Bacillus velezensis: A Treasure House of Bioactive Compounds of Medicinal, Biocontrol and Environmental Importance
Abstract
1. Introduction
2. Materials and Methods
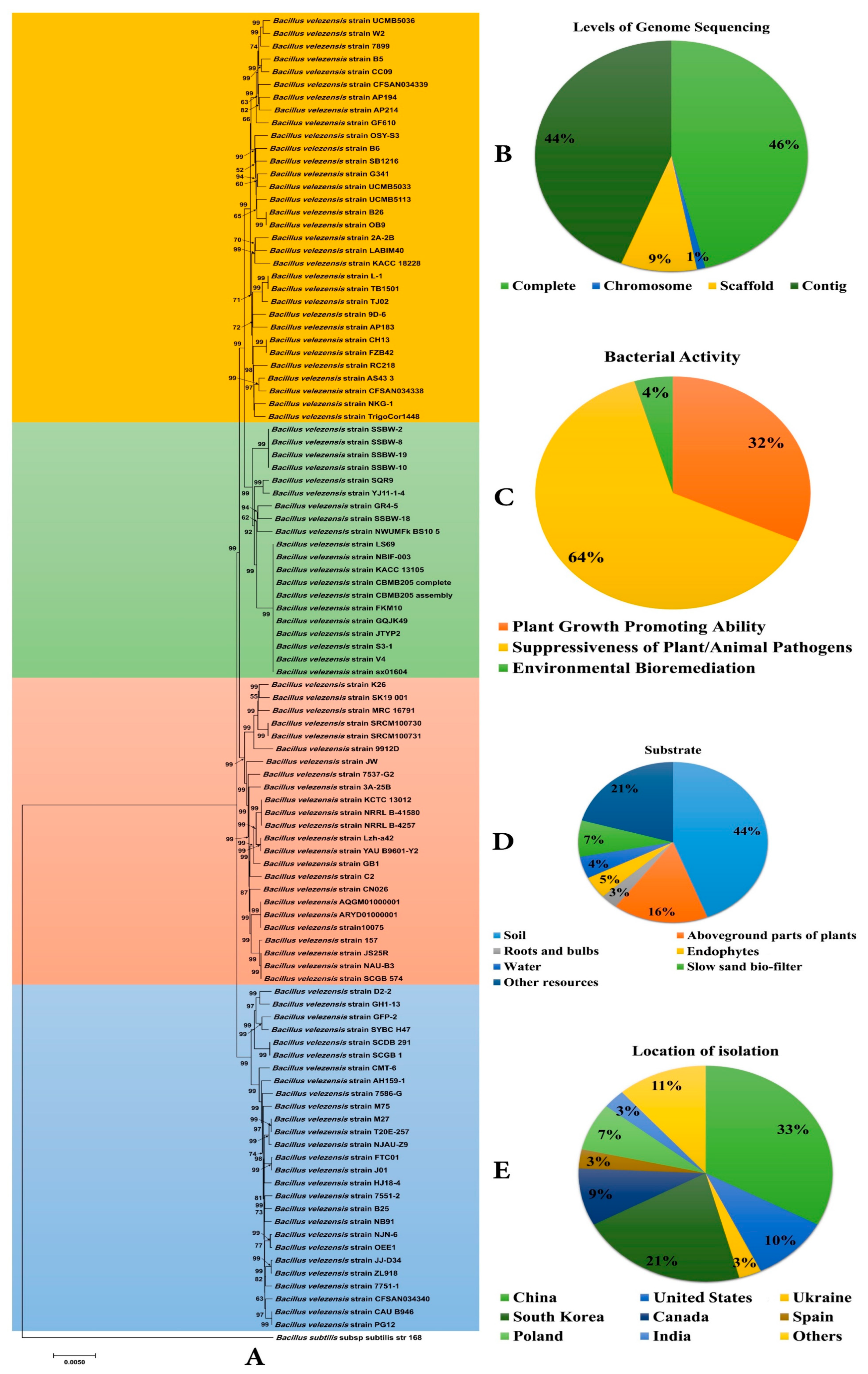
2.1. Selection and Phylogenomic Analysis of B. velezensis Genomes
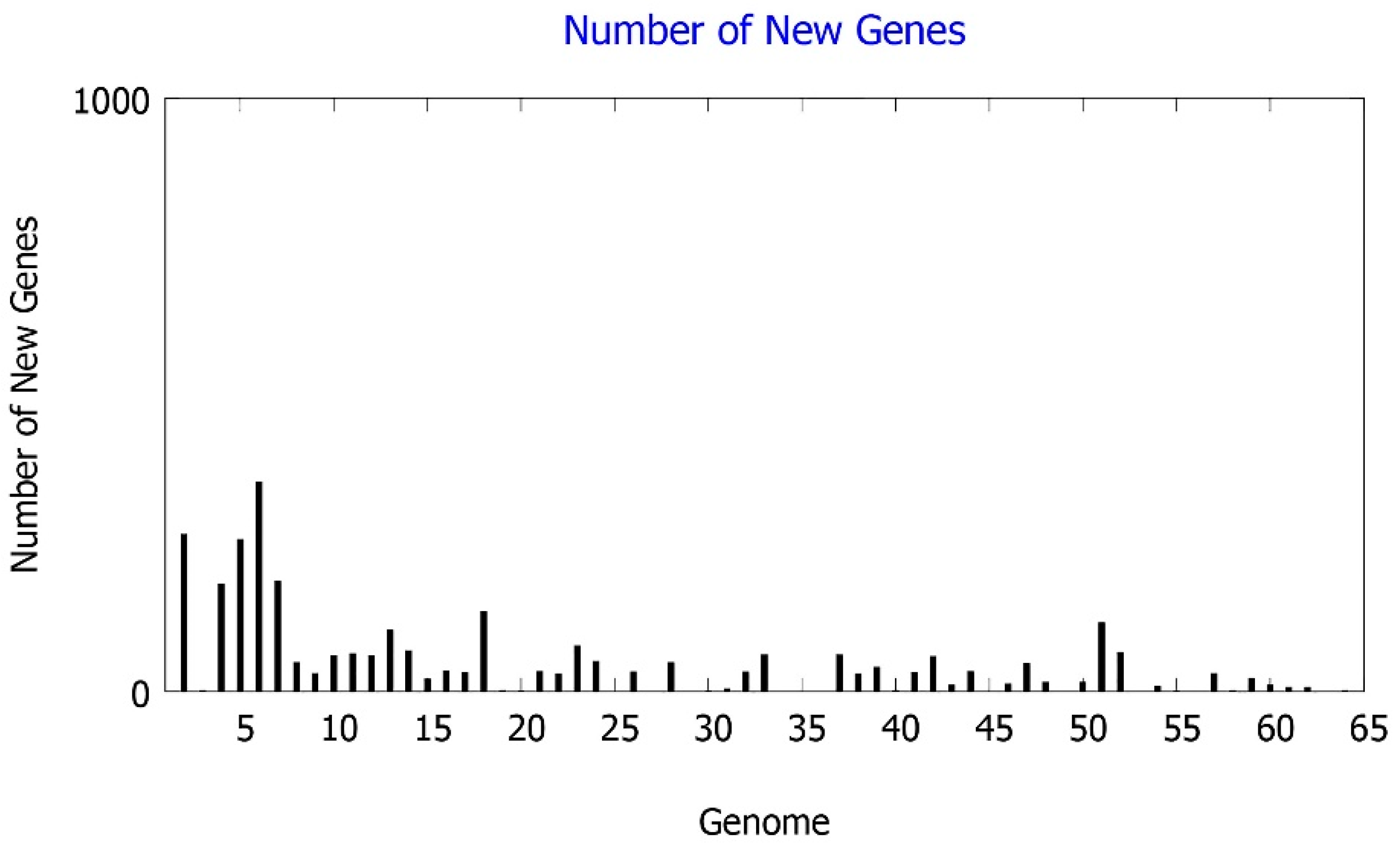
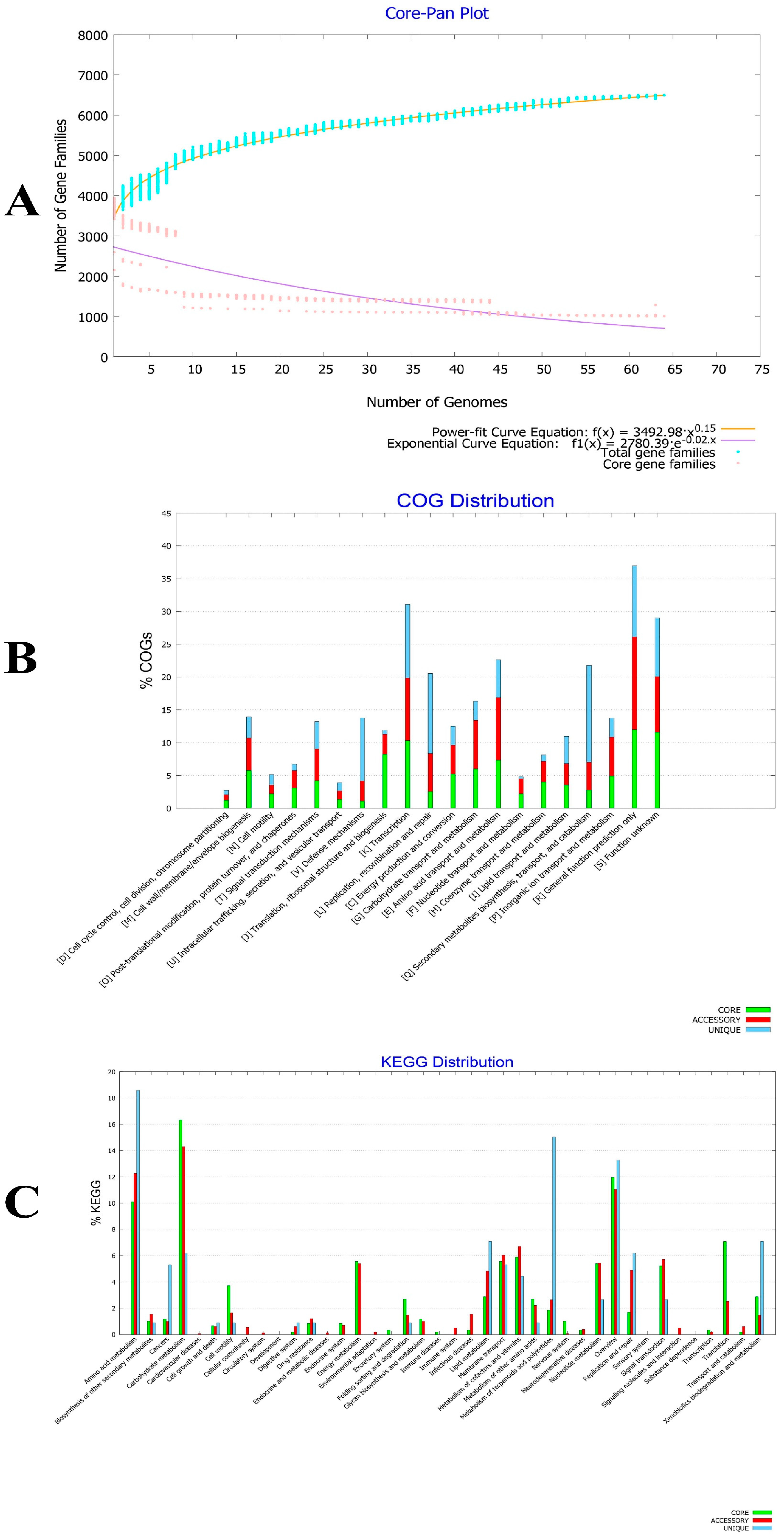
2.2. Comparative Genomics Study of B. velezensis Strains Assembled Genomes
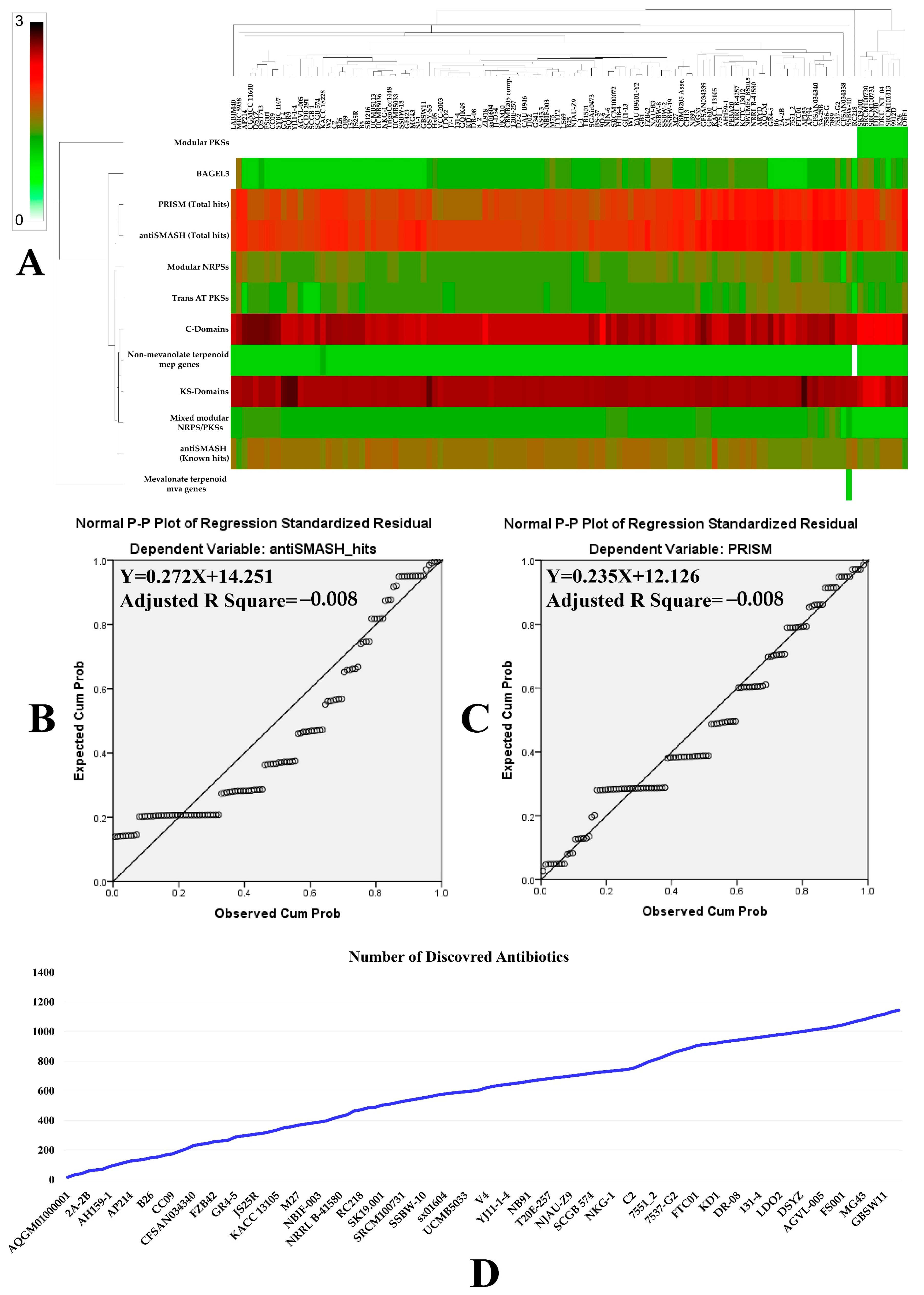
2.3. Genome Mining of Secondary Metabolites Genes Clusters
2.4. Mining of Genes Devoted to Plant Beneficial Functions on Homology Basis
Nutrient Acquisition
2.5. Genome Mining for Functional Genes Responsible for Growth Promotion and Root Colonization
2.6. Hormones of Plant Growth Promotion
2.7. Antioxidant Enzymes
2.8. Disease Resistance Induction in Plants
2.9. Antibiotics and Other Related Compounds
2.10. Drugs Resistance
2.11. Heavy Metals Tolerance
2.12. Aromatic Compounds Degradation
2.13. Determination of the Core and Accessory Genomes of the B. velezensis Isolates
3. Results
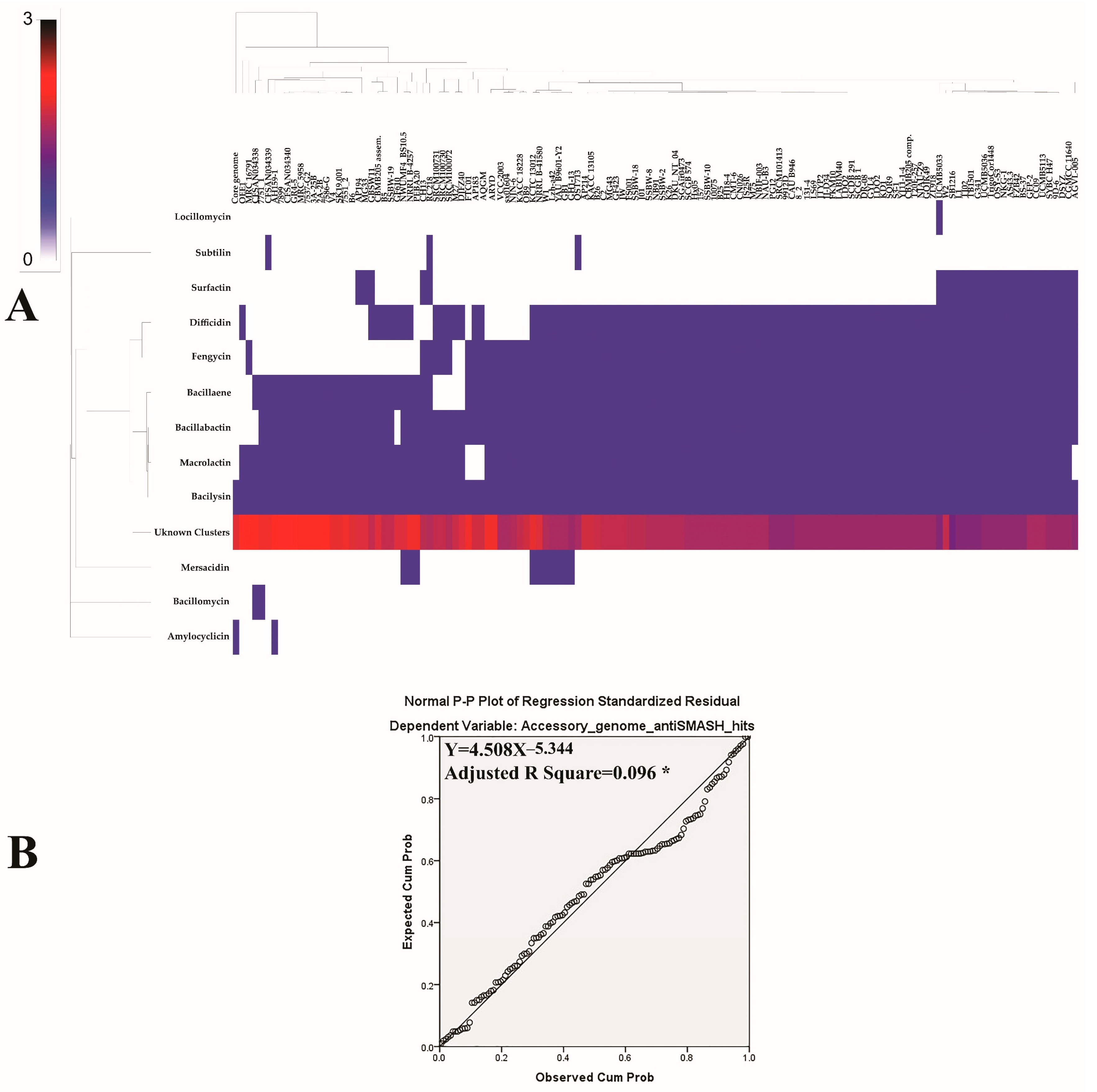
3.1. Secondary Metabolites Biosynthesis Capacities of B. velezensis Pan, Core and Accessory Genomes
3.2. Prediction of Secondary Metabolites Richness and Location within B. velezensis Genomes
3.3. Prediction of Secondary Metabolites Richness and Location within B. velezensis Genomes
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Borriss, R. Bacillus, a Plant-Beneficial Bacterium. In Principles of Plant-Microbe Interactions; Springer: Cham, Switzerland, 2015; pp. 379–391. [Google Scholar]
- Wu, B.; Wang, X.; Yang, L.; Yang, H.; Zeng, H.; Qiu, Y.; Wang, C.; Yu, J.; Li, J.; Xu, D. Effects of Bacillus amyloliquefaciens ZM9 on Bacterial Wilt and Rhizosphere Microbial Communities of Tobacco. Appl. Soil Ecol. 2016, 103, 1–12. [Google Scholar] [CrossRef]
- Al-Ali, A.; Deravel, J.; Krier, F.; Béchet, M.; Ongena, M.; Jacques, P. Biofilm Formation Is Determinant in Tomato Rhizosphere Colonization by Bacillus velezensis FZB42. Environ. Sci. Pollut. Res. 2018, 25, 29910–29920. [Google Scholar] [CrossRef] [PubMed]
- Belbahri, L.; Chenari Bouket, A.; Rekik, I.; Alenezi, F.N.; Vallat, A.; Luptakova, L.; Petrovova, E.; Oszako, T.; Cherrad, S.; Vacher, S. Comparative Genomics of Bacillus amyloliquefaciens Strains Reveals a Core Genome with Traits for Habitat Adaptation and a Secondary Metabolites Rich Accessory Genome. Front. Microbiol. 2017, 8, 1438. [Google Scholar] [CrossRef]
- Adeniji, A.A.; Babalola, O.O. Bacillus velezensis: Phylogeny, Useful Applications, and Avenues for Exploitation. Appl. Microbiol. Biotechnol. 2019, 103, 3669–3682. [Google Scholar] [CrossRef] [PubMed]
- Guo, Q.; Li, Y.; Lou, Y.; Shi, M.; Jiang, Y.; Zhou, J.; Sun, Y.; Xue, Q.; Lai, H. Bacillus amyloliquefaciens Ba13 Induces Plant Systemic Resistance and Improves Rhizosphere Microecology against Tomato Yellow Leaf Curl Virus Disease. Appl. Soil Ecol. 2019, 137, 154–166. [Google Scholar] [CrossRef]
- Borriss, R.; Chen, X.-H.; Rueckert, C.; Blom, J.; Becker, A.; Baumgarth, B.; Fan, B.; Pukall, R.; Schumann, P.; Spröer, C. Relationship of Bacillus amyloliquefaciens Clades Associated with Strains DSM 7T and FZB42T: A Proposal for Bacillus amyloliquefaciens Subsp. amyloliquefaciens Subsp. Nov. and Bacillus amyloliquefaciens Subsp. plantarum Subsp. Nov. Based on Complete Genome Sequence Comparisons. Int. J. Syst. Evol. Microbiol. 2011, 61, 1786–1801. [Google Scholar]
- Chowdhury, S.P.; Dietel, K.; Rändler, M.; Schmid, M.; Junge, H.; Borriss, R.; Hartmann, A.; Grosch, R. Effects of Bacillus amyloliquefaciens FZB42 on Lettuce Growth and Health under Pathogen Pressure and Its Impact on the Rhizosphere Bacterial Community. PLoS ONE 2013, 8, e68818. [Google Scholar] [CrossRef]
- Wu, L.; Wu, H.-J.; Qiao, J.; Gao, X.; Borriss, R. Novel Routes for Improving Biocontrol Activity of Bacillus Based Bioinoculants. Front. Microbiol. 2015, 6, 1395. [Google Scholar] [CrossRef]
