Genome-Wide Analysis of Poplar SQUAMOSA-Promoter-Binding Protein (SBP) Family under Salt Stress
Abstract
1. Introduction
2. Materials and Methods
2.1. Identification of SBP Genes
2.2. Phylogenetic Evolution and Gene Structure Analysis
2.3. Chromosome Location and Gene Duplication Analysis
2.4. Promoter and miRNA Analysis
2.5. Plant Materials
2.6. Gene Expression Analysis by RNA-Seq
2.7. Verification of Real-Time Quantitative PCR
2.8. Yeast Two-Hybrid Test
3. Results
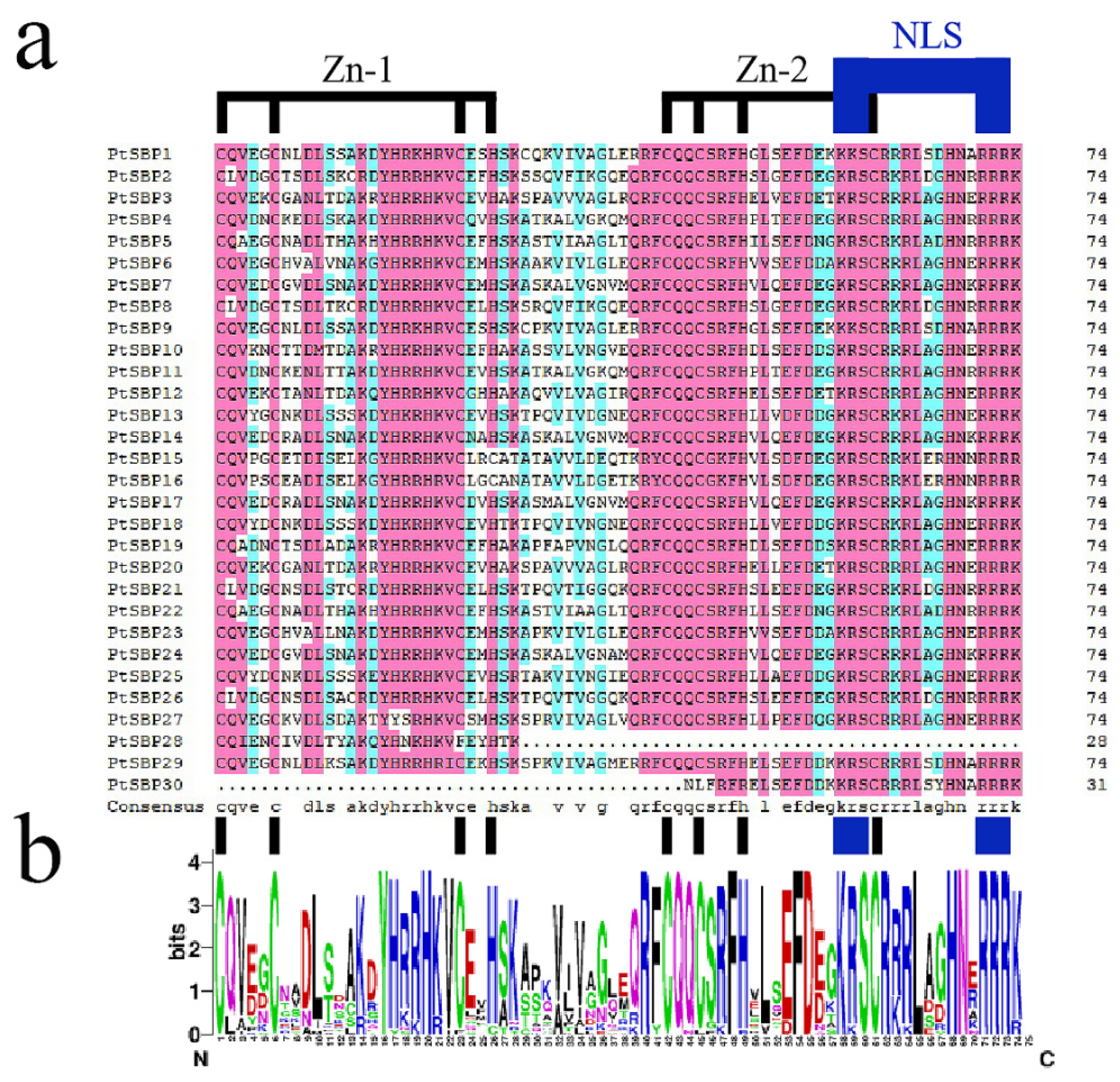
3.1. Identification and Sequence Alignment of PtSBPs
3.2. Phylogeny and Gene Structure Analysis
3.3. Chromosome Location and Gene Duplication Events
3.4. Cis-Acting Element and Target miRNA Analysis
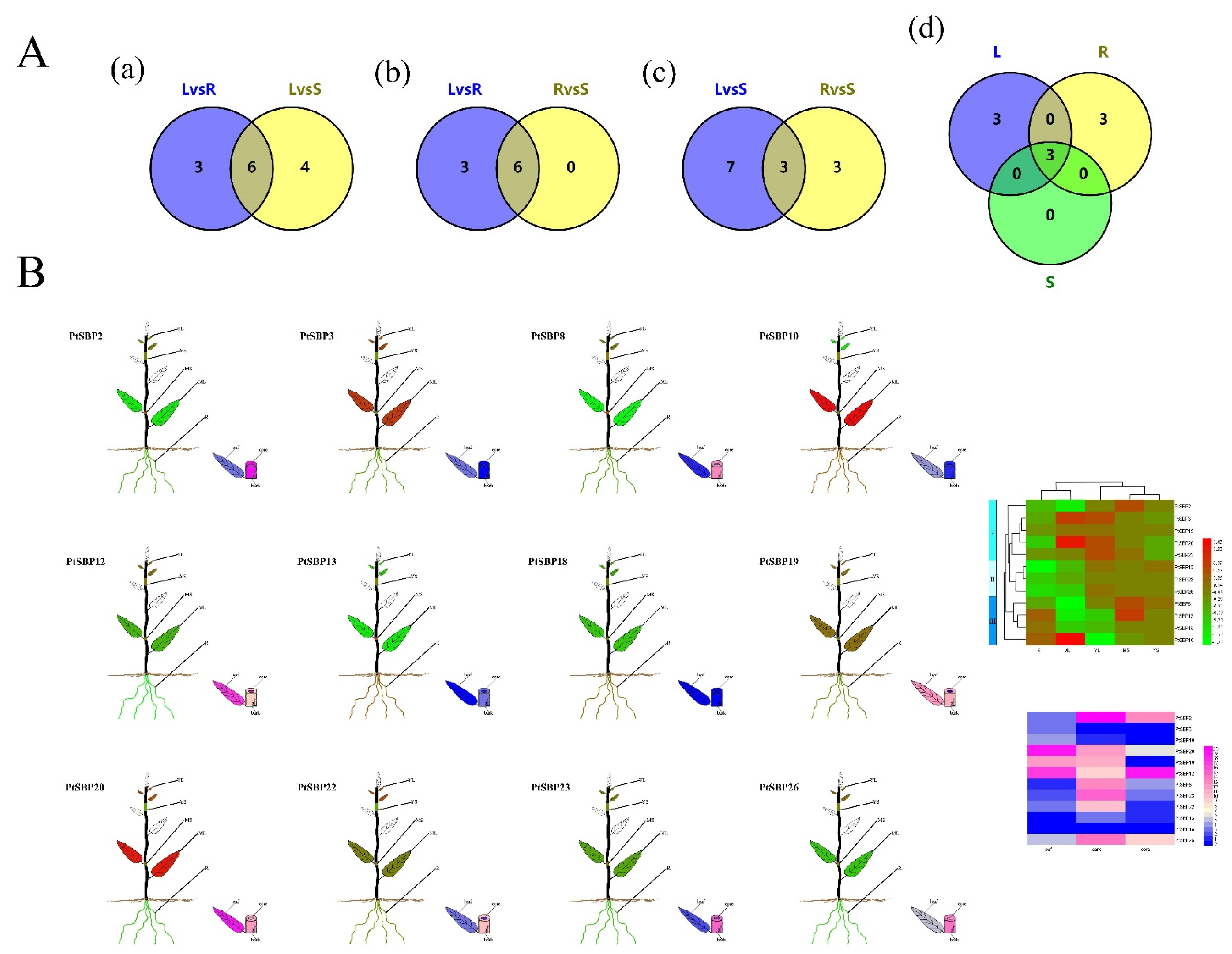
3.5. Gene Expression Analysis of PtSBPs in Different Tissues
3.6. Gene Expression Analysis of PtSBPs in Different Tissues under Salt Stress
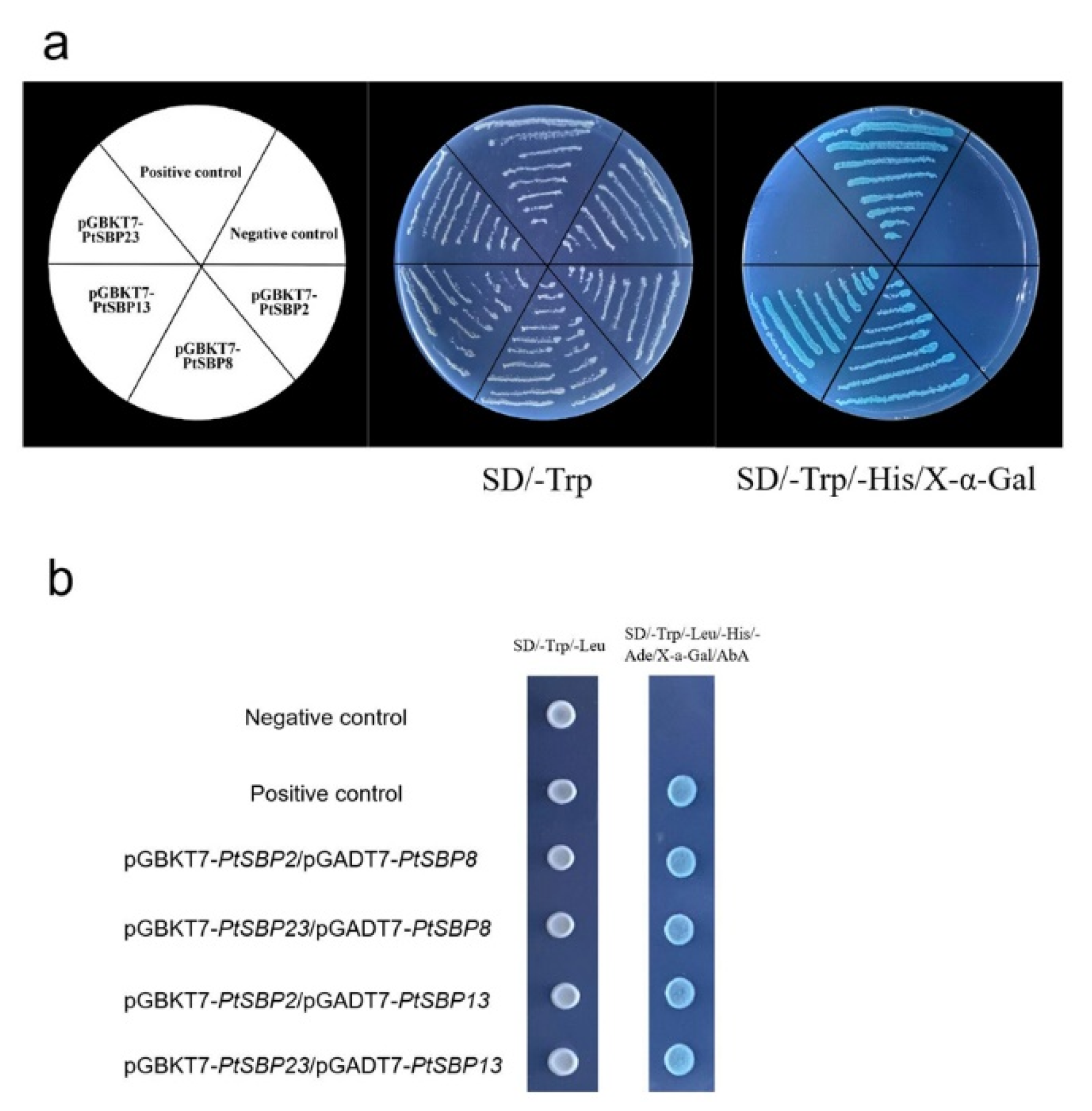
3.7. PtSBP Protein Interaction Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Yang, Z.; Wang, X.; Gu, S.; Hu, Z.; Xu, H.; Xu, C. Comparative study of SBP-box gene family in Arabidopsis and rice. Gene 2008, 407, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Gong, W.; Shen, Y.P.; Ma, L.G.; Pan, Y.; Du, Y.L.; Wang, D.H.; Yang, J.Y.; Hu, L.D.; Liu, X.F.; Dong, C.X.; et al. Genome-wide ORFeome cloning and analysis of Arabidopsis transcription factor genes. Plant. Physiol. 2004, 135, 773–782. [Google Scholar] [CrossRef] [PubMed]
- Klein, J.; Saedler, H.; Huijser, P. A new family of DNA binding proteins includes putative transcriptional regulators of the Antirrhinum majus floral meristem identity gene SQUAMOSA. Mol. Gen. Genet. MGG 1996, 250, 7–16. [Google Scholar] [CrossRef] [PubMed]
- Birkenbihl, R.P.; Jach, G.; Saedler, H.; Huijser, P. Functional dissection of the plant-specific SBP-domain: Overlap of the DNA-binding and nuclear localization domains. J. Mol. Biol. 2005, 352, 585–596. [Google Scholar] [CrossRef]
- Lakhotia, N.; Joshi, G.; Bhardwaj, A.R.; Katiyar-Agarwal, S.; Agarwal, M.; Jagannath, A.; Goel, S.; Kumar, A. Identification and characterization of miRNAome in root, stem, leaf and tuber developmental stages of potato (Solanum tuberosum L.) by high-throughput sequencing. BMC Plant. Biol. 2014, 14, 16. [Google Scholar] [CrossRef]
- Li, J.; Hou, H.; Li, X.; Xiang, J.; Yin, X.; Gao, H.; Zheng, Y.; Bassett, C.L.; Wang, X. Genome-wide identification and analysis of the SBP-box family genes in apple (Malus × domestica Borkh.). Plant. Physiol. Biochem. PPB 2013, 70, 100–114. [Google Scholar] [CrossRef]
- Cardon, G.H.; Höhmann, S.; Nettesheim, K.; Saedler, H.; Huijser, P. Functional analysis of the Arabidopsis thaliana SBP-box gene SPL3: A novel gene involved in the floral transition. Plant. J. Cell Mol. Biol. 1997, 12, 367–377. [Google Scholar] [CrossRef]
- Yamasaki, H.; Hayashi, M.; Fukazawa, M.; Kobayashi, Y.; Shikanai, T. SQUAMOSA Promoter Binding Protein-Like7 Is a Central Regulator for Copper Homeostasis in Arabidopsis. Plant. Cell 2009, 21, 347–361. [Google Scholar] [CrossRef]
- Unte, U.S.; Sorensen, A.M.; Pesaresi, P.; Gandikota, M.; Leister, D.; Saedler, H.; Huijser, P. SPL8, an SBP-box gene that affects pollen sac development in Arabidopsis. Plant. Cell 2003, 15, 1009–1019. [Google Scholar] [CrossRef]
