DNA Origami as Emerging Technology for the Engineering of Fluorescent and Plasmonic-Based Biosensors
Abstract
1. Introduction
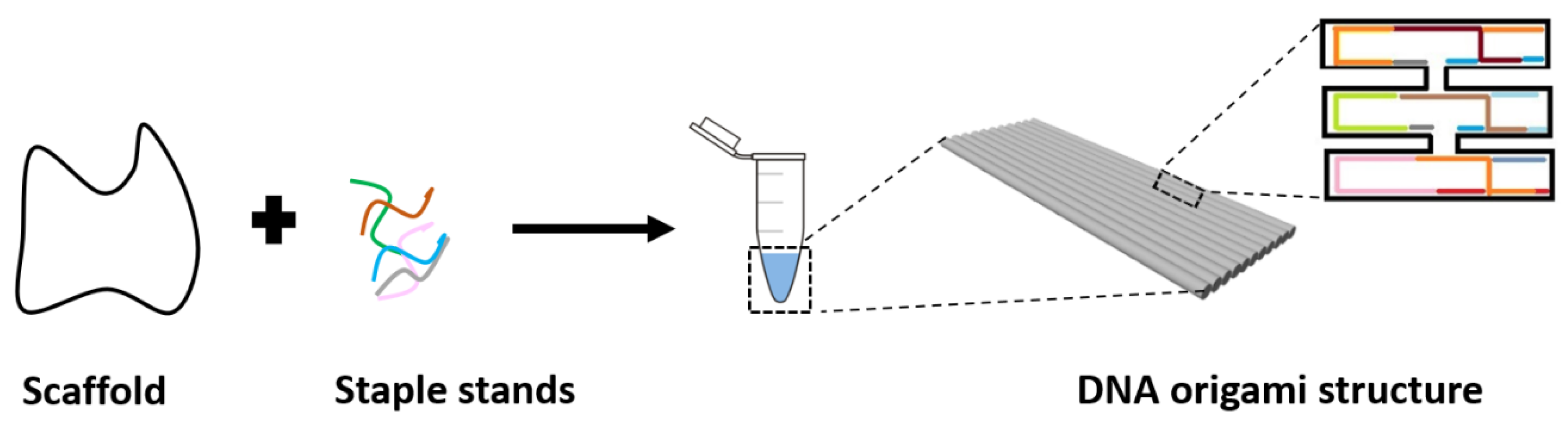
1.1. DNA Nanotechnology
1.2. DNA Origami-Based Structures Used for Biomolecular Sensing
2. DNA Origami Complexes Used in Optical Biomolecular Sensing
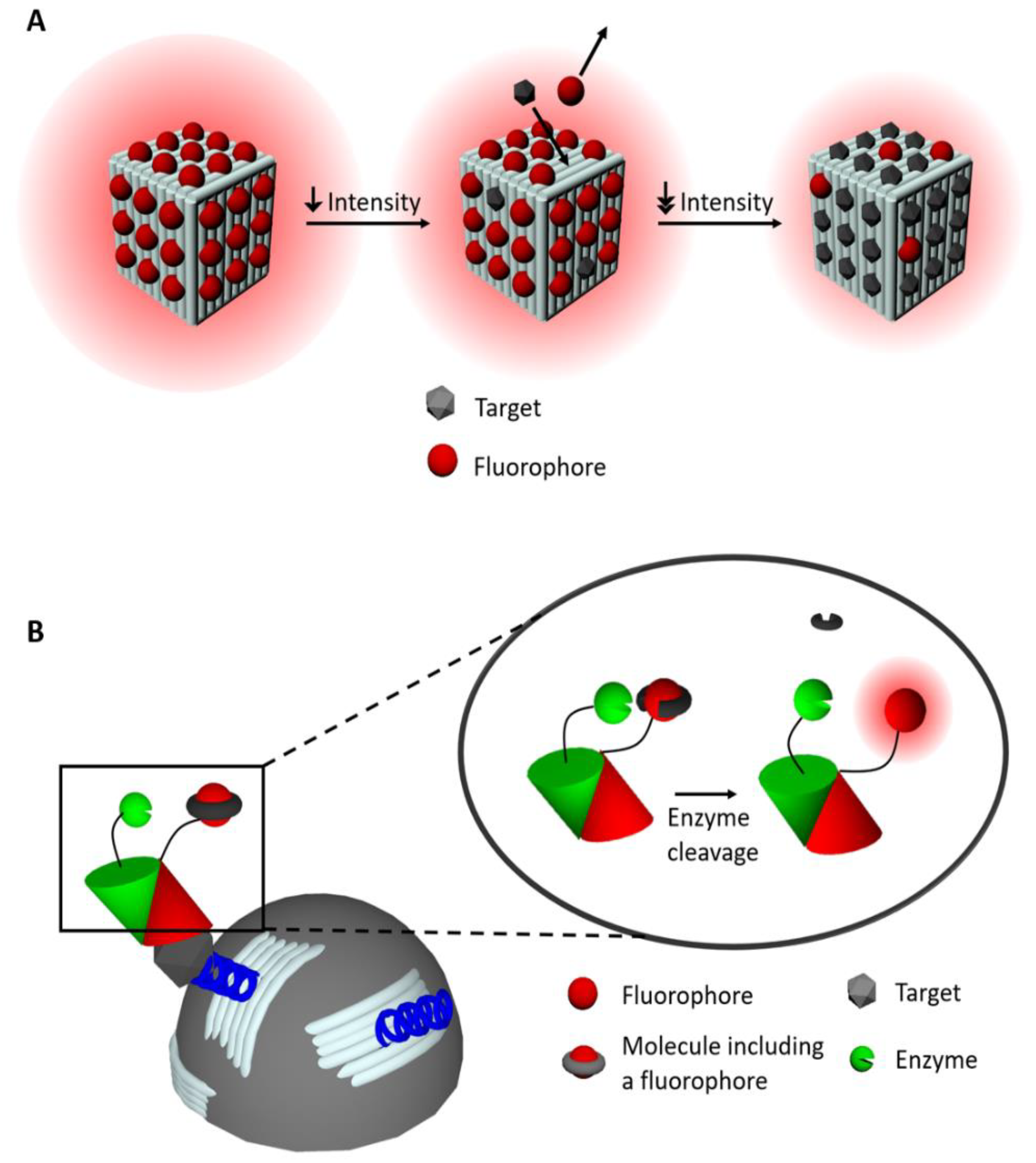
2.1. Fluorescence-Based Sensors
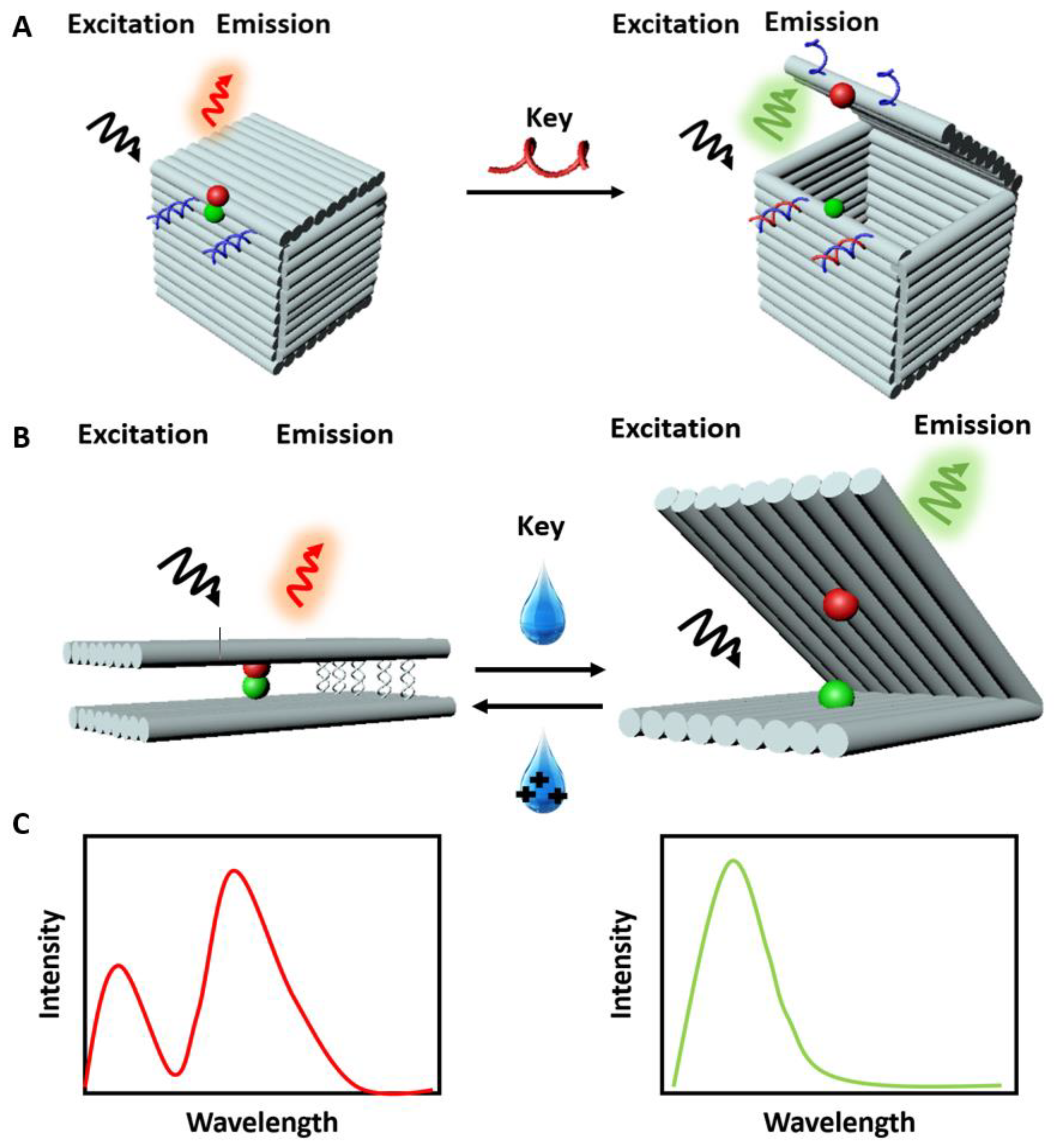
2.2. Fluorescence (Förster) Resonance Energy Transfer (FRET)-Based Sensors
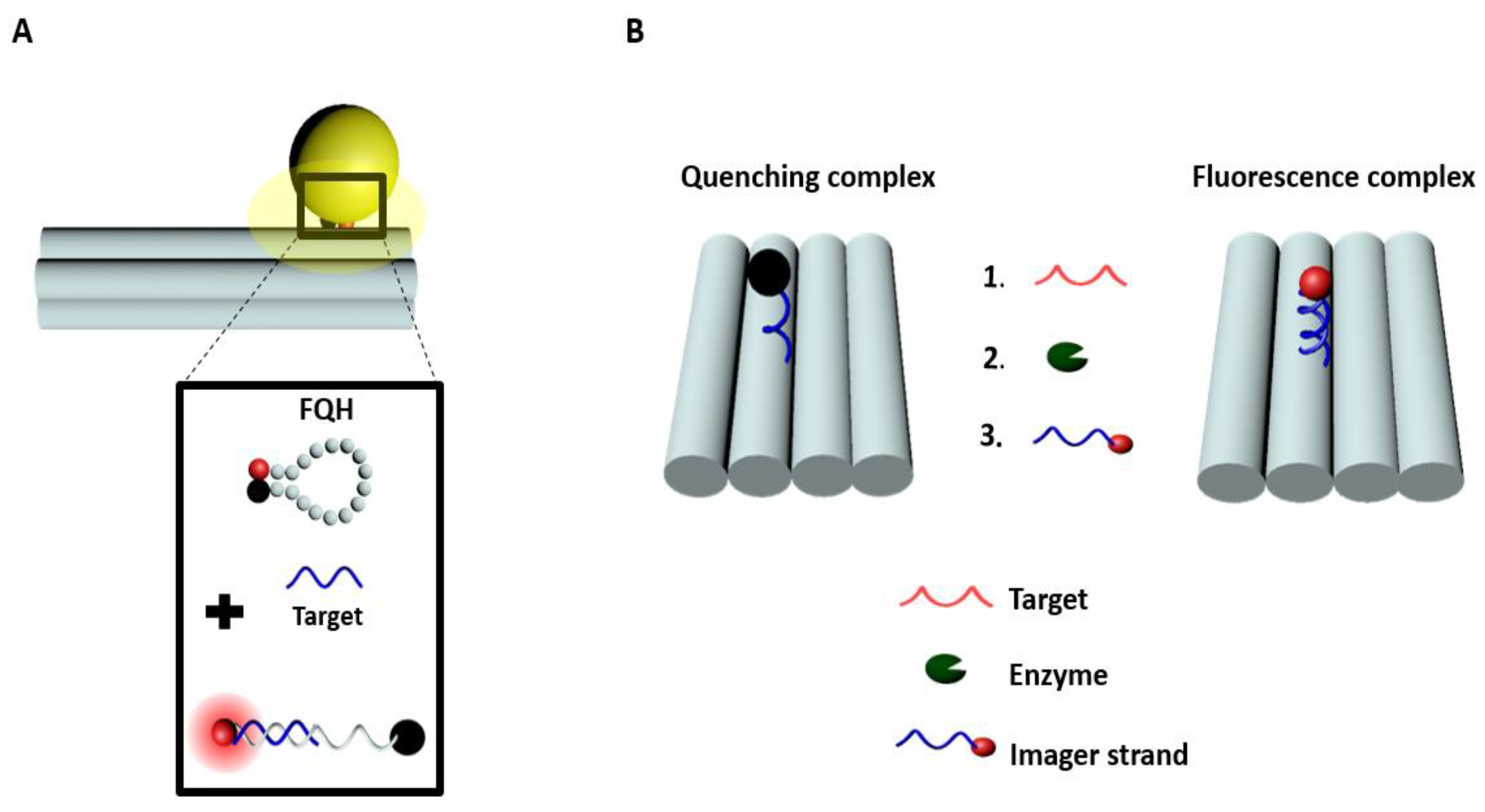
2.3. Quenching-Based Sensors
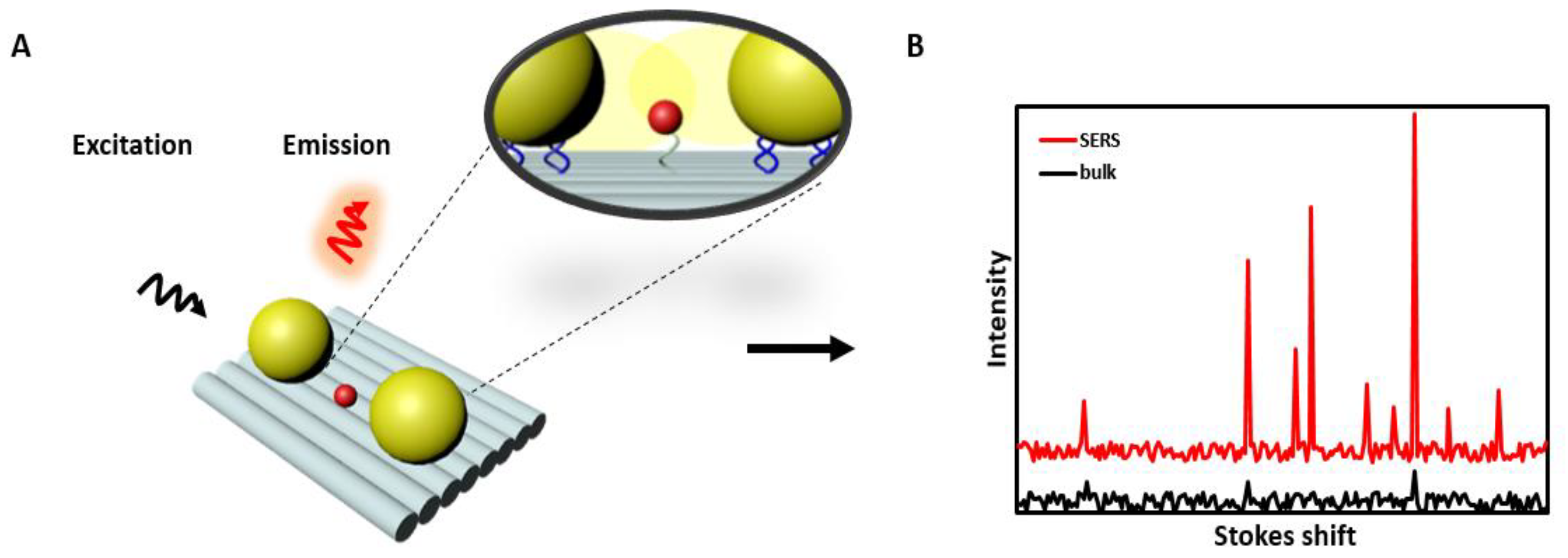
2.4. Surface-Enhanced Raman Scattering-Based Sensors
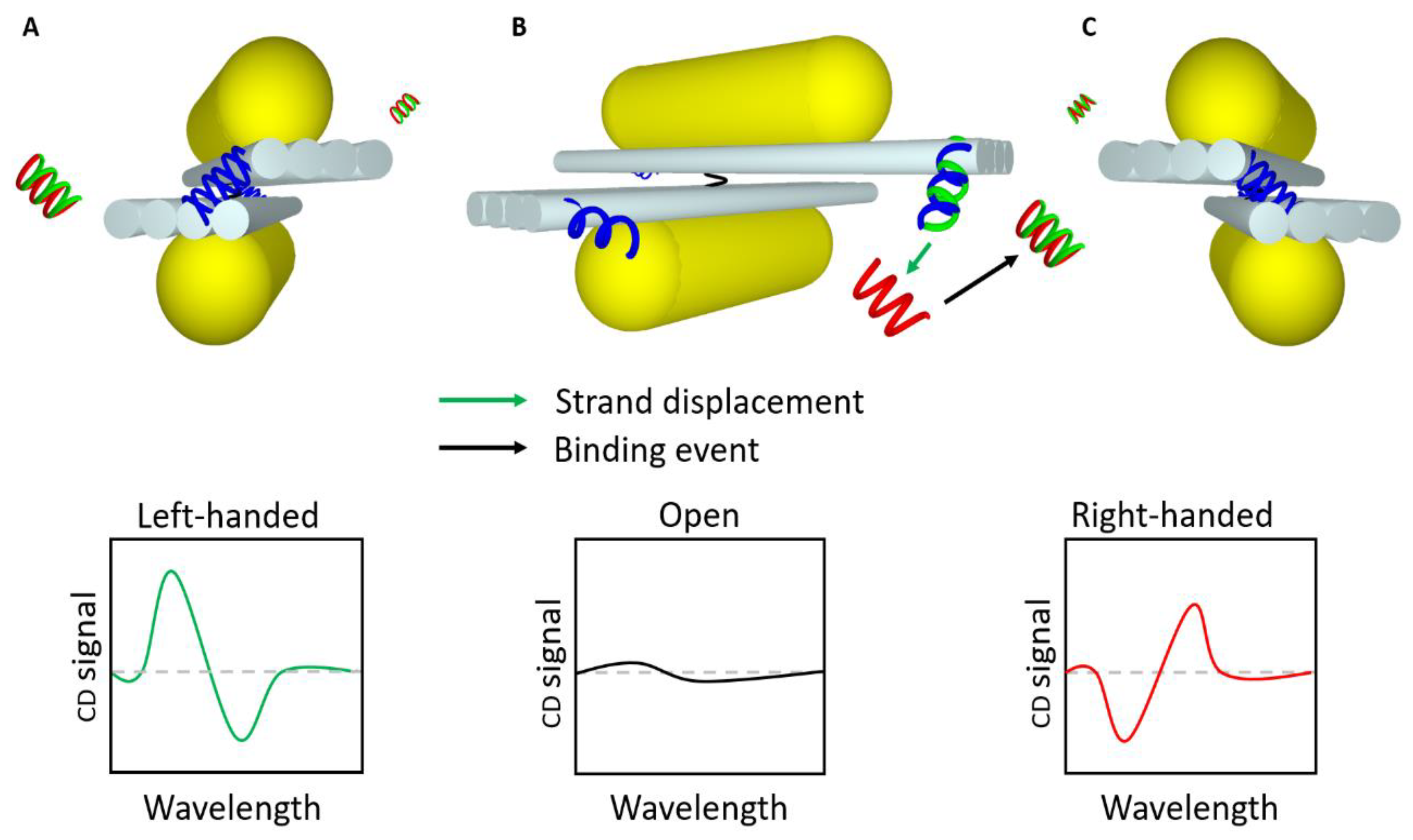
2.5. Circular Dichroism-Based Sensors
3. Conclusions and Perspectives
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| ATP | Adenosine Triphosphate |
| AuNPs | Gold Nanoparticles |
| BG | Background |
| CD | Circular Dichroism Spectroscopy |
| DNA-Box | DNA Origami Box |
| DNA-OS | DNA Origami Structure |
| dsDNA | Double-stranded DNA |
| DV | DNA Nanovault |
| EF | Enhancement Factor |
| FQH | Fluorescence-Quenching Hairpin |
| FRET | Fluorescence (Förster) Resonance Energy Transfer |
| LH | Left-Handed |
| NP | Nanoparticle |
| O/C | Open-Close States Change |
| PfLDH | Plasmodium Falciparum Lactate Dehydrogenase |
| POP | Proof of Principle |
| RH | Right-Handed |
| SERS | Surface-Enhanced Raman Spectroscopy |
| SiNW | Single Silicon Nanowire |
| ssDNA | Single-stranded DNA |
| T1/2 | time that takes to reach 50% of maximum FRET difference |
| TMSD | Toehold Mediated Strand Displacement |
References
- Pitchiaya, S.; Krishnan, Y. First blueprint, now bricks: DNA as construction material on the nanoscale. Chem. Soc. Rev. 2006, 35, 1111–1121. [Google Scholar] [CrossRef] [PubMed]
- Seeman, N.C. Nucleic acid junctions and lattices. J. Theor. Biol. 1982, 99, 237–247. [Google Scholar] [CrossRef]
- Kallenbach, N.R.; Ma, R.-I.; Seeman, N.C. An immobile nucleic acid junction constructed from oligonucleotides. Nature 1983, 305, 829–831. [Google Scholar] [CrossRef]
- He, Y.; Ye, T.; Su, M.; Zhang, C.; Ribbe, A.E.; Jiang, W.; Mao, C. Hierarchical self-assembly of DNA into symmetric supramolecular polyhedra. Nature 2008, 452, 198–201. [Google Scholar] [CrossRef]
- Aldaye, F.A.; Sleiman, H.F. Modular Access to Structurally Switchable 3D Discrete DNA Assemblies. J. Am. Chem. Soc. 2007, 129, 13376–13377. [Google Scholar] [CrossRef]
- Mathieu, F.; Liao, S.; Kopatsch, J.; Wang, T.; Mao, C.; Seeman, N.C. Six-Helix Bundles Designed from DNA. Nano Lett. 2005, 5, 661–665. [Google Scholar] [CrossRef]
- Winfree, E.; Liu, F.; Wenzler, L.A.; Seeman, N.C. Design and self-assembly of two-dimensional DNA crystals. Nature 1998, 394, 539–544. [Google Scholar] [CrossRef]
- Zheng, J.; Birktoft, J.J.; Chen, Y.; Wang, T.; Sha, R.; Constantinou, P.E.; Ginell, S.L.; Mao, C.; Seeman, N.C. From molecular to macroscopic via the rational design of a self-assembled 3D DNA crystal. Nature 2009, 461, 74–77. [Google Scholar] [CrossRef]
- Wei, B.; Dai, M.; Yin, P. Complex shapes self-assembled from single-stranded DNA tiles. Nature 2012, 485, 623–626. [Google Scholar] [CrossRef]
- Ke, Y.; Ong, L.L.; Shih, W.M.; Yin, P. Three-Dimensional Structures Self-Assembled from DNA Bricks. Science 2012, 338, 1177–1183. [Google Scholar] [CrossRef]
- Chandrasekaran, A.R. Programmable DNA scaffolds for spatially-ordered protein assembly. Nanoscale 2016, 8, 4436–4446. [Google Scholar] [CrossRef] [PubMed]
- Rothemund, P.W.K. Folding DNA to create nanoscale shapes and patterns. Nature 2006, 440, 297–302. [Google Scholar] [CrossRef] [PubMed]
- Chandrasekaran, A.R.; Punnoose, J.A.; Zhou, L.; Dey, P.; Dey, B.K.; Halvorsen, K. DNA nanotechnology approaches for microRNA detection and diagnosis. Nucleic Acids Res. 2019, 47, 10489–10505. [Google Scholar] [CrossRef] [PubMed]
- Shen, B.; Kostiainen, M.A.; Linko, V. DNA Origami Nanophotonics and Plasmonics at Interfaces. Langmuir 2018, 34, 14911–14920. [Google Scholar] [CrossRef]
- Xavier, P.L.; Chandrasekaran, A.R. DNA-based construction at the nanoscale: Emerging trends and applications. Nanotechnology 2018, 29, 062001. [Google Scholar] [CrossRef]
- Liu, X.; Zhao, Y.; Liu, P.; Wang, L.; Lin, J.; Fan, C. Biomimetic DNA Nanotubes: Nanoscale Channel Design and Applications. Angew. Chem. Int. Ed. 2019, 58, 8996–9011. [Google Scholar] [CrossRef]
- Uprety, B.; Jensen, J.; Aryal, B.R.; Davis, R.C.; Woolley, A.T.; Harb, J.N. Directional Growth of DNA-Functionalized Nanorods to Enable Continuous, Site-Specific Metallization of DNA Origami Templates. Langmuir 2017, 33, 10143–10152. [Google Scholar] [CrossRef]
- Andersen, E.S.; Dong, M.; Nielsen, M.M.; Jahn, K.; Subramani, R.; Mamdouh, W.; Golas, M.M.; Sander, B.; Stark, H.; Oliveira, C.L.; et al. Self-assembly of a nanoscale DNA box with a controllable lid. Nature 2009, 459, 73–76. [Google Scholar] [CrossRef]
- Funck, T.; Nicoli, F.; Kuzyk, A.; Liedl, T. Sensing Picomolar Concentrations of RNA Using Switchable Plasmonic Chirality. Angew. Chem. Int. Ed. 2018, 57, 13495–13498. [Google Scholar] [CrossRef]
