Differentiation of Environmental Conditions Promotes Variation of Two Quercus wutaishanica Community Assembly Patterns
Abstract
1. Introduction
2. Materials and Methods
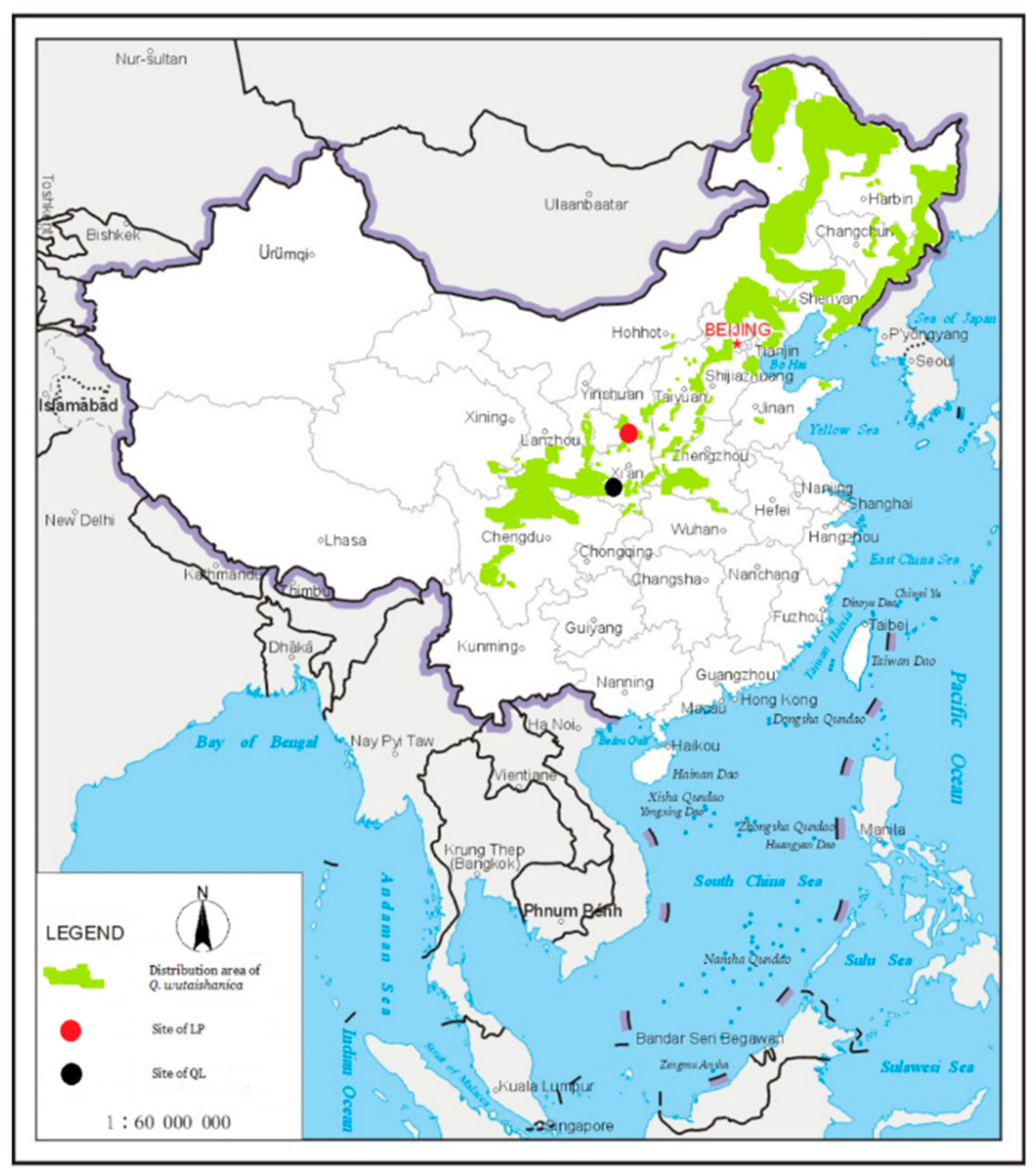
2.1. Study Site
2.2. Floristic Surveys
2.3. Construction of Phylogenetic Tree
2.4. Community Phylogenetic Structure
2.5. Phylogenetic Diversity
3. Results
3.1. Two Different Regional Species Pools
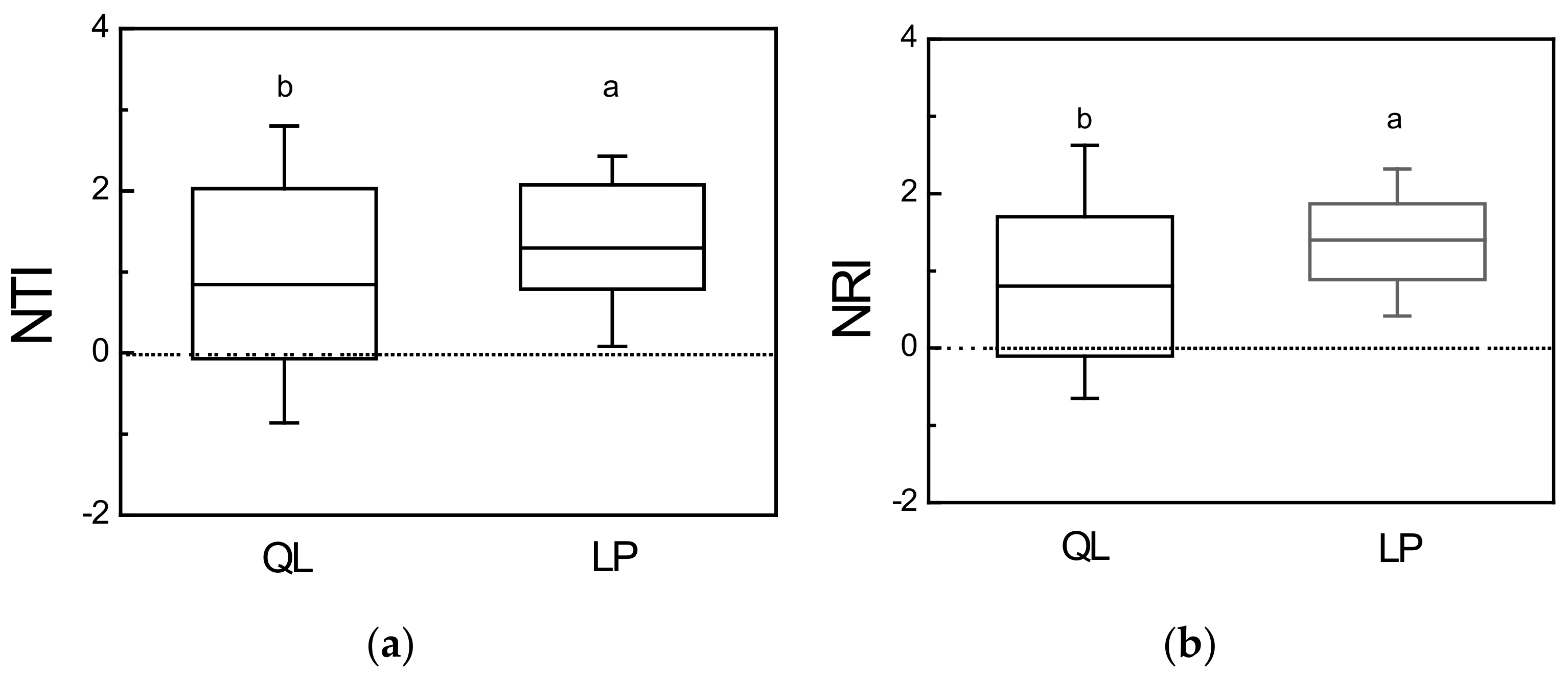
3.2. Phylogenetic Structure
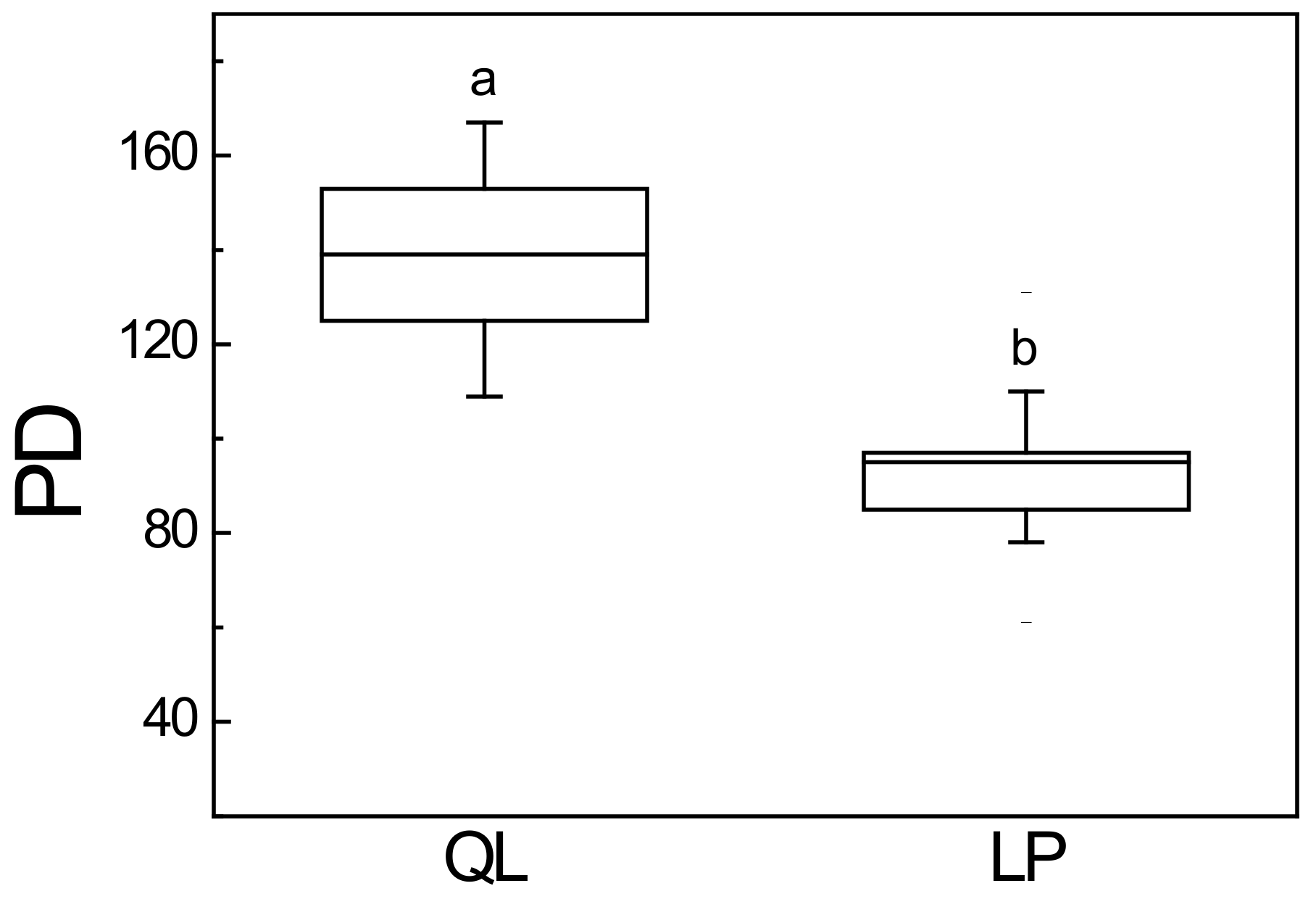
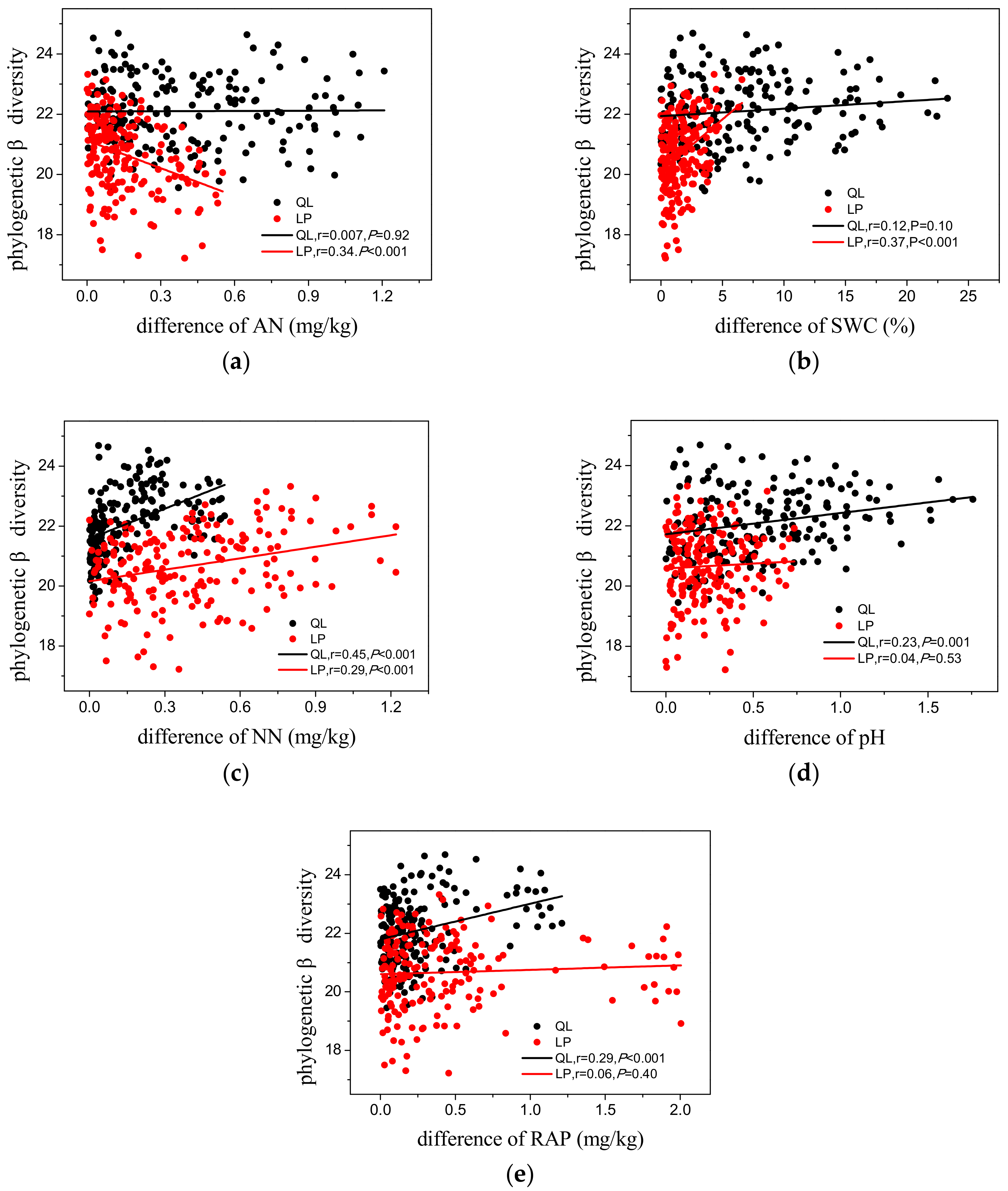
3.3. Phylogenetic Diversity
4. Discussion
4.1. Phylogenetic Structure Reveals Dissimilar Patterns of Community Assembly
4.2. Phylogenetic Patterns of Phylogenetic Diversity
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Weiher, E.; Keddy, P.A. Community assembly rules, morphological dispersion, and the coexistence of plant species. Oikos 1998, 81, 309–322. [Google Scholar] [CrossRef]
- Cornwell, W.K.; Ackerly, D.D. A link between plant traits and abundance: Evidence from coastal California woody plants. J. Ecol. 2010, 98, 814–821. [Google Scholar] [CrossRef]
- Hubbell, S.P. The Unified Neutral Theory of Biodiversity and Biogeography; Princeton University Press: Princeton, NJ, USA, 2001; pp. 32, 76–100. [Google Scholar]
- Zobel, M.; Otto, R.; Laanisto, L.; Naranjo-Cigala, A.; Pärtel, M.; Fernández-Palacios, J.M. The formation of species pools: Historical habitat abundance affects current local diversity. Glob. Ecol. Biogeogr. 2011, 20, 251–259. [Google Scholar] [CrossRef]
- Kraft, N.J.; Valencia, R.; Ackerly, D.D. Functional traits and niche-based tree community assembly in an Amazonian forest. Science 2008, 322, 580–582. [Google Scholar] [CrossRef] [PubMed]
- Kembel, S.W. Disentangling niche and neutral influences on community assembly: Assessing the performance of community phylogenetic structure tests. Ecol. Lett. 2009, 12, 949–960. [Google Scholar] [CrossRef] [PubMed]
