Affinity Propagation Clustering Using Path Based Similarity
Abstract
:1. Introduction
2. Method
2.1. Affinity Propagation Clustering
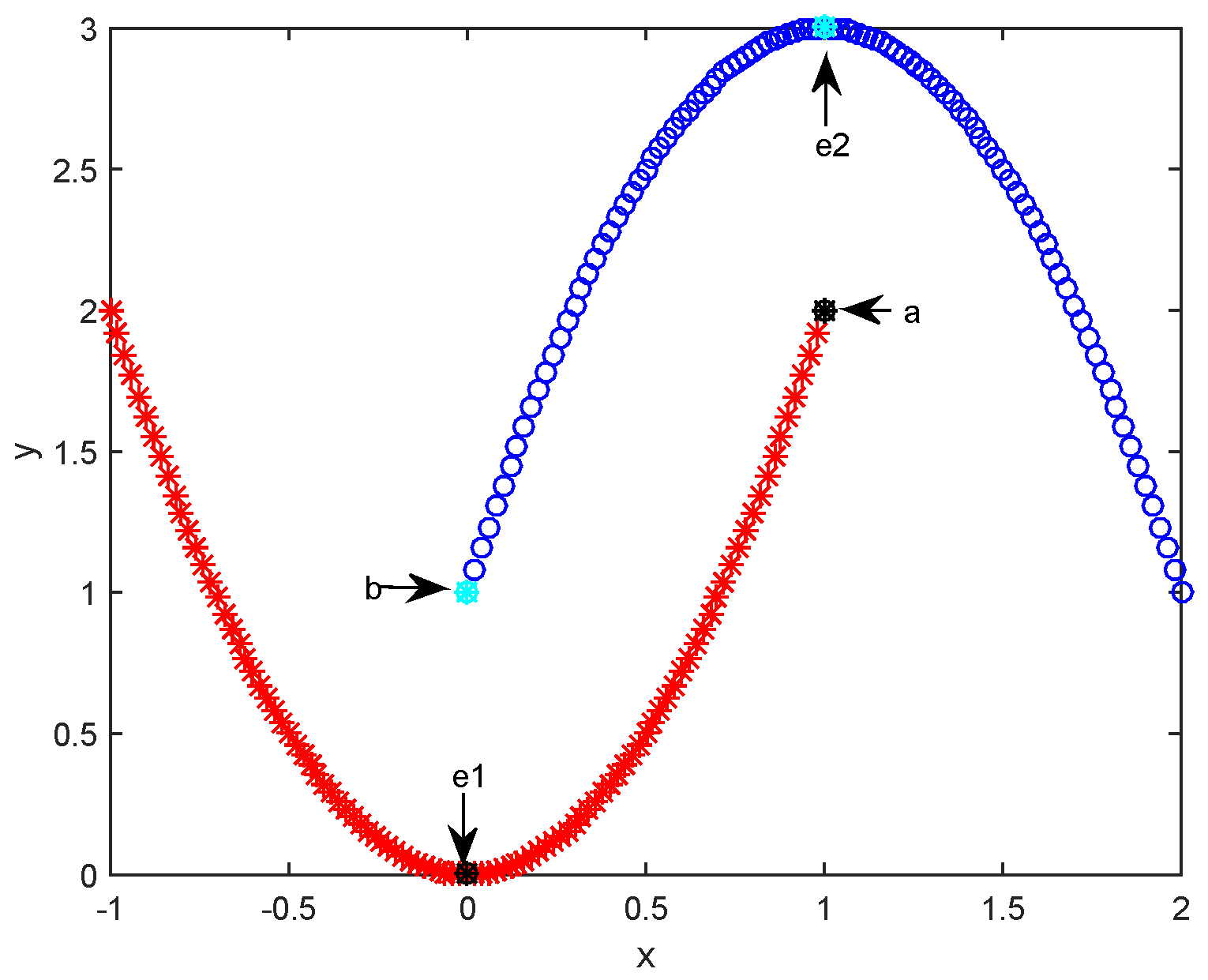
2.2. Affinity Propagation Clustering Using Path-Based Similarity
| Algorithm 1: APC-PS: APC Based on Path-Based Similarity |
| Input: |
| : negative Euclidean distance based similarity between n samples |
| : preference value of a sample chosen as the cluster center (or exemplar) |
| λ: damping factor for APC |
