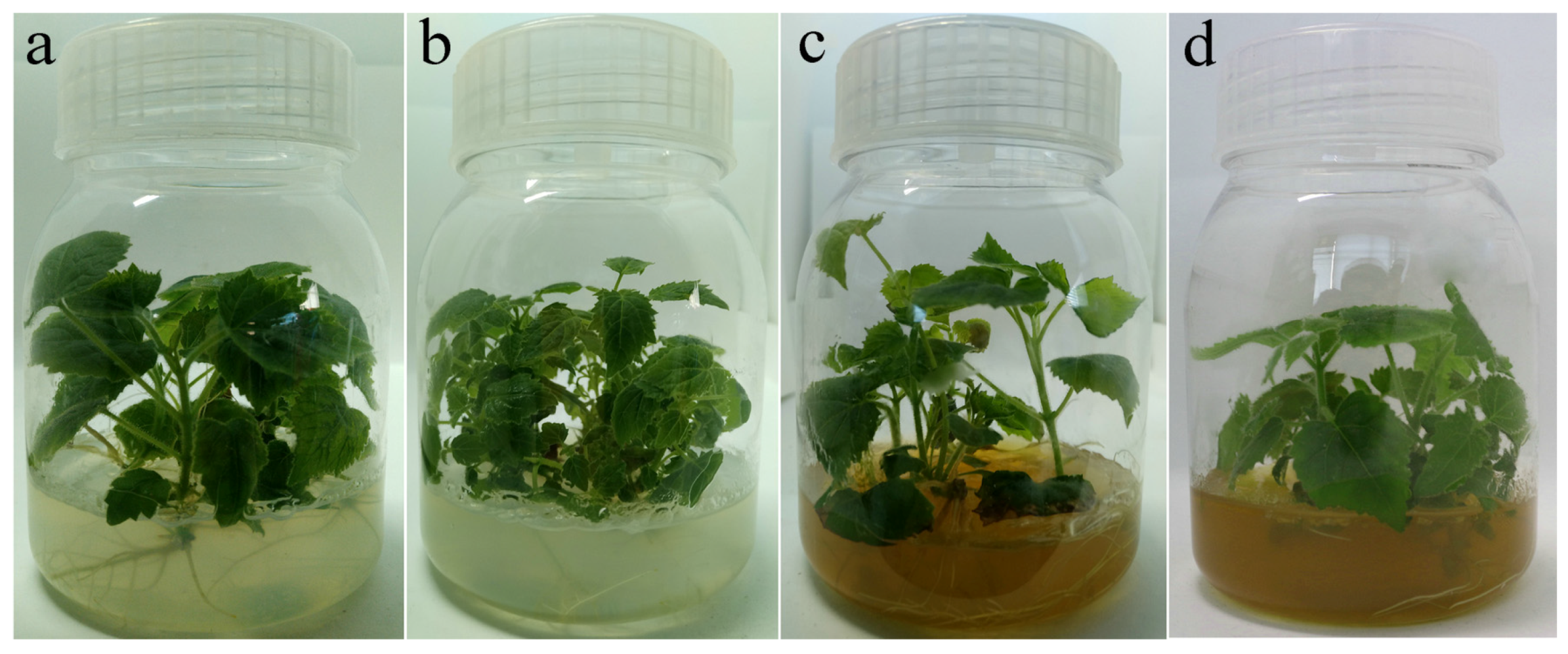
Compared with PT (Figure 2a), PTI (Figure 2b) showed typical PaWB symptom, such as witches’ broom, and yellowing and relatively small leaves. In the rifampicin treated-samples (PT-100 and PTI-100), PT-100 (Figure 2c) showed no significant differences compared with PT. PTI-100 (Figure 2d) showed asymptomatic morphology. We applied nest-PCR to detect phytoplasma DNA in the four samples, and found that phytoplasma DNA was detected only in PTI (Figure S1).

Figure 2.
Rifampin treated plantlets morphology. (a) PT; (b) PTI; (c) PT-100; (d) PTI-100.
3.2. High-Throughput Sequencing and Differentially Expressed Genes Analysis
Four cDNA libraries were constructed using the samples, and sequenced on an Illumina HiSeq 2000 platform. Among the clean reads, we identified 26,155, 27,017, 26,334, and 26,826 unique mRNAs from the PT, PTI, PT-100, and PTI-100 libraries, respectively (Table 1 and Table S2). A comparative summary of the mRNAs in the four libraries is provided as a Venn diagram (Figure S2a); 24,282 unique mRNAs were common among the four libraries, whereas 367, 337, 229, and 320 unique mRNAs were only presented in PT, PTI, PT-100, and PTI-100 libraries, respectively. In the PT/PT-100, PTI/PTI-100, and PT/PTI comparisons, we detected 1491 DEGs (706 upregulated and 785 downregulated), 1802 DEGs (725 upregulated and 1077 downregulated), and 3744 DEGs (2208 upregulated and 1536 downregulated), respectively (Table S2). Based on our comparison scheme (Figure 1), 2481 specific DEGs from comparison 1 (PT/PT-100) and comparison 2 (PTI/PTI-100) were identified (comparison 3), and 1063 common DEGs from comparison 3 and comparison 4 (PT/PTI) were PaWB-related genes. (Figure S3a and Table S2).

Table 1.
Statistical data of the RNA-Seq reads.