- Pandin, C.; Le Coq, D.; Deschamps, J.; Védie, R.; Rousseau, T.; Aymerich, S.; Briandet, R. Complete Genome Sequence of Bacillus velezensis QST713: A Biocontrol Agent That Protects Agaricus bisporus Crops against the Green Mould Disease. J. Biotechnol. 2018, 278, 10–19. [Google Scholar] [CrossRef]
- Cao, Y.; Pi, H.; Chandrangsu, P.; Li, Y.; Wang, Y.; Zhou, H.; Xiong, H.; Helmann, J.D.; Cai, Y. Antagonism of Two Plant-Growth Promoting Bacillus velezensis Isolates against Ralstonia solanacearum and Fusarium oxysporum. Sci. Rep. 2018, 8, 4360. [Google Scholar] [CrossRef] [PubMed]
- Ben Slama, H.; Triki, M.A.; Chenari Bouket, A.; Ben Mefteh, F.; Alenezi, F.N.; Luptakova, L.; Cherif-Silini, H.; Vallat, A.; Oszako, T.; Gharsallah, N.; et al. Screening of the High-Rhizosphere Competent Limoniastrum monopetalum’ Culturable Endophyte Microbiota Allows the Recovery of Multifaceted and Versatile Biocontrol Agents. Microorganisms 2019, 7, 249. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.-H.; Park, J.; Lim, J.Y.; Kim, H.; Choi, G.J.; Kim, J.-C.; Seo, Y.-S. Complete Genome Sequence of Bacillus velezensis G341, a Strain with a Broad Inhibitory Spectrum against Plant Pathogens. J. Biotechnol. 2015, 211, 97–98. [Google Scholar] [CrossRef] [PubMed]
- Chen, L. Complete Genome Sequence of Bacillus velezensis LM2303, a Biocontrol Strain Isolated from the Dung of Wild Yak Inhabited Qinghai-Tibet Plateau. J. Biotechnol. 2017, 251, 124–127. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.; Koh, I.; Lim, M.Y.; Chung, W.-H.; Rho, M. Pan-Genome Analysis of Bacillus for Microbiome Profiling. Sci. Rep. 2017, 7, 10984. [Google Scholar] [CrossRef] [PubMed]
- Baptista, J.P.; Sanches, P.P.; Teixeira, G.M.; Morey, A.T.; Tavares, E.R.; Yamada-Ogatta, S.F.; Da Rocha, S.P.D.; Hungria, M.; Ribeiro, R.A.; Balbi-Peña, M.I. Complete Genome Sequence of Bacillus velezensis LABIM40, an Effective Antagonist of Fungal Plant Pathogens. Genome Announc. 2018, 6, e00595-18. [Google Scholar] [CrossRef]
- Rabbee, M.F.; Ali, M.; Choi, J.; Hwang, B.S.; Jeong, S.C.; Baek, K. Bacillus velezensis: A Valuable Member of Bioactive Molecules within Plant Microbiomes. Molecules 2019, 24, 1046. [Google Scholar] [CrossRef]
- Cheffi, M.; Chenari Bouket, A.; Alenezi, F.N.; Luptakova, L.; Belka, M.; Vallat, A.; Rateb, M.E.; Tounsi, S.; Triki, M.A.; Belbahri, L. Olea europaea L. Root Endophyte Bacillus velezensis OEE1 Counteracts Oomycete and Fungal Harmful Pathogens and Harbours a Large Repertoire of Secreted and Volatile Metabolites and Beneficial Functional Genes. Microorganisms 2019, 7, 314. [Google Scholar] [CrossRef]
- Yao, A.V.; Bochow, H.; Karimov, S.; Boturov, U.; Sanginboy, S.; Sharipov, A.K. Effect of FZB 24(R) Bacillus subtilis as a Biofertilizer on Cotton Yields in Field Tests. Arch. Phytopathol. Plant Prot. 2006, 39, 323–328. [Google Scholar] [CrossRef]
- Idris, E.E.; Iglesias, D.J.; Talon, M.; Borriss, R. Tryptophan-Dependent Production of Indole-3-Acetic Acid (IAA) Affects Level of Plant Growth Promotion by Bacillus amyloliquefaciens FZB42. Mol. Plant Microbe Interact. 2007, 20, 619–626. [Google Scholar] [CrossRef]
- Le Mire, G.; Nguyen, M.; Fassotte, B.; du Jardin, P.; Verheggen, F.; Delaplace, P.; Jijakli, H. Implementing Biostimulants and Biocontrol Strategies in the Agroecological Management of Cultivated Ecosystems. Biotechnol. Agron. Soc. Environ. 2016, 20, 299–313. [Google Scholar]
- Ye, M.; Tang, X.; Yang, R.; Zhang, H.; Li, F.; Tao, F.; Li, F.; Wang, Z. Characteristics and Application of a Novel Species of Bacillus: Bacillus velezensis. ACS Chem. Biol. 2018, 13, 500–505. [Google Scholar] [CrossRef]
- Nguyen, M.L.; Glaes, J.; Spaepen, S.; Bodson, B.; Du Jardin, P.; Delaplace, P. Biostimulant Effects of Bacillus Strains on Wheat from In Vitro towards Field Conditions Are Modulated by Nitrogen Supply. J. Plant Nutr. Soil Sci. 2019, 182, 325–334. [Google Scholar] [CrossRef]