- Kuzyk, A.; Urban, M.J.; Idili, A.; Ricci, F.; Liu, N. Selective control of reconfigurable chiral plasmonic metamolecules. Sci. Adv. 2017, 3, e1602803. [Google Scholar] [CrossRef]
- Selnihhin, D.; Sparvath, S.M.; Preus, S.; Birkedal, V.; Andersen, E.S. Multifluorophore DNA Origami Beacon as a Biosensing Platform. ACS Nano 2018, 12, 5699–5708. [Google Scholar] [CrossRef] [PubMed]
- Douglas, S.M.; Bachelet, I.; Church, G.M. A Logic-Gated Nanorobot for Targeted Transport of Molecular Payloads. Science 2012, 335, 831–834. [Google Scholar] [CrossRef] [PubMed]
- Ding, S.Y.; You, E.-M.; Tian, Z.Q.; Moskovits, M. Electromagnetic theories of surface-enhanced Raman spectroscopy. Chem. Soc. Rev. 2017, 46, 4042–4076. [Google Scholar] [CrossRef] [PubMed]
- Zadegan, R.M.; Jepsen, M.D.; Thomsen, K.E.; Okholm, A.H.; Schaffert, D.H.; Andersen, E.S.; Birkedal, V.; Kjems, J. Construction of a 4 zeptoliters switchable 3D DNA box origami. ACS Nano 2012, 6, 10050–10053. [Google Scholar] [CrossRef] [PubMed]
- Strauss, M.T.; Schueder, F.; Haas, D.; Nickels, P.C.; Jungmann, R. Quantifying absolute addressability in DNA origami with molecular resolution. Nat. Commun. 2018, 9, 1600. [Google Scholar] [CrossRef] [PubMed]
- Kumar, V.; Palazzolo, S.; Bayda, S.; Corona, G.; Toffoli, G.; Rizzolio, F. DNA Nanotechnology for Cancer Therapy. Theranostics 2016, 6, 710–725. [Google Scholar] [CrossRef]
- Zhang, F.; Nangreave, J.; Liu, Y.; Yan, H. Structural DNA Nanotechnology: State of the Art and Future Perspective. J. Am. Chem. Soc. 2014, 136, 11198–11211. [Google Scholar] [CrossRef]
- Mei, Q.; Wei, X.; Su, F.; Liu, Y.; Youngbull, C.; Johnson, R.; Lindsay, S.; Yan, H.; Meldrum, D. Stability of DNA Origami Nanoarrays in Cell Lysate. Nano Lett. 2011, 11, 1477–1482. [Google Scholar] [CrossRef]
- Ijas, H.; Nummelin, S.; Shen, B.; Kostiainen, M.A.; Linko, V. Dynamic DNA Origami Devices: From Strand-Displacement Reactions to External-Stimuli Responsive Systems. Int. J. Mol. Sci. 2018, 19, 2114. [Google Scholar] [CrossRef]
- Goetzfried, M.A.; Vogele, K.; Mückl, A.; Kaiser, M.; Holland, N.B.; Simmel, F.C.; Pirzer, T. Periodic Operation of a Dynamic DNA Origami Structure Utilizing the Hydrophilic–Hydrophobic Phase-Transition of Stimulus-Sensitive Polypeptides. Small 2019, 15, 1903541. [Google Scholar] [CrossRef]
- Linko, V.; Ora, A.; Kostiainen, M.A. DNA Nanostructures as Smart Drug-Delivery Vehicles and Molecular Devices. Trends Biotechnol. 2015, 33, 586–594. [Google Scholar] [CrossRef] [PubMed]
- Bald, I.; Keller, A. Molecular processes studied at a single-molecule level using DNA origami nanostructures and atomic force microscopy. Molecules 2014, 19, 13803–13823. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Meyer, T.A.; Pan, V.; Dutta, P.K.; Ke, Y. The Beauty and Utility of DNA Origami. Chem 2017, 2, 359–382. [Google Scholar] [CrossRef]
- Vietz, C.; Lalkens, B.; Acuna, G.P.; Tinnefeld, P. Synergistic Combination of Unquenching and Plasmonic Fluorescence Enhancement in Fluorogenic Nucleic Acid Hybridization Probes. Nano Lett. 2017, 17, 6496–6500. [Google Scholar] [CrossRef]
- Acuna, G.P.; Moller, F.M.; Holzmeister, P.; Beater, S.; Lalkens, B.; Tinnefeld, P. Fluorescence enhancement at docking sites of DNA-directed self-assembled nanoantennas. Science 2012, 338, 506–510. [Google Scholar] [CrossRef]
- Prinz, J.; Schreiber, B.; Olejko, L.; Oertel, J.; Rackwitz, J.; Keller, A.; Bald, I. DNA Origami Substrates for Highly Sensitive Surface-Enhanced Raman Scattering. J. Phys. Chem. Lett. 2013, 4, 4140–4145. [Google Scholar] [CrossRef]
- Roller, E.-M.; Besteiro, L.V.; Pupp, C.; Khorashad, L.K.; Govorov, A.O.; Liedl, T. Hotspot-mediated non-dissipative and ultrafast plasmon passage. Nat. Phys. 2017, 13, 761–765. [Google Scholar] [CrossRef]
- Kuzyk, A.; Jungmann, R.; Acuna, G.P.; Liu, N. DNA Origami Route for Nanophotonics. ACS Photonics 2018, 5, 1151–1163. [Google Scholar] [CrossRef]
- Farran, B.; Dyson, G.; Craig, D.; Dombkowski, A.; Beebe-Dimmer, J.L.; Powell, I.J.; Podgorski, I.; Heilbrun, L.; Bolton, S.; Bock, C.H. A study of circulating microRNAs identifies a new potential biomarker panel to distinguish aggressive prostate cancer. Carcinogenesis 2018, 39, 556–561. [Google Scholar] [CrossRef]
- Kim, S.W.; Li, Z.; Moore, P.S.; Monaghan, A.P.; Chang, Y.; Nichols, M.; John, B. A sensitive non-radioactive northern blot method to detect small RNAs. Nucleic Acids Res. 2010, 38, e98. [Google Scholar] [CrossRef]
- Gasperskaja, E.; Kučinskas, V. The most common technologies and tools for functional genome analysis. Acta Med. Litu. 2017, 24, 1–11. [Google Scholar] [CrossRef]
- Salek-Maghsoudi, A.; Vakhshiteh, F.; Torabi, R.; Hassani, S.; Ganjali, M.R.; Norouzi, P.; Hosseini, M.; Abdollahi, M. Recent advances in biosensor technology in assessment of early diabetes biomarkers. Biosens. Bioelectron. 2018, 99, 122–135. [Google Scholar] [CrossRef]
- Rutten, I.; Daems, D.; Lammertyn, J. Boosting biomolecular interactions through DNA origami nano-tailored biosensing interfaces. J. Mater. Chem. B. 2020. [Google Scholar] [CrossRef]
