Reconstructing Clonal Evolution—A Systematic Evaluation of Current Bioinformatics Approaches
Abstract
1. Introduction
2. Materials and Methods
2.1. clevRsim
2.1.1. Simulating the Phylogeny
2.1.2. Simulating Variants
2.2. Simulated Data Sets
2.3. Tools for Variant Clustering
2.4. Tools for Clonal Evolution Tree Reconstruction
2.5. Statistical Analysis
3. Results
3.1. Reliability of clevRsim
3.2. Variant Clustering in the Absence of CNVs
3.3. Variant Clustering in the Presence of CNVs
3.4. Clonal Evolution Tree Reconstruction
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| ALL | Acute lymphoblastic leukemia |
| AML | Acute myeloid leukemia |
| BL | Burkitt lymphoma |
| CCF | Cancer cell fraction |
| CNV | Copy number variant |
| DSD | Discrete spectral distance |
| FISH | Fluorescence in situ hybridization |
| IPSS-R | Revised International Prognostic Scoring System |
| LOH | Loss of heterozygosity |
| MDS | Myeoldysplastic syndromes |
| scDNA-seq | Single-cell DNA sequencing |
| SNV | Single-nucleotide variant |
| VI | Variation of information |
| WES | Whole-exome sequencing |
Appendix A. Additional Materials and Methods
Appendix A.1. clevRsim
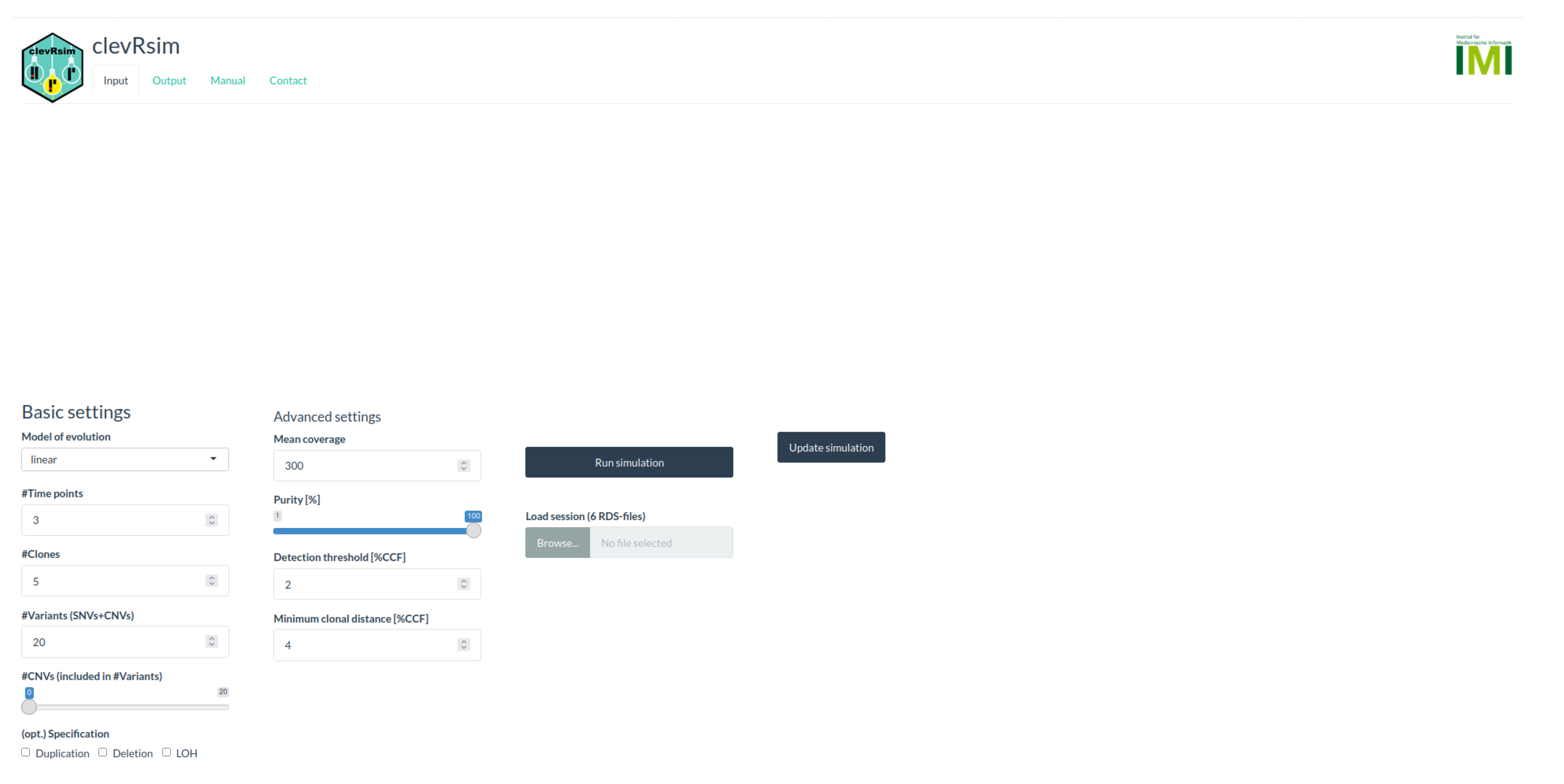

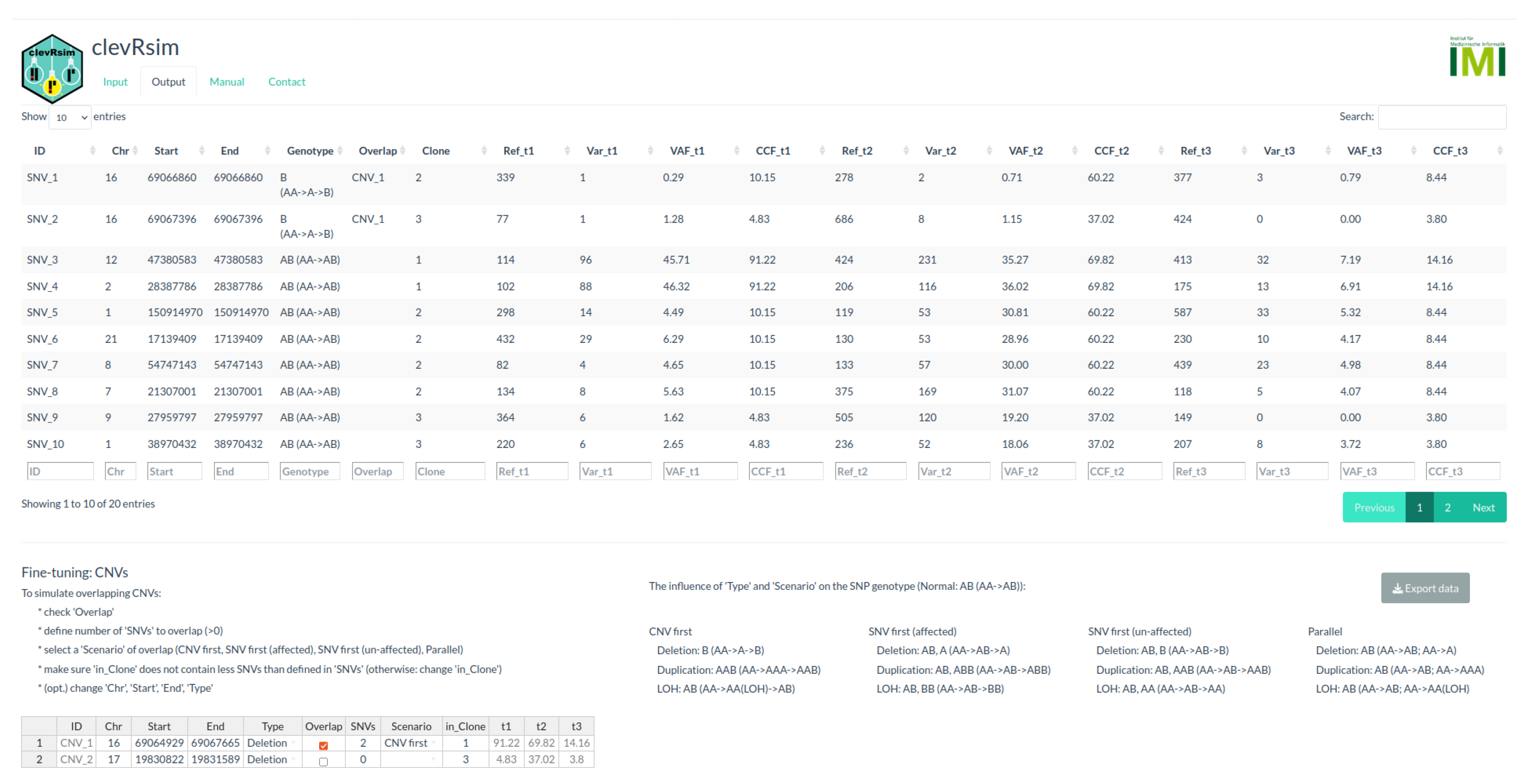

| Scenario | CNV Type | Genotype | ||||||
|---|---|---|---|---|---|---|---|---|
| CNV first | Deletion | AA | → | A | → | B | ||
| Duplication | AA | → | AAA | → | AAB | |||
| LOH | AA | → | AA | → | AB | |||
| SNV first (affected) | Deletion | AA | → | AB | → | A | ||
| Duplication | AA | → | AB | → | ABB | |||
| LOH | AA | → | AB | → | BB | |||
| SNV first (un-affected) | Deletion | AA | → | AB | → | B | ||
| Duplication | AA | → | AB | → | AAB | |||
| LOH | AA | → | AB | → | AA | |||
| Parallel | Deletion | AA | → | AB | AA | → | A | |
| Duplication | AA | → | AB | AA | → | AAA | ||
| LOH | AA | → | AB | AA | → | AA | ||
Appendix A.2. Tools for Variant Clustering
- category<-"Timepoints"
- values<-c(1:10)
- for(i in values){
- message("i=",i)
- for(j in 1:10){
- message("j=",j)
- y<-read.table(paste0(dir,category,"/",i,"/Patient",
- j,"/y.txt"),
- header=T,sep="\t")
- y<-as.matrix(y)
- n<-read.table(paste0(dir,category,"/",i,"/Patient",
- j,"/n.txt"),