- Xu, Z.; Zhang, H.; Sun, X.; Liu, Y.; Yan, W.; Xun, W.; Shen, Q.; Zhang, R. Bacillus velezensis Wall Teichoic Acids Are Required for Biofilm Formation and Root Colonization. Appl. Environ. Microbiol. 2019, 85, e02116–e02118. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Garcia, C.; Bejar, V.; Martinez-Checa, F.; Llamas, I.; Quesada, E. Bacillus velezensis Sp. Nov., a Surfactant-Producing Bacterium Isolated from the River Velez in Malaga, Southern Spain. Int. J. Syst. Evol. Microbiol. 2005, 55, 191–195. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.H.; Koumoutsi, A.; Scholz, R.; Eisenreich, A.; Schneider, K.; Heinemeyer, I.; Morgenstern, B.; Voss, B.; Hess, W.R.; Reva, O. Comparative Analysis of the Complete Genome Sequence of the Plant Growth-Promoting Bacterium Bacillus amyloliquefaciens FZB42. Nat. Biotechnol. 2007, 25, 1007–1014. [Google Scholar] [CrossRef]
- Dunlap, C.A.; Kim, S.-J.; Kwon, S.-W.; Rooney, A.P. Phylogenomic Analysis Shows That Bacillus amyloliquefaciens Subsp. Plantarum Is a Later Heterotypic Synonym of Bacillus methylotrophicus. Int. J. Syst. Evol. Microbiol. 2015, 65, 2104–2109. [Google Scholar] [CrossRef]
- Palazzini, J.M.; Dunlap, C.A.; Bowman, M.J.; Chulze, S.N. Bacillus velezensis RC 218 as a Biocontrol Agent to Reduce Fusarium Head Blight and Deoxynivalenol Accumulation: Genome Sequencing and Secondary Metabolite Cluster Profiles. Microbiol. Res. 2016, 192, 30–36. [Google Scholar] [CrossRef]
- Cai, X.-C.; Liu, C.-H.; Wang, B.-T.; Xue, Y.-R. Genomic and Metabolic Traits Endow Bacillus velezensis CC09 with a Potential Biocontrol Agent in Control of Wheat Powdery Mildew Disease. Microbiol. Res. 2017, 196, 89–94. [Google Scholar] [CrossRef]
- Stein, T. Bacillus Subtilis Antibiotics: Structures, Syntheses and Specific Functions. Mol. Microbiol. 2005, 56, 845–857. [Google Scholar] [CrossRef]
- Arguelles-Arias, A.; Ongena, M.; Halimi, B.; Lara, Y.; Brans, A.; Joris, B.; Fickers, P. Bacillus amyloliquefaciens GA1 as a Source of Potent Antibiotics and Other Secondary Metabolites for Biocontrol of Plant Pathogens. Microb. Cell Fact 2009, 8, 63. [Google Scholar] [CrossRef]
- Rückert, C.; Blom, J.; Chen, X.; Reva, O.; Borriss, R. Genome Sequence of B. amyloliquefaciens Type Strain DSM7T Reveals Differences to Plant-Associated B. amyloliquefaciens FZB42. J. Biotechnol. 2011, 155, 78–85. [Google Scholar] [CrossRef]
- He, P.; Hao, K.; Blom, J.; Rückert, C.; Vater, J.; Mao, Z.; Wu, Y.; Hou, M.; He, P.; He, Y. Genome Sequence of the Plant Growth Promoting Strain Bacillus amyloliquefaciens Subsp. plantarum B9601-Y2 and Expression of Mersacidin and Other Secondary Metabolites. J. Biotechnol. 2013, 164, 281–291. [Google Scholar] [CrossRef]
- Zhao, X.; Kuipers, O.P. Identification and Classification of Known and Putative Antimicrobial Compounds Produced by a Wide Variety of Bacillales Species. BMC Genom. 2016, 17, 882. [Google Scholar] [CrossRef]
- Liu, G.; Kong, Y.; Fan, Y.; Geng, C.; Peng, D.; Sun, M. Whole-Genome Sequencing of Bacillus velezensis LS69, a Strain with a Broad Inhibitory Spectrum against Pathogenic Bacteria. J. Biotechnol. 2017, 249, 20–24. [Google Scholar] [CrossRef]
- Fan, B.; Blom, J.; Klenk, H.-P.; Borriss, R. Bacillus amyloliquefaciens, Bacillus velezensis, and Bacillus siamensis Form an “Operational Group B. amyloliquefaciens” within the B. subtilis Species Complex. Front. Microbiol. 2017, 8, 22. [Google Scholar] [CrossRef]
- Wang, J.; Xing, J.; Lu, J.; Sun, Y.; Zhao, J.; Miao, S.; Xiong, Q.; Zhang, Y.; Zhang, G. Complete Genome Sequencing of Bacillus velezensis WRN014, and Comparison with Genome Sequences of Other Bacillus velezensis Strains. J. Microbiol. Biotechnol. 2019, 29, 794–808. [Google Scholar] [CrossRef]
- Rooney, A.P.; Price, N.P.; Ehrhardt, C.; Swezey, J.L.; Bannan, J.D. Phylogeny and Molecular Taxonomy of the Bacillus subtilis Species Complex and Description of Bacillus subtilis Subsp. inaquosorum Subsp. Nov. Int. J. Syst. Evol. Microbiol. 2009, 59, 2429–2436. [Google Scholar] [CrossRef]
- Dunlap, C.A.; Kim, S.-J.; Kwon, S.-W.; Rooney, A.P. Bacillus velezensis Is Not a Later Heterotypic Synonym of Bacillus amyloliquefaciens; Bacillus methylotrophicus, Bacillus amyloliquefaciens Subsp. plantarum and ‘Bacillus oryzicola’ Are Later Heterotypic Synonyms of Bacillus velezensis Based on Phylogenomics. Int. J. Syst. Evol. Microbiol. 2016, 66, 1212–1217. [Google Scholar] [PubMed]