- Lännenpää, M.; Jänönen, I.; Hölttä-Vuori, M.; Gardemeister, M.; Porali, I.; Sopanen, T. A new SBP-box gene BpSPL1 in silver birch (Betula pendula). Physiol. Plant. 2004, 120, 491–500. [Google Scholar] [CrossRef]
- Zeng, R.F.; Zhou, J.J.; Liu, S.R.; Gan, Z.M.; Zhang, J.Z.; Hu, C.G. Genome-Wide Identification and Characterization of SQUAMOSA-Promoter-Binding Protein (SBP) Genes Involved in the Flowering Development of Citrus Clementina. Biomolecules 2019, 9, 66. [Google Scholar] [CrossRef]
- Xie, K.; Wu, C.; Xiong, L. Genomic organization, differential expression, and interaction of SQUAMOSA promoter-binding-like transcription factors and microRNA156 in rice. Plant. Physiol. 2006, 142, 280–293. [Google Scholar] [CrossRef]
- Wang, S.; Wu, K.; Yuan, Q.; Liu, X.; Liu, Z.; Lin, X.; Zeng, R.; Zhu, H.; Dong, G.; Qian, Q.; et al. Control of grain size, shape and quality by OsSPL16 in rice. Nat. Genet. 2012, 44, 950–954. [Google Scholar] [CrossRef] [PubMed]
- Jiao, Y.; Wang, Y.; Xue, D.; Wang, J.; Yan, M.; Liu, G.; Dong, G.; Zeng, D.; Lu, Z.; Zhu, X.; et al. Regulation of OsSPL14 by OsmiR156 defines ideal plant architecture in rice. Nat. Genet. 2010, 42, 541–544. [Google Scholar] [CrossRef] [PubMed]
- Hou, H.; Li, J.; Gao, M.; Singer, S.D.; Wang, H.; Mao, L.; Fei, Z.; Wang, X. Genomic organization, phylogenetic comparison and differential expression of the SBP-box family genes in grape. PLoS ONE 2013, 8, e59358. [Google Scholar] [CrossRef] [PubMed]
- Arshad, M.; Feyissa, B.A.; Amyot, L.; Aung, B.; Hannoufa, A. MicroRNA156 improves drought stress tolerance in alfalfa (Medicago sativa) by silencing SPL13. Plant. Sci. Int. J. Exp. Plant. Biol. 2017, 258, 122–136. [Google Scholar] [CrossRef] [PubMed]
- Ali, H.; Liu, Y.; Azam, S.M.; Rahman, Z.U.; Priyadarshani, S.; Li, W.; Huang, X.; Hu, B.; Xiong, J.; Ali, U.; et al. Genomic Survey, Characterization, and Expression Profile Analysis of the SBP Genes in Pineapple (Ananas comosus L.). Int. J. Genom. 2017, 2017, 1032846. [Google Scholar] [CrossRef]
- Agalou, A.; Spaink, H.P.; Roussis, A. Novel interaction of selenium-binding protein with glyceraldehyde-3-phosphate dehydrogenase and fructose-bisphosphate aldolase of Arabidopsis thaliana. Funct. Plant. Biol. FPB 2006, 33, 847–856. [Google Scholar] [CrossRef] [PubMed]
- Kessens, R.; Sorensen, N.; Kabbage, M. An inhibitor of apoptosis (SfIAP) interacts with SQUAMOSA promoter-binding protein (SBP) transcription factors that exhibit pro-cell death characteristics. Plant. Direct 2018, 2, e00081. [Google Scholar] [CrossRef]
- Li, X.Y.; Lin, E.P.; Huang, H.H.; Niu, M.Y.; Tong, Z.K.; Zhang, J.H. Molecular Characterization of Squamosa Promoter Binding PROTEIN-LIKE (SPL) Gene Family in Betula luminifera. Front. Plant. Sci. 2018, 9, 608. [Google Scholar] [CrossRef]
- Yao, W.; Zhou, B.; Zhang, X.; Zhao, K.; Cheng, Z.; Jiang, T. Transcriptome analysis of transcription factor genes under multiple abiotic stresses in Populus simonii × P. nigra. Gene 2019, 707, 189–197. [Google Scholar] [CrossRef] [PubMed]
- Jansson, S.; Douglas, C.J. Populus: A model system for plant biology. Annu. Rev. Plant. Biol. 2007, 58, 435–458. [Google Scholar] [CrossRef] [PubMed]
- Lu, S.; Yang, C.; Chiang, V.L. Conservation and diversity of microRNA-associated copper-regulatory networks in Populus trichocarpa. J. Integr. Plant. Biol. 2011, 53, 879–891. [Google Scholar] [CrossRef]
