Developments on Core Collections of Plant Genetic Resources: Do We Know Enough?
Abstract
1. What Is a Core Collection?
2. The Progress of Core Collection
- Have core collections been formed for a diversity of plants?
- How can we effectively construct a representative core collection?
- How well can the core collection be utilized?
2.1. Diversity of Core Collections
2.2. Procedure of Constructing a Core Collection
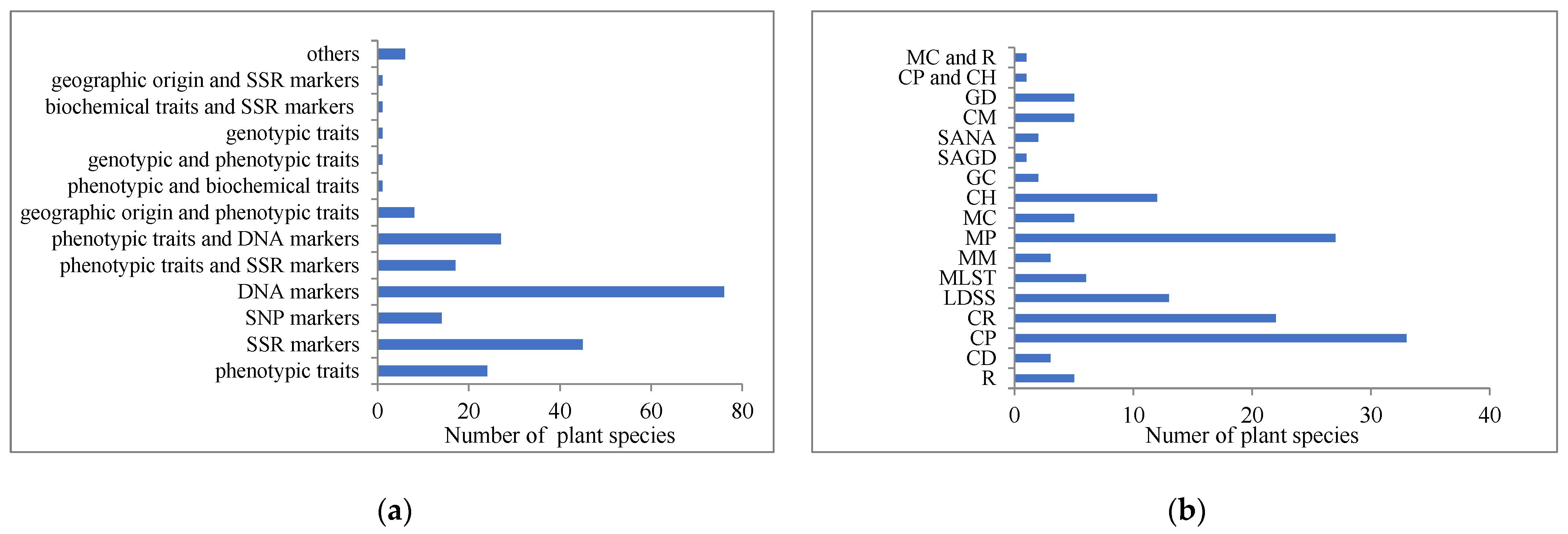
2.2.1. Data Selection for Core Collection
2.2.2. Separating the Accessions into Meaningful Groups
2.2.3. Sampling Strategies of Core Collection
2.2.4. Sampling Proportion of Core Collection
2.3. Evaluation of Core Collection
3. The Use of Core Collection
3.1. Genomic Study and Marker Development
3.1.1. Evaluation of Genetic Diversity
3.1.2. Classification of Species and Varieties
3.1.3. Construction of Genetic Maps
3.1.4. Discovery of Specific Genes
3.2. Identification of Disease or Pest Resistance
3.3. Gene Discovery and Allele Mining
4. Conclusions and Perspectives
4.1. Effective Data Integration and Accurate Phenotyping Methods
4.2. Appropriate Sampling Strategy and Software
4.3. Broaden the Scope of Core Collection
4.4. Identify Abundant Functional Genes and Markers
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Liu, Y.L.; Geng, Y.P.; Xie, X.D.; Zhang, P.F.; Hou, J.L.; Wang, W. Core collection construction and evaluation of the genetic structure of Glycyrrhiza in China using markers for genomic simple sequence repeats. Genet. Resour. Crop Evol. 2020, 67, 1839–1852. [Google Scholar] [CrossRef]
- Belaj, A.; del Carmen Dominguez-García, M.; Atienza, S.G.; Urdíroz, N.M.; De la Rosa, R.; Satovic, Z.; Martín, A.; Kilian, A.; Trujillo, I.; Valpuesta, V.; et al. Developing a core collection of olive (Olea europaea L.) based on molecular markers (DArTs, SSRs, SNPs) and agronomic traits. Tree Genet. Genomes 2012, 8, 365–378. [Google Scholar] [CrossRef]
- Van Hintum, T.J.L.; Brown, A.H.D.; Spillane, C.; Hodgkin, T. Core Collections of Plant Genetic Resources; IPGRI Technical Bulletin No. 3; International Plant Genetic Resources Institute: Rome, Italy, 2000. [Google Scholar]
- Brown, A.H.D. Core collections—A practical approach to genetic-resources management. Genome 1989, 31, 818–824. [Google Scholar] [CrossRef]
- Brown, A.H.D. The case for core collections. In The Use of Plant Genetic Resources; Brown, A.H.D., Frankel, O.H., Marshal, D.R., Eds.; University Cambridge Press: Cambridge, UK, 1989; pp. 136–156. [Google Scholar]
- Frankel, O.H. Genetic perspectives of germplasm conservation. In Genetic Manipulation: Impact on Man and Society; Arber, W.K., Illmensee, K., Peacock, W.J., Starlinger, P., Ehrlich, S.D., Dagert, M., Romac, S., Michel, B., Levy, S.B., Goebel, W., et al., Eds.; Cambridge University Press: Cambridge, UK, 1984; pp. 161–170. [Google Scholar]
- Li, Z.C.; Zhang, H.L.; Cao, Y.S.; Qiu, Z.E.; Wei, X.H.; Tang, S.X.; Yu, P.; Wang, X.K. Studies on the sampling strategy for primary rice. Acta Agron. Sin. 2003, 29, 20–24. [Google Scholar]
- Odong, T.L.; Jansen, J.; van Eeuwijk, F.A.; van Hintum, T.J.L. Quality of core collections for effective utilisation of genetic resources review, discussion and interpretation. Theor. Appl. Genet. 2013, 126, 289–305. [Google Scholar] [CrossRef] [PubMed]
- Balfourier, F.; Roussel, V.; Strelchenko, P.; Exbrayat-Vinson, F.; Sourdille, P.; Boutet, G.; Koenig, J.; Ravel, C.; Mitrofanova, O.; Beckert, M.; et al. A worldwide bread wheat core collection arrayed in a 384-well plate. Theor. Appl. Genet. 2007, 11, 1265–1275. [Google Scholar] [CrossRef] [PubMed]
- Richards, C.M.; Volk, G.M.; Reeves, P.A.; Reilley, A.A.; Henk, D.; Forsline, P.L.; Aldwinckle, H.S. Selection of stratified core sets representing wild apple (Malus sieversii). J. Am. Soc. Hortic. Sci. 2009, 134, 228–235. [Google Scholar] [CrossRef]
- Razieh, M.; Mohammad, R.D.; Darab, H.; Mehrshad, Z.; Elisa, V.; Sabrina, M.; Fariborz, Z.N. Development of a core collection in Iranian walnut (Juglans regia L.) germplasm using the phenotypic diversity. Sci. Hortic. 2019, 249, 439–448. [Google Scholar]
- Li, X.X.; Fang, Z.Y. From the research of core germplasm, the mining and utilization of excellent genes of crops are carried out. China Veg. 2005, S1, 1–7. (In Chinese) [Google Scholar]
- Jiang, M. Researches on Constructing Strategy of Core Germplasm Construction Based on Phenotype and Genetic Diversity of Buchloe dactyloides (Nutt.) Engelm. Master’s Thesis, Chinese Academy of Forestry, Beijing, China, 2020. [Google Scholar]
- Huang, C.Q.; Long, T.; Bai, C.J.; Wang, W.Q.; Tang, J.; Liu, G.D. Establishment of a core collection of Cynodon based on morphological data. Trop. Grassl.-Forrajes Trop. 2020, 8, 203–213. [Google Scholar] [CrossRef]
- Zhou, Y.C. Genetic Diversity Analysis and Construction of Core Germplasm in Bromus Inermis Leyss. Ph.D. Thesis, Northeast Normal University, Changchun, China, 2020. [Google Scholar]
- Wu, H.; Duan, A.; Wang, X.; Chen, Z.; Zhang, X.; He, G.; Zhang, J. Construction of a Core Collection of Germplasms from Chinese Fir Seed Orchards. Forests 2023, 14, 305. [Google Scholar] [CrossRef]
- Guo, Q.; Liu, J.; Li, J.K.; Cao, S.; Zhang, Z.J.; Zhang, J.T.; Zhang, Y.S.; Deng, Y.P.; Niu, D.S.; Su, L.Z.; et al. Genetic diversity and core collection extraction of Robinia pseudoacacia L. germplasm resources based on phenotype, physiology, and genotyping markers. Ind. Crops Prod. 2022, 178, 114627. [Google Scholar] [CrossRef]
- Mao, X.H.; Zhu, S.L.; Li, S.W.; Hua, H.; Tian, S.Y.; Zhong, W.G.; Dong, Y.F.; An, X.M. Core germplasm construction of Populus tomentosa based on the fluorescent SSR markers. J. Beijing For. Univ. 2020, 42, 40–47. (In Chinese) [Google Scholar]
- Feng, Y.; Yang, Z.; Tan, J.; Li, H.; Chen, X. Selection of first generation nucleus population of Pinus massoniana in Guangxi. J. Northeast For. Univ. 2018, 46, 20–24. [Google Scholar]
- Shashidhara, G.; Hema, M.V.; Koshy, B.; Farooqi, A.A. Assessment of genetic diversity and identification of core collection in sandalwood germplasm using RAPDs. J. Hortic. Sci. Biotechnol. 2003, 78, 528–536. [Google Scholar] [CrossRef]
- Li, R.; Shang, X.; Shang, C.S.; Chang, L.F.; Yan, L.; Bai, J.R. Core germplasm construction of Shanxi maize inbred line based on SSR markers. Seed 2021, 9, 72–76. (In Chinese) [Google Scholar]
- Chang, L.F.; Bai, J.R.; Li, R.; Zhang, C.Z.; Zhang, X.M.; Yang, R.J. Construction of a Core Collection of Sweet Corn Populations Based on SSR Markers. J. Maize Sci. 2018, 26, 40–49. (In Chinese) [Google Scholar]
- Li, Y.; Shi, Y.S.; Cao, Y.S.; Wang, T.Y. Establishment of a core collection for maize germplasm preserved in Chinese National Genebank using geographic distribution and characterization data. Genet. Resour. Crop Evol. 2004, 51, 845–852. [Google Scholar] [CrossRef]
- Ronaldo, R.C.; Glauco, V.M.; Cosme, D.C. Development of a Brazilian maize core collection. Genet. Mol. Biol. 2009, 32, 538–545. [Google Scholar]
- Li, M.; Qin, H.B.; Wang, Y.N.; Mu, Z.X.; Du, H.L. A Core Collection of Sorghum Landraces Formed by Taking Use of Agronomic Traits in Shanxi Province. J. Plant Genet. Resour. 2021, 22, 174–182. (In Chinese) [Google Scholar]
- Li, X.S.; Fu, Y.H.; Zhou, X.; Li, Q.; Liu, F.Z.; Yang, C.L.; Zhou, M.Q. Establishment of Coix lacryma-jobi L. Core Germplasm Collection Based on Phenotypic Characters. Chin. J. Trop. Crops 2020, 41, 669–675. (In Chinese) [Google Scholar]
- Yuan, H.J.; Zeng, X.Q.; Xu, Q.J.; Wang, Y.L.; Zhang, S.; Nyima, T.S. Genetic diversity of germplasm resources and core construction development in Hulless Barley. J. Triticeae Crops 2018, 38, 922–928. (In Chinese) [Google Scholar]
- Zhang, H.L.; Zhang, D.L.; Wang, M.X.; Sun, J.L.; Qi, Y.W.; Li, J.J.; Wei, X.H.; Han, L.Z.; Qiu, Z.E.; Tang, S.X.; et al. A core collection and mini core collection of Oryza sativa L. in China. Theor. Appl. Genet. 2011, 122, 49–61. [Google Scholar] [CrossRef]
- Gao, X.F.; Zhou, Y.; Song, D.Y.; Li, J.X.; Chen, W.; Li, S.P. Construction of Core Collection of Chinese Aegilops tauschii Coss. Germplasm Resource Based on Spike Morphological Traits and Molecular Markers. J. Plant Genet. Resour. 2021, 22, 361–370. (In Chinese) [Google Scholar]
- Boczkowska, M.; Lapinski, B.; Kordulasinska, I.; Dostatny, D.F.; Czembor, J.H. Promoting the use of common oat genetic resources through diversity analysis and core collection construction. PLoS ONE 2016, 11, e0167855. [Google Scholar] [CrossRef]