3.3. Genome-Wide Identification and Characterization of lncRNAs in P. tomentosa
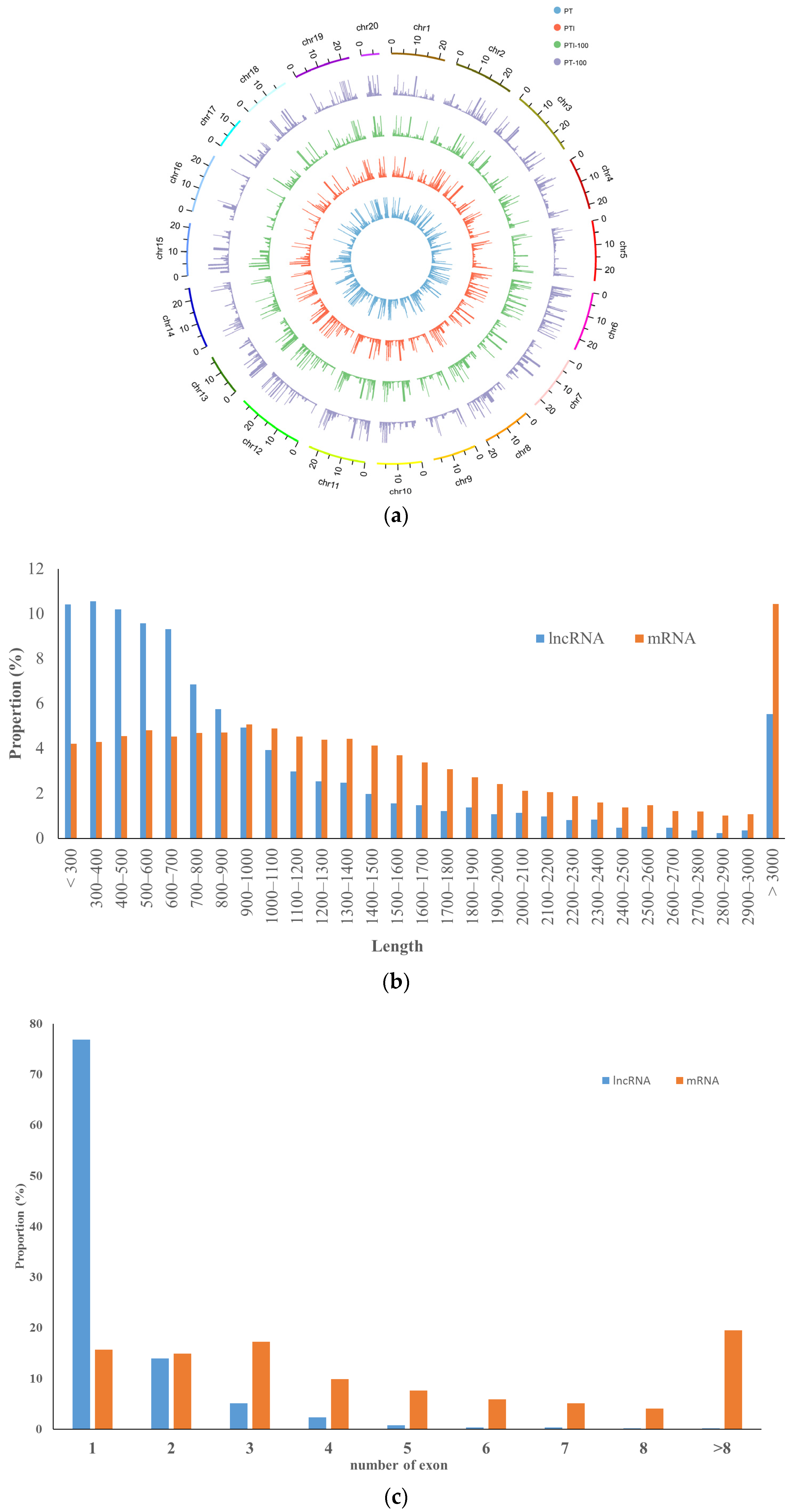
Based on our analysis, we identified 3434, 3504, 3520, and 3479 lncRNAs in the PT, PTI, PT-100, and PTI-100 libraries, respectively (Table S3). A comparative summary of the mRNAs in the four libraries is provided as a Venn diagram (Figure S2b); 3102 unique lncRNAs were common among the four libraries, whereas 10, 2, 1 and 3 unique lncRNAs were only presented in PT, PTI, PT-100, and PTI-100 libraries, respectively. A total of 3693 lncRNAs were revealed from the four libraries and classified into four categories: 41 into classcode “j”, novel long noncoding isoforms with at least one splice junction shared with the reference genes; 73 into classcode “o”, exonic overlap with a known transcript; 578 into classcode “x”, exonic overlap with reference on the opposite strand; and 3001 into classcode “u”, intergenic lncRNAs (Figure S4a). The INFERNAL analysis results showed that 493 of the unique lncRNAs belonged to 155 conserved lncRNAs families (Table S4). About half of the lncRNAs belonged to miRNA families, such as MIR159, MIR169, and MIR171, and five lncRNAs belonged to two small nucleolar RNA families (SNORD25 and SNORD27). Some non-plant-specific lncRNA families (X inactive specific transcript, small cajal body-specific RNA 8, fungal small nucleolar RNA, bacterial small transcript non-coding 150) were also found. Almost all the lncRNAs were distributed among the chromosomes and did not show significant chromosome location preferences (Figure 3a). The lengths of the lncRNAs were usually shorter than the lengths of the mRNAs, and more than 10% of all lncRNAs were ≤300, 300–400, or 400–500 nucleotides long, whereas about 10% of the mRNA were ≥3000 nucleotides in length (Figure 3b). Most lncRNAs have only exons, while most mRNAs have more than one exon (Figure 3c). All the unique lncRNAs were searched against the genomes of Arabidopsis thaliana, Oryza sativa, Glycine max, Selaginella, Chlamydomonas, Physcomitrella, Amborella, potato, Vitis vinifera, and Zea mays using BLAST. The results showed that only a small number of the P. tomentosa lncRNAs were conserved across the ten species; the highest number on lncRNAs matched V. vinifera lncRNAs, likely because Paulownia and V. vinifera are the most closely related in evolution (Table S5).
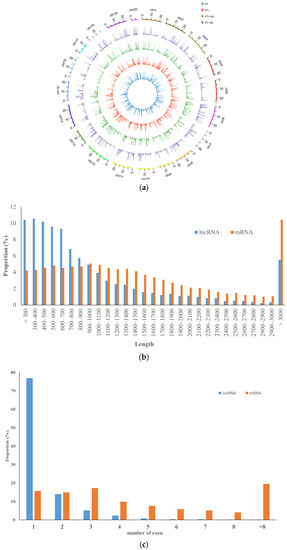
Figure 3.
Characteristics of Paulownia lncRNAs. (a) Distribution of lncRNAs along each chromosome; (b) Transcript length distribution of lncRNAs and mRNAs; (c) Number of exons per transcripts for lncRNAs and mRNAs.
3.4. Differential Expression of lncRNAs
The expression levels of the 3693 lncRNAs in the four libraries were measured by FPKM. Based on log2 fold change >1 and p < 0.05, we identified 750 (200 upregulated and 550 downregulated), 123 (63 upregulated and 60 downregulated), and 173 (98 upregulated and 75 downregulated) differentially expressed lncRNAs (DELs) in PT/PTI, PT/PT-100, and PTI/PTI-100, respectively (Table S3).
3.5. Prediction of lncRNA Candidate Target Genes
Because lncRNAs play important roles in regulating gene expression, identification and analysis of their candidate target genes could help understand their functions. A total of 6746 lncRNA–mRNA pairs were identified (4175 were cis-regulated and 2571 were trans-regulated, Table S6). In total, we detected 5333 candidate target genes for 3222 lncRNAs; among them, 3218 were cis target genes for 2290 lncRNAs, and 2401 were trans target genes for 2571 lncRNAs. We found that one lncRNA could have more than one target gene, and one target gene could be targeted by one or more lncRNAs. Among the 3222 lncRNAs, 1246 had a single candidate target gene, while nine candidate target genes were predicted for each of two lncRNAs (TCONS_00019479 and TCONS_00030009). Furthermore, among the 5333 candidate target genes, 4313 were targeted by one lncRNA, while the other two target genes (TCONS_00013387 and TCONS_00019818) were targeted by more than one lncRNA (each of the target genes were targeted by 11 lncRNAs) (Table S6).