- Chao, J.; Zhu, D.; Zhang, Y.; Wang, L.; Fan, C. DNA nanotechnology-enabled biosensors. Biosens. Bioelectron. 2016, 76, 68–79. [Google Scholar] [CrossRef] [PubMed]
- Gerard, M.; Chaubey, A.; Malhotra, B.D. Application of conducting polymers to biosensors. Biosens. Bioelectron. 2002, 17, 345–359. [Google Scholar] [CrossRef]
- Kuzuya, A.; Sakai, Y.; Yamazaki, T.; Xu, Y.; Komiyama, M. Nanomechanical DNA origami ’single-molecule beacons’ directly imaged by atomic force microscopy. Nat. Commun. 2011, 2, 449. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Li, Y.; Tian, C.; Mao, C. A Smart DNA Tetrahedron That Isothermally Assembles or Dissociates in Response to the Solution pH Value Changes. Biomacromolecules 2013, 14, 1711–1714. [Google Scholar] [CrossRef]
- Gerling, T.; Wagenbauer, K.F.; Neuner, A.; Dietz, H. Dynamic DNA devices and assemblies formed by shape-complementary, non-basepairing 3D components. Science 2015, 347, 1446–1452. [Google Scholar] [CrossRef]
- Juul, S.; Iacovelli, F.; Falconi, M.; Kragh, S.L.; Christensen, B.; Frøhlich, R.; Franch, O.; Kristoffersen, E.L.; Stougaard, M.; Leong, K.W.; et al. Temperature-Controlled Encapsulation and Release of an Active Enzyme in the Cavity of a Self-Assembled DNA Nanocage. ACS Nano 2013, 7, 9724–9734. [Google Scholar] [CrossRef]
- Li, M.; Cushing, S.K.; Wu, N. Plasmon-enhanced optical sensors: A review. Analyst 2015, 140, 386–406. [Google Scholar] [CrossRef]
- Zhang, X.; Hu, Y.; Yang, X.; Tang, Y.; Han, S.; Kang, A.; Deng, H.; Chi, Y.; Zhu, D.; Lu, Y. Förster resonance energy transfer (FRET)-based biosensors for biological applications. Biosens. Bioelectron. 2019, 138. [Google Scholar] [CrossRef] [PubMed]
- Funck, T.; Liedl, T.; Bae, W. Dual Aptamer-Functionalized 3D Plasmonic Metamolecule for Thrombin Sensing. Appl. Sci. 2019, 9, 3006. [Google Scholar] [CrossRef]
- Simoncelli, S.; Roller, E.-M.; Urban, P.; Schreiber, R.; Turberfield, A.J.; Liedl, T.; Lohmüller, T. Quantitative Single-Molecule Surface-Enhanced Raman Scattering by Optothermal Tuning of DNA Origami-Assembled Plasmonic Nanoantennas. ACS Nano 2016, 10, 9809–9815. [Google Scholar] [CrossRef] [PubMed]
- Daems, D.; Rutten, I.; Bath, J.; Decrop, D.; Van Gorp, H.; Ruiz, E.P.; De Feyter, S.; Turberfield, A.J.; Lammertyn, J. Controlling the Bioreceptor Spatial Distribution at the Nanoscale for Single Molecule Counting in Microwell Arrays. ACS Sens. 2019, 4, 2327–2335. [Google Scholar] [CrossRef] [PubMed]
- Olejko, L.; Cywiński, P.J.; Bald, I. An ion-controlled four-color fluorescent telomeric switch on DNA origami structures. Nanoscale 2016, 8, 10339–10347. [Google Scholar] [CrossRef] [PubMed]
- Suo, Z.; Chen, J.; Hou, X.; Hu, Z.; Xing, F.; Feng, L. Growing prospects of DNA nanomaterials in novel biomedical applications. RSC Adv. 2019, 9, 16479–16491. [Google Scholar] [CrossRef]
- Lakowicz, J.R. Principles of Fluorescence Spectroscopy, 3rd ed.; Springer: Boston, MA, USA, 2006. [Google Scholar]
- Hecht, B.; Novotny, L. Principles of Nano-Optics, 2nd ed.; Cambridge University Press: Cambridge, UK, 2006. [Google Scholar]
- Albinsson, B.; Hannestad, J.K.; Börjesson, K. Functionalized DNA nanostructures for light harvesting and charge separation. Coord. Chem. Rev. 2012, 256, 2399–2413. [Google Scholar] [CrossRef]
- Strunk, J.J.; Gregor, I.; Becker, Y.; Lamken, P.; Lata, S.; Reichel, A.; Enderlein, J.; Piehler, J. Probing Protein Conformations by in Situ Non-Covalent Fluorescence Labeling. Bioconjug. Chem. 2009, 20, 41–46. [Google Scholar] [CrossRef]
- Suei, S.; Raudsepp, A.; Kent, L.M.; Keen, S.A.J.; Filichev, V.V.; Williams, M.A.K. DNA visualization in single molecule studies carried out with optical tweezers: Covalent versus non-covalent attachment of fluorophores. Biochem. Biophys. Res. 2015, 466, 226–231. [Google Scholar] [CrossRef]
- Domljanovic, I.; Rexen Ulven, E.; Ulven, T.; Thomsen, R.P.; Okholm, A.H.; Kjems, J.; Voss, A.; Taskova, M.; Astakhova, K. Dihydropyridine Fluorophores Allow for Specific Detection of Human Antibodies in Serum. ACS Omega 2018, 3, 7580–7586. [Google Scholar] [CrossRef]
- Ihmels, H.; Otto, D. Intercalation of Organic Dye Molecules into Double-Stranded DNA—General Principles and Recent Developments. In Supermolecular Dye Chemistry; Würthner, F., Ed.; Springer: Berlin, Germany, 2005; pp. 161–204. [Google Scholar]
- Stassi, S.; Marini, M.; Allione, M.; Lopatin, S.; Marson, D.; Laurini, E.; Pricl, S.; Pirri, C.F.; Ricciardi, C.; Di Fabrizio, E. Nanomechanical DNA resonators for sensing and structural analysis of DNA-ligand complexes. Nat. Commun. 2019, 10, 1690. [Google Scholar] [CrossRef] [PubMed]
- Csapo, Z.; Gerstner, A.; Sasvari-Szekely, M.; Guttman, A. Automated Ultra-Thin-Layer SDS Gel Electrophoresis of Proteins Using Noncovalent Fluorescent Labeling. Anal. Chem. 2000, 72, 2519–2525. [Google Scholar] [CrossRef] [PubMed]
- Neto, B.; Lapis, A. Recent Developments in the Chemistry of Deoxyribonucleic Acid (DNA) Intercalators: Principles, Design, Synthesis, Applications and Trends. Molecules 2009, 14, 1725–1746. [Google Scholar] [CrossRef] [PubMed]