- header=T,sep="\t")
- n<-as.matrix(n)
- purity<-rep(1,length(y[1,]))
- tcn<-m<-y
- for(k in 1:length(y[,1])){
- for(l in 1:length(y[1,])){
- tcn[k,l]<-2
- m[k,l]<-1
- }
- }
- input.data<-list(y=y,
- n=n,
- purity=purity,
- tcn=tcn,
- m=m,
- I=length(y[,1]),
- S=length(y[1,]))
- clustering = tryCatch({
- all.set.results <- clusterSep(input.data,
- n.iter = 1000,
- n.burn = 100,
- thin = 1,
- max_K = 10)
- set.k.choices <- writeSetKTable(all.set.results)
- best.set.chains <- collectBestKChains(all.set.results,
- chosen_K = set.k.choices$chosen_K[!is.na(
- set.k.choices$chosen_K)])
- chains <- mergeSetChains(best.set.chains, input.data)
- cluster.table<-writeClusterCCFsTable(chains$w_chain)
- cluster.mutations<-writeClusterAssignmentsTable(
- chains$z_chain)
- }, error = function(e) {
- message("fail")
- NULL
- })
- }
- }
Appendix A.3. Tools for Clonal Evolution Tree Reconstruction
- python3 ~/Downloads/submarine/submarine.py --basic_version \
- --freq_file $dir/Timepoints/$sample/Sample.csv \
- --output_prefix $dir/Timepoints/$sample/sample_basic
Appendix B. Additional Results
Appendix B.1. Reliability of clevRsim
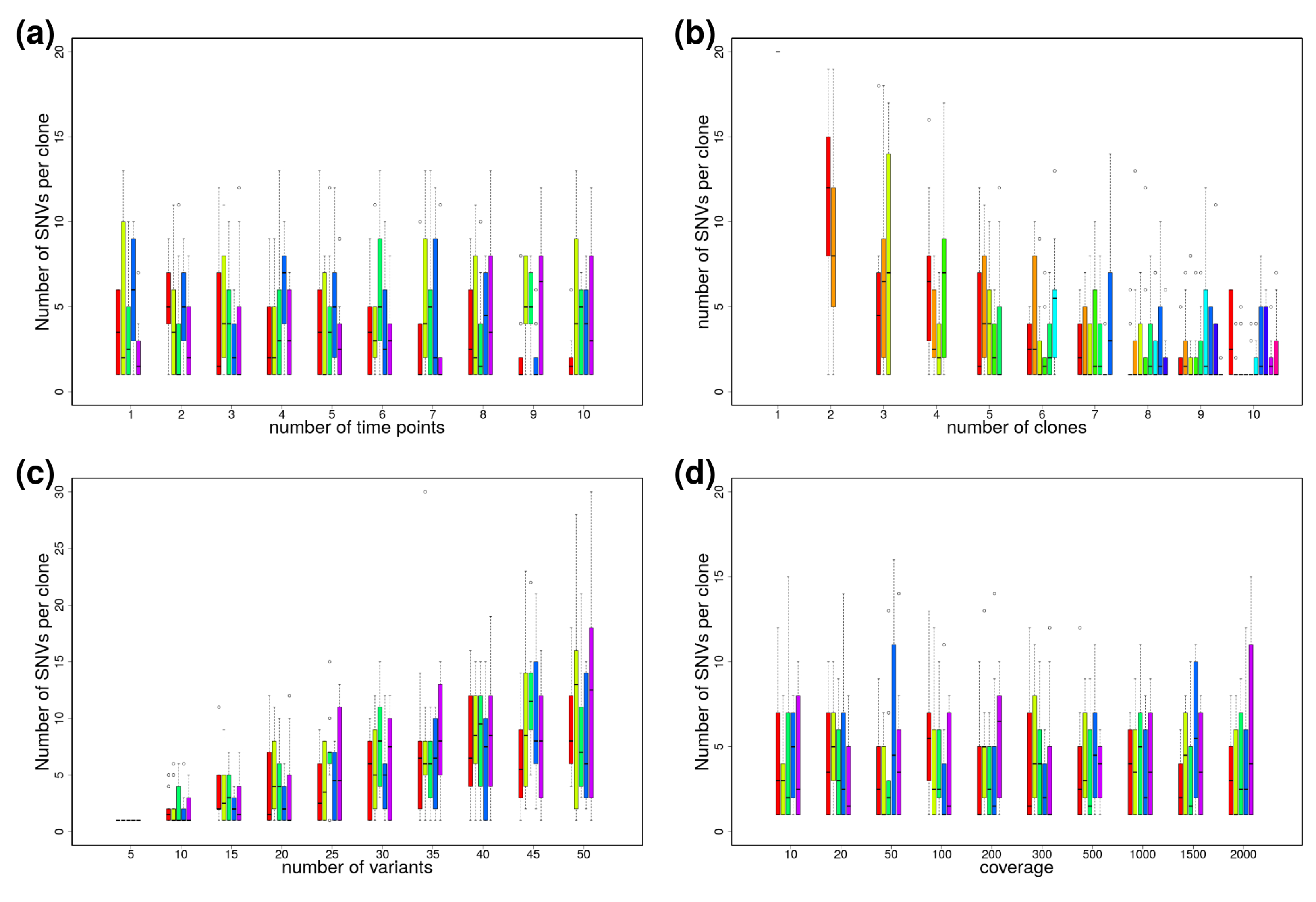
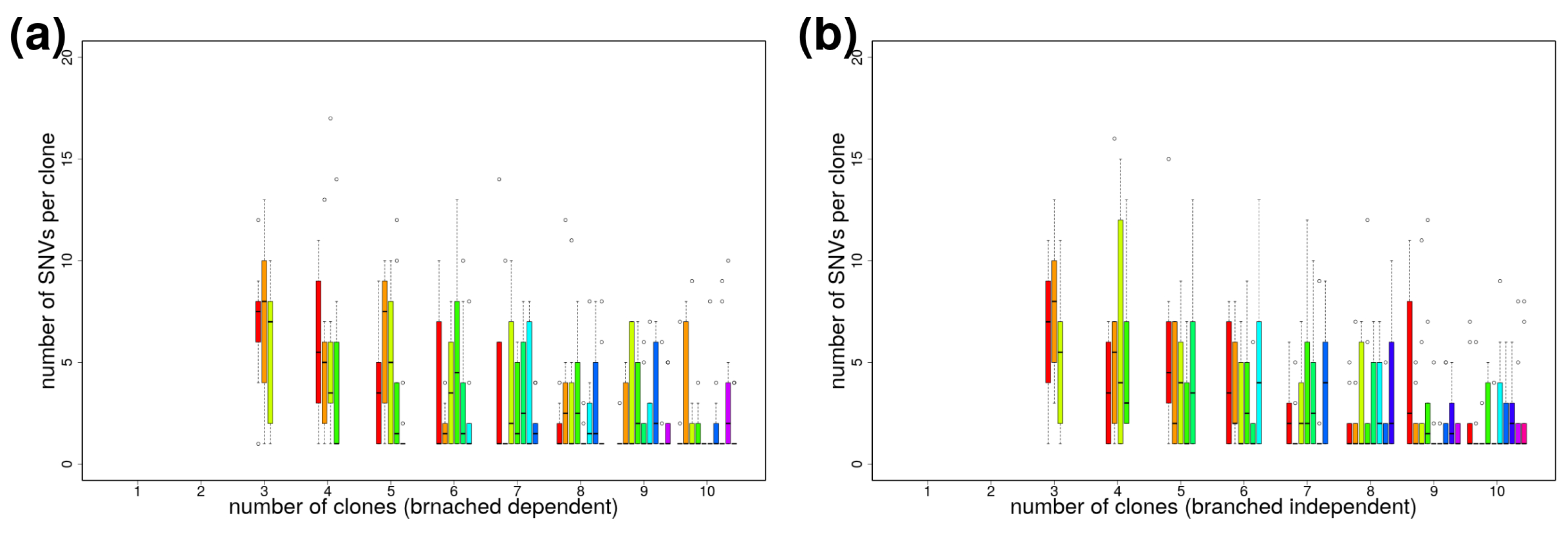
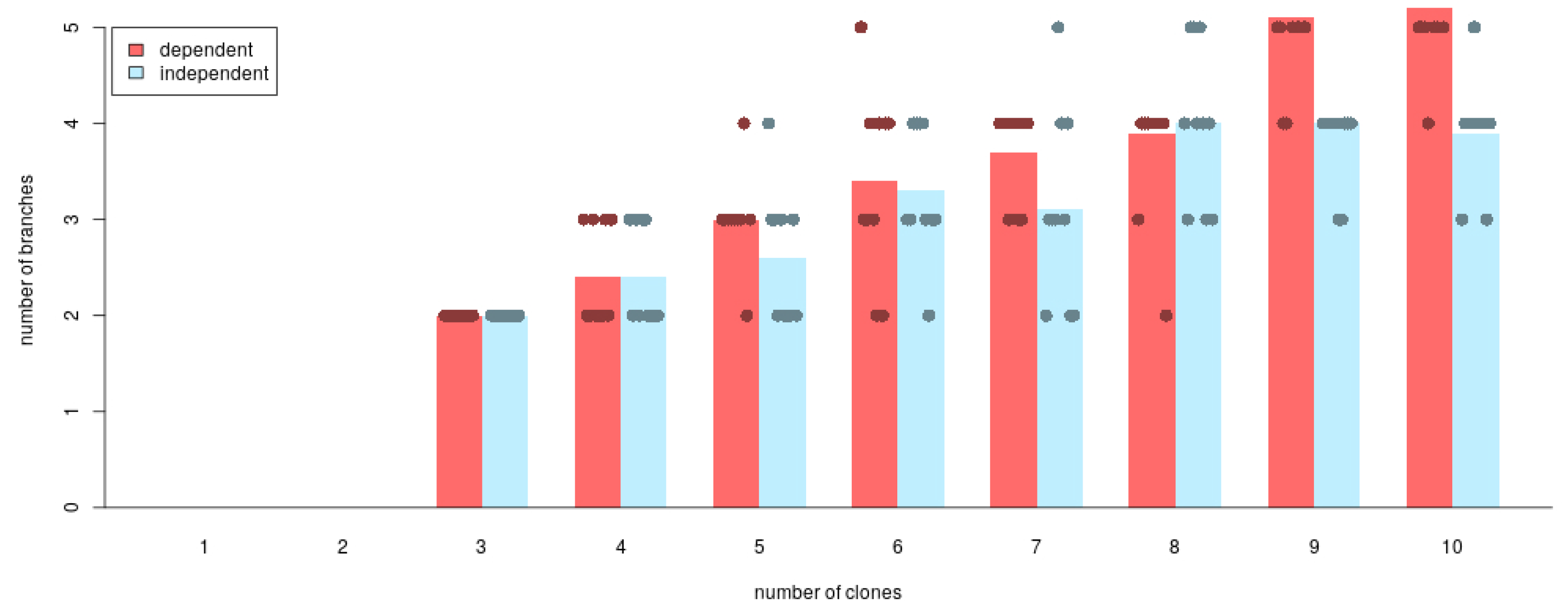
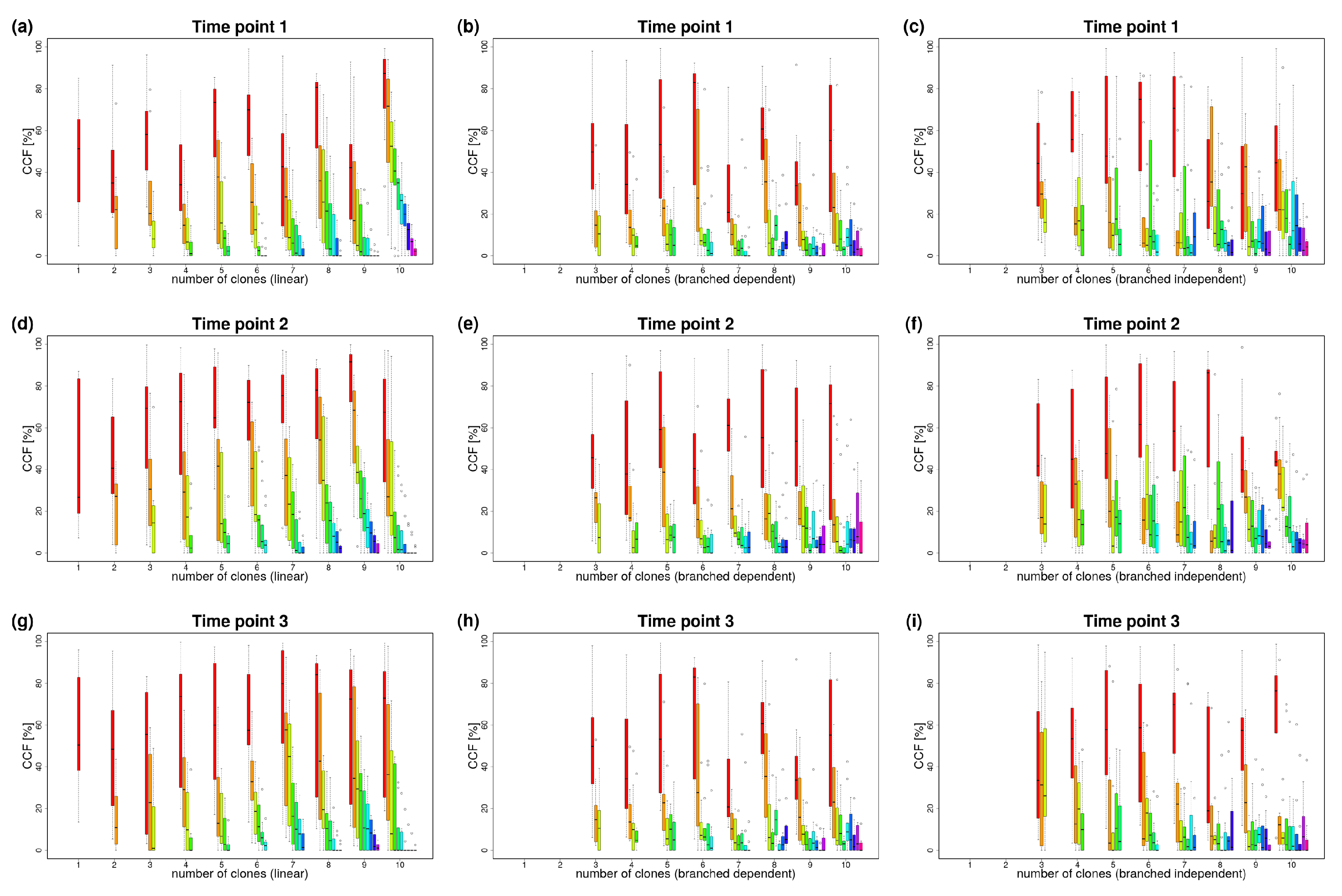
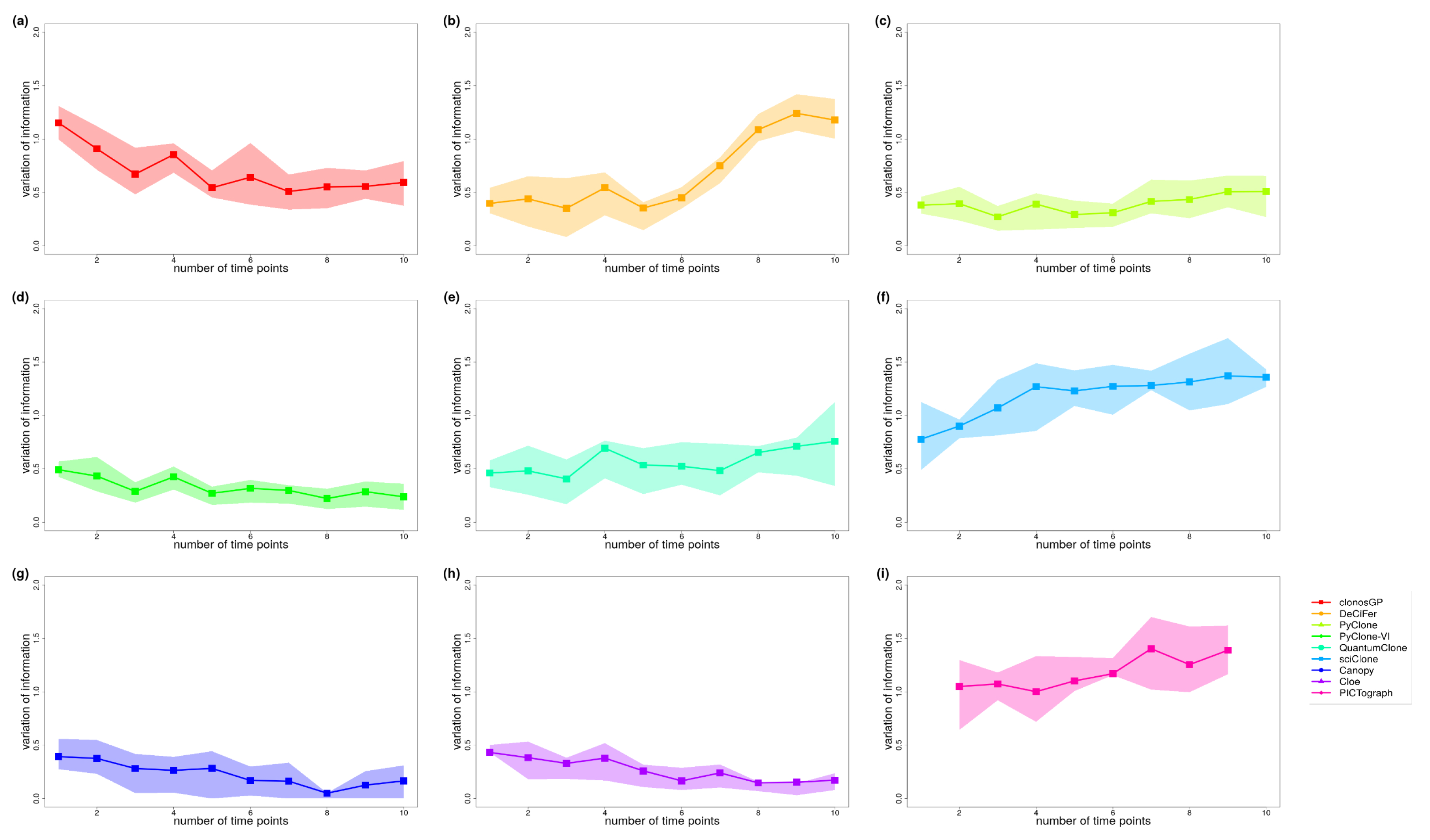
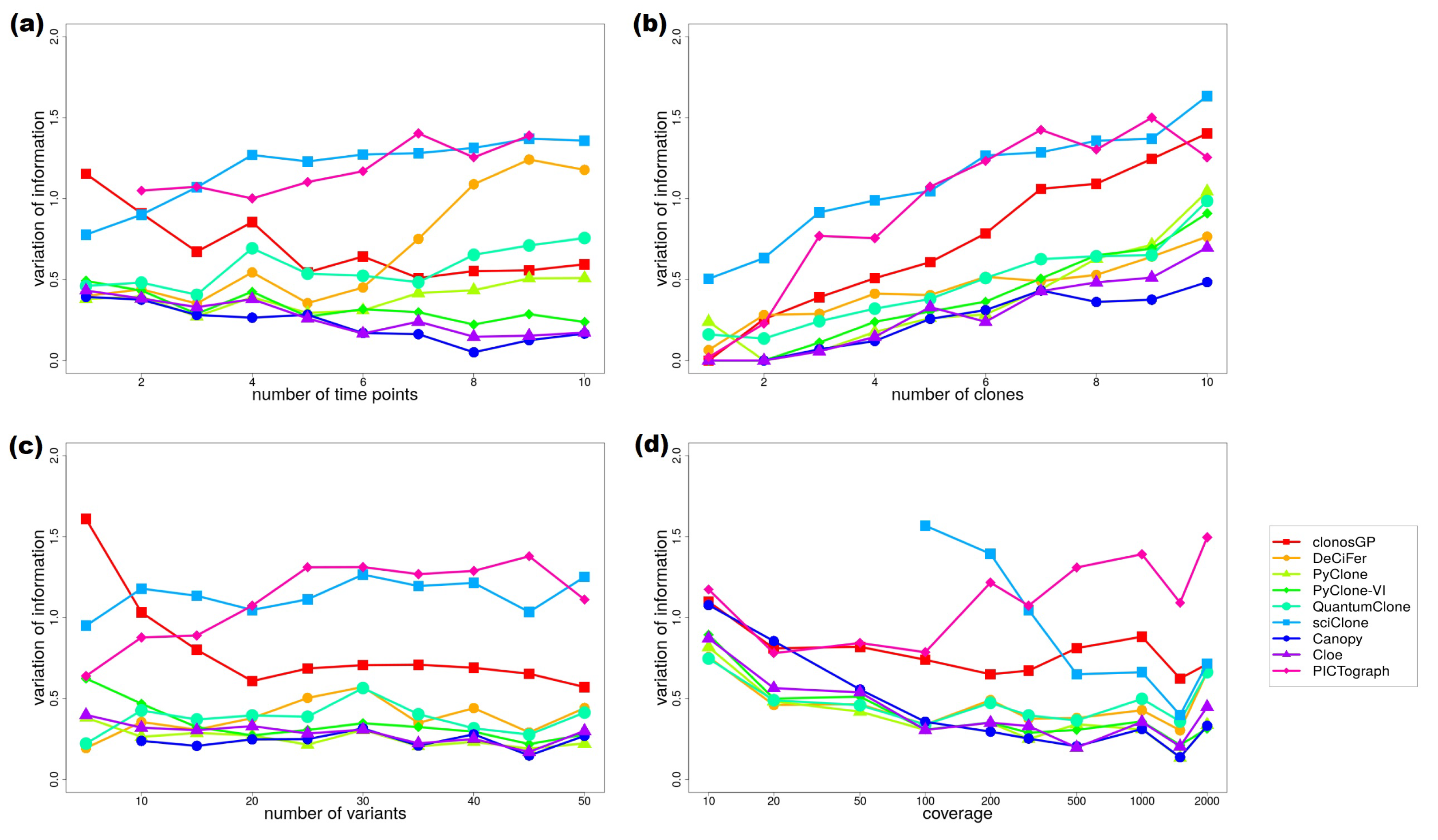
Appendix B.2. Variant Clustering in the Absence of CNVs
| Time Points | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Values | clonosGP | DeCiFer | PyClone | PyClone-VI | QuantumClone | sciClone | Canopy | Cloe | PICTograph |
| 1 | 1.15 | 0.40 | 0.38 | 0.49 | 0.46 | 0.78 | 0.39 | 0.43 | NA |
| 2 | 0.91 | 0.44 | 0.40 | 0.43 | 0.48 | 0.90 | 0.38 | 0.38 | 1.05 |
| 3 | 0.67 | 0.35 | 0.27 | 0.29 | 0.41 | 1.07 | 0.28 | 0.33 | 1.07 |
| 4 | 0.86 | 0.54 | 0.39 | 0.42 | 0.69 | 1.27 | 0.26 | 0.38 | 1.00 |
| 5 | 0.54 | 0.35 | 0.29 | 0.27 | 0.54 | 1.23 | 0.28 | 0.26 | 1.10 |
| 6 | 0.64 | 0.45 | 0.31 | 0.32 | 0.52 | 1.27 | 0.17 | 0.17 | 1.17 |
| 7 | 0.51 | 0.75 | 0.42 | 0.30 | 0.48 | 1.28 | 0.16 | 0.24 | 1.40 |
| 8 | 0.55 | 1.09 | 0.43 | 0.22 | 0.65 | 1.31 | 0.05 | 0.15 | 1.26 |
| 9 | 0.56 | 1.24 | 0.51 | 0.29 | 0.71 | 1.37 | 0.13 | 0.15 | 1.39 |
