The Synthesis and Evaluation of Aminocoumarin Peptidomimetics as Cytotoxic Agents on Model Bacterial E. coli Strains
Abstract
:1. Introduction
2. Materials and Methods
3. Experimental Section
3.1. Synthesis of p-Nitrophenylhydrogenglutarate
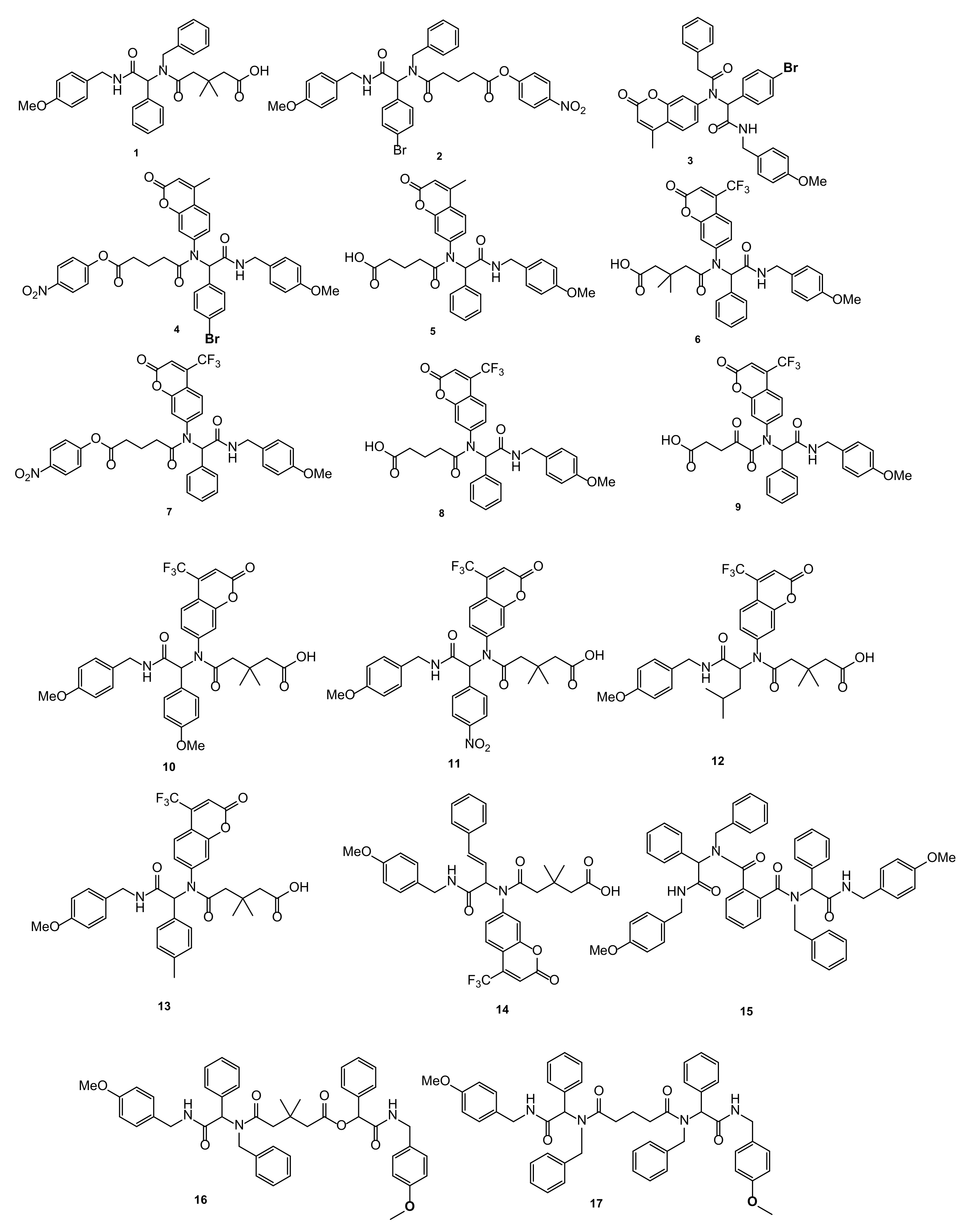
3.2. General Procedure for Synthesis of Compounds 1–17
3.2.1. (2-[(4-Methoxyphenyl)benzyl]amino-2-oxo-1-phenylethyl)-3,3-methyl-5-oxopentanoic acid (1)
3.2.2. (2-[(4-Methoxphenyl)-4-bromobenzyl]amino-2-oxo-1-phenylethyl)-5-(4-nitrophenoxy)-5-oxopentanoate (2)
3.2.3. (2-[(4-Methoxyphenyl)-4-bromobenzyl]amino-5-[(4-methyl-2-oxo-2H-1-benzopyran-7-yl)amino]-phenylacetamide (3)
3.2.4. (2-[(4-Methoxyphenyl)benzyl]amino-5-[(4-methyl-2-oxo-2H-1-benzopyran-7-yl)amino]-5-(4-nitrophenoxy)-5-oxopentanoate (4)
3.2.5. (2-[(4-Methoxyphenyl)benzyl]amino-5-[(4-methyl-2-oxo-2H-1-benzopyran-7-yl)amino]-5-oxopentanoic acid (5)
3.2.6. [5-((2-((4-Methoxybenzyl)amino)-2-oxo-1-phenylethyl)(2-oxo-4-(trifluoromethyl)-2H-chromen-7-yl)amino)-3,3-dimethyl-5-oxopentanoic acid] (6)
3.2.7. [4-Nitrophenyl-5-((2-((4-methoxybenzyl)amino)-2-oxo-1-phenylethyl)(2-oxo-4-(trifluoromethyl)-2H-chromen-7-yl)amino)-5-oxopentanoate] (7)
3.2.8. [5-((2-((4-Methoxybenzyl)amino)-2-oxo-1-phenylethyl)(2-oxo-4-(trifluoromethyl)-2H-chromen-7-yl)amino)-5-oxopentanoic acid] (8)
3.2.9. [5-((2-((4-Methoxybenzyl)amino)-2-oxo-1-phenylethyl)(2-oxo-4-(trifluoromethyl)-2H-chromen-7-yl)amino)-4,5-dioxopentanoic acid] (9)
3.2.10. [5-((2-((4-Methoxybenzyl)amino)-1-(4-methoxyphenyl)-2-oxoethyl)(2-oxo-4-(trifluoromethyl)-2H-chromen-7-yl)amino)-3,3-dimethyl-5-oxopentanoic acid] (10)
3.2.11. [5-((2-((4-Methoxybenzyl)amino)-1-(4-nitrophenyl)-2-oxoethyl)(2-oxo-4-(trifluoromethyl)-2H-chromen-7-yl)amino)-3,3-dimethyl-5-oxopentanoic acid] (11)
3.2.12. [5-((1-((4-Methoxybenzyl)amino)-4-methyl-1-oxopentan-2-yl)(2-oxo-4-(trifluoromethyl)-2H-chromen-7-yl)amino)-3,3-dimethyl-5-oxopentanoic acid] (12)
3.2.13. [5-((2-((4-Methoxybenzyl)amino)-2-oxo-1-(p-tolyl)ethyl)(2-oxo-4-(trifluoromethyl)-2H-chromen-7-yl)amino)-3,3-dimethyl-5-oxopentanoic acid] (13)
3.2.14. [(E)-5-((1-((4-Methoxybenzyl)amino)-1-oxo-4-phenylbut-3-en-2-yl)(2-oxo-4-(trifluoromethyl)-2H-chromen-7-yl)amino)-3,3-dimethyl-5-oxopentanoic acid] (14)
3.2.15. N1,N2-Dibenzyl-N1,N2-bis(2-((4-methoxybenzyl)amino)-2-oxo-1-phenylethyl)phthalamide (15)
3.2.16. {2-((4-Methoxybenzyl)amino)-2-oxo-1-phenylethyl-5-(benzyl(2-((4-methoxybenzyl)amino)-2-oxo-1-phenylethyl)amino)-3,3-dimethyl-5-oxopentanoate} (16)
3.2.17. {N1,N5-Dibenzyl-N1,N5-bis(2-((4-methoxybenzyl)amino)-2-oxo-1-phenylethyl)glutaramide} (17)