- Zhou, B.; Kang, Y.; Leng, J.; Xu, Q. Genome-Wide Analysis of the miRNA-mRNAs Network Involved in Cold Tolerance in Populus simonii x P. nigra. Genes 2019, 10, 430. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Liu, W.; Wang, X.; Yang, R.; Wu, Z.; Wang, H.; Wang, L.; Hu, Z.; Guo, S.; Zhang, H.; et al. MiR156 regulates anthocyanin biosynthesis through SPL targets and other microRNAs in poplar. Hortic. Res. 2020, 7, 118. [Google Scholar] [CrossRef]
- Jin, J.; Zhang, H.; Kong, L.; Gao, G.; Luo, J. PlantTFDB 3.0: A portal for the functional and evolutionary study of plant transcription factors. Nucleic Acids Res. 2014, 42, D1182–D1187. [Google Scholar] [CrossRef]
- Goodstein, D.M.; Shu, S.; Howson, R.; Neupane, R.; Hayes, R.D.; Fazo, J.; Mitros, T.; Dirks, W.; Hellsten, U.; Putnam, N.; et al. Phytozome: A comparative platform for green plant genomics. Nucleic Acids Res. 2012, 40, D1178–D1186. [Google Scholar] [CrossRef]
- Finn, R.D.; Bateman, A.; Clements, J.; Coggill, P.; Eberhardt, R.Y.; Eddy, S.R.; Heger, A.; Hetherington, K.; Holm, L.; Mistry, J.; et al. Pfam: The protein families database. Nucleic Acids Res. 2014, 42, D222–D230. [Google Scholar] [CrossRef]
- Crooks, G.E.; Hon, G.; Chandonia, J.M.; Brenner, S.E. WebLogo: A sequence logo generator. Genome Res. 2004, 14, 1188–1190. [Google Scholar] [CrossRef]
- Artimo, P.; Jonnalagedda, M.; Arnold, K.; Baratin, D.; Csardi, G.; de Castro, E.; Duvaud, S.; Flegel, V.; Fortier, A.; Gasteiger, E.; et al. ExPASy: SIB bioinformatics resource portal. Nucleic Acids Res. 2012, 40, W597–W603. [Google Scholar] [CrossRef]
- Horton, P.; Park, K.J.; Obayashi, T.; Fujita, N.; Harada, H.; Adams-Collier, C.J.; Nakai, K. WoLF PSORT: Protein localization predictor. Nucleic Acids Res. 2007, 35, W585–W587. [Google Scholar] [CrossRef] [PubMed]
- Letunic, I.; Bork, P. Interactive tree of life (iTOL) v3: An online tool for the display and annotation of phylogenetic and other trees. Nucleic Acids Res. 2016, 44, W242–W245. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef]
- Bailey, T.L.; Boden, M.; Buske, F.A.; Frith, M.; Grant, C.E.; Clementi, L.; Ren, J.; Li, W.W.; Noble, W.S. MEME SUITE: Tools for motif discovery and searching. Nucleic Acids Res. 2009, 37, W202–W208. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Chen, H.; Zhang, Y.; Thomas, H.R.; Frank, M.H.; He, Y.; Xia, R. TBtools: An Integrative Toolkit Developed for Interactive Analyses of Big Biological Data. Mol. Plant. 2020, 13, 1194–1202. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Tang, H.; Debarry, J.D.; Tan, X.; Li, J.; Wang, X.; Lee, T.H.; Jin, H.; Marler, B.; Guo, H.; et al. MCScanX: A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 2012, 40, e49. [Google Scholar] [CrossRef] [PubMed]
- Tuskan, G.A.; Difazio, S.; Jansson, S.; Bohlmann, J.; Grigoriev, I.; Hellsten, U.; Putnam, N.; Ralph, S.; Rombauts, S.; Salamov, A.; et al. The genome of black cottonwood, Populus trichocarpa (Torr. & Gray). Science 2006, 313, 1596–1604. [Google Scholar] [CrossRef]
- Zhang, Y. miRU: An automated plant miRNA target prediction server. Nucleic Acids Res. 2005, 33, W701–W704. [Google Scholar] [CrossRef]
- Dai, X.; Zhao, P.X. psRNATarget: A plant small RNA target analysis server. Nucleic Acids Res. 2011, 39, W155–W159. [Google Scholar] [CrossRef]
- Yao, W.; Wang, S.; Zhou, B.; Jiang, T. Transgenic poplar overexpressing the endogenous transcription factor ERF76 gene improves salinity tolerance. Tree Physiol. 2016, 36, 896–908. [Google Scholar] [CrossRef]