- Li, J.L.; Fan, Y.; Zhao, M.Y. Construction of Primary Core Collection of Buckwheat Germplasm Resources Based on Phenotypic Traits and SSR. J. Plant Genet. Resour. 2021, 22, 1240–1247. (In Chinese) [Google Scholar]
- Bhattacharjee, R.; Khairwal, I.S.; Bramel, P.J.; Reddy, K.N. Establishment of a pearl millet [Pennisetum glaucum (L.) R. Br.] core collection based on geographical distribution and quantitative traits. Euphytica 2007, 155, 35–45. [Google Scholar] [CrossRef]
- Choi, Y.M.; Kim, K.M.; Lee, S.; Oh, S.; Lee, M.C. Development of a core collection based on EST-SSR markers and phenotypic traits in foxtail millet [Setaria italica (L.) P. Beauv.]. J. Crop Sci. Biotechnol. 2019, 21, 395–405. [Google Scholar] [CrossRef]
- Upadhyaya, H.D.; Bramel, P.J.; Ortiz, R.; Singh, S. Developing a mini core of peanut for utilization of genetic resources. Crop Sci. 2002, 42, 2150–2156. [Google Scholar] [CrossRef]
- Su, W.; Wang, L.; Lei, J.; Chai, S.; Liu, Y.; Yang, Y.; Yang, X.; Jiao, C. Genome-wide assessment of population structure and genetic diversity and development of a core germplasm set for sweet potato based on specific length amplified fragment (SLAF) sequencing. PLoS ONE 2017, 12, e0172066. [Google Scholar] [CrossRef]
- Oliveira, E.J.; Ferreira, C.F.; Santos, V.S.; Oliveira, G.A.F. Development of a cassava core collection based on single nucleotide polymorphism markers. Genet. Mol. Res. 2014, 13, 6472–6485. [Google Scholar] [CrossRef] [PubMed]
- Upadhyaya, H.D.; Bramel, P.J.; Singh, S. Development of a chickpea core subset using geographic distribution and quantitative traits. Crop Sci. 2001, 41, 206–210. [Google Scholar] [CrossRef]
- Reddy, L.J.; Upadhyaya, H.D.; Gowda, C.L.L.; Singh, S. Development of core collection in pigeonpea [Cajanus cajan (L.) Millspaugh] using geographic and qualitativemorphological descriptors. Genet. Resour. Crop Evol. 2005, 52, M1049–M1056. [Google Scholar] [CrossRef]
- Regina, L.F.G.; Marcones, F.C.; Alessandro, A.-P. A lima bean core collection based on molecular markers. Sci. Agric. 2020, 77, e20180140. [Google Scholar]
- Wang, L.X.; Guan, Y.; Guan, R.X.; Li, Y.; Ma, Y.; Dong, Z.; Liu, X.; Zhang, H.; Zhang, Y.; Liu, Z.; et al. Establishment of Chinese soybean (Glycine max) core collections with agronomic traits and SSR markers. Euphytica 2006, 151, 215–223. [Google Scholar] [CrossRef]
- Jeong, N.; Kim, K.S.; Jeong, S.; Kim, J.Y.; Park, S.K.; Lee, J.S.; Jeong, S.C.; Kang, S.T.; Ha, B.K.; Kim, D.Y.; et al. Korean soybean core collection: Genotypic and phenotypic diversity population structure and genomewide association study. PLoS ONE 2019, 14, e0224074. [Google Scholar] [CrossRef]
- Wang, L.X.; Cheng, X.Z.; Wang, S.H. Genetic Diversity Analysis and a Core Collection of Rice Bean (Vigna umbellata) in China. J. Plant Genet. Resour. 2014, 15, 242–247. (In Chinese) [Google Scholar]
- Hao, X.P.; Wang, Y.; Tian, X. Construction of Primary Core Collection of Common Bean (Phaseolus vulgaris L.) Based on Agronomic Traits in Shanxi Province. J. Plant Genet. Resour. 2016, 17, 815–823. (In Chinese) [Google Scholar]
- Jiang, J.Y.; Yang, T.; Wang, F.G. Diversity Analysis of Germplasm Resources and Construction of Mini-core Collections for Vicia faba L. at Home and Abroad. Acta Agron. Sin. 2014, 40, 1311–1319. (In Chinese) [Google Scholar] [CrossRef]
- Schafleitner, R.; Nair, R.M.; Rathore, A.; Wang, Y.W.; Lin, C.Y.; Chu, S.H.; Lin, P.Y.; Chang, J.C.; Ebert, A.W. The AVRDC-The World Vegetable Center mungbean (Vigna radiata) core and mini core collections. BMC Genom. 2015, 16, 0111. [Google Scholar] [CrossRef]
- Wang, M.M.; Sun, D.L.; Chen, R.; Yang, Y.; Zhang, G.; Lv, M.; Wang, Q.; Xie, T.; Niu, G.; Shan, X.; et al. Construction and Evaluation of Cauliflower Core Collection. Acta Hortic. Sin. 2023, 50, 421–431. (In Chinese) [Google Scholar]
- Zhang, G.G.; Liu, H.D.; Du, D.Z. Population structure analysis and core germplasm construction of Brassica napus restorer lines. Chin. J. Oil Crop Sci. 2022, 175, 1–9. (In Chinese) [Google Scholar]
- Li, L.; He, W.M.; Ma, L.P.; Liu, P.; Xu, H.; Xu, J.; Zheng, X. Construction Chinese Cabbage (Brassica rapa L.) Core Collection and its EST-SSR Fingerprint Database by EST-SSR Molecular Markers. Genom. Appl. Biol. 2009, 28, 76–88. (In Chinese) [Google Scholar]
- Zheng, F.S.; Wang, X.M.; Li, G.H. Core collection construction of Ningxia tomato germplasm resources based on phenotypic traits. J. Zhejiang Univ. 2021, 47, 171–181. (In Chinese) [Google Scholar]
- Deng, X.B.; Liu, L.; Yan, Z.; Li, T.; Liu, X.Y.; Feng, J.J.; Chi, H.J.; Zheng, Z.; Li, J.M. Development of a Core Collection of Processing Tomato Germplasms and Analysis of Genetic Background. Acta Hortic. Sin. 2015, 42, 1299–1312. (In Chinese) [Google Scholar]
- Peng, F.; Li, Y.; Dai, Y.R.; Wang, X.M.; Yang, J.; Wang, Q.H.; Cai, X.F. Construction of spinach’s core germplasms based on its phenotypic traits. J. Shanghai Norm. Univ. 2022, 51, 9–19. (In Chinese) [Google Scholar]
- Hoshikawa, K.; Lin, Y.P.; Schafleitner, R.; Shirasawa, K.; Isobe, S.; Nguyen, D.C.; Ohsawa, R.; Yoshioka, Y. Genetic diversity analysis and core collection construction for Amaranthus tricolor germplasm based on genome-wide single-nucleotide polymorphisms. Sci. Hortic. 2023, 307, 111428. [Google Scholar] [CrossRef]
- Cui, J.J.; Cheng, J.W.; Cao, Y. Studies on Construction of Bitter Gourd Core Collection Based on SSR Markers and Phenotypic Traits. China Veg. 2022, 2, 25–32. (In Chinese) [Google Scholar]
- Hou, Z.Q.; Wang, L.H.; Zhao, M.L.; Yang, S.P.; Sun, X.M.; Gao, J.M.; Zhong, Q.W. Preliminary Construction of Core Collection of Jerusalem Artichoke Based on Phenotypic Data. Mol. Plant Breed. 2021, 19, 3463–3472. (In Chinese) [Google Scholar]
- Yang, F.; Zhang, Y.F.; Wang, J.H. SSR analysis of Chinese yam (Dioscorea opposita Thunb.) germplasm resources and construction of primary core collection. J. North. Agric. 2022, 50, 18–27. (In Chinese) [Google Scholar]
- Wang, X.; Bao, K.; Reddy, U.K.; Bai, Y.; Hammar, S.A.; Jiao, C.; Wehner, T.C.; Ramírez-Madera, A.O.; Weng, Y.Q.; Grumet, R.; et al. The USDA cucumber (Cucumis sativus L.) collection: Genetic diversity, population structure, genome-wide association studies, and core collection development. Hortic. Res. 2018, 5, 64. [Google Scholar] [CrossRef]
- Ge, R.D. Construction of Pumpkin Core Germplasm for Anvils. Master’s Thesis, Nanjing Agricultural University, Nanjing, China, 2020. [Google Scholar]
- Jiao, X.X. Construction of Core Collections of Wax Gourd. Master’s Thesis, Guangxi University, Nanning, China, 2018. [Google Scholar]
- Lee, H.Y.; Ro, N.Y.; Jeong, H.J.; Kwon, J.K.; Jo, J.; Ha, Y. Genetic diversity and population structure analysis to construct a core collection from a large Capsicum germplasm. BMC Genet. 2016, 17, 142. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.J.; Cao, Z.M.; Zhu, J.; Shen, L.B. Comparative study on the construction of sweet pepper core collection. J. Northeast Agric. Univ. 2016, 47, 21–29. (In Chinese) [Google Scholar]
- Miyatake, K.; Shinmura, Y.; Matsunaga, H.; Fukuoka, H.; Saito, T. Construction of a core collection of eggplant (Solanum melongena L.) based on genome-wide SNP and SSR genotypes. Breed. Sci. Preview 2019, 69, 498–502. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Cui, L.; Zhang, L.; Huang, Y.; Zhang, S.; Chen, W.; Deng, X.; Jiao, Z.; Yang, W.; Qiu, Z.; et al. Genetic Diversity Analysis and Core Germplasm Collection Construction of Radish Cultivars Based on Structure Variation Markers. Int. J. Mol. Sci. 2023, 24, 2554. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Zhou, F.; Gao, Y.; Liu, C.; Yu, S.; Zhao, K.; Gong, W.; Lang, J.; Zhang, H.; Yu, X. Genetic Diversity and Primary Core Collection Construction of Turnip (Brassica rapa L. ssp. rapifera Matzg) Landraces in Tibet Revealed via Morphological and SSR Markers. Agronomy 2021, 11, 1901. [Google Scholar] [CrossRef]
- Li, J.; Liu, X.B.; Zhao, Z.W.; Yang, Z.L. Genetic diversity, core collection and breeding history of Pleurotus ostreatus in China. Mycoscience 2019, 60, 14–24. [Google Scholar] [CrossRef]