- Swenson, N.G.; Enquist, B.J. Opposing assembly mechanisms in a neotropical dry forest: Implications for phylogenetic and functional community ecology. Ecology 2009, 90, 2161–2170. [Google Scholar] [CrossRef]
- Yang, Z.; Powell, J.R.; Zhang, C.; Du, G. The effect of environmental and phylogenetic drivers on community assembly in an alpine meadow community. Ecology 2012, 93, 2321–2328. [Google Scholar] [CrossRef]
- Wang, X.; Swenson, N.G.; Wiegand, T.; Wolf, A.; Howe, R.; Lin, F.; Ye, J.; Yuan, Z.; Shi, S.; Bai, X.; et al. Phylogenetic and functional diversity area relationships in two temperate forests. Ecography 2013, 36, 883–893. [Google Scholar] [CrossRef]
- Grandcolas, P. Phylogenetic analysis and the study of community structure. Oikos 1998, 82, 397–400. [Google Scholar] [CrossRef]
- Andersen, K.M.; Endara, M.J.; Turner, B.L.; Dalling, J.W. Trait-based community assembly of understory palms along a soil nutrient gradient in a lower montane tropical forest. Oecologia 2012, 168, 519–531. [Google Scholar] [CrossRef]
- Kraft, N.J.; Cornwell, W.K.; Webb, C.O.; Ackerly, D.D. Trait evolution, community assembly, and the phylogenetic structure of ecological communities. Am. Nat. 2007, 170, 271–283. [Google Scholar] [CrossRef] [PubMed]
- Schleicher, A.; Peppler-Lisbach, C.; Kleyer, M. Functional traits during succession: Is plant community assembly trait-driven? Preslia 2011, 83, 347–370. [Google Scholar]
- Krishnadas, M.S.; Beckman, N.G.; Penagos-Zuluaga, J.C.; Zhu, Y. Patterns of community assembly and the relative roles of space, environment, phylogeny, and disturbance in explaining functional diversity in a temperate forest. In Proceedings of the 99th ESA Annual Convention, Sacramento, CA, USA, 10–15 August 2014. [Google Scholar]
- Webb, C.O.; Ackerly, D.D.; McPeek, M.A.; Donoghue, M.J. Phylogenies and community ecology. Annu. Rev. Ecol. Syst. 2002, 8, 475–505. [Google Scholar] [CrossRef]
- Kembel, S.W.; Hubbell, S.P. The phylogenetic structure of a neotropical forest tree community. Ecology 2006, 87 (Suppl. 7), 86–99. [Google Scholar] [CrossRef]
