Male Parent Identification of Triploid Rubber Trees (Hevea brasiliensis) and the Mechanism of 2n Gametes Formation
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Materials
2.2. Collection of Seeds and Sowing
2.3. Ploidy Analysis by Flow Cytometry
2.4. Chromosome Counting
2.5. DNA Extraction and SSR Analysis
3. Results
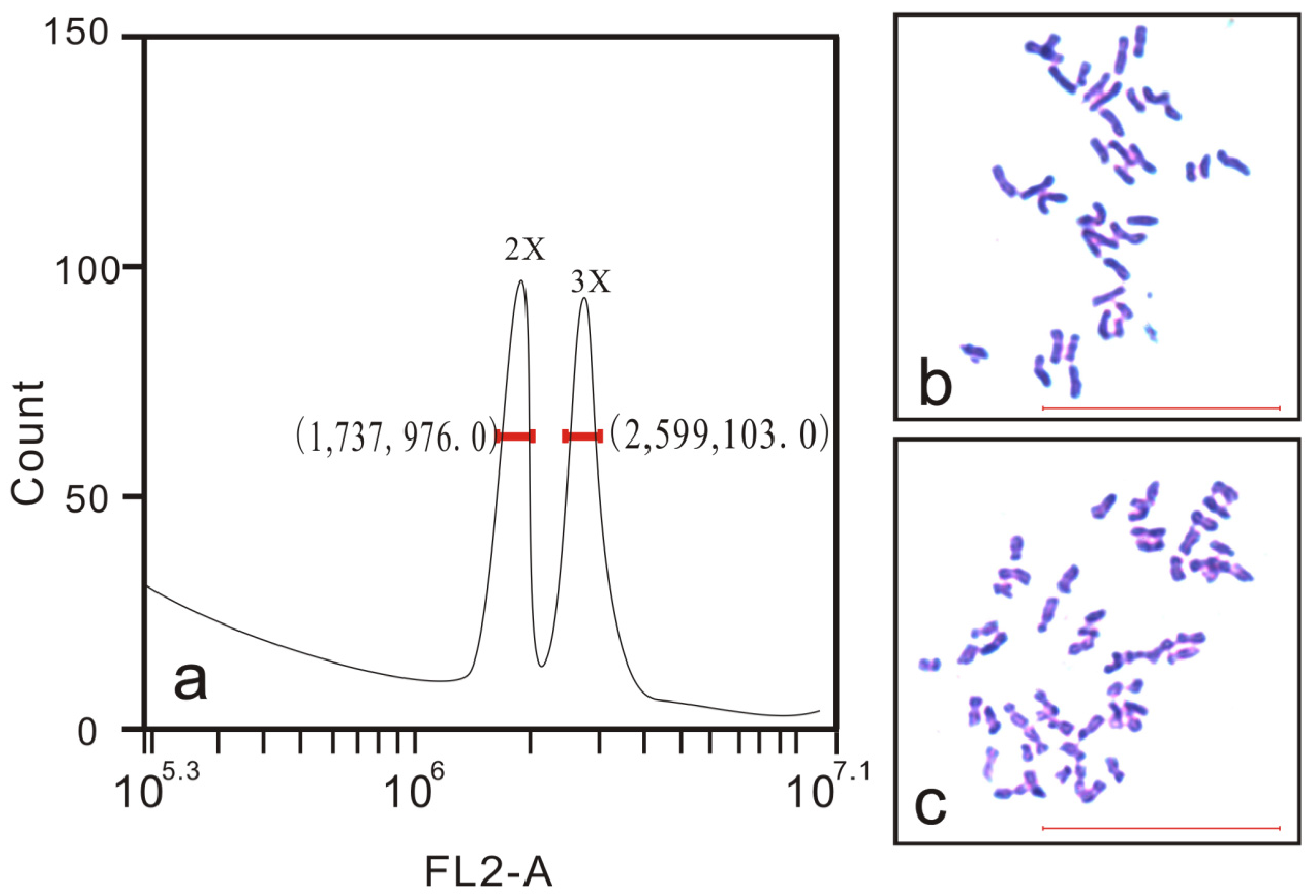
3.1. Determination of Hybrid Ploidy Levels
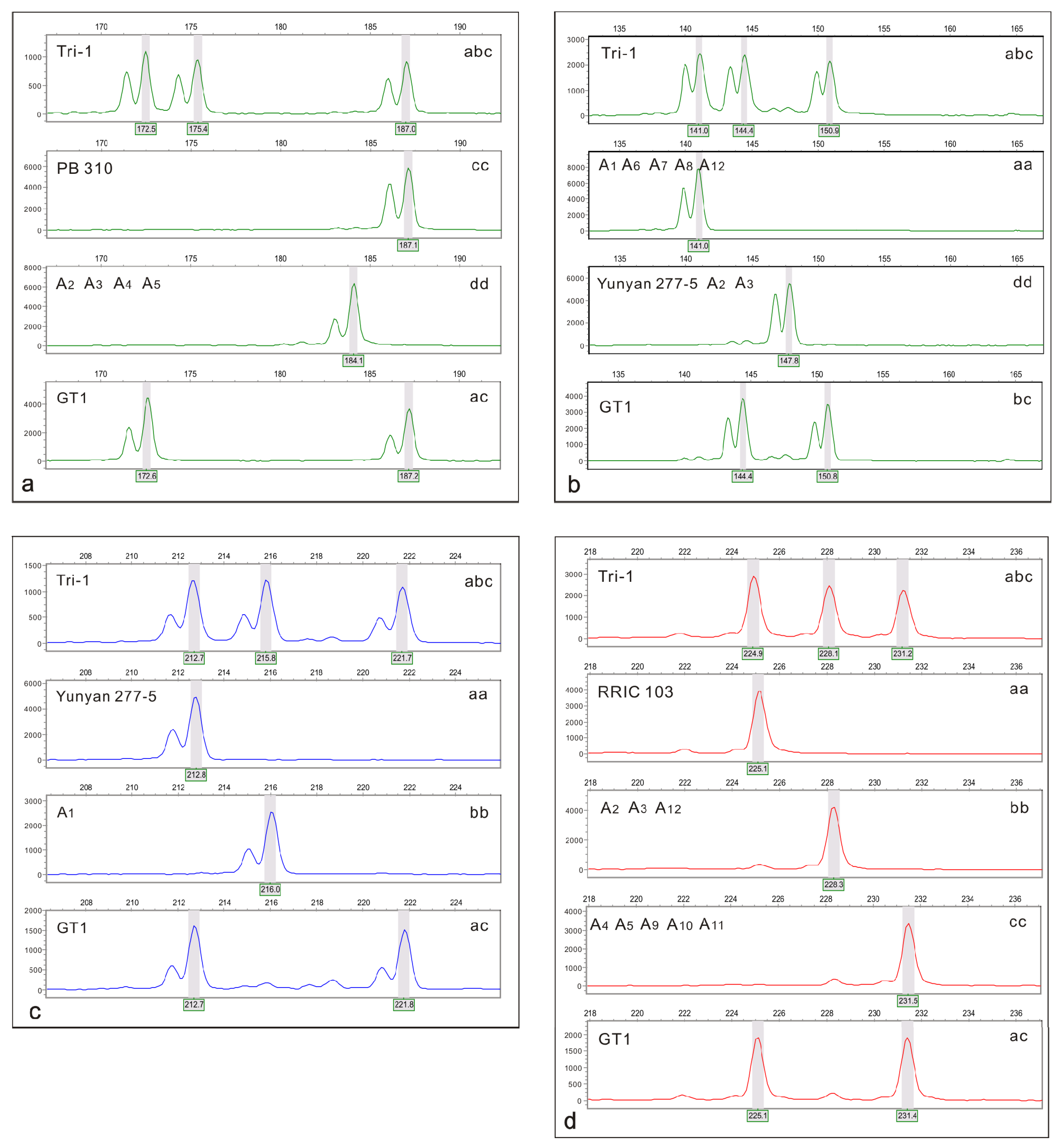
3.2. Screen of SSR Markers
3.3. Origin of 2n Gametes
3.4. Male Parent Identification of Triploids
3.5. Mechanism of 2n Gamete Formation
4. Discussion
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Lespinasse, D.; Rodier-Goud, M.; Grivet, L.; Leconte, A.; Legnate, H.; Seguin, M. A saturated genetic linkage map of rubber tree (Hevea spp.) based on RFLP, AFLP, microsatellite, and isozyme markers. Theor. Appl. Genet. 2000, 100, 127–138. [Google Scholar] [CrossRef]
- Li, X.; Chen, Q.; Xu, B.; Ren, X.; Huang, H.; Li, Q. An Analysis of the Patents concerning Hevea brasiliensis. Asian Agric. Res. 2015, 7, 58–61. [Google Scholar]
- Rao, B.S. Pollinating of hevea in Malaya. J. Rubber Res. Inst. Malaya 1961, 17, 14–18. [Google Scholar]
- Hamzah, S.; Chan, J.L.; Yeang, H.Y. Pollen tube growth and fruit-set success in Hevea brasiliensis hand-pollination influenced by the choice of clone and female flower. Euphytica 2002, 123, 1–8. [Google Scholar] [CrossRef]
- Yu, F.; Wang, B.-H.; Feng, S.-P.; Wang, J.-Y.; Li, W.-G.; Wu, Y.-T. Development, characterization, and cross-species/genera transferability of SSR markers for rubber tree (Hevea brasiliensis). Plant Cell Rep. 2011, 30, 335–344. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Fox, J.M. Mapping rubber tree growth in mainland Southeast Asia using time-series MODIS 250 m NDVI and statistical data. Appl. Geogr. 2012, 32, 420–432. [Google Scholar] [CrossRef]
- Raj, S.; Das, G.; Pothen, J.; Dey, S.K. Relationship between latex yield of Hevea brasiliensis and antecedent environmental paremeters. Int. J. Biometeorol. 2005, 49, 189–196. [Google Scholar] [CrossRef] [PubMed]
- Cornish, K. Similarities and differences in rubber biochemistry among plant species. Phytochemistry 2001, 57, 1123–1134. [Google Scholar] [CrossRef]
