Decellularized Extracellular Matrix for Cancer Research
Abstract
1. Introduction
2. dECM as Reconstituted Native ECM In Vitro
2.1. ECM Composition and Structure
2.2. ECM in Cancer Pathology
2.3. Decellularized ECM (dECM)
2.3.1. Comparison of Sources for dECM
2.3.2. Preparation
2.3.3. Characterization
3. Examples of dECM Utilized for Cancer Research
3.1. Cell Behaviors in/on dECM as ECM Models at Primary Sites
3.1.1. dECM Derived from Normal Tissues
3.1.2. dECM Derived from Cancer Tissues
3.1.3. Comparison of Cell Behavior in dECM Derived from Normal and Cancer Tissues
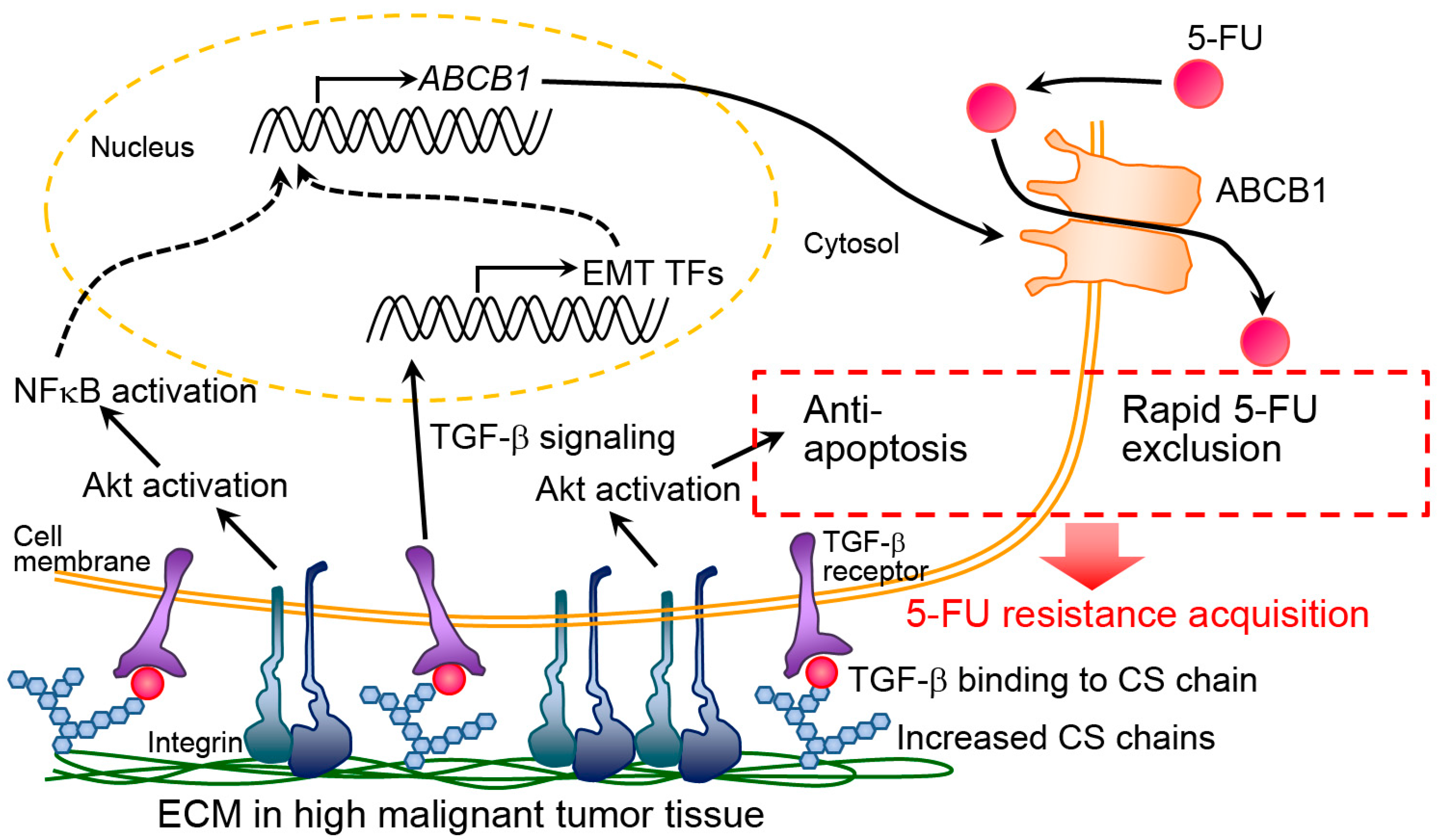
3.2. Mechanism Analysis of Chemoresistance
3.3. Cancer Cell Colonization at Metastatic Sites
4. Future Perspectives
4.1. Availability of dECM as In Vitro ECM Models in Cancer
4.2. Proper Selection of dECM Preparation Methods
4.3. Possibility of Contributions to Cancer Therapies
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer: The next generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. Hallmarks of cancer. Cell 2000, 100, 57–70. [Google Scholar] [CrossRef]
- King, T.D.; Suto, M.J.; Li, Y. The Wnt/β-Catenin Signaling Pathway: A Potential Therapeutic Target in the Treatment of Triple Negative Breast Cancer. J. Cell. Biochem. 2012, 113, 13–18. [Google Scholar] [CrossRef]
- Zheng, G.; Xiong, Y.; Yi, S.; Zhang, W.; Peng, B.; Zhang, Q.; He, Z. 14-3-3σ regulation by p53 mediates a chemotherapy response to 5-fluorouracil in MCF-7 breast cancer cells via Akt inactivation. FEBS Lett. 2012, 586, 163–168. [Google Scholar] [CrossRef]
- Longley, D.B.; Harkin, D.P.; Johnston, P.G. 5-Fluorouracil: Mechanisms of action and clinical strategies. Nat. Rev. Cancer 2003, 3, 330–338. [Google Scholar] [CrossRef] [PubMed]
- Ghajar, C.M.; Bissell, M.J. Tumor engineering: The other face of tissue engineering. Tissue Eng. Part A 2010, 16, 2153–2156. [Google Scholar] [CrossRef] [PubMed]
- Ghajar, C.M.; Bissell, M.J. Extracellular matrix control of mammary gland morphogenesis and tumorigenesis: Insights from imaging. Histochem. Cell Biol. 2008, 130, 1105–1118. [Google Scholar] [CrossRef]
- Lu, P.; Weaver, V.M.; Werb, Z. The extracellular matrix: A dynamic niche in cancer progression. J. Cell Biol. 2012, 196, 395–406. [Google Scholar] [CrossRef]
- Hynes, R.O.; Naba, A. Overview of the Matrisome-An Inventory of Extracellular Matrix Constituents and Functions. Cold Spring Harb. Perspect. Biol. 2012, 4, a004903. [Google Scholar] [CrossRef] [PubMed]
- Geiger, B.; Yamada, K.M. Molecular Architecture and Function of Matrix Adhesions. Cold Spring Harb. Perspect. Biol. 2011, 3, a005003. [Google Scholar] [CrossRef]
- Hynes, R.O. Integrins: Bidirectional, allosteric signaling machines. Cell 2002, 110, 673–687. [Google Scholar] [CrossRef]
- Harburger, D.S.; Calderwood, D.A. Integrin signaling at a glance. J. Cell Sci. 2009, 122, 159–163. [Google Scholar] [CrossRef] [PubMed]
- Rathinam, R.; Alahari, S.K. Important role of integrins in the cancer biology. Cancer Metastasis Rev. 2010, 29, 223–237. [Google Scholar] [CrossRef]
- Chia, J.; Kusuma, N.; Anderson, R.; Parker, B.; Bidwell, B.; Zamurs, L.; Nice, E.; Pouliot, N. Evidence for a Role of Tumor-Derived Laminin-511 in the Metastatic Progression of Breast Cancer. Am. J. Pathol. 2007, 170, 2135–2148. [Google Scholar] [CrossRef] [PubMed]