4. Microorganisms and Media
5. Results and Discussion
5.1. Chemistry
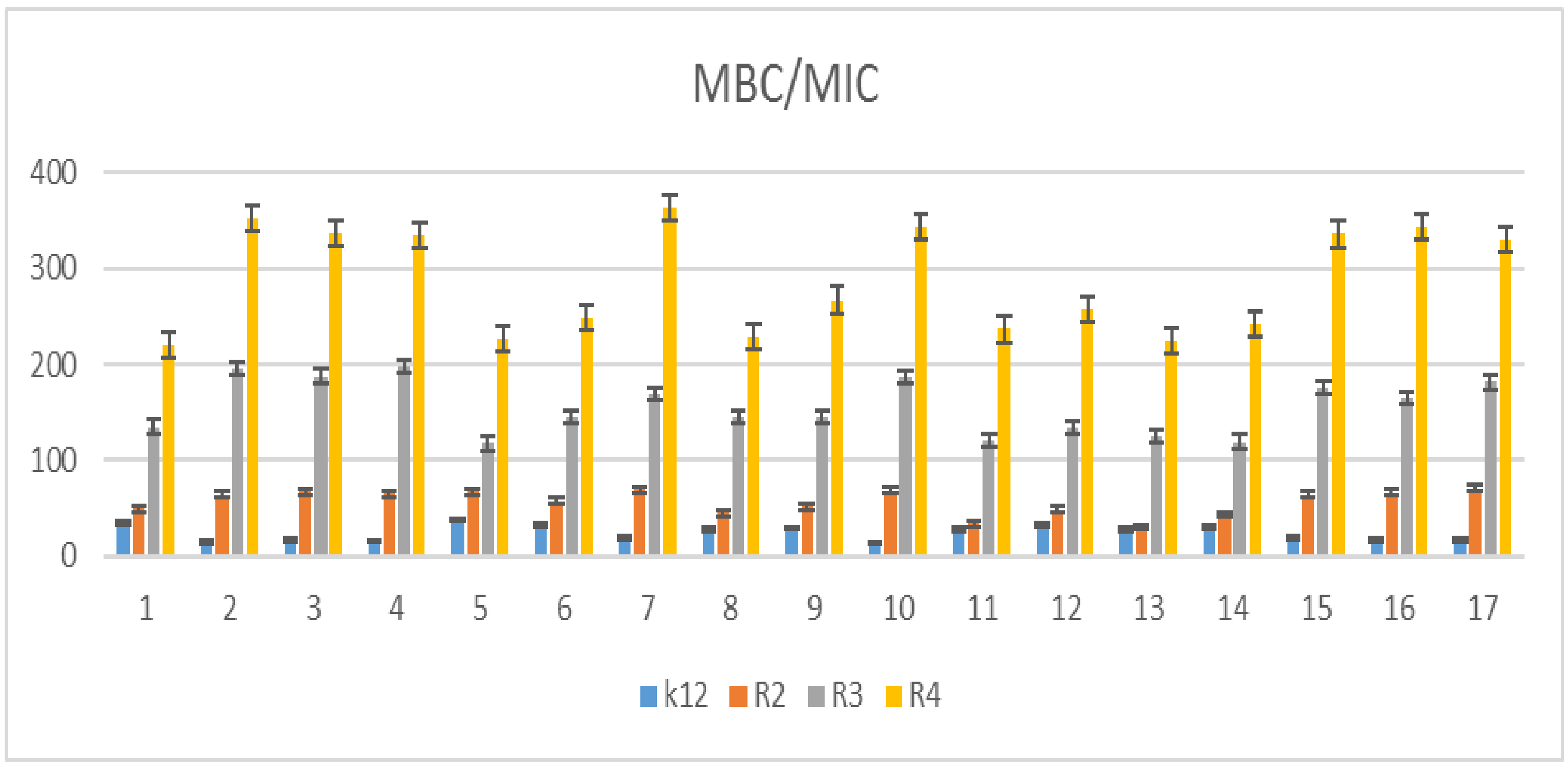
5.2. Cytotoxic Studies of the Library of Peptidomimetics
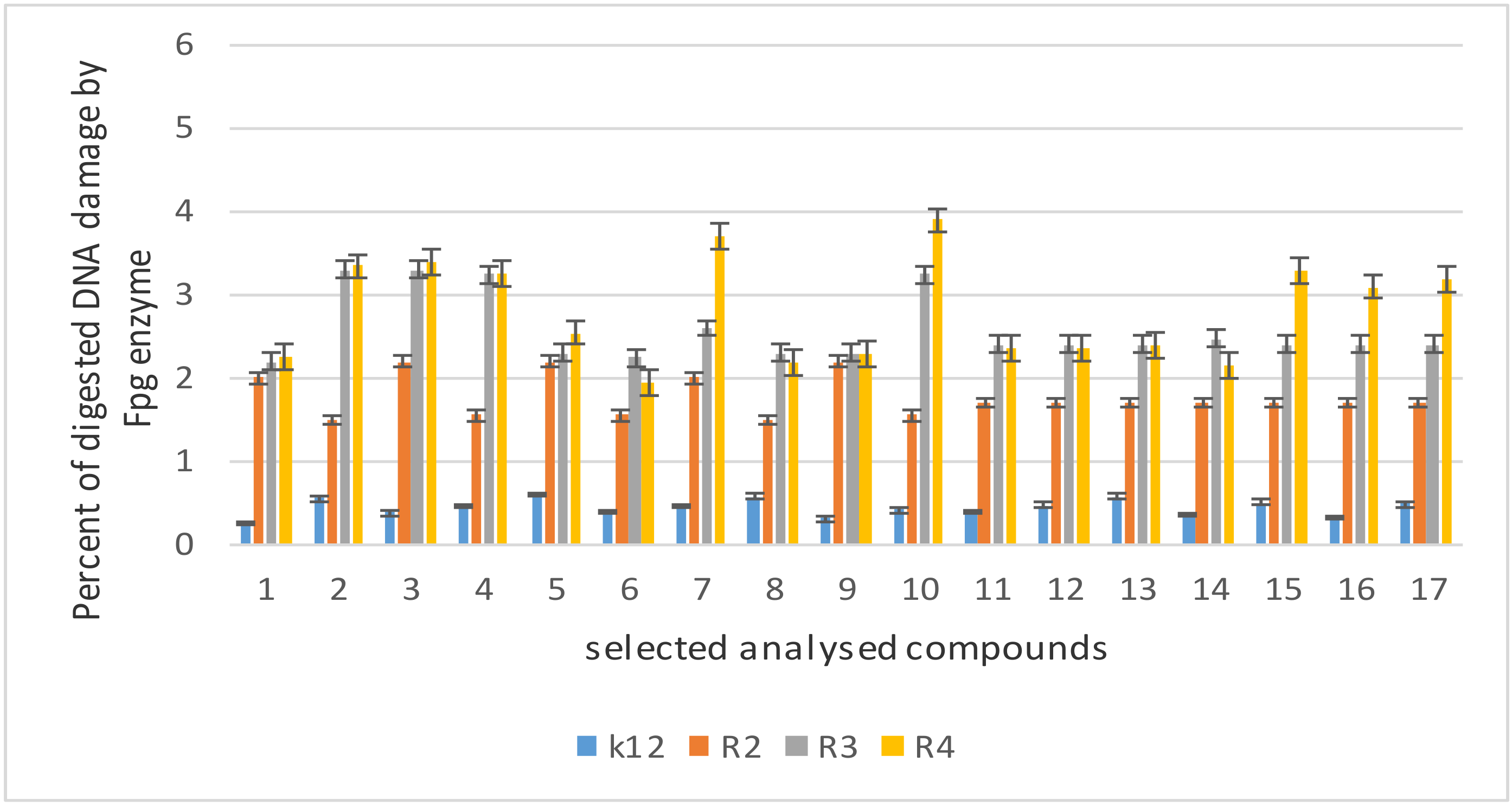
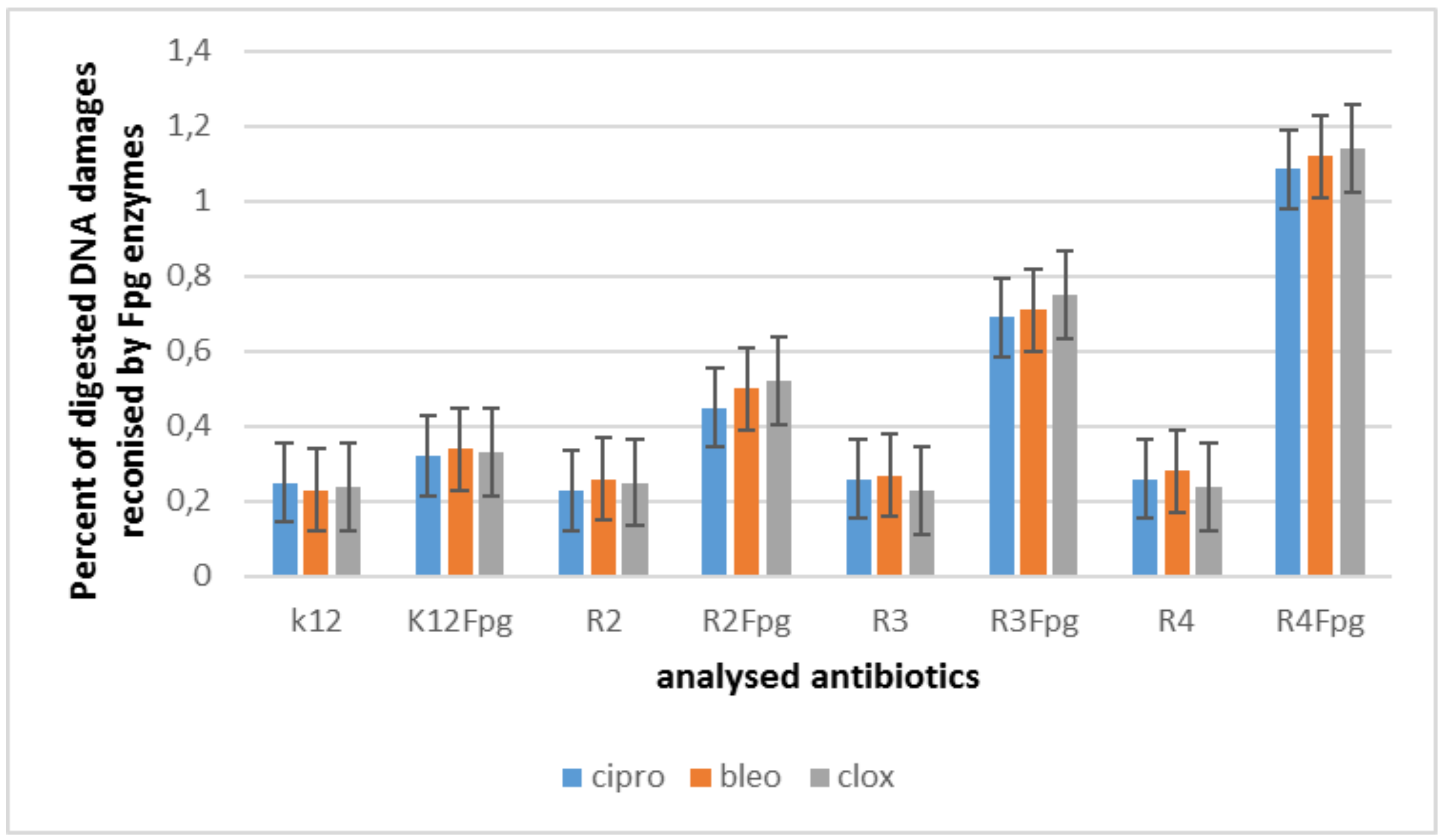
5.3. Modification of Bacterial DNA Isolated from E. coli R2–R4 Strains with Tested Coumarin Derivatives
6. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| MIC | minimum inhibitory concentration |
| MBC | minimum bactericidal concentration |
| Oc | open circle |
| Ccc | covalently closed circle |
References
- Dömling, A.; Ugi, I. Multicomponent Reactions with Isocyanides. Angew. Chem. Int. Ed. 2000, 39, 3168–3210. [Google Scholar] [CrossRef]
- Das, K.; Jain, B.; Gupta, P. Photophysics of Coumarin 500 and Coumarin 151 in AOT reverse micelles. Chem. Phys. Lett. 2005, 410, 160–164. [Google Scholar] [CrossRef]
- Kumar, S.; Mukesh, K.; Harjai, K.; Singh, V. Synthesis of coumarin based Knoevenagel-Ugi adducts by a sequentialone pot five-component reaction and their biological evaluation asanti-bacterial agents. Tetrahedron Lett. 2019, 60, 8–12. [Google Scholar] [CrossRef]
- Marquarding, D.; Gokel, G.; Hoffmann, P.; Ugi, I. Chapter 7—The Passerini Reaction and Related Reactions. Org. React. 1971, 20, 133. [Google Scholar] [CrossRef]
- Bos, M.; Riguet, E. Synthesis of Chiral γ-Lactones by One-Pot Sequential Enantioselective Organocatalytic Michael Addition of Boronic Acids and Diastereoselective Intramolecular Passerini Reaction. J. Org. Chem. 2014, 79, 10881–10889. [Google Scholar] [CrossRef]
- Bossio, R.; Marcaccini, S.; Pepino, R.; Torroba, T. Synthesis Studies on Isocyanides and Related Compounds: A Novel Syn-thetic Route to Furan Derivatives. Heterocycles 1993, 1983, 783. [Google Scholar] [CrossRef]
- Szymanski, W.; Ostaszewski, R. Toward stereocontrolled, chemoenzymatic synthesis of unnatural peptides. Tetrahedron 2008, 64, 3197–3203. [Google Scholar] [CrossRef]
- Szymanski, W.; Zwolinska, M.; Ostaszewski, R. Studies on the application of the Passerini reaction and enzymatic procedures to the synthesis of tripeptide mimetics. Tetrahedron 2007, 63, 7647–7653. [Google Scholar] [CrossRef]
- Szymanski, W.; Ostaszewski, R. Multicomponent diversity and enzymatic enantioselectivity as a route towards both enantiomers of α-amino acids—A model study. Tetrahedron Asymmetry 2006, 17, 2667–2671. [Google Scholar] [CrossRef]
- Devji, T.; Reddy, C.; Woo, C.; Awale, S.; Kadota, S.; Carrico-Moniz, D. Pancreatic anticancer activity of a novel geranylgeranylated coumarin derivative. Bioorganic Med. Chem. Lett. 2011, 21, 5770–5773. [Google Scholar] [CrossRef]
