Anaerobic Co-Digestion of Sludge and Organic Food Waste—Performance, Inhibition, and Impact on the Microbial Community
Abstract
1. Introduction
2. Materials and Methods
2.1. Experimental Methods
2.1.1. Substrate Characterisation
2.1.2. Biomethane Potential Tests
2.1.3. Continuous Experiments
2.1.4. Identification and Quantification of Methanogenic Archaea
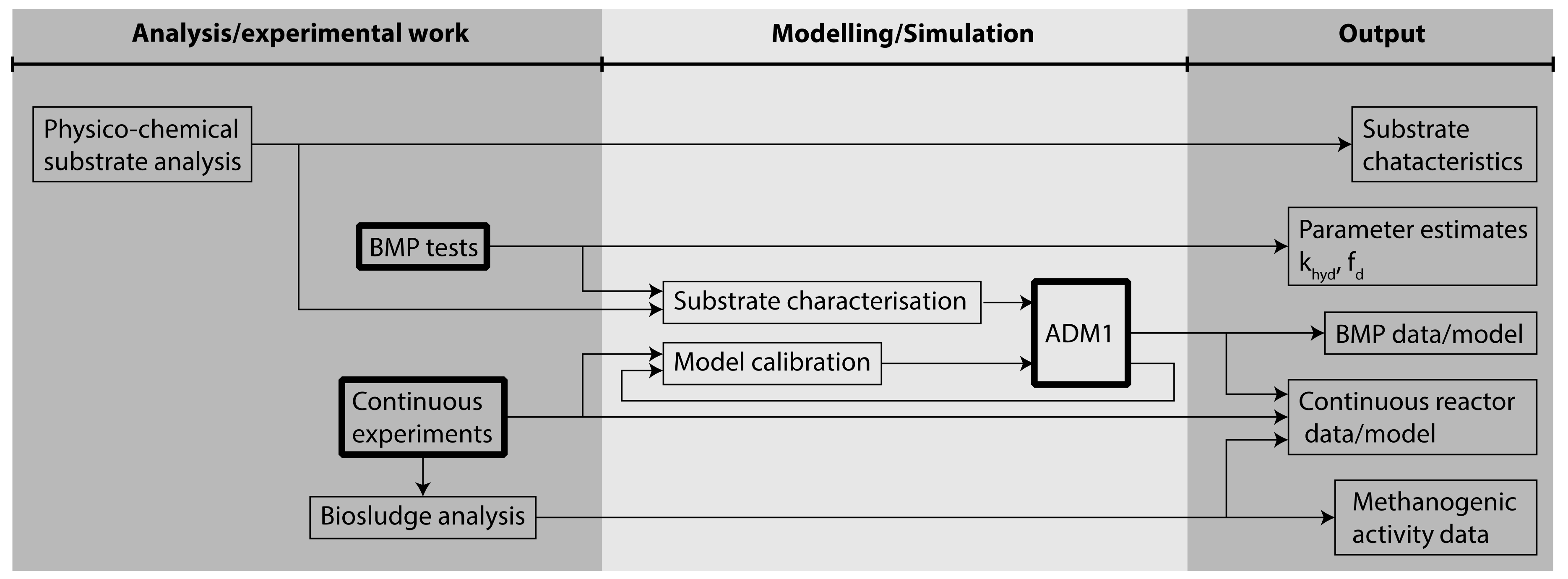
2.2. Modelling Anaerobic Digestion Processes
3. Results and Discussions
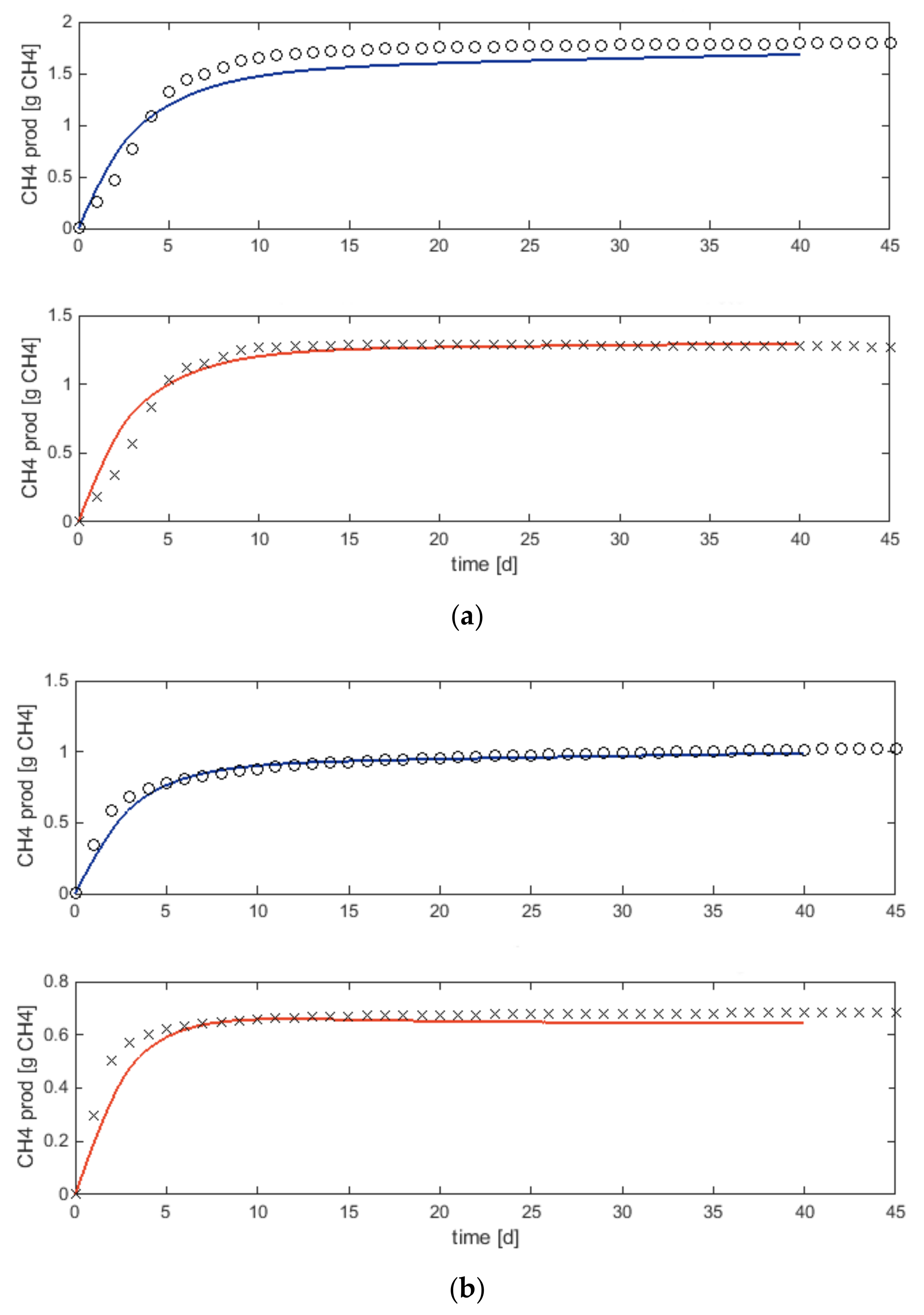