- Carlsson, C.; Larsson, A.; Björkman, M.; Jonsson, M.; Albinsson, B. Experimental and simulated fluorescence depolarization due to energy transfer as tools to study DNA-dye interactions. Biopolymers 1997, 41, 481–494. [Google Scholar] [CrossRef]
- Xiao, M.; Lai, W.; Man, T.; Chang, B.; Li, L.; Chandrasekaran, A.R.; Pei, H. Rationally Engineered Nucleic Acid Architectures for Biosensing Applications. Chem. Rev. 2019, 119, 11631–11717. [Google Scholar] [CrossRef]
- Domljanovic, I.; Carstens, A.; Okholm, A.; Kjems, J.; Nielsen, C.T.; Heegaard, N.H.H.; Astakhova, K. Complexes of DNA with fluorescent dyes are effective reagents for detection of autoimmune antibodies. Sci. Rep. 2017, 7, 1925. [Google Scholar] [CrossRef]
- Chen, G.; Liu, D.; He, C.; Gannett, T.R.; Lin, W.; Weizmann, Y. Enzymatic Synthesis of Periodic DNA Nanoribbons for Intracellular pH Sensing and Gene Silencing. J. Am. Chem. Soc. 2015, 137, 3844–3851. [Google Scholar] [CrossRef]
- Vietz, C.; Schutte, M.L.; Wei, Q.; Richter, L.; Lalkens, B.; Ozcan, A.; Tinnefeld, P.; Acuna, G.P. Benchmarking Smartphone Fluorescence-Based Microscopy with DNA Origami Nanobeads: Reducing the Gap toward Single-Molecule Sensitivity. ACS Omega 2019, 4, 637–642. [Google Scholar] [CrossRef]
- Trofymchuk, K.; Glembockyte, V.; Grabenhorst, L.; Steiner, F.; Vietz, C.; Close, C.; Pfeiffer, M.; Richter, L.; Schütte, M.L.; Selbach, F.; et al. Addressable Nanoantennas with Cleared Hotspots for Single-Molecule Detection on a Portable Smartphone Microscope. bioRxiv 2020. [Google Scholar] [CrossRef]
- Puchkova, A.; Vietz, C.; Pibiri, E.; Wünsch, B.; Sanz Paz, M.; Acuna, G.P.; Tinnefeld, P. DNA Origami Nanoantennas with over 5000-fold Fluorescence Enhancement and Single-Molecule Detection at 25 μM. Nano Lett. 2015, 15, 8354–8359. [Google Scholar] [CrossRef]
- Bui, H.; Díaz, S.A.; Fontana, J.; Chiriboga, M.; Veneziano, R.; Medintz, I.L. Utilizing the Organizational Power of DNA Scaffolds for New Nanophotonic Applications. Adv. Opt. Mater. 2019, 7, 1900562. [Google Scholar] [CrossRef]
- Stryer, L. Fluorescence energy transfer as a spectroscopic ruler. Ann. Rev. Biochem. 1978, 47, 819–846. [Google Scholar] [CrossRef] [PubMed]
- Buckhout-White, S.; Spillmann, C.M.; Algar, W.R.; Khachatrian, A.; Melinger, J.S.; Goldman, E.R.; Ancona, M.G.; Medintz, I.L. Assembling programmable FRET-based photonic networks using designer DNA scaffolds. Nat. Commun. 2014, 5, 5615. [Google Scholar] [CrossRef] [PubMed]
- Khrenova, M.; Topol, I.; Collins, J.; Nemukhin, A. Estimating orientation factors in the FRET theory of fluorescent proteins: The TagRFP-KFP pair and beyond. Biophys. J. 2015, 108, 126–132. [Google Scholar] [CrossRef]
- Li, Y.; Du, H.; Wang, W.; Zhang, P.; Xu, L.; Wen, Y.; Zhang, X. A Versatile Multiple Target Detection System Based on DNA Nano-assembled Linear FRET Arrays. Sci. Rep. 2016, 6, 26879. [Google Scholar] [CrossRef]
- Díaz, S.A.; Buckhout-White, S.; Ancona, M.G.; Spillmann, C.M.; Goldman, E.R.; Melinger, J.S.; Medintz, I.L. Extending DNA-Based Molecular Photonic Wires with Homogeneous Förster Resonance Energy Transfer. Adv. Opt. Mater. 2016, 4, 399–412. [Google Scholar] [CrossRef]
- Choi, Y.; Kotthoff, L.; Olejko, L.; Resch-Genger, U.; Bald, I. DNA Origami-Based Förster Resonance Energy-Transfer Nanoarrays and Their Application as Ratiometric Sensors. ACS Appl. Mater. Interfaces 2018, 10, 23295–23302. [Google Scholar] [CrossRef]
- Xu, P.; Huang, F.; Liang, H. Real-time study of a DNA strand displacement reaction using dual polarization interferometry. Biosens. Bioelectron. 2013, 41, 505–510. [Google Scholar] [CrossRef]
- Laios, A.; O’Toole, S.; Flavin, R.; Martin, C.; Kelly, L.; Ring, M.; Finn, S.P.; Barrett, C.; Loda, M.; Gleeson, N.; et al. Potential role of miR-9 and miR-223 in recurrent ovarian cancer. Mol. Cancer 2008, 7, 35. [Google Scholar] [CrossRef]
- Shen, J.; Ambrosone, C.B.; Zhao, H. Novel genetic variants in microRNA genes and familial breast cancer. Int. J. Cancer 2009, 124, 1178–1182. [Google Scholar] [CrossRef]
- Grossi, G.; Jepsen, M.D.E.; Kjems, J.; Andersen, E.S. Control of enzyme reactions by a reconfigurable DNA nanovault. Nat. Commun. 2017, 8, 992. [Google Scholar] [CrossRef] [PubMed]
- Ijäs, H.; Hakaste, I.; Shen, B.; Kostiainen, M.A.; Linko, V. Reconfigurable DNA Origami Nanocapsule for pH-Controlled Encapsulation and Display of Cargo. ACS Nano 2019, 13, 5959–5967. [Google Scholar] [CrossRef]
- Walter, H.-K.; Bauer, J.; Steinmeyer, J.; Kuzuya, A.; Niemeyer, C.M.; Wagenknecht, H.-A. “DNA Origami Traffic Lights” with a Split Aptamer Sensor for a Bicolor Fluorescence Readout. Nano Lett. 2017, 17, 2467–2472. [Google Scholar] [CrossRef] [PubMed]
- Tang, M.S.L.; Shiu, S.C.-C.; Godonoga, M.; Cheung, Y.-W.; Liang, S.; Dirkzwager, R.M.; Kinghorn, A.B.; Fraser, L.A.; Heddle, J.G.; Tanner, J.A. An aptamer-enabled DNA nanobox for protein sensing. Nanomed. NBM 2018, 14, 1161–1168. [Google Scholar] [CrossRef] [PubMed]
- Choi, Y.; Choi, H.; Lee, A.C.; Lee, H.; Kwon, S. A Reconfigurable DNA Accordion Rack. Angew. Chem. Int. Ed. 2018, 57, 2811–2815. [Google Scholar] [CrossRef] [PubMed]
- Mathur, D.; Henderson, E.R. Programmable DNA Nanosystem for Molecular Interrogation. Sci. Rep. 2016, 6, 27413. [Google Scholar] [CrossRef]