- Carpenter, P.M.; Dao, A.V.; Arain, Z.S.; Chang, M.K.; Nguyen, H.P.; Arain, S.; Wang-Rodriguez, J.; Kwon, S.-Y.; Wilczynski, S.P. Motility Induction in Breast Carcinoma by Mammary Epithelial Laminin 332 (Laminin 5). Mol. Cancer Res. 2009, 7, 462–475. [Google Scholar] [CrossRef] [PubMed]
- O’Connell, J.T.; Sugimoto, H.; Cooke, V.G.; MacDonald, B.A.; Mehta, A.I.; LeBleu, V.S.; Dewar, R.; Rocha, R.M.; Brentani, R.R.; Resnick, M.B.; et al. VEGF-A and Tenascin-C produced by S100A4+ stromal cells are important for metastatic colonization. Proc. Natl. Acad. Sci. U.S.A. 2011, 108, 16002–16007. [Google Scholar] [CrossRef]
- Uhm, J.H.; Dooley, N.P.; Kyritsis, A.P.; Rao, J.S.; Gladson, C.L. Vitronectin, a Glioma-derived Extracellular Matrix Protein, Protects Tumor Cells from Apoptotic Death. Clin. Cancer Res. 1999, 5, 1587–1594. [Google Scholar]
- Manabe, R.; Tsutsui, K.; Yamada, T.; Kimura, M.; Nakano, I.; Shimono, C.; Sanzen, N.; Furutani, Y.; Fukuda, T.; Oguri, Y.; et al. Transcriptome-based systematic identification of extracellular matrix proteins. Proc. Natl. Acad. Sci. U.S.A. 2008, 105, 12849–12854. [Google Scholar] [CrossRef]
- Hoshiba, T.; Lu, H.; Kawazoe, N.; Chen, G. Decellularized matrices for tissue engineering. Expert Opin. Biol. Ther. 2010, 10, 1717–1728. [Google Scholar] [CrossRef]
- Hoshiba, T.; Chen, G.; Endo, C.; Maruyama, H.; Wakui, M.; Nemoto, E.; Kawazoe, N.; Tanaka, M. Decellularized extracellular matrix (ECM) as an in vitro model to study the comprehensive roles of the ECM in stem cell differentiation. Stem Cells Int. 2016, 2016, 6397820. [Google Scholar] [CrossRef]
- Hoshiba, T. Cultured cell-derived decellularized matrices: A review toward the next decade. J. Mater. Chem. B 2017, 5, 4322–4331. [Google Scholar] [CrossRef]
- Badylak, S.F. The extracellular matrix as a biologic scaffold material. Biomaterials 2007, 28, 3587–3593. [Google Scholar] [CrossRef]
- Taylor, D.A.; Sampaio, L.C.; Ferdous, Z.; Gobin, A.S.; Taite, L.J. Decellularized matrices in regenerative medicine. Acta Biomater. 2018, 74, 74–89. [Google Scholar] [CrossRef]
- Naba, A.; Clauser, K.R.; Hoersch, S.; Liu, H.; Carr, S.A.; Hynes, R.O. The Matrisome: In Silico Definition and In Vivo Characterization by Proteomics of Normal and Tumor Extracellular Matrices. Mol. Cell. Proteomics 2012, 11, M111.014647. [Google Scholar] [CrossRef] [PubMed]
- Hayashi, T.; Mizuno, K. Collagen. In Encyclopedia of Molecular Biology; Creighton, T.E., Ed.; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 1999; pp. 500–511. [Google Scholar]
- Hynes, R.O. The Extracellular Matrix: Not Just Pretty Fibrils. Science 2009, 326, 1216–1219. [Google Scholar] [CrossRef]
- Chen, X.-D.; Fisher, L.W.; Robey, P.G.; Young, M.F. The small leucine-rich proteoglycan biglycan modulates BMP-4-induced osteoblast differentiation. FASEB J. 2004, 18, 948–958. [Google Scholar] [CrossRef] [PubMed]
- Colognato, H.; Yurchenco, P.D. Form and Function: The Laminin Family of Heterotrimers. Dev. Dyn. 2000, 218, 213–234. [Google Scholar] [CrossRef]