- Anders, S.; Huber, W. Differential expression analysis for sequence count data. Genome Biol. 2010, 11, R106. [Google Scholar] [CrossRef]
- Regier, N.; Frey, B. Experimental comparison of relative RT-qPCR quantification approaches for gene expression studies in poplar. BMC Mol. Biol. 2010, 11, 57. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods (San Diegocalif.) 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.-X.; Jin, J.-H.; He, Y.-M.; Lu, B.-Y.; Li, D.-W.; Chai, W.-G.; Khan, A.; Gong, Z.-H. Genome-Wide Identification and Analysis of the SBP-Box Family Genes under Phytophthora capsici Stress in Pepper (Capsicum annuum L.). Front. Plant. Sci. 2016, 7. [Google Scholar] [CrossRef]
- Zhang, L.; Wu, B.; Zhao, D.; Li, C.; Shao, F.; Lu, S. Genome-wide analysis and molecular dissection of the SPL gene family in Salvia miltiorrhiza. J. Integr. Plant. Biol. 2014, 56, 38–50. [Google Scholar] [CrossRef]
- Abdullah, M.; Cao, Y.P.; Cheng, X.; Shakoor, A.; Su, X.Q.; Gao, J.S.; Cai, Y.P. Genome-Wide Analysis Characterization and Evolution of SBP Genes in Fragaria vesca, Pyrus bretschneideri, Prunus persica and Prunus mume. Front. Genet. 2018, 9, 12. [Google Scholar] [CrossRef] [PubMed]
- Salinas, M.; Xing, S.; Höhmann, S.; Berndtgen, R.; Huijser, P. Genomic organization, phylogenetic comparison and differential expression of the SBP-box family of transcription factors in tomato. Planta 2012, 235, 1171–1184. [Google Scholar] [CrossRef]
- Gang, H.; Li, R.; Zhao, Y.; Liu, G.; Chen, S.; Jiang, J. Loss of GLK1 transcription factor function reveals new insights in chlorophyll biosynthesis and chloroplast development. J. Exp. Bot. 2019, 70, 3125–3138. [Google Scholar] [CrossRef]
- Wang, W.; Zheng, H.; Yang, S.; Yu, H.; Li, J.; Jiang, H.; Su, J.; Yang, L.; Zhang, J.; McDermott, J.; et al. Origin and evolution of new exons in rodents. Genome Res. 2005, 15, 1258–1264. [Google Scholar] [CrossRef]
- Shikata, M.; Koyama, T.; Mitsuda, N.; Ohme-Takagi, M. Arabidopsis SBP-box genes SPL10, SPL11 and SPL2 control morphological change in association with shoot maturation in the reproductive phase. Plant. Cell Physiol. 2009, 50, 2133–2145. [Google Scholar] [CrossRef]
- Dung Tien, L.; Nishiyama, R.; Watanabe, Y.; Vankova, R.; Tanaka, M.; Seki, M.; Le Huy, H.; Yamaguchi-Shinozaki, K.; Shinozaki, K.; Lam-Son Phan, T. Identification and Expression Analysis of Cytokinin Metabolic Genes in Soybean under Normal and Drought Conditions in Relation to Cytokinin Levels. PLoS ONE 2012, 7, e42411. [Google Scholar] [CrossRef]
- Eldem, V.; Okay, S.; Unver, T. Plant microRNAs: New players in functional genomics. Turk. J. Agric. For. 2013, 37, 1–21. [Google Scholar] [CrossRef]
- Wu, G.; Park, M.Y.; Conway, S.R.; Wang, J.W.; Weigel, D.; Poethig, R.S. The sequential action of miR156 and miR172 regulates developmental timing in Arabidopsis. Cell 2009, 138, 750–759. [Google Scholar] [CrossRef]
- Yin, D.D.; Li, S.S.; Shu, Q.Y.; Gu, Z.Y.; Wu, Q.; Feng, C.Y.; Xu, W.Z.; Wang, L.S. Identification of microRNAs and long non-coding RNAs involved in fatty acid biosynthesis in tree peony seeds. Gene 2018, 666, 72–82. [Google Scholar] [CrossRef]
- Geng, Y.; Jian, C.; Xu, W.; Liu, H.; Hao, C.; Hou, J.; Liu, H.; Zhang, X.; Li, T. miR164-targeted TaPSK5 encodes a phytosulfokine precursor that regulates root growth and yield traits in common wheat (Triticum aestivum L.). Plant. Mol. Biol. 2020, 104, 615–628. [Google Scholar] [CrossRef] [PubMed]
