Antagonistic Potential of Native Trichoderma spp. against Phytophthora cinnamomi in the Control of Holm Oak Decline in Dehesas Ecosystems
Abstract
1. Introduction
2. Materials and Methods
2.1. Experimental Design and Sample Collection
2.2. Trichoderma and Phytophthora Isolation
2.3. Mycelial Growth Inhibition
2.4. Mycoparasitism
2.5. Molecular Identification of Trichoderma and Phytophthora Strains
2.6. Tree Defoliation Data Collection
2.7. Statistical Analysis
3. Results
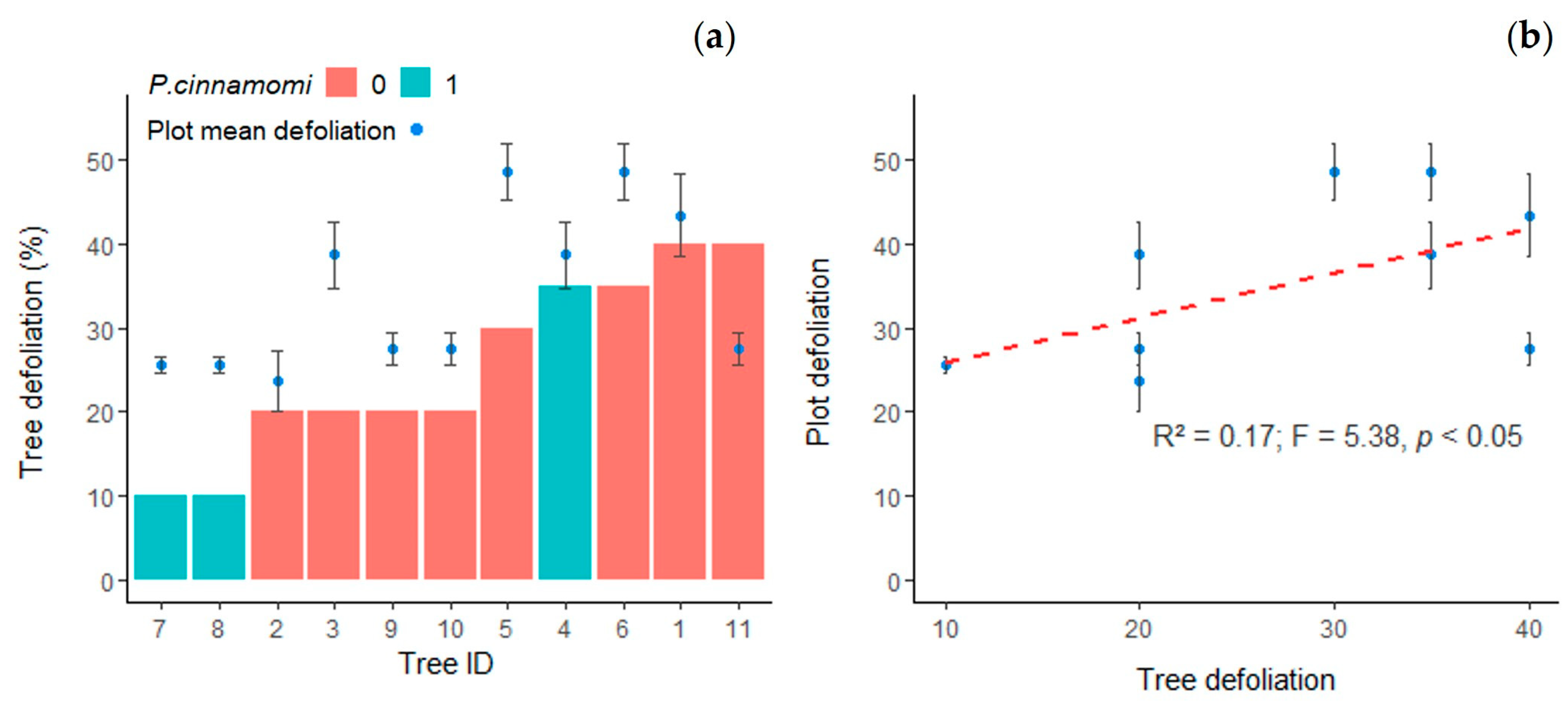
3.1. Health Status of Selected Trees
3.2. Isolation of Trichoderma spp. and Oomycete Strains
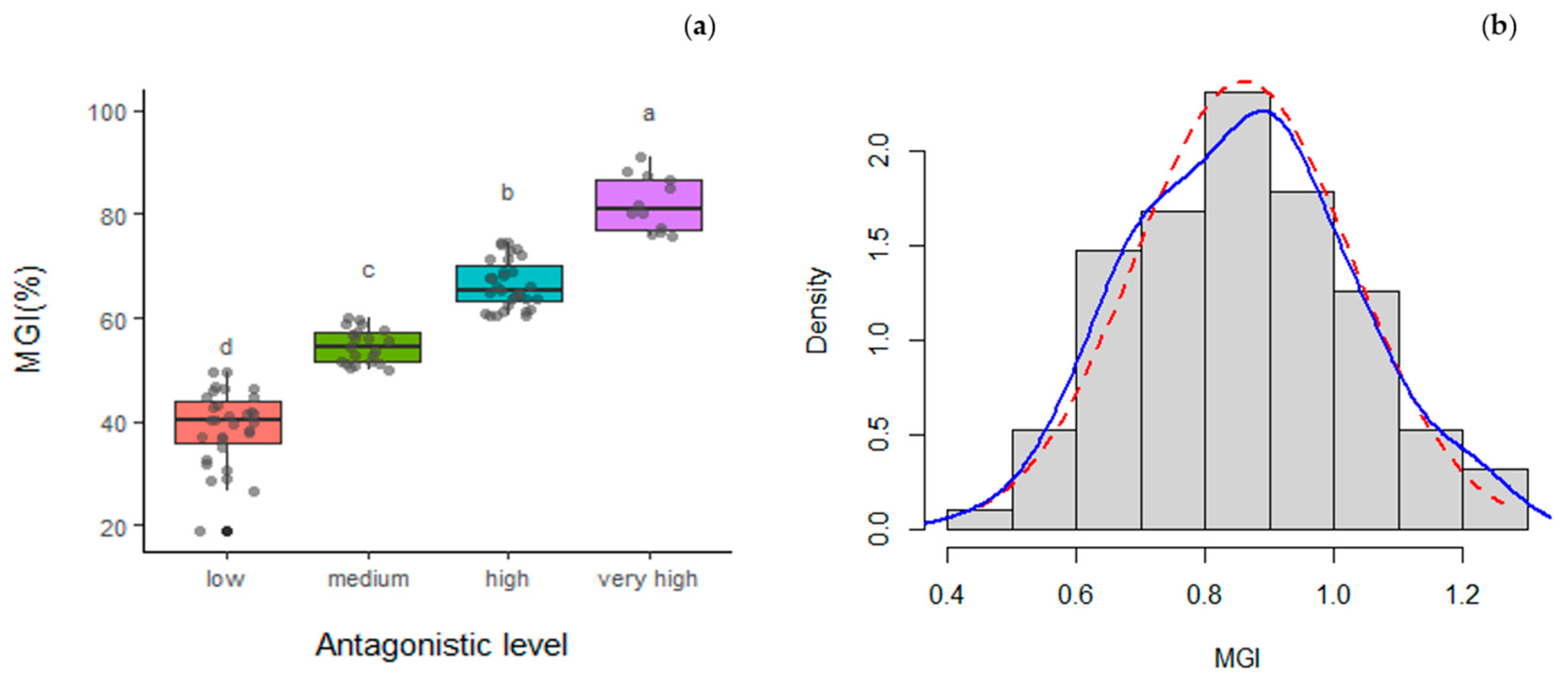
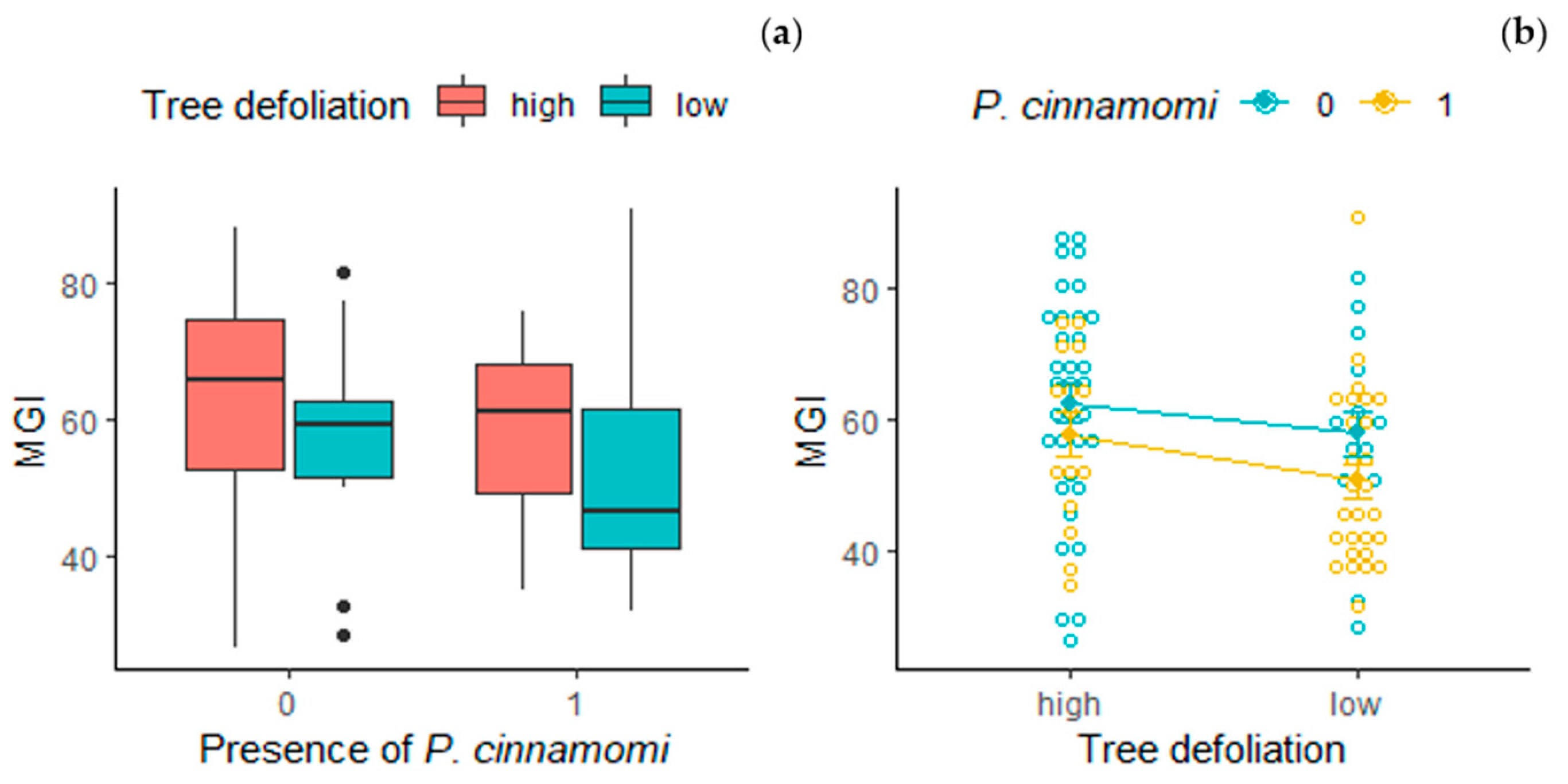
3.3. Mycelial Growth Inhibition
3.4. Molecular Identification of Trichoderma spp. and the Phytophthora Pathogen
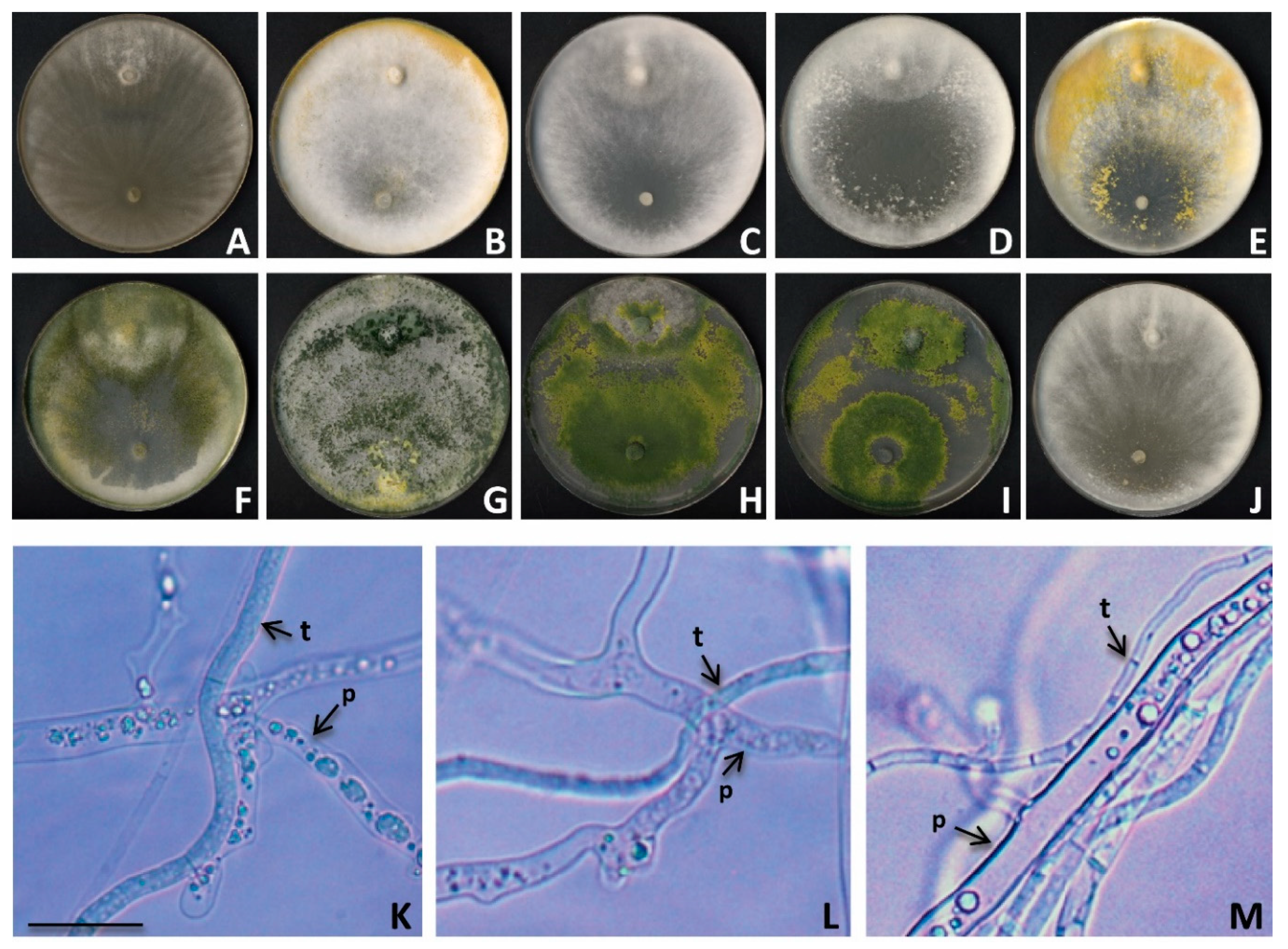
3.5. Mycoparasitism
4. Discussion
4.1. Molecular Characterization
4.2. Trichoderma spp. Inhibiting P. cinnamomi
4.3. Tree Health Status and Soil Microbial Community
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Delgado-Baquerizo, M.; Reich, P.B.; Trivedi, C.; Eldridge, D.J.; Abades, S.; Alfaro, F.D.; Bastida, F.; Berhe, A.A.; Cutler, N.A.; Gallardo, A.; et al. Multiple elements of soil biodiversity drive ecosystem functions across biomes. Nat. Ecol. Evol. 2020, 4, 210–220. [Google Scholar] [CrossRef] [PubMed]
- Deiner, K.; Bik, H.M.; Mächler, E.; Seymour, M.; Lacoursière-Roussel, A.; Altermatt, F.; Creer, S.; Bista, I.; Lodge, D.M.; De Vere, N.; et al. Environmental DNA metabarcoding: Transforming how we survey animal and plant communities. Mol. Ecol. 2017, 26, 5872–5895. [Google Scholar] [CrossRef]
- Tedersoo, L.; Bahram, M.; Zobel, M. How mycorrhizal associations drive plant population and community biology. Science 2020, 367, eaba1223. [Google Scholar] [CrossRef] [PubMed]
- Pérez-De-Luque, A.; Tille, S.; Johnson, I.; Pascual-Pardo, D.; Ton, J.; Cameron, D.D. The interactive effects of arbuscular mycorrhiza and plant growth-promoting rhizobacteria synergistically enhance host plant defences against pathogens. Sci. Rep. 2017, 7, 16409. [Google Scholar] [CrossRef]
- Tamayo-Vélez, Á.; Osorio, N.W. Soil fertility improvement by litter decomposition and inoculation with the fungus Mortierella sp. in avocado plantations of Colombia. Commun. Soil Sci. Plant Anal. 2018, 49, 139–147. [Google Scholar] [CrossRef]
- Turbé, A.; de Toni, A.; Benito, P.; Lavelle, P.; Lavelle, P.; Ruiz Camacho, N.; van der Putten, W.H.; Labouze, E.; Mudgal, S. Soil Biodiversity: Functions, Threats and Tools for Policy Makers; European Commission DG ENV. Bioemco: Paris, France, 2010; p. 251. [Google Scholar]
- Wall, D.H.; Nielsen, U.N.; Six, J. Soil biodiversity and human health. Nature 2015, 528, 69–76. [Google Scholar] [CrossRef] [PubMed]
- Belhaj, R.; McComb, J.; Burgess, T.I.; Hardy, G.E.S.J. Pathogenicity of 21 newly described Phytophthora species against seven Western Australian native plant species. Plant Pathol. 2018, 67, 1140–1149. [Google Scholar] [CrossRef]