- Reddy, N.S.; Mallireddigari, M.R.; Cosenza, S.; Gumireddy, K.; Bell, S.C.; Reddy, E.P.; Reddy, M.R. Synthesis of new coumarin 3-(N-aryl) sulfonamides and their anticancer activity. Bioorganic Med. Chem. Lett. 2004, 14, 4093–4097. [Google Scholar] [CrossRef]
- Hadjipavlou-Litina, D.J.; Litinas, K.E.; Kontogiorgis, C. The Anti-inflammatory Effect of Coumarin and its Derivatives. Anti-Inflamm. Anti-Allergy Agents Med. Chem. 2007, 6, 293–306. [Google Scholar] [CrossRef]
- Xue, H.; Lu, X.; Zheng, P.; Liu, L.; Han, C.; Hu, J.; Liu, Z.; Ma, T.; Li, Y.; Wang, L.; et al. Highly Suppressing Wild-Type HIV-1 and Y181C Mutant HIV-1 Strains by 10-Chloromethyl-11-demethyl-12-oxo-calanolide A with Druggable Profile. J. Med. Chem. 2010, 53, 1397–1401. [Google Scholar] [CrossRef]
- Curini, M.; Epifano, F.; Maltese, F.; Marcotullio, M.C.; Gonzales, S.P.; Rodriguez, J.C. Synthesis of Collinin, an Antiviral Coumarin. Aust. J. Chem. 2003, 56, 59–60. [Google Scholar] [CrossRef]
- Hwang, C.H.; Jaki, B.U.; Klein, L.L.; Lankin, D.C.; McAlpine, J.B.; Napolitano, J.G.; Fryling, N.A.; Franzblau, S.G.; Cho, S.H.; Stamets, P.E.; et al. Chlorinated Coumarins from the Polypore Mushroom Fomitopsis officinalis and Their Activity against Mycobacterium tuberculosis. J. Nat. Prod. 2013, 76, 1916–1922. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Arshad, A.; Osman, H.; Bagley, M.C.; Lam, C.K.; Mohamad, S.; Zahariluddin, A.S.M. Synthesis and antimicrobial properties of some new thiazolyl coumarin derivatives. Eur. J. Med. Chem. 2011, 46, 3788–3794. [Google Scholar] [CrossRef] [PubMed]
- Laurin, P.; Ferroud, D.; Klich, M.; Dupuis-Hamelin, C.; Mauvais, P.; Lassaigne, P.; Bonnefoy, A.; Musicki, B. Synthesis and in vitro evaluation of novel highly potent coumarin inhibitors of gyrase B. Bioorganic Med. Chem. Lett. 1999, 9, 2079–2084. [Google Scholar] [CrossRef]
- Tolomelli, A.; Squassabia, F. Peptides and Peptidomimetics in Medicine, Surgery and Biotechnology. Curr. Med. Chem. 2006, 13, 2449–2466. [Google Scholar] [CrossRef]
- Cottiglia, F.; Loy, G.; Garau, D.; Floris, C.; Casu, M.; Pompei, R.; Bonsignore, L. Antimicrobial evaluation of coumarins and flavonoids from the stems of Daphne gnidium L. Phytomedicine 2001, 8, 302–305. [Google Scholar] [CrossRef]
- Smyth, T.; Ramachandran, V.; Smyth, W. A study of the antimicrobial activity of selected naturally occurring and synthetic coumarins. Int. J. Antimicrob. Agents 2009, 33, 421–426. [Google Scholar] [CrossRef]