3.1. Biomethane Potential Tests and Substrate Characterisation
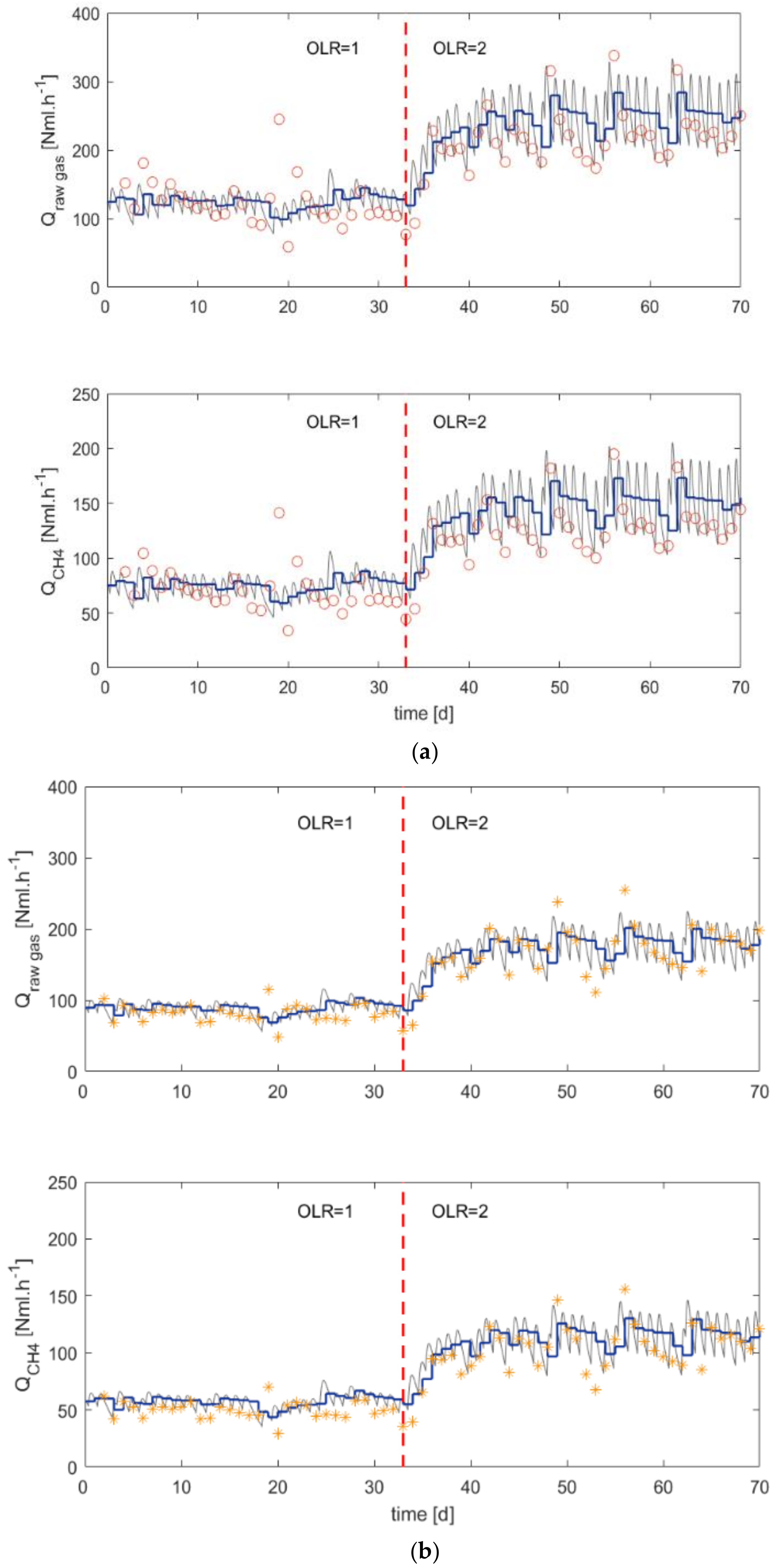
3.2. Continuous Reactors
4. Conclusions
- Applying anaerobic co-digestion (AcoD) with organic fractions of municipal solid waste (OFMSW) and mixed sludge increases the gas production at equivalent loads. With a feed composition 50/50% of the two substrates, the experiments show 22–42% more biogas production than for a reference reactor fed with only mixed sludge.
- Implementation of co-digestion of the sewage sludge and OFMSW shows rapid adaptation. In the biomethane potential tests (i.e., high substrate to inoculum ratio) a short lag, indicating adaptation, appears. However, in continuous lab-scale experiments at reasonable loading rates (1 to 2 kg/m3/d of VS) the response in gas production was immediate, showing that no adaptation was needed. This conclusion is supported by the equally rapid increase in methanogenic microbial population when co-digesting sludge and OFMSW.