| k: number of clusters |
| Output: |
| : the cluster membership for sample |
| 1: Initialize and set each diagonal entry of as p. |
| 2: Iteratively update , and using Equations (5)–(7), respectively. |
| 3: Identify exemplars based on the summation of and . |
| 4: If the number of exemplars is larger (or smaller) than k, then reduce (or increase) p and goto line 1. |
| 5: Compute the path-based similarity matrix using Equations (8) and (9). |
| 6: Assign non-exemplar sample i to the closest cluster (i.e., e) based on and set . |
3. Results
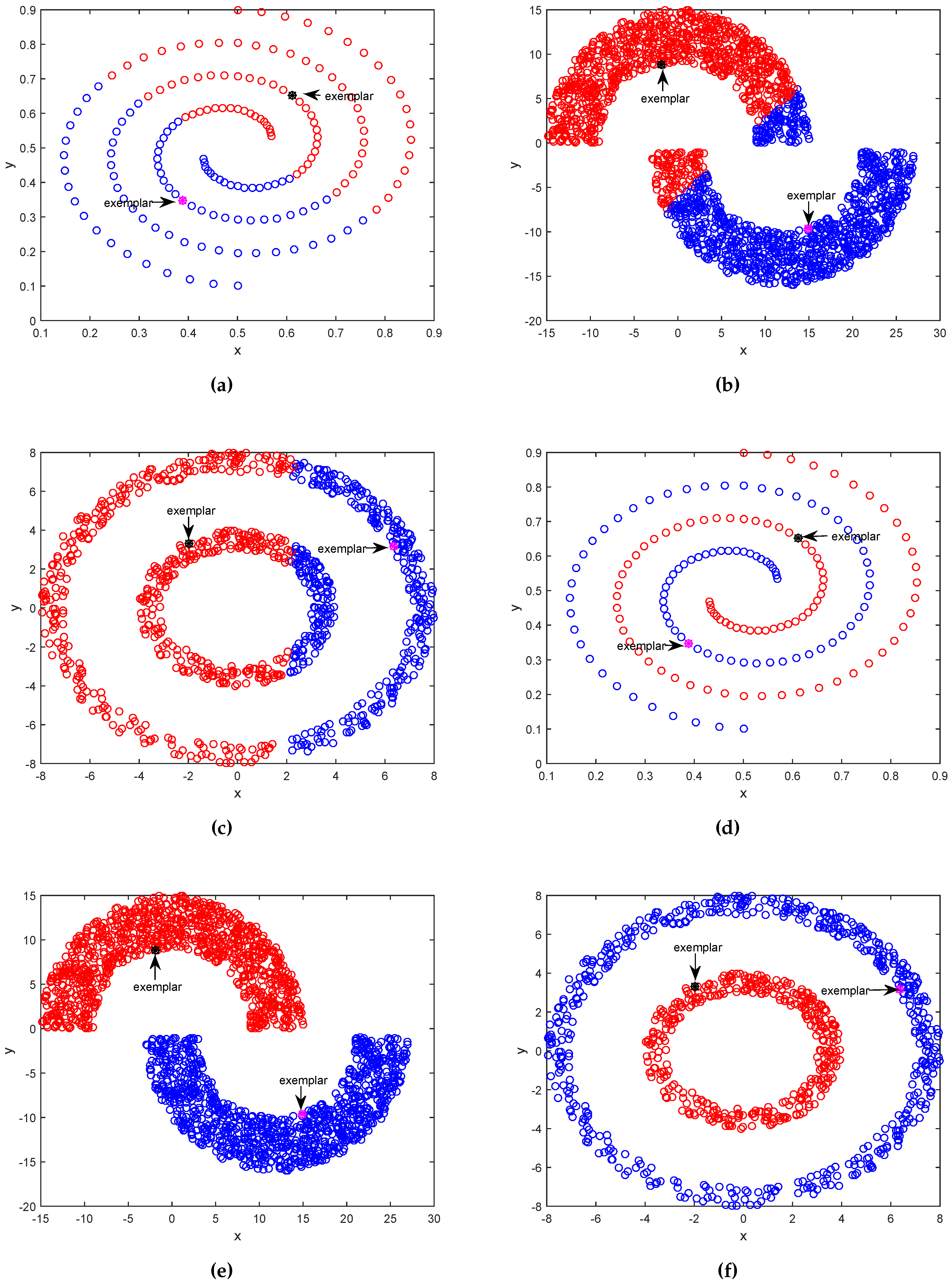
3.1. Results on Synthetic Datasets
3.2. Results on UCI Datasets
3.2.1. Clustering results on UCI datasets
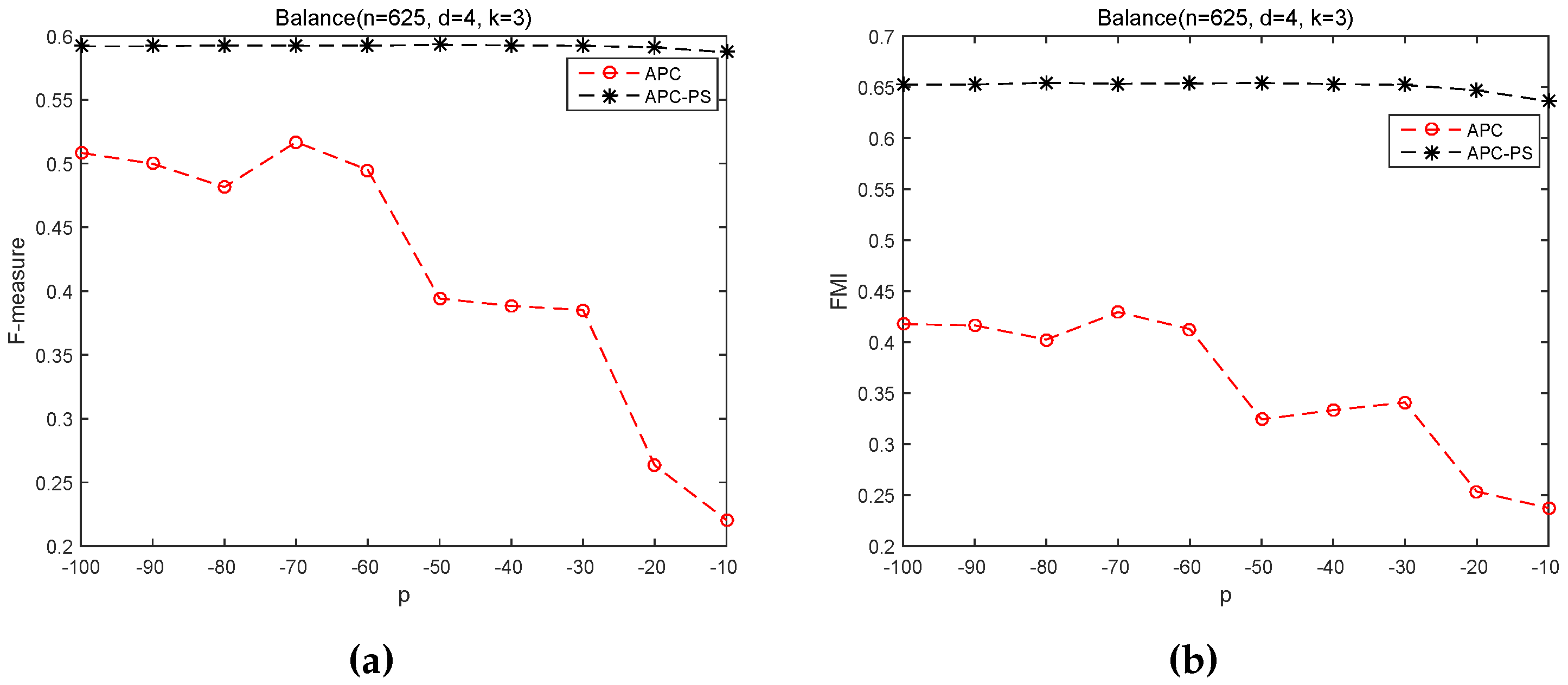
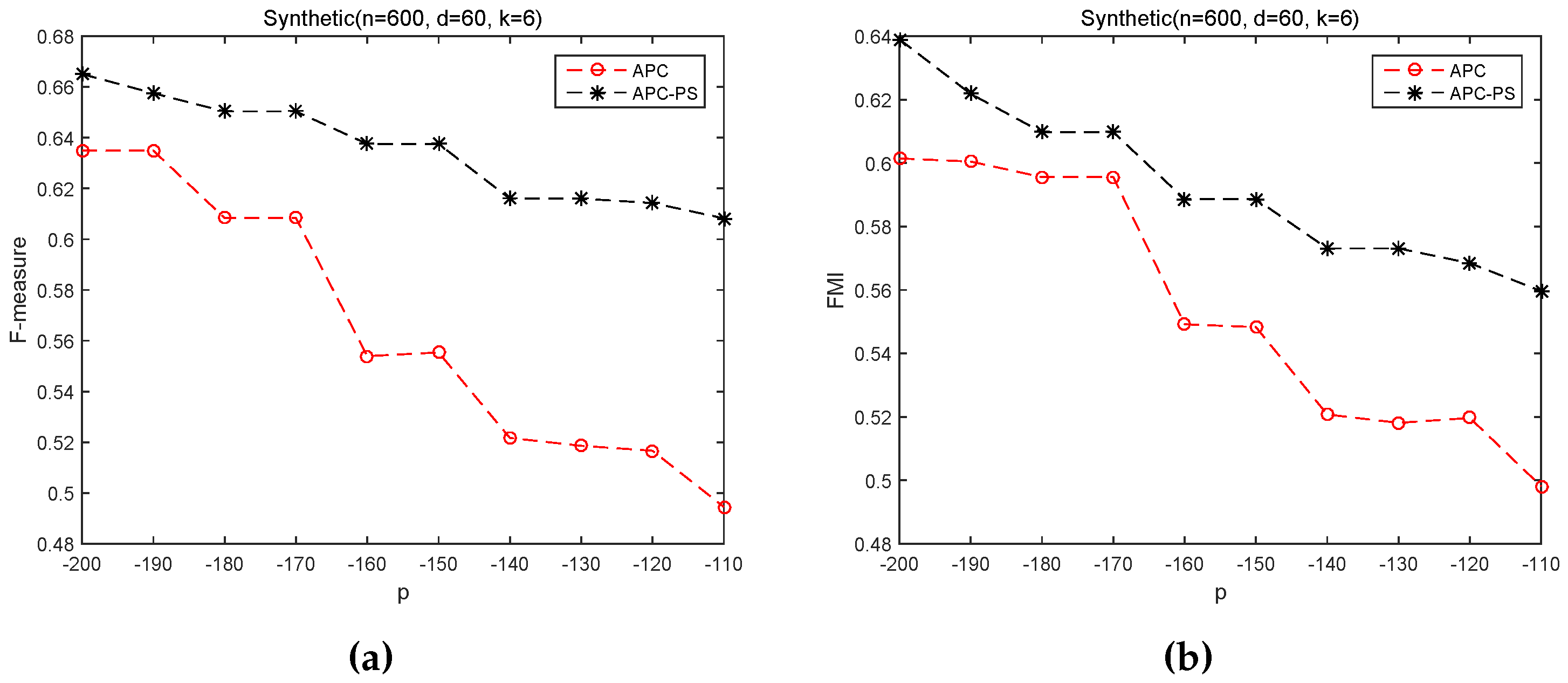
3.2.2. Sensitivity of p on APC and APC-PS
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Napolitano, F.; Raiconi, G.; Tagliaferri, R.; Ciaramella, A.; Staiano, A.; Miele, G. Clustering and visualization approaches for human cell cycle gene expression data analysis. Int. J. Approx. Reason. 2008, 47, 70–84. [Google Scholar] [CrossRef]
- Peng, W.F.; Du, S.; Li, F.X. Unsupervised image segmentation via affinity propagation. Appl. Mech. Mater. 2014, 610, 464–470. [Google Scholar] [CrossRef]
- Kang, J.H.; Lerman, K.; Plangprasopchok, A. Analyzing microblogs with affinity propagation. In Proceedings of the First Workshop on Social Media Analytics (SOMA ‘10), Washington, DC, USA, 25 July 2010; pp. 67–70.