- Zheng, L.; Nagpal, P.; Villarino, G.; Trinidad, B.; Bird, L.; Huang, Y.; Reed, J.W. miR167 limits anther growth to potentiate anther dehiscence. Development 2019, 146. [Google Scholar] [CrossRef] [PubMed]
- Stone, J.M.; Liang, X.; Nekl, E.R.; Stiers, J.J. Arabidopsis AtSPL14, a plant-specific SBP-domain transcription factor, participates in plant development and sensitivity to fumonisin B1. Plant. J. Cell Mol. Biol. 2005, 41, 744–754. [Google Scholar] [CrossRef]
- Wan, H.; Yuan, W.; Yu, K.; Liu, Y.; Li, Z.; Ye, Q.; Wang, R.; Ruan, M.; Zhou, G.; Yao, Z.; et al. Geno me-wide Identification, Structure Characterization and Expression Analysis of SBP Gene Family in Tomato. Mol. Plant. Breed. 2013, 11, 299–306. [Google Scholar]
- Aukerman, M.J.; Sakai, H. Regulation of flowering time and floral organ identity by a MicroRNA and its APETALA2-like target genes. Plant. Cell 2003, 15, 2730–2741. [Google Scholar] [CrossRef]
- Rhoades, M.W.; Reinhart, B.J.; Lim, L.P.; Burge, C.B.; Bartel, B.; Bartel, D.P. Prediction of plant microRNA targets. Cell 2002, 110, 513–520. [Google Scholar] [CrossRef]
- Cui, L.G.; Shan, J.X.; Shi, M.; Gao, J.P.; Lin, H.X. The miR156-SPL9-DFR pathway coordinates the relationship between development and abiotic stress tolerance in plants. Plant. J. Cell Mol. Biol. 2014, 80, 1108–1117. [Google Scholar] [CrossRef]
- Lee, H.; Yoo, S.J.; Lee, J.H.; Kim, W.; Yoo, S.K.; Fitzgerald, H.; Carrington, J.C.; Ahn, J.H. Genetic framework for flowering-time regulation by ambient temperature-responsive miRNAs in Arabidopsis. Nucleic Acids Res. 2010, 38, 3081–3093. [Google Scholar] [CrossRef]
- Kantar, M.; Lucas, S.J.; Budak, H. miRNA expression patterns of Triticum dicoccoides in response to shock drought stress. Planta 2011, 233, 471–484. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Bao, J.; Zhou, B.; Li, M.; Li, X.; Jin, J. The osa-miR164 target OsCUC1 functions redundantly with OsCUC3 in controlling rice meristem/organ boundary specification. New Phytol. 2020, 229, 1566–1581. [Google Scholar] [CrossRef] [PubMed]
- Hu, G.; Lei, Y.; Liu, J.; Hao, M.; Zhang, Z.; Tang, Y.; Chen, A.; Wu, J. The ghr-miR164 and GhNAC100 modulate cotton plant resistance against Verticillium dahlia. Plant. Sci. Int. J. Exp. Plant. Biol. 2020, 293, 110438. [Google Scholar] [CrossRef] [PubMed]
- Chai, G.; Wang, Z.; Tang, X.; Yu, L.; Qi, G.; Wang, D.; Yan, X.; Kong, Y.; Zhou, G. R2R3-MYB gene pairs in Populus: Evolution and contribution to secondary wall formation and flowering time. J. Exp. Bot. 2014, 65, 4255–4269. [Google Scholar] [CrossRef]
- Jung, J.H.; Lee, H.J.; Ryu, J.Y.; Park, C.M. SPL3/4/5 Integrate Developmental Aging and Photoperiodic Signals into the FT-FD Module in Arabidopsis Flowering. Mol. Plant. 2016, 9, 1647–1659. [Google Scholar] [CrossRef] [PubMed]
- Padmanabhan, M.S.; Ma, S.; Burch-Smith, T.M.; Czymmek, K.; Huijser, P.; Dinesh-Kumar, S.P. Novel positive regulatory role for the SPL6 transcription factor in the N TIR-NB-LRR receptor-mediated plant innate immunity. PLoS Pathog. 2013, 9, e1003235. [Google Scholar] [CrossRef]
- Chao, L.M.; Liu, Y.Q.; Chen, D.Y.; Xue, X.Y.; Mao, Y.B.; Chen, X.Y. Arabidopsis Transcription Factors SPL1 and SPL12 Confer Plant Thermotolerance at Reproductive Stage. Mol. Plant. 2017, 10, 735–748. [Google Scholar] [CrossRef]