- Kowalczyk, P.; Madej, A.; Paprocki, D.; Szymczak, M.; Ostaszewski, R. Coumarin Derivatives as New Toxic Compounds to Selected K12, R1–R4 E. coli Strains. Materials 2020, 13, 2499. [Google Scholar] [CrossRef]
- Emami, S.; Dadashpour, S. Current developments of coumarin-based anti-cancer agents in medicinal chemistry. Eur. J. Med. Chem. 2015, 102, 611–630. [Google Scholar] [CrossRef]
- Wilk, M.; Brodzka, A.; Koszelewski, D.; Madej, A.; Paprocki, D.; Żądło-Dobrowolska, A.; Ostaszewski, R. The influence of the isocyanoesters structure on the course of enzymatic Ugi reactions. Bioorganic Chem. 2019, 93, 102817. [Google Scholar] [CrossRef]
- Wilk, M.; Brodzka, A.; Koszelewski, D.; Samsonowicz-Górski, J.; Ostaszewski, R. Model Studies on the Enzyme-Regulated Diastereodivergent Cascade Passerini Reaction. Eur. J. Org. Chem. 2021, 2021, 4161–4165. [Google Scholar] [CrossRef]
- Madej, A.; Paprocki, D.; Koszelewski, D.; Żądło-Dobrowolska, A.; Brzozowska, A.; Walde, P.; Ostaszewski, R. Efficient Ugi reactions in an aqueous vesicle system. RSC Adv. 2017, 7, 33344. [Google Scholar] [CrossRef] [Green Version]
- Paprocki, D.; Koszelewski, D.; Walde, P.; Ostaszewski, R. Efficient Passerini reactions in an aqueous vesicle system. RSC Adv. 2015, 5, 102828–102835. [Google Scholar] [CrossRef]
- Agban, A.; Ounanian, M.; Luu-Duc, C.; Monget, D. Synthesis of new fluorogenic substrates derivated of 7-amino-4-trifluoromethylcoumarin. Detection of gram-negative bacteria, Streptococci group A and Enterococci. Ann. Pharm. Fr. 1990, 48, 326–334. [Google Scholar] [PubMed]
- Lukasiewicz, J.; Jachymek, W.; Niedziela, T.; Dzieciatkowska, M.; Lakomska, J.; Międzybrodzki, R.; Fortuna, W.; Szymaniec, S.; Misiuk Hojlo, M.; Lugowski, C. Serological characterization of anti-endotoxin serumdirected against the conjugate of oligosacharide core of Escherichia coli type R4 with tetanus toxoid. FEMS Immunol. Med. Microbiol. 2003, 37, 59–67. [Google Scholar] [CrossRef] [Green Version]
- Nnalue, N.A.; Khan, G.N.; Mustafa, N. Cross-reactivity between six Enterobacteriaceae complete lipopolysaccharide core chemotypes. J. Med. Microbiol. 1999, 48, 433–441. [Google Scholar] [CrossRef] [PubMed]
- Heinrichs, D.E.; Yethon, J.A.; Amor, P.A.; Whitfield, C. The assembly system for the outer core portion of R1 and R4 type lipopolysaccharides of Escherichia coli the r1 core specific β glucosyltransferase provides a novel attachment site for O-polysaccharides. J. Biol. Chem. 1998, 273, 29497–29505. [Google Scholar] [CrossRef] [Green Version]