- Sa, K.J.; Kim, D.M.; Oh, J.S.; Park, H.; Hyun, D.Y.; Lee, S.; Rhee, J.H.; Lee, J.K. Construction of a core collection of native Perilla germplasm collected from South Korea based on SSR markers and morphological characteristics. Sci. Rep. 2021, 11, 23891. [Google Scholar] [CrossRef]
- Xu, Y.; Chen, C.S.; Ji, D.H.; Xu, K.; Xie, X.X.; Xie, C.T. Developing a core collection of Pyropia haitanensis using simple sequence repeat markers. Aquaculture 2016, 452, 351–356. [Google Scholar] [CrossRef]
- Sun, Y.Q.; Chen, J.H.; Zhang, J. Construction A Core Collection of Armeniaca sibirica Based on Phenotypic Traits. J. Shenyang Agric. Univ. 2022, 53, 43–54. (In Chinese) [Google Scholar]
- Liu, J.; Liao, K.; Zhao, S.R. The Core Collection Construction of Xinjiang Wild Apricot Based on ISSR Molecular Markers. Sci. Agric. Sin. 2015, 48, 2017–2028. (In Chinese) [Google Scholar]
- Wang, Y.Z.; Zhang, J.H.; Sun, H.Y.; Ning, N. Construction and evaluation of a primary core collection of apricot germplasm in China. Sci. Hortic. 2011, 128, 311–319. [Google Scholar] [CrossRef]
- Sun, Y.; Dong, S.; Liu, Q.; Chen, J.; Pan, J.; Zhang, J. Selection of a core collection of Prunus sibirica L. germplasm by a stepwise clustering method using simple sequence repeat markers. PLoS ONE 2021, 16, e0260097. [Google Scholar] [CrossRef] [PubMed]
- Lu, M.; Zhang, H.; Bai, J. Genetic Diversity Analysis and Core Collection Construction in Rosa roxburghii Based on Fruits Quality and EST-SSR Markers. Mol. Plant Breed. 2020, 18, 3098–3106. (In Chinese) [Google Scholar]
- Bu, H.D.; Zhang, B.B.; Song, H.W.; Liang, Y.H.; Liu, Y.J.; Cheng, X.M.; Gu, G.J.; Liu, C. Construction Core Collections of Pear Germplasms in Cold Region by SSR and Phenotypic Traits. Acta Hortic. Sin. 2012, 39, 2113–2123. (In Chinese) [Google Scholar]
- Sun, Y.Q. Genetic Diversity Analysis and Core Collection Construction of Ziziphus jujuba var. spinosa (Bunge) Hu ex H. F. Chow. Master’s Thesis, Tarim University, Aral, China, 2016. [Google Scholar]
- Xu, C.; Gao, J.; Du, Z.; Li, D.; Wang, Z.; Li, Y.; Pang, X. Identifying the genetic diversity, genetic structure and a core collection of Ziziphus jujube Mill. Var. jujube accessions using microsatellite markers. Nat. Sci. Rep. 2016, 6, 31503. [Google Scholar]
- Sivalingam, P.N.; Singh, D.; Chauhan, S.; Changal, H.K.; Bhan, C.; Mohapatra, T.; More, T.A.; Sharma, S.K. Establishment of the core collection of Ziziphus mauritiana Lam. from India. Plant Genet. Resour. 2013, 12, 140–142. [Google Scholar] [CrossRef]
- Le Cunff, L.; Fournier-Level, A.; Laucou, V.; Vezzulli, S.; Lacombe, T.; Adam-Blondon, A.F.; Boursiquot, J.-M.; This, P. Construction of nested genetic core collections to optimize the exploitation of natural diversity in Vitis vinifera L. subsp. sativa. BMC Plant Biol. 2008, 8, 31. [Google Scholar] [CrossRef]
- Hu, J.; Wang, P.; Su, Y.; Wang, R.; Li, Q.; Sun, K. Microsatellite diversity, population structure, and core collection formation in Melon germplasm. Plant Mol. Biol. Rep. 2015, 33, 439–447. [Google Scholar] [CrossRef]
- Pal, S.; Revadi, M.; Thontadarya, R.N.; Reddy, D.C.L.; Varalakshmi, B.; Pandey, C.D.; Rao, E.S. Understanding genetic diversity, population structure and development of a core collectionof Indian accessions of watermelon (Citrullus lanatus (Thunb.) Matsum and Nakai). Plant Genet. Resour. Charact. Util. 2020, 18, 359–368. [Google Scholar] [CrossRef]
- Hu, G.G.; Jiang, Q.; Wang, Z. Genetic Diversity Analysis and Core Collection Construction of the Actinidia chinensis Complex (Kiwifruit) Based on SSR Markers. Agronomy 2022, 12, 3078. [Google Scholar] [CrossRef]
- Razi, S.; Soleimani, A.; Zeinalabedini, M.; Vazifeshenas, M.R.; Martinez-Gomez, P.; Kermani, A.M.; Raiszadeh, A.R.; Tayari, M.; Martinez-Garcia, P.J. Development of a Multipurpose Core Collection of New Promising Iranian Pomegranate (Punica granatum L.) Genotypes Based on Morphological and Pomological Traits. Horticulturae 2021, 7, 350. [Google Scholar] [CrossRef]
- Huang, X.F.; Wei, Y.L.; Yuan, Y. Genetic diversity analysis and core collection construction of 221 cultivars (strains) of Litchi chinensis based on SNP molecular markers. J. Plant Resour. Environ. 2022, 31, 74–84. (In Chinese) [Google Scholar]
- Sun, Q.; Bai, L.; Ke, L.; Xiang, X.; Zhao, J.; Ou, L. Developing a core collection of litchi (Litchi chinensis Sonn.) based on EST-SSR genotype data and agronomic traits. Sci. Hortic. 2012, 146, 29–38. [Google Scholar] [CrossRef]
- Dervishi, A.; Jakse, J.; Ismaili, H.; Javornik, B.; Stajner, N. Genetic structure and core collection of Olive germplasm from Albania revealed by microsatellite markers. Genes 2021, 12, 256. [Google Scholar] [CrossRef]
- Zhang, C.Y.; Chen, X.S.; Zhang, Y.M.; Yuan, Z.H.; Liu, Z.C.; Wang, Y.L.; Lin, Q. Method of Constructing Core Collection for Malus sieversii in Xinjiang, China Using Molecular Markers. Agric. Sci. China 2009, 8, 267–284. [Google Scholar] [CrossRef]
- Liang, W.; Dondini, L.; Franceschi, P.D.; Paris, R.; Sansavini, S.; Tartarini, S. Genetic diversity, population structure and construction of a core collection of apple cultivars from Italian germplasm. Plant Mol. Biol. Rep. 2015, 33, 458–473. [Google Scholar] [CrossRef]
- Kim, J.H.; Oh, Y.; Lee, G.A.; Kwon, Y.S.; Kim, S.A.; Kwon, S.I.; Do, Y.S.; Choi, C. Genetic Diversity, Structure, and Core Collection of Korean Apple Germplasm Using Simple Sequence Repeat Markers. Hortic. J. 2019, 88, 329–337. [Google Scholar] [CrossRef]
- Li, Y.X.; Gao, Q.J.; Li, T.H. Sampling strategy based on fruit char acteristics for a primary core collection of peach cultivars. J. Fruit Sci. 2006, 23, 359–364. [Google Scholar]
- Escribano, P.; Viruel, M.A.; Hormaza, J.I. Comparison of different methods to construct a core germplasm collection in woody perennial species with simple sequence repeat markers. A case study in cherimoya (Annona cherimola, Annonaceae), an underutilised subtropical fruit tree species. Ann. Appl. Biol. 2008, 153, 25–32. [Google Scholar] [CrossRef]
- Balas, F.C.; Osuna, M.D.; Domínguez, G.; Pérez-Gragera, F.; López-Corrales, M. Ex situ conservation of underutilised fruit tree species: Establishment of a core collection for Ficus carica L. using microsatellite markers (SSRs). Tree Genet. Genomes 2014, 10, 703–710. [Google Scholar] [CrossRef]
- Campoy, J.A.; Lerigoleur-Balsemin, E.; Christmann, H.; Beauvieux, R.; Girollet, N.; Quero-Garcia, J.; Dirlewanger, E.; Barreneche, T. Genetic diversity, linkage disequilibrium, population structure and construction of a core collection of Prunus avium L. landraces and bred cultivars. BMC Plant Biol. 2016, 16, 49. [Google Scholar] [CrossRef]
- Liu, Y.; Sun, Z.; Liu, D.; Wu, B.; Zhou, Q. Screening the core collection of pomelo germplasm based on molecular marker. J. Fruit Sci. 2006, 23, 339–345. (In Chinese) [Google Scholar]
- Zhang, Y.F.; Zhang, Q.L.; Yang, Y.; Luo, Z.R. Development of primary core collection for Japanese persimmon originated in China (Diospyros kaki Thunb.) by stepwise clustering. Acta Hortic. 2007, 760, 6976. [Google Scholar]
- Liu, X.L.; Liu, H.B.; Ma, L.; Li, X.J.; Xu, C.H.; Su, H.S.; Ying, X.M.; Cai, Q.; Fan, Y.H. Construction of sugarcane hybrids core collection by using stepwise clustering sampling approach with molecular marker data. Acta Agron. Sin. 2014, 40, 1885–1894. [Google Scholar] [CrossRef]
- Chen, M.K.; Chen, L.; Sun, W.H.; Ma, S.H.; Lan, S.; Peng, D.H.; Liu, Z.J.; Ai, Y. Genetic Diversity Analysis and Core Collection of Cymbidium ensifolium Germplasm Resources. Acta Hortic. Sin. 2022, 49, 175–186. (In Chinese) [Google Scholar]
- Li, J.W.; Su, J.S.; Zhang, F.; Fang, W.M.; Guan, Z.Y.; Chen, S.M.; Chen, F.D. Construction of Core Collection of Traditional Chrysanthemum morifolium Based on Phenotypic Traits. Sci. Agric. Sin. 2021, 54, 3514–3526. (In Chinese) [Google Scholar]
- Zhao, L.M.; Li, J.W.; Zhang, F.; Su, J.S.; Fang, W.M.; Wang, H.B.; Jiang, J.F.; Chen, S.M.; Chen, F.D.; Guan, Z.Y. Construction of a Core Collection of Spray Cut Chrysanthemum Based on Phenotypic Data. Acta Hortic. Sin. 2022, 49, 2273–2284. (In Chinese) [Google Scholar]
- Ming, J.; Zhang, Q.X.; Lan, Y.P. Core collection of Prunus mume Sieb. et Zucc. J. Beijing For. Univ. 2005, 27, 65–69. (In Chinese) [Google Scholar]
- Hu, H. Studies on the Construction of Core Collection for Wintersweet (Chimonanthus praeco) and Its Vegetative Propagation Technology. Master’s Thesis, Southwest University, Chongqing, China, 2020. [Google Scholar]
