Abstract
Understanding the capacity of forest tree species to adapt to climate change is of increasing importance for managing forest genetic resources. Through a genomics approach, we modelled spatial variation in climate adaptation within the Australian temperate forest tree Eucalyptus globulus, identified putative climate drivers of this genomic variation, and predicted locations of future climate refugia and populations at-risk of future maladaptation. Using 812,158 SNPs across 130 individuals from 30 populations (i.e., localities) spanning the species’ natural range, a gradientForest algorithm found 1177 SNPs associated with locality variation in home-site climate (climate-SNPs), putatively linking them to climate adaptation. Very few climate-SNPs were associated with population-level variation in drought susceptibility, signalling the multi-faceted nature and complexity of climate adaptation. Redundancy analysis (RDA) showed 24% of the climate-SNP variation could be explained by annual precipitation, isothermality, and maximum temperature of the warmest month. Spatial predictions of the RDA climate vectors associated with climate-SNPs allowed mapping of genomically informed climate selective surfaces across the species’ range under contemporary and projected future climates. These surfaces suggest over 50% of the current distribution of E. globulus will be outside the modelled adaptive range by 2070 and at risk of climate maladaptation. Such surfaces present a new integrated approach for natural resource managers to capture adaptive genetic variation and plan translocations in the face of climate change.
1. Introduction
Understanding the patterns of local adaptation to climate variation that occurs within tree species is important for managing our forests and forest genetic resources in the face of rapid climate change [1,2,3]. Increasingly, genomic approaches are being used to study such adaptations in plants [4,5]. One commonly applied approach focuses on dissection of the genetic architecture of functional traits thought to be responsible for adaptation. This involves the identification of quantitative trait loci (QTL) through either linkage mapping or genome wide association studies GWAS [6,7]; usually, these are based on phenotypic data derived from pedigreed families growing in common garden field trials [8]. Another approach, used predominantly for wild populations, involves the identification of DNA markers that exhibit signals of divergent selection. These signals include extreme differentiation in allele frequencies between populations/ecotypes [7,9], or significant genotype–environment association (GEA) [10,11]. These various approaches have advantages, disadvantages, and requirements [12,13]. For example, studies of trait-based adaptation depend on the strength of evidence for the direct fitness consequences of a trait [14]. In contrast, DNA marker-based outlier and GEA approaches do not require knowledge of trait variation associated with adaptation. Outlier approaches based on the degree of marker differentiation among populations, using FST for example, may be too conservative to detect loci other than those with a large effect [15]. GEA analysis is now increasingly being used to study adaptation in non-model species, due to the potential to identify loci of both small and large effects. Depending on the specific GEA method, genetic and spatial structure can also be taken into account at the same time that population-level allele frequencies are compared to population home-site (i.e., the geographic locality of the original seed collection within the native range), environmental, or climatic data [10].
The investigation of adaptation in non-model species such as forest trees has several associated challenges. For instance, due to the large size, long generation times [16], and often large ontogenetic changes between juvenile and adult forms of many forest tree species [17], studies employing common garden approaches can often only examine early life-cycle traits [18,19]. On the other hand, forest trees frequently exhibit clinal variation along climate gradients [20,21], making them highly amenable to GEA [22,23] and applying ‘space-for-time’ approaches for prediction of climate change responses at the population level [21,24].
In Australia, environmental adaptation has been studied in several forest tree genera [25,26], with the majority of studies focusing on Eucalyptus L’Hér, reviewed by [21,27,28]. This iconic genus is renowned for the adaptive variation that resides within species [27,29,30,31,32], which is increasingly being explored with genomics [33]. While early studies of adaptation genomics have focused on identifying loci based on FST outliers [9,31,34], GEA analyses that account for geography/distance and population structure are now being implemented [35,36,37,38,39]. Both styles of analysis have implicated heat, aridity, and seasonality as major drivers of adaptive genetic variation in Eucalyptus species such as E. melliodora [36], E. microcarpa [40], E. pauciflora [18,41], and E. grandis [38], as well as in species in the related genus Corymbia [35,42].
The present study focused on the temperate Australian tree, Eucalyptus globulus Labill., a foundation tree species of south-eastern Australia that provides critical habitat and food for numerous organisms, including the endangered swift parrot [43,44]. It was also one of the first eucalypt species to be grown outside Australia and is now one of the major eucalypts grown in industrial plantations in temperate regions of the world, including Australia [45]. Given its economic and ecological importance, it is critical to understand the variation in climate adaptation and identify the components of the native gene-pool at risk of maladaptation under future climates [46,47].
We here used range-wide sampling and a genome-wide marker set to identify climate-associated single nucleotide polymorphisms (termed ‘climate SNPs’) and climate vectors associated with this putatively adaptive genomic variation. These genomic informed climate vectors were then used to model the contemporary and future climatic selective surfaces of E. globulus across its native range in order to provide a framework to guide gene-pool conservation and seed transfer decisions. We also studied whether drought adaptation was an important facet of climate adaptation.
2. Materials and Methods
2.1. Study Species
Eucalyptus globulus is native to south-eastern Australia, including the islands of Tasmania, where forests dominated by the species occur over a latitudinal range from −38.4° S to −43.5° S, and an altitudinal range from sea-level to 830 m above sea-level [48,49,50]. During the domestication of E. globulus, large common garden trials were established throughout the world from open-pollinated seed collected from mother trees in native forests [45,49,51]. Quantitative genetic studies of these trials identified significant population differentiation in numerous traits, which was summarised by partitioning the gene pool into 13 geographic races [49]. While there are likely evolutionary constraints [52], population differences have been shown to: occur at multiple geographic scales, from hundreds of metres [53,54] to hundreds of kilometres [49,55], be adaptive (i.e., significant population × environment interactions for growth in common garden trials, [32]), and even to have extended community-level consequences [56,57]. Numerous traits exhibit clines with latitude and climate as well as signals of divergent selection (QST > FST) [47,49,58,59]. The patterns of molecular diversity in the gene pool have been characterised extensively using putatively neutral molecular markers, showing significant spatial structure from family groups [60,61] to broad-scale geographic clines [50,62,63]. Despite current large contemporary disjunctions in its natural distribution (e.g., Bass Strait), the population differentiation in microsatellite markers is relatively low (FST of 0.08 with 95% CI equal to 0.074–0.092, [64]), consistent with recent post-glacial fragmentation [65,66] and ongoing long-distance pollen dispersal [63,67].
2.2. Sampling Strategy and SNP Discovery
Trees sampled for DNA were from an E. globulus provenance–progeny trial, planted in 1989 near Latrobe in Tasmania, Australia, in land owned by Forico Ltd. The trial was established with 570 open-pollinated E. globulus families collected from wild mother trees, with each family represented by a two-tree plot in each of five replicates. To ensure a wide geographic spread of collecting localities and mother trees represented in the sampling, families were sampled from the trial within the races of the Furneaux Group, Gippsland/Strzelecki Ranges, King Island, north-eastern Tasmania, south-eastern Tasmania, southern Tasmania and, western/eastern Otways as described in [49], with an approximate number of 20 trees sampled per race or race combination. Families were randomly chosen from the main collecting localities originally sampled within each race, with one tree sampled per family in the trial. The localities represented from each race, the number of trees sampled, and their latitude, longitude, and elevation are indicated in Figure 1 and detailed in Table S1. DNA was extracted from cambial scrapings as described by Tibbits, et al. [68]. This DNA was a subsample of that collected by Thavamanikumar, et al. [69]. DNA was sequenced with short reads, aligned to a draft E. globulus genome, and SNPs were called and filtered for quality (see Appendix A.1).
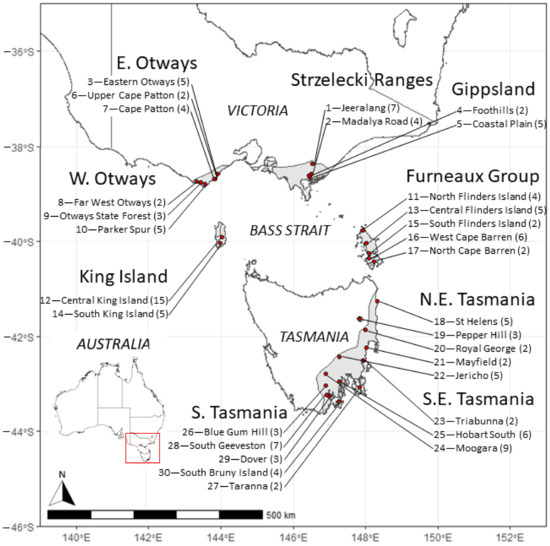
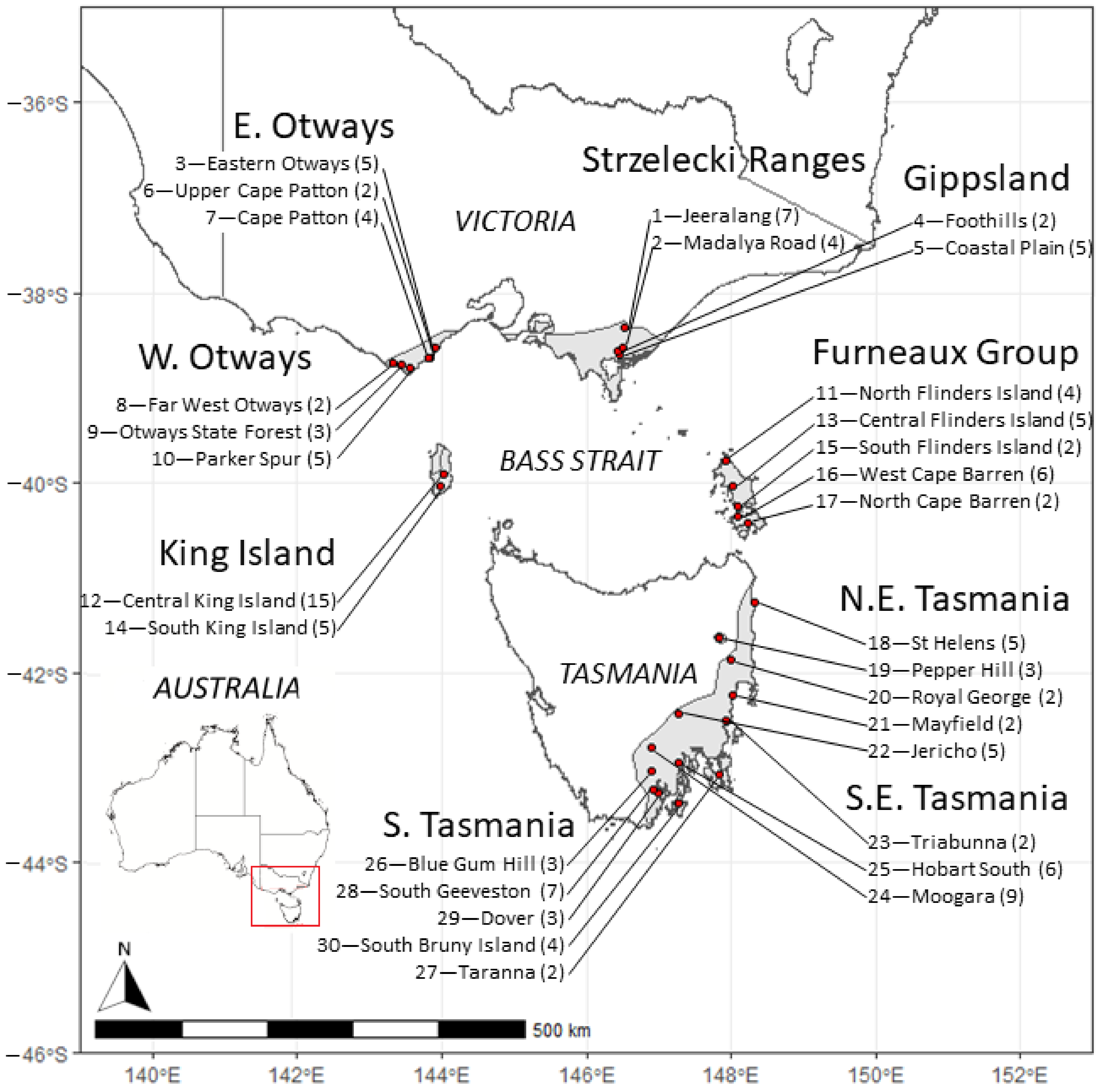
Figure 1.
Geographical distribution of Eucalyptus globulus mother trees of the open-pollinated families sampled from each race in the genetic trial (also see Table S1). The inset plot indicates the region of Australia examined in this study. The grey area indicates the natural range of E. globulus in south-eastern Australia. The small disjunct occurrences on the west coast of Tasmania are not shown, as they were not included in the present study. The race (bold) and locality (below) names follow Dutkowski and Potts [49], with the first number indicating a locality identifier, and the number in parentheses indicating the number of open-pollinated progeny/mother trees sourced from each locality used in this study. The mapped area is shown on the inserted map of Australia showing state boundaries.
2.3. GEA Analysis
As an indicator of the historic pattern of climate variation across the range of E. globulus, which would have shaped the patterns of adaptation, we obtained contemporary climate data (1979–2013) from CHELSA [70], describing 19 BIOCLIM variables [71] for temperature, rainfall, and seasonality at 30 arcsecond resolution (ca. 0.8 km). Variables were extracted for each locality using the R package ‘raster’ [72]. The gradientForest algorithm [73] was then used as a GEA method [46] to model the SNP data as a function of the climatic variables in a set of divided analyses (see Appendix A.2). The efficacy of gradientForests as an approach to detect SNPs under putative selection was established by running the gradientForest on an isolation-by-distance simulated population dataset, with SNPs with varying strengths of climate-associations as generated by Lotterhos and Whitlock [74] (Table S2). As gradientForests cannot directly accommodate spatial effects, we also calculated Moran’s eigenvector map variables (MEMs) based on geographic coordinates of the native locality from which the mother trees were originally sampled (i.e., their home-site) using the R package ‘adespatial’ [75]. These MEMs are orthogonal vectors that maximise the spatial autocorrelation between locations based on the eigenvectors of a neighbourhood network matrix, in our case based on the Gabriel connection graph. These MEMs can also be used as proxies for possible unmeasured environmental variation, as environmental and geographical variation often covary at broad spatial scales [76,77]. Of the total MEMs calculated, we included the first half of the positive MEMs (in this case seven) as spatial covariates, similar to what has been done in other studies [23,74,78,79].
SNPs identified through modelling the 19 climate variables exceeding an importance threshold of 0.15 (identified through visual analysis of the distribution of importance scores) were retained (hereafter referred to as climate-SNPs), excluding those solely important to the MEMs. This was done to select those SNPs that remained associated with climate after removing the effect of space. This set of ‘climate-SNPs’ is conceptually the same as the ‘climate-aligned adaptively enriched genetic space’ of Steane, et al. [9] or ‘environment-associated gene space’ of Mostert-O’Neill, et al. [38]. To test the effect of removing SNPs associated with spatial variables, a second gradientForest analysis was performed following the same procedure, but with no MEMs included as predictor variables. Patterns of allele frequency change across the associated climate gradients, after partialling out the effect of all other climate and spatial variables, were examined using the R package ‘extendedForest’ [73]. Distribution of climate-SNPs across chromosomes were examined using a Chi-squared test. Those climate-SNPs associated with annotated genes (within ±1000 bp) were tested for enrichment of gene ontology (GO) terms versus the original dataset (via SNP alignment against the E. grandis reference genome, [80]) using the R package ‘SNP2GO’ [81].
2.4. SNPs Associated with Drought Adaptation
To investigate the intersection between the climate-SNPs and SNPs associated with a natural climate-driven selective pressure, a partitioned extendedForest analysis (see Appendix A.3) was performed on the original SNP data using the locality-level drought damage scores obtained from four common-garden field trials established in Western Australia [47], which is outside the native range of the species. The field trials were established from families derived from open-pollinated single-tree seed collections from wild stands of E. globulus, which included most of the families that are represented in our SNP study. Dutkowski and Potts [47] assessed the percentage of leaf death on 5–6 year-old trees in 1994, following two years of below average summer rainfall. At assessment, damage was severe, with only 0 to 4% of the trees showing no damage across the four trials. The damage trait was heritable, with most of the genetic variation distributed between populations and the level of population differentiation among the highest reported for traits studied in E. globulus (QST = 0.39 ± 0.09 [47]). The race-level geographic pattern in drought damage was similar to that reported for drought mortality in field trials in Spain [82] and growth on dry sites in Australia [32]. The drought damage scores used in the present study were estimated for each sampling locality (following Dutkowski and Potts [49]), by averaging across the breeding value predictions for mothers from each locality [47]. The SNPs associated with the drought damage scores were then compared to the climate-SNPs.