- Cavender-Bares, J.; Kozak, K.H.; Fine, P.V.A.; Kembel, S.W. The merging of community ecology and phylogenetic biology. Ecol. Lett. 2009, 12, 693–715. [Google Scholar] [CrossRef]
- Lebrija-Trejos, E.; Meave, J.A.; Bongers, F.; Poorter, L. Functional traits and environmental filtering drive community assembly in a species-rich tropical system. Ecology 2010, 91, 386–398. [Google Scholar] [CrossRef]
- Webb, C.O. Exploring the phylogenetic structure of ecological communities: An example for rain forest trees. Am. Nat. 2000, 156, 145–155. [Google Scholar] [CrossRef]
- Cavender-Bares, J.; Kitajima, K.; Bazzaz, F.A. Multiple trait associations in relation to habitat differentiation among 17 Floridian oak species. Ecol. Monogr. 2004, 74, 635–662. [Google Scholar] [CrossRef]
- Satdichanh, M.; Millet, J.; Heinimann, A.; Nanthavong, K.; Harrison, R.D. Using plant functional traits and phylogenies to understand patterns of plant community assembly in a seasonal tropical forest in Lao PDR. PLoS ONE 2015, 10, e0130151. [Google Scholar] [CrossRef]
- Condit, R.; Pitman, N.; Leigh, E.G., Jr.; Chave, J.; Terborgh, J.; Foster, R.B.; Núnez, P.; Aguilar, S.; Valencia, R.; Villa, G.; et al. Beta-diversity in tropical forest trees. Science 2002, 295, 666–669. [Google Scholar] [CrossRef]
- Legendre, P.; Borcard, D.; Peres-Neto, P.R. Analyzing beta diversity: Partitioning the spatial variation of community composition data. Ecol. Monogr. 2005, 75, 435–450. [Google Scholar] [CrossRef]
- Faith, D.P. Conservation evaluation and phylogenetic diversity. Biol. Conserv. 1992, 61, 1–10. [Google Scholar] [CrossRef]
- Petchey, O.L.; Gaston, K.J. Functional diversity: Back to basics and looking forward. Ecol. Lett. 2006, 9, 741–758. [Google Scholar] [CrossRef]
- Swenson, N.G. The role of evolutionary processes in producing biodiversity patterns, and the interrelationships between taxonomic, functional and phylogenetic biodiversity. Am. J. Bot. 2011, 98, 472–480. [Google Scholar] [CrossRef]