3.6. Analysis of PaWB-Related lncRNAs
Based on our comparisons (Figure 1), 234 specific DELs from comparison 1 (PT/PT-100) and comparison 2 (PTI/PTI-100) were identified (comparison 3), 110 common DELs from comparison 3 and comparison 4 (PT/PTI) were PaWB-related lncRNAs (Figure S3b and Table S3). Among the 110 PaWB-related DELs, 5, 30, and 75 lncRNAs were assigned classcode “o”, “x”, and “u”, respectively (Figure S4b). Cis- or trans-regulated target genes were predicted for 80 of the 110 PaWB-related lncRNAs, and 3, 23, and 54 of them were assigned classcode “o”, “x”, and “u”, respectively. A total of 147 lncRNA–mRNA pairs were identified for the 80 PaWB-related lncRNAs (91 pairs were cis-regulated and 56 were trans-regulated). In total, 138 candidate target genes were predicted for the 80 lncRNAs: 82 cis target genes for 58 lncRNAs, and 56 trans target genes for 56 lncRNAs.
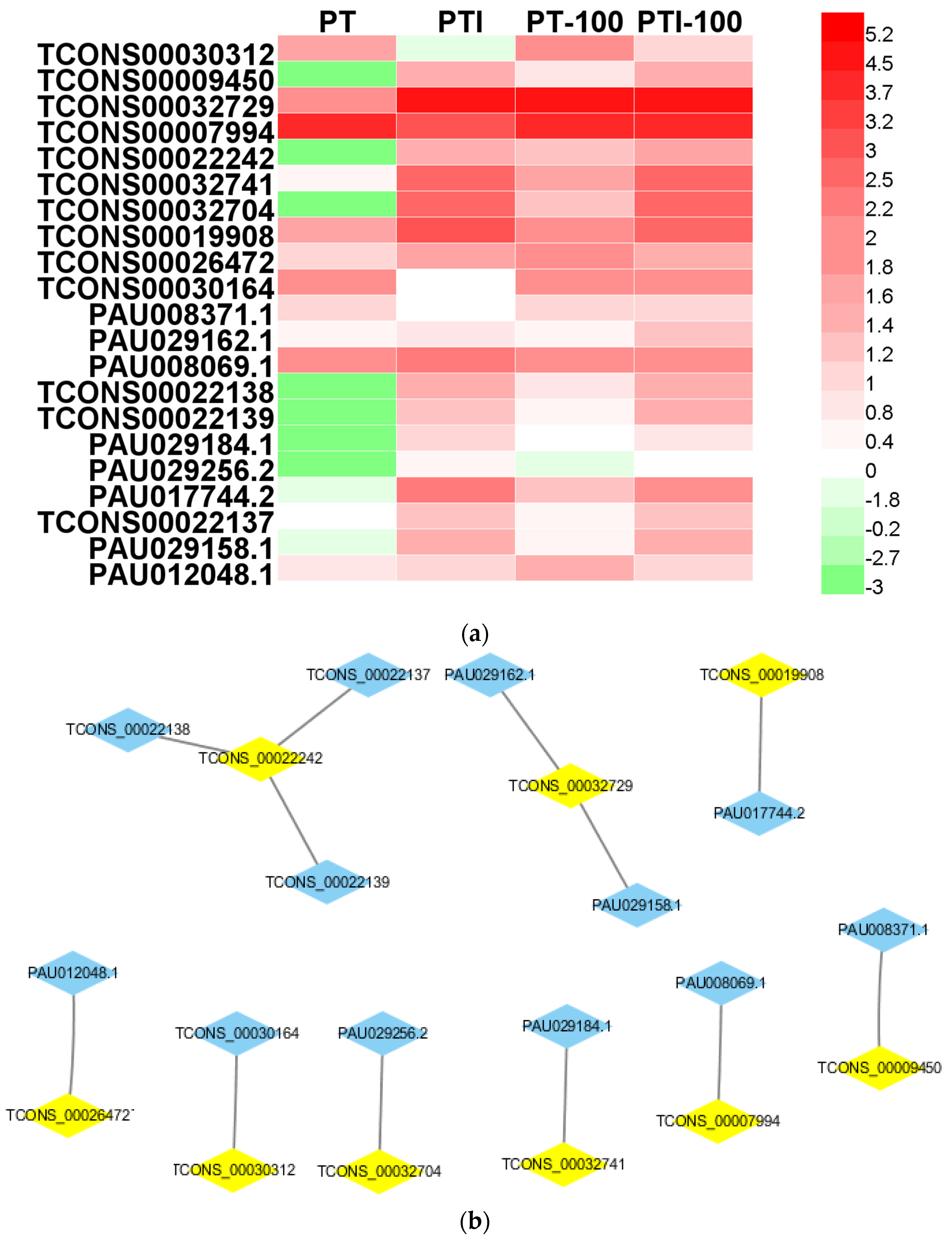
We compared the 138 candidate targets of the 80 PaWB-related lncRNAs with the 1063 PaWB-related DEGs, and identified 12 PaWB-related candidate target genes among the DEGs that were regulated by nine PaWB-related lncRNAs (Table 2 and Figure 4). The KEGG (Kyoto Encyclopedia of Genes and Genomes) analysis results indicated that the 12 candidate target genes were mapped to eight pathways, namely “Phenazine biosynthesis”, “RNA transport”, “Cysteine and methionine metabolism”, “Amino sugar and nucleotide sugar metabolism”, “Glycerolipid metabolism”, “Monoterpenoid biosynthesis”.

Table 2.
Paulownia witches’ broom (PaWB)-related lncRNAs targeting PaWB-related mRNAs.
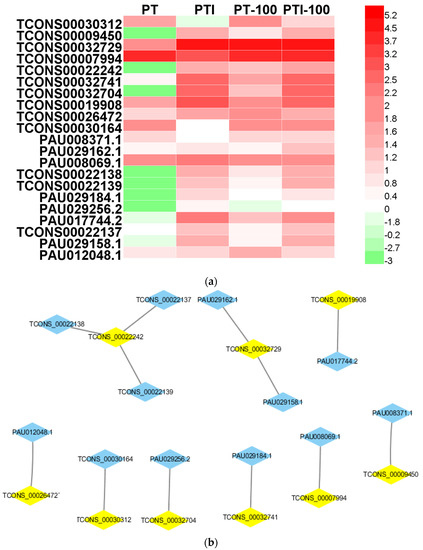
Figure 4.
Characteristics of PaWB-related lncRNAs and mRNAs. (a) Expression profiling in PT, PTI PT-100, and PTI-100. The data were based on lg(FPKM); (b) LncRNA-mRNA networks. The blue polygons represent mRNAs, the yellow polygons represent lncRNAs. FPKM, fragments per kilobase of exon model per million mapped reads.