2.5. Redundancy Modelling and Projection of Future Maladaptation
To summarise the influence of the climate variables on the climate-SNPs, a redundancy analysis (RDA) was performed using the R package ‘vegan’ [83]. Redundancy analysis is a form of multivariate regression that identifies a set of linear predictors that explain as much variation as possible in a set of linear response variables [84]. Prior to performing the RDA, climate variables were normalised to a unit variance. Forward selection was then used to choose the set of predictor climate variables that had the most explanatory power, removing variables with high (e.g., >3.5) variance inflation factors [85]. This approach served to verify the important climate variables identified through the gradientForest analysis and to identify the set of climate variables to use in the final RDA in which the response variables were the climate-SNPs. The MEMs originally used in the identification of the climate-SNPs were not included as predictors in this analysis, to enable the capture of the broad-scale variation in the climate-SNPs across the range of E. globulus. Each locality was fitted to the ordination space and an R2 value was calculated and corrected for multiple regressions [85]. Significant RDA axes were determined using a permutation test (1000 permutations) and retained for further analysis.
To visualise the broad-scale climate selection surface, the climate indices associated with the main directions of variation in climate-SNPs (i.e., putative adaptive variation) among localities described by the significant RDA axes were extrapolated across the native distribution of E. globulus in Tasmania and southern Victoria, using the contemporary climate data obtained from CHELSA, mentioned above (see Appendix A.4). To predict the influence of future climate on the current distribution of E. globulus, future climate projections were obtained from CHELSA for six global circulation models that covered a broad range of future climate projected values (ACCESS1-0, CSIRO-Mk3-6-0, HadGEM2-AO, MIROC5, MPI-ESM-MR, and GFDL-CM3), assuming the representative concentration pathway 8.5 (RCP8.5), a high emissions scenario [86] for the 2070s (mean of the 2061–2080 period). Each future climate layer was normalised and scaled using the mean and standard deviation from the contemporary layers, and the cell scores for each prediction were then combined and averaged to create a multi-model mean prediction [87].
Mapped grid cells falling within the range of the sampled localities on the three significant RDA axes (considered individually and in combination) were quantified for both contemporary and 2070s climate surfaces. To assess the ‘genomic vulnerability’ [23,36,88] of E. globulus populations within a grid cell to future climate change, we estimated the similarity between a grid cells’ contemporary and future (2070s) climate position in the three-dimensional RDA space using Euclidean distance. Given the anticipated low generational turnover and seed migration [61,89] of this long-lived species, areas with low Euclidean displacement were deemed to be candidates for future climate refugia, assuming the genotype–environment relationships modelled across contemporary populations are retained [88]. In addition to mapping the surface of predicted displacement (i.e., genomic vulnerability) across the geographic range of E. globulus, we calculated the percentage of occurrence records within decile bins of the predicted displacement measure for four regions: mainland Australia (Victoria), mainland Tasmania, the Furneaux Group of islands, and King Island. This was achieved using the Tasmanian distribution records for E. globulus, obtained from the Tasmanian Natural Values Atlas (http://www.naturalvaluesatlas.tas.gov.au/; accessed on 6 February 2015) and mainland Australia distribution records obtained from the Atlas of Living Australia (https://www.ala.org.au/; accessed on 5 March 2017). Spatial and environmental outlying occurrence records were identified using a modified Z-score test following Jordan, et al. [90], were manually checked against known distribution descriptions, and removed where appropriate. Focusing on the islands of Tasmania, where most records reside, we also determined the proportion of occurrence records of E. globulus that were predicted to remain within the locality ranges on all three RDA axes under the current and 2070s climate.
3. Results
3.1. gradientForest Analysis and Climate SNP Investigation
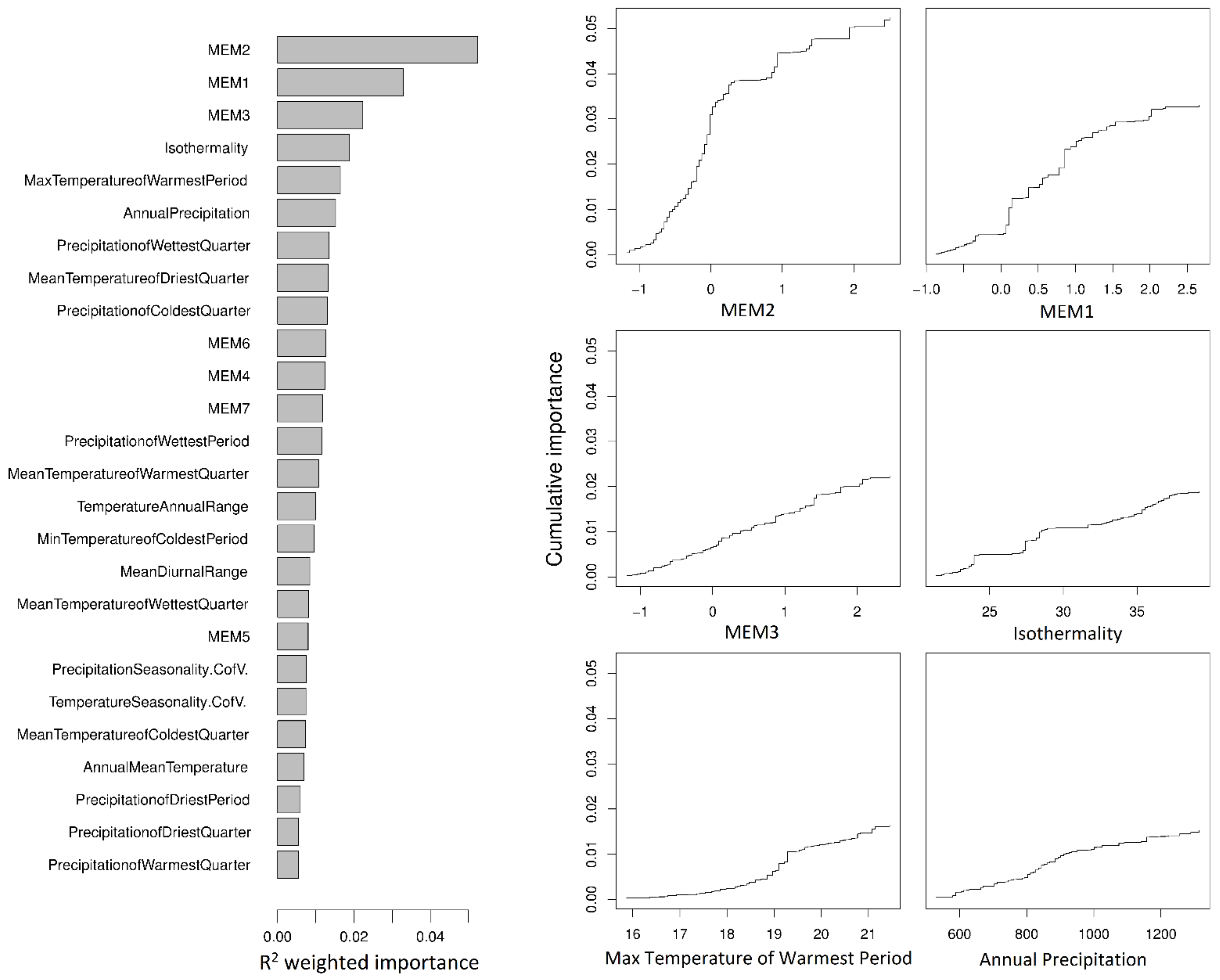
A total of 812,158 high-quality, genome-wide SNPs were called from whole genome shotgun sequence data, representing 130 individuals from 30 localities (Figure 1). Variation in SNP allele frequencies were used as response variables against 19 climate variables and 7 Moran’s eigenvector map variables (MEMs) in partitioned gradientForest analyses. These analyses identified 71,183 SNPs of positive importance, which were rerun in a final gradientForest analysis to assess their relative importance. The most important variables describing the allelic variation in SNP frequencies among the localities were MEM2, MEM1, and MEM3 (Figure 2). MEM2 explained 15% of the variation and the MEMs combined explained 44%, indicating a strong spatial component to the genetic differences among localities. After accounting for the MEM-associated variation, the most important climate variables were isothermality, maximum temperature of the warmest period (with period being one month), and annual precipitation, each explaining 4.3–5.4% of the SNP variation. While the change in cumulative importance of SNP allele frequencies was fairly rapid for MEM2 (representing rapid allelic turnover between mainland and Tasmanian localities), for all other variables, allele replacement occurred gradually or with small step-changes (Figure 2).
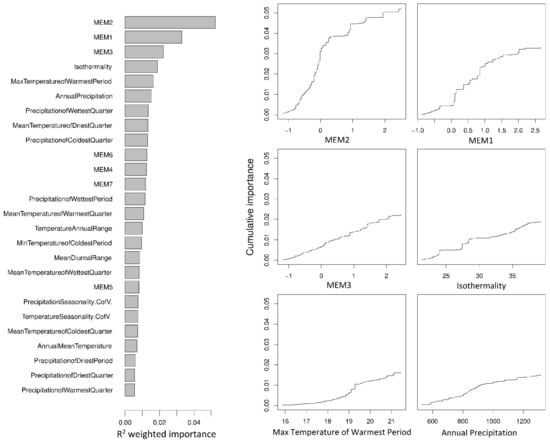
Figure 2.
Relative importance of predictor variables (R2 weighted, left) and cumulative importance of allele replacement in 71,183 SNPs across spatial (MEM1, MEM2, and MEM3) and climate gradients (right) as calculated by gradientForest. The three most important climate variables after accounting for spatial variation are shown. Values along the y axis represent change itself and have no directionality. Values along the x axis show the range of each variable. The modelled function shows how the total importance (or amount of allele replacement) associated with a variable is distributed across the range of that variable (e.g., the majority of the SNP differences associated with MEM2 are found between −1 and 0.5). MEM1 was strongly correlated with longitude (Pearson’s r = −0.84, p < 0.001), MEM2 with latitude (Pearson’s r = 0.79, p < 0.001) and MEM3 differentiated Furneaux Group island localities from all other localities.
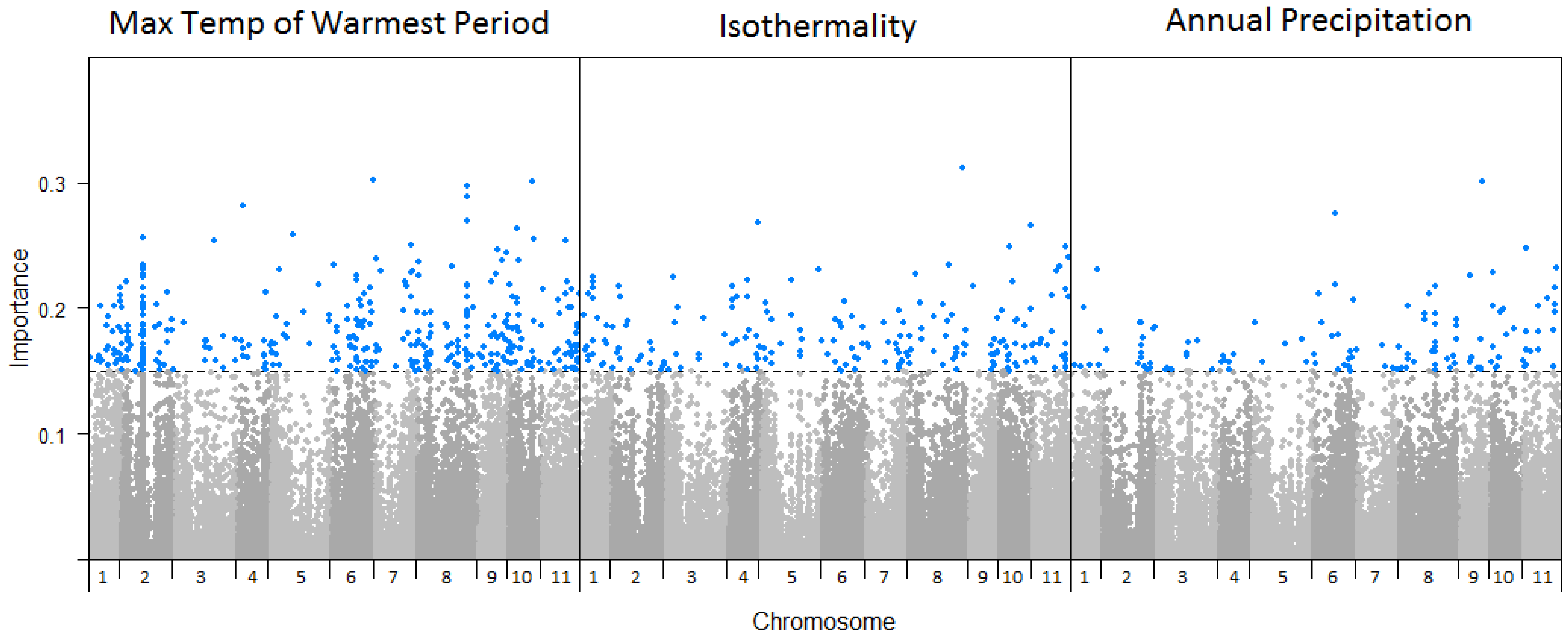
Of the 71,183 SNPs analysed, 1177 were found to exhibit importance values > 0.15 for one or more of the 19 climate variables (climate-SNPs) (Table S3), and were selected for further analysis. The climate variable associated with the greatest number of SNPs was maximum temperature of the warmest period (n = 366 climate-SNPs), followed by isothermality (n = 205 climate-SNPs), and annual precipitation (n = 126 climate-SNPs) (Figure 3). These climate-SNPs were distributed across the genome (Figure 3), although there were higher concentrations on some chromosomes for those associated with maximum temperature of the warmest period (χ210 = 89.6, p < 0.001; highest number on chromosome 2) and annual precipitation (χ210 = 35.1, p < 0.001; highest number on chromosome 8). There was no overlap in climate-SNPs between the broad categories of climate variables (i.e., precipitation, temperature, or seasonality), but some SNPs reached the importance threshold for multiple climate variables within these broad categories.
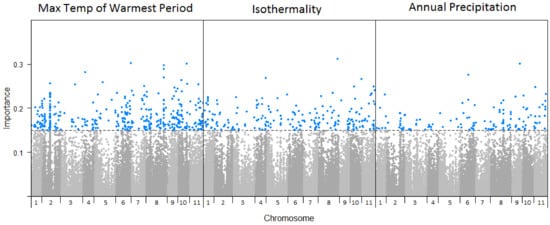
Figure 3.
Manhattan plots of SNP importance for maximum temperature of the warmest period, isothermality, and annual precipitation across the 11 Eucalyptus globulus chromosomes. Each of the 71,183 SNPs are shown (points) across the 11 chromosomes, where the blue points correspond to those associated with the climate variable above the 0.15 cumulative importance threshold.
As many of the climate variables are inherently spatially autocorrelated and covary with geography, we repeated the gradientForest analyses without MEMs as predictors. The comparison of these two analyses (i.e., including and excluding MEMs) allowed evaluation of the effect of not accounting for spatial variance on the SNPs detected or, in other words, contrasting the broad-scale associations with those fine-scale associations detected when spatial components are removed. In this case, we found 63,509 SNPs of positive importance, with an ordering of the most important climate variables (isothermality, maximum temperature of the warmest month, and annual precipitation) identical to that found when MEMs were included. Of the 63,509 SNPs, 1073 were found to have importance of >0.18 (chosen to achieve a similar proportion to the original set) with any climate variable, of which 827 (70%) were climate-SNPs previously identified in the analysis, which included MEMs. This indicates that these 827 SNPs show both fine- and broad-scale associations with one or more climate variables. However, the analysis without MEMs resulted in a higher proportion of the SNP variation being explained by the climate variables (R2 weighted importance range: 7.5–9.1%).