- Daley, W.P.; Peters, S.B.; Larsen, M. Extracellular matrix dynamics in development and regenerative medicine. J. Cell Sci. 2008, 121, 255–264. [Google Scholar] [CrossRef]
- Page-McCaw, A.; Ewald, A.J.; Werb, Z. Matrix metalloproteinases and the regulation of tissue remodeling. Nat. Rev. Mol. Cell. Biol. 2007, 8, 221–233. [Google Scholar] [CrossRef]
- Ioachim, E.; Charchanti, A.; Briasoulis, E.; Karavasilis, V.; Tsanou, H.; Arvanitis, D.L.; Agnantis, N.J.; Pavidis, N. Immunohistochemical expression of extracellular matrix components tenascin, fibronectin, collagen type IV and laminin in breast cancer: Their prognostic value and role in tumour invasion and progression. Eur. J. Cancer. 2002, 38, 2362–2370. [Google Scholar] [CrossRef]
- Kharaishvili, G.; Cizkova, M.; Bouchalova, K.; Mgebrishvili, G.; Kolar, Z.; Bouchal, J. Collagen triple helix repeat containing 1 protein, periostin and versican in primary and metastatic breast cancer: An immunohistochemical study. J. Clin. Pathol. 2011, 64, 977–982. [Google Scholar] [CrossRef]
- Ricciardelli, C.; Quinn, D.I.; Raymond, W.A.; McCaul, K.; Sutherland, P.D.; Stricker, P.D.; Grygiel, J.J.; Sutherland, R.L.; Marshall, V.R.; Tilley, W.D.; et al. Elevated levels of peritumoral chondroitin sulfate are predictive of poor prognosis in patients treated by radical prostatectomy for early-stage prostate cancer. Cancer Res. 1999, 59, 2324–2328. [Google Scholar]
- Kiewe, P.; Bechrakis, N.E.; Schmittel, A.; Ruf, P.; Lindhofer, H.; Thiel, E.; Nagorsen, D. Increased chondroitin sulphate proteoglycan expression (B5 immunoreactivity) in metastases of uveal melanoma. Ann. Oncol. 2006, 17, 1830–1834. [Google Scholar] [CrossRef]
- Soucy, P.A.; Werbin, J.; Heinz, W.; Hoh, J.H.; Romer, L.H. Microelastic properties of lung cell-derived extracellular matrix. Acta Biomater. 2011, 7, 96–105. [Google Scholar] [CrossRef] [PubMed]
- Satyam, A.; Kumar, P.; Fan, X.; Gorelov, A.; Rochev, Y.; Joshi, L.; Peinado, H.; Lyden, D.; Thomas, B.; Rodriguez, B.; et al. Macromolecular crowding meets tissue engineering by self-assembly: A paradigm shift in regenerative medicine. Adv. Mater. 2014, 26, 3024–3034. [Google Scholar] [CrossRef]
- Furuyama, A.; Kimata, K.; Mochitate, K. Assembly of basement membrane in vitro by cooperation between alveolar epithelial cells and pulmonary fibroblasts. Cell Struct. Funct. 1997, 22, 603–614. [Google Scholar] [CrossRef]
- Prewitz, M.C.; Seib, F.P.; von Bonin, M.; Friedrichs, J.; Stißel, A.; Niehage, C.; Müller, K.; Anastassiadis, K.; Waskow, C.; Hoflack, B.; et al. Tightly anchored tissue-mimetic matrices as instructive stem cell microenvironments. Nat. Methods 2013, 10, 788–794. [Google Scholar] [CrossRef] [PubMed]
- Hoshiba, T.; Tanaka, M. Optimization of the tissue source, malignancy, and initial substrate of tumor cell-derived matrices to increase cancer cell chemoresistance against 5-fluorouracil. Biochem. Biophys. Res. Commun. 2015, 457, 353–357. [Google Scholar] [CrossRef]
- Hoshiba, T.; Lu, H.; Yamada, T.; Kawazoe, N.; Tateishi, T.; Chen, G. Effects of Extracellular Matrices Derived from Different Cell Sources on Chondrocyte Functions. Biotechnol. Prog. 2011, 27, 788–795. [Google Scholar] [CrossRef]