3.7. Quantitative RT-PCR Analysis of DEGs and DELs
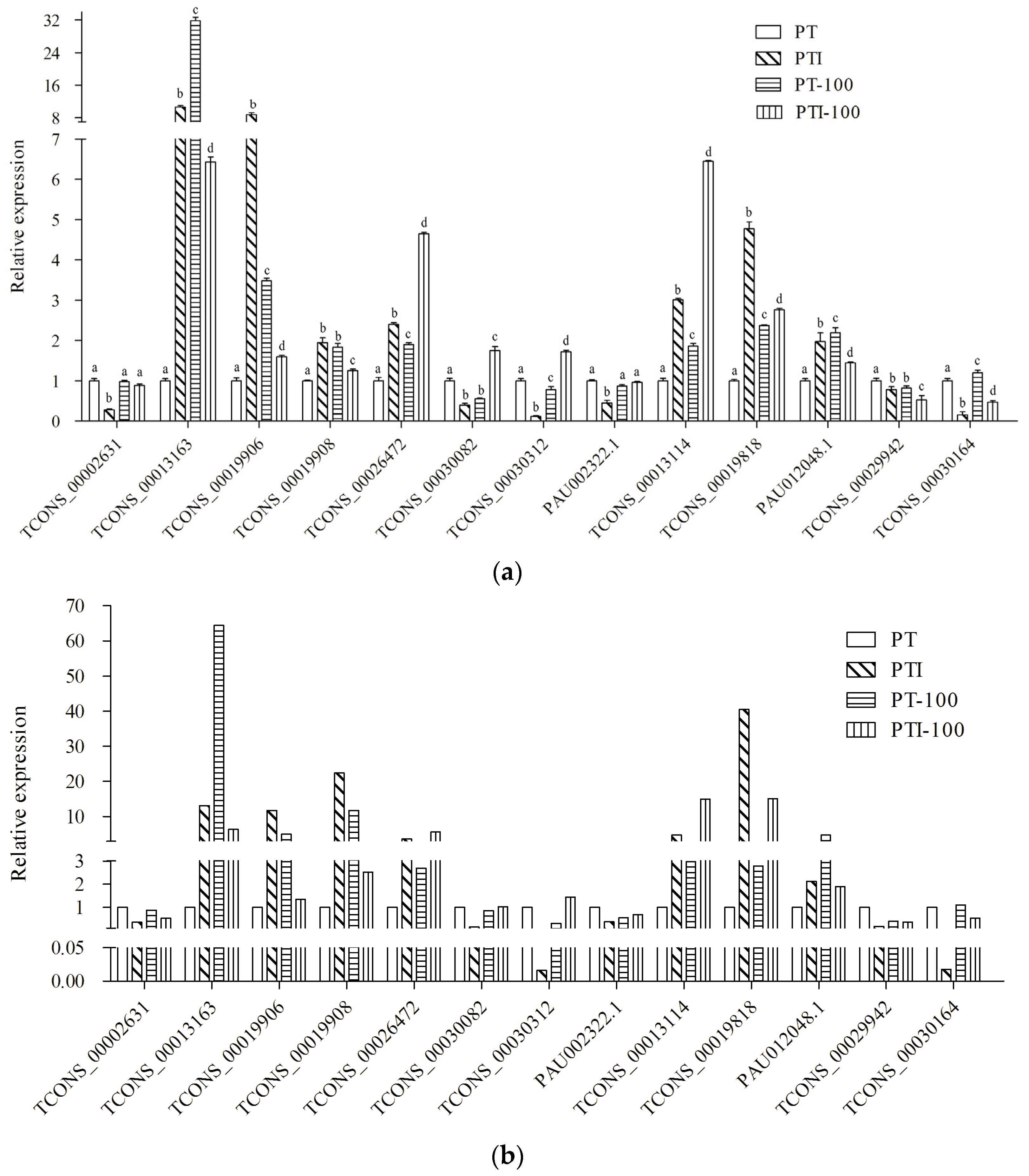
To confirm the expression of the P. tomentosa lncRNAs and their response to PaWB, qRT-PCR analysis was performed to verify the results of the high-throughput RNA-seq. We randomly selected seven lncRNAs and six mRNAs for qRT-PCR validation. The results demonstrated that, except TCONS_00029942, the qRT-PCR results were consistent with those from the RNA-seq analysis (Figure 5). This correlation confirmed that the RNA-seq data were reliable.
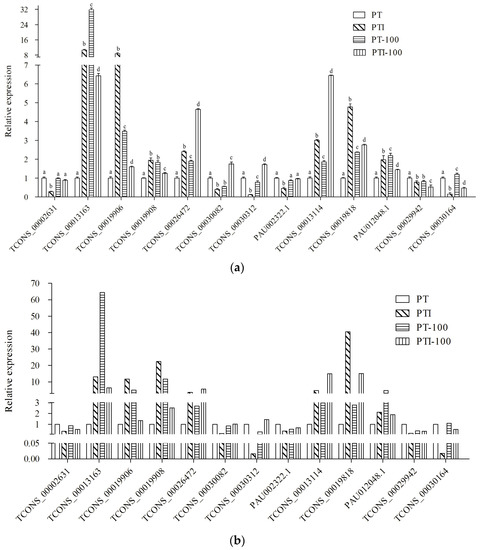
Figure 5.
Expression pattern confirmation by qRT-PCR. (a) Changes in the relative expression levels as determined by qRT-PCR. Standard error of the mean for three technical replicates is represented by the error bars. Samples marked with various letters show a significant difference at p < 0.05; (b) Changes in the relative expression levels as determined by RNA-seq. The expression level of PT was set to 1.
4. Discussion
LncRNAs play key roles in various biological pathways in animals and plants. There have been recent advances in RNA-seq which, when combined with genome-wide mapping, have resulted in the identification of lncRNAs in plants. The identification of plant lncRNAs has opened up the investigation of novel regulatory pathways in plants. For example, 7655 lncRNAs from control and gibberellin-treated Populus tomentosa have been identified, indicating that lncRNAs may participate in auxin signal transduction and synthesis of cellulose and pectin, and may influence growth and wood properties. In the present study, we investigated transcriptomic changes in PT, PTI, PT-100, and PTI-100 seedlings, and systematically identified genes and lncRNAs associated with PaWB. Our results represent the first comprehensive analysis of lncRNAs in Paulownia. The identified lncRNAs will be useful for other woody plants researchers and provide an important resource for future functional genomics and regulatory expression studies.
The Paulownia lncRNAs had fewer exons and were shorter than mRNAs in length, which is consistent with other studies [35,44]. BLAST searches of the Paulownia lncRNAs against the genomes of species such as Arabidopsis thaliana, Oryza sativa, and Glycine max revealed that the majority of Paulownia lncRNAs were not conserved with lncRNAs in other species. The low conservation suggested that lncRNAs may undergo rapid evolution. It is not surprising that lncRNAs are not well conserved for various reasons. First, lncRNAs are not constrained by codon usage. Second, although lncRNAs may possess short conserved motifs, these short motifs are not easily identifiable by local alignment software such as BLAST. Third, some lncRNAs may interact directly with RNA-binding proteins through conserved secondary structures.
Xyloglucan (XG) is the main hemicellulose of the primary cell wall in dicotyledons. Xylosyl transferases (XT, EC:2.4.1.207) are a family of enzymes that catalyze xyloglucan endotransglucosylase and/or xyloglucan endohydrolase [49]. XTs can cut xyloglucan chains and transfer the fragment with the new reducing end either to another XG or to water, and play an important role in cell wall remodeling, affecting plant growth and development [50]. Transglycosylation can contribute to both cell wall assembly and cell wall loosening [51]. In this study, XT was the predicted target gene of lncRNA (TCONS_00022242), and it was upregulated in PT/PTI. XT may be related to witches’ broom and hyperplasia because of its roles in cell wall assembly and loosening.