- The organic loading rate and substrate composition have an impact on the composition of the microbial community in the reactor. The methanogenic microbial population increases when commencing co-digestion of sewage sludge and OFMSW on a WWTP inoculum. This effect is further pronounced at an increased load, which also promotes a change in the methanogenic microorganisms towards the acetate production pathway.
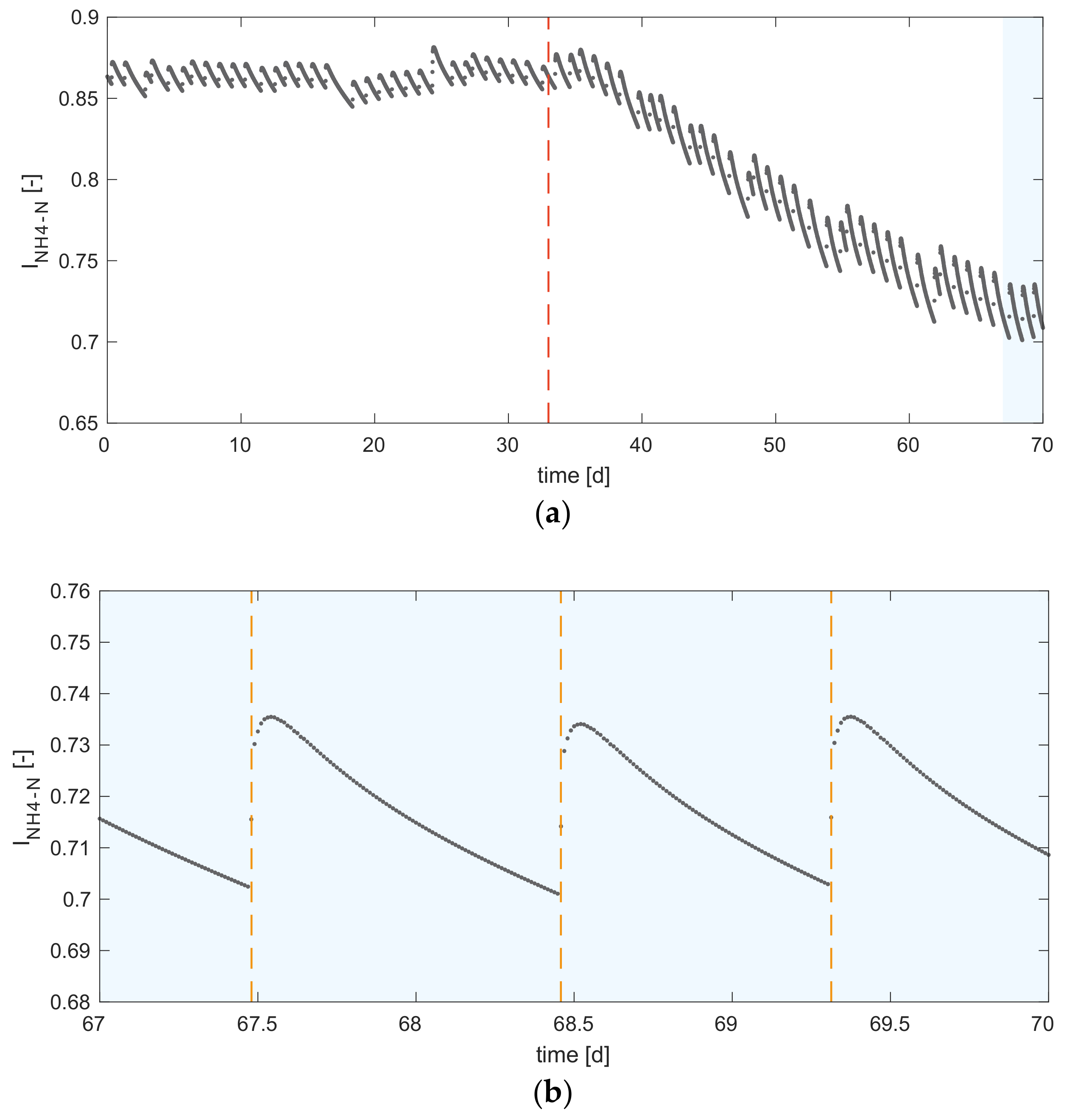
- The feeding strategy of continuous lab-scale digestion experiments has an impact on the instantaneous digester performance. The simulation results show that intermittent feeding leads to short-term ammonium inhibition of the process.