- Jiang, L.Y. The Wild Germplasm Resources Evaluation and Core Collection Establishment of Rosa rugosa. Master’s Thesis, Shandong Agricultural University, Tai’an, China, 2018. [Google Scholar]
- Li, M.Q.; Shi, G.Y.; Ye, S.H.; Huang, Y.W.; Bian, X.R. Methods of establishing Lanzhou lily core collection based on ISSR molecular marker data. J. Desert Res. 2015, 35, 1573–1578. (In Chinese) [Google Scholar]
- Li, B.Y. Studies on Genetic Diversity and Construction of Core Collection of Tree Peony Cultivars from Chiese Central Plains. Master’s Thesis, Beijing Forestry University, Beijing, China, 2007. [Google Scholar]
- Wang, Z.Y.; Li, Z.F.; Peng, C.; Chen, Y.; Zhang, X.Y. Construction of Lagerstroemia indica Core Collection based on SSR Markers and Comparison of Sampling Strategies. Mol. Plant Breed. 2022, 1, 1–13. (In Chinese) [Google Scholar]
- Wang, L.; Wang, J.M.; Wang, W.; Wang, L.; Wang, L.J.; Yan, X.C.; Tan, M.L. Development of a core collection in sunflower (Helianthus annuus L.) germplasm using phenotypic diversity. Chin. J. Oil Crop Sci. 2021, 43, 1052–1060. (In Chinese) [Google Scholar]
- Yang, L.; Li, H.; Guo, Q.Q. Construction of core collections of Sophora moorcroftiana endemic to the Qinghai–Tibet Plateau. Plant Physiol. J. 2022, 58, 1133–1144. (In Chinese) [Google Scholar]
- Li, J.H.; Ou, X.H.; Deng, W.J.; Luo, K.K.; Zhang, H.Y.; He, M.L.; Yan, H.J. Construction of core germplasm bank of Fallopia multiflora using SRAP molecular markers. Guihaia 2021, 41, 1920–1930. (In Chinese) [Google Scholar]
- Gong, F.S.; Geng, Y.P.; Zhang, P.F.; Zhang, F.; Fan, X.F.; Liu, Y.L. Genetic Diversity and Structure of Core Collection of Huangqi (Astragalus) Developed by Genomic Simple Sequence Repeat Markers. Res. Sq. 2022, 1, 1–12. [Google Scholar]
- Liu, R.X. Study on Construction of Scutellaria baicalensis Core Collection and Screening of Excellent Germplasm. Master’s Thesis, Beijing University of Chinese Medicine, Beijing, China, 2016. [Google Scholar]
- Liu, M.; Hu, X.; Wang, X.; Zhang, J.; Peng, X.; Hu, Z.; Liu, Y. Constructing a Core Collection of the Medicinal Plant Angelica biserrata Using Genetic and Metabolic Data. Front. Plant Sci. 2020, 11, 600249. [Google Scholar] [CrossRef] [PubMed]
- Yang, M.L. The Germplasm Evaluation and Core Collection Construction of Cornus officinalis Based on ISSR Markers. Master’s Thesis, Henan Agricultural University, Zhengzhou, China, 2016. [Google Scholar]
- Liu, F.M.; Zhang, N.N.; Liu, X.J.; Yang, Z.J.; Jia, H.Y.; Xu, D.P. Genetic diversity and population structure analysis of Dalbergia Odorifera germplasm and development of a core collection using microsatellite markers. Genes 2019, 10, 281. [Google Scholar] [CrossRef]
- Niu, S.; Koiwa, H.; Song, Q.; Qiao, D.; Chen, J.; Zhao, D.; Chen, Z.; Wang, Y.; Zhang, T. Development of core collections for Guizhou tea genetic resources and GWAS of leaf size using SNP developed by genotyping-by-sequencing. PeerJ 2020, 8, e8572. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Z.F. Analysis of Genetic Diversity and Core Collection Construction of Cultivated Tea Germplasm Resources in Guizhou. Master’s Thesis, Guizhou University, Guizhou, China, 2022. [Google Scholar]
- Wang, X.C.; Chen, L.; Yang, Y.J. Establishment of core collection for Chinese tea germplasm based on cultivated region grouping and phenotypic data. Front. Agric. China 2011, 5, 344–350. [Google Scholar] [CrossRef]
- Leroy, T.; de Bellis, F.; Legnate, H.; Musoli, P.; Kalonji, A.; Loor Solórzano, R.G.; Cubry, P. Developing core collections to optimize the management and the exploitation of diversity of the coffee Coffea canephora. Genetica 2014, 142, 185–199. [Google Scholar] [CrossRef]
- Martínez, I.B.; de la Cruz, V.M.; Nelson, M.R.; Bertin, P. Establishment of a core collection of traditional Cuban Theobroma cacao plants for conservation and utilization purposes. Plant Mol. Biol. Report. 2014, 35, 47–60. [Google Scholar] [CrossRef]
- Qian, Y.Y.; Liu, Y.; Cui, S.F.; Wang, G.G.; Zhang, X.; Jin, W.P.; Li, J.L. Analysis of Genetic Diversity of Cotton Germplasm Resources and Extraction of Core Germplasm Based on Phenotypic Traits. Acta Agric. Boreali-Simiga 2019, 34, 29–35. (In Chinese) [Google Scholar]
- Han, P.; Tian, X.M.; Wang, Y.; Huang, C.; Ma, Y.Z.; Zhou, X.F.; Yu, Y.; Zhang, D.W.; Xu, H.J.; Cao, Y.; et al. Construction of a core germplasm bank of upland cotton (Gossypium hirsutum L.) based on phenotype, genotype and favorable alleles. Genet. Resour. Crop Evol. 2022, 69, 2399–2411. [Google Scholar] [CrossRef]
- Mei, Y.J.; Zhou, J.P.; Xu, H.M. Development of Sea Island cotton (Gossypium barbadense L.) core collection using genotypic values. AJCS 2012, 6, 673–680. [Google Scholar]
- Luan, M.B.; Zou, Z.Z.; Zhu, J.J. Development of a core collection for ramie by heuristic search based on SSR markers. Biotechnol. Biotechnol. Equip. 2014, 28, 798–804. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Ambreen, H.; Variath, M.T.; Rao, A.R.; Agarwal, M.; Kumar, A.; Goel, S.; Jagannath, A. Utilization of molecular, phenotypic, and geographical diversity to develop compact composite core collection in the Oilseed Crop, Safflower (Carthamus tinctorius L.) through maximization strategy. Front. Plant Sci. 2016, 7, 1554. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.Y.; Mei, H.X.; Du, Z.W.; Wu, K.; Zheng, Y.; Cui, X.; Zheng, L. Construction of Core Collection of Sesame Based on Phenotype and Molecular Markers. Sci. Agric. Sin. 2017, 50, 2433–2441. (In Chinese) [Google Scholar]
- Ronfort, J.; Bataillon, T.; Santoni, S.; Delalande, M.; David, J.L.; Prosperi, J.M. Microsatellite diversity and broad scale geographic structure in a model legume: Building a set of nested core collection for studying naturally occurring variation in Medicago truncatula. BMC Plant Biol. 2006, 6, 28. [Google Scholar] [CrossRef]
- Fang, L.C.; Xia, H.M.; Ma, W.J.; Zhang, X.Y. Genetic Diversity Analysis and Primary Core Collection of Catalpa bungei Germplasm with SSR Markers. J. Northeast. For. Univ. 2017, 45, 1–5. (In Chinese) [Google Scholar]
- Xue, H.; Yu, X.; Fu, P.; Liu, B.; Zhang, S.; Li, J.; Zhai, W.; Lu, N.; Zhao, X.; Wang, J.; et al. Construction of the Core Collection of Catalpa fargesii f. duclouxii (Huangxinzimu) Based on Molecular Markers and Phenotypic Traits. Forests 2021, 12, 1518. [Google Scholar] [CrossRef]
- Xu, C.H.; Liu, X.L.; Mao, J.; Liu, H.B.; Lin, X.Q.; Lu, X.; Su, H.S. Construction of a core-collection of Saccharum spontaneum based on SSR molecular markers. J. Hunan Agric. Univ. 2020, 46, 657–663. (In Chinese) [Google Scholar]
- Peng, C.; Fan, X.P.; Su, X.H.; Long, K.L.; Zhang, X.Y. Construction of core collection in southern type of Populus deltoides using SSR makers. Acta Bot. Boreali-Occident. Sin. 2019, 39, 250–257. (In Chinese) [Google Scholar]
- Zhong, Y.D.; Liu, L.P.; Li, Y.Q.; Wu, Z.X.; Yang, A.; Liu, S.J.; Liu, Q.L.; Liu, T.Y.; Yu, F. Sampling Strategy of Primary Core Germplasm of Cinnamomum camphora in China. J. Southwest For. Univ. 2020, 40, 1–13. (In Chinese) [Google Scholar]
- Huang, Y.Q.; Yin, G.G.; Yang, J.C.; Yu, N.; Zou, W.T.; Li, R.S. Developing a Mini Core Germplasm of Phoebe bournei Based on SSR Molecular Marker. Mol. Plant Breed. 2020, 18, 2641–2648. (In Chinese) [Google Scholar]
- Wu, H.; Yu, W.W.; Wu, H.; Dong, L.M.; Dai, W.S.; Ye, S.Y.; Shen, D.F. Determination of Core Germplasm in Torreya grandis Fort. ex Lindl. Based on Seed Traits and Molecular Markers. Mol. Plant Breed. 2019, 17, 5513–5520. (In Chinese) [Google Scholar]
- Duan, F.; Zhang, H.; Li, S.; Tian, Z.Q.; Gan, X.H. Core Collection Construction of Endangered Plant Tetracentron sinense Based on ISSR Molecular Markers. Subtrop. Plant Sci. 2018, 47, 101–106. (In Chinese) [Google Scholar]
- Shen, Z.; Ma, L.Y.; Ao, Y.; Duan, J. Analysis of the genetic diversity and construction of core collection of Xanthoceras sorbifolia Bunge. Using microsatellite marker data. Mol. Plant Breed. 2017, 15, 3341–3350. (In Chinese) [Google Scholar]
- Yang, H.; Zhang, R.; Wang, B.; Xu, Z.; Zhou, Z. Construction of core collection of Schima superba based on SSR molecular markers. Sci. Silvae Sin. 2017, 53, 37–46. (In Chinese) [Google Scholar]
- Peng, C.; Li, Z.F.; Xiang, S.S.; Zhang, X.Y. Molecular Marker Evaluation and Construction of Primary Core Collection of Sapium sebiferum. Mol. Plant Breed. 2017, 15, 1455–1460. (In Chinese) [Google Scholar]