- Ahrends, A.; Hollingsworth, P.M.; Ziegler, A.D.; Fox, J.M.; Chen, H.; Su, Y.; Xu, J. Current trends of rubber plantation expansion may threaten biodiversity and livelihoods. Glob. Environ. Chang. 2015, 34, 48–58. [Google Scholar] [CrossRef]
- Priyadarshan, P.M.; Hoa, T.T.T.; Huasun, H.; Gonçalves, P.S. Yielding potential of rubber (Hevea brasiliensis) in sub-optimal environments. J. Crop Improv. 2005, 14, 221–247. [Google Scholar] [CrossRef]
- Li, H.B.; Zhou, T.Y.; Ning, L.Y.; Li, G.H. Cytological Identification and Breeding Course of Hevea ‘Yunyan 77–2’ and ‘Yunyan 77–4’. J. Trop. Subtrop. Bot. 2009, 17, 602–605. [Google Scholar]
- Ao, S.C.; He, L.G.; Xiao, G.X.; Chen, J.B.; He, C.G. High yield and cold resistance Hevea. of Yunyan 77–2 and Yunyan 77–4 Selection and Breeding. J. Yunnan Trop. Crops 1998, 21, 3–8. [Google Scholar]
- Yang, L.Q. Forepart study on new varieties Yunyan 77–2 and Yunyan 77–4 of Hevea brasiliensis. J. Yunnan Trop. Crops 2002, 25, 1–4. [Google Scholar]
- Feng, S.P.; Li, W.G.; Huang, H.S.; Wang, J.Y.; Wu, C.T. Development, characterization and cross-species/genera transferability of EST-SSR markers for rubber tree (Hevea brasiliensis). Mol. Breed. 2009, 23, 87–97. [Google Scholar]
- Carputo, D.; Barone, A.; Frusciante, L. 2n gametes in the potato: Essential ingredients for breeding and germplasm transfer. Theor. Appl. Genet. 2000, 101, 805–813. [Google Scholar] [CrossRef]
- Lopez-Lavalle, L.A.B.; Orjeda, G. Occurrence and cytological mechanism of 2n pollen formation in a tetraploid accession of Ipomoea batatas (sweet potato). J. Hered. 2002, 93, 185–192. [Google Scholar] [CrossRef]
- Zhang, X.-Z.; Liu, G.-J.; Yan, L.-Y.; Zhao, Y.-B.; Chang, R.-F.; Wu, L.-P. Creating triploid germplasm via induced 2n pollen in Capsicum annuum L. J. Hortic. Sci. Biotech. 2002, 78, 84–88. [Google Scholar]
- Przybyla, A.; Behrend, A.; Bornhake, C.; Hohe, A. Breeding of polyploid heather (Calluna vulgaris). Euphytica 2014, 199, 273–282. [Google Scholar] [CrossRef]
- Veilleux, R. Diploid and polyploidy gametes in crop pants: Mechanisms of formation and utilization in plant breeding. Plant Breed. Rev. 1985, 3, 253–288. [Google Scholar]
- Bretagnolle, F.; Thompson, J.D. Gametes with the somatic chromosome number: Mechanisms of their formation and role in the evolution of autopolyploid pants. New Phytol. 1995, 129, 1–22. [Google Scholar] [CrossRef]