- Hoshiba, T.; Yamada, T.; Lu, H.; Kawazoe, N.; Chen, G. Maintenance of cartilaginous gene expression on extracellular matrix derived from serially passaged chondrocytes during in vitro chondrocyte expansion. J. Biomed. Mater. Res. Part A 2012, 100, 694–702. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.D.; Dusevich, V.; Feng, J.Q.; Manolagas, S.C.; Jilka, R.L. Extracellular matrix made by bone marrow cells facilitates expansion of marrow-derived mesenchymal progenitor cells and prevents their differentiation into osteoblasts. J. Bone Miner. Res. 2007, 22, 1943–1956. [Google Scholar] [CrossRef]
- Hoshiba, T.; Sugano, Y.; Yokoyama, N. Murine neural stem cell (NSC) niche line, MEB5-derived decellularized matrix as an in vitro extracellular matrix model in NSC niche. Chem. Lett. 2018, 47, 1498–1501. [Google Scholar] [CrossRef]
- Gilbert, T.W.; Sellaro, T.L.; Badylak, S.F. Decellularization of tissues and organs. Biomaterials 2006, 27, 3675–3683. [Google Scholar] [CrossRef] [PubMed]
- Keane, T.J.; Swinehart, I.T.; Badylak, S.F. Methods of tissue decellularization used for preparation of biologic scaffolds and in vivo relevance. Methods 2015, 84, 25–34. [Google Scholar] [CrossRef]
- Crapo, P.M.; Gilbert, T.W.; Badylak, S.F. An overview of tissue and whole organ decellularization processes. Biomaterials 2011, 32, 3233–3243. [Google Scholar] [CrossRef] [PubMed]
- Lu, H.; Hoshiba, T.; Kawazoe, N.; Chen, G. Autologous extracellular matrix scaffolds for tissue engineering. Biomaterials 2011, 32, 2489–2499. [Google Scholar] [CrossRef]
- Kim, I.G.; Gil, C.-H.; Seo, J.; Park, S.-J.; Subbiah, R.; Jung, T.-H.; Kim, J.S.; Jeong, Y.-H.; Chung, H.-M.; Lee, J.H.; et al. Mechanotransduction of human pluripotent stem cells culticated on tunable cell-derived extracellular matrix. Biomaterials 2018, 150, 100–111. [Google Scholar] [CrossRef] [PubMed]
- Freytes, D.O.; Martin, J.; Velankar, S.S.; Lee, A.S.; Badylak, S.F. Preparation and rheological characterization of a gel form of the porcine urinary bladder matrix. Biomaterials 2008, 29, 1630–1637. [Google Scholar] [CrossRef]
- Singelyn, J.M.; Christman, K.L. Injectable Materials for the Treatment of Myocardial Infarction and Heart Failure: The Promise of Decellularized Matrices. J. Cardiovasc. Trans. Res. 2010, 3, 478–486. [Google Scholar] [CrossRef]
- Harris, G.M.; Raitman, I.; Schwarzbauer, J.E. Cell-derived decellularized extracellular matrices. Methods Cell Biol. 2018, 143, 97–114. [Google Scholar]
- Li, Y.; Foss, C.A.; Summerfield, D.D.; Doyle, J.J.; Torok, C.M.; Dietz, H.C.; Pomper, M.G.; Yu, M. Targeting collagen strands by photo-triggered triple-helix hybridization. Proc. Natl. Acad. Sci. USA 2012, 109, 14767–14772. [Google Scholar] [CrossRef]
- Hwang, J.; Huang, Y.; Burwell, T.J.; Peterson, N.C.; Connor, J.; Weiss, S.J.; Yu, S.M.; Li, Y. In Situ Imaging of Tissue Remodeling with Collagen Hybridizing Peptides. ACS Nano 2017, 11, 9825–9835. [Google Scholar] [CrossRef]
- Svensson, K.J.; Christianson, H.C.; Kucharzewska, P.; Fagerström, V.; Lundstedt, L.; Borgquist, S.; Jirström, K.; Belting, M. Chondroitin sulfate expression predicts poor outcome in breast cancer. Int. J. Oncol. 2011, 39, 1421–1428. [Google Scholar]