| Gene Name | Accession Number | AA | Molecular Weight | Theoretical PI | Instability Index | Aliphatic Index | GRAVY | AtSBP Ortholog | Subcellular Locatioon |
|---|---|---|---|---|---|---|---|---|---|
| PtSBP1 | Potri.001G055900.1 | 446 | 49008.35 | 8.67 | 52.71 | 61.95 | −0.724 | AT5G43270.3 | Nucleus |
| PtSBP2 | Potri.001G058600.1 | 376 | 41280.85 | 9.3 | 54.07 | 56.81 | −0.707 | AT5G50670.1 | Cytoplasm; Nucleus |
| PtSBP3 | Potri.001G398200.1 | 244 | 27427.02 | 9.07 | 63.32 | 61.56 | −0.784 | AT3G15270.1 | Nucleus |
| PtSBP4 | Potri.002G002400.1 | 1073 | 119091.39 | 8.53 | 59.07 | 76.59 | −0.528 | AT1G20980.1 | Nucleus |
| PtSBP5 | Potri.002G142200.1 | 325 | 36132.89 | 9.11 | 57.79 | 58.95 | −0.705 | AT1G02065.1 | Cytoplasm; Nucleus |
| PtSBP6 | Potri.002G142400.1 | 398 | 43784.56 | 8.53 | 53.54 | 65.73 | −0.606 | AT1G27360.4 | Nucleus |
| PtSBP7 | Potri.002G188700.4 | 1002 | 110900.96 | 5.87 | 53.09 | 79.63 | −0.365 | AT2G47070.1 | Nucleus |
| PtSBP8 | Potri.003G169400.1 | 313 | 34232.95 | 9.23 | 53.55 | 58.27 | −0.699 | AT5G50570.2 | Nucleus |
| PtSBP9 | Potri.003G172600.1 | 411 | 45630.63 | 9.24 | 47.86 | 60.07 | −0.818 | AT5G43270.3 | Cytoplasm; Nucleus |
| PtSBP10 | Potri.004G046700.1 | 148 | 16466.96 | 7.6 | 80.6 | 36.89 | −1.316 | AT2G33810.1 | Nucleus |
| PtSBP11 | Potri.005G258700.1 | 1072 | 118587.67 | 8.14 | 58.2 | 77.31 | −0.475 | AT1G20980.1 | Nucleus |
| PtSBP12 | Potri.007G138800.1 | 202 | 22709.64 | 9.49 | 47.34 | 51.14 | −0.916 | AT1G53160.1 | Nucleus |
| PtSBP13 | Potri.008G097900.1 | 510 | 55865.53 | 7.25 | 46.81 | 68.63 | −0.492 | AT1G69170.1 | Nucleus |
| PtSBP14 | Potri.008G098600.1 | 1035 | 115390.06 | 7.3 | 43.54 | 84.11 | −0.373 | AT2G47070.1 | Nucleus |
| PtSBP15 | Potri.008G197000.1 | 793 | 88909.58 | 6.43 | 46.93 | 78.89 | −0.358 | AT5G18830.3 | Cytoplasm; Nucleus |
| PtSBP16 | Potri.010G026200.1 | 782 | 87604.3 | 6.49 | 49.47 | 81.24 | −0.298 | AT5G18830.3 | Cytoplasm; Nucleus |
| PtSBP17 | Potri.010G154000.1 | 1030 | 114785.39 | 8.13 | 45.62 | 82.24 | −0.409 | AT2G47070.1 | Nucleus |
| PtSBP18 | Potri.010G154300.1 | 600 | 66181.99 | 8.18 | 47.49 | 75.27 | −0.335 | AT1G69170.1 | Nucleus |
| PtSBP19 | Potri.011G055900.1 | 144 | 16188.76 | 6.53 | 85.47 | 44.79 | −1.206 | AT2G33810.1 | Nucleus |
| PtSBP20 | Potri.011G116800.1 | 242 | 27239.63 | 8.93 | 70.78 | 56.03 | −0.888 | AT3G15270.1 | Nucleus |
| PtSBP21 | Potri.012G100700.1 | 346 | 38302.62 | 8.68 | 55.2 | 51.3 | −0.707 | AT5G50570.2 | Nucleus |
| PtSBP22 | Potri.014G057700.1 | 328 | 36449.12 | 8.85 | 59.69 | 55.7 | −0.745 | AT1G02065.1 | Nucleus |
| PtSBP23 | Potri.014G057800.1 | 408 | 45197.94 | 6.81 | 60.25 | 62.4 | −0.694 | AT1G27370.4 | Nucleus |
| PtSBP24 | Potri.014G114300.1 | 1004 | 111261.28 | 6.13 | 51.82 | 80.3 | −0.393 | AT3G60030.1 | Nucleus |
| PtSBP25 | Potri.015G060400.1 | 561 | 61613.62 | 8.39 | 53.58 | 63.62 | −0.562 | AT1G69170.1 | Cytoplasm; Nucleus |
| PtSBP26 | Potri.015G098900.1 | 381 | 41809.37 | 8.68 | 56.67 | 60.37 | −0.644 | AT5G50570.2 | Nucleus |
| PtSBP27 | Potri.016G048500.1 | 382 | 41034.42 | 9.03 | 59.25 | 57.75 | −0.614 | AT2G42200.1 | Nucleus |
| PtSBP28 | Potri.017G012900.1 | 110 | 12592.56 | 9.49 | 35.42 | 56.55 | −0.776 | AT1G53160.2 | Cytoplasm; Nucleus |
| PtSBP29 | Potri.018G149900.1 | 472 | 51720.59 | 8.58 | 61.98 | 62.46 | −0.569 | AT5G43270.3 | Cytoplasm; Nucleus |