- Jung, T.; Pérez-Sierra, A.; Durán, A.; Jung, M.H.; Balci, Y.; Scanu, B. Canker and decline diseases caused by soil- and airborne Phytophthora species in forests and woodlands. Pers. Mol. Phylogeny Evol. Fungi 2018, 40, 182–220. [Google Scholar] [CrossRef]
- Trumbore, S.E.; Brando, P.M.; Hartmann, H. Forest health and global change. Science 2015, 349, 814–818. [Google Scholar] [CrossRef]
- Rodríguez-Calcerrada, J.; Sancho-Knapik, D.; Martin-StPaul, N.K.; Limousin, J.-M.; McDowell, N.G.; Gil-Pelegrín, E. Drought-Induced Oak Decline—Factors Involved, Physiological Dysfunctions, and Potential Attenuation by Forestry Practices. In Oaks Physiological Ecology. Exploring the Functional Diversity of Genus Quercus L.; Gil-Pelegrín, E., Peguero-Pina, J.J., Sancho-Knapik, D., Eds.; Tree Physiology; Springer International Publishing: Cham, Switzerland, 2017; pp. 419–451. ISBN 978-3-319-69099-5. [Google Scholar]
- Ruiz-Gómez, F.J.; Pérez-De-Luque, A.; Navarro-Cerrillo, R.M. The involvement of Phytophthora root rot and drought stress in holm oak decline: From ecophysiology to microbiome influence. Curr. For. Rep. 2019, 5, 251–266. [Google Scholar] [CrossRef]
- Carrasco Gotarredona, Á.; Fernández Cancio, Á.; Trapero Casas, A.; López Pantoja, G.; Sánchez Osorio, I.; Ruiz Navarro, J.M.; Jiménez Molina, J.J.; Domínguez Nevado, L.; Romero Martín, M.Á.; Carbonero Muñoz, M.D.; et al. Procesos de Decaimiento Forestal (la Seca): Situación del Conocimiento, 1st ed.; Consejería de Medio Ambiente: Córdoba, Spain, 2009; ISBN 978-84-92807-29-1. [Google Scholar]
- Sánchez-Cuesta, R.; Navarro-Cerrillo, R.M.; Quero, J.L.; Ruiz-Gómez, F.J. Small-scale abiotic factors influencing the spatial distribution of Phytophthora cinnamomi under declining Quercus ilex trees. Forests 2020, 11, 375. [Google Scholar] [CrossRef]
- Ruiz-Gómez, F.J.; Navarro-Cerrillo, R.M.; Sánchez-Cuesta, R.; Pérez-De-Luque, A. Histopathology of infection and colonization of Quercus ilex fine roots by Phytophthora cinnamomi. Plant Pathol. 2015, 64, 605–616. [Google Scholar] [CrossRef]
- Burgess, T.I.; Scott, J.; McDougall, K.L.; Stukely, M.J.C.; Crane, C.; Dunstan, W.A.; Brigg, F.; Andjic, V.; White, D.; Rudman, T.; et al. Current and projected global distribution of Phytophthora cinnamomi, one of the world’s worst plant pathogens. Glob. Chang. Biol. 2016, 23, 1661–1674. [Google Scholar] [CrossRef]
- Hardham, A.R.; Blackman, L.M. Phytophthora cinnamomi. Mol. Plant Pathol. 2018, 19, 260–285. [Google Scholar] [CrossRef]
- Jiggins, J.; Roling, N. Adaptive management: Potential and limitations for ecological governance. Int. J. Agric. Resour. Gov. Ecol. 2000, 1, 28. [Google Scholar] [CrossRef]
- Sharma, A.; Sharma, P. Trichoderma: Host Pathogen Interactions and Applications, 1st ed.; Rhizosphere Biology; Springer: Singapore, 2020; p. 319. ISBN 9789811533204. [Google Scholar]
- Vinale, F.; Sivasithamparam, K.; Ghisalberti, E.L.; Marra, R.; Woo, S.L.; Lorito, M. Trichoderma–plant–pathogen interactions. Soil Biol. Biochem. 2008, 40, 1–10. [Google Scholar] [CrossRef]