- Gaston, K.J. Global patterns in biodiversity. Nature 2000, 405, 220–227. [Google Scholar] [CrossRef] [PubMed]
- Yue, M. Species diversity of higher plant of Quercus liaotungensis forest in Qingling Mountains and the loess plateau. Acta Bot. Bor. Occid. Sin. 1998, 18, 124–131, (In Chinese with English Abstract). [Google Scholar]
- Wiens, J.J.; Ackerly, D.D.; Allen, A.P.; Anacker, B.L.; Buckley, L.B.; Cornell, H.V.; Damschen, E.I.; Davies, T.J.; Grytnes, J.A.; Harrison, S.P.; et al. Niche conservatism as an emerging principle in ecology and conservation biology. Ecol. Lett. 2010, 13, 1310–1324. [Google Scholar] [CrossRef]
- Xu, J.; Chen, Y.; Wang, G.; Chai, Y.; Wang, M.; Liu, X.; Yue, M. Interspecific Association of Dominant Woody Plant Communities in Qiaoshan Forest Region, Shaanxi. Acta Bot. Bor. Occid. Sin. 2014, 34, 1467–1475, (In Chinese with English Abstract). [Google Scholar]
- Zhu, Z. A preliminary study on the Quercus wutaishanica forests in the Qinling mountain and the Loess Plateau of Northern part of the Shaanxi province. Acta Phytoecol. Geobot. Sin. 1982, 6, 11–20, (In Chinese with English Abstract). [Google Scholar]
- Xu, J.; Chen, Y.; Zhang, L.; Chai, Y.; Wang, M.; Guo, Y.; Li, T.; Yue, M. Using phylogeny and functional traits for assessing community assembly along environmental gradients: A deterministic process driven by elevation. Ecol. Evol. 2017, 7, 5056–5069. [Google Scholar] [CrossRef]
- Webb, C.O.; Donoghue, M.J. Phylomatic: Tree assembly for applied phylogenetics. Mol. Ecol. Notes 2005, 5, 181–183. [Google Scholar] [CrossRef]
- Wikström, N.; Savolainen, V.; Chase, M.W. Evolution of the angiosperms: Calibrating the family tree. Proc. R. Soc. B Biol. Sci. 2001, 268, 2211–2220. [Google Scholar] [CrossRef] [PubMed]
- Webb, C.O.; Ackerly, D.D.; Kembel, S.W. Phylocom: Software for the analysis of phylogenetic community structure and trait evolution. Bioinformatics 2008, 24, 2098–2100. [Google Scholar] [CrossRef] [PubMed]