- Yan, L.P.; Wu, D.J.; Mao, X.H.; Yao, J.X.; Ren, F.; Li, S.W.; Wang, K.F.; Wang, Y.H.; Liu, C.L. Construction of core collection of Fraxinus based on SSR molecular markers. J. Cent. South Univ. For. Technol. 2019, 39, 1–9. (In Chinese) [Google Scholar]
- Li, H.G. Genetic Diversity Analysis, Core Collection Construction and Molecular Identification of Eucommia ulmoides Oliver. Ph.D. Thesis, Chinese Academy of Forestry, Beijing, China, 2017. [Google Scholar]
- Mao, J.; Liu, X.L.; Su, H.S.; Lu, X.; Lin, X.Q.; Cai, Q.; Fan, Y.H. Constructing Core Collection of Saccharum arundinaceum L. Based on Phenotype and Molecular Markers. J. Plant Genet. Resour. 2016, 17, 607–615. (In Chinese) [Google Scholar]
- Boccacci, P.; Aramini, M.; Ordidge, M.; van Hintum, T.J.L.; Marinoni, T.D.; Valentini, N.; Sarraquigne, J.-P.; Solar, A.; Rovira, M.; Bacchetta, L.; et al. Comparison of selection methods for the establishment of a core collection using SSR markers for hazelnut (Corylus avellana L.) accessions from European germplasm repositories. Tree Genet. Genomes 2021, 17, 48. [Google Scholar] [CrossRef]
- Bernard, A.; Barreneche, T.; Donkpegan, A.; Lheureux, F.; Dirlewanger, E. Comparison of structure analyses and core collections for the management of walnut genetic resources. Tree Genet. Genomes 2020, 8, 365–378. [Google Scholar] [CrossRef]
- Mahmoodi, R.; Dadpour, M.R.; Hassani, D.; Zeinalabedini, M.; Vendramin, E.; Leslie, C.A. Composite coreset construction and diversity analysis of Iranian walnut germplasm using molecular markers and phenotypic traits. PLoS ONE 2021, 3, e0248623. [Google Scholar]
- Wei, Z.G.; Gao, Y.C.; Yang, C.P. Sampling method for constructing germplasm core collections of Betula platyphylla. J. Northeast For. Univ. 2009, 37, 1–4. (In Chinese) [Google Scholar]
- Li, H.Y.; Liang, Z.W.; Chen, C.; Huang, H.H.; Tong, Z.K.; Lu, Y.Q. Core Germplasm of Betula luminifera Screened by Molecular Markers. J. Zhejiang For. Sci. Technol. 2011, 3, 1–4. (In Chinese) [Google Scholar]
- Zhao, J.; Tong, Y.Q.; Ge, T.; Ge, J. Genetic diversity estimation and core collection construction of Sinojackia huangmeiensis based on novel microsatellite markers. Biochem. Syst. Ecol. 2016, 64, 74e80. [Google Scholar] [CrossRef]
- Li, N.; Yang, Y.; Xu, F.; Chen, X.; Wei, R.; Li, Z.; Pan, W.; Zhang, W. Genetic Diversity and Population Structure Analysis of Castanopsis hystrix and Construction of a Core Collection Using Phenotypic Traits and Molecular Markers. Genes 2022, 13, 2383. [Google Scholar] [CrossRef]
- Zhang, Y.; Hu, D.; Zuo, J.; Zhang, P.; Wang, Z.; Chen, C. Development of a mulberry core collection originated in China to enhance germplasm conservation. Crop Breed. Appl. Biotechnol. 2019, 19, 55–61. [Google Scholar]
- Nie, X.H.; Wang, Z.H.; Liu, N.W.; Song, L.; Yan, B.Q.; Xing, Y.; Zhang, Q.; Fang, K.F.; Zhao, Y.L.; Chen, X.; et al. Fingerprinting 146 Chinese chestnut (Castanea mollissima Blume) accessions and selecting a core collection using SSR markers. J. Integr. Agric. 2021, 20, 1277–1286. [Google Scholar] [CrossRef]
- Pereira-Lorenzo, S.; Ramos-Cabrer, A.M.; Barreneche, T.; Mattioni, C.; Villani, F.; Díaz-Hernández, M.B.; Martín, L.M.; Martín, A. Database of European chestnut cultivars and definition of a core collection using simple sequence repeats. Tree Genet. Genomes 2017, 13, 114. [Google Scholar] [CrossRef]
- Miyamoto, N.; Ono, M.; Watanabe, A. Construction of a core collection and evaluation of genetic resources for Cryptomeria japonica (Japanese cedar). J. For. Res. 2015, 20, 186–196. [Google Scholar] [CrossRef]
- Lv, J.B.; Li, C.R.; Zhou, C.P.; Chen, J.B.; Li, F.G.; Wang, Q.G.; Li, M.; Wang, Y.; Chen, S.; Chen, J.; et al. Genetic diversity analysis of a breeding population of Eucalyptu scloeziana F.Muell. (Myrtaceae) and extraction ofacore germplasm collection using microsatellite markers. Ind. Crops Prod. 2020, 145, 112157. [Google Scholar] [CrossRef]
- Liu, D.H.; Zhang, F.Q.; Zhang, W.H. Establishment of Eucalyptus urophylla Core Collection Based on Geographical Distribution and Phenotypic Data. J. Southwest For. Univ. 2013, 33, 1–8. (In Chinese) [Google Scholar]
- Di Guardo, M.; Scollo, F.; Ninot, A.; Rovira, M.; Hermoso, J.F.; Distefano, G.; La Malfa, S.; Batlle, I. Genetic structure analysis and selection of a core collection for carob tree germplasm conservation and management. Tree Genet. Genomes 2019, 15, 41. [Google Scholar] [CrossRef]
- Ghazal, H.; Mouhaddab, J.; Ait Aabd, N.; Msanda, F.; Filali-Maltouf, F.; Belkadi, B.; Ferradouss, A.; El Modafar, C.; Ibnsouda Koraichi, S.; Ghazal, H.; et al. Assessing genetic diversity and constructing a core collection of an endangered Moroccan endemic tree [Argania spinosa (L.) Skeels]. Moroc. J. Biol. 2016, 13, 1–12. [Google Scholar]
- Wang, X.; Cao, Z.; Gao, C.; Li, K. Strategy for the construction of a core collection for Pinus yunnanensis Franch. to optimize timber based on combined phenotype and molecular marker data. Genet. Resour. Crop Evol. 2021, 68, 3219–3240. [Google Scholar] [CrossRef]
- Wang, X.; Liu, X.; Xing, S.; Kong, Q.; Zhang, Y.; Sun, L.; Gao, Y. AFLP analysis of genetic diversity and a construction of the core collection of partial ancient ginkgo trees in China. Acta Hortic. Sin. 2016, 43, 249–260. (In Chinese) [Google Scholar]
- Zhang, Z.; Yang, Q.; Niu, Y.; Zhang, Y.; Dong, S.; Zhang, W.; Wang, Z. Diversity analysis and establishment of core collection among Akebia trifoliata (Thunb.) Koidz. in Qinba mountain area of China using ISSR and SRAP markers. Genet. Resour. Crop Evol. 2020, 68, 1085–1102. [Google Scholar] [CrossRef]
- Zhu, Y.; Liang, D.; Song, Z.; Tan, Y.; Guo, X.; Wang, D. Genetic Diversity Analysis and Core Germplasm Collection Construction of Camellia oleifera Based on Fruit Phenotype and SSR Data. Genes 2022, 13, 2351. [Google Scholar] [CrossRef]
- Zhang, L.B. Analysis of Population Genetic Variation of Plus Trees and Construction of Core Collection in Cornus wilsoniana. Master’s Thesis, Beijing Forestry University, Beijing, China, 2020. [Google Scholar]
- Amalraj, V.A.; Balakrishnan, R.; Jebadhas, A.W.; Balasundaram, N. Constituting a core collection of Saccharum spontaneum L. and comparison of three stratified random sampling procedures. Genet. Resour. Crop Evol. 2006, 53, 1563–1572. [Google Scholar] [CrossRef]
- Chandra, S.; Huaman, Z.; Krishna, S.H.; Ortiz, R. Optimal sampling strategy and core collection size of Andean tetraploid potato based on isozyme data: A simulation study. Theor. Appl. Genet. 2002, 104, 1325–1334. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.; Zhu, J.; Xu, H.M. Methods of constructing core collections by stepwise clustering with three sampling strategies based on the genotypic values of crops. Theor. Appl. Genet. 2000, 101, 264–268. [Google Scholar] [CrossRef]
- Adal, A.M.; Demissie, Z.A.; Mahmoud, S.S. Identification, validation and cross-species transferability of novel Lavandula EST-SSRs. Planta 2015, 241, 987–1004. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.C.; Hu, J.; Xu, H.M.; Zhang, S. A strategy on constructing core collections by least distance stepwise sampling. Theor. Appl. Genet. 2007, 115, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Zhang, G.Y.; Wang, X.F.; Liu, S.J.; Ma, Z.Y. Cluster analysis and sampling methods for core collection construction in glandless cotton. Cotton Sci. 2004, 16, 8–12. [Google Scholar]
- Zhang, X.; Zhao, Y.; Cheng, Y.; Feng, X.; Guo, Q.; Zhou, M.; Hodgkin, T. Establishment of sesame germplasm core collection in China. Genet. Resour. Crop Evol. 2000, 47, 273–279. [Google Scholar]
- Li, C.T.; Shi, C.H.; Wu, J.G.; Xu, H.M.; Zhang, H.Z.; Ren, Y.L. Methods of developing core collections based on the predicted genotypic value of rice (Oryza sativa L.). Theor. Appl. Genet. 2004, 108, 1172–1176. [Google Scholar] [CrossRef]
- Xu, H.; Qiu, Y.; Hu, J.; Wang, J. Mothods of constructing core collection of crop germplasm by comparing different genetic distances, cluster methods and sampling strategies. Acta Agron. Sin. 2004, 30, 932–936. [Google Scholar]
- Schoen, D.J.; Brown, A.H.D. Conservation of allelic richness in wild crop relatives is aided by assessment of genetic markers. Proc. Natl. Acad. Sci. USA 1993, 90, 10623–10627. [Google Scholar] [CrossRef]