We used extendedForest to model the partial dependency of each climate-SNP after averaging the effect of all other climate variables and MEMs. This analysis revealed only slight changes in allele frequency in most SNPs in response to the variation in the climate variable, with allele frequency changes of 0.06–0.20 (for examples, see Figure S1). The turnover function of almost every climate-SNP was broadly monotonic, with most exhibiting a steep inflection in allele frequency often associated with a geographical feature (for example, island versus other localities in isothermality-associated SNPs). Excluding MEMs from this analysis resulted in almost identical patterns of change in allele frequency.
Gene ontology (GO) analysis revealed two GO terms enriched in genes associated with the 1177 climate-SNPs relative to the entire set, GO:0016788 (p < 0.001), which is associated with the hydrolysis of ester bonds, and its child term GO:0004518 (p < 0.001), which is linked with the specific hydrolysis of ester bonds in nucleic acids. These GO terms (and their child terms) are associated with 18 genes that contain or are in proximity to 23 climate-SNPs, with annotated functions including DNA repair endonucleases, ribonucleases, protein phosphatases, and general ester hydrolysis (Table S4). Of these 23 SNPs, 19 were important to temperature-related climate variables (with 15 important to the maximum temperature of the warmest period).
3.2. Adaptive SNPs for Drought Damage and Their Importance to Climatic Adaptation
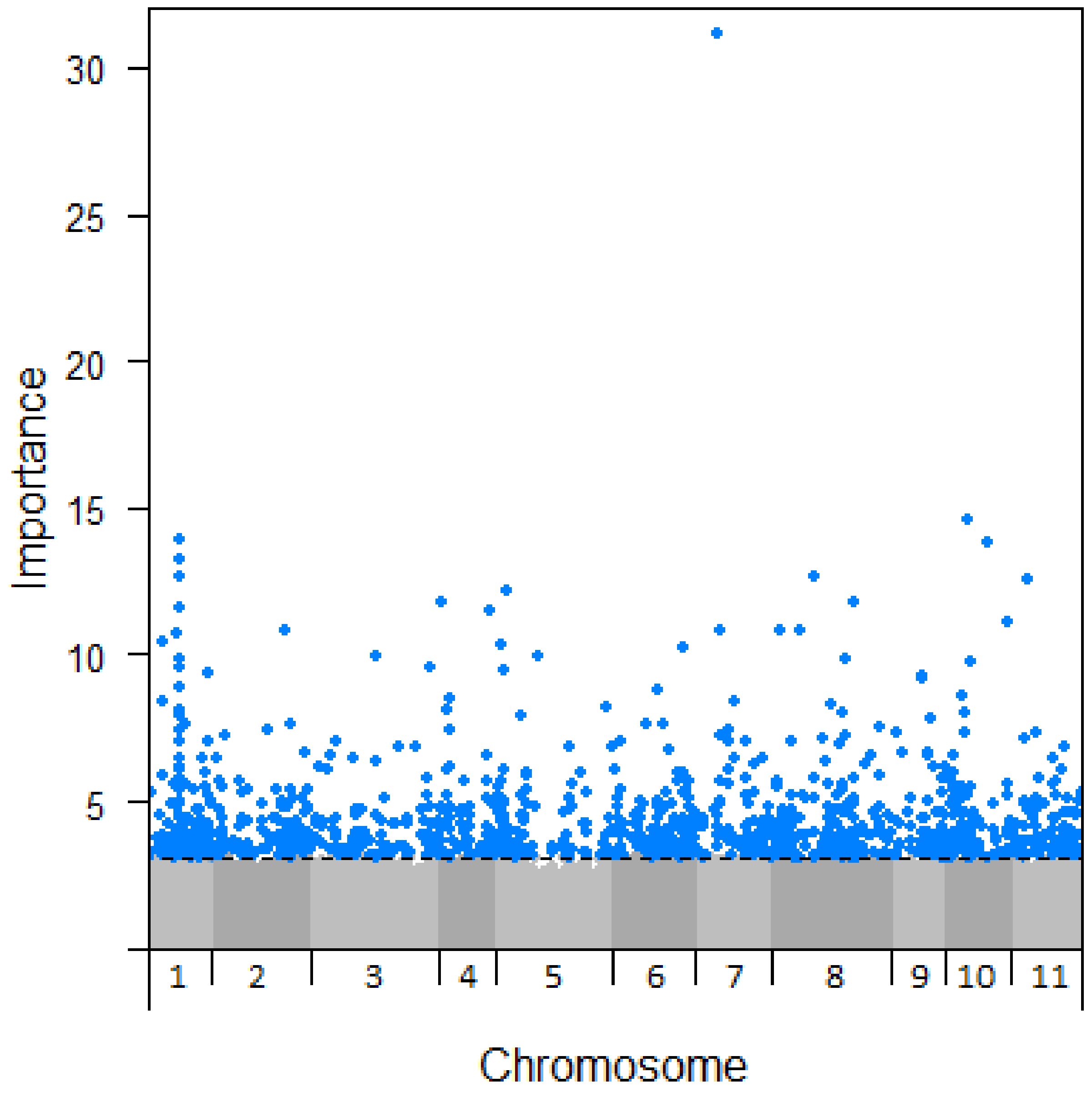
With significant genetic-based variation in drought susceptibility known to occur across the geographic range of E. globulus, we explored the extent to which the climate-SNPs identified were likely to contribute to drought adaptation. This was achieved using existing locality-level drought damage data for trees from the same seed collection used in this genomics study (see Methods). Drought damage for each locality was modelled with extendedForest using the 812,158 SNPs and MEMs as predictors, divided into 33 separate runs of 25,000 SNPs due to computational limitations. Of these, the approximately 800 most important SNPs in each run (25,000 in total) were combined and extendedForest re-run with this SNP set to rank their relative importance. To maintain a sample size comparable to the climate-SNPs, the 1192 most important SNPs (importance > 3.1) in predicting drought damage were retained (note that the ‘importance’ scale of the two analyses are not directly comparable). Repeating the procedure but excluding MEMs as predictors found 1160 SNPs with importance > 3.1, with 759 SNPs (64%) shared between the analyses with and without MEMs. Given that the two sets of SNPs were similar and the likelihood that the analysis with MEMs may have excluded relevant SNPs, we present the set of 1160 SNPs identified without using MEMs (Figure 4, Table S5). These SNPs are our best candidates for further study of drought adaptation and were distributed across all chromosomes, although the distribution was skewed (χ210 = 80.8, p < 0.001), most notably towards chromosome 1 (with 197 SNPs). The most important SNP, “G_0337_23205” (Figure 4), was also highest in importance when accounting for spatial variation. This SNP was not associated with any annotated genes in the E. grandis genome and was not identified as a climate-SNP. Gene ontology analysis did not show any terms enriched in this set of 1160 SNPs.
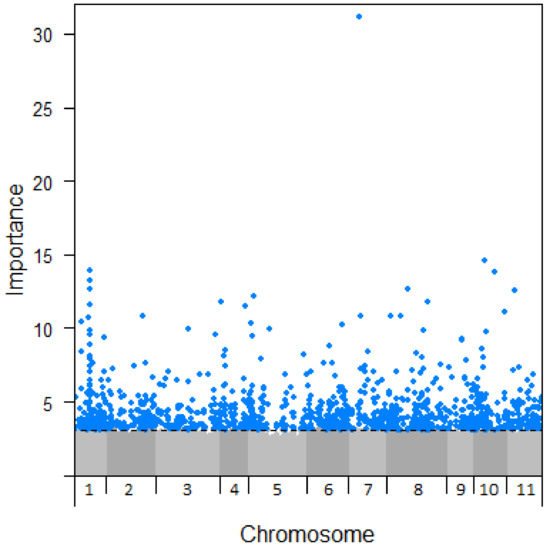
Figure 4.
Manhattan plot of SNP importance to drought damage (not accounting for spatial variance) across Eucalyptus globulus chromosomes. A threshold of importance >3.1 was set in order to capture a number of SNPs similar to that found to be important in the analysis using climate variables.
When comparing the climate-SNPs and those strongly associated with drought damage (ignoring spatial variation), only six SNPs were shared between the two sets. Three of these were associated with precipitation of the wettest quarter, and one each with precipitation of the coldest quarter, mean temperature of the wettest quarter, and maximum temperature of the warmest period (Tables S3 and S5). These six SNPs demonstrated typical patterns of partial dependency with the climate variables, with sharp inflection in allele frequency near the middle of the climatic range.
3.3. Redundancy Analysis and Climate Projection
The 1177 climate-SNPs were taken into a redundancy analysis to generate vectors describing the major axes of climate-associated SNP variation. MEMs were not included in this analysis. Using forward selection of variables, the redundancy analysis showed that most of the variation in the climate-SNPs could be explained by the three most important climate variables detected in the gradientForest model: isothermality, maximum temperature of the warmest period, and annual precipitation (Figure 5). The redundancy model based on these three climate variables was highly significant (F3,26 = 4.03, p < 0.001) and explained 23.9% of the genomic variation among localities. The first axis explained 4.0% of the overall adjusted variance in the climate-SNPs (RDA1, F1,26 = 6.44, p < 0.001) and the second axis 2.5% (RDA2, F1,26 = 4.05, p < 0.001). RDA1 was dominated by both maximum temperature of the warmest period and annual rainfall, which varied in the same direction, while RDA2 was dominated by isothermality. RDA3 was also significant (F1,26 = 1.61, p < 0.01), explained 1.0% of the overall adjusted variation in the climate-SNPs, and was again dominated by maximum temperature and annual rainfall, but in opposing directions. This axis represented an aridity gradient, which was confirmed by fitting the locality-level drought damage scores (Table S1) into the RDA ordination. RDA3 was the only SNP-aligned climate vector significantly correlated with these drought damage scores (r = 0.61, p < 0.01).
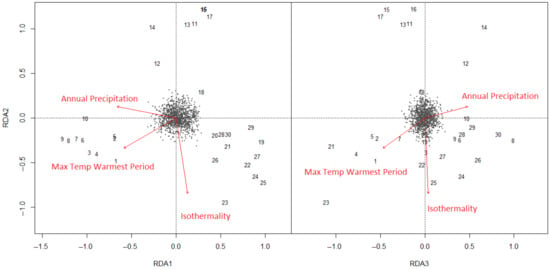
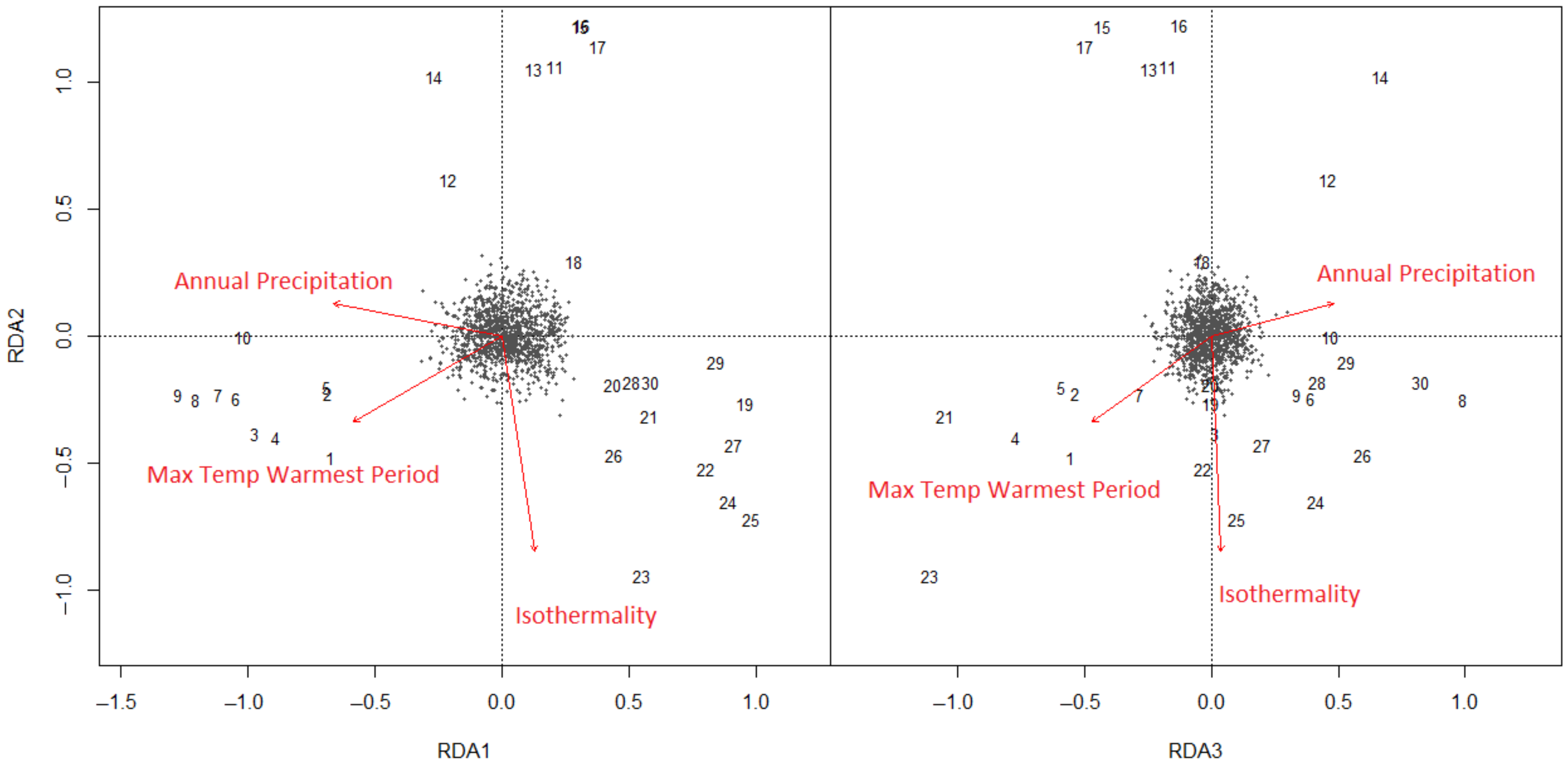
Figure 5.
Redundancy analysis of climate-SNPs in E. globulus, showing the position of climate SNPs (black points), localities (numbers referring to the locality code in Figure 1) and climate vectors along RDA2 versus RDA1, and RDA2 versus RDA3. The vectors (red arrows) show the direction of variation in the climate variables when fitted into the three-dimensional RDA space. The RDA axes represent independent climate vectors, which are most closely aligned with the locality variation in the 1177 climate-SNPs.