- Lim, K.-B.; Ramanna, M.S.; de Jong, J.H.; Jacobsen, E.; van Tuyl, J.M. Indeterminate meiotic restitution (IMR): A novel type of meiotic nuclear restitution mechanism detected in interspecific lily hybrids by GISH. Theor. Appl. Genet. 2001, 103, 219–230. [Google Scholar] [CrossRef]
- Mok, D.W.S.; Peloquin, S.J. The inheritance of three mechanisms of diplandroid (2n pollen) formation in diploid potatoes. Heredity 1975, 35, 295–302. [Google Scholar] [CrossRef]
- Hermsen, J.G. Mechanisms and genetic implications of 2n-gamete formation. Iowa State J. Res. 1984, 58, 421–434. [Google Scholar]
- Douches, D.S.; Maas, D.L. Comparison of FDR- and SDR-derived tetraploid progeny from 2x × 4x crosses using haploids of Solanum tuberosum L. that produce mixed modes of 2n eggs. Theor. Appl. Genet. 1998, 97, 1307–1313. [Google Scholar] [CrossRef]
- Carputo, D.; Frusciante, L.; Peloquin, S.J. The role of 2n gametes and endosperm balance number in the origin and evolution of polyploids in the tuber-bearing Solanums. Genetics 2003, 163, 287–294. [Google Scholar] [PubMed]
- Chow, K.S.; Wan, K.L.; Isa, M.N.M.; Bahari, A.; Tan, S.H.; Harikrishna, K.; Yeang, H.Y. Insights into rubber biosynthesis from transcriptome analysis of Hevea brasiliensis latex. J. Exp. Bot. 2007, 58, 2429–2440. [Google Scholar] [CrossRef] [PubMed]
- Souza, L.M.; Mantello, C.C.; Santos, M.O.; de Souza, G.P.; Souza, A.P. Microsatellites from rubber tree (Hevea brasiliensis) for genetic diversity analysis and cross-amplification in six Hevea wild species. Conserv. Genet. Resour. 2009, 1, 75–79. [Google Scholar] [CrossRef]
- Triwitayakorn, T.; Chatkulkawin, P.; Kanjanawattanawong, S.; Sraphet, S.; Yoocha, T.; Sangsrakru, D.; Chanprasert, J.; Ngamphiw, C.; Jomchai, N.; Therawattanasuk, K.; et al. Transcriptome sequencing of Hevea brasiliensis for development of microsatellite markers and construction of a genetic linkage map. DNA Res. 2011, 18, 471–482. [Google Scholar] [CrossRef] [PubMed]
- Schuelke, M. An economic method for the fluorescent labeling of PCR fragments. Nat. Biotechnol. 2000, 18, 233–234. [Google Scholar] [CrossRef] [PubMed]
- Esselink, G.D.; Smulders, M.J.M.; Vosman, B. Identification of cut rose (Rosa hybrida) and rootstock varieties using robust sequence tagged microsatellite site markers. Theor. Appl. Genet. 2003, 106, 277–286. [Google Scholar] [CrossRef] [PubMed]