- Amor, K.; Heinrichs, D.E.; Frirdich, E.; Ziebell, K.; Johnson, R.P.; Whitfield, C. Distribution of Core Oligosaccharide Types in Lipopolysaccharides from Escherichia coli. Infect. Immun. 2000, 68, 1116–1124. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Appelmelk, B.J.; An, Y.; Hekker, T.A.M.; Thijs, L.G.; MacLaren, D.M.; De Graaf, J. Frequencies of lipopolysaccharide core types in Escherichia coli strains from bacteraemic patients. Microbiology 1994, 140, 1119–1124. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Borkowski, A.; Ławniczak, L.; Cłapa, T.; Narożna, D.; Selwet, M.; Pęziak, D.; Markiewicz, B.; Chrzanowski, L. Different antibacterial activity of novel theophylline-based ionic liquids—Growth kinetic and cytotoxicity studies. Ecotoxicol. Environ. Saf. 2016, 130, 54–64. [Google Scholar] [CrossRef]
- Otaibi, A.A.; Sherwani, S.; Al-Zahrani, S.A.; Alshammari, E.M.; Khan, W.A.; Alsukaibi, A.K.D.; Khan, S.N.; Khan, M.W.A. Biologically Active α-Amino Amide Analogs and γδ T Cells-A Unique Anticancer Approach for Leukemia. Front Oncol. 2021, 11, 706586. [Google Scholar] [CrossRef] [PubMed]
- Krishnamoorthy, K.; Manivannan, G.; Kim, S.J.; Jeyasubramanian, K.; Premanathan, M. Antibacterial activity of MgO nanoparticles based on lipid peroxidation by oxygen vacancy. J. Nanopart. Res. 2012, 14, 1063. [Google Scholar] [CrossRef]
- Hough-Troutman, W.L.; Smiglak, M.; Griffin, S.; Reichert, W.M.; Mirska, I.; Jodynis-Liebert, J.; Adamska, T.; Nawrot, J.; Stasiewicz, M.; Rogers, R.D.; et al. Ionic liquids with dual biological function: Sweet and anti-microbial, hydrophobic quaternary ammonium-based salts. New J. Chem. 2009, 33, 26–33. [Google Scholar] [CrossRef]
- Inácio, Â.S.; Domingues, N.; Nunes, A.; Martins, P.; Moreno, M.J.; Estronca, L.; Fernandes, R.; Moreno, A.; Borrego, M.J.; Gomes, J.P.; et al. Quaternary ammonium surfactant structure determines selective toxicity towards bacteria: Mechanisms of action and clinical implications in antibacterial prophylaxis. J. Antimicrob. Chemother. 2015, 71, 641–654. [Google Scholar] [CrossRef] [Green Version]
- Jurado, J.; Saparbaev, M.; Matray, T.J.; Greenberg, M.M.; Laval, J. The Ring Fragmentation Product of Thymidine C5-Hydrate When Present in DNA Is Repaired by the Escherichia coli Fpg and Nth Proteins†. Biochemistry 1998, 37, 7757–7763. [Google Scholar] [CrossRef]
- Cussac, C.; Laval, F. Reduction of the Toxicity and Mutagenicity of Aziridine in Mammalian Cells Harboring the Escherichia Coli fpg Gene. Nucleic Acids Res. 1996, 24, 1742–1746. [Google Scholar] [CrossRef] [Green Version]
- Kawase, M.; Varu, B.; Shah, A.; Motohashi, N.; Tani, S.; Saito, S.; Debnath, S.; Mahapatra, S.; Dastidar, S.G.; Chakrabarty, A.N. Antimicrobial Activity of New Coumarin Derivatives. Arzneimittelforschung 2001, 51, 67–71. [Google Scholar] [CrossRef]
