Heterologous Expression of the Phytochelatin Synthase CaPCS2 from Chlamydomonas acidophila and Its Effect on Different Stress Factors in Escherichia coli
Abstract
:1. Introduction
2. Materials and Methods
2.1. Bacterial Strains and Growth Conditions
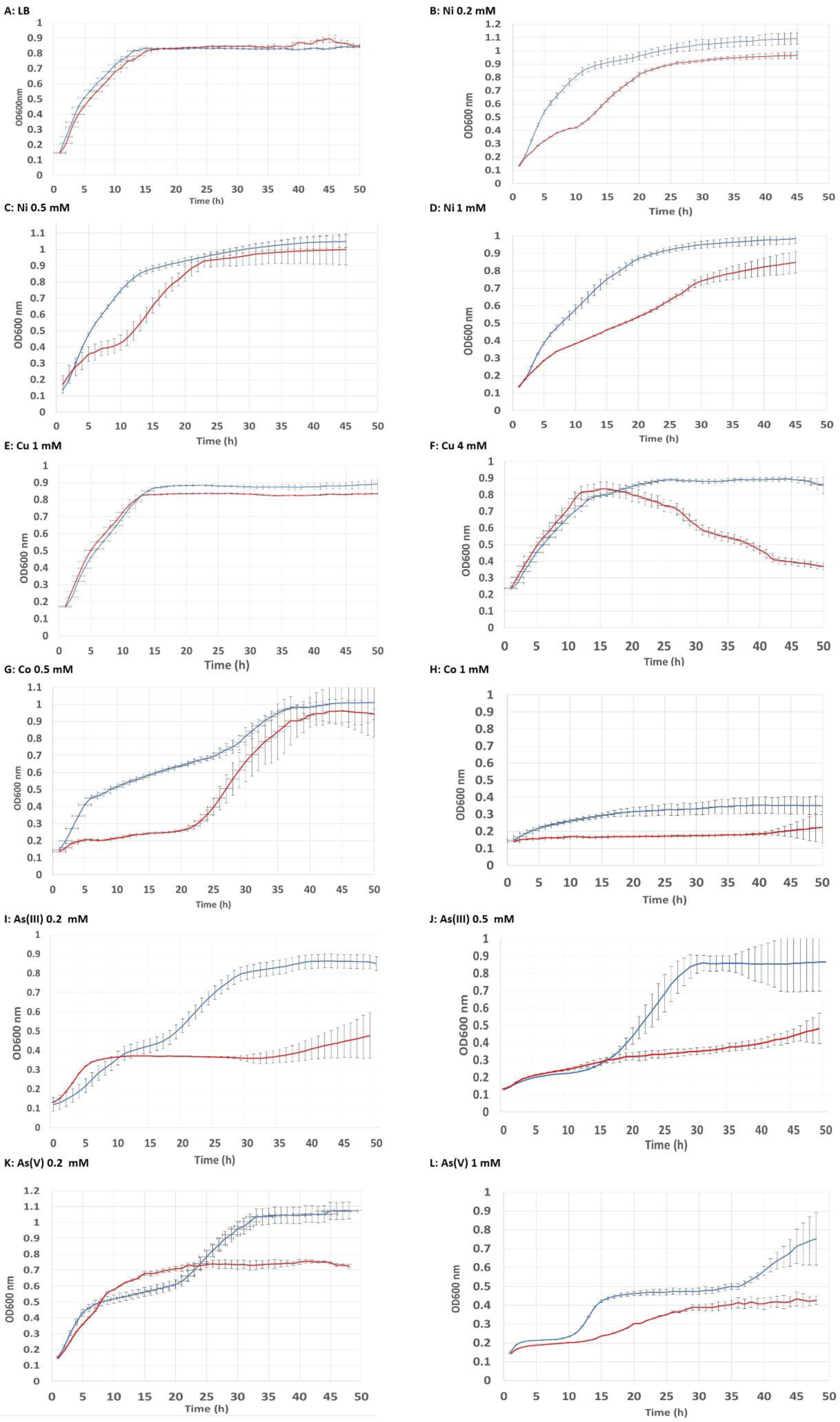