- Nybom, H.; Esselink, G.D.; Werlemark, G.; Vosman, B. Microsatellite DNA marker inheritance indicates preferential paring between two highly homologous genomes in polyploid and hemisexual dog-roses, Rosa L. Sect. Caninae DC. Heredity 2004, 92, 139–150. [Google Scholar] [CrossRef] [PubMed]
- Xie, K.-D.; Wang, X.-P.; Biswas, M.K.; Liang, W.-J.; Xu, Q.; Grosser, J.W.; Guo, W.-W. 2n megagametophyte formed via SDR contributes to tetraploidization in polyembryonic ‘Nadorcott’ tangor crossed by citrus allotetraploids. Plant Cell Rep. 2014, 33, 1641–1650. [Google Scholar]
- Raboin, L.-M.; Carreel, F.; Noyer, J.-L.; Baurens, F.-C.; Horry, J.-P.; Bakry, F.; Du Montcel, H.T.; Ganry, J.; Lanaud, C.; Lagoda, P.J.L. Diploid ancestors of triploid export banana cultivars: Molecular identification of 2n restitution gamete donors and n gamete donors. Mol. Breed. 2005, 16, 333–341. [Google Scholar]
- Chen, C.; Lyon, M.T.; O’Malley, D.; Federici, C.T.; Gmitter, J.; Grosser, J.W.; Chaparro, J.X.; Roose, M.L.; Gmitter, F.G., Jr. Origin and frequency of 2n gametes in Citrus sinensis × Poncirustrifoliata and their reciprocal crosses. Plant Sci. 2008, 174, 1–8. [Google Scholar] [CrossRef]
- Xu, M.; Sun, Y.; Li, H. EST-SSRs development and paternity analysis for Liriodendron spp. New For. 2010, 40, 361–382. [Google Scholar] [CrossRef]
- Xavier, M.A.; Pinto, L.R.; Fávero, T.M.; Perecin, D.; Carlini-Garcia, L.A.; Landell, M.G.A. Paternity identification in sugarcane polycrosses by using microsatellite markers. Genet. Mol. Res. 2014, 13, 2268–2277. [Google Scholar] [CrossRef] [PubMed]
- Park, T.-H.; Kim, J.-B.; Hutten, R.C.B.; van Eck, H.J.; Jacobsen, E.; Visser, R.G.F. Genetic positioning of centromeres using half-tetrad analysis in a 4x-2x cross population of potato. Genetics 2007, 176, 85–94. [Google Scholar]
- Cuenca, J.; Froelicher, Y.; Aleza, P.; Juárez, J.; Navarro, L.; Ollitrault, P. Multilocus half-tetrad analysis and centromere mapping in citrus: Evidence of SDR mechanism for 2n megagametophyte production and partial chiasma interference in mandarin cv ‘Fortune’. Heredity 2011, 107, 462–470. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Li, D.-L.; Kang, X.-Y. Induction of unreduced megaspores with high temperature during megasporogenesis in Populus. Ann. For. Sci. 2012, 69, 59–67. [Google Scholar] [CrossRef]