- Aleandri, M.P.; Chilosi, G.; Bruni, N.; Tomassini, A.; Vettraino, A.M.; Vannini, A. Use of nursery potting mixes amended with local Trichoderma strains with multiple complementary mechanisms to control soil-borne diseases. Crop. Prot. 2015, 67, 269–278. [Google Scholar] [CrossRef]
- Chemeltorit, P.P.; Mutaqin, K.; Widodo, W. Combining Trichoderma hamatum THSW13 and Pseudomonas aeruginosa BJ10–86: A synergistic chili pepper seed treatment for Phytophthora capsici infested soil. Eur. J. Plant Pathol. 2016, 147, 157–166. [Google Scholar] [CrossRef]
- García-Núñez, H.G.; Martínez-Campos, Á.R.; Hermosa-Prieto, M.R.; Monte-Vázquez, E.; Aguilar-Ortigoza, C.J.; González-Esquivel, C.E. Caracterización morfológica y molecular de cepas nativas de Trichoderma y su potencial de biocontrol sobre Phytophthora infestans. Mex. J. Phytopathol. 2017, 35, 58–79. [Google Scholar] [CrossRef]
- Promwee, A.; Yenjit, P.; Issarakraisila, M.; Intana, W.; Chamswarng, C. Efficacy of indigenous Trichoderma harzianum in controlling Phytophthora leaf fall (Phytophthora palmivora) in Thai rubber trees. J. Plant Dis. Prot. 2017, 124, 41–50. [Google Scholar] [CrossRef]
- Widmer, T.L.; Johnson-Brousseau, S.; Kosta, K.; Ghosh, S.; Schweigkofler, W.; Sharma, S.; Suslow, K. Remediation of Phytophthora ramorum-infested soil with Trichoderma asperellum isolate 04-22 under ornamental nursery conditions. Biol. Control. 2018, 118, 67–73. [Google Scholar] [CrossRef]
- Ros, M.; Raut, I.; Santisima-Trinidad, A.B.; Pascual, J.A. Relationship of microbial communities and suppressiveness of Trichoderma fortified composts for pepper seedlings infected by Phytophthora nicotianae. PLoS ONE 2017, 12, e0174069. [Google Scholar] [CrossRef] [PubMed]
- Atanasova, L.; Le Crom, S.; Gruber, S.; Coulpier, F.; Seidl-Seiboth, V.; Kubicek, C.P.; Druzhinina, I.S. Comparative transcriptomics reveals different strategies of Trichoderma mycoparasitism. BMC Genom. 2013, 14, 121. [Google Scholar] [CrossRef] [PubMed]
- Reithner, B.; Ibarra-Laclette, E.; Mach, R.; Herrera-Estrella, A. Identification of mycoparasitism-related genes in Trichoderma atroviride. Appl. Environ. Microbiol. 2011, 77, 4361–4370. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, A.D.; Ousset, M.J.; Sosa, M.C. Biological control of Phytophthora collar rot of pear using regional Trichoderma strains with multiple mechanisms. Biol. Control. 2019, 135, 124–134. [Google Scholar] [CrossRef]
- Zeilinger, S.; Omann, M. Trichoderma biocontrol: Signal transduction pathways involved in host sensing and mycoparasitism. Gene Regul. Syst. Biol. 2007, 1, 227–234. [Google Scholar] [CrossRef]
- Dunstan, W.A.; Rudman, T.; Shearer, B.L.; Moore, N.A.; Paap, T.; Calver, M.C.; Dell, B.; Hardy, G.E.S.J. Containment and spot eradication of a highly destructive, invasive plant pathogen (Phytophthora cinnamomi) in natural ecosystems. Biol. Invasions 2009, 12, 913–925. [Google Scholar] [CrossRef]
- Corcobado, T.; Vivas, M.; Moreno, G.; Solla, A. Ectomycorrhizal symbiosis in declining and non-declining Quercus ilex trees infected with or free of Phytophthora cinnamomi. For. Ecol. Manag. 2014, 324, 72–80. [Google Scholar] [CrossRef]