| 10 | 0.59 | 1.18 | 0.51 | 0.24 | 0.76 | 1.36 | 0.17 | 0.17 | NA |
| Clones | |||||||||
| Values | clonosGP | DeCiFer | PyClone | PyClone-VI | QuantumClone | sciClone | Canopy | Cloe | PICTograph |
| 1 | 0.00 | 0.06 | 0.24 | 0.00 | 0.16 | 0.50 | NA | 0.00 | 0.02 |
| 2 | 0.26 | 0.28 | 0.00 | 0.00 | 0.14 | 0.63 | 0.00 | 0.00 | 0.23 |
| 3 | 0.39 | 0.29 | 0.06 | 0.11 | 0.24 | 0.92 | 0.07 | 0.06 | 0.77 |
| 4 | 0.51 | 0.41 | 0.18 | 0.24 | 0.32 | 0.99 | 0.12 | 0.15 | 0.76 |
| 5 | 0.61 | 0.40 | 0.26 | 0.30 | 0.38 | 1.05 | 0.26 | 0.33 | 1.07 |
| 6 | 0.78 | 0.52 | 0.28 | 0.36 | 0.51 | 1.27 | 0.31 | 0.24 | 1.23 |
| 7 | 1.06 | 0.49 | 0.44 | 0.51 | 0.63 | 1.29 | 0.43 | 0.43 | 1.43 |
| 8 | 1.09 | 0.53 | 0.63 | 0.65 | 0.64 | 1.36 | 0.36 | 0.48 | 1.30 |
| 9 | 1.25 | 0.64 | 0.71 | 0.69 | 0.65 | 1.37 | 0.38 | 0.51 | 1.50 |
| 10 | 1.40 | 0.77 | 1.04 | 0.91 | 0.99 | 1.63 | 0.48 | 0.70 | 1.25 |
| Variants | |||||||||
| Values | clonosGP | DeCiFer | PyClone | PyClone-VI | QuantumClone | sciClone | Canopy | Cloe | PICTograph |
| 5 | 1.61 | 0.19 | 0.38 | 0.62 | 0.22 | 0.95 | NA | 0.40 | 0.64 |
| 10 | 1.03 | 0.35 | 0.26 | 0.47 | 0.43 | 1.18 | 0.24 | 0.32 | 0.88 |
| 15 | 0.80 | 0.31 | 0.29 | 0.32 | 0.37 | 1.13 | 0.21 | 0.30 | 0.89 |
| 20 | 0.61 | 0.38 | 0.27 | 0.27 | 0.40 | 1.05 | 0.25 | 0.33 | 1.07 |
| 25 | 0.69 | 0.50 | 0.21 | 0.31 | 0.39 | 1.11 | 0.25 | 0.28 | 1.31 |
| 30 | 0.71 | 0.57 | 0.31 | 0.35 | 0.56 | 1.27 | 0.32 | 0.31 | 1.31 |
| 35 | 0.71 | 0.35 | 0.20 | 0.32 | 0.40 | 1.20 | 0.21 | 0.22 | 1.27 |
| 40 | 0.69 | 0.44 | 0.23 | 0.29 | 0.32 | 1.21 | 0.28 | 0.25 | 1.29 |
| 45 | 0.65 | 0.29 | 0.19 | 0.22 | 0.28 | 1.03 | 0.15 | 0.17 | 1.38 |
| 50 | 0.57 | 0.44 | 0.22 | 0.28 | 0.41 | 1.25 | 0.27 | 0.30 | 1.11 |
| Coverage | |||||||||
| Values | clonosGP | DeCiFer | PyClone | PyClone-VI | QuantumClone | sciClone | Canopy | Cloe | PICTograph |
| 10 | 1.10 | 0.76 | 0.82 | 0.89 | 0.75 | NA | 1.08 | 0.87 | 1.17 |
| 20 | 0.81 | 0.46 | 0.48 | 0.50 | 0.49 | NA | 0.85 | 0.56 | 0.78 |
| 50 | 0.82 | 0.47 | 0.42 | 0.51 | 0.46 | NA | 0.56 | 0.54 | 0.84 |
| 100 | 0.74 | 0.34 | 0.31 | 0.31 | 0.34 | 1.57 | 0.36 | 0.31 | 0.79 |
| 200 | 0.65 | 0.49 | 0.35 | 0.35 | 0.47 | 1.39 | 0.30 | 0.35 | 1.22 |