- Upadhyay, K.; Manvar, A.; Rawal, K.; Joshi, S.; Trivedi, J.; Chaniyara, R.; Shah, A. Evaluation of Structurally Diverse Benzoazepines Clubbed with Coumarins as Mycobacterium tuberculosis Agents. Chem. Biol. Drug Des. 2012, 80, 1003–1008. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, A.K. ChemInform Abstract: Potent HIV Protease Inhibitors Incorporating High-Affinity P2-Ligands and (R)-(Hydroxyethylamino)sulfonamide Isostere. ChemInform 2010, 29, 27. [Google Scholar] [CrossRef]
- Zhao, H.; Neamati, N.; Hong, H.; Mazumder, A.; Wang, S.; Sunder, S.; Milne, G.W.A.; Pommier, Y.; Burke, T.R. Coumarin-Based Inhibitors of HIV Integrase. J. Med. Chem. 1997, 40, 242–249. [Google Scholar] [CrossRef] [PubMed]
- Stromberg, Z.R.; Van Goor, A.; Redweik, G.A.J.; Brand, M.J.W.; Wannemuehler, M.J.; Mellata, M. Pathogenic and non-pathogenic Escherichia coli colonization and host inflammatory response in a defined microbiota mouse model. Dis. Models Mech. 2018, 11, dmm035063. [Google Scholar] [CrossRef] [Green Version]
- Larry, K.L.; Tanka, S.K. Therapy of Helicobacter pylori Infections: Current Status and Future Directions. In Annual Reports in Medicinal Chemistry; Academic Press: Cambridge, MA, USA, 1995; Volume 30, pp. 151–158. [Google Scholar]
- Kowalczyk, P.; Trzepizur, D.; Szymczak, M.; Skiba, G.; Kramkowski, K.; Ostaszewski, R. 1,2-Diarylethanols—A New Class of Compounds That Are Toxic to E. coli K12, R2–R4 Strains. Materials 2021, 14, 1025. [Google Scholar] [CrossRef]
- Kowalczyk, P.; Madej, A.; Szymczak, M.; Ostaszewski, R. α-Amidoamids as New Replacements of Antibiotics—Research on the Chosen K12, R2–R4 E. coli Strains. Materials 2020, 13, 5169. [Google Scholar] [CrossRef]
- Kowalczyk, P.; Borkowski, A.; Czerwonka, G.; Cłapa, T.; Cieśla, J.; Misiewicz, A.; Borowiec, M.; Szala, M. The microbial tox-icity of quaternary ammonium ionic liquids is dependent on the type of lipopolysaccharide. J. Mol. Liq. 2018, 266, 540–547. [Google Scholar] [CrossRef]
- Borkowski, A.; Kowalczyk, P.; Czerwonka, G.; Cieśla, J.; Cłapa, T.; Misiewicz, A.; Szala, M.; Drabik, M. Interaction of qua-ternary ammonium ionic liquids with bacterial membranes—Studies with Escherichia coli R1–R4-type lipopolysaccharides. J. Mol. Liq. 2017, 246, 282–289. [Google Scholar] [CrossRef]
- Kowalczyk, P.; Gawdzik, B.; Trzepizur, D.; Szymczak, M.; Skiba, G.; Raj, S.; Kramkowski, K.; Lizut, R.; Ostaszewski, R. δ-Lactones—A New Class of Compounds That Are Toxic to E. coli K12 and R2–R4 Strains. Materials 2021, 14, 2956. [Google Scholar] [CrossRef]