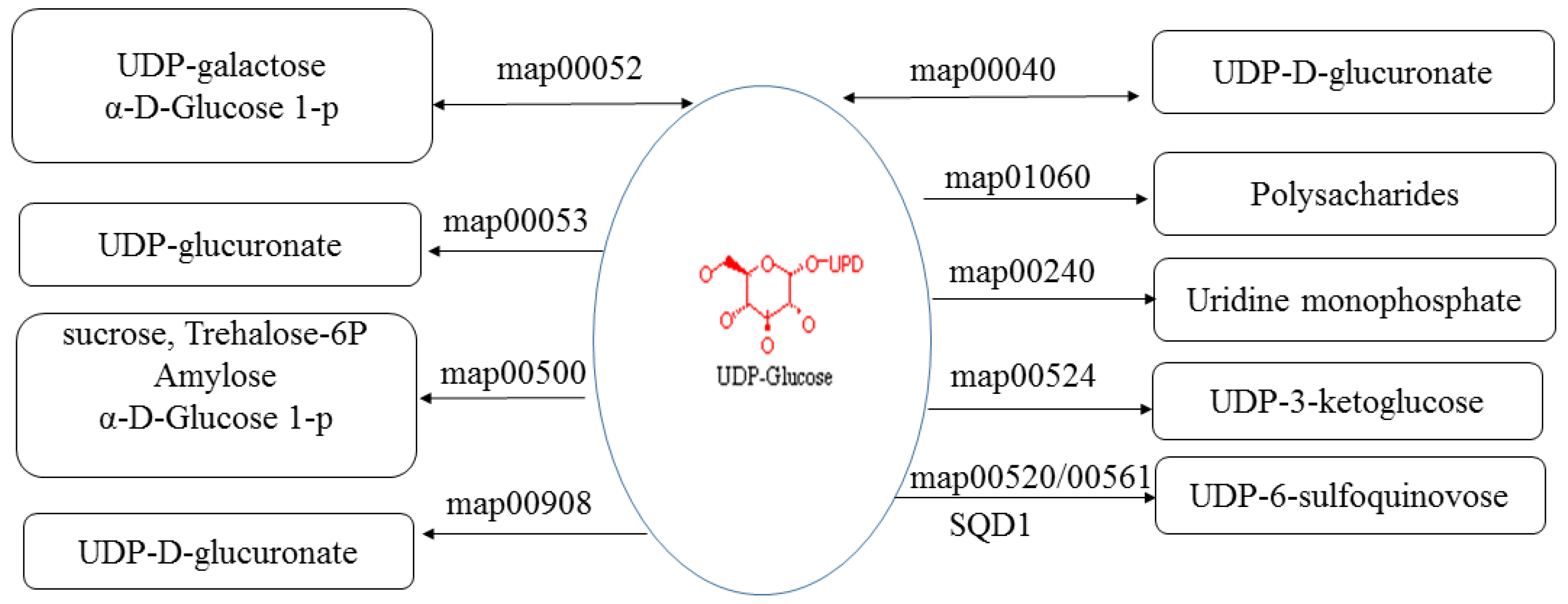
UDP-sulfoquinovose synthase (SQD1) catalyzes the transfer of sulfite to UDP-glucose to produce UDP-sulfoquinovose, which is the head group donor for the biosynthesis of the plant sulfolipid sulfoquinovosyldiacylglyerol (SQDG), a unique nonphosphorous lipid found in the photosynthetic membranes of plants [52]. SQD1 (PAU029158.1) was the candidate target of PaWB-related lncRNA (TCONS_00032729), and it was upregulated in PT/PTI and downregulated in PTI/PTI-100. The changes in SQD1 expression may influence SQDG and then photosynthesis. In addition, UDP-glucose is involved in many pathways (Figure 6), including Pentose and glucuronate interconversions (map00040), Galactose metabolism (map00052), Ascorbate and aldarate metabolism (map00053), Pyrimidine metabolism (map00240), Starch and sucrose metabolism (map00500), Amino sugar and nucleotide sugar metabolism (map00520), Neomycin, kanamycin and gentamicin biosynthesis (map00524), Glycerolipid metabolism (map00561), Zeatin biosynthesis (map00908), Biosynthesis of plant secondary metabolites (map01060), and Metabolic pathways (map01100). Therefore, changes in SQD1 expression may influence UDP-glucose content and the pathways in which UDP-glucose is involved.
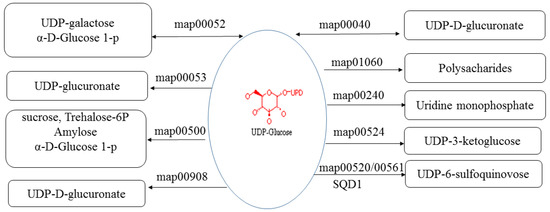
Figure 6.
KEGG (Kyoto Encyclopedia of Genes and Genomes) pathways which UDP-glucose were involved in. map00052: Galactose metabolism; map00040: Pentose and glucuronate interconversions; map00053: Ascorbate and aldarate metabolism; map01060: Biosynthesis of plant secondary metabolites; map00240: Pyrimidine metabolism; map00500: Starch and sucrose metabolism; map00524: Neomycin, kanamycin and gentamicin biosynthesis; map00520: Amino sugar and nucleotide sugar metabolism; map00561: Glycerolipid metabolism; SQD1: UDP-sulfoquinovose synthase.
Plants constitutively synthesize a wide variety of secondary metabolites to aid fitness by preventing pathogen invasion [53]. Among these metabolites, terpenoids are frequently described as natural products that are active against pathogens [54]. Neomenthol dehydrogenase (MNR, PAU012048.1), which participates in the monoterpene synthesis of neomenthol and isomenthol, is a monoterpenoid dehydrogenase [55]. In Capsicum annuum, the menthone reductase gene CaMNR1 regulates plant defenses against a broad spectrum of pathogens [56]. In our study, MNR was the target of a PaWB-related lncRNA (TCONS_00026472), and was upregulated in PT/PTI. The change in MNR expression may be in response to PaWB-phytoplasmas infection.
DNA methylation plays an essential role in regulating plant development [57]. DNA methyltransferase (Dnmt), an important enzyme for DNA methylation, is not only associated with DNA methylation, but is also linked to many important biological activities, including cell proliferation and senescence [58]. In previous studies, we found that PaWB may be related to changes in DNA methylation [59,60]. In this study, a PaWB-related gene, DNA (cytosine-5-)-methyltransferase (PAU029256.2, Dnmt1), was identified as the target gene of a PaWB-related lncRNA (TCONS_00032704), and was upregulated in PT/PTI. After infection, the change in Dnmt1 expression might influence DNA methylation and result in the phenotype of infected Paulownia.