- Schoen, D.J.; Brown, A.H.D. Maximising genetic diversity in core collections of wild relatives of crop species. In Core Collections of Plant Genetic Resources; Hodgkin, T., Bronw, A.H.D., van Hintum, T.J.L., Morales, E.A.V., Eds.; Wiley: New York, NY, USA, 1995; pp. 55–76. [Google Scholar]
- Gouesnard, B.; Bataillon, T.M.; Decoux, G.; Rozale, C.; Schoen, D.J.; David, J.L. MSTRAT: An algorithm for building germplasm core collections by maximizing allelic or phenotypic richness. J. Hered. 2001, 92, 93–94. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.-W.; Chung, H.-K.; Cho, G.-T.; Ma, K.-H.; Chandrabalan, D.; Gwag, J.-G.; Kim, T.-S.; Cho, E.-G.; Park, Y. PowerCore: A program applying the advanced M strategy with a heuristic search for establishing core sets. Bioinformatics 2007, 23, 2155–2162. [Google Scholar] [CrossRef]
- Cipriani, G.; Spadotto, A.; Jurman, I.; Di Gaspero, G.; Crespan, M.; Meneghetti, S.; Frare, E.; Vignani, R.; Cresti, M.; Morgante, M. The SSR-Based Molecular Profile of 1005 Grapevine (Vitis vinifera L.) Accessions Uncovers New Synonymy and Parentages, and Reveals a Large Admixture amongst Varieties of Different Geographic Origin. Theor. Appl. Genet. 2010, 121, 1569–1585. [Google Scholar] [CrossRef]
- De Beukelaer, H.; Smykal, P.; Davenport, G.F.; Fack, V. Core Hunter II: Fast core subset selection based on multiple genetic diversity measures using Mixed Replica search. BMC Bioinform. 2012, 13, 312. [Google Scholar] [CrossRef] [PubMed]
- De Beukelaer, H.; Davenport, G.F.; Fack, V. Core Hunter 3: Flexible core subset selection. BMC Bioinform. 2018, 19, 203. [Google Scholar] [CrossRef]
- Chen, R.; Hara, T.; Ohsawa, R.; Yoshioka, Y. Analysis of genetic diversity of rapeseed genetic resources in Japan and core collection construction. Breed. Sci. 2017, 67, 239–247. [Google Scholar] [CrossRef]
- Yoneazwa, K.; Nomura, T.; Morish, H. Sampling strategies for use in stratified germplasm collection. In Core Collection of Plant Genetic Resources; Hodgkin, T., Brown, A.H.D., van Hintum, T.H.L., Eds.; John Wily & Sons: Chichester, UK, 1995; pp. 35–53. [Google Scholar]
- Qiu, L.J.; Xing, L.L.; Guo, Y.; Wang, J.; Jackson, S.A.; Chang, R.-Z. A platform for soybean molecular breeding: The utilization of core collections for food security. Plant Mol. Biol. 2013, 83, 41–50. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; Hou, L.; Zhang, Z.; Pang, X.; Li, Y. Genetic Diversity, Population Structure, and Linkage Disequilibrium of a Core Collection of Ziziphus jujuba Assessed with Genome-wide SNPs Developed by Genotyping-by-sequencing and SSR Markers. Front. Plant Sci. 2017, 8, 575. [Google Scholar] [CrossRef]
- Lu, B.Y.; Ma, M.L.; Zhang, W.; Meng, H.L.; Lei, E.; Wang, T.; Li, C. Development of 23 novel microsatellite markers of Amomum tsao-ko (Zingiberaceae) based on restriction-site-associated DNA sequencing. Mol. Biol. Rep. 2021, 48, 1943–1949. [Google Scholar] [CrossRef] [PubMed]
- Li, S.G.; Shen, D.; Liu, B.; Qiu, Y.; Zhang, X.H.; Zhang, Z.H.; Wang, H.P.; Li, X.X. Development and Application of Cucumber InDel Markers Based on Genome Re-sequencing. J. Plant Genet. Resour. 2013, 14, 278–283. [Google Scholar]
- Guo, G.J.; Zhang, G.L.; Pan, B.G.; Diao, W.P.; Liu, J.B.; Ge, W.; Gao, C.; Zhang, Y.; Jiang, C.; Wang, S. Development and Application of InDel Markers for Capsicum spp. Based on Whole-Genome Re-Sequencing. Sci. Rep. 2019, 9, 3691–3794. [Google Scholar] [CrossRef]
- Meng, L.; Liu, B.; Lin, L.B.; Cheng, F.; Wang, X.W.; Wu, J. Development of InDel Markers for Brassica campestris and Genetic Linkage Map Construction of the RILs Population. Acta Hortic. Sin. 2012, 8, 1491–1500. [Google Scholar]
- Yu, X.; Wang, H.; Zhong, W.L.; Bai, J.J.; Liu, P.L. QTL mapping of leafy heads by genome resequencing in the RIL population of Brassica rapa. PLoS ONE 2013, 8, e76059. [Google Scholar] [CrossRef] [PubMed]
- Song, X.; Ge, T.; Li, Y.; Hou, X. Genome-wide identification of SSR and SNP markers from the non-heading Chinese cabbage for comparative genomic analyses. BMC Genom. 2015, 16, 328–346. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.; Park, Y.; Lee, J.; Sim, S.C. Development of molecular markers for Ty-2 and Ty-3 selection in tomato breeding. Sci. Hortic. 2020, 265, 109230. [Google Scholar] [CrossRef]
- Qing, D.J.; Dai, G.X.; Zhou, W.Y.; Huang, S.S.; Liang, H.F.; Gao, L.; Gao, J.; Huang, J.; Zhou, M.; Chen, R.; et al. Development of molecular marker and introgression of Bph3 into elite rice cultivars by marker-assisted selection. Breed. Sci. 2019, 69, 40–46. [Google Scholar] [CrossRef]
- Kroc, M.; Czepiel, K.; Wilczura, P.; Mokrzycka, M.; Święcicki, W. Development and Validation of a Gene-Targeted dCAPS Marker for Marker-Assisted Selection of Low-Alkaloid Content in Seeds of Narrow-Leafed Lupin (Lupinus angustifolius L.). Genes 2019, 10, 428. [Google Scholar] [CrossRef]
- Liu, S.L.; Gu, X.F.; Miao, H.; Wang, Y.; Weng, Y.Q.; Wehner, T.C.; Zhang, S.P. Molecular Mapping and Candidate Gene Analysis of Black Fruit Spine Gene in Cucumber (Cucumis sativus L.). Sci. Agric. Sin. 2014, 47, 122–132. [Google Scholar]
- Wang, M.; Gu, X.F.; Miao, H.; Liu, S.L.; Wang, Y.; Todd C, W.; Zhang, S.P. Molecular Mapping and Candidate Gene Analysis for Heavy Netting Gene (H) of Mature Fruit of Cucumber (Cucumis sativus L.). Sci. Agric. Sin. 2014, 47, 1550–1557. [Google Scholar]
- Fu, D.; Zhong, K.Z.; Zhong, Z.Z.; Hu, G.C.; Zhang, P.; Tong, H. Genome-Wide Association Study of Sheath Blight Resistance within a Core Collection of Rice (Oryza sativa L.). Agronomy 2022, 12, 1493. [Google Scholar] [CrossRef]
- Xie, X.F.; Zheng, Y.; Lu, L.B.; Yuan, J.Z.; Hu, J.; Bu, S.; Lin, Y.; Liu, Y.; Guan, H.; Wu, W. Genome-Wide Association Study of QTLs Conferring Resistance to Bacterial Leaf Streak in Rice. Plants 2021, 10, 2039. [Google Scholar] [CrossRef]
- Coan, M.M.D.; Senhorinho, H.J.C.; Pinto, R.J.B.; Scapim, C.A.; Tessmann, D.J.; Williams, W.P.; Warburton, M.L. Genome-Wide Association Study of Resistance to Ear Rot by in a Tropical Field Maize and Popcorn Core Collection. Crop Sci. 2018, 58, 564–578. [Google Scholar] [CrossRef]
- Halder, J.; Zhang, J.F.; Ali, S.; Sidhu, J.S.; Gill, H.S.; Talukder, S.K.; Kleinjan, J.; Turnipseed, B.; Sehgal, S.K. Mining and genomic characterization of resistance to tan spot, Stagonospora nodorum blotch (SNB), and Fusarium head blight in Watkins core collection of wheat landraces. BMC Plant Biol. 2019, 19, 480. [Google Scholar] [CrossRef] [PubMed]
- Sapkota, S.; Hao, Y.F.; Johnson, J.; Buck, J.; Aoun, M.; Mergoum, M. Genome-Wide Association Study of a Worldwide Collection of Wheat Genotypes Reveals Novel Quantitative Trait Loci for Leaf Rust Resistance. Plant Genome 2019, 12, 190033. [Google Scholar] [CrossRef] [PubMed]
- Silvar, C.; Casas, A.M.; Kopahnke, D.; Habekuß, A.; Schweizer, G.; Gracia, M.P.; Lasa, J.M.; Ciudad, F.J.; Molina-Cano, J.L.; Igartua, E.; et al. Screening the Spanish Barley Core Collection for disease resistance. Plant Breed. 2010, 129, 45–52. [Google Scholar] [CrossRef]
- Cuevas, H.E.; Prom, L.K. Evaluation of genetic diversity, agronomic traits, and anthracnose resistance in the NPGS Sudan Sorghum Core collection. BMC Genom. 2020, 21, 88. [Google Scholar] [CrossRef]
- Sharma, R.; Upadhyaya, H.D.; Manjunatha, S.V.; Rao, V.P.; Thakur, R.P. Resistance to Foliar Diseases in a Mini-Core Collection of Sorghum Germplasm. Plant Dis. 2012, 96, 1629–1633. [Google Scholar] [CrossRef]
- Binagwa, P.H.; Traore, S.M.; Egnin, M.; Bernard, G.C.; Ritte, I.; Mortley, D.; Kamfwa, K.; He, G.; Bonsi, C. Genome-Wide Identification of Powdery Mildew Resistance in Common Bean (Phaseolus vulgaris L.). Front. Genet. 2021, 12, 673069. [Google Scholar] [CrossRef]
- Shi, A.; Gepts, P.; Song, Q.; Xiong, H.; Michaels, T.E.; Chen, S. Genome-Wide Association Study and Genomic Prediction for Soybean Cyst Nematode Resistance in USDA Common Bean (Phaseolus vulgaris) Core Collection. Front. Plant Sci. 2021, 12, 624156. [Google Scholar] [CrossRef]
- Zia, B.; Shi, A.; Olaoye, D.; Xiong, H.Z.; Ravelombola, W.; Gepts, P.; Schwartz, H.F.; Brick, M.A.; Otto, K.; Ogg, B.; et al. Genome-Wide Association Study and Genomic Prediction for Bacterial Wilt Resistance in Common Bean (Phaseolus vulgaris) Core Collection. Front. Genet. 2022, 13, 853114. [Google Scholar] [CrossRef]