- Jahkola, T.; Toivonen, T.; Virtanen, I.; von Smitten, K.; Nordling, S.; von Boguslawski, K.; Haglund, C.; Nevanlinna, H.; Blomqvist, C. Tenascin-C expression in invasion border of early breast cancer: A predictor of local and distant recurrence. Br. J. Cancer 1998, 78, 1507–1513. [Google Scholar] [CrossRef] [PubMed]
- Mishra, D.K.; Thrall, M.J.; Baird, B.N.; Ott, H.C.; Blackmon, S.H.; Kurie, J.M.; Kim, M.P. Human Lung Cancer Cells Grown on Acellular Rat Lung Matrix Create Perfusable Tumor Nodules. Ann. Thorac. Surg. 2012, 93, 1075–1081. [Google Scholar] [CrossRef] [PubMed]
- Xiong, G.; Flynn, T.J.; Chen, J.; Trinkle, C.; Xu, R. Development of an ex vivo breast cancer lung colonization model utilizing a decellularized lung matrix. Integr. Biol. 2015, 7, 1518–1525. [Google Scholar] [CrossRef]
- Dunne, L.W.; Huang, Z.; Meng, W.; Fan, X.; Zhang, N.; Zhang, Q.; An, Z. Human decellularized adipose tissue scaffold as a model for breast cancer cell growth and drug treatments. Biomaterials 2014, 35, 4940–4949. [Google Scholar] [CrossRef] [PubMed]
- Sun, D.; Liu, Y.; Wang, H.; Deng, F.; Zhang, Y.; Zhao, S.; Ma, X.; Wu, H.; Sun, G. Novel decellularized liver matrix-alginate hybrid gel beads for the 3D culture of hepatocellular carcinoma cells. Int. J. Biol. Macromol. 2018, 109, 1154–1163. [Google Scholar] [CrossRef]
- Hussein, K.H.; Park, K.M.; Ghim, J.H.; Yang, S.R.; Woo, H.M. Three dimensional culture of HepG2 liver cells on a rat decellularized liver matrix for pharmacological studies. J. Biomed. Mater. Res. Part B 2016, 104, 263–273. [Google Scholar] [CrossRef]
- Tian, X.; Werner, M.E.; Roche, K.C.; Hanson, A.D.; Foote, H.P.; Yu, S.K.; Warner, S.B.; Copp, J.A.; Lara, H.; Wauthier, E.L.; et al. Organ-specific metastases obtained by culturing colorectal cancer cells on tissue-specific decellularized scaffolds. Nat. Biomed. Eng. 2018, 2, 443–452. [Google Scholar] [CrossRef]
- Liu, G.; Wang, B.; Li, S.; Jin, Q.; Dai, Y. Human breast cancer decellularized scaffolds promote epithelial-to-mesenchymal transitions and stemness of breast cancer cells in vitro. J. Cell. Physiol. 2019, 234, 9447–9456. [Google Scholar] [CrossRef] [PubMed]
- Koh, I.; Cha, J.; Park, J.; Choi, J.; Kang, S.-G.; Kim, P. The mode and dynamics of glioblastoma cell invasion into a decellularized tissue-derived extracellular matrix-based three-dimensional tumor model. Sci. Rep. 2018, 8, 4608. [Google Scholar] [CrossRef]
- Lü, W.-D.; Zhang, L.; Wu, C.-L.; Liu, Z.-G.; Lei, G.-Y.; Liu, J.; Gao, W.; Hu, Y.-R. Development of an Acellular Tumor Extracellular Matrix as a Three-Dimensional Scaffold for Tumor Engineering. PLoS ONE 2014, 9, e103672. [Google Scholar] [CrossRef]
- Romero-López, M.; Trinh, A.L.; Sobrino, A.; Hatch, M.M.S.; Keating, M.T.; Fimbres, C.; Lewis, D.E.; Gershon, P.D.; Botvinick, E.L.; Digman, M.; et al. Recapitulating the human tumor microenvironment: Colon tumor-derived extracellular matrix promotes angiogenesis and tumor cell growth. Biomaterials 2017, 116, 118–129. [Google Scholar] [CrossRef] [PubMed]
- Pinto, M.L.; Rios, E.; Silva, A.C.; Neves, S.C.; Caires, H.R.; Pinto, A.T.; Durães, C.; Carvalho, F.A.; Cardoso, A.P.; Santos, N.C.; et al. Decellularized human colorectal cancer matrices polarize macrophages towards an anti-inflammatory phenotype promoting cancer cell invasion via CCL18. Biomaterials 2017, 124, 211–224. [Google Scholar] [CrossRef] [PubMed]