- Marras, A.E.; Shi, Z.; Lindell, M.G.; Patton, R.A.; Huang, C.-M.; Zhou, L.; Su, H.-J.; Arya, G.; Castro, C.E. Cation-Activated Avidity for Rapid Reconfiguration of DNA Nanodevices. ACS Nano 2018, 12, 9484–9494. [Google Scholar] [CrossRef]
- Lund, K.; Manzo, A.J.; Dabby, N.; Michelotti, N.; Johnson-Buck, A.; Nangreave, J.; Taylor, S.; Pei, R.; Stojanovic, M.N.; Walter, N.G.; et al. Molecular robots guided by prescriptive landscapes. Nature 2010, 465, 206–210. [Google Scholar] [CrossRef]
- Hemmig, E.A.; Fitzgerald, C.; Maffeo, C.; Hecker, L.; Ochmann, S.E.; Aksimentiev, A.; Tinnefeld, P.; Keyser, U.F. Optical Voltage Sensing Using DNA Origami. Nano Lett. 2018, 18, 1962–1971. [Google Scholar] [CrossRef]
- Hudoba, M.W.; Luo, Y.; Zacharias, A.; Poirier, M.G.; Castro, C.E. Dynamic DNA Origami Device for Measuring Compressive Depletion Forces. ACS Nano 2017, 11, 6566–6573. [Google Scholar] [CrossRef]
- Fraiji, L.K.; Hayes, D.M.; Werner, T.C. Static and dynamic fluorescence quenching experiments for the physical chemistry laboratory. J. Chem. Educ. 1992, 69. [Google Scholar] [CrossRef]
- Zu, F.; Yan, F.; Bai, Z.; Xu, J.; Wang, Y.; Huang, Y.; Zhou, X. The quenching of the fluorescence of carbon dots: A review on mechanisms and applications. Microchim. Acta 2017, 184, 1899–1914. [Google Scholar] [CrossRef]
- Olley, D.A.; Wren, E.J.; Vamvounis, G.; Fernee, M.J.; Wang, X.; Burn, P.L.; Meredith, P.; Shaw, P.E. Explosive Sensing with Fluorescent Dendrimers: The Role of Collisional Quenching. Chem. Mater. 2011, 23, 789–794. [Google Scholar] [CrossRef]
- Ke, Y.; Meyer, T.; Shih, W.M.; Bellot, G. Regulation at a distance of biomolecular interactions using a DNA origami nanoactuator. Nat. Commun. 2016, 7, 10935. [Google Scholar] [CrossRef] [PubMed]
- Koussa, M.A.; Halvorsen, K.; Ward, A.; Wong, W.P. DNA nanoswitches: A quantitative platform for gel-based biomolecular interaction analysis. Nat. Methods 2015, 12, 123–126. [Google Scholar] [CrossRef]
- Torelli, E.; Marini, M.; Palmano, S.; Piantanida, L.; Polano, C.; Scarpellini, A.; Lazzarino, M.; Firrao, G. A DNA origami nanorobot controlled by nucleic acid hybridization. Small 2014, 10, 2918–2926. [Google Scholar] [CrossRef]
- Horn, T.; Chang, C.A.; Urdea, M.S. Chemical synthesis and characterization of branched oligodeoxyribonucleotides (bDNA) for use as signal amplifiers in nucleic acid quantification assays. Nucleic Acids Res. 1997, 25, 4842–4849. [Google Scholar] [CrossRef][Green Version]
- Ochmann, S.E.; Vietz, C.; Trofymchuk, K.; Acuna, G.P.; Lalkens, B.; Tinnefeld, P. Optical Nanoantenna for Single Molecule-Based Detection of Zika Virus Nucleic Acids without Molecular Multiplication. Anal. Chem. 2017, 89, 13000–13007. [Google Scholar] [CrossRef]
- Khatua, S.; Paulo, P.M.R.; Yuan, H.; Gupta, A.; Zijlstra, P.; Orrit, M. Resonant Plasmonic Enhancement of Single-Molecule Fluorescence by Individual Gold Nanorods. ACS Nano 2014, 8, 4440–4449. [Google Scholar] [CrossRef]
- Bidault, S.; Devilez, A.; Maillard, V.; Lermusiaux, L.; Guigner, J.-M.; Bonod, N.; Wenger, J. Picosecond Lifetimes with High Quantum Yields from Single-Photon-Emitting Colloidal Nanostructures at Room Temperature. ACS Nano 2016, 10, 4806–4815. [Google Scholar] [CrossRef]
- Yehl, K.; Mugler, A.; Vivek, S.; Liu, Y.; Zhang, Y.; Fan, M.; Weeks, E.R.; Salaita, K. High-speed DNA-based rolling motors powered by RNase H. Nat. Nanotech. 2016, 11, 184–190. [Google Scholar] [CrossRef] [PubMed]
- Del Grosso, E.; Dallaire, A.-M.; Vallée-Bélisle, A.; Ricci, F. Enzyme-Operated DNA-Based Nanodevices. Nano Lett. 2015, 15, 8407–8411. [Google Scholar] [CrossRef] [PubMed]
- Masoud, R.; Tsukanov, R.; Tomov, T.E.; Plavner, N.; Liber, M.; Nir, E. Studying the Structural Dynamics of Bipedal DNA Motors with Single-Molecule Fluorescence Spectroscopy. ACS Nano 2012, 6, 6272–6283. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Vietz, C.; Schroder, T.; Acuna, G.; Lalkens, B.; Tinnefeld, P. A DNA Walker as a Fluorescence Signal Amplifier. Nano Lett. 2017, 17, 5368–5374. [Google Scholar] [CrossRef]
- Wickham, S.F.; Bath, J.; Katsuda, Y.; Endo, M.; Hidaka, K.; Sugiyama, H.; Turberfield, A.J. A DNA-based molecular motor that can navigate a network of tracks. Nat. Nanotech. 2012, 7, 169–173. [Google Scholar] [CrossRef]
- Kroener, F.; Heerwig, A.; Kaiser, W.; Mertig, M.; Rant, U. Electrical Actuation of a DNA Origami Nanolever on an Electrode. J. Am. Chem. Soc. 2017, 139, 16510–16513. [Google Scholar] [CrossRef]
- Kroener, F.; Traxler, L.; Heerwig, A.; Rant, U.; Mertig, M. Magnesium-Dependent Electrical Actuation and Stability of DNA Origami Rods. ACS Appl. Mater. Interfaces 2019, 11, 2295–2301. [Google Scholar] [CrossRef]
- Jain, P.K.; Huang, W.; El-Sayed, M.A. On the Universal Scaling Behavior of the Distance Decay of Plasmon Coupling in Metal Nanoparticle Pairs: A Plasmon Ruler Equation. Nano Lett. 2007, 7, 2080–2088. [Google Scholar] [CrossRef]
- Jensen, T.R.; Malinsky, M.D.; Haynes, C.L.; Van Duyne, R.P. Nanosphere Lithography: Tunable Localized Surface Plasmon Resonance Spectra of Silver Nanoparticles. J. Phys. Chem. B 2000, 104, 10549–10556. [Google Scholar] [CrossRef]