| 300 | 0.67 | 0.38 | 0.25 | 0.29 | 0.40 | 1.05 | 0.25 | 0.33 | 1.07 |
| 500 | 0.81 | 0.38 | 0.34 | 0.31 | 0.36 | 0.65 | 0.20 | 0.20 | 1.31 |
| 1000 | 0.88 | 0.43 | 0.31 | 0.36 | 0.50 | 0.66 | 0.31 | 0.36 | 1.39 |
| 1500 | 0.62 | 0.30 | 0.13 | 0.21 | 0.36 | 0.40 | 0.14 | 0.20 | 1.09 |
| 2000 | 0.71 | 0.66 | 0.34 | 0.31 | 0.66 | 0.72 | 0.33 | 0.45 | 1.50 |
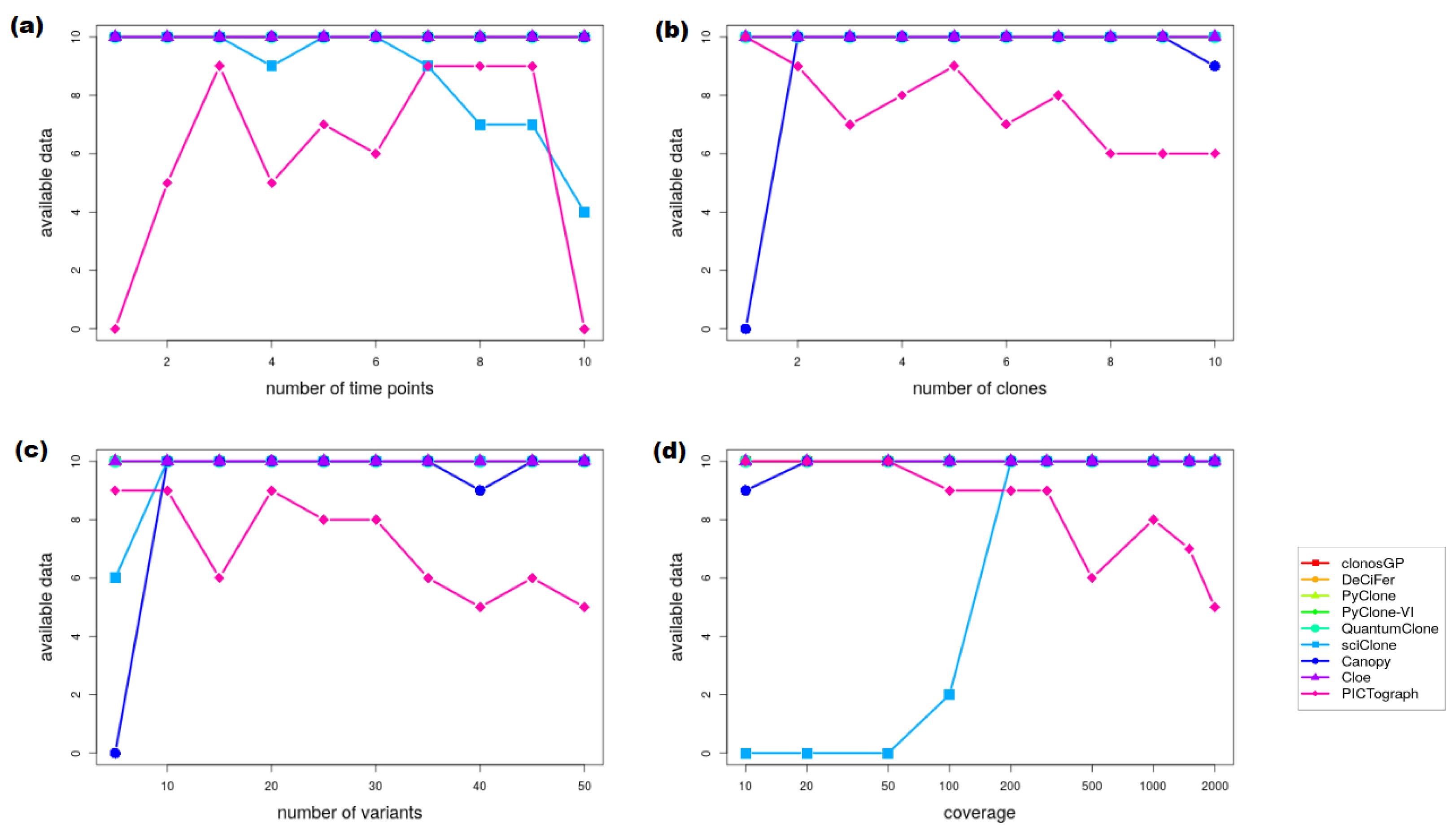
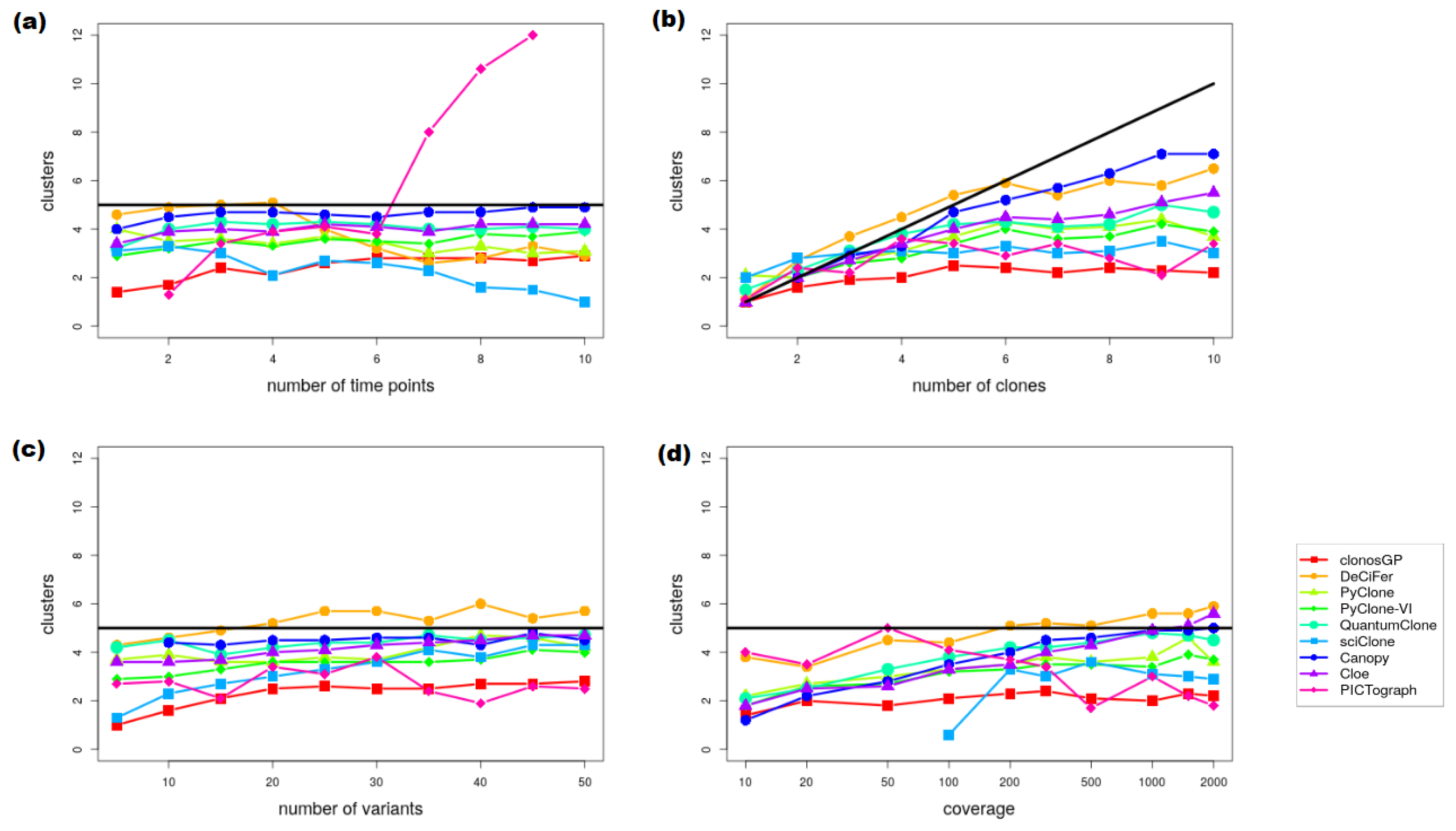
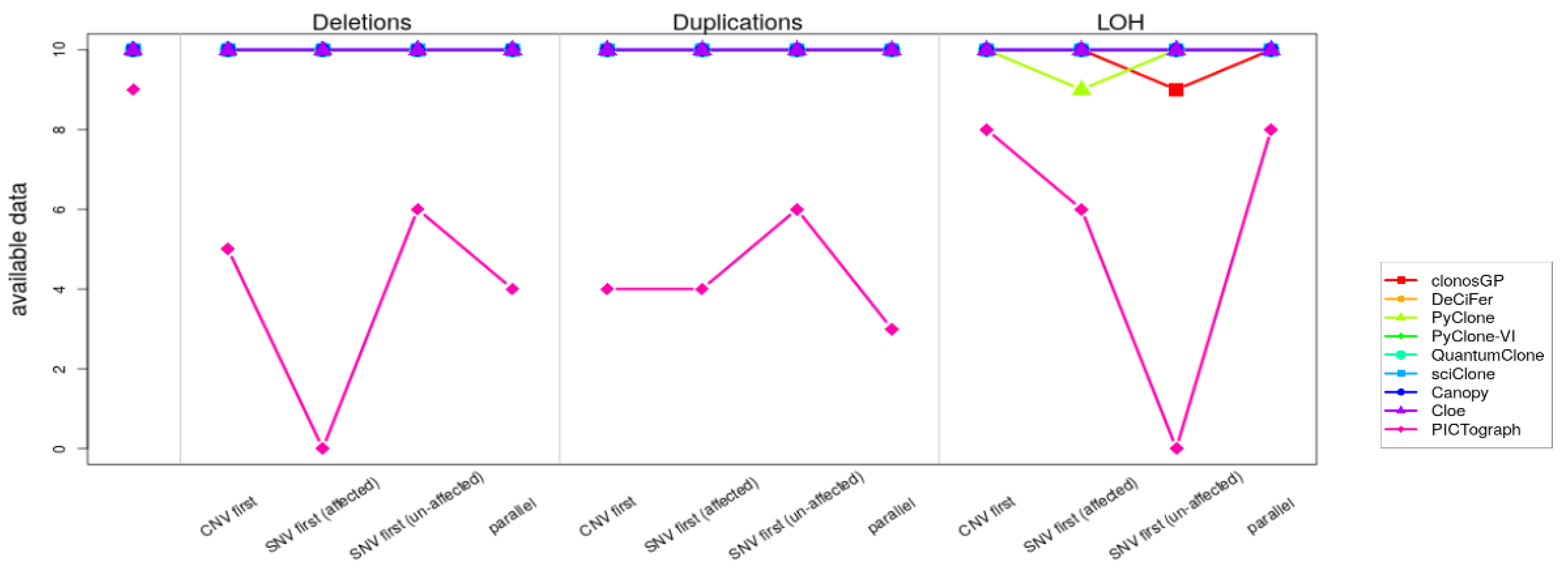
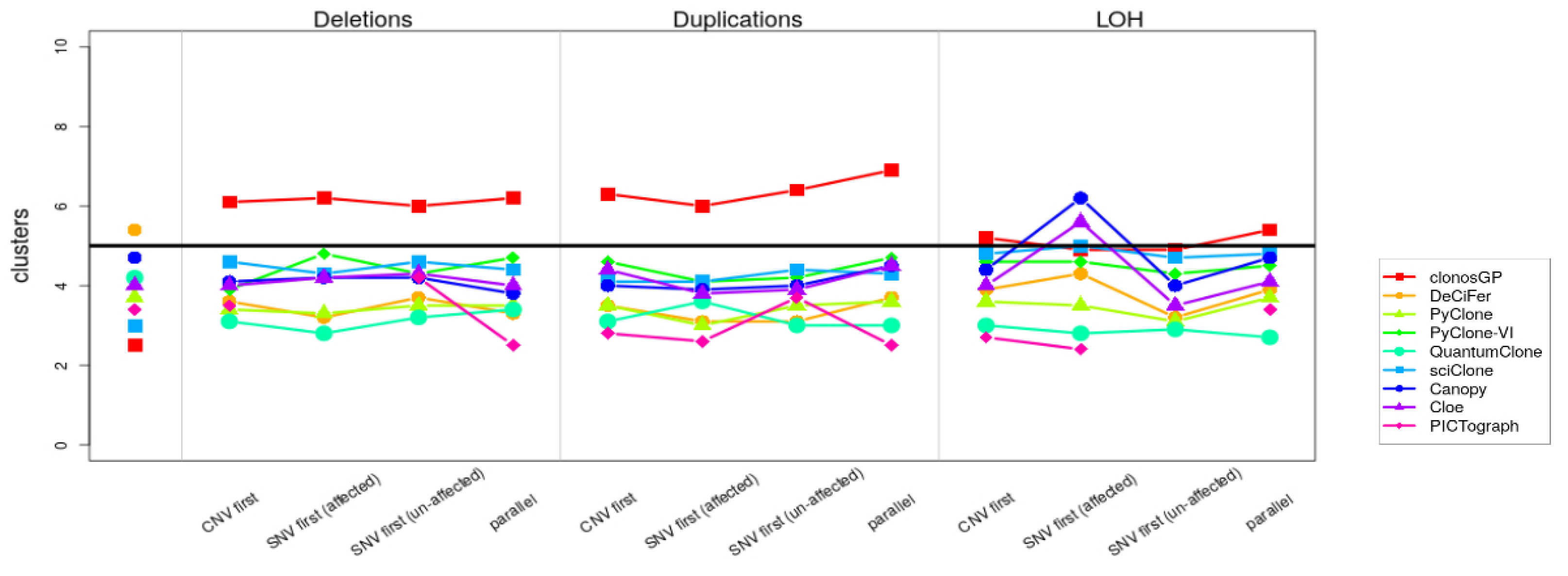
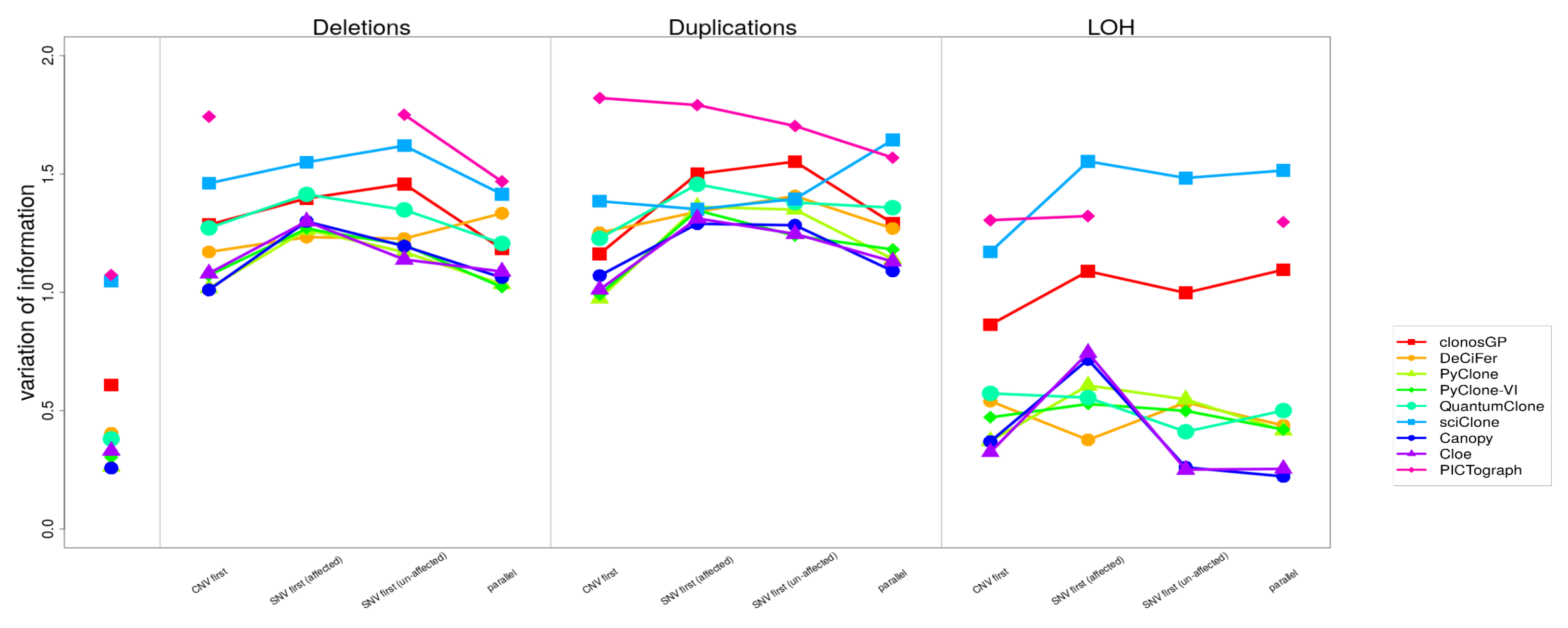
Appendix B.3. Variant Clustering in the Presence of CNVs
| Deletions | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Scenario of Overlap | clonosGP | DeCiFer | PyClone | PyClone-VI | QuantumClone | sciClone | Canopy | Cloe | PICTograph |
| CNV first | 1.29 | 1.17 | 1.02 | 1.07 | 1.27 | 1.46 | 1.01 | 1.08 | 1.74 |
| SNV first (affected) | 1.40 | 1.23 | 1.26 | 1.27 | 1.41 | 1.55 | 1.30 | 1.30 | NA |
| SNV first (un-affected) | 1.46 | 1.23 | 1.17 | 1.20 | 1.35 | 1.62 | 1.19 | 1.14 | 1.75 |
| Parallel | 1.18 | 1.33 | 1.03 | 1.02 | 1.21 | 1.41 | 1.06 | 1.09 | 1.47 |
| Duplications | |||||||||
| Scenario of Overlap | clonosGP | DeCiFer | PyClone | PyClone-VI | QuantumClone | sciClone | Canopy | Cloe | PICTograph |
| CNV first | 1.16 | 1.25 | 0.97 | 0.99 | 1.23 | 1.39 | 1.07 | 1.01 | 1.82 |
| SNV first (affected) | 1.50 | 1.34 | 1.36 | 1.35 | 1.46 | 1.35 | 1.29 | 1.31 | 1.79 |
| SNV first (un-affected) | 1.55 | 1.41 | 1.35 | 1.24 | 1.38 | 1.39 | 1.28 | 1.25 | 1.70 |
| Parallel | 1.29 | 1.27 | 1.14 | 1.18 | 1.36 | 1.64 | 1.09 | 1.13 | 1.57 |
| LOH | |||||||||
| Scenario of Overlap | clonosGP | DeCiFer | PyClone | PyClone-VI | QuantumClone | sciClone | Canopy | Cloe | PICTograph |
| CNV first | 0.86 | 0.54 | 0.37 | 0.47 | 0.57 | 1.17 | 0.37 | 0.32 | 1.30 |
| SNV first (affected) | 1.09 | 0.38 | 0.61 | 0.53 | 0.55 | 1.55 | 0.72 | 0.74 | 1.32 |
| SNV first (un-affected) | 1.00 | 0.53 | 0.55 | 0.50 | 0.41 | 1.48 | 0.26 | 0.25 | NA |
| Parallel | 1.10 | 0.44 | 0.42 | 0.42 | 0.50 | 1.51 | 0.22 | 0.25 | 1.30 |
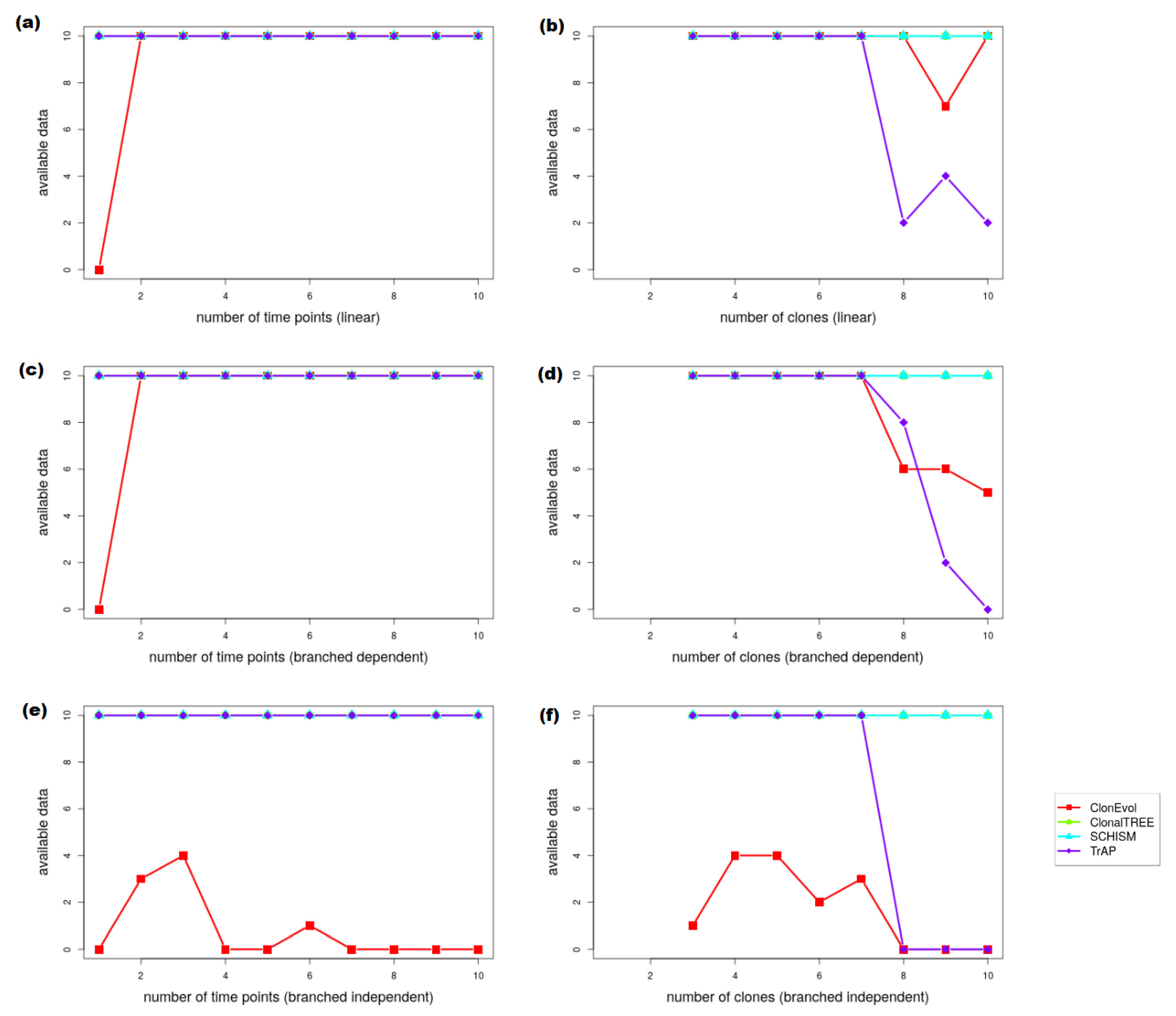
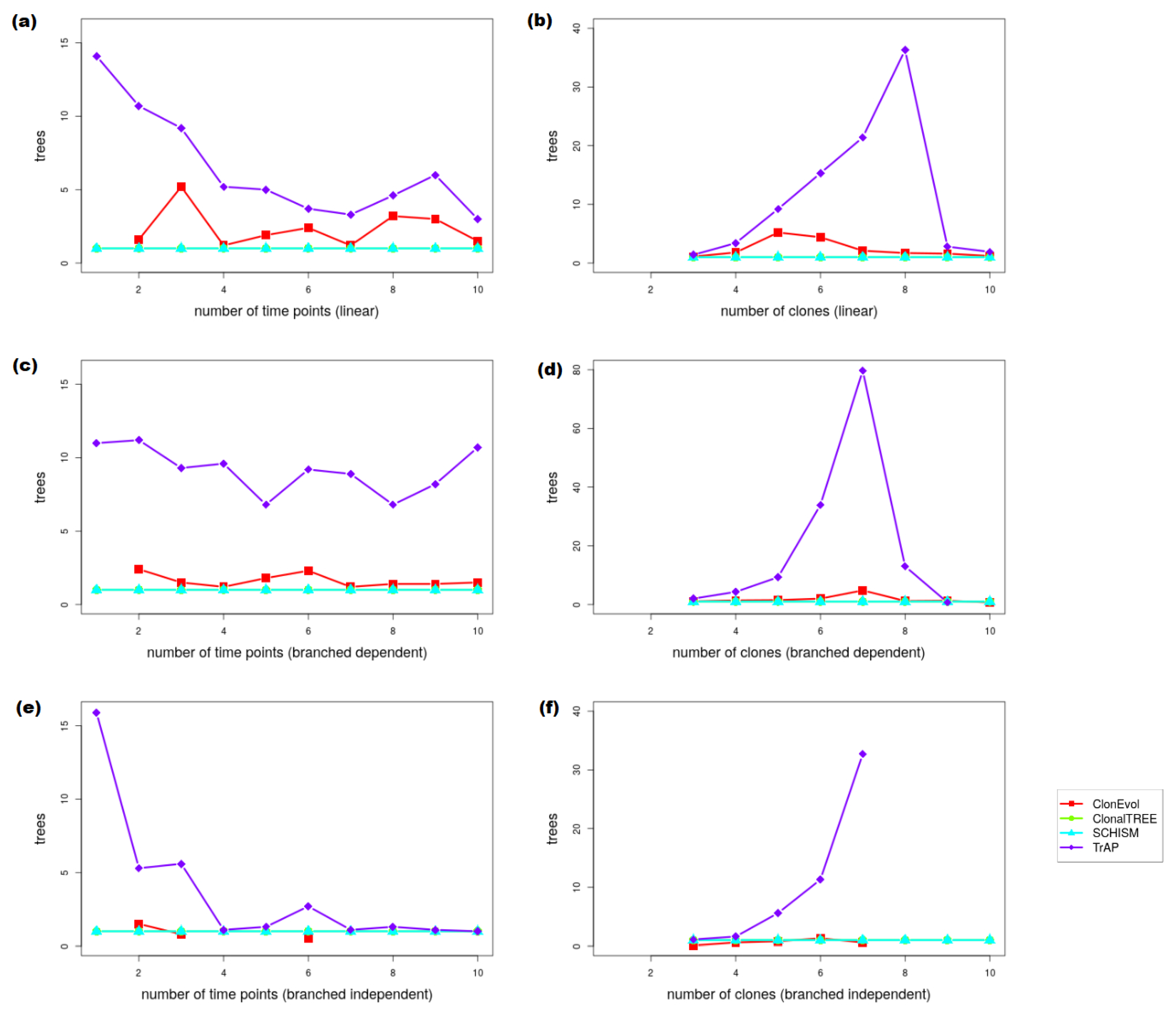
Appendix B.4. Clonal Evolution Tree Reconstruction