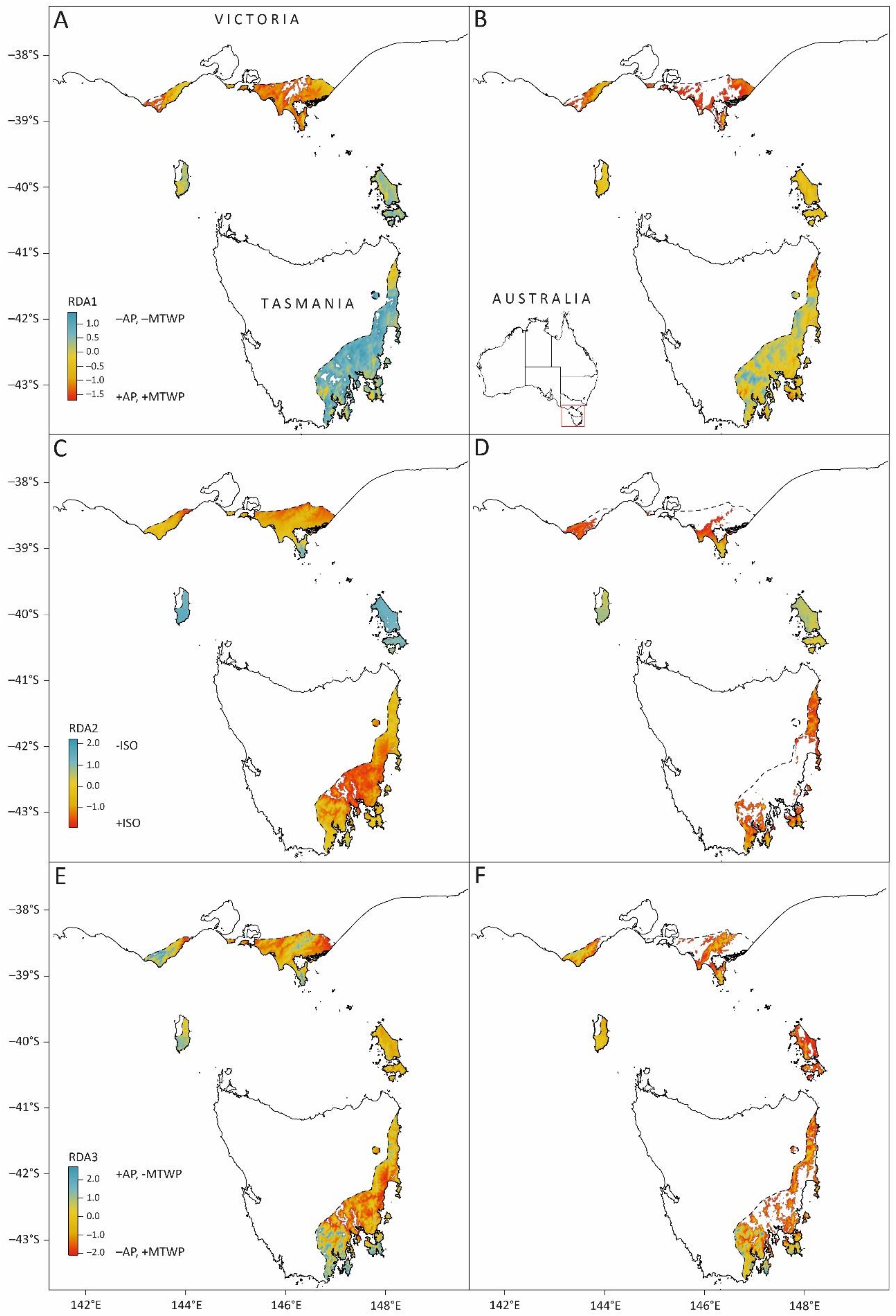
Using the climate variable loadings of the three significant RDA axes (Table S6) and contemporary rasters for the same climate variables, we developed genomics-informed climate indices that were spatially predicted across the native range of E. globulus (Figure 6). These spatial predictions represent the underlying climate-based selective surface to which the E. globulus gene pool has potentially adapted historically. The contemporary RDA1 projections indicate a latitudinal cline, with northern populations being warmer and wetter than southern populations (blue to red surface in Figure 6A). Projecting RDA1 using the 2070s climate predictions suggests that most of the southern E. globulus distribution, including the Bass Strait Islands, will experience conditions closer to the centre of the current RDA1 range of E. globulus (Figure 6B). In southern Victoria, which is already close to the high end of the RDA1 range, large portions of the eastern distribution will move beyond the current locality range in a more negative direction (not shown). A similar situation was found for the prediction based on RDA2, where the Bass Strait Island populations experience the highest values of RDA2 (i.e., lowest isothermality) under the contemporary climate, and the lowest values (i.e., high isothermality) occur inland in southern Victoria and in the central east coast of Tasmania (Figure 6C). Under the projected 2070s climate, the northern part of the Victorian distribution of E. globulus and the centre of the Tasmanian distribution are predicted to be beyond the current locality range of RDA2 (Figure 6D), in a negative direction, which reflects increasing isothermality (not shown). The contemporary aridity gradient defined by RDA3 shows aridity increasing from west to east in the Otway Ranges region, inland in eastern Victoria (Gippsland), and from south to central regions in eastern Tasmania (Figure 6E). Under the 2070s predictions, again, southern Victoria, the centre of the Tasmanian distribution and parts of the Furneaux Group are projected to move outside the current climatic range of the E. globulus localities on RDA3 as the climate becomes more arid (increasing negative values of RDA3) (Figure 6F).
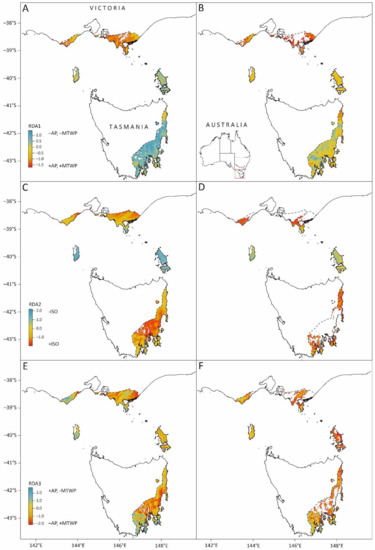
Figure 6.
Maps of the contemporary (left—(A,C,E)) and projected 2070s (right—(B,D,F)) climate scaled to loadings of the Table S1. (A,B); RDA2: (C,D); RDA3: (E,F)) are shown in Figure 5 and represent independent facets of the climate variation, which is associated with the identified set of climate-SNPs (i.e., SNPs indicated in blue in Figure 3).
For the contemporary climate, the variation along these axes represents the climate-selective surface, which has most contributed to the variation in climate SNPs among the E. globulus localities. The 2070’s projection shows how this surface will spatially change given the future climate projections for the climate variables contributing to each RDA axis. The key climate variables positively or negatively contributing to variation along each RDA axis are indicated on the legend (annual precipitation (AP), maximum temperature of the warmest period (MTWP) and isothermality (ISO)). The colour scheme of each grid cell represents the three climate variables (AP, MTWP, and ISO) for that cell scaled to the loadings of the indicated RDA axis (Table S6) and summed, to indicate its position along that axis (e.g., in Figure 6A, most of the cells covering the Tasmanian distribution of E. globulus are close to the positive limit of RDA1, indicating a lower combination of annual precipitation and maximum temperature when compared to those in the Victorian distribution). This scheme is based on the contemporary range of variation across the E. globulus range and is maintained in the future projection. Projections are limited to the current distribution of E. globulus, while white areas within that distribution are grid cells that fall outside of the range of RDA values defined by the localities used for modelling (Figure 5).
Based on 0.8 km grid cells (the resolution of the underlying climate rasters) falling within the locality range of variation on all three RDA axes (Figure 5) under the contemporary climate (hereafter referred to as ‘modelled suitable climate habitat’), 90% of the geographic envelope of E. globulus outlined in Figure 6 (20,703 km2, excluding west coast E. globulus) is predicted to be within genomically informed, modelled, suitable climate habitat for the species. Under future climates (2070s), this area reduces to 34%. Following projected future change in climate (regardless of being within or outside of the contemporary range on the RDA axes), the populations predicted to be least vulnerable are those on King Island and in the western Otways (shown in blue, Figure 7). Most of the King Island distribution was also predicted to be within modelled suitable climate habitat (as shown in Figure 6). In contrast, the E. globulus populations on the central and northern east coast of mainland Tasmania and Furneaux Islands are predicted to be most vulnerable (shown in red, Figure 7). A large portion of the central area on the Tasmanian east coast was also predicted to be outside the range of contemporary locality variation on both RDA2 and RDA3 in the 2070s climate (Figure 6). However, despite exhibiting high genomic vulnerability in the projected 2070s climate (Figure 7), the far north-eastern and south-eastern areas of mainland Tasmania and most of the Furneaux Islands were still predicted to be within modelled suitable climate habitat (Figure 6).
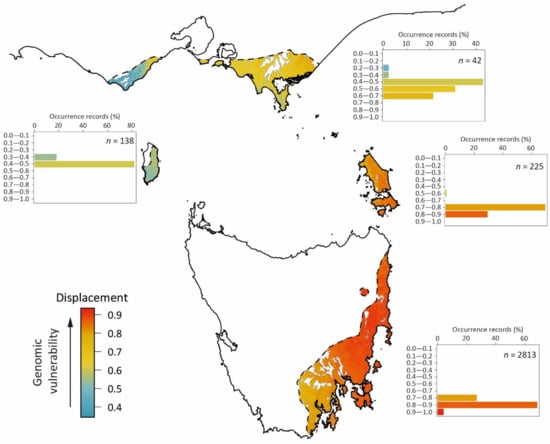
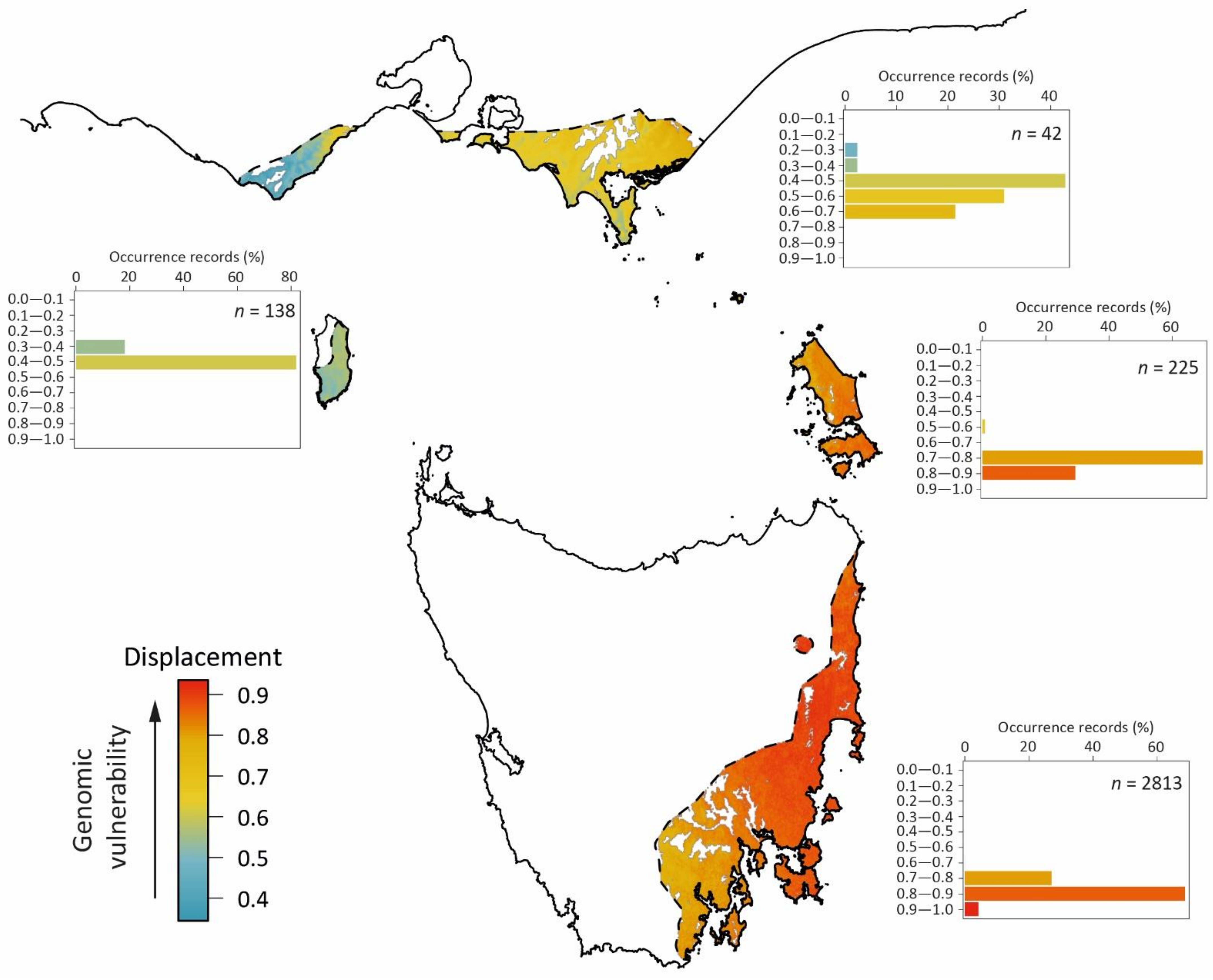
Figure 7.
The predicted genomic vulnerability of E. globulus populations to maladaptation under climate change. The main map shows the genomic vulnerability (colour scaled from blue to red) over the native geographic range of E. globulus, quantified as the displacement (Euclidean distance) predicted for grid cells in the three-dimensional RDA space shown in Figure 5 from the contemporary to the 2070s climate. White areas within the geographic range are outside the contemporary range for the species in the RDA space. The predicted least vulnerable populations are those on King Island and in the western Otways (shown in blue), and the most vulnerable are populations on the central and northern east coast of Tasmania (shown in red). Inset bar-charts show the percent of occurrence records within each decile bin of the predicted genomic vulnerability index for four regions (clockwise): mainland Australia (Victoria), Furneaux Islands, Tasmania, and King Island (e.g., of the recorded occurrences of E. globulus on King Island, 80% are located in grid cells that are displaced by 0.4–0.5 from the contemporary to predicted 2070s climate, indicating moderate genomic vulnerability). The number of records assessed for each region (n) is also shown in these bar-charts. The area indicated to be outside of the range of E. globulus on King Island has been extensively cleared for agriculture, but the occurrence of an isolated remnant tree within this area suggests that it may have been previously within the past range of E. globulus.
In terms of the recorded occurrences of E. globulus on the islands of Tasmania (n = 2813), 92% currently fall within the modelled suitable climate habitat, but this decreases to 40% by the 2070s. As expected, most of the recorded occurrences that were maintained within modelled suitable climate habitat were on King Island, the Furneaux Islands, and in areas in the far north-eastern and south-eastern regions of mainland Tasmania (Figure 6). In terms of genomic vulnerability (Figure 7), 67% of the recorded occurrences had a displacement between the contemporary and 2070s projected climate which exceeded 0.8 units, and would be considered at highest risk of maladaptation. Of these high risk occurrences, 68% were also predicted to be outside of the current modelled suitable climate habitat of E. globulus (i.e., 46% of all recorded occurrences in Tasmania) and were considered most at risk. However, populations in south-eastern Tasmania were predicted to be less vulnerable to climate maladaptation than those in the north-east (Figure 7). For the populations in the far north-east of Tasmania, the modelling suggests that their 2070s climate would favour contemporary SNP frequencies of populations located in more central areas of eastern Tasmania (visualised by matching colours of the areas in Figure 6). These central eastern populations are not only predicted to be the more genomically vulnerable by the 2070s (Figure 7), but outside the current modelled suitable climate for the species (Figure 6). Accordingly, our modelling suggest that the least vulnerable future climate refugia for E. globulus on mainland Tasmania are those in southern Tasmania.
4. Discussion
4.1. The Genomic Nature of Climate Adaptation
Our study clearly suggests that climate adaptation is polygenic and genome-wide, consistent with other SNP-climate association studies of plants [91,92], including forest trees [93,94] and more specifically eucalypts [29,35,37,38,40]. This is expected, given the numerous traits likely to be under direct and indirect selection along environmental gradients. In addition, the environmental gradients themselves may involve parallel changes in multiple biotic and abiotic selective agents [41]. The effect of climate variation on allele frequencies was not as intense as expected, with the maximum change in the frequency of climate-SNP alleles not exceeding 0.2 (after accounting for space). These small changes in allele frequencies, combined with the large number of climate-SNPs, suggests that the adaptative response involves adjustments across multiple genes and/or regulatory elements [42], likely modulating a suite of responses to climate variation. Other studies in forest trees have presented similar results. For instance, Eucalyptus microcarpa climate adaptation appeared to be characterised by subtle shifts in allele frequencies, with no consistent fixed differences identified [40]. Likewise, in various species of spruce (Picea spp.), oak (Quercus spp.), and in Pinus taeda and Fagus sylvatica, only small to moderate shifts in allele frequencies have been observed in genes associated with climatic gradients [22,93,94,95,96].
The genomic control of drought adaptation is also clearly complex in terms of the various interacting traits and genes, given the numerous SNPs associated with the locality-level drought damage. Variants associated with adaptation to aridity and drought traits are known to be numerous and occur across the eucalypt genome [42,97], with a recent study in Corymbia callophylla observing that drought-associated SNPs occur more often in regulatory elements than in protein-coding genes, and show signals of epistatic interaction [42]. Consistent with that observation is the lack of annotated genes near our most important drought-associated SNP, “G_0337_23205”, potentially suggesting the presence of long-distance regulatory elements [98].