- Fan, B.; Wang, C.; Ding, X.L.; Song, X.; Wu, L.; Wu, H.; Xuewen, G.; Borriss, R. Corrigendum: Bacillus velezensis FZB42 in 2018: The Gram-Positive Model Strain for Plant Growth Promotion and Biocontrol. Front. Microbiol. 2019, 10, 1279. [Google Scholar] [CrossRef]
- Parks, D.H.; Imelfort, M.; Skennerton, C.T.; Hugenholtz, P.; Tyson, G.W. CheckM: Assessing the Quality of Microbial Genomes Recovered from Isolates, Single Cells, and Metagenomes. Genome Res. 2015, 25, 1043–1055. [Google Scholar] [CrossRef]
- Bertels, F.; Silander, O.K.; Pachkov, M.; Rainey, P.B.; Van Nimwegen, E. Automated Reconstruction of Whole-Genome Phylogenies from Short-Sequence Reads. Mol. Biol. Evol. 2014, 31, 1077–1088. [Google Scholar] [CrossRef]
- Felsenstein, J. Evolutionary Trees from DNA Sequences: A Maximum Likelihood Approach. J. Mol. Evol. 1981, 17, 368–376. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis Version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef] [PubMed]
- Kimura, M. A Simple Method for Estimating Evolutionary Rates of Base Substitutions through Comparative Studies of Nucleotide Sequences. J. Mol. Evol. 1980, 16, 111–120. [Google Scholar] [CrossRef]
- Goris, J.; Konstantinidis, K.T.; Klappenbach, J.A.; Coenye, T.; Vandamme, P.; Tiedje, J.M. DNA–DNA Hybridization Values and Their Relationship to Whole-Genome Sequence Similarities. Int. J. Syst. Evol. Microbiol. 2007, 57, 81–91. [Google Scholar] [CrossRef]
- Richter, M.; Rosselló-Móra, R. Shifting the Genomic Gold Standard for the Prokaryotic Species Definition. Proc. Natl. Acad. Sci. USA 2009, 106, 19126–19131. [Google Scholar] [CrossRef]
- Yoon, S.-H.; Ha, S.-M.; Kwon, S.; Lim, J.; Kim, Y.; Seo, H.; Chun, J. Introducing EzBioCloud: A Taxonomically United Database of 16S rRNA Gene Sequences and Whole-Genome Assemblies. Int. J. Syst. Evol. Microbiol. 2017, 67, 1613–1617. [Google Scholar] [CrossRef] [PubMed]
- Meier-Kolthoff, J.P.; Auch, A.F.; Klenk, H.-P.; Göker, M. Genome Sequence-Based Species Delimitation with Confidence Intervals and Improved Distance Functions. BMC Bioinf. 2013, 14, 60. [Google Scholar] [CrossRef]
- Chaudhari, N.M.; Gupta, V.K.; Dutta, C. BPGA-an Ultra-Fast Pan-Genome Analysis Pipeline. Sci. Rep. 2016, 6, 24373. [Google Scholar] [CrossRef]
- Kanehisa, M.; Sato, Y.; Morishima, K. BlastKOALA and GhostKOALA: KEGG Tools for Functional Characterization of Genome and Metagenome Sequences. J. Mol. Biol. 2016, 428, 726–731. [Google Scholar] [CrossRef]
- Weber, T.; Blin, K.; Duddela, S.; Krug, D.; Kim, H.U.; Bruccoleri, R.; Lee, S.Y.; Fischbach, M.A.; Müller, R.; Wohlleben, W. antiSMASH 3.0—A Comprehensive Resource for the Genome Mining of Biosynthetic Gene Clusters. Nucleic Acids Res. 2015, 43, W237–W243. [Google Scholar] [CrossRef]
- Ziemert, N.; Podell, S.; Penn, K.; Badger, J.H.; Allen, E.; Jensen, P.R. The Natural Product Domain Seeker NaPDoS: A Phylogeny Based Bioinformatic Tool to Classify Secondary Metabolite Gene Diversity. PLoS ONE 2012, 7, e34064. [Google Scholar] [CrossRef] [PubMed]
- Li, M.H.; Ung, P.M.; Zajkowski, J.; Garneau-Tsodikova, S.; Sherman, D.H. Automated Genome Mining for Natural Products. BMC Bioinf. 2009, 10, 185. [Google Scholar] [CrossRef]
- Van Heel, A.J.; De Jong, A.; Montalbán-López, M.; Kok, J.; Kuipers, O.P. BAGEL3: Automated Identification of Genes Encoding Bacteriocins and (Non-) Bactericidal Post-Translationally Modified Peptides. Nucleic Acids Res. 2013, 41, W448–W453. [Google Scholar] [CrossRef] [PubMed]
- Bruto, M.; Prigent-Combaret, C.; Muller, D.; Moënne-Loccoz, Y. Analysis of Genes Contributing to Plant-Beneficial Functions in Plant Growth-Promoting Rhizobacteria and Related Proteobacteria. Sci. Rep. 2014, 4, 6261. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.T.; Lee, L.Y.; Tai, C.Y.; Hung, C.H.; Chang, Y.S.; Wolfram, J.H.; Rogers, R.; Goldstein, A.H. Cloning of an Erwinia herbicola Gene Necessary for Gluconic Acid Production and Enhanced Mineral Phosphate Solubilization in Escherichia coli HB101: Nucleotide Sequence and Probable Involvement in Biosynthesis of the Coenzyme Pyrroloquinoline Quinone. J. Bacteriol. 1992, 174, 5814–5819. [Google Scholar] [CrossRef][Green Version]