- Kelly, K.L.; Coronado, E.; Zhao, L.L.; Schatz, G.C. The Optical Properties of Metal Nanoparticles: The Influence of Size, Shape, and Dielectric Environment. J. Phys. Chem. B 2003, 107, 668–677. [Google Scholar] [CrossRef]
- Kneipp, K.; Kneipp, H.; Itzkan, I.; Dasari, R.R.; Feld, M.S. Ultrasensitive chemical analysis by Raman spectroscopy. Chem. Rev. 1999, 99, 2957–2976. [Google Scholar] [CrossRef] [PubMed]
- Le Ru, E.C.; Etchegoin, P.G. Chapter 4—SERS enhancement factors and related topics. In Principles of Surface-Enhanced Raman Spectroscopy, 1st ed.; Le Ru, E.C., Etchegoin, P.G., Eds.; Elsevier: Amsterdam, The Netherlands, 2009; pp. 185–264. [Google Scholar]
- Thacker, V.V.; Herrmann, L.O.; Sigle, D.O.; Zhang, T.; Liedl, T.; Baumberg, J.J.; Keyser, U.F. DNA origami based assembly of gold nanoparticle dimers for surface-enhanced Raman scattering. Nat. Commun. 2014, 5. [Google Scholar] [CrossRef] [PubMed]
- Kühler, P.; Roller, E.-M.; Schreiber, R.; Liedl, T.; Lohmüller, T.; Feldmann, J. Plasmonic DNA-Origami Nanoantennas for Surface-Enhanced Raman Spectroscopy. Nano Lett. 2014, 14, 2914–2919. [Google Scholar] [CrossRef] [PubMed]
- Tanwar, S.; Haldar, K.K.; Sen, T. DNA Origami Directed Au Nanostar Dimers for Single-Molecule Surface-Enhanced Raman Scattering. J. Am. Chem. Soc. 2017, 139, 17639–17648. [Google Scholar] [CrossRef] [PubMed]
- Zhan, P.; Wen, T.; Wang, Z.; He, Y.; Shi, J.; Wang, T.; Liu, X.; Lu, G.; Ding, B. DNA Origami Directed Assembly of Gold Bowtie Nanoantennas for Single-Molecule Surface-Enhanced Raman Scattering. Angew. Chem. Int. Ed. 2018, 57, 2846–2850. [Google Scholar] [CrossRef] [PubMed]
- Prinz, J.; Matković, A.; Pešić, J.; Gajić, R.; Bald, I. Hybrid Structures for Surface-Enhanced Raman Scattering: DNA Origami/Gold Nanoparticle Dimer/Graphene. Small 2016, 12, 5458–5467. [Google Scholar] [CrossRef] [PubMed]
- Prinz, J.; Heck, C.; Ellerik, L.; Merk, V.; Bald, I. DNA origami based Au–Ag-core–shell nanoparticle dimers with single-molecule SERS sensitivity. Nanoscale 2016, 8, 5612–5620. [Google Scholar] [CrossRef]
- Moeinian, A.; Gür, F.N.; Gonzalez-Torres, J.; Zhou, L.; Murugesan, V.D.; Dashtestani, A.D.; Guo, H.; Schmidt, T.L.; Strehle, S. Highly Localized SERS Measurements Using Single Silicon Nanowires Decorated with DNA Origami-Based SERS Probe. Nano Lett. 2019, 19, 1061–1066. [Google Scholar] [CrossRef]
- Heck, C.; Prinz, J.; Dathe, A.; Merk, V.; Stranik, O.; Fritzsche, W.; Kneipp, J.; Bald, I. Gold Nanolenses Self-Assembled by DNA Origami. ACS Photonics 2017, 4, 1123–1130. [Google Scholar] [CrossRef]
- Pilo-Pais, M.; Watson, A.; Demers, S.; LaBean, T.H.; Finkelstein, G. Surface-Enhanced Raman Scattering Plasmonic Enhancement Using DNA Origami-Based Complex Metallic Nanostructures. Nano Lett. 2014, 14, 2099–2104. [Google Scholar] [CrossRef] [PubMed]
- Heck, C.; Kanehira, Y.; Kneipp, J.; Bald, I. Placement of Single Proteins within the SERS Hot Spots of Self-Assembled Silver Nanolenses. Angew. Chem. Int. Ed. 2018, 57, 7444–7447. [Google Scholar] [CrossRef]
- Putz, A.-M.; Putz, M.V. Spectral inverse quantum (Spectral-IQ) method for modeling mesoporous systems: Application on silica films by FTIR. Int. J. Mol. Sci. 2012, 13, 15925–15941. [Google Scholar] [CrossRef]
- Fan, Z.; Govorov, A.O. Plasmonic Circular Dichroism of Chiral Metal Nanoparticle Assemblies. Nano Lett. 2010, 10, 2580–2587. [Google Scholar] [CrossRef]
- Schreiber, R. Plasmonic DNA Nanostructures with Tailored Optical Response; Ludwig-Maximilians-Universität München: München, Germany, 2013. [Google Scholar]
- Kuzyk, A.; Schreiber, R.; Fan, Z.; Pardatscher, G.; Roller, E.-M.; Högele, A.; Simmel, F.C.; Govorov, A.O.; Liedl, T. DNA-based self-assembly of chiral plasmonic nanostructures with tailored optical response. Nature 2012, 483, 311–314. [Google Scholar] [CrossRef]
- Schreiber, R.; Luong, N.; Fan, Z.; Kuzyk, A.; Nickels, P.C.; Zhang, T.; Smith, D.M.; Yurke, B.; Kuang, W.; Govorov, A.O.; et al. Chiral plasmonic DNA nanostructures with switchable circular dichroism. Nat. Commun. 2013, 4, 2948. [Google Scholar] [CrossRef]
- Lieberman, I.; Shemer, G.; Fried, T.; Kosower, E.M.; Markovich, G. Plasmon-Resonance-Enhanced Absorption and Circular Dichroism. Angew. Chem. Int. Ed. 2008, 47, 4855–4857. [Google Scholar] [CrossRef]
- Kneer, L.M.; Roller, E.-M.; Besteiro, L.V.; Schreiber, R.; Govorov, A.O.; Liedl, T. Circular Dichroism of Chiral Molecules in DNA-Assembled Plasmonic Hotspots. ACS Nano 2018, 12, 9110–9115. [Google Scholar] [CrossRef]
- Shen, X.; Song, C.; Wang, J.; Shi, D.; Wang, Z.; Liu, N.; Ding, B. Rolling Up Gold Nanoparticle-Dressed DNA Origami into Three-Dimensional Plasmonic Chiral Nanostructures. J. Am. Chem. Soc. 2012, 134, 146–149. [Google Scholar] [CrossRef]
- Urban, M.J.; Dutta, P.K.; Wang, P.; Duan, X.; Shen, X.; Ding, B.; Ke, Y.; Liu, N. Plasmonic Toroidal Metamolecules Assembled by DNA Origami. J. Am. Chem. Soc. 2016, 138, 5495–5498. [Google Scholar] [CrossRef]
- Man, T.; Ji, W.; Liu, X.; Zhang, C.; Li, L.; Pei, H.; Fan, C. Chiral Metamolecules with Active Plasmonic Transition. ACS Nano 2019, 13, 4826–4833. [Google Scholar] [CrossRef]