Very few SNPs were identified as being important to both drought and climate adaptation. Although this result was a little unexpected, it may be explained by the fact that both climate and drought adaptation are complex traits involving multiple morphological, anatomical, and physiological adaptations [99]. The drought SNPs identified in our analysis may also be relatively specific to the drought conditions experienced by the field trials in Western Australia [47], and are unlikely to be an exhaustive list of SNPs putatively associated with drought adaptation in E. globulus. There are numerous mechanisms that affect a tree’s risk of drought-induced mortality, including susceptibility to hydraulic failure [99]. Responses to drought are likely different depending on the extent and duration of the water deficit. For instance, responses to short-term drought (hours to days) have been shown to minimise water loss or protect against effects of dehydration, while adaptations to long-term (days to weeks/months) droughts include shortening the life cycle or optimising resource gain through specific acclimatisation reactions [100]. Soil water content is often a more accurate predictor for drought adaptation patterns than climatic water predictors [101], implying that previous seasonal rainfall will influence the impact of a particular water deficit [102]. Phloem and xylem growth and repair are also seasonal, suggesting windows of increased vulnerability to drought damage [102]. However, the physiological response to water stress is only a small component of the breadth of adaptations required under drought conditions [103]. Water deficit is often confounded and intensified by heat stress [21,104] and their impacts can be amplified by biotic threats such as pests and disease [105,106]. In addition, from an evolutionary perspective, drought adaptation strategies can vary between resistance and recovery depending on climate or species, as seen in the varied responses to drought of co-occurring Pinus ponderosa and Populus tremuloides populations [107]. In E. globulus, drought resistance and recovery are largely under independent genetic control [108]. All things considered, the SNPs associated with drought damage are likely to be only a small component of the historic genome-wide adaptation to climate across the range of E. globulus.
In a similar vein, variants associated with climate may not only reflect direct adaptation to the abiotic factors that change with climate (e.g., heat, cold stress etc. [109]), but also the (co)adaptation associated with complex changes in biotic interactions that also occur across climate gradients (as proposed in the geographic mosaic theory [110]). Such changes have been hypothesised for E. globulus [111] and are no better seen than in the disease adaptation that occurs across its geographic range [112,113]. Such biotic interactions are expected to be embodied in our genomic signals of climate adaptation. Climate-responsive biotic factors are likely to contribute to the complexity of the genomic changes, which has been observed in a study of the sub-tropical E. grandis, where Mostert-O’Neill, Reynolds [38] showed GO terms for defense responses were the terms consistently enriched in their sets of environmentally associated SNPs, although this was not evident in our enrichment analysis.
4.2. Climate Variables Driving Genomic Adaptation
The current adaptive differences among the E. globulus provenances are a result of historical selection and constraints [52]. Our RDA analysis allowed the identification of the independent climate vectors most likely to have historically contributed to shaping the genomic adaptation in these provenances. The home-site climate variables contributing to these vectors and exhibiting the most important associations with genomic adaptation were isothermality, annual precipitation, and the maximum temperature of the warmest month. Specifically, the majority of climate-SNPs (and the majority of those associated with enriched functional groups, identified through GO analysis) were associated with variation in maximum temperature. Temperature appears to be a ubiquitous driver of adaptive variation, both in a quantitative genetic [114,115,116,117] and genomic [118,119] sense in many forest tree species, including eucalypts [9,31,35,37,120]. Functional traits that have been demonstrated to vary significantly with home-site maximum temperature include morphological variation in the leaf traits of Quercus berberidifolia [121], and lignotuber development in E. ovata [122] and E. pauciflora [18]. Maximum temperature of the warmest month is also one of the most important variables driving the eucalypt species distributions in temperate Australia [123]. Any changes in future temperatures are likely to be closely correlated with shifts in tree species distributions; indeed, this has already been observed [124,125].
The interaction of precipitation with temperature is an important consideration. Annual precipitation was found to be one of the most important climactic variables in explaining the genetic variation observed in E. globulus, and is a key selective force in many systems [126]. However, studies in Australia often find aridity, the ratio of precipitation to evapotranspiration (in turn influenced by temperature), to be an equal or better predictor of adaptive differences [9,37,97,127], which is unsurprising given that much of the continent is subtropical or tropical with extreme gradients of aridity. For example, a study of Corymbia callophylla populations in Western Australia found, using RDA, that an aridity gradient (along with maximum temperature of the warmest month and annual precipitation) explained 65% of the variation in climate-associated SNPs [37]. While E. globulus is a temperate species, the climate variables driving adaptation were largely the same (although the greater variance explained in Corymbia is likely due to less variance associated with spatial components, given its more continuous distribution), and an aridity gradient is reflected in RDA3. Trees endemic to the northern hemisphere also experience selective pressure from aridity, but in addition need to survive the often much lower temperatures experienced in temperate and boreal regions [128].
Isothermality was the climate variable that explained the largest proportion of SNP variation among the E. globulus localities studied, and is generally considered a measure of temperature seasonality or “evenness” [96,129]. It represents the ratio of mean diurnal temperature range to annual temperature range [71]. While it has been associated with tree species’ distributions [130,131], species richness [132,133], historical change in leaf morphology [134], and with SNPs associated with divergent selection within species [135], including eucalypts [36], it is difficult to link isothermality to specific physiological processes. Similar to the MEMs, it may be correlated with more proximal climatic measures [136]. A component of isothermality is continentality (often defined as the difference between the mean temperatures of the warmest month and the coldest month), which is known to influence tree species distributions in the northern hemisphere [137]. Indeed, the isothermality gradient in our study is highly correlated with annual temperature range (r = −0.80, p < 0.001) and mainly reflected a continentality effect separating E. globulus populations on the Bass Strait Islands from those of mainland Australia and Tasmania. It is likely that continental populations are adapted to greater seasonal variation in energy availability when compared to their coastal counterparts [132]. As large changes in isothermality are predicted across the distribution of E. globulus on mainland Australia and Tasmania (see below), its influence as a potential selective force warrants consideration.
4.3. Predicting Future Selective Surfaces
Understanding the extent to which E. globulus is at risk of future climate maladaptation within its native distribution is important for the conservation of genetic resources and guiding seed transfer decisions for this globally important plantation tree species that also provides critical habitat and food resource for endangered animal species [43,44]. There are clearly challenges and assumptions associated with modelling the future distribution of suitable climate habitat, whether it involves traditional species distribution modelling or is guided by intra-specific variation in climate adaptation [138,139], as done with our genomics-informed modelling. Not least are the issues of accounting for extreme events rather than climate averages [140], competitive and other biotic interactions [141,142], and resilient vegetative recovery mechanisms that many forest tree species, such as eucalypts, possess [143].
Bearing these qualifications in mind, our predictions of genomic vulnerability suggest that the E. globulus populations most at risk of future climate maladaptation are those in the central and northern parts of the eastern range of E. globulus in Tasmania. It is predicted that occurrences within these limits will be subject to directional selection due to changes in climate variables, which in our analyses translate into independent, adaptively relevant changes in annual precipitation and temperatures (RDA1), isothermality (RDA2), and aridity (RDA3). While increased temperatures have the potential to lead to increased photosynthesis and productivity, which would likely be enhanced by elevated CO2 levels, this may not be realised in drier areas [144]. Increased temperatures and precipitation are also expected to increase the risk of selective pressures such as wood decay and foliar disease [52,112,145]. In Australia, fungal pathogen load in native eucalypt forest increases with increasing annual rainfall [146]. Specifically, the risk of one of the major native diseases of E. globulus (Teratosphaeria sp.) increases in warmer and wetter regions [112,145], which is consistent with increasing negative values of RDA1. Indeed, as climates change, fungal pathogens are predicted to increase up to 100-fold in coastal eucalypt forests when compared with current abundance [146]. This finding is of direct relevance to E. globulus as it is a common dominant tree in the coastal forest of south-eastern Australia, including Tasmania. However, the extent to which predictions of genomic vulnerability translate into future performance and fitness is a key research challenge [147], requiring lines of evidence from multiple approaches, including common garden experiments [148,149].
Eucalyptus globulus is one of the few forest tree species where direct insights into the effects of climate change on Darwinian fitness can be gauged. This is possible by using a space-for-time interpolation of the climate modelling of naturalisation/invasion risk from (i) plantations of the species established over the last few decades in Australia (a large component of which is outside the natural distribution), and (ii) plantations in other countries where the species has long been introduced, grows well, and reproduces outside the realised climate experienced in the natural range [150]. While provenance differences in reproduction [151] and germination responses [152] exist, at a species level, these exotic plantings indicate clear climatic limits to the reproductive fitness of E. globulus, as assessed in terms of wildling establishment. Such naturally established seedling recruits are a good indicator of the potential of the proximally planted E. globulus trees to contribute to the next generation. In the case of E. globulus plantations in Australia, Larcombe, et al. [153] found that wilding abundance increased in areas that had environmental conditions similar to those in the native range, reducing with increasing annual temperatures and rainfall seasonality, and with decreasing mean annual precipitation (<1400 mm). In Portugal, where plantations extended into areas with higher rainfall than experienced by the species in Australia, mean annual precipitation was shown to be the most important predictor of wildling densities, which peaked in areas with 1300–1500 mm annual precipitation, close to the maximum for the native distribution, but declined rapidly in lower and higher rainfall areas [154]. It was proposed that this optimum could be defined by a balance between drought stress and factors such as competition and disease susceptibility in higher rainfall areas. Together, these studies of wildling establishment argue that climate change along the RDA axes identified will directly impact the fitness of E. globulus in native forest. However, it is important to note that these ‘space-for-time’ comparisons do not account for interacting factors or novel selection forces that may alter future selection surfaces.
Multiple selective forces that impact future adaptative responses may or may not be directly related to climate adaptation, and may result in more than a simple reshuffling of the extant genetic variation across the landscape. A factor that will diminish our power to predict selective trajectories is the possibility of novel future climates, accentuated by elevated CO2, which may alter growth and drought responses [144,155]. When grown in temperatures exceeding those of its native range, E. globulus is limited in its photosynthetic capacity, which may be key in defining the future climate zones where E. globulus can continue to be a productive plantation species [156]. Novel climates may also subject tree species to numerous novel biotic interactions that may challenge tree hosts, a likely occurrence given the increasing frequency of exotic pests globally [157,158]. Exotic pests may differentially affect performance at the species and provenance levels and potentially alter the selection landscape for native eucalypts in Australia [113,159]. Changing species distributions may also change mating and hybridisation patterns, which in turn may change the predictability of adaptation trajectories based on historic patterns [38,160].
5. Conclusions
The present study serves to highlight the importance of understanding the limits to plasticity and the extent to which predicted changes in climate will impact the vegetative and reproductive performance and, ultimately, the fitness of sedentary, long-lived foundation species such as trees. A priority action for threatened and economically important species such as E. globulus will be the collection and long-term storage of seed resources from populations identified as most at risk of climate maladaptation before reproductive capacity is lost. Some populations most at risk are already likely to be at the environmental limits of the species and thus habour adaptative variations that are of value for the implementation of climate-adjusted restoration and translocation strategies, and enriching the genetic diversity of breeding populations. Quantifying the predicted climate of current and future plantation forestry sites in a similar manner to this study will give the best indication of which provenances are most important to incorporate into these breeding populations, to maximise future performance and prevent climate-induced failure.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/f13040575/s1, Figure S1: Examples of partial dependency patterns observed among climate-SNPs; Table S1: Eucalyptus globulus localities sampled; Table S2: Validation of gradientForest GEA method using IBD simulation; Table S3: List of climate-SNPs; Table S4: Enriched gene ontology (GO) terms associated with climate-SNPs; Table S5: List of SNPs associated with drought damage; Table S6: RDA loadings of major axes of climate variation.
Author Contributions
Conceptualisation, all authors; formal analysis, J.B.B., P.A.H.; data curation, J.B.B., J.F.G.T.; writing—original draft preparation, J.B.B.; writing—review and editing, P.A.H., J.F.G.T., D.A.S., R.E.V., B.M.P. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported by the Australian Research Council (grant number DP190102053).
Data Availability Statement
The read data used to generate the SNPs in this study are accessible on the European Nucleotide Archive (accession PRJEB47881). The E. globulus draft genome v33 is available upon reasonable request to the authors.
Acknowledgments
The authors wish to acknowledge Greg Dutkowski for his assistance in collating the drought damage data. We also thank Forico Pty Ltd. for access to the E. globulus trial studied and Tree Breeding Australia Limited for access to the E. globulus drought damage information. We thank Hossein Kahrood and Peter Ades (University of Melbourne) for their contribution towards the generation of the sequence data used in this study. We also acknowledge the use of the high performance computing facilities provided by Digital Research Services, IT Services at the University of Tasmania.
Conflicts of Interest
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Appendix A
Appendix A.1. Genotyping by Whole Genome Shotgun Sequencing
Genomic libraries were prepared using Nextera DNA Library Preparation Kit (Illumina). Double-stranded genomic DNA was used as input for the Nextera ‘tagmentation’ reaction (DNA fragmentation and tagging). The reaction was performed as per the manufacturer’s protocol, replacing the column clean-up (disassociation) step with a 0.6% SDS (sodium dodecyl sulfate) treatment. Library amplification was performed using Nextera Index 1 (i7) and Index 2 (i5) adapters on a Stratagene Mx3005PTM qPCR system (Agilent Technologies), with a thermal profile as follows: 72 °C for 3 min; 98 °C for 30 s; 5 cycles of 98 °C for 10 s, 63 °C for 30 s, and 72 °C for 3 min. SYBR Green I (Thermo Fisher Scientific) in 1:1000 dilution was added to the PCR mixtures for monitoring the amplification process. Amplified libraries were pooled and purified with MinElute PCR Purification Kit (Qiagen) or AMPure XP SPRI (Solid Phase Reversible Immobilisation) paramagnetic beads (Agencourt, Beckman Coulter). Purified library pools were then subjected to size selection using gel electrophoresis by excising the DNA fragments within the desired size range (300–450 bp). The excised libraries were extracted from the gel with QIAquick Gel Extraction Kit (Qiagen) prior to quality assessment with High Sensitivity D1000 ScreenTape using a 2200 TapeStation system (Agilent Technologies, CA, USA).
These libraries were sequenced using a HiSeq 2000 Sequencing System (Illumina) at the Victorian Government Department of Economic Development, Jobs, Transport, and Resources (DEDJTR), Centre for AgriBioscience (AgriBio), Bundoora, Australia. The Illumina paired-read data generated from whole genome sequencing (available at the European Nucleotide Archive, accession PRJEB47881) were quality controlled and aligned to an E. globulus draft genome assembly for full specifications of genome assembly, see [161]. Variants were discovered using find-snp v2.6.16 (Gydle Inc., Québec, Canada) with output in Variant Call Format v.4.2 (VCF).
The raw SNP data were filtered using PLINK v1.9 [162] to biallelic SNPs with <30% missing data and a minor allele frequency (MAF) >0.05. To minimise the impact of genotyping errors and paralogous loci, SNPs that deviated from HWE in an extreme manner were also removed (p < 1 × 10−7) [163]. To reduce the computational burden, pairs of SNPs were also assessed for linkage disequilibrium (LD), with the less informative (lower MAF) member of the pair removed if found to be in high LD (r2 > 0.7). Once finalised, the minor allele frequency of each SNP in each locality was calculated for use in subsequent analysis.
Appendix A.2. gradientForest Analysis of Climate Variables
The machine learning algorithm gradientForest [97] was used to fit the SNP data to the environmental and spatial variables, using 5000 regression trees per SNP and a variable correlation threshold of 0.5. Due to computational limitations, gradientForest was fit to blocks of only 100,000 SNPs at a single time, repeated until the entire dataset was covered. In brief, the gradientForest algorithm gives each SNP an importance score, both globally and to each individual predictor variable, which indicates how relevant that SNP is to delineating the randomForest regression [97]. Those SNPs from the separate analyses deemed important to the overall model (in the upper quartile for importance within the set of 100,000) were then collated and gradientForest was refit to this set, to ensure each importance score was scaled correctly among the SNPs. Visual analysis of the distribution of importance scores led to the selection of an importance threshold of 0.15 for SNP selection. SNPs above this threshold within the 19 climate variables were chosen as climate-SNPs for further analysis.