- Miller, S.H.; Browne, P.; Prigent-Combaret, C.; Combes-Meynet, E.; Morrissey, J.P.; O’Gara, F. Biochemical and Genomic Comparison of Inorganic Phosphate Solubilization in Pseudomonas Species. Environ. Microbiol. Rep. 2010, 2, 403–411. [Google Scholar] [CrossRef]
- Shao, J.; Li, S.; Zhang, N.; Cui, X.; Zhou, X.; Zhang, G.; Shen, Q.; Zhang, R. Analysis and Cloning of the Synthetic Pathway of the Phytohormone Indole-3-Acetic Acid in the Plant-Beneficial Bacillus amyloliquefaciens SQR9. Microb. Cell Fact 2015, 14, 130. [Google Scholar] [CrossRef] [PubMed]
- Niazi, A.; Manzoor, S.; Asari, S.; Bejai, S.; Meijer, J.; Bongcam-Rudloff, E. Genome Analysis of Bacillus amyloliquefaciens Subsp. plantarum UCMB5113: A Rhizobacterium That Improves Plant Growth and Stress Management. PLoS ONE 2014, 9, e104651. [Google Scholar] [CrossRef]
- Petersen, T.N.; Brunak, S.; Von Heijne, G.; Nielsen, H. SignalP 4.0: Discriminating Signal Peptides from Transmembrane Regions. Nat. Methods 2011, 8, 785–786. [Google Scholar] [CrossRef] [PubMed]
- Gupta, A.; Gopal, M.; Thomas, G.V.; Manikandan, V.; Gajewski, J.; Thomas, G.; Seshagiri, S.; Schuster, S.C.; Rajesh, P.; Gupta, R. Whole Genome Sequencing and Analysis of Plant Growth Promoting Bacteria Isolated from the Rhizosphere of Plantation Crops Coconut, Cocoa and Arecanut. PLoS ONE 2014, 9, e104259. [Google Scholar] [CrossRef] [PubMed]
- Boch, J.; Kempf, B.; Schmid, R.; Bremer, E. Synthesis of the Osmoprotectant Glycine Betaine in Bacillus subtilis: Characterization of the gbsAB Genes. J. Bacteriol. 1996, 178, 5121–5129. [Google Scholar] [CrossRef]
- Mazzola, M.; Cook, R.J.; Thomashow, L.S.; Weller, D.M.; Pierson, L.S. Contribution of Phenazine Antibiotic Biosynthesis to the Ecological Competence of Fluorescent Pseudomonads in Soil Habitats. Appl. Environ. Microbiol. 1992, 58, 2616–2624. [Google Scholar] [CrossRef] [PubMed]
- Ghelardi, E.; Salvetti, S.; Ceragioli, M.; Gueye, S.A.; Celandroni, F.; Senesi, S. Contribution of Surfactin and swrA to Flagellin Expression, Swimming, and Surface Motility in Bacillus subtilis. Appl. Environ. Microbiol. 2012, 78, 6540–6544. [Google Scholar] [CrossRef]
- Shen, X.; Hu, H.; Peng, H.; Wang, W.; Zhang, X. Comparative Genomic Analysis of Four Representative Plant Growth-Promoting Rhizobacteria in Pseudomonas. BMC Genom. 2013, 14, 271. [Google Scholar] [CrossRef] [PubMed]
- Vlamakis, H.; Chai, Y.; Beauregard, P.; Losick, R.; Kolter, R. Sticking Together: Building a Biofilm the Bacillus subtilis Way. Nat. Rev. Microbiol. 2013, 11, 157–168. [Google Scholar] [CrossRef]
- Palacios, O.A.; Choix, F.J.; Bashan, Y.; de-Bashan, L.E. Influence of tryptophan and indole-3-acetic acid on starch accumulation in the synthetic mutualistic Chlorella sorokiniana-Azospirillum brasilense system under heterotrophic conditions. Res Microbiol. 2016, 167, 367–379. [Google Scholar] [CrossRef]
- Whistler, C.A.; Corbell, N.A.; Sarniguet, A.; Ream, W.; Loper, J.E. The Two-Component Regulators GacS and GacA Influence Accumulation of the Stationary-Phase Sigma Factor ςS and the Stress Response in Pseudomonas fluorescens Pf-5. J. Bacteriol. 1998, 180, 6635–6641. [Google Scholar] [CrossRef]
- Ochsner, U.A.; Vasil, M.L.; Alsabbagh, E.; Parvatiyar, K.; Hassett, D.J. Role of the Pseudomonas aeruginosa oxyR-recG Operon in Oxidative Stress Defense and DNA Repair: OxyR-Dependent Regulation of katB-ankB, ahpB, and ahpC-ahpF. J. Bacteriol. 2000, 182, 4533–4544. [Google Scholar] [CrossRef]
- Loper, J.E.; Hassan, K.A.; Mavrodi, D.V.; Ii, E.W.D.; Lim, C.K.; Shaffer, B.T.; Elbourne, L.D.H.; Stockwell, V.O.; Hartney, S.L.; Breakwell, K.; et al. Comparative Genomics of Plant-Associated Pseudomonas Spp.: Insights into Diversity and Inheritance of Traits Involved in Multitrophic Interactions. PLoS Genet. 2012, 8, e1002784. [Google Scholar] [CrossRef]