- Ruiz-Gómez, F.J.; Navarro-Cerrillo, R.M.; Pérez-De-Luque, A.; Oβwald, W.; Vannini, A.; Morales-Rodríguez, C. Assessment of functional and structural changes of soil fungal and oomycete communities in holm oak declined dehesas through metabarcoding analysis. Sci. Rep. 2019, 9, 5315. [Google Scholar] [CrossRef]
- Marti, A.F.I.; Romero-Rodríguez, C.; Navarro-Cerrillo, R.M.; Abril, N.; Jorrín-Novo, J.V.; Dodd, R.S. Population genetic diversity of Quercus ilex subsp. ballota (Desf.) Samp. reveals divergence in recent and evolutionary migration rates in the Spanish dehesas. Forests 2018, 9, 337. [Google Scholar] [CrossRef]
- Valero Galván, J.; Jorrin Novo, J.J.; Cabrera, A.G.; Ariza, D.; García-Olmo, J.; Navarro-Cerrillo, R.M. Population variability based on the morphometry and chemical composition of the acorn in Holm oak (Quercus ilex subsp. ballota [Desf.] Samp.). Eur. J. For. Res. 2011, 131, 893–904. [Google Scholar] [CrossRef]
- Hart, M.M.; Antunes, P.M.; Chaudhary, V.B.; Abbott, L.K. Fungal inoculants in the field: Is the reward greater than the risk? Funct. Ecol. 2018, 32, 126–135. [Google Scholar] [CrossRef]
- Eichhorn, J.; Roskams, P.; Potočić, N.; Timmermann, V.; Ferretti, M.; Mues, V.; Szepesi, A.; Durrant, D.; Seletković, I.; Schroeck, H.-W.; et al. Part IV: Visual assessment of crown condition and damaging agents. In UNECE ICP Forests Programme Co-ordinating Centre. Manual on Methods and Criteria for Harmonized Sampling, Assessment, Monitoring and Analysis of the Effects of Air Pollution on Forests; Thünen Institute of Forest Ecosystems: Eberswalde, Germany, 2017; p. 54. ISBN 978-3-86576-162-0. [Google Scholar]
- Zamora Rojas, E.; Andicoberry de los Reyes, S.; Sánchez Clemente, M.E. Anexo A.1. IV El decaimiento y la podredumbre radical en las dehesas andaluzas. In Ecosistemas de Dehesa: Desarrollo de Políticas y Herramientas para la Gestión y Conservación de la Biodiversidad (Life Biodehesa Project); Consejería de Medio Ambiente y Ordenación del Territorio: Sevilla, Spain, 2014. [Google Scholar]
- Hirte, W.F. The use of dilution plate method for the determination of soil microflora. 2. The qualitative demonstration of bacteria and actinomycetes. Zent. Bakteriol. Parasitenkd. Infekt. Hyg. 1969, 123, 167–178. [Google Scholar]
- Gil, S.V.; Pastor, S.; March, G. Quantitative isolation of biocontrol agents Trichoderma spp., Gliocladium spp. and actinomycetes from soil with culture media. Microbiol. Res. 2009, 164, 196–205. [Google Scholar] [CrossRef]
- Williams, J.; Clarkson, J.M.; Mills, P.R.; Cooper, R.M. A selective medium for quantitative reisolation of Trichoderma harzianum from Agaricus bisporus compost. Appl. Environ. Microbiol. 2003, 69, 4190–4191. [Google Scholar] [CrossRef]
- Erwin, D.C.; Ribeiro, O.K. Phytophthora Diseases Worldwide; APS Press: Washington, DC, USA, 1996; p. 592. ISBN 978-0-89054-212-5. [Google Scholar]