Appendix A.3. extendedForest Analysis of Drought Damage
The drought-SNP association used the initial set of 812,158 SNPs as predictor variables for drought damage rather than as response variables, and thus a larger amount of regression trees (100,000) per response was required. Due to computational limitations, SNPs were divided into sets of 25,000 and used in multiple extendedForest runs. The SNPs with highest importance from each run were collated into a single dataset and run again to ensure each importance score was scaled correctly among the SNPs. The top 4% in importance were then selected to achieve a similar number of SNPs to those reported for climate.
Appendix A.4. Visualising Climate Selection Surface
The climate indices associated with the adaptive changes among provenances described by the significant RDA axes were extrapolated across Tasmania and southern Victoria. This was undertaken by normalising the raster grid cells from the RDA-selected climate layers obtained from CHELSA [70], using the mean and standard deviations of the climate data for the 30 localities of E. globulus sampled in Tasmania and southern Victoria. The values of the climate selection surface were truncated to the predicted range for the E. globulus localities used in this study, using the 95% confidence intervals around the mean gradient value. Each climate layer was then scaled by its associated weighting in the most significant RDA axis, and the value of each layer was combined and visualised. This was repeated for all significant RDA axes.
References
- Loarie, S.R.; Duffy, P.B.; Hamilton, H.; Asner, G.P.; Field, C.B.; Ackerly, D.D. The velocity of climate change. Nature 2009, 462, 1052–1055. [Google Scholar] [CrossRef]
- Sáenz-Romero, C.; O’Neill, G.; Aitken, S.N.; Lindig-Cisneros, R. Assisted migration field tests in Canada and Mexico: Lessons, limitations, and challenges. Forests 2021, 12, 9. [Google Scholar] [CrossRef]
- Franks, S.J.; Weber, J.J.; Aitken, S.N. Evolutionary and plastic responses to climate change in terrestrial plant populations. Evol. Appl. 2014, 7, 123–139. [Google Scholar] [CrossRef]
- Isabel, N.; Holliday, J.A.; Aitken, S.N. Forest genomics: Advancing climate adaptation, forest health, productivity, and conservation. Evol. Appl. 2020, 13, 3–10. [Google Scholar] [CrossRef] [PubMed]
- Flood, P.J.; Hancock, A.M. The genomic basis of adaptation in plants. Curr. Opin. Plant Biol. 2017, 36, 88–94. [Google Scholar] [CrossRef]
- Orr, H.A. The genetic theory of adaptation: A brief history. Nat. Rev. Genet 2005, 6, 119–127. [Google Scholar] [CrossRef] [PubMed]
- Stapley, J.; Reger, J.; Feulner, P.G.D.; Smadja, C.; Galindo, J.; Ekblom, R.; Bennison, C.; Ball, A.D.; Beckerman, A.P.; Slate, J. Adaptation genomics: The next generation. Trends Ecol. Evol. 2010, 25, 705–712. [Google Scholar] [CrossRef]
- Sáenz-Romero, C.; Kremer, A.; Nagy, L.; Újvári-Jármay, É.; Ducousso, A.; Kóczán-Horváth, A.; Hansen, J.K.; Mátyás, C. Common garden comparisons confirm inherited differences in sensitivity to climate change between forest tree species. PeerJ 2019, 7, e6213. [Google Scholar] [CrossRef] [PubMed]
- Steane, D.A.; Potts, B.M.; McLean, E.; Prober, S.M.; Stock, W.D.; Vaillancourt, R.E.; Byrne, M. Genome-wide scans detect adaptation to aridity in a widespread forest tree species. Mol. Ecol. 2014, 23, 2500–2513. [Google Scholar] [CrossRef] [PubMed]
- Rellstab, C.; Gugerli, F.; Eckert, A.J.; Hancock, A.M.; Holderegger, R. A practical guide to environmental association analysis in landscape genomics. Mol. Ecol. 2015, 24, 4348–4370. [Google Scholar] [CrossRef]
- Hoban, S.; Kelley, J.L.; Lotterhos, K.E.; Antolin, M.F.; Bradburd, G.; Lowry, D.B.; Poss, M.L.; Reed, L.K.; Storfer, A.; Whitlock, M.C. Finding the genomic basis of local adaptation: Pitfalls, practical solutions, and future directions. Am. Nat 2016, 188, 379–397. [Google Scholar] [CrossRef]
- Bierne, N.; Roze, D.; Welch, J.J. Pervasive selection or is it…? why are FST outliers sometimes so frequent? Mol. Ecol. 2013, 22, 2061–2064. [Google Scholar] [CrossRef]
- Forester, B.R.; Lasky, J.R.; Wagner, H.H.; Urban, D.L. Comparing methods for detecting multilocus adaptation with multivariate genotype–environment associations. Mol. Ecol. 2018, 27, 2215–2233. [Google Scholar] [CrossRef]
- Mitchell-Olds, T.; Shaw, R.G. Regression analysis of natural selection: Statistical inference and biological interpretation. Evolution 1987, 41, 1149–1161. [Google Scholar] [CrossRef]
- Whitlock, M.C.; Lotterhos, K.E. Reliable detection of loci responsible for local adaptation: Inference of a null model through trimming the distribution of FST. Am. Nat. 2015, 186, S24–S36. [Google Scholar] [CrossRef]
- Petit, R.J.; Hampe, A. Some evolutionary consequences of being a tree. Annu. Rev. Ecol. Evol. Syst. 2006, 37, 187–214. [Google Scholar] [CrossRef]
- Hudson, C.J.; Freeman, J.S.; Jones, R.C.; Potts, B.M.; Wong, M.M.; Weller, J.L.; Hecht, V.F.; Poethig, R.S.; Vaillancourt, R.E. Genetic control of heterochrony in Eucalyptus globulus. G3 Genes Genomes Genet. 2014, 4, 1235–1245. [Google Scholar] [CrossRef]
- Gauli, A.; Vaillancourt, R.E.; Bailey, T.G.; Steane, D.A.; Potts, B.M. Evidence for local climate adaptation in early-life traits of Tasmanian populations of Eucalyptus pauciflora. Tree Genet. Genom. 2015, 11, 104. [Google Scholar] [CrossRef]
- Liepe, K.J.; Hamann, A.; Smets, P.; Fitzpatrick, C.R.; Aitken, S.N. Adaptation of lodgepole pine and interior spruce to climate: Implications for reforestation in a warming world. Evol. Appl. 2016, 9, 409–419. [Google Scholar] [CrossRef]
- Alberto, F.J.; Derory, J.; Boury, C.; Frigerio, J.-M.; Zimmermann, N.E.; Kremer, A. Imprints of natural selection along environmental gradients in phenology-related genes of Quercus petraea. Genetics 2013, 195, 495–512. [Google Scholar] [CrossRef]
- Kremer, A.; Potts, B.M.; Delzon, S.; Bailey, J. Genetic divergence in forest trees: Understanding the consequences of climate change. Funct. Ecol. 2014, 28, 22–36. [Google Scholar] [CrossRef]
- Rellstab, C.; Zoller, S.; Walthert, L.; Lesur, I.; Pluess, A.R.; Graf, R.; Bodénès, C.; Sperisen, C.; Kremer, A.; Gugerli, F. Signatures of local adaptation in candidate genes of oaks (Quercus spp.) with respect to present and future climatic conditions. Mol. Ecol. 2016, 25, 5907–5924. [Google Scholar] [CrossRef] [PubMed]
- Fitzpatrick, M.C.; Keller, S.R. Ecological genomics meets community-level modelling of biodiversity: Mapping the genomic landscape of current and future environmental adaptation. Ecol. Lett. 2015, 18, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Wogan, G.O.U.; Wang, I.J. The value of space-for-time substitution for studying fine-scale microevolutionary processes. Ecography 2018, 41, 1456–1468. [Google Scholar] [CrossRef]
- Naidu, B.P.; Paleg, L.G.; Jones, G.P. Accumulation of proline analogues and adaptation of Melaleuca species to diverse environments in Australia. Aust. J. Bot. 2000, 48, 611–620. [Google Scholar] [CrossRef]
- Álvarez, S.; Navarro, A.; Nicolás, E.; Sánchez-Blanco, M.J. Transpiration, photosynthetic responses, tissue water relations and dry mass partitioning in Callistemon plants during drought conditions. Sci. Hortic. 2011, 129, 306–312. [Google Scholar] [CrossRef]
- Potts, B.; Wiltshire, R. Eucalypt genetics and genecology. In Eucalypt Ecology: Individuals to Ecosystems; Williams, J., Woinarski, J., Eds.; Cambridge University Press: Cambridge, UK, 1997. [Google Scholar]
- Prober, S.M.; Potts, B.M.; Bailey, T.; Byrne, M.; Dillon, S.; Harrison, P.A.; Hoffmann, A.A.; Jordan, R.; McLean, E.H.; Steane, D.A.; et al. Climate adaptation and ecological restoration in eucalypts. Proc. R. Soc. Vic. 2016, 128, 40–53. [Google Scholar] [CrossRef]
- Steane, D.A.; Mclean, E.H.; Potts, B.M.; Prober, S.M.; Stock, W.D.; Stylianou, V.M.; Vaillancourt, R.E.; Byrne, M. Evidence for adaptation and acclimation in a widespread eucalypt of semi-arid Australia. Biol. J. Linn. Soc. 2017, 121, 484–500. [Google Scholar] [CrossRef]
- Drake, J.E.; Aspinwall, M.J.; Pfautsch, S.; Rymer, P.D.; Reich, P.B.; Smith, R.A.; Crous, K.Y.; Tissue, D.T.; Ghannoum, O.; Tjoelker, M.G. The capacity to cope with climate warming declines from temperate to tropical latitudes in two widely distributed Eucalyptus species. Glob. Chang. Biol. 2015, 21, 459–472. [Google Scholar] [CrossRef]
- Dillon, S.; McEvoy, R.; Baldwin, D.S.; Rees, G.N.; Parsons, Y.; Southerton, S. Characterisation of adaptive genetic diversity in environmentally contrasted populations of Eucalyptus camaldulensis Dehnh. (river red gum). PLoS ONE 2014, 9, e103515. [Google Scholar] [CrossRef]
- Costa e Silva, J.; Potts, B.M.; Dutkowski, G.W. Genotype by environment interaction for growth of Eucalyptus globulus in Australia. Tree Genet. Genom. 2006, 2, 61–75. [Google Scholar] [CrossRef]
- Grattapaglia, D.; Vaillancourt, R.E.; Shepherd, M.; Thumma, B.R.; Foley, W.; Külheim, C.; Potts, B.M.; Myburg, A.A. Progress in Myrtaceae genetics and genomics: Eucalyptus as the pivotal genus. Tree Genet. Genom. 2012, 8, 463–508. [Google Scholar] [CrossRef]
- Byrne, M.; Prober, S.; McLean, E.; Steane, D.; Stock, W.; Potts, B.; Vaillancourt, R. Adaptation to Climate in Widespread Eucalypt Species; National Climate Change Adaptation Research Facility: Gold Coast, Australia, 2013; p. 86. [Google Scholar]
- Murray, K.D.; Janes, J.K.; Jones, A.; Bothwell, H.M.; Andrew, R.L.; Borevitz, J.O. Landscape drivers of genomic diversity and divergence in woodland Eucalyptus. Mol. Ecol. 2019, 28, 5232–5247. [Google Scholar] [CrossRef]
- Supple, M.A.; Bragg, J.G.; Broadhurst, L.M.; Nicotra, A.B.; Byrne, M.; Andrew, R.L.; Widdup, A.; Aitken, N.C.; Borevitz, J.O. Landscape genomic prediction for restoration of a Eucalyptus foundation species under climate change. Elife 2018, 7, e31835. [Google Scholar] [CrossRef]
- Ahrens, C.W.; Byrne, M.; Rymer, P.D. Standing genomic variation within coding and regulatory regions contributes to the adaptive capacity to climate in a foundation tree species. Mol. Ecol. 2019, 28, 2502–2516. [Google Scholar] [CrossRef]
- Mostert-O’Neill, M.M.; Reynolds, S.M.; Acosta, J.J.; Lee, D.J.; Borevitz, J.O.; Myburg, A.A. Genomic evidence of introgression and adaptation in a model subtropical tree species, Eucalyptus grandis. Mol. Ecol. 2021, 30, 625–638. [Google Scholar] [CrossRef]
- von Takach, B.; Ahrens, C.W.; Lindenmayer, D.B.; Banks, S.C. Scale-dependent signatures of local adaptation in a foundation tree species. Mol. Ecol. 2021, 30, 2248–2261. [Google Scholar] [CrossRef]
- Jordan, R.; Hoffmann, A.A.; Dillon, S.K.; Prober, S.M. Evidence of genomic adaptation to climate in Eucalyptus microcarpa: Implications for adaptive potential to projected climate change. Mol. Ecol. 2017, 26, 6002–6020. [Google Scholar] [CrossRef]
- Costa e Silva, J.; Potts, B.; Harrison, P.A.; Bailey, T. Temperature and rainfall are separate agents of selection shaping population differentiation in a forest tree. Forests 2019, 10, 1145. [Google Scholar] [CrossRef]
- Ahrens, C.W.; Murray, K.; Mazanec, R.A.; Ferguson, S.; Bragg, J.; Jones, A.; Tissue, D.T.; Byrne, M.; Borevitz, J.O.; Rymer, P.D. Genomic constraints to drought adaptation. bioRxiv 2021. [Google Scholar] [CrossRef]
- Hingston, A.B.; Gartrell, B.D.; Pinchbeck, G. How specialized is the plant–pollinator association between Eucalyptus globulus ssp. globulus and the swift parrot Lathamus discolor? Aust. Ecol. 2004, 29, 624–630. [Google Scholar] [CrossRef]
- Porfirio, L.L.; Harris, R.M.B.; Stojanovic, D.; Webb, M.H.; Mackey, B. Projected direct and indirect effects of climate change on the Swift Parrot, an endangered migratory species. Emu Aust. Ornithol. 2016, 116, 273–283. [Google Scholar] [CrossRef]
- Potts, B.M.; Vaillancourt, R.E.; Jordan, G.; Dutkowski, G.; Costa e Silva, J.; McKinnon, G.; Steane, D.; Volker, P.; Lopez, G.; Apiolaza, L.; et al. Exploration of the Eucalyptus globulus gene pool. In Proceedings of the Eucalyptus in a Changing World, International IUFRO Conference of the WP2.08.03 on Silviculture and Improvement of Eucalypts, Aveiro, Portugal, 11–15 October 2004; pp. 46–61. [Google Scholar]
- Fitzpatrick, M.C.; Chhatre, V.E.; Soolanayakanahally, R.Y.; Keller, S.R. Experimental support for genomic prediction of climate maladaptation using the machine learning approach Gradient Forests. Mol. Ecol. Resour. 2021, 21, 2749–2765. [Google Scholar] [CrossRef] [PubMed]
- Dutkowski, G.W.; Potts, B.M. Genetic variation in the susceptibility of Eucalyptus globulus to drought damage. Tree Genet. Genom. 2012, 8, 757–773. [Google Scholar] [CrossRef]
- Williams, K.; Potts, B. The natural distribution of Eucalyptus species in Tasmania. Tasforests 1996, 8, 39–165. [Google Scholar]
- Dutkowski, G.W.; Potts, B.M. Geographic patterns of genetic variation in Eucalyptus globulus ssp. globulus and a revised racial classification. Aust. J. Bot. 1999, 47, 237–263. [Google Scholar] [CrossRef]
- Jones, R.C.; Steane, D.A.; Lavery, M.; Vaillancourt, R.E.; Potts, B.M. Multiple evolutionary processes drive the patterns of genetic differentiation in a forest tree species complex. Ecol. Evol. 2013, 3, 1–17. [Google Scholar] [CrossRef]