- Modelling is a suitable tool to evaluate the experimental design of AD. The BMP tests of the substrates were simulated using ADM1, and a characterisation procedure featuring both influent fractionation and parameter estimation was used. The estimated input fractions and parameters were successfully used also for modelling continuous experiments. It can be concluded that model characterisation based on BMP data is applicable for modelling continuous reactors. Furthermore, the study shows that modelling provides insights into inhibition phenomena which cannot be observed in the continuous experimental results.
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Nomenclature
| AcoD | Anaerobic co-digestion |
| AD | Anaerobic digestion |
| ADM1 | Anaerobic Digestion Model No. 1 |
| AMPTS | Automatic methane potential test system |
| BMP | Biomethane potential |
| COD | Chemical oxygen demand (g COD/m3) |
| CSTR | Continuous stirred tank reactor |
| DS | Dry solids (%) |
| fd | Model parameter for biodegradable fraction of COD (-) |
| HRT | Hydraulic retention time (d) |
| IIN | Model variable for ammonium inhibition (-) |
| khyd | Model parameter for hydrolysis rate (d−1) |
| LCFA | Long-chain fatty acids |
| OFMSW | Organic fraction of municipal solid waste |
| OLR | Organic loading rate (kg/m3/d) |
| SIN | Model state variable for inorganic nitrogen (mol N/l) |
| TN | Total nitrogen (g N/m3) |
| VAD | Reactor volume for anaerobic digestion |
| VFA | Volatile fatty acids (g COD/m3) |
| VS | Volatile solids (%) |
| WWTP | Wastewater treatment plant |
| Xch | Model state variable for carbohydrates (g COD/m3) |
| Xpr | Model state variable for proteins (g COD/m3) |
| Xli | Model state variable for lipids (g COD/m3) |
References
- Olsson, G. Water and Energy: Threats and Opportunities; IWA Publishing: London, UK, 2012. [Google Scholar]
- Batstone, D.J.; Virdis, B. The role of anaerobic digestion in the emerging energy economy. Curr. Opion. Biotechnol. 2014, 27, 142–149. [Google Scholar] [CrossRef] [PubMed]
- Krupp, M.; Schubert, J.; Widmann, R. Feasibility study for Co-Digestion of Sewage Sludge with OFMSW on Two Wastewater Treatment Plants in Germany. Waste Manag. 2005, 25, 393–399. [Google Scholar] [CrossRef] [PubMed]
- Lundkvist, M. Energieffektivisering–Rapport till möte med Energimyndigheten; Report Sweco Report 2005-01-14T; Sweco Viak AB: Halmstad, Sweden, 2005. (In Swedish) [Google Scholar]
- Mata-Alvarez, J.; Dosta, J.; Macé, S.; Astals, S. Codigestion of solid wastes: A review of its uses and perspectives including modeling. Crit. Rev. Biotechnol. 2011, 31, 99–111. [Google Scholar] [CrossRef] [PubMed]
- Mata-Alvarez, J.; Dosta, J.; Romero-Güiza, M.S.; Fonoll, X.; Peces, M.; Astals, S. A critical review on anaerobic co-digestion achievements between 2010 and 2013. Renew. Sustain. Energy Rev. 2014, 36, 412–427. [Google Scholar] [CrossRef]
- Ren, Y.Y.; Yu, M.; Wu, C.F.; Wang, Q.H.; Gao, M.; Huang, Q.Q.; Liu, Y.A. Comprehensive review on food waste anaerobic digestion: Research updates and tendencies. Bioresour. Technol. 2018, 247, 1069–1076. [Google Scholar] [CrossRef] [PubMed]
- Sosnowski, P.; Wieczorek, A.; Ledakowicz, S. Anaerobic co-digestion of sewage sludge and organic fraction of municipal solid wastes. Adv. Environ. Res. 2003, 7, 609–616. [Google Scholar] [CrossRef]