- Piccoli, M.; D’Angelo, E.; Crotti, S.; Sensi, F.; Urbani, L.; Maghin, E.; Burns, A.; Coppi, P.D.; Fassan, M.; Rugge, M.; Rizzolio, F.; et al. Decellularized colorectal cancer matrix as bioactive microenvironment for in vitro 3D cancer research. J. Cell. Physiol. 2018, 233, 5937–5948. [Google Scholar] [CrossRef]
- Jin, Q.; Liu, G.; Li, S.; Yuan, H.; Yun, Z.; Zhang, W.; Zhang, S.; Dai, Y.; Ma, Y. Decellularized breast matrix as bioactive microenvironment for in vitro three-dimensional cancer culture. J. Cell. Physiol. 2019, 234, 3425–3435. [Google Scholar] [CrossRef]
- Aguado, B.A.; Caffe, J.R.; Nanavati, D.; Rao, S.S.; Bushnell, G.G.; Azarin, S.M.; Shea, L.D. Extracellular matrix mediators of metastatic cell colonization characterized using scaffold mimics of the pre-metastatic niche. Acta Biomater. 2016, 33, 13–24. [Google Scholar] [CrossRef] [PubMed]
- Miyauchi, Y.; Yasuchika, K.; Fukumitsu, K.; Ishii, T.; Ogiso, S.; Minami, T.; Kojima, H.; Yamaoka, R.; Katayama, H.; Kawai, T.; et al. A novel three-dimensional culture system maintaining the physiological extracellular matrix of fibrotic model livers accelerates progression of hepatocellular carcinoma cells. Sci. Rep. 2017, 7, 9827. [Google Scholar] [CrossRef]
- Sansing, H.A.; Sarkeshik, A.; Yates, J.R.; Patel, V.; Gutkind, J.S.; Yamada, K.M.; Berrier, A.L. Integrin αβ1, αvβ, α6β effectors p130Cas, Src and talin regulate carcinoma invasion and chemoresistance. Biochem. Biophys. Res. Commun. 2011, 406, 171–176. [Google Scholar] [CrossRef]
- Eberle, K.E.; Sansing, H.A.; Szaniszlo, P.; Resto, V.A.; Berrier, A.L. Carcinoma Matrix Controls Resistance to Cisplatin through Talin Regulation of NF-κB. PLoS ONE 2011, 6, e21496. [Google Scholar] [CrossRef]
- Serebriiskii, I.; Castelló-Cros, R.; Lamb, A.; Golemis, E.A.; Cukierman, E. Fibroblast-derived 3D matrix differentially regulates the growth and drug-responsiveness of human cancer cells. Matrix Biol. 2008, 27, 573–585. [Google Scholar] [CrossRef] [PubMed]
- Castelló-Cros, R.; Khan, D.R.; Simons, J.; Valianou, M.; Cukierman, E. Staged stromal extracellular 3D matrices differentially regulate breast cancer cell responses through PI3K and beta1-integrins. BMC Cancer 2009, 9, 94. [Google Scholar] [CrossRef] [PubMed]
- Hoshiba, T.; Tanaka, M. Breast cancer cell behaviors on staged tumorigenesis-mimicking matrices derived from tumor cells at various malignant stages. Biochem. Biophys. Res. Commun. 2013, 439, 291–296. [Google Scholar] [CrossRef]
- Hoshiba, T.; Tanaka, M. Decellularized matrices as in vitro models of extracellular matrix in tumor tissues at different malignant levels: Mechanism of 5-fluorouracil resistance in colorectal tumor cells. BBA-Mol. Cell Res. 2016, 1863, 2749–2757. [Google Scholar] [CrossRef]
- Hoshiba, T. An extracellular matrix (ECM) model at high malignant colorectal tumor increases chondroitin sulfate chains to promote epithelial-mesenchymal transition and chemoresistance acquisition. Exp. Cell Res. 2018, 370, 571–578. [Google Scholar] [CrossRef] [PubMed]