5. Conclusions
Although the functions of most lncRNAs are still unknown, the evidence suggests that lncRNAs may play regulatory roles by interacting with RNA, DNA, and protein-coding genes. For example, it has been reported that lncRNAs can act as cis- or trans-regulators of protein-coding gene expression in animals and plants. The lncRNAs detected in this study will lay the groundwork for future functional studies of lncRNAs in Paulownia. We revealed 26,155 and 27,017 genes, and 3434 and 3504 lncRNAs in healthy and PaWB-infected P. tomentosa, respectively. A total of 6746 lncRNA–mRNA pairs were identified (4175 were cis-regulated and 2571 were trans-regulated). In PT/PTI, 3744 DEGs (2208 upregulated and 1536 downregulated) and 750 (200 upregulated and 550 downregulated) were identified. According to our analysis, 1063 mRNAs and 110 lncRNAs were related to PaWB, in which 12 mRNAs were candidate target genes of nine lncRNAs. To understand the specific biological role of lncRNAs and their regulatory mechanisms in Paulownia, future research should include functional analyses of the lncRNA candidate target genes using overexpression, CRISPR-Cas9, or RNA interference gene silencing strategies.
Supplementary Materials
The following are available online at www.mdpi.com/1999-4907/8/9/348/s1, Figure S1: Detection of phytoplasma in Paulownia seedings. 1: PTI, 2: PTI -100, 3: PT, 4: PT-100, 5: ddH2O, D: DNA maker. Figure S2: A Venn diagram showing mRNAs (a) and lncRNAs (b) that are commonly expressed in PT, PTI, PT-100, and PTI-100 samples as well as those specifically expressed under one but not the other. Figure S3: Details of the comparison schemes. (a) PaWB-related mRNAs; (b) PaWB-related lncRNAs. Figure S4: Classification of lncRNAs, (a) all-lncRNAs; (b) PaWB-related lncRNAs. class code ‘u’: intergenic lncRNAs, class code ‘o’: lncRNAs that had exonic overlap with a known transcript, class code ‘x’: lncRNAs that had exonic overlap with reference on the opposite strand, class code ‘j’: novel long noncoding isoforms with at least one splice junction shared with the reference genes; Table S1: Primer sequence used in this study. Table S2: The revealed mRNAs in this study. Table S3: The revealed lncRNAs in this study. Table S4: Classification of predicted lncRNAs into different ncRNA families. Table S5: The conservation of Paulownia lncRNAs. Table S6: The identified target genes in this study.
Acknowledgments
The authors wish to acknowledge the support received from Distinguished Talents Foundation of Henan Province of China (174200510001) and the Natural Science Foundation of Henan Province of China (162300410158).
Author Contributions
Z.W. and G.F. conceived and designed the experiments; Z.W. performed the experiments; Y.C. and Y.D. analyzed the data; X.Z. contributed reagents/materials/analysis tools; Z.W. wrote the paper.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Fan, G.; Zhai, X.; Qin, H.; Yang, X.; Dong, Z.; Jiang, J. Study on In Vitro plantlet regeneration of paulownia plant with witches’ broom. J. Henan Agric. Univ. 2005, 39, 254–258. [Google Scholar]
- Hogenhout, S.A.; Oshima, K.; Ammar el, D.; Kakizawa, S.; Kingdom, H.N.; Namba, S. Phytoplasmas: Bacteria that manipulate plants and insects. Mol. Plant Pathol. 2008, 9, 403–423. [Google Scholar] [CrossRef] [PubMed]
- Fan, G.; Dong, Y.; Deng, M.; Zhao, Z.; Niu, S.; Xu, E. Plant-pathogen interaction, circadian rhythm, and hormone-related gene expression provide indicators of phytoplasma infection in Paulownia fortunei. Int. J. Mol. Sci. 2014, 15, 23141–23162. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Liu, W.; Fan, G.; Zhai, X.; Zhao, Z.; Dong, Y.; Deng, M.; Cao, Y. Quantitative proteome-level analysis of paulownia witches’ broom disease with methyl methane sulfonate assistance reveals diverse metabolic changes during the infection and recovery processes. PeerJ 2017, 5, e3495. [Google Scholar] [CrossRef] [PubMed]
- Ipekci, Z.; Gozukirmizi, N. Direct somatic embryogenesis and synthetic seed production from paulownia elongata. Plant Cell Rep. 2003, 22, 16–24. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Fan, G.; Dong, Y.; Zhai, X.; Deng, M.; Zhao, Z.; Liu, W.; Cao, Y. Implications of polyploidy events on the phenotype, microstructure, and proteome of paulownia australis. PLoS ONE 2017, 12, e0172633. [Google Scholar] [CrossRef] [PubMed]
- Ju, G.; Wang, R.; Zhou, Y.; Dajin, R. Study on the relationship between the content of vitamin C and resistance of paulownia spp. To witches’ broom. For. Res. 1996, 9, 421–424. [Google Scholar]
- Wang, R.; Wang, S.; Sun, X. Effects of hormones on the morphogenesis of witches broom of Paulownia spp. Sci. Silvae Sin. 1981, 17, 281–286. [Google Scholar]
- Yue, H.; Wu, K.; Wu, Y.; Zhang, J.; Sun, R. Cloning and Characterization of Three Subunits of the Phytoplasma Sec Protein Translocation System Associated with the Paulownia Witches’-Broom. Plant Prot. 2009, 35, 25–31. [Google Scholar]
- Wang, J.; Zhu, X.; Gao, R.; Lin, C.; Li, Y.; Xu, Q.; Piao, C.; Li, X.; Li, H.; Tian, G. Genetic and serological analyses of elongation factor EF-TU of paulownia witches’-broom phytoplasma (16sri-d). Plant Pathol. 2010, 59, 972–981. [Google Scholar] [CrossRef]
- Lin, C.; Zhou, T.; Li, H.; Fan, Z.; Li, Y.; Piao, C.; Tian, G. Molecular characterisation of two plasmids from paulownia witches’-broom phytoplasma and detection of a plasmid-encoded protein in infected plants. Eur. J. Plant Pathol. 2008, 123, 321–330. [Google Scholar] [CrossRef]
- Du, T.; Wang, Y.; Hu, Q.; Chen, J.; Liu, S.; Huang, W.; Lin, M. Transgenic paulownia expressing shiva-1 gene has increased resistance to paulownia witches’ broom disease. J. Integr. Plant Biol. 2005, 47, 1500–1506. [Google Scholar] [CrossRef]
- Fan, G.; Zhang, B.; Zhai, X.; Liu, F.; Ma, Y.B.; Kan, S. Effects of rifampin on the changes of morphology and plant endogenous hormones of paulownia seedlings with witches’ broom. J. Henan Agric. Univ. 2007, 41, 387–391. [Google Scholar]