- Dissanayake, D.R.; Wijewardana, T.G.; Gunawardena, G.A.; Poxton, I.R. Distribution of lipopolysaccharide core types among avian pathogenic Escherichia coli in relation to the major phylogenetic groups. Vet. Microbiol. 2008, 132, 355–363. [Google Scholar] [CrossRef]
- Maciejewska, A.; Kaszowska, M.; Jachymek, W.; Lugowski, C.; Lukasiewicz, J. Lipopolysaccharide-linked Enterobacterial Common Antigen (ECALPS) Occurs in Rough Strains of Escherichia coli R1, R2, and R4. Int. J. Mol. Sci. 2020, 21, 6038. [Google Scholar] [CrossRef] [PubMed]
- Prost, M.E.; Prost, R. Basic parameters of evaluation of the effectiveness of antibiotic therapy. OphthaTherapy 2017, 4, 233–236. [Google Scholar] [CrossRef]
- D’Souza, L.J.; Gigant, B.; Knossow, M.; Green, B.S. Remarkable remote chiral recognition in a reaction mediated by a catalytic antibody. J. Am. Chem. Soc. 2002, 124, 2114–2115. [Google Scholar] [CrossRef] [PubMed]
- Castonguay, R.; Lherbet, C.; Keillor, J.W. Mapping of the active site of rat kidney γ-glutamyl transpeptidase using activated esters and their amide derivatives. Bioorganic Med. Chem. 2002, 10, 4185–4191. [Google Scholar] [CrossRef]
- Vrinceanu, D.; Dumitru, M.; Stefan, A.; Neagos, A.; Musat, G.; Nica, E.A. Severe DRESS syndrome after carbamazepine intake in a case with multiple addictions: A case report. Exp. Ther. Med. 2020, 20, 2377–2380. [Google Scholar] [CrossRef]
- Tamayo, M.; Santiso, R.; Gosálvez, J.; Bou, G.; del Carmen Fernandez, M.; Fernández, J.L. Cell wall active antibiotics re-duce chromosomal DNA fragmentation by peptidoglycan hydrolysis in Staphylococcus aureus. Arch. Microbiol. 2012, 194, 967–975. [Google Scholar] [CrossRef]
- Opoku-Temeng, C.; Onyedibe, K.I.; Aryal, U.K.; Sintim, H.O. Proteomic analysis of bacterial response to a 4-hydroxybenzylidene indolinone compound, which re-sensitizes bacteria to traditional antibiotics. J. Proteom. 2019, 202, 103368. [Google Scholar] [CrossRef]
- Osano, E.; Arakawa, Y.; Wacharotayankun, R.; Ohta, M.; Horii, T.; Ito, H.; Yoshimura, F.; Kato, N. Molecular characterization of an enterobacterial metallo beta-lactamase found in a clinical isolate of Serratia marcescens that shows imipenem resistance. Antimicrob. Agents Chemother. 1994, 38, 71–78. [Google Scholar] [CrossRef] [Green Version]



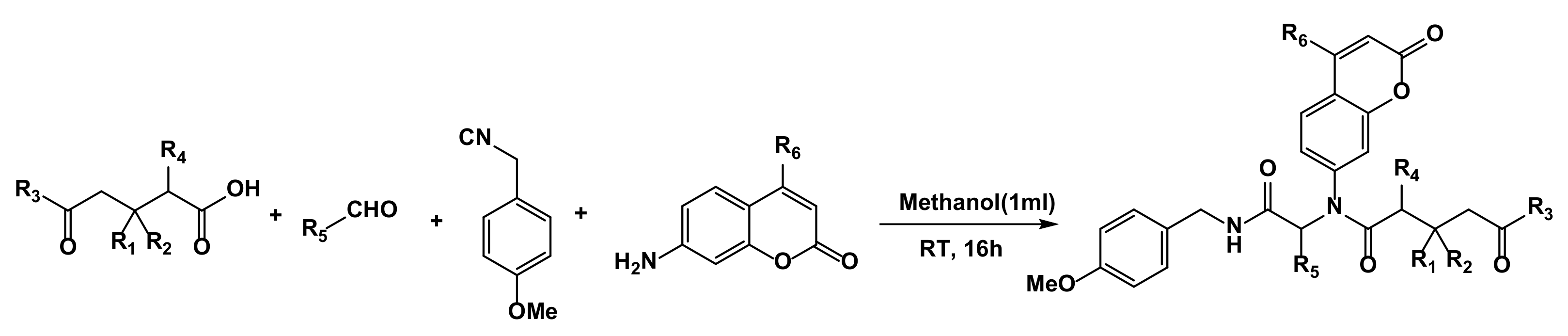


| Entry | R1 | R2 | R3 | R4 | R5 | R6 | Product Symbol | Yield (%) |
|---|---|---|---|---|---|---|---|---|
| 1 | H | H | 4-NO2-C6H4OH | H | C6H5 | CH3 | 4 | 25 |
| 2 | H | H | OH | H | C6H5 | CH3 | 5 | 90 |