- Pigot, A.L.; Etienne, R.S. A new dynamic null model for phylogenetic community structure. Ecol. Lett. 2014, 18, 153–163. [Google Scholar] [CrossRef]
- Forest, F.; Grenyer, R.; Rouget, M.; Davies, T.J.; Cowling, R.M.; Faith, D.P.; Balmford, A.; Manning, J.C.; Procheş, Ş.; van der Bank, M.; et al. Preserving the evolutionary potential of floras in biodiversity hotspots. Nature 2007, 445, 757–760. [Google Scholar] [CrossRef]
- Anderson, M.J.; Crist, T.O.; Chase, J.M.; Vellend, M.; Inouye, B.D.; Freestone, A.L.; Sanders, N.J.; Cornell, H.V.; Comita, L.S.; Davies, K.F.; et al. Navigating the multiple meanings of β diversity: A roadmap for the practicing ecologist. Ecol. Lett. 2011, 14, 19–28. [Google Scholar] [CrossRef]
- Myers, J.A.; Chase, J.M.; Jimenez, I.; Jorgensen, P.M.; Araujo-Murakami, A.; Paniagua-Zambrana, N.; Seidel, R. Beta-diversity in temperate and tropical forests reflects dissimilar mechanisms of community assembly. Ecol. Lett. 2013, 16, 151–157. [Google Scholar] [CrossRef]
- Siefert, A.; Ravenscroft, C.; Weiser, M.D.; Swenson, N.G. Functional beta-diversity patterns reveal deterministic community assembly processes in eastern North American trees. Glob. Ecol. Biogeogr. 2013, 22, 682–691. [Google Scholar] [CrossRef]
- Letten, A.D.; Keith, D.A.; Tozer, M.G. Phylogenetic and functional dissimilarity does not increase during temporal heathland succession. Proc. R. Soc. Lond. B Biol. 2014, 281, 20142102. [Google Scholar] [CrossRef]
- Chai, Y.; Yue, M.; Liu, X.; Guo, Y.; Wang, M.; Xu, J.; Zhang, C.; Chen, Y.; Zhang, L.; Zhang, R. Patterns of taxonomic, phylogenetic diversity during a long-term succession of forest on the Loess Plateau, China: Insights into assembly process. Sci. Rep. 2016, 6, 27087. [Google Scholar] [CrossRef]
- Mayfield, M.M.; Levine, J.M. Opposing effects of competitive exclusion on the phylogenetic structure of communities. Ecol. Lett. 2010, 13, 1085–1093. [Google Scholar] [CrossRef] [PubMed]