| PtSBP30 | Potri.019G073600.1 | 69 | 8351.57 | 10.62 | 91.51 | 43.91 | −0.916 | AT3G57920.1 | Nucleus |
| No. | Paralogous Genes | Ks | Ka | Ka/Ks |
|---|---|---|---|---|
| 1 | PtSBP2 (Chr01)/PtSBP8 (Chr03) | 0.178443215 | 0.061591622 | 0.345160908 |
| 2 | PtSBP3 (Chr01)/PtSBP20 (Chr11) | 0.342751855 | 0.078286805 | 0.228406656 |
| 3 | PtSBP4 (Chr02)/PtSBP11 (Chr05) | 0.271895969 | 0.068274383 | 0.2511048 |
| 4 | PtSBP6 (Chr02)/PtSBP23 (Chr14) | 0.284902171 | 0.071482635 | 0.250902388 |
| 5 | PtSBP10 (Chr04)/PtSBP19 (Chr11) | 0.25806393 | 0.114291101 | 0.442879023 |
| 6 | PtSBP12 (Chr07)/PtSBP28 (Chr17) | 0.470500471 | 0.265439967 | 0.564165146 |
| 7 | PtSBP13 (Chr08)/PtSBP18 (Chr10) | 0.236628628 | 0.100342683 | 0.424051323 |
| 8 | PtSBP14 (Chr08)/PtSBP17 (Chr10) | 0.224605439 | 0.064208977 | 0.285874541 |
| 9 | PtSBP15 (Chr08)/PtSBP16 (Chr10) | 0.232984091 | 0.078535107 | 0.33708356 |
| 10 | PtSBP21 (Chr12)/PtSBP26 (Chr15) | 0.300891016 | 0.112241239 | 0.373029545 |
| Cis-element | Sequences | Function | Genes |
|---|---|---|---|
| 3-AF1 binding site | TAAGAGAGGAA | cis-acting regulatory element involved in light responsiveness | PtSBP30 |
| AACA_motif | TAACAAACTCCA | involved in endosperm-specific negative expression | PtSBP30 |
| ABRE | ACGTG,CACGTG/CGCACGTGTC,GCAACGTGTC,GCCGCGTGGC/CGTACGTGCA,TACGTGTC | cis-acting element involved in the abscisic acid responsiveness | PtSBP13, PtSBP14 |
| ACE | CTAACGTATT | cis-acting regulatory element involved in light responsiveness | PtSBP13 |
| AE-box | AGAAACAA,AGAAACTT | cis-acting regulatory element involved in light responsiveness | PtSBP17 |
| ARE | AAACCA | cis-acting regulatory element essential for the anaerobic induction | PtSBP14, PtSBP15 |
| AT1-motif | AATTATTTTTTATT | cis-acting regulatory element involved in light responsiveness | PtSBP16, PtSBP17 |
| ATCT-motif | AATCTAATCC | cis-acting regulatory element involved in light responsiveness | PtSBP25, PtSBP26 |
| AuxRR-core | GGTCCAT | cis-acting regulatory element involved in auxin responsiveness | PtSBP15 |
| Box 4 | ATTAAT | cis-acting regulatory element involved in light responsiveness | PtSBP24-27 |
| CAT-box | GCCACT | cis-acting regulatory element related to meristem expression | PtSBP11 |
| CGTCA-motif | CGTCA | cis-acting regulatory element involved in the MeJA-responsiveness | PtSBP11 |
| chs-CMA1a | TTACTTAA | cis-acting regulatory element involved in light responsiveness | PtSBP21 |
| chs-CMA2a | TCACTTGA | cis-acting regulatory element involved in light responsiveness | PtSBP19-20, PtSBP24 |
| GA-motif | ATAGATAA | cis-acting regulatory element involved in light responsiveness | PtSBP19, PtSBP20 |
| Gap-box | CAAATGAA(A/G)A | cis-acting regulatory element involved in light responsiveness | PtSBP16 |
| GARE-motif | TCTGTTG | cis-acting element involved in gibberellin-responsiveness | PtSBP4, PtSBP25, PtSBP30 |
| GATA-motif | AAGGATAAGG/GATAGGA/GATAGGG | cis-acting regulatory element involved in light responsiveness | PtSBP20 |
| G-box | CACGAC,CACGTC,CACGTT,GCCACGTGGA,TACGTG/TAACACGTAG,ACACGTG(G/T)CACC,TCCACATGGCA | cis-acting regulatory element involved in light responsiveness | PtSBP12, PtSBP15 |
| GC-motif | CCCCCG | cis-acting regulatory element essential for the anaerobic induction | PtSBP12, PtSBP13 |
| GCN4_motif | TGAGTCA | cis-regulatory element involved in endosperm expression | PtSBP11 |