- Lu, M.; Zhang, P.; Kang, X. Induction of 2n female gametes in Populus adenopoda Maxim by high temperature exposure during female gametophyte development. Breed. Sci. 2013, 63, 96–103. [Google Scholar] [CrossRef] [PubMed]
- De Storme, N.; Geelen, D. Sexual polyploidization in plants–cytological mechanisms and molecular regulation. New Phytol. 2013, 198, 670–684. [Google Scholar] [CrossRef] [PubMed]
- D’Erfurth, I.; Jolivet, S.; Froger, N.; Catrice, O.; Novatchkova, M.; Simon, M.; Jenczewski, E.; Mercier, R. Mutations in AtPS1 (Arabidopsis thaliana parallel spindle 1) lead to the production of diploid pollen grains. PLoS Genet. 2008, 4, e1000274. [Google Scholar] [CrossRef] [PubMed]
- De Storme, N.; Geelen, D. The Arabidopsis mutant jason produces unreduced first division restitution male gametes through a parallel/fused spindle mechanism in meiosis II. Plant Physiol. 2011, 155, 1403–1415. [Google Scholar] [CrossRef] [PubMed]
| Rubber Trees (Number) | Pedigree/Source |
|---|---|
| PB 310 (1) | PB 5/51 × RRIM 600 |
| Yunyan 277-5 (1) | PB 5/63 × Tjir 1 |
| RRIC 103 (3) | RRIC 52 × PB 86 |
| A1-A12 (12) | Original clones |
| Marker | Sequence ID | Number of Alleles in This Study | SSR Motif | Reference |
|---|---|---|---|---|
| RUB 9 | EC608804 | 4 | (AG)n | [26] |
| RUB 16 | EC607870 | 3 | (ATC)n | [26] |
| RUB 19 | EC606684 | 8 | (GAA)n | [26] |
| RUB 20 | EC606350 | 4 | (GAT)n | [26] |
| RUB 28 | EC606163 | 7 | (CT)n | [26] |
| RUB 33 | EC605199 | 5 | (CT)n | [26] |
| RUB 65 | EC605124 | 5 | (AAG)n | [26] |
| RUB 70 | EC601354 | 6 | (GGA)n | [26] |
| RUB 74 | EC606215 | 3 | (AGA)n | [26] |
| RUB 75 | EC606169 | 4 | (GGA)n | [26] |
| RUB 90 | CB376545 | 6 | (AGA)n | Unpublished |
| RUB 95 | FJ919795 | 5 | (AT)n(GT)m | [27] |
| RUB 98 | EC609548 | 5 | (CG)n | [26] |
| RUB 102 | EC606350 | 4 | (GAT)n | [26] |
| RUB 103 | EC604892 | 5 | (CT)n | [26] |
| RUB 112 | EC604918 | 10 | (ATT)n | [26] |
| RUB 138 | EC606085 | 3 | (TTA)n | [26] |
| RUB 156 | EC600469 | 6 | (AGA)n | [26] |
| RUB 161 | FJ919800 | 4 | (CT)n | [27] |
| RUB 178 | Pr012324089 | 10 | (GCTTCT)n(CTT)m | [28] |
| RUB 179 | Pr012324091 | 4 | (TCT)n | [28] |
| RUB 183 | Pr012324095 | 7 | (CAA)n | [28] |
| RUB 184 | Pr012324096 | 4 | (AGA)n | [28] |
| RUB 190 | Pr012324102 | 8 | (TTA)n | [28] |
| RUB 199 | Pr012324114 | 4 | (TCT)n | [28] |