- Someya, Y.; Yamaguchi, A.; Sawai, T. A Novel Glycylcycline, 9-(N,N-Dimethylglycylamido)-6-Demethyl-6-Deoxytetracycline, Is Neither Transported Nor Recognized by the Transposon Tn10-Encoded Metal-Tetracycline/H+ Antiporter. Antimicrob. Agents Chemother 1995, 39, 247–249. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Sakaguchi, R.; Amano, H.; Shishido, K. Nucleotide Sequence Homology of the Tetracycline-Resistance Determinant Naturally Maintained in Bacillus subtilis Marburg 168 Chromosome and the Tetracycline-Resistance Gene of B. subtilis Plasmid pNS1981. Biochim. Et Biophys. Acta (BBA)—Gene Struct. Expr. 1988, 950, 441–444. [Google Scholar] [CrossRef]
- Jacob, J.; Evers, S.; Bischoff, K.; Carlier, C.; Courvalin, P. Characterization of the Sat 4 Gene Encoding a Streptothricin Acetyltransferase in Campylobacter coli BE/G4. FEMS Microbiol. Lett. 1994, 120, 13–17. [Google Scholar] [PubMed]
- Fu, Z.; Liu, Y.; Chen, C.; Guo, Y.; Ma, Y.; Yang, Y.; Hu, F.; Xu, X.; Wang, M. Characterization of Fosfomycin Resistance Gene, fosB, in Methicillin-Resistant Staphylococcus aureus Isolates. PLoS ONE 2016, 11, e0154829. [Google Scholar] [CrossRef]
- Rey, M.W.; Ramaiya, P.; Nelson, B.A.; Brody-Karpin, S.D.; Zaretsky, E.J.; Tang, M.; De Leon, A.L.; Xiang, H.; Gusti, V.; Clausen, I.G.; et al. Complete Genome Sequence of the Industrial Bacterium Bacillus licheniformis and Comparisons with Closely Related Bacillus Species. Genome Biol. 2004, 5, r77. [Google Scholar] [CrossRef]
- Barbe, V.; Cruveiller, S.; Kunst, F.; Lenoble, P.; Meurice, G.; Sekowska, A.; Vallenet, D.; Wang, T.; Moszer, I.; Médigue, C.; et al. From a Consortium Sequence to a Unified Sequence: The Bacillus subtilis 168 Reference Genome a Decade Later. Microbiology 2009, 155, 1758–1775. [Google Scholar] [CrossRef]
- Neyfakh, A.A.; Borsch, C.M.; Kaatz, G.W. Fluoroquinolone Resistance Protein NorA of Staphylococcus aureus Is a Multidrug Efflux Transporter. Antimicrob. Agents Chemother 1993, 37, 128–129. [Google Scholar] [CrossRef] [PubMed]
- Doublet, B.; Schwarz, S.; Kehrenberg, C.; Cloeckaert, A. Florfenicol Resistance Gene floR Is Part of a Novel Transposon. Antimicrob. Agents Chemother 2005, 49, 2106–2108. [Google Scholar] [CrossRef]
- Noguchi, N.; Sasatsu, M.; Kono, M. Genetic Mapping in Bacillus subtilis 168 of the aadK Gene Which Encodes Aminoglycoside 6-Adenylyltransferase. FEMS Microbiol. Lett. 1993, 114, 47–52. [Google Scholar] [CrossRef]
- Hosoya, S.; Yamane, K.; Takeuchi, M.; Sato, T. Identification and Characterization of the Bacillus subtilis D-Glucarate/Galactarate Utilization Operon ycbCDEFGHJ. FEMS Microbiol. Lett. 2002, 210, 193–199. [Google Scholar] [CrossRef]
- Ohki, R.; Tateno, K.; Takizawa, T.; Aiso, T.; Murata, M. Transcriptional Termination Control of a Novel ABC Transporter Gene Involved in Antibiotic Resistance in Bacillus subtilis. J. Bacteriol. 2005, 187, 5946–5954. [Google Scholar] [CrossRef]
- Duan, J.; Jiang, W.; Cheng, Z.; Heikkila, J.J.; Glick, B.R. The Complete Genome Sequence of the Plant Growth-Promoting Bacterium Pseudomonas Sp. UW4. PLoS ONE 2013, 8, e58640. [Google Scholar] [CrossRef]
- Rademacher, C.; Masepohl, B. Copper-Responsive Gene Regulation in Bacteria. Microbiology 2012, 158, 2451–2464. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Wang, D.; Wang, Y.; Hasman, H.; Aarestrup, F.M.; Alwathnani, H.A.; Zhu, Y.-G.; Rensing, C. Genome Sequences of Copper Resistant and Sensitive Enterococcus faecalis Strains Isolated from Copper-Fed Pigs in Denmark. Stand. Genom. Sci. 2015, 10, 35. [Google Scholar] [CrossRef] [PubMed]
- Hu, K.H.; Liu, E.; Dean, K.; Gingas, M.; DeGraff, W.; Trun, N.J. Overproduction of Three Genes Leads to Camphor Resistance and Chromosome Condensation in Escherichia coli. Genetics 1996, 143, 1521–1532. [Google Scholar] [CrossRef] [PubMed]
- Franks, S.E.; Ebrahimi, C.; Hollands, A.; Okumura, C.Y.; Aroian, R.V.; Nizet, V.; McGillivray, S.M. Novel Role for the yceGH Tellurite Resistance Genes in the Pathogenesis of Bacillus anthracis. Infect. Immun. 2014, 82, 1132–1140. [Google Scholar] [CrossRef][Green Version]
- Moore, C.M.; Gaballa, A.; Hui, M.; Ye, R.W.; Helmann, J.D. Genetic and Physiological Responses of Bacillus subtilis to Metal Ion Stress. Mol. Microbiol. 2005, 57, 27–40. [Google Scholar] [CrossRef]