- Qian, B.Z. Inflammation fires up cancer metastasis. Semin. Cancer Biol. 2017, 47, 170–176. [Google Scholar] [CrossRef] [PubMed]
- Mitsumoto, M.; Kamura, T.; Kobayashi, H.; Sonoda, T.; Kaku, T.; Nakano, H. Emergence of higher levels of invasive and metastatic properties in the drug resistant cancer cell lines after the repeated administration of cisplatin in tumor-bearing mice. J. Cancer Res. Clin. Oncol. 1998, 124, 607–614. [Google Scholar] [CrossRef] [PubMed]
- Fidler, I.J. The pathogenesis of cancer metastasis: The “seed and soil” hypothesis revisited. Nat. Rev. Cancer 2003, 3, 1–6. [Google Scholar] [CrossRef]
| dECM Type | Advantages | Disadvantages |
|---|---|---|
| Tissue/organ-derived dECM | -Similar to native ECM composition and structure | -Limited ECM sources -Difficult for large-scale in vitro analyses -Large batch-to-batch differences due to cancer heterogeneity |
| Cultured cell-derived dECM | -Possible for large-scale in vitro analyses | -Difficult to prepare dECM that completely mimics native ECM composition and structure |
| Purposes | Principle | Methods |
|---|---|---|
| Confirmation of cell removal | DNA/cell nuclei detection | -Staining with hematoxylin and Hoechst 33258 -DNA content measurement |
| Intracellular protein detection | -Actin staining with fluorescent-labeled phalloidin -Immunocytochemistry of cytosolic proteins | |
| Compositional analysis | Detection of non-nucleic components | -Eosin staining |
| GAGs detection | -Alcian blue and toluidine blue stainings | |
| Collagens detection | -Sirius red and azan stainings | |
| Specific proteins/carbohydrates detection | -Immunohistochemical analysis with antibodies -Staining with lectins | |
| Proteomics (exhaustive research) | -Mass spectrometry | |
| Structural analysis | Structure observation | -SEM |
| Basement membrane detection | -TEM | |
| Fibril alignment | -fast Fourier transform analysis |
| dECM Type | Malignancy of dECM Source | Tissue/Cell of dECM Origin | Cells Cultured on dECM | Results | Reference |
|---|---|---|---|---|---|
| Tissue/organ- derived | Normal | Lung | Lung cancer A549, H460, H1299 cells | -Developed pattern of growth similar with original human lung cancer. | [56] |
| Breast cancer MDA-MB-231 and MCF-7 cells | -MDA-MB-231 cells undergone EMT can proliferation. -MCF-7 cells not undergone EMT died by apoptosis. | [57] | |||
| Adipose tissue | Breast cancer MCF-7, BT474, SKBR3 cells | -Proliferation, underwent EMT, and increased invasion. -Increased chemoresistance via Akt. | [58] | ||
| Liver | Hepatocellular carcinoma, HCCLM3 cells | -Increased uPA production and MMP-2 activity. -Decreased PAI-1 production. | [59] | ||
| Hepatocellular carcinoma, HepG2 cells | -Increased expression of genes relating to hepatic functions. | [60] | |||
| Liver and lung | Colorectal cancer HT-29, Caco2 and SW480 cells | -Exhibited morphology and gene expression pattern similar with metastatic sites of colorectal cancer. -The cells educated by dECM acquire metastatic ability. | [61] | ||
| Cancer | Mammary grand | Breast cancer MCF-7 cells | -Underwent EMT, increased stem cell marker expression and chemoresistance. | [62] | |