| Markers | Triploids | FP | MP | |||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Tri-1 | Tri-2 | Tri-3 | Tri-4 | Tri-5 | Tri-6 | Tri-7 | Tri-8 | GT1 | PB310 | Yunyan 277-5 | RRIC 103 | A1 | A2 | A3 | A4 | A5 | A6 | A7 | A8 | A9 | A10 | A11 | A12 | |
| RUB9 | bbc | bbc | bbc | bbc | abc | abc | bbc | bcc | bc | cc | cc | bb | aa | ab | ab | ab | ab | ad | ad | ad | aa | aa | aa | bb |
| RUB 16 | abc | abc | abc | abc | aab | abc | bcc | abc | ac | cc | ab | bc | bb | bb | bb | bc | bc | bc | bc | bc | bb | bb | bb | ab |
| RUB 19 | abd | bbc | bbf | bbf | aae | abf | bbf | abb | ab | ab | ab | bf | ef | ff | ff | ff | ff | fh | fh | fh | fg | fg | fg | cd |
| RUB 20 | aac | bbc | bbc | bbc | aac | abd | abb | aaa | ab | ab | aa | ab | cd | cc | cc | cc | cc | cd | cd | cd | aa | aa | aa | cc |
| RUB 28 | aab | aab | aab | bbe | abc | bbc | abb | abd | ab | ad | ad | aa | cc | ae | ae | af | af | bg | bg | bg | aa | aa | aa | aa |
| RUB 33 | aab | aab | abb | abb | aaa | aaa | abb | aaa | ab | ac | aa | ac | aa | aa | aa | ad | ad | ae | ae | ae | ad | ad | ad | ae |
| RUB 65 | abb | aab | aaa | aaa | abb | abb | aab | abb | ab | bc | ab | be | aa | aa | aa | aa | aa | aa | aa | aa | ad | ad | ad | aa |
| RUB 70 | aab | aaa | abc | aaa | bbb | aab | aac | aab | ab | ad | ab | cf | bb | ac | ac | ab | ab | bb | bb | bb | be | ee | ee | ab |
| RUB 74 | aab | aab | abc | aaa | abc | abc | aab | aab | ab | ab | bb | ab | cc | ac | ac | ac | ac | ac | ac | ac | aa | aa | aa | aa |
| RUB 75 | aab | abd | abc | aac | aab | aaa | aaa | abb | ab | bb | bb | ab | ab | ac | ac | ab | ab | ac | ac | ac | aa | aa | aa | ad |
| RUB 90 | aab | aaa | abc | aab | aab | aab | abf | aab | ab | ab | aa | bf | aa | ac | ac | ac | ac | bd | bd | bd | de | de | de | aa |
| RUB 95 | abc | abc | aab | aab | aab | aab | abc | abb | ab | ac | ac | cd | ad | aa | aa | ac | ac | ac | ac | be | dd | dd | dd | ce |
| RUB 98 | abc | aac | aab | abd | aab | aaa | aab | aab | ab | aa | ab | ab | aa | ad | ad | aa | aa | ae | ae | ae | aa | aa | aa | ac |
| RUB 102 | aaa | bbc | bbc | bbc | aac | abd | abb | aaa | ab | ab | aa | ab | cd | cc | cc | cc | cc | cd | cd | cd | aa | aa | aa | ac |
| RUB 103 | abd | abb | abd | bbd | aaa | abb | abc | aaa | ab | ac | ac | ac | aa | dd | dd | aa | aa | aa | aa | aa | ee | ee | ee | ad |