| 3 | H | H | 4-NO2-C6H4OH | H | C6H5 | CF3 | 7 | 32 |
| 4 | CH3 | CH3 | OH | H | C6H5 | CF3 | 6 | 13 |
| 5 | H | H | OH | H | C6H5 | CF3 | 8 | 25 |
| 6 | CH3 | CH3 | OH | H | p-OCH3-C6H4 | CF3 | 10 | 10 |
| 7 | CH3 | CH3 | OH | H | p-NO2-C6H4 | CF3 | 11 | 16 |
| 8 | CH3 | CH3 | OH | H | CH2CH(CH3)2 | CF3 | 12 | 3 |
| 9 | CH3 | CH3 | OH | H | p-CH3-C6H4 | CF3 | 13 | 20 |
| 10 | CH3 | CH3 | OH | H | CH=CH-C6H5 | CF3 | 14 | 25 |
| 11 | O | OH | O | C6H5 | CF3 | 9 | 25 | |
| No. of Samples | 2 | 3 | 4 | 7 | 10 | 15 | 16 | 17 | Type of Test |
|---|---|---|---|---|---|---|---|---|---|
| K12 | *** | ** | *** | ** | ** | * | ** | ** | MIC |
| R2 | *** | ** | *** | ** | ** | * | *** | ** | MIC |
| R3 | *** | ** | *** | ** | ** | * | ** | ** | MIC |
| R4 | *** | ** | *** | ** | ** | * | ** | ** | MIC |
| K12 | *** | ** | ** | ** | ** | * | ** | ** | MBC |
| R2 | ** | ** | ** | ** | ** | * | ** | ** | MBC |
| R3 | ** | ** | ** | ** | ** | * | ** | ** | MBC |
| R4 | ** | ** | ** | ** | ** | * | ** | ** | MBC |
| K12 | * | ** | * | * | * | * | ** | * | MBC/MIC |
| R2 | * | * | * | * | * | * | ** | * | MBC/MIC |
| R3 | * | * | * | * | * | * | ** | * | MBC/MIC |
| R4 | * | * | * | * | * | * | ** | * | MBC/MIC |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kowalczyk, P.; Wilk, M.; Parul, P.; Szymczak, M.; Kramkowski, K.; Raj, S.; Skiba, G.; Sulejczak, D.; Kleczkowska, P.; Ostaszewski, R. The Synthesis and Evaluation of Aminocoumarin Peptidomimetics as Cytotoxic Agents on Model Bacterial E. coli Strains. Materials 2021, 14, 5725. https://doi.org/10.3390/ma14195725
Kowalczyk P, Wilk M, Parul P, Szymczak M, Kramkowski K, Raj S, Skiba G, Sulejczak D, Kleczkowska P, Ostaszewski R. The Synthesis and Evaluation of Aminocoumarin Peptidomimetics as Cytotoxic Agents on Model Bacterial E. coli Strains. Materials. 2021; 14(19):5725. https://doi.org/10.3390/ma14195725
Chicago/Turabian StyleKowalczyk, Paweł, Monika Wilk, Parul Parul, Mateusz Szymczak, Karol Kramkowski, Stanisława Raj, Grzegorz Skiba, Dorota Sulejczak, Patrycja Kleczkowska, and Ryszard Ostaszewski. 2021. "The Synthesis and Evaluation of Aminocoumarin Peptidomimetics as Cytotoxic Agents on Model Bacterial E. coli Strains" Materials 14, no. 19: 5725. https://doi.org/10.3390/ma14195725
APA StyleKowalczyk, P., Wilk, M., Parul, P., Szymczak, M., Kramkowski, K., Raj, S., Skiba, G., Sulejczak, D., Kleczkowska, P., & Ostaszewski, R. (2021). The Synthesis and Evaluation of Aminocoumarin Peptidomimetics as Cytotoxic Agents on Model Bacterial E. coli Strains. Materials, 14(19), 5725. https://doi.org/10.3390/ma14195725