- Jeffers, S.N.; Martin, S.B. Comparison of two media selective for Phytophthora and Pythium species. Plant Dis. 1986, 70, 1038–1043. [Google Scholar] [CrossRef]
- Rahman, M.A.; Begum, M.F.; Alam, M.F. Screening of Trichoderma isolates as a biological control agent against Ceratocystis paradoxa causing pineapple disease of sugarcane. Mycobiology 2009, 37, 277–285. [Google Scholar] [CrossRef]
- Li, Y.; Sun, R.; Yu, J.; Saravanakumar, K.; Chen, J. Antagonistic and biocontrol potential of Trichoderma asperellum ZJSX5003 against the maize stalk rot pathogen Fusarium graminearum. Indian J. Microbiol. 2016, 56, 318–327. [Google Scholar] [CrossRef] [PubMed]
- Elías, R.; Arcos, O.; Arbeláez, G. Estudio del antagonismo de algunas especies de Trichoderma aisladas de suelos colombianos en el control de Fusarium oxysporum y Rhizoctonia solani. Agron. Colomb. 1993, 10, 52–61. [Google Scholar]
- Carbone, I.; Kohn, L.M. A Method for designing primer sets for speciation studies in filamentous ascomycetes. Mycologia 1999, 91, 553. [Google Scholar] [CrossRef]
- Jaklitsch, W.M.; Komon, M.; Kubicek, C.P.; Druzhinina, I.S. Hypocrea voglmayrii sp. nov. from the Austrian Alps represents a new phylogenetic clade in Hypocrea/Trichoderma. Mycologia 2005, 97, 1365–1378. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.J.; Whelen, S.; Hall, B.D. Phylogenetic relationships among ascomycetes: Evidence from an RNA polymerse II subunit. Mol. Biol. Evol. 1999, 16, 1799–1808. [Google Scholar] [CrossRef] [PubMed]
- Cooke, D.; Drenth, A.; Duncan, J.; Wagels, G.; Brasier, C. A Molecular Phylogeny of Phytophthora and related oomycetes. Fungal Genet. Biol. 2000, 30, 17–32. [Google Scholar] [CrossRef] [PubMed]
- Jaklitsch, W.M. European species of Hypocrea Part I. The green-spored species. Stud. Mycol. 2009, 63, 1–91. [Google Scholar] [CrossRef] [PubMed]
- Milton, J.S.; Tsokos, J.O. Statistical Methods in the Biological and Health Sciences, 3rd ed.; McGraw-Hill: New York, NY, USA, 1983; p. 600. ISBN 978-0-07-042359-6. [Google Scholar]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2020. [Google Scholar]
- R Studio Team. R Studio: Integrated Development for R; R Studio, PBC: Boston, MA, USA, 2020. [Google Scholar]
- De Mendiburu, F. Agricolae: Statistical Procedures for Agricultural Research; R Package Version 1.3-3. Available online: https://CRAN.R-project.org/package=agricolae (accessed on 11 January 2021).
- Jaklitsch, W.; Voglmayr, H. Biodiversity of Trichoderma (Hypocreaceae) in Southern Europe and Macaronesia. Stud. Mycol. 2015, 80, 1–87. [Google Scholar] [CrossRef]
- Bissett, J.; Szakacs, G.; Nolan, C.A.; Druzhinina, I.; Gradinger, C.; Kubicek, C.P. New species of Trichoderma from Asia. Can. J. Bot. 2003, 81, 570–586. [Google Scholar] [CrossRef]
- Chaverri, P.; Candoussau, F.; Samuels, G.J. Hypocrea phyllostachydis and its Trichoderma anamorph, a new bambusicolous species from France. Mycol. Prog. 2004, 3, 29–36. [Google Scholar] [CrossRef]