| RUB 112 | aab | aab | abb | abe | aac | abc | aab | abd | ab | fg | fg | aj | bc | be | be | ah | ah | ag | ag | ag | fi | fi | fi | aa |
| RUB 138 | bbc | bbb | abc | aac | bbc | aac | abb | aaa | ab | aa | aa | ab | ac | cc | cc | ac | ac | aa | aa | aa | ac | ac | ac | bc |
| RUB 156 | abc | aab | acd | abc | abc | abc | aac | aac | ac | ac | aa | ac | bb | bd | bd | be | be | ce | ce | ce | ef | ef | ef | ab |
| RUB 161 | aab | aaa | abc | abc | aac | abc | aaa | aab | ab | aa | aa | aa | cc | cc | cc | ac | ac | aa | aa | aa | cd | cd | cd | aa |
| RUB 178 | aab | bcc | cce | ccf | aad | bcc | ccc | acc | ac | aa | ac | cj | bd | ef | ef | dg | dg | dh | dh | dh | ii | ii | ii | bd |
| RUB 179 | aab | aab | bbc | abc | bbc | abc | abb | aab | ab | dd | aa | ad | cc | cc | cc | cc | cc | cc | cc | cc | cd | cd | cd | ac |
| RUB 183 | abc | ccg | acd | acd | aaf | ace | acc | aaa | ac | cc | ac | af | ef | dd | dd | dd | dd | ad | ad | ad | af | af | af | bg |
| RUB 184 | abc | abc | ccd | bcd | abc | abc | bcc | bcd | bc | cd | dd | bc | aa | dd | dd | ad | ad | aa | aa | aa | bd | bd | bd | aa |
| RUB 190 | bbc | bbb | abc | abc | abc | bbc | aaa | aaa | ab | ad | ah | aa | cc | cc | cc | ef | ef | dd | dd | dd | gg | gg | gg | bc |
| RUB 199 | abc | bcc | abc | abc | aab | acc | acc | aac | ac | ac | bc | aa | bc | bb | bb | cc | cc | bd | bd | bd | cc | cc | cc | bb |
| Rubber Trees | N | N/25 (%) |
|---|---|---|
| PB 310 | 11 | 44.00 |
| Yunyan 277-5 | 14 | 56.00 |
| RRIC 103 | 13 | 52.00 |
| A1 | 14 | 56.00 |
| A2 | 16 | 64.00 |
| A3 | 16 | 64.00 |
| A4 | 16 | 64.00 |
| A5 | 16 | 64.00 |
| A6 | 14 | 56.00 |
| A7 | 14 | 56.00 |
| A8 | 13 | 52.00 |
| A9 | 9 | 36.00 |
| A10 | 8 | 32.00 |
| A11 | 8 | 32.00 |
| A12 | 25 | 100.00 |
| Triploids | Male Parent | N1/25 (%) |
|---|---|---|
| Tri-1 | A12 | 100.00 |
| Tri-2 | A12 | 100.00 |
| Tri-3 | A2/A3 | 100.00 |
| Tri-4 | A2/A3 | 100.00 |
| Tri-5 | A1 | 100.00 |
| Tri-6 | A1 | 100.00 |
| Tri-7 | RRIC 103 | 100.00 |
| Tri-8 | Yunyan 277-5 | 96.00 |
| Markers | Triploid Samples | HR% | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Tri-1 | Tri-2 | Tri-3 | Tri-4 | Tri-5 | Tri-6 | Tri-7 | Tri-8 | ||
| RUB 9 | bc | bc | bc | bc | bc | bc | bc | bc | 100.00 |
| RUB 16 | ac | ac | ac | ac | aa | ac | cc | ac | 75.00 |
| RUB 19 | ab | bb | bb | bb | aa | ab | bb | - | 28.57 |
| RUB 20 | aa | bb | bb | bb | aa | ab | - | aa | 14.29 |