- Hooper, D.U.; Chapin, F.S.; Ewel, J.J.; Hector, A.; Inchausti, P.; Lavorel, S.; Lawton, J.H.; Lodge, D.M.; Loreau, M.; Naeem, S.; et al. Effects of biodiversity on ecosystem functioning: A consensus of current knowledge. Ecol. Monogr. 2005, 75, 3–35. [Google Scholar] [CrossRef]
- Vivanco, L.; Austin, A.T. Tree species identity alters forest litter decomposition through long-term plant and soil interactions in Patagonia, Argentina. J. Ecol. 2008, 96, 727–736. [Google Scholar] [CrossRef]
- Pennington, R.T.; Lavin, M.; Oliveira-Filho, A. Woody plant diversity, evolution, and ecology in the tropics: Perspectives from seasonally dry tropical forests. Ann. Rev. Ecol. Evol. Syst. 2009, 40, 437–457. [Google Scholar] [CrossRef]
- Chai, Y.; Yue, M.; Wang, M.; Xu, J.; Liu, X.; Zhang, R.; Wan, P. Plant functional traits suggest a change in novel ecological strategies for dominant species in the stages of forest succession. Oecologia 2015, 180, 771–783. [Google Scholar] [CrossRef] [PubMed]
- Yan, Y.; Xian, Y.; Tang, Z. Patterns of species diversity and phylogenetic structure of vascular plants on the Qinghai-Tibetan plateau. Ecol. Evol. 2013, 3, 4584–4595. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Chai, Y.; Wang, M.; Dang, H.; Guo, Y.; Chen, Y.; Zhang, C.; Li, T.; Zhang, L.; Yue, M. Shifts in Plant Community Assembly Processes across Growth Forms along a Habitat Severity Gradient: A Test of the Plant Functional Trait Approach. Front. Plant Sci. 2018, 9, 180. [Google Scholar] [CrossRef]
- Swenson, N.G.; Enquist, B.J.; Thompson, J.; Zimmerman, J.K. The influence of spatial and size scale on phylogenetic relatedness in tropical forest communities. Ecology 2007, 88, 1770–1780. [Google Scholar] [CrossRef]
- Culmsee, H.; Leuschner, C. Consistent patterns of elevational change in tree taxonomic and phylogenetic diversity across Malesian mountain forests. J. Biogeogr. 2013, 40, 1997–2010. [Google Scholar] [CrossRef]
- Gonzálezcaro, S.; Umaña, M.N.; Álvarez, E.; Stevenson, P.R.; Swenson, N.G. Phylogenetic alpha and beta diversity in tropical tree assemblages along regional-scale environmental gradients in northwest south America. J. Plant Ecol. 2014, 7, 145–153. [Google Scholar] [CrossRef]