- Wu, X.; Wang, X.; Drlica, K.; Zhao, X. A Toxin-Antitoxin Module in Bacillus subtilis Can Both Mitigate and Amplify Effects of Lethal Stress. PLoS ONE 2011, 6, e23909. [Google Scholar] [CrossRef] [PubMed]
- Osman, D.; Cavet, J.S. Bacterial Metal-Sensing Proteins Exemplified by ArsR–SmtB Family Repressors. Nat. Prod. Rep. 2010, 27, 668–680. [Google Scholar] [CrossRef]
- Graf, N.; Wenzel, M.; Altenbuchner, J. Identification and Characterization of the Vanillin Dehydrogenase YfmT in Bacillus subtilis 3NA. Appl. Microbiol. Biotechnol. 2016, 100, 3511–3521. [Google Scholar] [CrossRef] [PubMed]
- Piddington, C.S.; Kovacevich, B.R.; Rambosek, J. Sequence and Molecular Characterization of a DNA Region Encoding the Dibenzothiophene Desulfurization Operon of Rhodococcus Sp. Strain IGTS8. Appl. Environ. Microbiol. 1995, 61, 468–475. [Google Scholar] [CrossRef]
- Duy, N.V.; Wolf, C.; Mäder, U.; Lalk, M.; Langer, P.; Lindequist, U.; Hecker, M.; Antelmann, H. Transcriptome and Proteome Analyses in Response to 2-Methylhydroquinone and 6-Brom-2-Vinyl-Chroman-4-on Reveal Different Degradation Systems Involved in the Catabolism of Aromatic Compounds in Bacillus subtilis. Proteomics 2007, 7, 1391–1408. [Google Scholar]
- Ozer, E.A.; Allen, J.P.; Hauser, A.R. Characterization of the Core and Accessory Genomes of Pseudomonas aeruginosa Using Bioinformatic Tools Spine and AGEnt. BMC Genom. 2014, 15, 737. [Google Scholar] [CrossRef] [PubMed]
- Tettelin, H.; Riley, D.; Cattuto, C.; Medini, D. Comparative Genomics: The Bacterial Pan-Genome. Curr. Opin. Microbiol. 2008, 11, 472–477. [Google Scholar] [CrossRef] [PubMed]
- Skinnider, M.A.; Dejong, C.A.; Rees, P.N.; Johnston, C.W.; Li, H.; Webster, A.L.H.; Wyatt, M.A.; Magarvey, N.A. Genomes to Natural Products Prediction Informatics for Secondary Metabolomes (PRISM). Nucleic Acids Res. 2015, 43, 9645–9662. [Google Scholar] [CrossRef]
- Chun, B.H.; Kim, K.H.; Jeong, S.E.; Jeon, C.O. Genomic and Metabolic Features of the Bacillus amyloliquefaciens Group—B. amyloliquefaciens, B. velezensis, and B. siamensis—Revealed by Pan-Genome Analysis. Food Microbiol. 2019, 77, 146–157. [Google Scholar] [CrossRef] [PubMed]
- Yi, H.; Chun, J.; Cha, C.-J. Genomic Insights into the Taxonomic Status of the Three Subspecies of Bacillus subtilis. Syst. Appl. Microbiol. 2014, 37, 95–99. [Google Scholar] [CrossRef]
- Li, W.; Jia, M.-X.; Deng, J.; Wang, J.-H.; Lin, Q.-L.; Liu, C.; Wang, S.-S.; Tang, J.-X.; Zeng, X.-X.; Ma, L.; et al. Isolation, Genetic Identification and Degradation Characteristics of COD-Degrading Bacterial Strain in Slaughter Wastewater. Saudi J. Biol. Sci. 2018, 25, 1800–1805. [Google Scholar] [CrossRef]
- Luo, C.; Chen, Y.; Liu, X.; Wang, X.; Wang, X.; Li, X.; Zhao, Y.; Wei, L. Engineered Biosynthesis of Cyclic Lipopeptide Locillomycins in Surrogate Host Bacillus velezensis FZB42 and Derivative Strains Enhance Antibacterial Activity. Appl. Microbiol. Biotechnol. 2019, 103, 4467–4481. [Google Scholar] [CrossRef]
- Meng, Q.; Jiang, H.; Hao, J.J. Effects of Bacillus velezensis Strain BAC03 in Promoting Plant Growth. Biol. Control 2016, 98, 18–26. [Google Scholar] [CrossRef]
- Grady, E.N.; MacDonald, J.; Ho, M.T.; Weselowski, B.; McDowell, T.; Solomon, O.; Renaud, J.; Yuan, Z.-C. Characterization and Complete Genome Analysis of the Surfactin-Producing, Plant-Protecting Bacterium Bacillus velezensis 9D-6. BMC Microbiol. 2019, 19, 5. [Google Scholar] [CrossRef] [PubMed]
- Meena, K.R.; Tandon, T.; Sharma, A.; Kanwar, S.S. Lipopeptide Antibiotic Production by Bacillus velezensis KLP2016. J. Appl. Pharm. Sci. 2018, 8, 91–98. [Google Scholar]
- Bafana, A.; Chakrabarti, T.; Devi, S.S. Azoreductase and Dye Detoxification Activities of Bacillus velezensis Strain AB. Appl. Microbiol. Biotechnol. 2008, 77, 1139–1144. [Google Scholar] [CrossRef] [PubMed]
- Federhen, S. Type Material in the NCBI Taxonomy Database. Nucleic Acids Res. 2014, 43, D1086–D1098. [Google Scholar] [CrossRef] [PubMed]