- Lan, X.; Liu, T.; Wang, Z.; Govorov, A.O.; Yan, H.; Liu, Y. DNA-Guided Plasmonic Helix with Switchable Chirality. J. Am. Chem. Soc. 2018, 140, 11763–11770. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Nguyen, M.K.; Natarajan, A.K.; Nguyen, V.H.; Kuzyk, A. A DNA Origami-Based Chiral Plasmonic Sensing Device. ACS Appl. Mater. Interfaces 2018, 10, 44221–44225. [Google Scholar] [CrossRef] [PubMed]
- Zhou, C.; Xin, L.; Duan, X.; Urban, M.J.; Liu, N. Dynamic Plasmonic System That Responds to Thermal and Aptamer-Target Regulations. Nano Lett. 2018, 18, 7395–7399. [Google Scholar] [CrossRef] [PubMed]
- Kuzyk, A.; Schreiber, R.; Zhang, H.; Govorov, A.O.; Liedl, T.; Liu, N. Reconfigurable 3D plasmonic metamolecules. Nat. Mater. 2014, 13, 862–866. [Google Scholar] [CrossRef]
- Kuzyk, A.; Yang, Y.; Duan, X.; Stoll, S.; Govorov, A.O.; Sugiyama, H.; Endo, M.; Liu, N. A light-driven three-dimensional plasmonic nanosystem that translates molecular motion into reversible chiroptical function. Nat. Commun. 2016, 7, 10591. [Google Scholar] [CrossRef]
- Wu, Y.; Tilley, R.D.; Gooding, J.J. Challenges and Solutions in Developing Ultrasensitive Biosensors. J. Am. Chem. Soc. 2019, 141, 1162–1170. [Google Scholar] [CrossRef]
- Putz, M.V. On the reducible character of Haldane-Radić enzyme kinetics to conventional and logistic Michaelis-Menten models. Molecules 2011, 16, 3128–3145. [Google Scholar] [CrossRef]
- Anastassacos, F.M.; Zhao, Z.; Zeng, Y.; Shih, W.M. Glutaraldehyde Cross-Linking of Oligolysines Coating DNA Origami Greatly Reduces Susceptibility to Nuclease Degradation. J. Am. Chem. Soc. 2020, 142, 3311–3315. [Google Scholar] [CrossRef]
- Kopatz, I.; Zalk, R.; Levi-Kalisman, Y.; Zlotkin-Rivkin, E.; Frank, G.A.; Kler, S. Packaging of DNA origami in viral capsids. Nanoscale 2019, 11, 10160–10166. [Google Scholar] [CrossRef]
- Lee, A.J.; Walti, C. DNA nanostructures: A versatile lab-bench for interrogating biological reactions. Comput. Struct. Biotechnol. J. 2019, 17, 832–842. [Google Scholar] [CrossRef]
- Wang, W.; Yu, S.; Huang, S.; Bi, S.; Han, H.; Zhang, J.-R.; Lu, Y.; Zhu, J.-J. Bioapplications of DNA nanotechnology at the solid–liquid interface. Chem. Soc. Rev. 2019, 48, 4892–4920. [Google Scholar] [CrossRef]
- Zheng, Y.; Soeriyadi, A.H.; Rosa, L.; Ng, S.H.; Bach, U.; Justin Gooding, J. Reversible gating of smart plasmonic molecular traps using thermoresponsive polymers for single-molecule detection. Nat. Commun. 2015, 6, 8797. [Google Scholar] [CrossRef]
- Praetorius, F.; Kick, B.; Behler, K.L.; Honemann, M.N.; Weuster-Botz, D.; Dietz, H. Biotechnological mass production of DNA origami. Nature 2017, 552, 84–87. [Google Scholar] [CrossRef]
- Keller, A.; Linko, V. Challenges and Perspectives of DNA Nanostructures in Biomedicine. Angew. Chem. Int. Ed. 2020. [Google Scholar] [CrossRef]
- Shen, B.; Linko, V.; Tapio, K.; Pikker, S.; Lemma, T.; Gopinath, A.; Gothelf, K.V.; Kostiainen, M.A.; Toppari, J.J. Plasmonic nanostructures through DNA-assisted lithography. Sci. Adv. 2018, 4, eaap8978. [Google Scholar] [CrossRef]
- Gopinath, A.; Miyazono, E.; Faraon, A.; Rothemund, P.W.K. Engineering and mapping nanocavity emission via precision placement of DNA origami. Nature 2016, 535, 401–405. [Google Scholar] [CrossRef]
- Xin, Y.; Kielar, C.; Zhu, S.; Sikeler, C.; Xu, X.; Moser, C.; Grundmeier, G.; Liedl, T.; Heuer-Jungemann, A.; Smith, D.M.; et al. Cryopreservation of DNA Origami Nanostructures. Small 2020, 16, 1905959. [Google Scholar] [CrossRef]
| Signaling Procedure | Analyte | Sensitivity | Response Time | Publication Year | Reference |
|---|---|---|---|---|---|
| *Fluorescence enhancement | Single dye with 200 mM of NiCl2 | 25 μM BG fluorophores | Second range | 2015 | [73] |
| Change in pH (pH-sensitive dyes) | Intercellular pH | High sensitivity for pH from 4 to 8 | 30 min (pH change 6.4–5.6) | 2015 | [70] |
| Exchange of dye with target | Antibodies (systemic lupus erythematosus) | 10× higher (than ELISA) | 1.5 h | 2017 | [69] |
| Exchange of dye with target | Antibodies (systemic lupus erythematosus) | 7% false positives (lower than ELISA) | 1.5 h | 2018 | [62] |
| Enzyme cleaving | Peanut antigen Ara h1 | 232 fM (Aptamer) 18 fM (Aptamer + DNA-OS) | 3 h 30 min | 2019 | [54] |
| Aptamer binding + labeling dyes | Thrombin | 2 ± 0.2 nM (Aptamer + DNA-OS) 22 ± 3 nM (aptamer) | 25 min (fluorescence record) | 2020 | [43] |
| *Fluorescence enhancement | ssDNA (Oxa-48) | 2 nM | 2 h (incubation) | 2020 | [72] |
| Signaling Procedure | Analyte | Sensitivity | Response Time | Publication Year | Reference |
|---|---|---|---|---|---|
| O/C (strand displacement) | “Key”ssDNA (POP) | 200 μM | 40 s | 2009 | [18] |
| *Enzyme-assisted movement | ssDNA + Zn2+ | 1:1 (molar ratio complex: ssDNA) 1 mM (Zn2+) | 3 nm/min (50 cleavage steps) | 2010 | [91] |
| O/C (strand displacement) | ssDNA | 0.4 μM (10× molar excess) | Min range (1st opening) 4 h (2nd reopening) | 2012 | [24] |
| *Conformation Change | MgCl2, Temperature (POP) | 5–25 mM (MgCl2 concentration range) 11 to 47 °C (during O/C) | - | 2015 | [48] |