- Eldridge, K.G.; Davidson, J.; Harwood, C.; Wyk, G.v. Eucalypt Domestication and Breeding; Clarendon Press: Oxford, UK, 1994. [Google Scholar]
- Costa e Silva, J.; Potts, B.M.; Harrison, P.A. Population divergence along a genetic line of least resistance in the tree species Eucalyptus globulus. Genes 2020, 11, 1095. [Google Scholar] [CrossRef]
- Jordan, G.J.; Potts, B.M.; Chalmers, P.; Wiltshire, R.J. Quantitative genetic evidence that the timing of vegetative phase change in Eucalyptus globulus ssp. globulus is an adaptive trait. Aust. J. Bot. 2000, 48, 561–567. [Google Scholar] [CrossRef]
- Foster, S.A.; McKinnon, G.E.; Steane, D.A.; Potts, B.M.; Vaillancourt, R.E. Parallel evolution of dwarf ecotypes in the forest tree Eucalyptus globulus. New Phytol. 2007, 175, 370–380. [Google Scholar] [CrossRef]
- Hamilton, M.G.; Freeman, J.S.; Blackburn, D.P.; Downes, G.M.; Pilbeam, D.J.; Potts, B.M. Independent lines of evidence of a genetic relationship between acoustic wave velocity and kraft pulp yield in Eucalyptus globulus. Ann. For. Sci. 2017, 74, 17. [Google Scholar] [CrossRef]
- Barbour, R.C.; Forster, L.G.; Baker, S.C.; Steane, D.A.; Potts, B.M. Biodiversity consequences of genetic variation in bark characteristics within a foundation tree species. Conserv. Biol. 2009, 23, 1146–1155. [Google Scholar] [CrossRef]
- Gosney, B.J.; Potts, B.M.; Forster, L.G.; Whiteley, C.; O’Reilly-Wapstra, J.M. Consistent community genetic effects in the context of strong environmental and temporal variation in Eucalyptus. Oecologia 2021, 195, 367–382. [Google Scholar] [CrossRef]
- O’Reilly-Wapstra, J.M.; Miller, A.M.; Hamilton, M.G.; Williams, D.; Glancy-Dean, N.; Potts, B.M. Chemical variation in a dominant tree species: Population divergence, selection and genetic stability across environments. PLoS ONE 2013, 8, e58416. [Google Scholar] [CrossRef]
- Nickolas, H.; Williams, D.; Downes, G.; Harrison, P.A.; Vaillancourt, R.E.; Potts, B.M. Application of resistance drilling to genetic studies of growth, wood basic density and bark thickness in Eucalyptus globulus. Aust. For. 2020, 83, 172–179. [Google Scholar] [CrossRef]
- Skabo, S.; Vaillancourt, R.; Potts, B. Fine-scale genetic structure of Eucalyptus globulus ssp. globulus forest revealed by RAPDs. Aust. J. Bot. 1998, 46, 583–594. [Google Scholar] [CrossRef]
- Jones, T.H.; Vaillancourt, R.E.; Potts, B.M. Detection and visualization of spatial genetic structure in continuous Eucalyptus globulus forest. Mol. Ecol. 2007, 16, 697–707. [Google Scholar] [CrossRef]
- Steane, D.A.; Conod, N.; Jones, R.C.; Vaillancourt, R.E.; Potts, B.M. A comparative analysis of population structure of a forest tree, Eucalyptus globulus (Myrtaceae), using microsatellite markers and quantitative traits. Tree Genet. Genom. 2006, 2, 30–38. [Google Scholar] [CrossRef]
- Yeoh, S.H.; Bell, J.C.; Foley, W.J.; Wallis, I.R.; Moran, G.F. Estimating population boundaries using regional and local-scale spatial genetic structure: An example in Eucalyptus globulus. Tree Genet. Genom. 2012, 8, 695–708. [Google Scholar] [CrossRef]
- Costa e Silva, J.; Vaillancourt, R.E.; Steane, D.A.; Jones, R.C.; Marques, C. Microsatellite analysis of population structure in Eucalyptus globulus. Genome 2017, 60, 770–777. [Google Scholar] [CrossRef]
- Freeman, J.; Jackson, H.; Steane, D.; McKinnon, G.; Dutkowski, G.; Potts, B.; Vaillancourt, R. Chloroplast DNA phylogeography of Eucalyptus globulus. Aust. J. Bot. 2001, 49, 585–596. [Google Scholar] [CrossRef]
- McKinnon, G.E.; Jordan, G.J.; Vaillancourt, R.E.; Steane, D.A.; Potts, B.M. Glacial refugia and reticulate evolution: The case of the Tasmanian eucalypts. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2004, 359, 275–284. [Google Scholar] [CrossRef]
- Mimura, M.; Barbour, R.C.; Potts, B.M.; Vaillancourt, R.E.; Watanabe, K.N. Comparison of contemporary mating patterns in continuous and fragmented Eucalyptus globulus native forests. Mol. Ecol. 2009, 18, 4180–4192. [Google Scholar] [CrossRef]
- Tibbits, J.F.G.; McManus, L.J.; Spokevicius, A.V.; Bossinger, G. A rapid method for tissue collection and high-throughput isolation of genomic DNA from mature trees. Plant Mol. Bio. Rep. 2006, 24, 81–91. [Google Scholar] [CrossRef]
- Thavamanikumar, S.; McManus, L.J.; Ades, P.K.; Bossinger, G.; Stackpole, D.J.; Kerr, R.; Hadjigol, S.; Freeman, J.S.; Vaillancourt, R.E.; Zhu, P.; et al. Association mapping for wood quality and growth traits in Eucalyptus globulus ssp. globulus Labill identifies nine stable marker-trait associations for seven traits. Tree Genet. Genom. 2014, 10, 1661–1678. [Google Scholar] [CrossRef]
- Karger, D.N.; Conrad, O.; Böhner, J.; Kawohl, T.; Kreft, H.; Soria-Auza, R.W.; Zimmermann, N.E.; Linder, H.P.; Kessler, M. Climatologies at high resolution for the earth’s land surface areas. Sci. Data 2017, 4, 170122. [Google Scholar] [CrossRef]
- Busby, J.R. BIOCLIM-a bioclimate analysis and prediction system. Plant Prot Q 1991, 6, 8–9. [Google Scholar]
- Hijmans, R.J. Raster: Geographic Data Analysis and Modeling; R Package Version 2.9-22; R Foundation for Statistical Computing: Vienna, Austria, 2019. [Google Scholar]
- Ellis, N.; Smith, S.J.; Pitcher, C.R. Gradient forests: Calculating importance gradients on physical predictors. Ecology 2012, 93, 156–168. [Google Scholar] [CrossRef]
- Lotterhos, K.E.; Whitlock, M.C. Evaluation of demographic history and neutral parameterization on the performance of FST outlier tests. Mol. Ecol. 2014, 23, 2178–2192. [Google Scholar] [CrossRef]
- Dray, S.; Bauman, D.; Blanchet, G.; Borcard, D.C.S.; Guenard, G.; Jombart, T.; Larocque, G.; Legendre, P.; Madi, N.; Wagner, H. Adespatial: Multivariate Multiscale Spatial Analysis; R Package Version 0.3-4; R Foundation for Statistical Computing: Vienna, Austria, 2019. [Google Scholar]
- Manel, S.; Poncet, B.N.; Legendre, P.; Gugerli, F.; Holderegger, R. Common factors drive adaptive genetic variation at different spatial scales in Arabis alpina. Mol. Ecol. 2010, 19, 3824–3835. [Google Scholar] [CrossRef]
- Sork, V.L.; Aitken, S.N.; Dyer, R.J.; Eckert, A.J.; Legendre, P.; Neale, D.B. Putting the landscape into the genomics of trees: Approaches for understanding local adaptation and population responses to changing climate. Tree Genet. Genom. 2013, 9, 901–911. [Google Scholar] [CrossRef]
- van Rensburg, A.J.; Cortazar-Chinarro, M.; Laurila, A.; Van Buskirk, J. Adaptive genomic variation associated with environmental gradients along a latitudinal cline in Rana temporaria. bioRxiv 2018. [Google Scholar] [CrossRef]
- Gugger, P.F.; Liang, C.T.; Sork, V.L.; Hodgskiss, P.; Wright, J.W. Applying landscape genomic tools to forest management and restoration of Hawaiian koa (Acacia koa) in a changing environment. Evol. Appl. 2017, 11, 231–242. [Google Scholar] [CrossRef] [PubMed]
- Myburg, A.A.; Grattapaglia, D.; Tuskan, G.A.; Hellsten, U.; Hayes, R.D.; Grimwood, J.; Jenkins, J.; Lindquist, E.; Tice, H.; Bauer, D.; et al. The genome of Eucalyptus grandis. Nature 2014, 510, 356–362. [Google Scholar] [CrossRef]
- Szkiba, D.; Kapun, M.; Haeseler, A.v.; Gallach, M. SNP2GO: Functional analysis of genome-wide association studies. Genetics 2014, 197, 285–289. [Google Scholar] [CrossRef]
- Toro, M.; Silió, L.; Rodriguez, M.; Soria, F.; Toval, G. Genetic analysis of survival to drought in Eucalyptus globulus in Spain. In Proceedings of the 6th World Congress on Genetics Applied to Livestock Production, Armidale, Australia, 11–16 January 1998; pp. 499–502. [Google Scholar]
- Oksanen, J.; Blanchet, F.; Friendly, M.; Kindt, R.; Legendre, P.; McGlinn, D.; Minchin, P.; O’Hara, R.; Simpson, G.; Solymos, P.; et al. Vegan: Community Ecology Package; R Package Version 2.5-4; R Foundation for Statistical Computing: Vienna, Austria, 2019. [Google Scholar]
- Legendre, P.; Legendre, L.F. Numerical Ecology; Elsevier: Amsterdam, The Netherlands, 2012; Volume 24. [Google Scholar]
- Borcard, D.; Gillet, F.; Legendre, P. Numerical Ecology with R; Springer Science: New York, NY, USA, 2018. [Google Scholar]
- Riahi, K.; Rao, S.; Krey, V.; Cho, C.; Chirkov, V.; Fischer, G.; Kindermann, G.; Nakicenovic, N.; Rafaj, P. RCP 8.5—A scenario of comparatively high greenhouse gas emissions. Clim. Chang. 2011, 109, 33. [Google Scholar] [CrossRef]
- Harrison, P.A. Climate change and the suitability of local and non-local species for restoration. Ecol. Manage. Restor. 2021, 22, 75–91. [Google Scholar] [CrossRef]
- Bay, R.A.; Harrigan, R.J.; Underwood, V.L.; Gibbs, H.L.; Smith, T.B.; Ruegg, K. Genomic signals of selection predict climate-driven population declines in a migratory bird. Science 2018, 359, 83–86. [Google Scholar] [CrossRef]
- Booth, T.H. Going nowhere fast: A review of seed dispersal in eucalypts. Aust. J. Bot. 2017, 65, 401–410. [Google Scholar] [CrossRef]
- Jordan, G.J.; Harrison, P.A.; Worth, J.R.P.; Williamson, G.J.; Kirkpatrick, J.B. Palaeoendemic plants provide evidence for persistence of open, well-watered vegetation since the Cretaceous. Glob. Ecol. Biogeogr. 2016, 25, 127–140. [Google Scholar] [CrossRef]
- Hancock, A.M.; Brachi, B.; Faure, N.; Horton, M.W.; Jarymowycz, L.B.; Sperone, F.G.; Toomajian, C.; Roux, F.; Bergelson, J. Adaptation to climate across the Arabidopsis thaliana genome. Science 2011, 334, 83–86. [Google Scholar] [CrossRef]
- Westengen, O.T.; Berg, P.R.; Kent, M.P.; Brysting, A.K. Spatial structure and climatic adaptation in African maize revealed by surveying SNP diversity in relation to global breeding and landrace panels. PLoS ONE 2012, 7, e47832. [Google Scholar] [CrossRef]
- Hornoy, B.; Pavy, N.; Gérardi, S.; Beaulieu, J.; Bousquet, J. Genetic adaptation to climate in white spruce involves small to moderate allele frequency shifts in functionally diverse genes. Genome Biol. Evol. 2015, 7, 3269–3285. [Google Scholar] [CrossRef]
- Csilléry, K.; Lalagüe, H.; Vendramin, G.G.; González-Martínez, S.C.; Fady, B.; Oddou-Muratorio, S. Detecting short spatial scale local adaptation and epistatic selection in climate-related candidate genes in European beech (Fagus sylvatica) populations. Mol. Ecol. 2014, 23, 4696–4708. [Google Scholar] [CrossRef]
- Chen, J.; Källman, T.; Ma, X.; Gyllenstrand, N.; Zaina, G.; Morgante, M.; Bousquet, J.; Eckert, A.; Wegrzyn, J.; Neale, D.; et al. Disentangling the roles of history and local selection in shaping clinal variation of allele frequencies and gene expression in Norway spruce (Picea abies). Genetics 2012, 191, 865–881. [Google Scholar] [CrossRef]
- De La Torre, A.R.; Wilhite, B.; Neale, D.B. Environmental genome-wide association reveals climate adaptation is shaped by subtle to moderate allele frequency shifts in loblolly pine. Genome Biol. Evol. 2019, 11, 2976–2989. [Google Scholar] [CrossRef]
- Steane, D.A.; Potts, B.M.; McLean, E.H.; Collins, L.; Holland, B.R.; Prober, S.M.; Stock, W.D.; Vaillancourt, R.E.; Byrne, M. Genomic scans across three eucalypts suggest that adaptation to aridity is a genome-wide phenomenon. Genome Biol. Evol. 2017, 9, 253–265. [Google Scholar] [CrossRef]
- Noonan, J.P.; McCallion, A.S. Genomics of long-range regulatory elements. Annu. Rev. Genom. Hum. Genet. 2010, 11, 1–23. [Google Scholar] [CrossRef]
- Choat, B.; Brodribb, T.J.; Brodersen, C.R.; Duursma, R.A.; López, R.; Medlyn, B.E. Triggers of tree mortality under drought. Nature 2018, 558, 531–539. [Google Scholar] [CrossRef]
- Chaves, M.M.; Maroco, J.P.; Pereira, J.S. Understanding plant responses to drought—From genes to the whole plant. Funct. Plant Biol. 2003, 30, 239–264. [Google Scholar] [CrossRef]
- Piedallu, C.; Gégout, J.C.; Perez, V.; Lebourgeois, F. Soil water balance performs better than climatic water variables in tree species distribution modelling. Global Ecol. Biogeogr. 2013, 22, 470–482. [Google Scholar] [CrossRef]
- Anderegg, L.D.L.; Anderegg, W.R.L.; Berry, J.A. Not all droughts are created equal: Translating meteorological drought into woody plant mortality. Tree Physiol. 2013, 33, 672–683. [Google Scholar] [CrossRef]
- Mitchell, P.J.; O’Grady, A.P.; Tissue, D.T.; Worledge, D.; Pinkard, E.A. Co-ordination of growth, gas exchange and hydraulics define the carbon safety margin in tree species with contrasting drought strategies. Tree Physiol. 2014, 34, 443–458. [Google Scholar] [CrossRef]
- Allen, C.D.; Macalady, A.K.; Chenchouni, H.; Bachelet, D.; McDowell, N.; Vennetier, M.; Kitzberger, T.; Rigling, A.; Breshears, D.D.; Hogg, E.T. A global overview of drought and heat-induced tree mortality reveals emerging climate change risks for forests. For. Ecol. Manag. 2010, 259, 660–684. [Google Scholar] [CrossRef]
- Mitchell, P.J.; Battaglia, M.; Pinkard, E.A. Counting the costs of multiple stressors: Is the whole greater than the sum of the parts? Tree Physiol. 2013, 33, 447–450. [Google Scholar] [CrossRef]
- Logan, J.A.; Régnière, J.; Powell, J.A. Assessing the impacts of global warming on forest pest dynamics. Front. Ecol. Environ. 2003, 1, 130–137. [Google Scholar] [CrossRef]