- Slama, H.B.; Cherif-Silini, H.; Chenari Bouket, A.; Qader, M.; Silini, A.; Yahiaoui, B.; Alenezi, F.N.; Luptakova, L.; Triki, M.A.; Vallat, A.; et al. Screening for Fusarium Antagonistic Bacteria from Contrasting Niches Designated the Endophyte Bacillus halotolerans as Plant Warden against Fusarium. Front. Microbiol. 2019, 9. [Google Scholar] [CrossRef]
- Inglin, R.C.; Meile, L.; Stevens, M.J.A. Clustering of Pan-and Core-Genome of Lactobacillus Provides Novel Evolutionary Insights for Differentiation. BMC Genom. 2018, 19, 284. [Google Scholar] [CrossRef]
- Carlos Guimaraes, L.; Benevides de Jesus, L.; Vinicius Canario Viana, M.; Silva, A.; Thiago Juca Ramos, R.; De Castro Soares, S.; Azevedo, V. Inside the Pan-Genome-Methods and Software Overview. Curr. Genom. 2015, 16, 245–252. [Google Scholar] [CrossRef]
- Velivelli, S.L.; De Vos, P.; Kromann, P.; Declerck, S.; Prestwich, B.D. Biological Control Agents: From Field to Market, Problems, and Challenges. Trends Biotechnol. 2014, 32, 493–496. [Google Scholar] [CrossRef] [PubMed]
- Cawoy, H.; Debois, D.; Franzil, L.; De Pauw, E.; Thonart, P.; Ongena, M. Lipopeptides as Main Ingredients for Inhibition of Fungal Phytopathogens by Bacillus subtilis/amyloliquefaciens. Microb. Biotechnol. 2015, 8, 281–295. [Google Scholar] [CrossRef] [PubMed]
- Babalola, O.O.; Ayangbenro, A.S.; Olanrewaju, O.S. Draft Genome Sequences of Three Rhizospheric Plant Growth-Promoting Bacteria. Microbiol. Resour. Announc. 2019, 8, e00455-19. [Google Scholar] [CrossRef]
- Ma, Z.; Hu, J. Complete Genome Sequence of a Marine-Sediment-Derived Bacterial Strain Bacillus velezensis SH-B74, a Cyclic Lipopeptides Producer and a Biopesticide. 3 Biotech 2019, 9, 162. [Google Scholar] [CrossRef]
- Yongsawatdigul, J.; Rodtong, S.; Raksakulthai, N. Acceleration of Thai Fish Sauce Fermentation Using Proteinases and Bacterial Starter Cultures. J. Food Sci. 2007, 72, M382–M390. [Google Scholar] [CrossRef] [PubMed]
- Yao, Z.; Kim, J.A.; Kim, J.H. Characterization of a Fibrinolytic Enzyme Secreted by Bacillus velezensis BS2 Isolated from Sea Squirt Jeotgal. J. Microbiol. Biotechnol. 2019, 29, 347–356. [Google Scholar] [CrossRef]
- Yoshikawa, S.; Kurihara, H.; Kawai, Y.; Yamazaki, K.; Tanaka, A.; Nishikiori, T.; Ohta, T. Effect of Halotolerant Starter Microorganisms on Chemical Characteristics of Fermented Chum Salmon (Oncorhynchus keta) Sauce. J. Agric. Food Chem. 2010, 58, 6410–6417. [Google Scholar] [CrossRef] [PubMed]
- He, G.; Huang, J.; Zhou, R.; Wu, C.; Jin, Y. Effect of Fortified Daqu on the Microbial Community and Flavor in Chinese Strong-Flavor Liquor Brewing Process. Front. Microbiol. 2019, 10, 56. [Google Scholar] [CrossRef] [PubMed]
- Udomsil, N.; Rodtong, S.; Choi, Y.J.; Hua, Y.; Yongsawatdigul, J. Use of Tetragenococcus halophilus as a Starter Culture for Flavor Improvement in Fish Sauce Fermentation. J. Agric. Food Chem. 2011, 59, 8401–8408. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Alenezi, F.N.; Slama, H.B.; Bouket, A.C.; Cherif-Silini, H.; Silini, A.; Luptakova, L.; Nowakowska, J.A.; Oszako, T.; Belbahri, L. Bacillus velezensis: A Treasure House of Bioactive Compounds of Medicinal, Biocontrol and Environmental Importance. Forests 2021, 12, 1714. https://doi.org/10.3390/f12121714
Alenezi FN, Slama HB, Bouket AC, Cherif-Silini H, Silini A, Luptakova L, Nowakowska JA, Oszako T, Belbahri L. Bacillus velezensis: A Treasure House of Bioactive Compounds of Medicinal, Biocontrol and Environmental Importance. Forests. 2021; 12(12):1714. https://doi.org/10.3390/f12121714
Chicago/Turabian StyleAlenezi, Faizah N., Houda Ben Slama, Ali Chenari Bouket, Hafsa Cherif-Silini, Allaoua Silini, Lenka Luptakova, Justyna Anna Nowakowska, Tomasz Oszako, and Lassaad Belbahri. 2021. "Bacillus velezensis: A Treasure House of Bioactive Compounds of Medicinal, Biocontrol and Environmental Importance" Forests 12, no. 12: 1714. https://doi.org/10.3390/f12121714
APA StyleAlenezi, F. N., Slama, H. B., Bouket, A. C., Cherif-Silini, H., Silini, A., Luptakova, L., Nowakowska, J. A., Oszako, T., & Belbahri, L. (2021). Bacillus velezensis: A Treasure House of Bioactive Compounds of Medicinal, Biocontrol and Environmental Importance. Forests, 12(12), 1714. https://doi.org/10.3390/f12121714