| Glioblastoma | Isolated glioblastoma cells | -Increased invasive ability via HAS gene expression. | [63] | ||
| A549-derived lung cancer | Breast cancer MCF-7 cells | -Cell proliferation. -Increased IL-8, bFGF, and VEGF production. | [64] | ||
| Normal and cancer | Colon | Colorectal cancer SW620, SW480, HCT116 cells, normal lung fibroblasts, endothelial colony forming cells | -Increased angiogenesis and cancer cell proliferation in cancer tissue-derived dECM. | [65] | |
| Isolated monocytes | -Promoted monocyte differentiation and CCL18 production to accelerate cancer cell invasion. | [66] | |||
| Colorectal cancer HT-29 cells | -Increased IL-8 production in cancer tissue-derived dECM. | [67] | |||
| Breast | Breast cancer MCF-7 cells | -Suppressed proliferation, EMT and angiogenic gene expression and increased apoptosis in normal tissue-derived dECM. -Promoted MMP-9 production, proliferation, EMT, and angiogenic gene expression and suppressed apoptosis in cancer tissue-derived dECM. | [68] | ||
| Lung and liver | Breast cancer LM2-4 and 4T1 cells | -Promoted cell adhesion and colonization in cancer tissue-derived dECM. | [69] | ||
| Normal and fibrosis | Liver | Hepatocellular carcinoma HLF and HuH7 cells | -Promoted proliferation. -Promoted EMT via integrin-FAK signaling. | [70] | |
| Cultured-cell-derived | Cancer | Tongue (Oral carcinoma HN12 cells) | Oral carcinoma HN12 cells | -Increased chemoresistance via talin, FAK, and NF-κB-mediated signals | [71,72] |
| Normal | Fibroblasts (NIH-3T3) | Various cancer and benign cells (HCT116, NCI-H460, PA-1, COLO 205, PANC-1, MCF-7, SW620, HCT116/p53-, HS 578T, PA1/E6, MCF-10A | -Increased chemoresistance via integrin β1-dependent survival signal. | [73] | |
| Normal and cancer | Fibroblasts (NIH-3T3 cells and cancer associated fibroblasts) | Breast cancer MDA-MB-231, MCF-7, and MCF-10A cells | -Activated PI3K-Akt signaling via integrin β1. -Changed morphology and cell migration behaviors | [74] | |
| Benign tumor and cancer | Breast (MDA-MB-231, MCF-7, and MCF-10A cells) | Breast cancer MDA-MB-231, MCF-7, and MCF-10A cells | -Promoted proliferation on invasive MDA-MB-231 cell-derived dECM. -Suppressed proliferation on benign MCF-10A cell-derived dECM. -Increased chemoresistance on invasive MDA-MB-231 cell-derived dECM. | [75] | |
| Normal and cancer | Colon (HT-29, SW480, CCD-841-CoN cells) | Colon cancer HT-29 and SW480 cells | -Increased chemoresistance on invasive HT-29 cell-derived dECM via Akt activation and ABCB1 upregulation. -Promoted EMT on invasive HT-29 cell-derived dECM via TGF-β signaling. | [39,76,77] |
© 2019 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hoshiba, T. Decellularized Extracellular Matrix for Cancer Research. Materials 2019, 12, 1311. https://doi.org/10.3390/ma12081311
Hoshiba T. Decellularized Extracellular Matrix for Cancer Research. Materials. 2019; 12(8):1311. https://doi.org/10.3390/ma12081311
Chicago/Turabian StyleHoshiba, Takashi. 2019. "Decellularized Extracellular Matrix for Cancer Research" Materials 12, no. 8: 1311. https://doi.org/10.3390/ma12081311
APA StyleHoshiba, T. (2019). Decellularized Extracellular Matrix for Cancer Research. Materials, 12(8), 1311. https://doi.org/10.3390/ma12081311