| Time Points (Linear) | Clones (Linear) | |||||||
|---|---|---|---|---|---|---|---|---|
| Values | ClonEvol | ClonalTREE | SCHISM | TrAP | ClonEvol | ClonalTREE | SCHISM | TrAP |
| 1 | NA | 1.73 | 2.82 | 0.82 | ||||
| 2 | 0.91 | 1.69 | 2.18 | 0.62 | ||||
| 3 | 0.83 | 1.3 | 1.88 | 0.61 | 0.48 | 1.11 | 0.3 | 0.32 |
| 4 | 0.79 | 0.93 | 0.79 | 0.36 | 0.82 | 1.35 | 0.87 | 0.40 |
| 5 | 0.57 | 1.43 | 1.12 | 0.43 | 0.83 | 1.3 | 1.88 | 0.61 |
| 6 | 0.41 | 1.07 | 0.53 | 0.27 | 1.39 | 2.29 | 2.51 | 1.36 |
| 7 | 0.48 | 1.42 | 0.71 | 0.26 | 1.8 | 1.97 | 2.97 | 1.79 |
| 8 | 0.56 | 1.39 | 0.92 | 0.50 | 1.7 | 1.43 | 2.7 | 1.22 |
| 9 | 0.47 | 1.44 | 0.68 | 0.42 | 2.11 | 2.43 | 3.91 | 1.86 |
| 10 | 0.44 | 1.5 | 0.64 | 0.33 | 2.48 | 2.43 | 3.12 | 2.22 |
| Time Points (Branched Dependent) | Clones (Branched Dependent) | |||||||
| Values | ClonEvol | ClonalTREE | SCHISM | TrAP | ClonEvol | ClonalTREE | SCHISM | TrAP |
| 1 | NA | 2.17 | 2.43 | 0.84 | ||||
| 2 | 1.3 | 1.61 | 1.77 | 0.84 | ||||
| 3 | 1.29 | 1.82 | 1.51 | 1.21 | 0.69 | 1.68 | 0.31 | 0.79 |
| 4 | 1.24 | 1.94 | 0.69 | 1.02 | 0.91 | 2.22 | 0.93 | 0.71 |
| 5 | 0.88 | 1.67 | 1.18 | 0.88 | 1.29 | 1.82 | 1.51 | 1.21 |
| 6 | 1.22 | 1.85 | 1.08 | 0.90 | 1.66 | 2.3 | 2.48 | 1.39 |
| 7 | 1.15 | 1.59 | 0.78 | 1.12 | 1.48 | 1.8 | 2.36 | 1.26 |
| 8 | 0.8 | 2.01 | 0.52 | 0.91 | 1.77 | 2 | 2.03 | 1.29 |
| 9 | 1.03 | 1.76 | 0.72 | 0.91 | 2.58 | 2.48 | 1.93 | 1.84 |
| 10 | 1 | 1.38 | 0.88 | 1.16 | 3.84 | 1.74 | 1.81 | NA |
| Time Points (Branched Independent) | Clones (Branched Independent) | |||||||
| Values | ClonEvol | ClonalTREE | SCHISM | TrAP | ClonEvol | ClonalTREE | SCHISM | TrAP |
| 1 | NA | 1.37 | 3.81 | 2.02 | ||||
| 2 | 1.55 | 1.18 | 3.76 | 1.91 | ||||
| 3 | 1.2 | 1.47 | 2.98 | 1.98 | 1.02 | 0.4 | 0.98 | 1.07 |
| 4 | NA | 1.08 | 3.42 | 1.73 | 0.7 | 0.81 | 2.03 | 1.22 |
| 5 | NA | 1.23 | 4.45 | 1.31 | 1.2 | 1.47 | 2.98 | 1.98 |
| 6 | 1.16 | 1.17 | 3.1 | 1.46 | 1.54 | 1.32 | 5.63 | 2.69 |
| 7 | NA | 0.76 | 3.97 | 1.72 | 1.8 | 1.71 | 7 | 3.46 |
| 8 | NA | 1.07 | 4.13 | 1.60 | NA | 1.72 | 8.43 | NA |
| 9 | NA | 1.01 | 4.64 | 1.75 | NA | 1.78 | 12.54 | NA |
| 10 | NA | 0.92 | 4.39 | 1.55 | NA | 1.78 | 9.15 | NA |
References