- Hong, L.; Cai, S.M.; Fu, Z.Q.; Zhou, P.L. Community identification of financial market based on affinity propagation. In Recent Progress in Data Engineering and Internet Technology; Springer: Berlin, Germany, 2013; pp. 121–127. [Google Scholar]
- Papalexakis, E.E.; Beutel, A.; Steenkiste, P. Network anomaly detection using co-clustering. In Encyclopedia of Social Network Analysis and Mining; Springer: New York, NY, USA, 2014; pp. 1054–1068. [Google Scholar]
- Ester, M.; Kriegel, H.P.; Sander, J.; Xu, X. A density-based algorithm for discovering clusters in large spatial databases with noise. In Proceedings of the 2nd International Conference on Knowledge Discovery and Data Mining, Portland, OR, USA, 2–4 August 1996; Volume 96, pp. 226–231.
- Rodriguez, A.; Laio, A. Clustering by fast search and find of density peaks. Science 2014, 344, 1492–1496. [Google Scholar] [CrossRef] [PubMed]
- Frey, B.J.; Dueck, D. Clustering by passing messages between data points. Science 2007, 315, 972–976. [Google Scholar] [CrossRef] [PubMed]
- Frey, B.J.; Dueck, D. Response to comment on “Clustering by passing messages between data points”. Science 2008, 319, 726. [Google Scholar] [CrossRef]
- Zhang, R. Two similarity measure methods based on human vision properties for image segmentation based on affinity propagation clustering. In Proceedings of the International Conference on Measuring Technology and Mechatronics Automation (ICMTMA), Changsha, China, 13–14 March 2010; Volume 3, pp. 1054–1058.
- Du, H.; Wang, Y.P.; Duan, L.L. A new method for grayscale image segmentation based on affinity propagation clustering algorithm. In Proceedings of the IEEE 9th International Conference on Computational Intelligence and Security, Leshan, China, 14–15 Decebmer 2013; pp. 170–173.
- Leone, M.; Sumedha; Weigt, M. Clustering by soft-constraint affinity propagation: Applications to gene-expression data. Bioinformatics 2007, 23, 2708–2015. [Google Scholar] [CrossRef] [PubMed]
- Zhao, C.W.; Peng, Q.K.; Zhao, C.W.; Sun, S.H. Chinese text automatic summarization based on affinity propagation cluster. In Proceedings of the International Conference on Fuzzy Systems and Knowledge Discovery, Tianjin, China, 14–16 August 2009; Volume 1, pp. 425–429.
- Xiao, Y.; Yu, J. Semi-supervised clustering based on affinity propagation algorithm. J. Softw. 2008, 19, 2803–2813. [Google Scholar] [CrossRef]
- Wagstaff, K.; Cardie, C.; Rogers, S.; Schrödl, S. Constrained k-means clustering with background knowledge. In Proceedings of the 18th International Conference on Machine Learning, Williamstown, MA, USA, 28 June–1 July 2001; pp. 577–584.