- Chaverri, P.; Branco-Rocha, F.; Jaklitsch, W.M.; Gazis, R.; Degenkolb, T.; Samuels, G.J. Systematics of the Trichoderma harzianum species complex and the re-identification of commercial biocontrol strains. Mycologia 2015, 107, 558–590. [Google Scholar] [CrossRef]
- Devi, P.; Prabhakaran, N.; Kamil, D.; Pandey, P.; Borah, J.L. Characterization of Indian native isolates of Trichoderma spp. and assessment of their bio-control efficiency against plant pathogens. Afr. J. Biotechnol. 2012, 11, 15150–15160. [Google Scholar] [CrossRef]
- Gerin, D.; Pollastro, S.; Raguseo, C.; Angelini, R.M.D.M.; Faretra, F. A ready-to-use single- and duplex-TaqMan-qPCR assay to detect and quantify the biocontrol agents Trichoderma asperellum and Trichoderma gamsii. Front. Microbiol. 2018, 9, 2073. [Google Scholar] [CrossRef]
- Samuels, G.J. Trichoderma: Systematics, the sexual state, and ecology. Phytopathology 2006, 96, 195–206. [Google Scholar] [CrossRef] [PubMed]
- Bae, S.-J.; Mohanta, T.K.; Chung, J.Y.; Ryu, M.; Park, G.; Shim, S.; Hong, S.-B.; Seo, H.; Bae, D.-W.; Bae, I.; et al. Trichoderma metabolites as biological control agents against Phytophthora pathogens. Biol. Control. 2016, 92, 128–138. [Google Scholar] [CrossRef]
- Cuervo-Parra, J.A.; Sánchez López, V.; Romero-Cortes, T.; Ramírez-Lepe, M. Hypocrea/Trichoderma viridescens ITV43 with potential for biocontrol of Moniliophthora roreri Cif Par, Phytophthora megasperma and Phytophthora capsici. Afr. J. Microbiol. Res. 2014, 8, 1704–1712. [Google Scholar] [CrossRef]
- Ojaghian, S.; Wang, L.; Zhang, L. Expression of lytic enzyme genes involved in antagonistic activity of Trichoderma viridescens during interaction with Sclerotinia sclerotiorum. Eur. J. Plant Pathol. 2020, 157, 223–226. [Google Scholar] [CrossRef]
- Gu, X.; Wang, R.; Sun, Q.; Wu, B.; Sun, J.-Z. Four new species of Trichoderma in the Harzianum clade from northern China. MycoKeys 2020, 73, 109–132. [Google Scholar] [CrossRef] [PubMed]
- Benítez, T.; Rincón, A.M.; Limón, M.C.; Codón, A.C. Biocontrol mechanisms of Trichoderma strains. Int. Microbiol. 2004, 7, 249–260. [Google Scholar]
- Howell, C.R. Mechanisms employed by Trichoderma species in the biological control of plant diseases: The history and evolution of current concepts. Plant Dis. 2003, 87, 4–10. [Google Scholar] [CrossRef]
- Chen, J.-L.; Sun, S.-Z.; Miao, C.-P.; Wu, K.; Chen, Y.-W.; Xu, L.-H.; Guan, H.-L.; Zhao, L.-X. Endophytic Trichoderma gamsii YIM PH30019: A promising biocontrol agent with hyperosmolar, mycoparasitism, and antagonistic activities of induced volatile organic compounds on root-rot pathogenic fungi of Panax notoginseng. J. Ginseng Res. 2016, 40, 315–324. [Google Scholar] [CrossRef] [PubMed]
- Kubicek, C.P.; Herrera-Estrella, A.; Seidl-Seiboth, V.; Martinez, D.A.; Druzhinina, I.S.; Thon, M.; Zeilinger, S.; Casas-Flores, S.; Horwitz, B.A.; Mukherjee, P.K.; et al. Comparative genome sequence analysis underscores mycoparasitism as the ancestral life style of Trichoderma. Genome Biol. 2011, 12, R40. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Harvey, P.R.; Stummer, B.E.; Warren, R.A.; Zhang, G.; Guo, K.; Li, J.; Yang, H. Antibiosis functions during interactions of Trichoderma afroharzianum and Trichoderma gamsii with plant pathogenic Rhizoctonia and Pythium. Funct. Integr. Genom. 2015, 15, 599–610. [Google Scholar] [CrossRef]