- Isermann, M. Soil pH and species diversity in coastal dunes. Plant Ecol. 2005, 178, 111–120. [Google Scholar] [CrossRef]
- Michalcová, D.; Gilbert, J.C.; Lawson, C.S.; Gowing, D.J.G.; Marrs, R.H. The combined effect of waterlogging, extractable P and soil pH on alpha-diversity: A case study on mesotrophic grasslands in the UK. Plant Ecol. 2011, 212, 879–888. [Google Scholar] [CrossRef]
- Dinnage, R. Disturbance alters the phylogenetic composition and structure of plant communities in an old field system. PLoS ONE 2009, 4, e7071. [Google Scholar] [CrossRef] [PubMed]
- Helmus, M.R.; Keller, W.B.; Paterson, M.J.; Yan, N.D.; Cannon, C.H.; Rusak, J.A. Communities contain closely related species during ecosystem disturbance. Ecol. Lett. 2010, 13, 162–174. [Google Scholar] [CrossRef] [PubMed]
| Plot | SWC (%) | pH | RAP (mg/kg) | NN (mg/kg) | AN (mg/kg) |
|---|---|---|---|---|---|
| QL-1 | 22.017 | 6.215 | 26.530 | 12.350 | 7.570 |
| QL-2 | 21.000 | 6.330 | 5.110 | 6.930 | 5.550 |
| QL-3 | 27.500 | 5.858 | 9.620 | 3.690 | 11.150 |
| QL-4 | 23.600 | 6.134 | 13.753 | 7.657 | 8.090 |
| QL-5 | 30.500 | 5.920 | 13.720 | 2.540 | 14.690 |
| QL-6 | 20.000 | 5.534 | 8.360 | 2.640 | 18.170 |
| QL-7 | 20.200 | 5.359 | 5.550 | 1.950 | 27.600 |
| QL-8 | 23.567 | 5.604 | 9.210 | 2.377 | 20.153 |
| QL-9 | 38.000 | 6.085 | 8.290 | 2.930 | 23.250 |
| QL-10 | 31.100 | 5.163 | 8.410 | 4.890 | 29.710 |
| QL-11 | 22.600 | 6.090 | 6.840 | 10.710 | 10.320 |
| QL-12 | 30.567 | 5.779 | 7.847 | 6.177 | 21.093 |
| QL-13 | 20.200 | 5.410 | 4.960 | 3.950 | 11.220 |
| QL-14 | 22.000 | 5.515 | 7.040 | 2.510 | 9.510 |
| QL-15 | 18.500 | 5.050 | 5.400 | 1.840 | 14.190 |
| QL-16 | 20.233 | 5.325 | 5.800 | 2.767 | 11.640 |
| QL-17 | 15.700 | 5.394 | 4.600 | 2.030 | 9.060 |
| QL-18 | 14.700 | 4.572 | 3.860 | 2.010 | 11.700 |
| QL-19 | 16.300 | 5.186 | 2.310 | 1.590 | 7.460 |
| QL-20 | 15.567 | 5.051 | 3.590 | 1.877 | 9.407 |
| LP-1 | 18.434 | 7.990 | 11.670 | 15.780 | 12.760 |