- Fan, G.; Cao, X.; Niu, S.; Deng, M.; Zhao, Z.; Dong, Y. Transcriptome, microrna, and degradome analyses of the gene expression of paulownia with phytoplamsa. BMC Genom. 2015, 16, 896. [Google Scholar] [CrossRef] [PubMed]
- Fan, G.; Cao, X.; Zhao, Z.; Deng, M. Transcriptome analysis of the genes related to the morphological changes of paulownia tomentosa plantlets infected with phytoplasma. Acta Physiol. Plant. 2015, 37, 1–12. [Google Scholar] [CrossRef]
- Fan, G.; Xu, E.; Deng, M.; Zhao, Z.; Niu, S. Phenylpropanoid metabolism, hormone biosynthesis and signal transduction-related genes play crucial roles in the resistance of paulownia fortunei to paulownia witches’ broom phytoplasma infection. Genes Genom. 2015, 37, 913–929. [Google Scholar] [CrossRef]
- Liu, R.; Dong, Y.; Fan, G.; Zhao, Z.; Deng, M.; Cao, X.; Niu, S. Discovery of genes related to witches broom disease in paulownia tomentosa x paulownia fortunei by a de novo assembled transcriptome. PLoS ONE 2013, 8, e80238. [Google Scholar] [CrossRef] [PubMed]
- Mou, H.Q.; Lu, J.; Zhu, S.F.; Lin, C.L.; Tian, G.Z.; Xu, X.; Zhao, W.J. Transcriptomic analysis of paulownia infected by paulownia witches’-broom phytoplasma. PLoS ONE 2013, 8, e77217. [Google Scholar] [CrossRef] [PubMed]
- Fan, G.; Cao, Y.; Deng, M.; Zhai, X.; Zhao, Z.; Niu, S.; Ren, Y. Identification and dynamic expression profiling of micrornas and target genes of paulownia tomentosa in response to paulownia witches’ broom disease. Acta Physiol. Plant. 2016, 39, 28. [Google Scholar] [CrossRef]
- Fan, G.; Niu, S.; Xu, T.; Deng, M.; Zhao, Z.; Wang, Y.; Cao, L.; Wang, Z. Plant-pathogen interaction-related micrornas and their targets provide indicators of phytoplasma infection in paulownia tomentosa x paulownia fortunei. PLoS ONE 2015, 10, e0140590. [Google Scholar] [CrossRef] [PubMed]
- Fan, G.; Niu, S.; Zhao, Z.; Deng, M.; Xu, E.; Wang, Y.; Yang, L. Identification of micrornas and their targets in paulownia fortunei plants free from phytoplasma pathogen after methyl methane sulfonate treatment. Biochimie 2016, 127, 271–280. [Google Scholar] [CrossRef] [PubMed]
- Niu, S.; Fan, G.; Deng, M.; Zhao, Z.; Xu, E.; Cao, L. Discovery of micrornas and transcript targets related to witches’ broom disease in paulownia fortunei by high-throughput sequencing and degradome approach. Mol. Genet. Genom. 2016, 291, 181–191. [Google Scholar] [CrossRef] [PubMed]
- Cao, X.; Fan, G.; Dong, Y.; Zhao, Z.; Deng, M.; Wang, Z.; Liu, W. Proteome profiling of paulownia seedlings infected with phytoplasma. Front. Plant Sci. 2017, 8, 342. [Google Scholar] [CrossRef] [PubMed]
- Rinn, J.L.; Chang, H.Y. Genome regulation by long noncoding RNAs. Annu. Rev. Biochem. 2012, 81, 145–166. [Google Scholar] [CrossRef] [PubMed]
- Ben Amor, B.; Wirth, S.; Merchan, F.; Laporte, P.; d’Aubenton-Carafa, Y.; Hirsch, J.; Maizel, A.; Mallory, A.; Lucas, A.; Deragon, J.M.; et al. Novel long non-protein coding RNAs involved in arabidopsis differentiation and stress responses. Genome Res. 2009, 19, 57–69. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Jung, C.; Xu, J.; Wang, H.; Deng, S.; Bernad, L.; Arenas-Huertero, C.; Chua, N.H. Genome-wide analysis uncovers regulation of long intergenic noncoding RNAs in arabidopsis. Plant Cell 2012, 24, 4333–4345. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Chung, P.J.; Liu, J.; Jang, I.C.; Kean, M.J.; Xu, J.; Chua, N.H. Genome-wide identification of long noncoding natural antisense transcripts and their responses to light in arabidopsis. Genome Res. 2014, 24, 444–453. [Google Scholar] [CrossRef] [PubMed]
- Ding, J.; Lu, Q.; Ouyang, Y.; Mao, H.; Zhang, P.; Yao, J.; Xu, C.; Li, X.; Xiao, J.; Zhang, Q. A long noncoding rna regulates photoperiod-sensitive male sterility, an essential component of hybrid rice. Proc. Natl. Acad. Sci. USA 2012, 109, 2654–2659. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.C.; Liao, J.Y.; Li, Z.Y.; Yu, Y.; Zhang, J.P.; Li, Q.F.; Qu, L.H.; Shu, W.S.; Chen, Y.Q. Genome-wide screening and functional analysis identify a large number of long noncoding RNAs involved in the sexual reproduction of rice. Genome Biol. 2014, 15, 512. [Google Scholar] [CrossRef] [PubMed]
- Xin, M.; Wang, Y.; Yao, Y.; Song, N.; Hu, Z.; Qin, D.; Xie, C.; Peng, H.; Ni, Z.; Sun, Q. Identification and characterization of wheat long non-protein coding RNAs responsive to powdery mildew infection and heat stress by using microarray analysis and SBS sequencing. BMC Plant Biol. 2011, 11, 61. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Chen, X.; Wang, C.; Xu, Z.; Wang, Y.; Liu, X.; Kang, Z.; Ji, W. Long non-coding genes implicated in response to stripe rust pathogen stress in wheat (Triticum aestivum L.). Mol. Biol. Rep. 2013, 40, 6245–6253. [Google Scholar] [CrossRef] [PubMed]
- Fan, C.; Hao, Z.; Yan, J.; Li, G. Genome-wide identification and functional analysis of lincrnas acting as mirna targets or decoys in maize. BMC Genom. 2015, 16, 793. [Google Scholar] [CrossRef] [PubMed]
- Lu, X.; Chen, X.; Mu, M.; Wang, J.; Wang, X.; Wang, D.; Yin, Z.; Fan, W.; Wang, S.; Guo, L.; et al. Genome-wide analysis of long noncoding RNAs and their responses to drought stress in cotton (Gossypium hirsutum L.). PLoS ONE 2016, 11, e0156723. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Yuan, D.; Tu, L.; Gao, W.; He, Y.; Hu, H.; Wang, P.; Liu, N.; Lindsey, K.; Zhang, X. Long noncoding RNAs and their proposed functions in fibre development of cotton (Gossypium spp.). New Phytol. 2015, 207, 1181–1197. [Google Scholar] [CrossRef] [PubMed]
- Joshi, R.K.; Megha, S.; Basu, U.; Rahman, M.H.; Kav, N.N. Genome wide identification and functional prediction of long non-coding RNAs responsive to sclerotinia sclerotiorum infection in Brassica napus. PLoS ONE 2016, 11, e0158784. [Google Scholar] [CrossRef] [PubMed]