- Bolzonella, D.; Battistoni, P.; Susini, C.; Cecchi, F. Anaerobic codigestion of waste activated sludge and ofmsw: The experiences of viareggio and treviso plants (Italy). Water Sci. Technol. 2006, 53, 203–211. [Google Scholar] [CrossRef] [PubMed]
- Gómez, X.; Cuetos, M.J.; Cara, J.; Morán, A.; García, A.I. Anaerobic co-digestion of primary sludge and the fruit and vegetable fraction of the municipal solid wastes. Renew. Energy 2006, 31, 2017–2024. [Google Scholar] [CrossRef]
- Zhang, R.H.; El-Mashad, H.M.; Hartman, K.; Wang, F.Y.; Liu, G.Q.; Choate, C.; Gamble, P. Characterization of food waste as feedstock for anaerobic digestion. Bioresour. Technol. 2007, 98, 929–935. [Google Scholar] [CrossRef] [PubMed]
- Iacovidou, E.; Ohandja, D.-G.; Voulvoulis, N. Food waste co-digestion with sewage sludge–Realising its potential in the UK. J. Environ. Manag. 2012, 112, 267–274. [Google Scholar] [CrossRef] [PubMed]
- Fonoll, X.; Astals, S.; Dosta, J.; Mata-Alvarez, J. Anaerobic co-digestion of sewage sludge and fruit wastes: Evaluation of the transitory states when the co-substrate is changed. Chem. Eng. J. 2015, 262, 1268–1274. [Google Scholar] [CrossRef]
- Breunig, H.M.; Jin, L.; Robinson, A.; Scown, C.D. Bioenergy potential from food waste in California. Environ. Sci. Technol. 2017, 51, 1120–1128. [Google Scholar] [CrossRef] [PubMed]
- Yenigun, O.; Demirel, B. Ammonia inhibition in anaerobic digestion: A review. Process. Biochem. 2013, 48, 901–911. [Google Scholar] [CrossRef]
- Arnell, M. Performance Assessment of Wastewater Treatment Plants-Multi-Objective Analysis Using Plant-Wide Models. Ph.D. Thesis, Lund University, Lund, Sweden, 16 December 2016. [Google Scholar]
- Li, L.; Peng, X.Y.; Wang, X.M.; Di Wu, D. Anaerobic digestion of food waste: A review focusing on process stability. Bioresour. Technol. 2018, 248, 20–28. [Google Scholar] [CrossRef] [PubMed]
- Gou, C.L.; Yang, Z.H.; Huang, J.; Wang, H.L.; Xu, H.Y.; Wang, L.K. Effects of temperature and organic loading rate on the performance and microbial community of anaerobic co-digestion of waste activated sludge and food waste. Chemosphere 2014, 105, 146–151. [Google Scholar] [CrossRef] [PubMed]
- Derbal, K.; Bencheikh-Iehocine, M.; Cecchi, F.; Meniai, A.H.; Pavan, P. Application of the IWA ADM1 model to simulate anaerobic co-digestion of organic waste with waste activated sludge in mesophilic condition. Bioresour. Technol. 2009, 100, 1539–1543. [Google Scholar] [CrossRef] [PubMed]
- Zaher, U.; Li, R.; Jeppsson, U.; Steyer, J.P.; Chen, S. GISCOD: General integrated solid waste co-digestion model. Water Res. 2009, 43, 2717–2727. [Google Scholar] [CrossRef] [PubMed]
- Batstone, D.J.; Keller, J.; Angelidaki, R.I.; Kalyuzhnyi, S.V.; Pavlostathis, S.G.; Rozzi, A.; Sanders, W.T.M.; Siegrist, H.; Vavilin, V.A. Anaerobic Digestion Model No. 1 (ADM1); IWA Publishing: London, UK, 2002. [Google Scholar]
- Rosen, C.; Jeppsson, U. Aspects on ADM1 Implementation within the BSM2 Framework. Department of Industrial Electrical Engineering and Automation. Available online: http://portal.research.lu.se/ws/files/5715450/588082.pdf (accessed on 13 August 2018).