- Cucu, M.A.; Gilardi, G.; Pugliese, M.; Ferrocino, I.; Gullino, M.L. Effects of biocontrol agents and compost against the Phytophthora capsici of zucchini and their impact on the rhizosphere microbiota. Appl. Soil Ecol. 2020, 154, 103659. [Google Scholar] [CrossRef]
- Osorio-Hernández, E.; Hernández-Morales, J.; Conde-Martínez, V.; Michel-Aceves, A.C.; Lopez-Santillan, J.A.; Torres-Castillo, J.A. In vitro activities of Trichoderma species against Phytophthora parasitica and Fusarium oxysporum. Afr. J. Microbiol. Res. 2016, 10, 521–527. [Google Scholar] [CrossRef]
- Navarro-Cerrillo, R.M.; Varo-Martínez, M.Á.; Acosta-Muñoz, C.; Rodriguez, G.P.; Sánchez-Cuesta, R.; Ruiz-Gómez, F.J. Integration of WorldView-2 and airborne laser scanning data to classify defoliation levels in Quercus ilex L. dehesas affected by root rot mortality: Management implications. For. Ecol. Manag. 2019, 451, 117564. [Google Scholar] [CrossRef]
- Sánchez-Cuesta, R.; Ruiz-Gómez, F.J.; Duque-Lazo, J.; González-Moreno, P.; Navarro-Cerrillo, R.M. The environmental drivers influencing spatio-temporal dynamics of oak defoliation and mortality in dehesas of Southern Spain. For. Ecol. Manag. 2021, 485, 118946. [Google Scholar] [CrossRef]
| - | Average Plot Defoliation (%) | Average Plot Characteristics | - | Tree Defoliation (%) | Tree Characteristics | ||||
|---|---|---|---|---|---|---|---|---|---|
| Plot Code | 2015 | 2019 | H (m) | DBH (cm) | Tree Id | 2015 | 2019 | H (m) | DBH (cm) |
| CO1023 | 33.5 | 43.4 | 6.56 ± 0.19 | 34.17 ± 2.03 | 1 | 50 | 40 | 4.80 | 24.00 |
| CO1101 | 20.1 | 23.7 | 6.87 ± 0.25 | 38.67 ± 2.17 | 2 | 25 | 20 | 7.00 | 29.00 |
| HU1027 | 25.0 | 38.7 | 9.29 ± 0.45 | 38.76 ± 2.23 | 3 | 15 | 20 | 12.00 | 53.00 |
| - | - | - | - | 4 | 30 | 35 | 8.50 | 28.00 | |
| HU1028 | 37.1 | 48.7 | 5.05 ± 0.21 | 24.00 ± 1.91 | 5 | 10 | 30 | 5.00 | 32.00 |
| - | - | - | - | 6 | 20 | 30 | 5.60 | 28.00 | |
| SE1048 | 21.9 | 25.6 | 6.66 ± 0.24 | 38.03 ± 1.49 | 7 | 40 | 10 | 8.70 | 49.90 |
| - | - | - | - | 8 | 20 | 10 | 7.00 | 39.00 | |
| SE1050 | 21.7 | 27.5 | 9.43 ± 0.42 | 39.77 ± 2.34 | 9 | 20 | 20 | 11.50 | 42.70 |
| - | - | - | - | 10 | 25 | 20 | 10.50 | 39.00 | |
| - | - | - | - | 11 | 30 | 40 | 10.50 | 46.10 | |
| Trichoderma spp. | Phytophthora spp. | P. cinnamomi | |||||
|---|---|---|---|---|---|---|---|
| Plot | Tree | OTU#51 | Isolation | OTUs | Isolation | OTU#4 | Isolation |
| CO1023 | 1 | 165 | ✓ | 121 | ✓ | - | ✗ |
| CO1101 | 2 | 462 | ✓ | 3626 | ✓ | - | ✗ |
| HU1027 | 3 | 1639 | ✓ | 223 | ✓ | - | ✓ |
| 4 | 2199 | ✓ | 369 | ✓ | 10 | ✓ | |
| HU1028 | 5 | 518 | ✓ | 43 | ✓ | - | ✗ |
| 6 | 512 | ✓ | 195 | ✓ | - | ✗ | |
| SE1048 | 7 | 52 | ✓ | 6345 | ✓ | 1755 | ✓ |
| 8 | 1122 | ✓ | 9181 | ✓ | 167 | ✗ | |
| SE1050 | 9 | 886 | ✓ | 70 | ✓ | - | ✗ |
| 10 | 912 | ✓ | 20 | ✓ | - | ✗ | |
| 11 | 806 | ✓ | 21 | ✓ | - | ✗ | |