- Gazol, A.; Camarero, J.J.; Anderegg, W.R.L.; Vicente-Serrano, S.M. Impacts of droughts on the growth resilience of Northern Hemisphere forests. Global Ecol. Biogeogr. 2017, 26, 166–176. [Google Scholar] [CrossRef]
- Ammitzboll, H.; Vaillancourt, R.E.; Potts, B.M.; Harrison, P.A.; Brodribb, T.; Sussmilch, F.C.; Freeman, J.S. Independent genetic control of drought resistance, recovery, and growth of Eucalyptus globulus seedlings. Plant Cell Environ. 2020, 43, 103–115. [Google Scholar] [CrossRef]
- Teulières, C.; Bossinger, G.; Moran, G.; Marque, C. Stress studies in Eucalyptus. Plant Stress 2007, 1, 197–215. [Google Scholar]
- Gomulkiewicz, R.; Thompson, J.N.; Holt, R.D.; Nuismer, S.L.; Hochberg, M.E. Hot spots, cold spots, and the geographic mosaic theory of coevolution. Am. Nat. 2000, 156, 156–174. [Google Scholar] [CrossRef]
- Barbour, R.C.; O’Reilly-Wapstra, J.M.; Little, D.W.D.; Jordan, G.J.; Steane, D.A.; Humphreys, J.R.; Bailey, J.K.; Whitham, T.G.; Potts, B.M. A geographic mosaic of genetic variation within a foundation tree species and its community-level consequences. Ecology 2009, 90, 1762–1772. [Google Scholar] [CrossRef] [PubMed]
- Hamilton, M.; Williams, D.; Tilyard, P.; Pinkard, E.; Wardlaw, T.; Glen, M.; Vaillancourt, R.; Potts, B. A latitudinal cline in disease resistance of a host tree. Heredity 2013, 110, 372–379. [Google Scholar] [CrossRef] [PubMed]
- Freeman, J.S.; Hamilton, M.G.; Lee, D.J.; Pegg, G.S.; Brawner, J.T.; Tilyard, P.A.; Potts, B.M. Comparison of host susceptibility to native and exotic pathogens provides evidence for pathogen imposed selection in forest trees. New Phytol. 2018, 221, 2261–2272. [Google Scholar] [CrossRef] [PubMed]
- Carnicer, J.; Barbeta, A.; Sperlich, D.; Coll, M.; Peñuelas, J. Contrasting trait syndromes in angiosperms and conifers are associated with different responses of tree growth to temperature on a large scale. Front Plant Sci. 2013, 4, 409. [Google Scholar] [CrossRef]
- Mimura, M.; Aitken, S. Adaptive gradients and isolation-by-distance with postglacial migration in Picea sitchensis. Heredity 2007, 99, 224–232. [Google Scholar] [CrossRef]
- Rehfeldt, G.E.; Tchebakova, N.M.; Parfenova, Y.I.; Wykoff, W.R.; Kuzmina, N.A.; Milyutin, L.I. Intraspecific responses to climate in Pinus sylvestris. Glob. Chang. Biol. 2002, 8, 912–929. [Google Scholar] [CrossRef]
- Moles, A.T.; Perkins, S.E.; Laffan, S.W.; Flores-Moreno, H.; Awasthy, M.; Tindall, M.L.; Sack, L.; Pitman, A.; Kattge, J.; Aarssen, L.W. Which is a better predictor of plant traits: Temperature or precipitation? J. Veg. Sci. 2014, 25, 1167–1180. [Google Scholar] [CrossRef]
- Evans, L.M.; Slavov, G.T.; Rodgers-Melnick, E.; Martin, J.; Ranjan, P.; Muchero, W.; Brunner, A.M.; Schackwitz, W.; Gunter, L.; Chen, J.-G. Population genomics of Populus trichocarpa identifies signatures of selection and adaptive trait associations. Nat. Genet. 2014, 46, 1089. [Google Scholar] [CrossRef]
- De Kort, H.; Vandepitte, K.; Bruun, H.H.; Closset-Kopp, D.; Honnay, O.; Mergeay, J. Landscape genomics and a common garden trial reveal adaptive differentiation to temperature across Europe in the tree species Alnus glutinosa. Mol. Ecol. 2014, 23, 4709–4721. [Google Scholar] [CrossRef]
- Blackman, C.J.; Aspinwall, M.J.; Tissue, D.T.; Rymer, P.D. Genetic adaptation and phenotypic plasticity contribute to greater leaf hydraulic tolerance in response to drought in warmer climates. Tree Physiol. 2017, 37, 583–592. [Google Scholar] [CrossRef]
- Riordan, E.C.; Gugger, P.F.; Ortego, J.; Smith, C.; Gaddis, K.; Thompson, P.; Sork, V.L. Association of genetic and phenotypic variability with geography and climate in three southern California oaks. Am. J. Bot. 2016, 103, 73–85. [Google Scholar] [CrossRef]
- Harrison, P.A. Integrating Climate Change into Conservation and Restoration Strategies: The Case of the Tasmanian Eucalypts; University of Tasmania: Tasmania, Australia, 2017. [Google Scholar]
- Butt, N.; Pollock, L.J.; McAlpine, C.A. Eucalypts face increasing climate stress. Ecol. Evol. 2013, 3, 5011–5022. [Google Scholar] [CrossRef]
- Jump, A.S.; Peñuelas, J. Running to stand still: Adaptation and the response of plants to rapid climate change. Ecol. Lett. 2005, 8, 1010–1020. [Google Scholar] [CrossRef]
- Beckage, B.; Osborne, B.; Gavin, D.G.; Pucko, C.; Siccama, T.; Perkins, T. A rapid upward shift of a forest ecotone during 40 years of warming in the Green Mountains of Vermont. Proc. Natl. Acad. Sci. USA 2008, 105, 4197–4202. [Google Scholar] [CrossRef]
- Chapin, F.S.; Bloom, A.J.; Field, C.B.; Waring, R.H. Plant responses to multiple environmental factors. Bioscience 1987, 37, 49–57. [Google Scholar] [CrossRef]
- Larter, M.; Brodribb, T.J.; Pfautsch, S.; Burlett, R.; Cochard, H.; Delzon, S. Extreme aridity pushes trees to their physical limits. Plant Physiol. 2015, 168, 804–807. [Google Scholar] [CrossRef]
- Box, E.O. Vegetation analogs and differences in the northern and southern hemispheres: A global comparison. Plant Ecol. 2002, 163, 139–154. [Google Scholar] [CrossRef]
- Harris, R.M.B.; Carter, O.; Gilfedder, L.; Porfirio, L.L.; Lee, G.; Bindoff, N.L. Noah’s Ark conservation will not preserve threatened ecological communities under climate change. PLoS ONE 2015, 10, e0124014. [Google Scholar] [CrossRef]
- Trisurat, Y.; Shrestha, R.P.; Kjelgren, R. Plant species vulnerability to climate change in Peninsular Thailand. Appl. Geogr. 2011, 31, 1106–1114. [Google Scholar] [CrossRef]
- Bradbury, D.; Smithson, A.; Krauss, S.L. Signatures of diversifying selection at EST-SSR loci and association with climate in natural Eucalyptus populations. Mol. Ecol. 2013, 22, 5112–5129. [Google Scholar] [CrossRef]
- Ricklefs, R.E.; He, F. Region effects influence local tree species diversity. Proc. Natl. Acad. Sci. USA 2016, 113, 674–679. [Google Scholar] [CrossRef]
- Couvreur, T.L.P.; Porter-Morgan, H.; Wieringa, J.J.; Chatrou, L.W. Little ecological divergence associated with speciation in two African rain forest tree genera. BMC Evol. Biol. 2011, 11, 296. [Google Scholar] [CrossRef]
- Ortiz-Rodríguez, A.E.; Ramírez-Barahona, S.; González Hernández, D.; Lorea-Hernández, F. Past climatic fluctuations are associated with morphological differentiation in the cloud forest endemic tree Ocotea psychotrioides (Lauraceae). Plant Syst. Evol. 2018, 304, 607–617. [Google Scholar] [CrossRef]
- Song, Z.; Zhang, M.; Li, F.; Weng, Q.; Zhou, C.; Li, M.; Li, J.; Huang, H.; Mo, X.; Gan, S. Genome scans for divergent selection in natural populations of the widespread hardwood species Eucalyptus grandis (Myrtaceae) using microsatellites. Sci. Rep. 2016, 6, 34941. [Google Scholar] [CrossRef]
- Gardner, A.S.; Maclean, I.M.D.; Gaston, K.J. Climatic predictors of species distributions neglect biophysiologically meaningful variables. Divers. Distrib. 2019, 25, 1318–1333. [Google Scholar] [CrossRef]
- Caccianiga, M.; Andreis, C.; Armiraglio, S.; Leonelli, G.; Pelfini, M.; Sala, D. Climate continentality and treeline species distribution in the Alps. Plant Biosyst. 2008, 142, 66–78. [Google Scholar] [CrossRef]
- Ikeda, D.H.; Max, T.L.; Allan, G.J.; Lau, M.K.; Shuster, S.M.; Whitham, T.G. Genetically informed ecological niche models improve climate change predictions. Glob. Chang. Biol. 2017, 23, 164–176. [Google Scholar] [CrossRef]
- Razgour, O.; Forester, B.; Taggart, J.B.; Bekaert, M.; Juste, J.; Ibáñez, C.; Puechmaille, S.J.; Novella-Fernandez, R.; Alberdi, A.; Manel, S. Considering adaptive genetic variation in climate change vulnerability assessment reduces species range loss projections. Proc. Natl. Acad. Sci. USA 2019, 116, 10418–10423. [Google Scholar] [CrossRef]
- Perez-Navarro, M.A.; Broennimann, O.; Esteve, M.A.; Moya-Perez, J.M.; Carreño, M.F.; Guisan, A.; Lloret, F. Temporal variability is key to modelling the climatic niche. Divers. Distrib. 2021, 27, 473–484. [Google Scholar] [CrossRef]
- Gilman, S.E.; Urban, M.C.; Tewksbury, J.; Gilchrist, G.W.; Holt, R.D. A framework for community interactions under climate change. Trends Ecol. Evol. 2010, 25, 325–331. [Google Scholar] [CrossRef]
- Bolte, A.; Hilbrig, L.; Grundmann, B.; Kampf, F.; Brunet, J.; Roloff, A. Climate change impacts on stand structure and competitive interactions in a southern Swedish spruce–beech forest. Eur. J. For. Res. 2010, 129, 261–276. [Google Scholar] [CrossRef]
- Boucher, D.; Gauthier, S.; Thiffault, N.; Marchand, W.; Girardin, M.; Urli, M. How climate change might affect tree regeneration following fire at northern latitudes: A review. New For. 2020, 51, 543–571. [Google Scholar] [CrossRef]
- Battaglia, M.; Bruce, J. Direct climate change impacts on growth and drought risk in blue gum (Eucalyptus globulus) plantations in Australia. Aust. For. 2017, 80, 216–227. [Google Scholar] [CrossRef]
- Pinkard, E.A.; Paul, K.; Battaglia, M.; Bruce, J. Vulnerability of plantation carbon stocks to defoliation under current and future climates. Forests 2014, 5, 1224–1242. [Google Scholar] [CrossRef]
- Chen, Q.-L.; Hu, H.-W.; Yan, Z.-Z.; Li, C.-Y.; Nguyen, B.-A.T.; Zhu, Y.-G.; He, J.-Z. Precipitation increases the abundance of fungal plant pathogens in Eucalyptus phyllosphere. Environ. Microbiol. 2021, 23, 7688–7700. [Google Scholar] [CrossRef]
- Shaw, R.G. From the past to the future: Considering the value and limits of evolutionary prediction. Am. Nat. 2019, 193, 1–10. [Google Scholar] [CrossRef]
- Frank, A.; Howe, G.T.; Sperisen, C.; Brang, P.; Clair, J.B.S.; Schmatz, D.R.; Heiri, C. Risk of genetic maladaptation due to climate change in three major European tree species. Glob. Chang. Biol. 2017, 23, 5358–5371. [Google Scholar] [CrossRef]
- Bradley St Clair, J.; Howe, G.T. Genetic maladaptation of coastal Douglas-fir seedlings to future climates. Glob. Chang. Biol. 2007, 13, 1441–1454. [Google Scholar] [CrossRef]
- Booth, T.H.; Pryor, L.D. Climatic requirements of some commercially important eucalypt species. For. Ecol. Manag. 1991, 43, 47–60. [Google Scholar] [CrossRef]
- McGowen, M.; Potts, B.; Vaillancourt, R.; Gore, P.; Williams, D.; Pilbeam, D. The genetic control of sexual reproduction in Eucalyptus globulus. In Proceedings of the Eucalyptus in a Changing World. International IUFRO Conference, Aveiro, Portugal, 11–15 October 2004. [Google Scholar]
- Rix, K.D.; Gracie, A.J.; Potts, B.M.; Brown, P.H.; Gore, P.L. Genetic control of Eucalyptus globulus seed germination. Ann. For. Sci. 2015, 72, 457–467. [Google Scholar] [CrossRef]
- Larcombe, M.J.; Silva, J.S.; Vaillancourt, R.E.; Potts, B.M. Assessing the invasive potential of Eucalyptus globulus in Australia: Quantification of wildling establishment from plantations. Biol. Invasions 2013, 15, 2763–2781. [Google Scholar] [CrossRef]
- Catry, F.; Moreira, F.; Deus, E.; Silva, J.; Aguas, A. Assessing the extent and the environmental drivers of Eucalyptus globulus wildling establishment in Portugal: Results from a countrywide survey. Biol. Invasions 2015, 17, 3163–3181. [Google Scholar] [CrossRef]
- Jiang, M.; Kelly, J.W.G.; Atwell, B.J.; Tissue, D.T.; Medlyn, B.E. Drought by CO2 interactions in trees: A test of the water savings mechanism. New Phytol. 2021, 230, 1421–1434. [Google Scholar] [CrossRef]
- Crous, K.Y.; Quentin, A.G.; Lin, Y.-S.; Medlyn, B.E.; Williams, D.G.; Barton, C.V.M.; Ellsworth, D.S. Photosynthesis of temperate Eucalyptus globulus trees outside their native range has limited adjustment to elevated CO2 and climate warming. Glob. Chang. Biol. 2013, 19, 3790–3807. [Google Scholar] [CrossRef]
- Bebber, D.P. Range-expanding pests and pathogens in a warming world. Annu. Rev. Phytopathol. 2015, 53, 335–356. [Google Scholar] [CrossRef]
- Paine, T.D.; Steinbauer, M.J.; Lawson, S.A. Native and exotic pests of Eucalyptus: A worldwide perspective. Annu. Rev. Entomol. 2011, 56, 181–201. [Google Scholar] [CrossRef]
- Bailey, T.; Harrison, P.; Davidson, N.; Weller-Wong, A.; Tilyard, P.; Steane, D.; Vaillancourt, R.; Potts, B. Embedding genetics experiments in restoration to guide plant choice for a degraded landscape with a changing climate. Ecol. Manag. Restor. 2021, 22, 92–105. [Google Scholar] [CrossRef]
- Janes, J.K.; Hamilton, J.A. Mixing it up: The role of hybridization in forest management and conservation under climate change. Forests 2017, 8, 237. [Google Scholar] [CrossRef]
- Butler, J.B.; Freeman, J.S.; Potts, B.M.; Vaillancourt, R.E.; Kahrood, H.V.; Ades, P.K.; Rigault, P.; Tibbits, J.F.G. Patterns of Genomic Diversity and Linkage Disequilibrium across the Disjunct Range of the Australian Forest Tree Eucalyptus globulus. 2022; manuscript submitted for publication. [Google Scholar]
- Chang, C.C.; Chow, C.C.; Tellier, L.C.; Vattikuti, S.; Purcell, S.M.; Lee, J.J. Second-generation PLINK: Rising to the challenge of larger and richer datasets. Gigascience 2015, 4, 7. [Google Scholar] [CrossRef]
- Pavan, S.; Delvento, C.; Ricciardi, L.; Lotti, C.; Ciani, E.; D’Agostino, N. Recommendations for choosing the genotyping method and best practices for quality control in crop genome-wide association studies. Front. Genet 2020, 11, 447. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).