- Gernaey, K.V.; Jeppsson, U.; Vanrolleghem, P.A.; Copp, J.B. Benchmarking of Control Strategies for Wastewater Treatment Plants; IWA Publishing: London, UK, 2014. [Google Scholar]
- Wang, P.; Wang, H.T.; Qiu, Y.Q.; Ren, L.H.; Jiang, B. Microbial characteristics in anaerobic digestion process of food waste for methane production–A review. Bioresour. Technol. 2017, 248, 29–36. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Whitman, W.B. Metabolic, phylogenetic, and ecological diversity of the methanogenic archaea. Ann. N. Y. Acad. Sci. 2008, 1125, 171–189. [Google Scholar] [CrossRef] [PubMed]
- Angelidaki, I.; Alves, M.; Bolzonella, D.; Borzacconi, L.; Campos, J.L.; Guwy, A.J.; Kalyuzhnyi, S.; Jenicek, P.; van Lier, J.B. Defining the biomethane potential (bmp) of solid organic wastes and energy crops: a proposed protocol for batch assays. Water Sci. Technol. 2009, 59, 927. [Google Scholar] [CrossRef] [PubMed]
- Holliger, C.; Alves, M.; Andrade, D.; Angelidaki, I.; Astals, S.; Baier, U.; Bougrier, C.; Buffiere, P.; Carballa, M.; De Wilde, V.; et al. Towards a standardization of biomethane potential tests. Water Sci. Technol. 2016, 74, 2515–2522. [Google Scholar] [CrossRef] [PubMed]
- Carlsson, M.; Schnürer, A. Handbook Methane Potential. Available online: http://sgc.camero.se/ckfinder/userfiles/files/SGC237.pdf (accessed on 13 August 2018).
- Forgács, G.; Pourbafrani, M.; Niklasson, C.; Taherzadeh, M.J.; Hováth, I.S. Methane production from citrus wastes: process development and cost estimation. J. Chem. Technol. Biotechnol. 2011, 87, 250–255. [Google Scholar] [CrossRef]
- Snaidr, J.; Amann, R.; Huber, I.; Ludwig, W.; Schleifer, K.H. Phylogenetic analysis and in situ identification of bacteria in activated sludge. Appl. Environ. Microbiol. 1997, 63, 2884–2896. [Google Scholar] [PubMed]
- Raskin, L.; Stromley, J.M.; Rittmann, B.E.; Stahl, D.A. Group-specific 16s rRNA hybridization probes to describe natural communities of methanogens. Appl. Environ. Microbiol. 1994, 60, 1232–1240. [Google Scholar] [PubMed]
- Amann, R.I.; Ludwig, W.; Schleifer, K.H. Phylogenetic identification and in-situ detection of individual microbial-cells without cultivation. Microbiol. Rev. 1995, 59, 143–169. [Google Scholar] [PubMed]
- Daims, H.; Brühl, A.; Amann, R.; Schleifer, K.-H.; Wagner, M. The Domain-Specific Probe EUB338 is Insufficient for the Detection of all Bacteria: Development and Evaluation of a More Comprehensive Probe Set. Syst. Appl. Microbiol. 1999, 22, 434–444. [Google Scholar] [CrossRef]
- Arnell, M.; Astals, S.; Åmand, L.; Batstone, D.J.; Jensen, P.D.; Jeppsson, U. Modelling anaerobic co-digestion in benchmark simulation model No. 2: Parameter estimation, substrate characterisation and plant-wide integration. Water Res. 2016, 98, 138–146. [Google Scholar] [CrossRef] [PubMed]
- Neves, L.; Oliveira, R.; Alves, M.M. Co-digestion of cow manure, food waste and intermittent input of fat. Bioresour. Technol. 2009, 100, 1957–1962. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Chen, S.; Li, X. Biogas production from anaerobic co-digestion of food waste with dairy manure in a two-phase digestion system. Appl. Biochem. Biotechnol. 2009, 160, 643–654. [Google Scholar] [CrossRef] [PubMed]