| LP-2 | 17.088 | 8.300 | 19.260 | 21.590 | 7.050 |
| LP-3 | 20.594 | 8.115 | 12.760 | 19.590 | 6.580 |
| LP-4 | 16.802 | 7.960 | 6.080 | 12.980 | 3.710 |
| LP-5 | 16.630 | 8.330 | 3.100 | 9.990 | 3.560 |
| LP-6 | 17.788 | 8.370 | 3.490 | 17.700 | 6.110 |
| LP-7 | 18.060 | 8.330 | 2.540 | 8.660 | 4.810 |
| LP-8 | 17.449 | 8.677 | 2.880 | 7.160 | 3.160 |
| LP-9 | 17.545 | 8.397 | 4.160 | 4.800 | 14.190 |
| LP-10 | 17.751 | 8.325 | 5.940 | 4.590 | 8.980 |
| LP-11 | 16.259 | 8.238 | 4.890 | 3.570 | 6.640 |
| LP-12 | 18.594 | 8.427 | 15.570 | 5.500 | 9.450 |
| LP-13 | 17.753 | 8.638 | 6.830 | 13.710 | 6.960 |
| LP-14 | 17.954 | 8.550 | 7.430 | 13.250 | 5.340 |
| LP-15 | 15.891 | 8.628 | 9.080 | 27.970 | 5.910 |
| LP-16 | 20.306 | 8.420 | 42.630 | 19.060 | 4.530 |
| LP-17 | 18.500 | 8.495 | 14.930 | 10.373 | 6.710 |
| LP-18 | 18.500 | 8.495 | 4.803 | 18.880 | 5.050 |
| LP-19 | 16.600 | 8.544 | 5.807 | 8.330 | 5.117 |
| LP-20 | 14.000 | 8.695 | 4.430 | 5.507 | 5.040 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, M.; Xu, J.; Chai, Y.; Guo, Y.; Liu, X.; Yue, M. Differentiation of Environmental Conditions Promotes Variation of Two Quercus wutaishanica Community Assembly Patterns. Forests 2020, 11, 43. https://doi.org/10.3390/f11010043
Wang M, Xu J, Chai Y, Guo Y, Liu X, Yue M. Differentiation of Environmental Conditions Promotes Variation of Two Quercus wutaishanica Community Assembly Patterns. Forests. 2020; 11(1):43. https://doi.org/10.3390/f11010043
Chicago/Turabian StyleWang, Mao, Jinshi Xu, Yongfu Chai, Yaoxin Guo, Xiao Liu, and Ming Yue. 2020. "Differentiation of Environmental Conditions Promotes Variation of Two Quercus wutaishanica Community Assembly Patterns" Forests 11, no. 1: 43. https://doi.org/10.3390/f11010043
APA StyleWang, M., Xu, J., Chai, Y., Guo, Y., Liu, X., & Yue, M. (2020). Differentiation of Environmental Conditions Promotes Variation of Two Quercus wutaishanica Community Assembly Patterns. Forests, 11(1), 43. https://doi.org/10.3390/f11010043