- Tian, J.; Song, Y.; Du, Q.; Yang, X.; Ci, D.; Chen, J.; Xie, J.; Li, B.; Zhang, D. Population genomic analysis of gibberellin-responsive long non-coding RNAs in populus. J. Exp. Bot. 2016, 67, 2467–2482. [Google Scholar] [CrossRef] [PubMed]
- Shuai, P.; Liang, D.; Tang, S.; Zhang, Z.; Ye, C.Y.; Su, Y.; Xia, X.; Yin, W. Genome-wide identification and functional prediction of novel and drought-responsive lincrnas in populus trichocarpa. J. Exp. Bot. 2014, 65, 4975–4983. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Q.H.; Stephen, S.; Taylor, J.; Helliwell, C.A.; Wang, M.B. Long noncoding RNAs responsive to fusarium oxysporum infection in Arabidopsis thaliana. New Phytol. 2014, 201, 574–584. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Yu, W.; Yang, Y.; Li, X.; Chen, T.; Liu, T.; Ma, N.; Yang, X.; Liu, R.; Zhang, B. Genome-wide analysis of tomato long non-coding RNAs and identification as endogenous target mimic for microrna in response to tylcv infection. Sci. Rep. 2015, 5, 16946. [Google Scholar] [CrossRef] [PubMed]
- Cui, J.; Luan, Y.; Jiang, N.; Bao, H.; Meng, J. Comparative transcriptome analysis between resistant and susceptible tomato allows the identification of lncrna16397 conferring resistance to phytophthora infestans by co-expressing glutaredoxin. Plant J. 2017, 89, 577–589. [Google Scholar] [CrossRef] [PubMed]
- Trapnell, C.; Pachter, L.; Salzberg, S.L. Tophat: Discovering splice junctions with RNA-Seq. Bioinformatics 2009, 25, 1105–1111. [Google Scholar] [CrossRef] [PubMed]
- Trapnell, C.; Roberts, A.; Goff, L.; Pertea, G.; Kim, D.; Kelley, D.R.; Pimentel, H.; Salzberg, S.L.; Rinn, J.L.; Pachter, L. Differential gene and transcript expression analysis of RNA-seq experiments with tophat and cufflinks. Nat. Protoc. 2012, 7, 562–578. [Google Scholar] [CrossRef] [PubMed]
- Sun, L.; Zhang, Z.; Bailey, T.L.; Perkins, A.C.; Tallack, M.R.; Xu, Z.; Liu, H. Prediction of novel long non-coding RNAs based on RNA-Seq data of mouse klf1 knockout study. BMC Bioinform. 2012, 13, 331. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Wang, Y.; Chen, M.; Xiao, P.; Hu, C.; Zeng, Z.; Wang, C.; Wang, J.; Hu, Z. Genome-wide long non-coding RNA screening, identification and characterization in a model microorganism chlamydomonas reinhardtii. Sci. Rep. 2016, 6, 34109. [Google Scholar] [CrossRef] [PubMed]
- Nawrocki, E.P.; Kolbe, D.L.; Eddy, S.R. Infernal 1.0: Inference of rna alignments. Bioinformatics 2009, 25, 1335–1337. [Google Scholar] [CrossRef] [PubMed]
- Jia, H.; Osak, M.; Bogu, G.K.; Stanton, L.W.; Johnson, R.; Lipovich, L. Genome-wide computational identification and manual annotation of human long noncoding RNA genes. RNA 2010, 16, 1478–1487. [Google Scholar] [CrossRef] [PubMed]
- Tafer, H.; Hofacker, I.L. Rnaplex: A fast tool for RNA-RNA interaction search. Bioinformatics 2008, 24, 2657–2663. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-delta delta c(t)) method. Methods (San Diego, Calif.) 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Van Sandt, V.S.; Guisez, Y.; Verbelen, J.P.; Vissenberg, K. Analysis of a xyloglucan endotransglycosylase/hydrolase (xth) from the lycopodiophyte selaginella kraussiana suggests that xth sequence characteristics and function are highly conserved during the evolution of vascular plants. J. Exp. Bot. 2006, 57, 2909–2922. [Google Scholar] [CrossRef] [PubMed]
- Osato, Y.; Yokoyama, R.; Nishitani, K. A principal role for atxth18 in Arabidopsis thaliana root growth: A functional analysis using rnai plants. J. Plant Res. 2006, 119, 153–162. [Google Scholar] [CrossRef] [PubMed]
- Thompson, J.E.; Fry, S.C. Restructuring of wall-bound xyloglucan by transglycosylation in living plant cells. Plant J. 2001, 26, 23–34. [Google Scholar] [CrossRef] [PubMed]
- Shimojima, M.; Hoffmann-Benning, S.; Garavito, R.M.; Benning, C. Ferredoxin-dependent glutamate synthase moonlights in plant sulfolipid biosynthesis by forming a complex with SQD1. Arch. Biochem. Biophys. 2005, 436, 206–214. [Google Scholar] [CrossRef] [PubMed]
- Kliebenstein, D.J. Secondary metabolites and plant/environment interactions: A view through Arabidopsis thaliana tinged glasses. Plant Cell Environ. 2004, 27, 675–684. [Google Scholar] [CrossRef]
- Litvak, M.E.; Monson, R.K. Patterns of induced and constitutive monoterpene production in conifer needles in relation to insect herbivory. Oecologia 1998, 114, 531–540. [Google Scholar] [CrossRef] [PubMed]
- Ringer, K.L.; McConkey, M.E.; Davis, E.M.; Rushing, G.W.; Croteau, R. Monoterpene double-bond reductases of the (−)-menthol biosynthetic pathway: Isolation and characterization of cdnas encoding (−)-isopiperitenone reductase and (+)-pulegone reductase of peppermint. Arch. Biochem. Biophys. 2003, 418, 80–92. [Google Scholar] [CrossRef]
- Choi, H.W.; Lee, B.G.; Kim, N.H.; Park, Y.; Lim, C.W.; Song, H.K.; Hwang, B.K. A role for a menthone reductase in resistance against microbial pathogens in plants. Plant Physiol. 2008, 148, 383–401. [Google Scholar] [CrossRef] [PubMed]
- Finnegan, E.J.; Peacock, W.J.; Dennis, E.S. DNA methylation, a key regulator of plant development and other processes. Curr. Opin. Genet. Dev. 2000, 10, 217–223. [Google Scholar] [CrossRef]
- Bestor, T.H.; Verdine, G.L. DNA methyltransferases. Curr. Opin. Cell Biol. 1994, 6, 380–389. [Google Scholar] [CrossRef]
- Cao, X.; Fan, G.; Deng, M.; Zhao, Z.; Dong, Y. Identification of genes related to paulownia witches’ broom by AFLP and MSAP. Int. J. Mol. Sci. 2014, 15, 14669–14683. [Google Scholar] [CrossRef] [PubMed]
- Cao, X.; Fan, G.; Zhao, Z.; Deng, M.; Dong, Y. Morphological changes of paulownia seedlings infected phytoplasmas reveal the genes associated with witches’ broom through aflp and msap. PLoS ONE 2014, 9, e112533. [Google Scholar] [CrossRef] [PubMed]
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).