| Isolate | TEF1α | RPB2 | Clade | Gene Bank Accession Numbers | |
|---|---|---|---|---|---|
| TEF1 α | RPB2 | ||||
| T8 #1.1 | T. gamsii | T. gamsii | Viride | MZ552289 | MZ552272 |
| T8 #11.1 | T. viridarium | T. viridarium | Viride | MZ552291 | MZ552274 |
| T8 #10.1 | T. hamatum | T. hamatum | Hamatum | MZ552290 | MZ552273 |
| T9 #2.5 | T. olivascens | T. olivascens | Viridescens | MZ552292 | MZ552275 |
| T9 #5.5 | T. gamsii | T. gamsii | Viride | MZ552293 | MZ552276 |
| T10 #1.6 | T. virens | T. virens | Green | MZ552294 | MZ552277 |
| T9 #8.5 | T. olivascens | - | Viridescens | MZ552306 | - |
| T10 #8.6 | T. gamsii | T. gamsii | Viride | MZ552295 | MZ552278 |
| T10 #11.6 | T. paraviridescens | T. paraviridescens | Viridescens | MZ552296 | MZ552279 |
| T10 #13 | T. hamatum | - | Hamatum | MZ552307 | - |
| T11 #2.1 | T. linzhiense | - | Harzianum | MZ552308 | - |
| T11 #6.1 | T. hamatum | T. hamatum | Hamatum | MZ552297 | MZ552280 |
| T11 #9.1 | T. hirsutum | T. hirsutum | Harzianum | MZ552298 | MZ552281 |
| T11 #11.1 | T. gamsii | T. gamsii | Viride | MZ552300 | MZ552283 |
| T3 #16.2 | T. harzianum | - | Harzianum | MZ552309 | - |
| T3 #18.2 | T. gamsii | T. gamsii | Viride | MZ552301 | MZ552284 |
| T5 #2.3 | T. gamsii | T. gamsii | Viride | MZ552302 | MZ552285 |
| T5 #4.3 | T. gamsii | T. gamsii | Viride | MZ552303 | MZ552286 |
| T11 #10.1 | T. gamsii | T. gamsii | Viride | MZ552299 | MZ552282 |
| T5 #7.3 | T. gamsii | T. gamsii | Viride | MZ552304 | MZ552287 |
| T2 #15 | T. samuelsii | T. samuelsii | - | MZ552305 | MZ552288 |
| Isolate | Strain | MGI (%) | Mycoparasitism Scale 1 |
|---|---|---|---|
| T8 #11.1 | T. viridarium | 90.89 | 4 |
| T5 #2.3 | T. gamsii | 88.36 | 3 |
| T11 #10.1 | T. gamsii | 86.67 | 4 |
| T5 #4.3 | T. gamsii | 85.09 | 4 |
| T9 #8.5 | T. olivascens | 81.86 | 4 |
| T5 #7.3 | T. gamsii | 80.36 | 3 |
| T10 #13.6 | T. hamatum | 80.35 | 4 |
| T9 #5.5 | T. gamsii | 77.33 | 4 |
| T11 #2.1 | T. linzhiense | 76.53 | 4 |
| T11 #9.1 | T. hirsutum | 75.90 | 4 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ruiz-Gómez, F.J.; Miguel-Rojas, C. Antagonistic Potential of Native Trichoderma spp. against Phytophthora cinnamomi in the Control of Holm Oak Decline in Dehesas Ecosystems. Forests 2021, 12, 945. https://doi.org/10.3390/f12070945
Ruiz-Gómez FJ, Miguel-Rojas C. Antagonistic Potential of Native Trichoderma spp. against Phytophthora cinnamomi in the Control of Holm Oak Decline in Dehesas Ecosystems. Forests. 2021; 12(7):945. https://doi.org/10.3390/f12070945
Chicago/Turabian StyleRuiz-Gómez, Francisco J., and Cristina Miguel-Rojas. 2021. "Antagonistic Potential of Native Trichoderma spp. against Phytophthora cinnamomi in the Control of Holm Oak Decline in Dehesas Ecosystems" Forests 12, no. 7: 945. https://doi.org/10.3390/f12070945
APA StyleRuiz-Gómez, F. J., & Miguel-Rojas, C. (2021). Antagonistic Potential of Native Trichoderma spp. against Phytophthora cinnamomi in the Control of Holm Oak Decline in Dehesas Ecosystems. Forests, 12(7), 945. https://doi.org/10.3390/f12070945