- Wang, K.J.; Zhang, J.Y.; Li, D.; Zhang, X.N.; Guo, T. Adaptive affinity propagation clustering. Acta Autom. Sin. 2007, 33, 1242–1246. [Google Scholar]
- Xia, D.Y.; Fei, W.U.; Zhang, X.Q.; Zhuang, Y.T. Local and global approaches of affinity propagation clustering for large scale data. J. Zhejiang Univ. Sci. A 2008, 9, 1373–1381. [Google Scholar] [CrossRef]
- Serdah, A.M.; Ashour, W.M. Clustering large-scale data based on modified affinity propagation algorithm. J. Artif. Intell. Soft Comput. Res. 2016, 6, 23–33. [Google Scholar] [CrossRef]
- Zhang, X.L.; Wang, W.; Norvag, K.; Sebag, M. K-AP: Generating specified K clusters by efficient affinity propagation. In Proceedings of the IEEE Tenth International Conference on Data Mining (ICDM), Sydney, Australia, 13–17 December 2010; pp. 1187–1192.
- Barbakh, W.; Fyfe, C. Inverse weighted clustering algorithm. Comput. Inf. Syst. 2007, 11, 10–18. [Google Scholar]
- Walter, S.F. Clustering by Affinity Propagation. Ph.D. Thesis, ETH Zurich, Zürich, Switzerland, 2007. [Google Scholar]
- Zhang, L.; Du, Z. Affinity propagation clustering with geodesic distances. J. Computat. Inf. Syst. 2010, 6, 47–53. [Google Scholar]
- Guo, K.; Guo, W.; Chen, Y.; Qiu, Q.; Zhang, Q. Community discovery by propagating local and global information based on the MapReduce model. Inf. Sci. 2015, 323, 73–93. [Google Scholar] [CrossRef]
- Meo, P.D.; Ferrara, E.; Fiumara, G.; Ricciardello, A. A novel measure of edge centrality in social networks. Knowl.-Based Syst. 2012, 30, 136–150. [Google Scholar] [CrossRef]
- Lichman, M. UCI Machine Learning Repository. 2013. Available online: http://www.ics.uci.edu/ml (accessed on 21 July 2016).
- Jain, A.K. Data clustering: 50 years beyond k-means. Pattern Recognit. Lett. 2010, 31, 651–666. [Google Scholar] [CrossRef]
- Floyd, R.W. Algorithm 97: Shortest path. Commun. ACM 1962, 5, 345. [Google Scholar] [CrossRef]
- MacQueen, J. Some methods for classification and analysis of multivariate observations. In Proceedings of the 5th Berkeley Symposium on Mathematical Statistics and Probability, Oakland, CA, USA, 21 June–18 July 1965 and 27 December 1965–7 January 1966; Volume 1, pp. 281–297.
- Dempster, A.P.; Laird, N.M.; Rubin, D.B. Maximum likelihood from incomplete data via the EM algorithm. J. R. Stat.Soc. Ser. B (Methodol.) 1977, 39, 1–38. [Google Scholar]
- Bradley, P.S.; Fayyad, U.; Reina, C. Scaling EM (eXpectation-Maximization) Clustering to Large Databases; Technical Report, MSR-TR-98-35; Microsoft Research Redmond: Redmond, WA, USA, 1998. [Google Scholar]
- Elhamifar, E.; Vidal, R. Sparse subspace clustering: algorithm, theory, and applications. IEEE Trans. Pattern Anal. Mach. Intell. 2013, 35, 2765–2781. [Google Scholar] [CrossRef] [PubMed]
- Larsen, B.; Aone, C. Fast and effective text mining using linear-time document clustering. In Proceedings of the 5th ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, San Diego, CA, USA, 15–18 August 1999; pp. 16–22.
- Dalli, A. Adaptation of the F-measure to cluster based lexicon quality evaluation. In Proceedings of the EACL 2003 Workshop on Evaluation Initiatives in Natural Language Processing. Association for Computational Linguistics, Budapest, Hungary, 13 April 2003; pp. 51–56.