- Global Cancer Observatory: Cancer Today. International Agency for Research on Cancer. Available online: https://gco.iarc.fr/today (accessed on 29 January 2023).
- World Cancer Research Fund International. Cancer Survival Statistics. Available online: https://www.wcrf.org/cancer-trends/cancer-survival-statistics/ (accessed on 29 January 2023).
- Alexander, T.B.; Wang, L.; Inaba, H.; Triplett, B.M.; Pounds, S.; Ribeiro, R.C.; Pui, C.H.; Rubnitz, J.E. Decreased relapsed rate and treatment-related mortality contribute to improved outcomes for pediatric acute myeloid leukemia in successive clinical trials. Cancer 2017, 123, 3791–3798. [Google Scholar] [CrossRef]
- Rockberg, J.; Amelio, J.M.; Taylor, A.; Jörgensen, L.; Ragnhammar, P.; Hansson, J. Epidemiology of cutaneous melanoma in Sweden-Stage-specific survival and rate of recurrence. Int. J. Cancer 2016, 139, 2722–2729. [Google Scholar] [CrossRef]
- Das, S.; Kundu, M.; Jena, B.C.; Mandal, M. Chapter 25—Causes of cancer: Physical, chemical, biological carcinogens, and viruses. In Materials Today: Biomaterials for 3D Tumor Modeling; Kundu, S.C., Reis, R.L., Eds.; Elsevier: Amsterdam, The Netherlands, 2020; pp. 607–641. [Google Scholar]
- Greenberg, P.L.; Tuechler, H.; Schanz, J.; Sanz, G.; Garcia-Manero, G.; Solé, F.; Bennett, J.M.; Bowen, D.; Fenaux, P.; Dreyfus, F.; et al. Revised international prognostic scoring system for myelodysplastic syndromes. Blood 2012, 120, 2454–2465. [Google Scholar] [CrossRef]
- Malcovati, L.; Crouch, S.; De Graaf, A.O.; Sandmann, S.; Tobiasson, M.; Kosmider, O.; van der Reijden, B.A.; Painter, D.; Van de Loosdrecht, A.A.; Symeonidis, A.; et al. Mutation Profiles Identify Distinct Clusters of Lower Risk Myelodysplastic Syndromes with Unique Clinical and Biological Features and Clinical Endpoints. Blood 2020, 136, 29. [Google Scholar] [CrossRef]
- Reutter, K.; Sandmann, S.; Rohde, J.; Müller, S.; Wöste, M.; Khanam, T.; Michgehl, U.; Klapper, W.; Wößmann, W.; Seggewiß, J.; et al. Reconstructing clonal evolution in relapsed and non-relapsed Burkitt lymphoma. Leukemia 2021, 35, 639–643. [Google Scholar] [CrossRef] [PubMed]