| RUB 28 | ab | ab | ab | bb | ab | bb | bb | ab | 62.50 |
| RUB 33 | ab | ab | bb | bb | aa | aa | bb | aa | 25.00 |
| RUB 65 | bb | ab | aa | aa | bb | bb | aa | - | 14.29 |
| RUB 70 | - | aa | ab | aa | bb | aa | aa | - | 16.67 |
| RUB 74 | ab | ab | ab | aa | ab | ab | - | aa | 71.43 |
| RUB 75 | ab | ab | ab | aa | - | aa | aa | ab | 57.14 |
| RUB 90 | ab | aa | ab | ab | ab | ab | ab | ab | 87.50 |
| RUB 95 | ab | ab | ab | ab | ab | ab | ab | bb | 87.50 |
| RUB 98 | ab | aa | ab | ab | ab | aa | - | - | 66.67 |
| RUB 102 | aa | bb | bb | bb | aa | ab | - | aa | 14.29 |
| RUB 103 | ab | bb | ab | bb | aa | bb | ab | aa | 37.50 |
| RUB 112 | ab | ab | ab | ab | aa | ab | ab | - | 85.71 |
| RUB 138 | bb | bb | ab | aa | bb | aa | - | aa | 14.29 |
| RUB 156 | ac | aa | ac | ac | ac | ac | - | ac | 85.71 |
| RUB 161 | ab | aa | ab | ab | aa | ab | aa | ab | 62.50 |
| RUB 178 | aa | cc | cc | cc | aa | cc | cc | - | 0.00 |
| RUB179 | ab | ab | bb | ab | bb | ab | bb | ab | 62.50 |
| RUB 183 | ac | cc | ac | ac | aa | ac | cc | aa | 50.00 |
| RUB 184 | bc | bc | cc | bc | bc | bc | - | bc | 85.71 |
| RUB 190 | bb | bb | ab | ab | ab | bb | aa | aa | 37.50 |
| RUB 199 | ac | cc | ac | ac | aa | ac | cc | aa | 50.00 |
| HR% | 75.00 | 44.00 | 68.00 | 52.00 | 37.50 | 60.00 | 27.78 | 47.37 | |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yao, P.-Q.; Li, G.-H.; Long, Q.-Y.; He, L.-G.; Kang, X.-Y. Male Parent Identification of Triploid Rubber Trees (Hevea brasiliensis) and the Mechanism of 2n Gametes Formation. Forests 2016, 7, 301. https://doi.org/10.3390/f7120301
Yao P-Q, Li G-H, Long Q-Y, He L-G, Kang X-Y. Male Parent Identification of Triploid Rubber Trees (Hevea brasiliensis) and the Mechanism of 2n Gametes Formation. Forests. 2016; 7(12):301. https://doi.org/10.3390/f7120301
Chicago/Turabian StyleYao, Peng-Qiang, Guo-Hua Li, Qing-Yi Long, Li-Gang He, and Xiang-Yang Kang. 2016. "Male Parent Identification of Triploid Rubber Trees (Hevea brasiliensis) and the Mechanism of 2n Gametes Formation" Forests 7, no. 12: 301. https://doi.org/10.3390/f7120301
APA StyleYao, P.-Q., Li, G.-H., Long, Q.-Y., He, L.-G., & Kang, X.-Y. (2016). Male Parent Identification of Triploid Rubber Trees (Hevea brasiliensis) and the Mechanism of 2n Gametes Formation. Forests, 7(12), 301. https://doi.org/10.3390/f7120301