- Halkidi, M.; Batistakis, Y.; Vazirgiannis, M. On clustering validation techniques. J. Intell. Inf. Syst. 2001, 17, 107–145. [Google Scholar] [CrossRef]
| Dataset | n | d | k |
|---|---|---|---|
| Balance | 625 | 4 | 3 |
| Glass | 214 | 9 | 6 |
| Synthetic | 600 | 60 | 6 |
| Mushrooms | 8124 | 112 | 2 |
| Method | Balance Scale | Synthetic Control | Glass Identification | Mushrooms |
|---|---|---|---|---|
| EM | 0.5718 ± 0.0020 | 0.5950 ± 0.0531 | 0.5096 ± 0.0291 | 0.6798 ± 0.1030 |
| k-means | 0.5652 ± 0.0328 | 0.6662 ± 0.0240 | 0.5364 ± 0.0234 | 0.7740 ± 0.1368 |
| SSC | 0.5728 ± 0.0000 | 0.5926 ± 0.0000 | 0.4840 ± 0.0012 | 0.7934 ± 0.0000 |
| pAPC | 0.5638 ± 0.0987 | 0.6477 ± 0.0000 | 0.4971 ± 0.0000 | —————— |
| APCGD | 0.5829 ± 0.1021 | 0.4192 ± 0.0000 | 0.4248 ± 0.0297 | 0.5781 ± 0.0000 |
| APC | 0.5518 ± 0.0563 | 0.6350 ± 0.0000 | 0.5598 ± 0.0000 | 0.7616 ± 0.0000 |
| APC-PS | 0.5925 ± 0.0018 | 0.6650 ± 0.0000 | 0.5620 ± 0.0000 | 0.7634 ± 0.0000 |
| Method | Balance Scale | Synthetic Control | Glass Identification | Mushrooms |
|---|---|---|---|---|
| EM | 0.4708 ± 0.0130 | 0.5581 ± 0.0545 | 0.4516 ± 0.0397 | 0.6086 ± 0.0828 |
| k-means | 0.4567 ± 0.0241 | 0.6277 ± 0.0251 | 0.4775 ± 0.0464 | 0.7232 ± 0.1106 |
| SSC | 0.4675 ± 0.0000 | 0.5664 ± 0.0000 | 0.3778 ± 0.0005 | 0.6732 ± 0.0000 |
| pAPC | 0.5076 ± 0.1039 | 0.5664 ± 0.0000 | 0.3895 ± 0.0000 | —————— |
| APCGD | 0.5173 ± 0.0872 | 0.4750 ± 0.0000 | 0.4912 ± 0.0239 | 0.5198 ± 0.0000 |
| APC | 0.4294 ± 0.0449 | 0.6015 ± 0.0000 | 0.5606 ± 0.0000 | 0.7105 ± 0.0000 |
| APC-PS | 0.6539 ± 0.0046 | 0.6390 ± 0.0000 | 0.5477 ± 0.0000 | 0.7277 ± 0.0000 |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jiang, Y.; Liao, Y.; Yu, G. Affinity Propagation Clustering Using Path Based Similarity. Algorithms 2016, 9, 46. https://doi.org/10.3390/a9030046
Jiang Y, Liao Y, Yu G. Affinity Propagation Clustering Using Path Based Similarity. Algorithms. 2016; 9(3):46. https://doi.org/10.3390/a9030046
Chicago/Turabian StyleJiang, Yuan, Yuliang Liao, and Guoxian Yu. 2016. "Affinity Propagation Clustering Using Path Based Similarity" Algorithms 9, no. 3: 46. https://doi.org/10.3390/a9030046
APA StyleJiang, Y., Liao, Y., & Yu, G. (2016). Affinity Propagation Clustering Using Path Based Similarity. Algorithms, 9(3), 46. https://doi.org/10.3390/a9030046