- Byun, J.M.; Koh, Y.; Shin, D.Y.; Kim, I.; Yoon, S.S.; Lee, J.O.; Bang, S.M.; Kim, K.H.; Jung, S.H.; Lee, W.S.; et al. BCR-ABL translocation as a favorable prognostic factor in elderly patients with acute lymphoblastic leukemia in the era of potent tyrosine kinase inhibitors. Haematologica 2017, 102, e187–e190. [Google Scholar] [CrossRef]
- Gerlinger, M.; Rowan, A.J.; Horswell, S.; Math, M.; Larkin, J.; Endesfelder, D.; Gronroos, E.; Martinez, P.; Matthews, N.; Stewart, A.; et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med. 2012, 366, 883–892. [Google Scholar] [CrossRef]
- Zhang, J.; Fujimoto, J.; Zhang, J.; Wedge, D.C.; Song, X.; Zhang, J.; Seth, S.; Chow, C.W.; Cao, Y.; Gumbs, C.; et al. Intratumor heterogeneity in localized lung adenocarcinomas delineated by multiregion sequencing. Science 2014, 346, 256–259. [Google Scholar] [CrossRef] [PubMed]
- Luquette, L.J.; Bohrson, C.L.; Sherman, M.A.; Park, P.J. Identification of somatic mutations in single cell DNA-seq using a spatial model of allelic imbalance. Nat. Commun. 2019, 10, 3908. [Google Scholar] [CrossRef] [PubMed]
- Navin, N.E. Cancer genomics: One cell at a time. Genome Biol. 2014, 15, 452. [Google Scholar] [CrossRef]
- Sandmann, S.; Behrens, Y.L.; Davenport, C.; Thol, F.; Heuser, M.; Dörfel, D.; Löhr, F.; Castrup, A.; Steinemann, D.; Varghese, J.; et al. Clonal Evolution at First Sight: A Combined Visualization of Diverse Diagnostic Methods Improves Understanding of Leukemic Progression. Front. Oncol. 2022, 12, 888114. [Google Scholar] [CrossRef]
- Ding, L.; Raphael, B.J.; Chen, F.; Wendl, M.C. Advances for studying clonal evolution in cancer. Cancer Lett. 2013, 340, 212–219. [Google Scholar] [CrossRef] [PubMed]
- Da Silva-Coelho, P.; Kroeze, L.I.; Yoshida, K.; Koorenhof-Scheele, T.N.; Knops, R.; van de Locht, L.T.; de Graaf, A.O.; Massop, M.; Sandmann, S.; Dugas, M.; et al. Clonal evolution in myelodysplastic syndromes. Nat. Commun. 2017, 8, 15099. [Google Scholar] [CrossRef] [PubMed]
- Sandmann, S.; Richter, S.; Jiang, X.; Varghese, J. Exploring Current Challenges and Perspectives for Automatic Reconstruction of Clonal Evolution. Cancer Genom. Proteom. 2022, 19, 194–204. [Google Scholar] [CrossRef] [PubMed]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2021; Available online: https://www.R-project.org/ (accessed on 29 January 2023).