- Pandey, A.K.; Yee, M.; Win, M.M.; Moh, L.H.M.; Adapala, G.; Rathore, A.; Sheu, Z.-M.; Nair, R.M. Identification of new sources of resistance to dry root rot caused by Macrophomina phaseolina isolates from India and Myanmar in a mungbean mini-core collection. Crop Prot. 2021, 143, 105569. [Google Scholar] [CrossRef]
- Togola, A.; Boukar, O.; Servent, A.; Chamarthi, S.; Tamò, M.; Fatokun, C. Identification of sources of resistance in cowpea mini core accessions to Aphis craccivora Koch (Homoptera: Aphididae) and their biochemical characterization. Euphytica Neth. J. Plant Breed. 2020, 216, 88. [Google Scholar] [CrossRef]
- Sharma, M.; Rathore, A.; Mangala, U.N.; Ghosh, R.; Sharma, S.; Upadhyay, H.; Pande, S. New sources of resistance to Fusarium wilt and sterility mosaic disease in a mini-core collection of pigeonpea germplasm. Eur. J. Plant Pathol. 2012, 133, 707–714. [Google Scholar] [CrossRef][Green Version]
- Das, S.; Porter, L.D.; Ma, Y.; Coyne, C.J.; Chaves-Cordoba, B.; Naidu, R.A. Resistance in lentil (Lens culinaris) genetic resources to the pea aphid (Acyrthosiphon pisum). Entomol. Exp. Appl. 2022, 170, 755–769. [Google Scholar] [CrossRef]
- Achola, E.; Wasswa, P.; Fonceka, D.; Clevenger, J.P.; Bajaj, P.; Ozias-Akins, P.; Rami, J.-F.; Deom, C.M.; Hoisington, D.A.; Edema, R.; et al. Genome-wide association studies reveal novel loci for resistance to groundnut rosette disease in the African core groundnut collection. Theor. Appl. Genet. 2023, 136, 35. [Google Scholar] [CrossRef]
- Ding, Y.B.; Qiu, X.K.; Luo, H.Y.; Huang, L.; Guo, J.B.; Yu, B.; Sudini, H.; Pandey, M.; Kang, Y.; Liu, N.; et al. Comprehensive evaluation of Chinese peanut mini-mini core collection and QTL mapping for aflatoxin resistance. BMC Plant Biol. 2022, 22, 207. [Google Scholar] [CrossRef] [PubMed]
- Chen, B.; Shi, R.K.; Wang, Z.C.; Liu, S.; Li, Q.; Liu, Z.; Wang, G.; Wu, J.; Ma, S. Identification of Verticillium Wilt Resistance of Core Collection of Upland Cotton and Screening of Elite Resistant Germplasm. J. Agric. Sci. Technol. 2021, 23, 45–51. (In Chinese) [Google Scholar]
- You, F.M.; Rashid, K.Y.; Zheng, C.; Khan, N.; Li, P.; Xiao, J.; He, L.; Yao, Z.; Cloutier, S. Insights into the Genetic Architecture and Genomic Prediction of Powdery Mildew Resistance in Flax (Linum usitatissimum L.). Int. J. Mol. Sci. 2022, 23, 4960. [Google Scholar] [CrossRef] [PubMed]
- Kumar, V.; Paillard, S.; Fopa-Fomeju, B.; Falentin, C.; Deniot, G.; Baron, C.; Vallée, P.; Manzanares-Dauleux, M.J.; Delourme, R. Multi-year linkage and association mapping confirm the high number of genomic regions involved in oilseed rape quantitative resistance to blackleg. Theor. Appl. Genet. 2018, 131, 1627–1643. [Google Scholar] [CrossRef] [PubMed]
- McClure, K.A.; Gardner, K.M.; Douglas, G.M.; Song, J.; Forney, C.F.; DeLong, J.; Fan, L.; Du, L.; Toivonen, P.M.; Somers, D.J.; et al. A Genome-Wide Association Study of Apple Quality and Scab Resistance. Plant Genome 2018, 11, 170075. [Google Scholar] [CrossRef]
- Yang, X.P.; Liu, G.; Hou, X.L.; Xu, J.H.; Zhang, M. Association Analysis of Fusarium wilt Resistance of Core Collection of Watermelon Germplasms Based on SRAP Markers. Acta Hortic. Sin. 2013, 40, 1298–1308. (In Chinese) [Google Scholar]
- Natarajan, S.; Kim, H.T.; Thamilarasan, S.K.; Veerappan, K.; Park, J.I.; Nou, I.S. WholeGenome Re-Sequencing and Characterization of Powdery Mildew Disease-Associated Allelic Variationin Melon. PLoS ONE 2016, 11, e0157524. [Google Scholar] [CrossRef]
- Singh, K.N.; Rawat, S.; Kumar, K.; Agarwal, S.K.; Goel, S.; Jagannath, A.; Agarwal, M. Identification of significant marker-trait associations for Fusarium wilt resistance in a genetically diverse core collection of safflower using AFLP and SSR markers. J. Appl. Genet. 2022, 63, 447–462. [Google Scholar] [CrossRef]
- Wang, X.L.; Li, J.Q.; Sun, J.; Gu, S.; Wang, J.B.; Su, C.; Li, Y.; Ma, D.; Zhao, M.; Chen, W. Mining Beneficial Genes for Salt Tolerance From a Core Collection of Rice Landraces at the Seedling Stage Through Genome-Wide Association Mapping. Front. Plant Sci. 2022, 13, 847863. [Google Scholar] [CrossRef]
- Liu, S.; Zhong, H.; Meng, X.X.; Sun, H.; Li, Y.S.; Pinson, S.R.M.; Chang, S.K.C.; Peng, Z. Genome-wide association studies of ionomic and agronomic traits in USDA mini core collection of rice and comparative analyses of different mapping methods. BMC Plant Biol. 2020, 20, 441. [Google Scholar] [CrossRef]
- Zhang, P.; Zhong, K.Z.; Zhong, Z.Z.; Tong, H.H. Genome-wide association study of important agronomic traits within a core collection of rice (Oryza sativa L.). BMC Plant Biol. 2019, 19, 259. [Google Scholar] [CrossRef]
- Song, J.; Li, J.; Sun, J.; Hu, T.; Wu, A.; Liu, S.; Wang, W.; Ma, D.; Zhao, M. Genome-Wide Association Mapping for Cold Tolerance in a Core Collection of Rice (Oryza sativa L.) Landraces by Using High-Density Single Nucleotide Polymorphism Markers From Specific-Locus Amplified Fragment Sequencing. Front. Plant Sci. 2018, 9, 875. [Google Scholar] [CrossRef]
- Li, X.N.; Zhou, Y.; Bu, Y.P.; Wang, X.F.; Zhang, Y.M.; Guo, N.; Zhao, J.; Xing, H. Genome-wide association analysis for yield-related traits at the R6 stage in a Chinese soybean mini core collection. Genes Genom. 2021, 43, 897–912. [Google Scholar] [CrossRef]
- Hao, D.; Cheng, H.; Yin, Z.; Cui, S.; Zhang, D.; Wang, H.; Yu, D. Identification of single nucleotide polymorphisms and haplotypes associated with yield and yield components in soybean (Glycine max) landraces across multiple environments. Theor. Appl. Genet. 2012, 124, 447–458. [Google Scholar] [CrossRef]
- Qiu, P.C.; Zhang, W.B.; Li, C.D.; Jiang, H.W.; Liu, C.Y.; Fan, D.M.; Zeng, Q.L.; Hu, G.H.; Chen, Q.S. Genetic overlap of droughttolerance loci between germination stage and seedling stage analyzed using introgression lines in soybean. Acta Agron. Sin. 2011, 37, 477–483. [Google Scholar] [CrossRef]
- Li, Y.H.; Li, W.; Zhang, C.; Yang, L.; Chang, R.Z.; Gaut, B.S.; Qiu, L.J. Genetic diversity in domesticated soybean (Glycine max) and its wild progenitor (Glycine soja) for simple sequence repeat and single-nucleotide polymorphism loci. New Phytol. 2010, 188, 242–253. [Google Scholar] [CrossRef]
- Jiang, H.W.; Li, C.D.; Liu, C.Y.; Zhang, W.B.; Qiu, P.C.; Li, W.F.; Gao, Y.L.; Hu, G.H.; Chen, Q.S. Genotype analysis and QTL mapping for tolerance to low temperature in germination by introgression lines in soybean. Acta Agron. Sin. 2009, 35, 1268–1273. [Google Scholar] [CrossRef]
- Garcia, M.; Eckermann, P.; Haefele, S.; Satija, S.; Sznajder, B.; Timmin, A.; Baumann, U.; Wolters, P.; Mather, D.E.; Fleury, D. Genome wide association mapping of grain yield in adiverse collection of spring wheat (Triticum aestivum L.) evaluated in southern Australia. PLoS ONE 2019, 14, e0211730. [Google Scholar] [CrossRef]
- Ma, Z.Y.; He, S.P.; Wang, X.F.; Sun, J.L.; Zhang, Y.; Zhang, G.; Wu, L.; Li, Z.; Liu, Z.; Sun, G.; et al. Resequencing a core collection of upland cotton identifies genomic variation and loci influencing fiber quality and yield. Nat. Genet. 2018, 50, 803–813. [Google Scholar] [CrossRef] [PubMed]
- Lin, T.; Zhu, G.; Zhang, J.; Xu, X.; Yu, Q.; Zheng, Z.; Zhang, Z.H.; Lun, Y.Y.; Li, S.; Wang, X.X.; et al. Genomic analyses provide insights into the history of tomato breeding. Nat. Genet. 2014, 46, 1220–1226. [Google Scholar] [CrossRef]
- Galpaz, N.; Gonda, I.; Shem-Tov, D.; Barad, O.; Tzuri, G.; Lev, S.; Fei, Z.; Xu, Y.; Mao, L.; Jiao, C.; et al. Deciphering genetic factors that determine melon fruit-quality traits using RNA-Seq-based high-resolution QTL and eQTL mapping. Plant J. 2018, 94, 169–191. [Google Scholar] [CrossRef]
- Nathanael, F.; Andres, G.; Mohit, V.; Michael, P.; Anna, H.; Collins, K.; Niranjan, B. Genome-wide association mapping identifies markers associated with cane yield components and sucrose traits in the Louisiana sugarcane core collection. Genomics 2019, 111, 1794–1801. [Google Scholar]
- Jiao, Y.; Wang, Y.; Xue, D.; Wang, J.; Yan, M.; Liu, G.; Dong, G.; Zeng, D.; Lu, Z.; Zhu, X.; et al. Regulation of OsSPL14 by OsmiR156 defines ideal plant architecture in rice. Nat. Genet. 2010, 42, 541–544. [Google Scholar] [CrossRef]
- Jin, J.; Huang, W.; Gao, J.P.; Yang, J.; Shi, M.; Zhu, M.Z.; Luo, D.; Lin, H.X. Genetic control of rice plant architecture under domestication. Nat. Genet. 2008, 40, 1365–1369. [Google Scholar] [CrossRef]