| GT1-motif | GGTTAA,GGTTAAT | cis-acting regulatory element involved in light responsiveness | PtSBP29, PtSBP30 |
| HD-Zip 1 | CAAT(A/T)ATTG | element involved in differentiation of the palisade mesophyll cells | PtSBP13, PtSBP16 |
| I-box | AAGATAAGGCT/CCATATCCAAT/GGATAAGGTG,TGATAATGT | cis-acting regulatory element involved in light responsiveness | PtSBP19, PtSBP20, PtSBP24 |
| LAMP-element | CTTTATCA | cis-acting regulatory element involved in light responsiveness | PtSBP20 |
| L-box | ATTCTCACCTACCA | cis-acting regulatory element involved in light responsiveness | PtSBP24 |
| LS7 | CAGATTTATTTTTA | cis-acting regulatory element involved in light responsiveness | PtSBP21 |
| LTR | CCGAAA | cis-acting element involved in low-temperature responsiveness | PtSBP13 |
| MBS | CAACTG | MYB binding site involved in drought-inducibility | PtSBP27, PtSBP28 |
| MBSI | AAAAAAC(G/C)GTTA,TTTTTACGGTTA | MYB binding site involved in flavonoid biosynthetic genes regulation | PtSBP27 |
| MRE | AACCTAA | MYB binding site involved in light responsiveness | PtSBP27 |
| MSA-like | (T/C)C(T/C)AACGG(T/C)(T/C)A | cis-acting element involved in cell cycle regulation | PtSBP18 |
| O2-site | GATGATGTGG,GTTGACGTGA | cis-acting regulatory element involved in zein metabolism regulation | PtSBP11 |
| P-box | CCTTTTG | cis-acting element involved in gibberellin-responsiveness | PtSBP1, PtSBP4, PtSBP5, PtSBP7, PtSBP9, PtSBP10, PtSBP11, PtSBP24, PtSBP30 |
| RY-element | CATGCATG | cis-acting regulatory element involved in seed-specific regulation | PtSBP12 |
| Sp1 | GGGCGG | cis-acting regulatory element involved in light responsiveness | PtSBP29 |
| TATC-box | TATCCCA | cis-acting element involved in gibberellin-responsiveness | PtSBP13 |
| TCA-element | CCATCTTTTT | cis-acting element involved in salicylic acid responsiveness | PtSBP13 |
| TCCC-motif | TCTCCCT | cis-acting regulatory element involved in light responsiveness | PtSBP20 |
| TC-rich repeats | ATTCTCTAAC/GTTTTCTTAC | cis-acting element involved in defense and stress responsiveness | PtSBP13, PtSBP18 |
| TCT-motif | TCTTAC | cis-acting regulatory element involved in light responsiveness | PtSBP16, PtSBP19-21 |
| TGACG-motif | TGACG | cis-acting regulatory element involved in the MeJA-responsiveness | PtSBP11, PtSBP12 |
| TGA-element | AACGAC | cis-acting regulatory element involved in auxin responsiveness | PtSBP18 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Guo, Q.; Li, L.; Zhao, K.; Yao, W.; Cheng, Z.; Zhou, B.; Jiang, T. Genome-Wide Analysis of Poplar SQUAMOSA-Promoter-Binding Protein (SBP) Family under Salt Stress. Forests 2021, 12, 413. https://doi.org/10.3390/f12040413
Guo Q, Li L, Zhao K, Yao W, Cheng Z, Zhou B, Jiang T. Genome-Wide Analysis of Poplar SQUAMOSA-Promoter-Binding Protein (SBP) Family under Salt Stress. Forests. 2021; 12(4):413. https://doi.org/10.3390/f12040413
Chicago/Turabian StyleGuo, Qing, Li Li, Kai Zhao, Wenjing Yao, Zihan Cheng, Boru Zhou, and Tingbo Jiang. 2021. "Genome-Wide Analysis of Poplar SQUAMOSA-Promoter-Binding Protein (SBP) Family under Salt Stress" Forests 12, no. 4: 413. https://doi.org/10.3390/f12040413
APA StyleGuo, Q., Li, L., Zhao, K., Yao, W., Cheng, Z., Zhou, B., & Jiang, T. (2021). Genome-Wide Analysis of Poplar SQUAMOSA-Promoter-Binding Protein (SBP) Family under Salt Stress. Forests, 12(4), 413. https://doi.org/10.3390/f12040413