- Davies, A.; Gao, R.; Navin, N. Tumor evolution: Linear, branching, neutral or punctuated? Biochim. Biophys. Acta Rev. Cancer 2017, 1867, 151–161. [Google Scholar] [CrossRef]
- Sandmann, S.; Inserte, C.; Varghese, J. clevRvis: Visualization Techniques for Clonal Evolution. GigaScience 2022, accepted. [Google Scholar]
- Kimura, M.; Crow, J.F. He number of alleles that can be maintained in a finite population. Genetics 1964, 49, 725–738. [Google Scholar] [CrossRef]
- Fischer, A.; Vázquez-García, I.; Illingworth, C.J.R.; Mustonen, V. High-definition reconstruction of clonal composition in cancer. Cell Rep. 2014, 7, 1740–1752. [Google Scholar] [CrossRef]
- Vavoulis, D.V.; Cutts, A.; Taylor, J.C.; Schuh, A. A statistical approach for tracking clonal dynamics in cancer using longitudinal next-generation sequencing data. Bioinformatics 2021, 37, 147–154. [Google Scholar] [CrossRef]
- Satas, G.; Zaccaria, S.; El-Kebir, M.; Raphael, B. DeCiFering the elusive cancer cell fraction in tumor heterogeneity and evolution. Cell Syst. 2021, 12, 1004–1018.e10. [Google Scholar] [CrossRef] [PubMed]
- Roth, A.; Khattra, J.; Yap, D.; Wan, A.; Laks, E.; Biele, J.; Ha, G.; Aparicio, S.; Bouchard-Côté, A.; Shah, S.P. PyClone: Statistical inference of clonal population structure in cancer. Nat. Methods 2014, 11, 396–398. [Google Scholar] [CrossRef] [PubMed]
- Gillis, S.; Roth, A. PyClone-VI: Scalable inference of clonal population structures using whole genome data. BMC Bioinform. 2020, 21, 571. [Google Scholar] [CrossRef] [PubMed]
- Deveau, P.; Colmet Daage, L.; Oldridge, D.; Bernard, V.; Bellini, A.; Chicard, M.; Clement, N.; Lapouble, E.; Combaret, V.; Boland, A.; et al. QuantumClone: Clonal assessment of functional mutations in cancer based on a genotype-aware method for clonal reconstruction. Bioinformatics 2018, 34, 1808–1816. [Google Scholar] [CrossRef]
- Miller, C.A.; White, B.S.; Dees, N.D.; Griffith, M.; Welch, J.S.; Griffith, O.L.; Vij, R.; Tomasson, M.H.; Graubert, T.A.; Walter, M.J.; et al. SciClone: Inferring Clonal Architecture and Tracking the Spatial and Temporal Patterns of Tumor Evolution. PLoS Comput Biol. 2014, 10, e1003665. [Google Scholar] [CrossRef]
- Jiang, Y.; Qiu, Y.; Minn, A.J.; Zhang, N.R. Assessing intratumor heterogeneity and tracking longitudinal and spatial clonal evolutionary history by next-generation sequencing. Proc. Natl. Acad. Sci. USA 2016, 113, E5528–E5537. [Google Scholar] [CrossRef]
- Marass, F.; Mouliere, F.; Yuan, K.; Rosenfeld, N.; Markowetz, F. A phylogenetic latent feature model for clonal deconvolution. Ann. Appl. Stat. 2016, 10, 2377–2404. [Google Scholar] [CrossRef]
- Popic, V.; Salari, R.; Hajirasouliha, I.; Kashef-Haghighi, D.; West, R.B.; Batzoglou, S. Fast and scalable inference of multi-sample cancer lineages. Genome Biol. 2015, 16, 91. [Google Scholar] [CrossRef]
- El-Kebir, M.; Satas, G.; Oesper, L.; Raphael, B.J. Inferring the Mutational History of a Tumor Using Multi-state Perfect Phylogeny Mixtures. Cell Syst. 2016, 3, 43–53. [Google Scholar] [CrossRef] [PubMed]
- Zheng, L.; Niknafs, N.; Wood, L.D.; Karchin, R.; Scharpf, R.B. Estimation of cancer cell fractions and clone trees from multi-region sequencing of tumors. Bioinformatics 2022, 38, 3677–3683. [Google Scholar] [CrossRef] [PubMed]
- Dang, H.X.; White, B.S.; Foltz, S.M.; Miller, C.A.; Luo, J.; Fields, R.C.; Maher, C.A. ClonEvol: Clonal ordering and visualization in cancer sequencing. Ann. Oncol. 2017, 28, 3076–3082. [Google Scholar] [CrossRef] [PubMed]
- Ismail, W.M.; Tang, H. Clonal reconstruction from time course genomic sequencing data. BMC Genom. 2019, 20, 1002. [Google Scholar] [CrossRef] [PubMed]
- Niknafs, N.; Beleva-Guthrie, V.; Naiman, D.Q.; Karchin, R. SubClonal Hierarchy Inference from Somatic Mutations: Automatic Reconstruction of Cancer Evolutionary Trees from Multi-region Next Generation Sequencing. PLoS Comput Biol. 2015, 11, e1004416. [Google Scholar] [CrossRef]
- Strino, F.; Parisi, F.; Micsinai, M.; Kluger, Y. TrAp: A tree approach for fingerprinting subclonal tumor composition. Nucleic Acids Res. 2013, 41, e165. [Google Scholar] [CrossRef]
- Ismail, W.M.; Tang, H. A scalable algorithm for clonal reconstruction from sparse time course genomic sequencing data. bioRxiv 2021. [Google Scholar] [CrossRef]
- Sundermann, L.K.; Wintersinger, J.; Rätsch, G.; Stoye, J.; Morris, Q. Reconstructing tumor evolutionary histories and clone trees in polynomial-time with SubMARine. PLoS Comput Biol. 2021, 17, e1008400. [Google Scholar] [CrossRef] [PubMed]
- Marchant, N.; Steorts, R. clevr: Clustering and Link Prediction Evaluation in R. version 0.1.1. 2020. Available online: https://CRAN.R-project.org/package=clevr (accessed on 29 January 2023).
- You, K. NetworkDistance: Distance Measures for Networks. version 0.3.4. 2021. Available online: https://CRAN.R-project.org/package=NetworkDistance (accessed on 29 January 2023).
- Csardi, G.; Nepusz, T. The igraph software package for complex network research. Inter J. Complex Syst. 2006, 1695, 1–9. [Google Scholar]


| Data Set | Model of | #Time Points | #Clones | #SNVs | Mean Coverage | #CNVs |
|---|---|---|---|---|---|---|
| Clonal Evolution | ||||||
| sim01–sim10 | linear | 1–10 | 5 | 20 | 300x | 0 |
| sim11–sim20 | linear | 3 | 1–10 | 20 | 300x | 0 |
| sim21–sim30 | linear | 3 | 5 | 5–50 (step 5) | 300x | 0 |
| sim31–sim40 | linear | 3 | 5 | 20 | 10x, 20x, 50x, 100x, 200x, | 0 |
| 300x, 500x, 1000x, 1500x, 2000x | ||||||
| sim41–sim44 | linear | 3 | 5 | 20 | 300x | 6 (del) |
| sim45–sim48 | linear | 3 | 5 | 20 | 300x | 6 (dup) |
| sim49–sim52 | linear | 3 | 5 | 20 | 300x | 6 (LOH) |
| sim53–sim62 | branched dependent | 1–10 | 5 | 20 | 300x | 0 |
| sim63–sim70 | branched dependent | 3 | 3–10 | 20 | 300x | 0 |
| sim71–sim80 | branched independent | 1–10 | 5 | 20 | 300x | 0 |
| sim81–sim88 | branched independent | 3 | 3–10 | 20 | 300x | 0 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sandmann, S.; Richter, S.; Jiang, X.; Varghese, J. Reconstructing Clonal Evolution—A Systematic Evaluation of Current Bioinformatics Approaches. Int. J. Environ. Res. Public Health 2023, 20, 5128. https://doi.org/10.3390/ijerph20065128
Sandmann S, Richter S, Jiang X, Varghese J. Reconstructing Clonal Evolution—A Systematic Evaluation of Current Bioinformatics Approaches. International Journal of Environmental Research and Public Health. 2023; 20(6):5128. https://doi.org/10.3390/ijerph20065128
Chicago/Turabian StyleSandmann, Sarah, Silja Richter, Xiaoyi Jiang, and Julian Varghese. 2023. "Reconstructing Clonal Evolution—A Systematic Evaluation of Current Bioinformatics Approaches" International Journal of Environmental Research and Public Health 20, no. 6: 5128. https://doi.org/10.3390/ijerph20065128
APA StyleSandmann, S., Richter, S., Jiang, X., & Varghese, J. (2023). Reconstructing Clonal Evolution—A Systematic Evaluation of Current Bioinformatics Approaches. International Journal of Environmental Research and Public Health, 20(6), 5128. https://doi.org/10.3390/ijerph20065128