- Tan, L.; Li, X.; Liu, F.; Sun, X.; Li, C.; Zhu, Z.; Fu, Y.; Cai, H.; Wang, X.; Xie, D.; et al. Control of a key transition from prostrate to erect growth in rice domestication. Nat. Genet. 2008, 40, 1360–1364. [Google Scholar] [CrossRef]
- Wei, X.; Xu, J.; Guo, H.; Jiang, L.; Chen, S.; Yu, C.; Zhou, Z.; Hu, P.; Zhai, H.; Wan, J. DTH8 suppresses flowering in rice, influencing plant height and yield potential simultaneously. Plant Physiol. 2010, 153, 1747–1758. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, N.; Shenton, M.; Kawahara, Y.; Kumagai, M.; Sakai, H.; Kanamori, H.; Yonemaru, J.; Fukuoka, S.; Sugimoto, K.; Ishimoto, M.; et al. Whole-Genome Sequencing of the NARO World Rice Core Collection (WRC) as the Basis for Diversity and Association Studies. Plant Cell Physiol. 2020, 61, 922–932. [Google Scholar] [CrossRef]
- Qi, J.J.; Liu, X.; Shen, D.; Miao, H.; Xie, B.Y.; Li, X.X.; Zeng, P.; Wang, S.; Shang, Y.; Gu, X.F.; et al. A genomic variation map provides insights into the genetic basis of cucumber domestication and diversity. Nat. Genet. 2013, 45, 1510–1515. [Google Scholar] [CrossRef] [PubMed]
- Guo, S.G.; Zhang, J.G.; Sun, H.H.; Salse, J.; Lucas, W.J.; Zhang, H.Y.; Zheng, Y.; Mao, L.Y.; Ren, Y.; Wang, Z.W.; et al. The draft genome of watermelon (Citrullus lanatus) and resequencing of 20 diverse accessions. Nat. Genet. 2013, 45, 51–58. [Google Scholar] [CrossRef] [PubMed]

| Species Category | Name | |
|---|---|---|
| Grain crops | Cereals | maize [21,22,23,24], sorghum [25], coix [26], hulless barley [27], rice [28], wheat [9,29], oat [30], buckwheat [31], pearl millet [32], foxtail millet [33], peanut [34] |
| Potatos | sweet potato [35], cassava [36] | |
| Pulses | chickpea [37], Pigeonpea [38], lima bean [39], soybean [40,41], rice bean [42], commom bean [43], faba bean [44], mung bean [45] | |
| Horticultural crops | Vegetables | cauliflower [46], rapeseed [47], Cabbage [48], tomato [49,50], spinach [51], amaranth [52], bitter gourd [53], Jerusalem artichoke [54], yam [55], cucumber [56], pumpkin [57], white gourd [58], pepper [59], sweet pepper [60], eggplant [61], radish [62], Turnip [63], oyster mushroom [64], perilla [65], Pyropia haitanensis [66] |
| Fruits | pricot [67,68,69,70], pear [71,72], jujube [73,74,75], grape [76], melon [77], watermelon [78], kiwifruit [79], pomegranate [80], litchi [81,82], olive [83], apple [84,85,86], peach [87], cherimoya [88], fig [89], sweet cherry [90], pomelo [91], persimmon [92], sugarcane [93] | |
| Ornamental plants | Cymbidium ensifolium [94], Chrysanthemum morifolium [95,96], Prunus mume [97], Chimonanthus praecox [98], Rosa rugosa [99], Lilium brownii [100], Paeonia suffruticosa [101], Lagerstroemia indica [102], Helianthus annuus [103], Sophora moorcroftiana [104] | |
| Herbs | Fallopia multiflora [105], Astragalus [106], Scutellaria baicalensis [107], Angelica biserrata [108], Glycyrrhiza [1], Cornus officinalis [109], Dalbergia Odorifera [110] | |
| Spice | Santalum album [20] | |
| Teas | Guizhou tea [111], Chinese tea [112,113] | |
| Beverages | Coffee [114], Theobroma cacao [115] | |
| Fibers | cotton [116], upland cotton [117], island cotton [118], ramie [119] | |
| Oilseeds | safflower [120], sesame [121] | |
| Forages | Buchloe dactyloides [13], Cynodon [14], Medicago truncatula [122], Bromus inermis [15] | |
| Trees | Catalpa bungei [123], Catalpa fargesii [124], Saccharum spontaneum [125], Populus deltoides [126], Populus tomentosa [18], Cinnamomum camphora [127], Phoebe bournei [128], Robinia pseudoacacia [17], Torreya grandis [129], Tetracentron sinense [130], Xanthoceras sorbifolia [131], schima superba [132], Sapium sebiferum [133], Fraxinus chinensis [134], Eucommia ulmoides [135], Saccharum arundinaceum [136], Corylus avellana [137], Juglans regia [138,139], Betula platyphylla [140], Betula luminifera [141], Sinojackia huangmeiensis [142], Castanopsis hystrix [143], Morus alba [144], Castanea mollissima [145], Castanea sativa [146], Cunninghamia lanceolata [16], Cryptomeria japonica [147], Eucalyptus cloeziana [148], Eucalyptus urophylla [149], Ceratonia siliqua [150], Argania spinosa [151], Pinus massoniana [19], Pinus yunnanensis [152], Ginkgo biloba [153], Akebia trifoliata [154], Camellia oleifera [155], Cornus wilsoniana [156] | |
| Core Collection | No. of Sample | Type of Disease | No. of QTLs/Genes/ Resistant Samples | Reference |
|---|---|---|---|---|
| rice | 150 | sheath blight | 13 genes | [188] |
| rice | 510 | bacterial leaf streak | 69 QTLs | [189] |
| maize | 183 | fusarium ear rots | 4 genes | [190] |
| wheat | 121 | tan spot Pt race 1 | 10 QTLs | [191] |
| tan spot Pt race 5 | 5 QTLs | |||
| stagonospora nodorum blotch | 5 QTLs | |||
| wheat | 331 | leaf rust | 11 QTLs | [192] |
| barley | 159 | powdery mildew | 58 resistant accessions | [193] |
| leaf rust | 42 resistant accessions | |||
| net blotch | 2 resistant accessions | |||
| mild mosaic virus | 13 resistant accessions | |||
| yellow dwarf virus | 32 resistant accessions | |||
| sorghum | 318 | anthracnose | 4 QTLs | [194] |
| sorghum | 242 | anthracnose | 13 resistant accessions | [195] |
| leaf blight | 27 resistant accessions | |||
| leaf rust | 6 resistant accessions | |||
| anthracnose, leaf blight, leaf rust | 3 resistant accessions | |||
| common bean | 211 | powdery mildew | 4 genes | [196] |
| common bean | 315 | cyst nematode HG Type 0 | 11 SNPs | [197] |
| common bean | 168 | bacterial wilt | 14 genes | [198] |
| mungbean | 296 | dry root rot | 29 resistant accessions | [199] |
| cowpea | 375 | Aphis craccivora Koch pest | 3 resistant accessions | [200] |
| pigeonpea | 146 | fusarium wilt | 6 resistant accessions | [201] |
| sterility mosaic | 24 resistant accessions | |||
| fusarium wilt and sterility mosaic | 5 resistant accessions | |||
| lentils | 188 | pea aphid | 14 genes and 13 SNPs | [202] |
| groundnut | 213 | groundnut rosette | 32 SNPs | [203] |
| peanut | 99 | aflatoxin | 16 SNPs | [204] |
| upland cotton | 419 | verticillium wilt | 12 resistant accessions | [205] |
| flax | 447 | powdery mildew | 1 gene and 3 QTLs | [206] |
| oilseed rape | 166 | blackleg | 8 QTLs | [207] |
| tomato | 171 | yellow leaf curl virus | 2 genes | [183] |
| apple | 176 | apple scab | 10 genes | [208] |
| watermelon | 35 | fusarium wilt | 1 SRAPs | [209] |
| melon | 4 | powdery mildew | 112 SNPs and 12 InDels | [210] |
| safflower | 84 | fusarium wilt | 3 AFLPs and 1 SSRs | [211] |
| Core Collection | No. of Sample | Trait | No. of QTLs/Genes | Reference |
|---|---|---|---|---|
| rice | 150 | salt tolerance | 65 QTLs | [212] |
| rice | 191 | yield and heavy metal content | 250 QTLs | [213] |
| agronomic traits | 97 QTLs | |||
| rice | 150 | agronomic | 32 QTLs | [214] |
| rice | 150 | cold tolerance | 26 QTLs | [215] |
| soybean | 224 | agronomic | 16 QTLs | [216] |
| soybean | 189 | yield and yield component | 19 QTLs | [217] |
| soybean | 23 | salt tolerance | 22 QTLs | [218] |
| low temperature tolerance | 15 QTLs | |||
| soybean | 159 | high oil content | 6 QTLs | [219] |
| high protein content | 1 QTLs | |||
| drought tolerance | 5 QTLs | |||
| soybean | 46 | low temperature resistant | 13 QTLs | [220] |
| wheat | 568 | yield and yield components | 17 QTLs | [221] |
| upland cotton | 419 | fiber-related | 5 genes | [222] |
| leaf heads of cabbage | 150 | agronomic | 18 QTLs | [181] |
| tomato | 360 | agronomic | 2 QTLs | [223] |
| melon | 79 | fruit quality | 241 QTLs | [224] |
| sugarcane | 97 | yield components and sucrose | 56 QTLs | [225] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gu, R.; Fan, S.; Wei, S.; Li, J.; Zheng, S.; Liu, G. Developments on Core Collections of Plant Genetic Resources: Do We Know Enough? Forests 2023, 14, 926. https://doi.org/10.3390/f14050926
Gu R, Fan S, Wei S, Li J, Zheng S, Liu G. Developments on Core Collections of Plant Genetic Resources: Do We Know Enough? Forests. 2023; 14(5):926. https://doi.org/10.3390/f14050926
Chicago/Turabian StyleGu, Rui, Shaohui Fan, Songpo Wei, Jiarui Li, Shihui Zheng, and Guanglu Liu. 2023. "Developments on Core Collections of Plant Genetic Resources: Do We Know Enough?" Forests 14, no. 5: 926. https://doi.org/10.3390/f14050926
APA StyleGu, R., Fan, S., Wei, S., Li, J., Zheng, S., & Liu, G. (2023). Developments on Core Collections of Plant Genetic Resources: Do We Know Enough? Forests, 14(5), 926. https://doi.org/10.3390/f14050926