| Change of telomeric DNA into guanine quadruplexes | Na+, K+ | 1 mM K+, 25 mM NaCl | - | 2016 | [55] |
| Conformation change (strand displacement + adhesive force) | ssDNA (POP) | - | - | 2016 | [89] |
| O/C (strand displacement) | ssDNA (POP) | 1:1.3 molar excess (DV + closing lock) 1:1.5 molar excess (DV + opening lock) | 15 min | 2017 | [84] |
| O/C (split aptamer) | ATP | 0.10–1.00 mM (Range of sensitivity) | 15–25 min (Observation of the fluorescence) | 2017 | [86] |
| *Interaction with environment | Depletion force | ~100 fN (Resolution) | ms range (Unspecified) | 2017 | [93] |
| *Electric potential change | Optical voltage change | 200 mV (Minimum before to be sensitive) | ∼50 s | 2018 | [92] |
| *O/C (strand displacement) | ssDNA | 10–100 pM | 100 s (Efficiency with t1/2) | 2018 | [21] |
| *O/C by environment change | Cation | ∼200–1000 mM (Monovalent ions) ∼5–40 mM (Divalent ions) ∼0.06–0.14 mM (Trivalent ions) | ≤ 200 ms (O/C Transitions) | 2018 | [90] |
| Conformation change (strand displacement) | ssDNA | POP | POP | 2018 | [88] |
| O/C (aptamer) | PfLDH (protein) | 100 nM | 0–20 min | 2018 | [87] |
| Change in pH (pH sensitive dyes) | pH | 6–8 (pH range to be sensitive) | - | 2018 | [80] |
| O/C (pH-latches) | pH | 0.5 pH | 30 s (opening) Hours (closing) | 2019 | [85] |
| Signaling Procedure | Analyte | Sensitivity | Response Time | Publication Year | Reference |
|---|---|---|---|---|---|
| Hybridization of target | ssDNA | 20 pmol | 1 h (incubation) | 2014 | [99] |
| O/C (change in environment) | K+, miR-210 (miRNA) BamHI (Enzyme) | 100 mM KCl 1 equivalent miR-210 (equivalents to the locking strand) 5 units of BamHI | 2 h (miRNA) 10 min (BamHI) | 2016 | [97] |
| *O/C (hairpin + optical antenna) | ssDNA | POP | POP | 2017 | [34] |
| *O/C (hairpin + optical antenna) | Zika DNA/RNA | 1 nM (RNA and DNA) | 18 h | 2017 | [101] |
| *Enzyme-assisted moment of complex (molecular motors) | ssDNA (with mismatches) | 0, 1, 2, or 3 mismatches | 2 h (no Mismatch) + 4 h (with Mismatch) | 2017 | [107] |
| Signaling Procedure | Analyte | Enhancement Factor | Response Time | Publication Year | Reference |
|---|---|---|---|---|---|
| Hotspot (two AuNPs) | TAMRA | - | - | 2013 | [36] |
| Hotspot (two AuNPs) | SYBR gold (25 dyes) | 1.4 × 105 | - | 2014 | [117] |
| *Hotspot (two AuNPs) | Rhodamine 6G, ssDNA | 107 (dye) 105 (ssDNA) | - | 2014 | [116] |
| Hotspot (four AuNPs) | aminobenzenethiol (4-ABT) | 102/nanoparticle | - | 2014 | [124] |
| *Hotspot (two AuNPs) | Single Cy3.5 | 102 (for gap 1.4 nm vs. 2.5 nm) | - | 2016 | [53] |
| *Hotspot (two AuNPs) | TAMRA and Cy3 | 1010 | - | 2016 | [121] |
| Hotspot (two AuNPs + graphene) | TAMRA | - | - | 2016 | [120] |
| *Hotspot (two gold nanostars) | Single Texas red | 2.0 × 1010 (particles gap of 7 nm) 8.0 × 109 (particles gap of 13 nm) | - | 2017 | [118] |
| Hotspot (gold nanolenses) | TAMRA | 1.4 × 106 | - | 2017 | [123] |
| *Hotspot (two gold nanoprisms) | Cy5 and Cy3 | 109 to 1010 | - | 2018 | [119] |
| Hotspot (silver nanolenses) | Streptavidin | 101 (in blue region, vs. gold nanolenses) 4.0 × 100 (at 532 nm, vs. gold nanolenses) | - | 2018 | [125] |
| Single silicon nanowire | Methylene blue | 1.1 × 105 | - | 2019 | [122] |
| Signaling Procedure | Analyte | Sensitivity | Response Time | Publication Year | Reference |
|---|---|---|---|---|---|
| Measurement of CD signal | POP | - | - | 2012 | [133] |
| Measurement of CD signal | POP | - | - | 2012 | [129] |
| O/C structures | Fuel | 70 nM (first cycle) | - | 2014 | [139] |
| RH, LH structures proportion | pH | Range depends on the percentage of LH or RH molecules | Few minutes | 2017 | [20] |
| O/C structure | Viral RNA (Hepatitis C virus) | 100 pM (Buffer) 1 nM (Human serum) | 30 min (incubation) 0.01 s (each wavelength) | 2018 | [19] |
| O/C structure | Adenosine | 20 µM (Lock style 1) 65 µM (Lock style 2) | 1 min (Lock style 1) | 2018 | [137] |
| O/C structure | ATP and Cocaine | mM to µM range (ATP or Cocaine) | - | 2018 | [138] |
| O/C structure | Human α-thrombin | 100 pM | - | 2019 | [52] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Loretan, M.; Domljanovic, I.; Lakatos, M.; Rüegg, C.; Acuna, G.P. DNA Origami as Emerging Technology for the Engineering of Fluorescent and Plasmonic-Based Biosensors. Materials 2020, 13, 2185. https://doi.org/10.3390/ma13092185
Loretan M, Domljanovic I, Lakatos M, Rüegg C, Acuna GP. DNA Origami as Emerging Technology for the Engineering of Fluorescent and Plasmonic-Based Biosensors. Materials. 2020; 13(9):2185. https://doi.org/10.3390/ma13092185
Chicago/Turabian StyleLoretan, Morgane, Ivana Domljanovic, Mathias Lakatos, Curzio Rüegg, and Guillermo P. Acuna. 2020. "DNA Origami as Emerging Technology for the Engineering of Fluorescent and Plasmonic-Based Biosensors" Materials 13, no. 9: 2185. https://doi.org/10.3390/ma13092185
APA StyleLoretan, M., Domljanovic, I., Lakatos, M., Rüegg, C., & Acuna, G. P. (2020). DNA Origami as Emerging Technology for the Engineering of Fluorescent and Plasmonic-Based Biosensors. Materials, 13(9), 2185. https://doi.org/10.3390/ma13092185