- La Cour Jansen, J.; Gruvberger, C.; Hanner, N.; Aspegren, H.; Svard, A. Digestion of sludge and organic waste in the sustainability concept for Malmo, Sweden. Water Sci. Technol. 2004, 49, 163–169. [Google Scholar] [CrossRef] [PubMed]
- Angelidaki, I.; Ellegaard, L.; Ahring, B.K. A comprehensive model of anaerobic bioconversion of complex substrates to biogas. Biotechnol. Bioeng. 1999, 63, 363–372. [Google Scholar] [CrossRef]
| Measurements | Mixed Sludge | OFMSW |
|---|---|---|
| DS (kg DS/t) | 73.6 | 186 |
| VS (kg VS/t) | 59.6 | 173 |
| COD (g/L) | 57.2 | 235* |
| NH4 (mg/L) | 99.8 | 511.2 |
| TN (mg/L) | 3 320 | 5 184 |
| VFA (mg/L) | 1 597 | 10 040 |
| Raw protein (N × 6.25) (% of DS) | 21 | 20 |
| Raw lipids (g/100 g) | 1.24 | 2.98 |
| Estimated model parameters and state variables | ||
| Hydrolysis rate (khyd) (d−1) | 0.34 | 0.25 |
| Carbohydrates (Xch) (kg COD/m3) | 2.60 | 69.9 |
| Protein (Xpr) (kg COD/m3) | 20.2 | 56.7 |
| Lipids (Xli) (kg COD/m3) | 24.6 | 85.6 |
| Ultimate methane potential (B0) (m3 CH4/t VS) | 287 | 475 |
| Percentage of Taxonomic Group | |||||
|---|---|---|---|---|---|
| Orders and Families of Methanogenic Archaea | Inoculum and Start Point of both Reactors | R1 OLR 1.0 | R2 OLR 1.0 | R1 OLR 2.0 | R2 OLR 2.0 |
| Methanomicrobiales total | 8 | 12 | 9 | 12 | 15 |
| Thereof: Methanocorpusculaceae | n.d. | n.d. | n.d. | n.d. | n.d. |
| Thereof: Methanospirillaceae | n.d. | n.d. | n.d. | 2 | 1 |
| Methanobacteriales | n.d. | n.d. | n.d. | n.d. | n.d. |
| Methanosarcinales total | 8 | 15 | 10 | 20 | 15 |
| Thereof: Methanosaetaceae | 8 | 15 | 10 | 20 | 15 |
| Methanococcaceae | n.d. | n.d. | n.d. | n.d. | n.d. |
| All methanogenic archaea | 16 | 27 | 19 | 32 | 30 |
| Measurements | Average Day 0–30 | Average Day 50–70 | ||
|---|---|---|---|---|
| Data | Model | Data | Model | |
| VFA (g/L) | 181 | 108 | 121 | 220 |
| Alkalinity (mg/L) | 3420 | 2990 | 2628 | 3890 |
| NH4-N (mg/L) | 1100 | 700 | 794 | 880 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Keucken, A.; Habagil, M.; Batstone, D.; Jeppsson, U.; Arnell, M. Anaerobic Co-Digestion of Sludge and Organic Food Waste—Performance, Inhibition, and Impact on the Microbial Community. Energies 2018, 11, 2325. https://doi.org/10.3390/en11092325
Keucken A, Habagil M, Batstone D, Jeppsson U, Arnell M. Anaerobic Co-Digestion of Sludge and Organic Food Waste—Performance, Inhibition, and Impact on the Microbial Community. Energies. 2018; 11(9):2325. https://doi.org/10.3390/en11092325
Chicago/Turabian StyleKeucken, Alexander, Moshe Habagil, Damien Batstone, Ulf Jeppsson, and Magnus Arnell. 2018. "Anaerobic Co-Digestion of Sludge and Organic Food Waste—Performance, Inhibition, and Impact on the Microbial Community" Energies 11, no. 9: 2325. https://doi.org/10.3390/en11092325
APA StyleKeucken, A., Habagil, M., Batstone, D., Jeppsson, U., & Arnell, M. (2018). Anaerobic Co-Digestion of Sludge and Organic Food Waste—Performance, Inhibition, and Impact on the Microbial Community. Energies, 11(9), 2325. https://doi.org/10.3390/en11092325