2.2. Analysis of Exposure to Metal(loid)s, Salt, and Acid pH by Growth Curve Tests
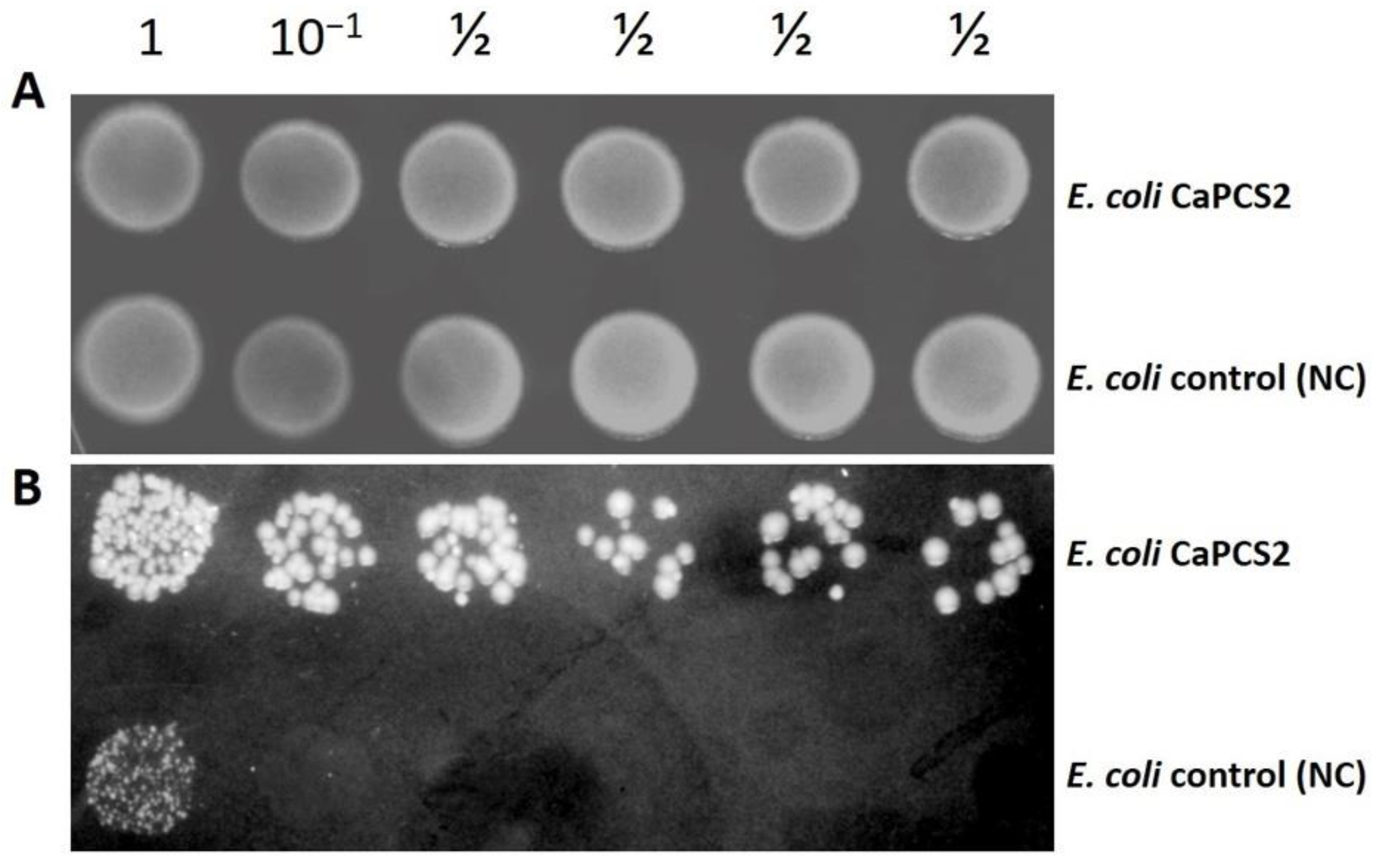
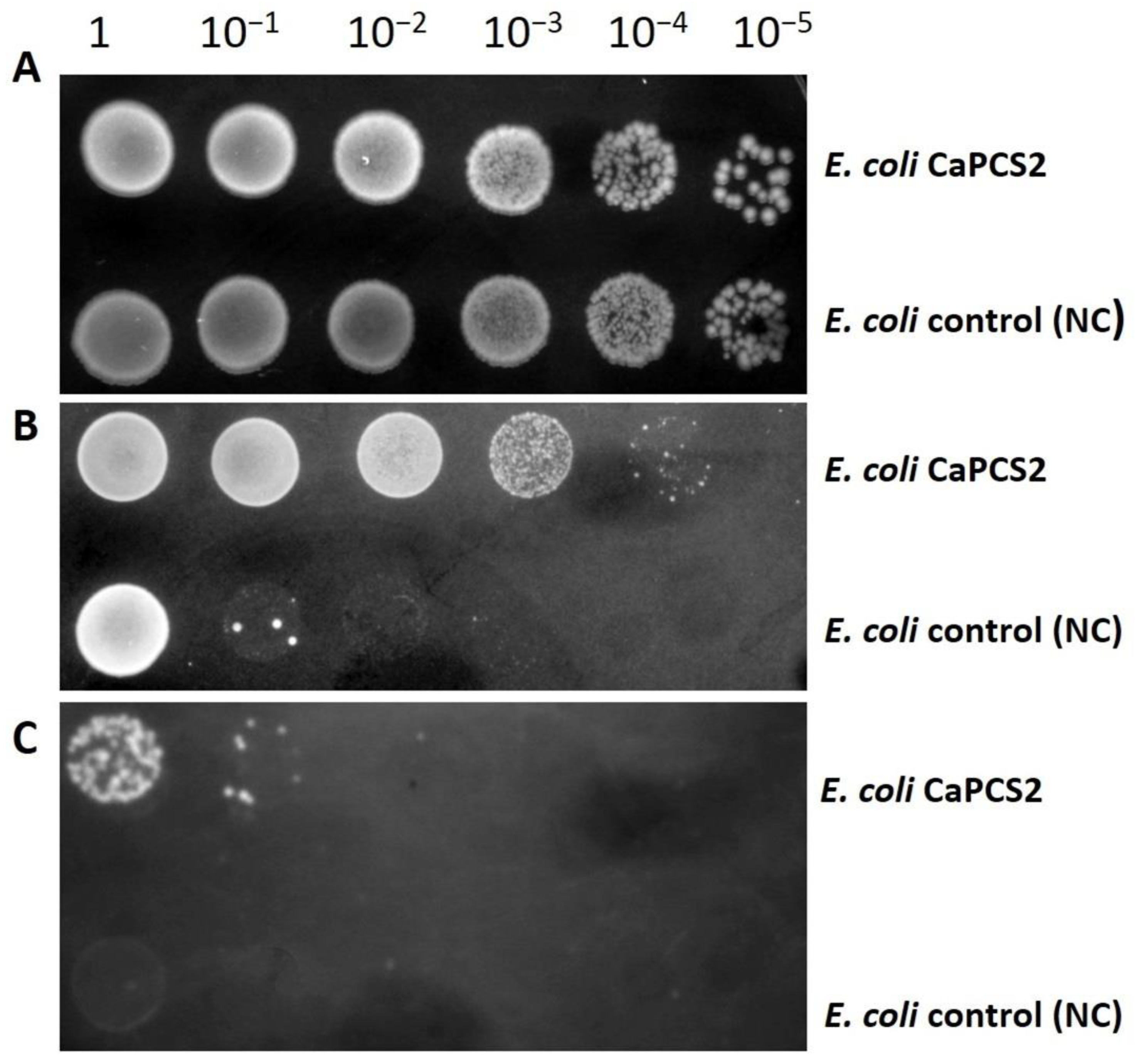
2.3. Test for Resistance to UVB Radiation and Exposure to Perchlorate Using the Droplet Test
2.4. Determination of Cellular Metal Concentration Using Inductively Coupled Plasma Spectroscopy-Mass Spectrometry (ICP-MS) Quantitative Analysis
2.5. Statistics
2.6. Gene Characterization
2.7. Phylogenetic Analysis
3. Results
3.1. Tolerance to Metal(loid) Exposure
3.2. Tolerance to UVB Radiation and Exposure to Perchlorate by Droplet Test
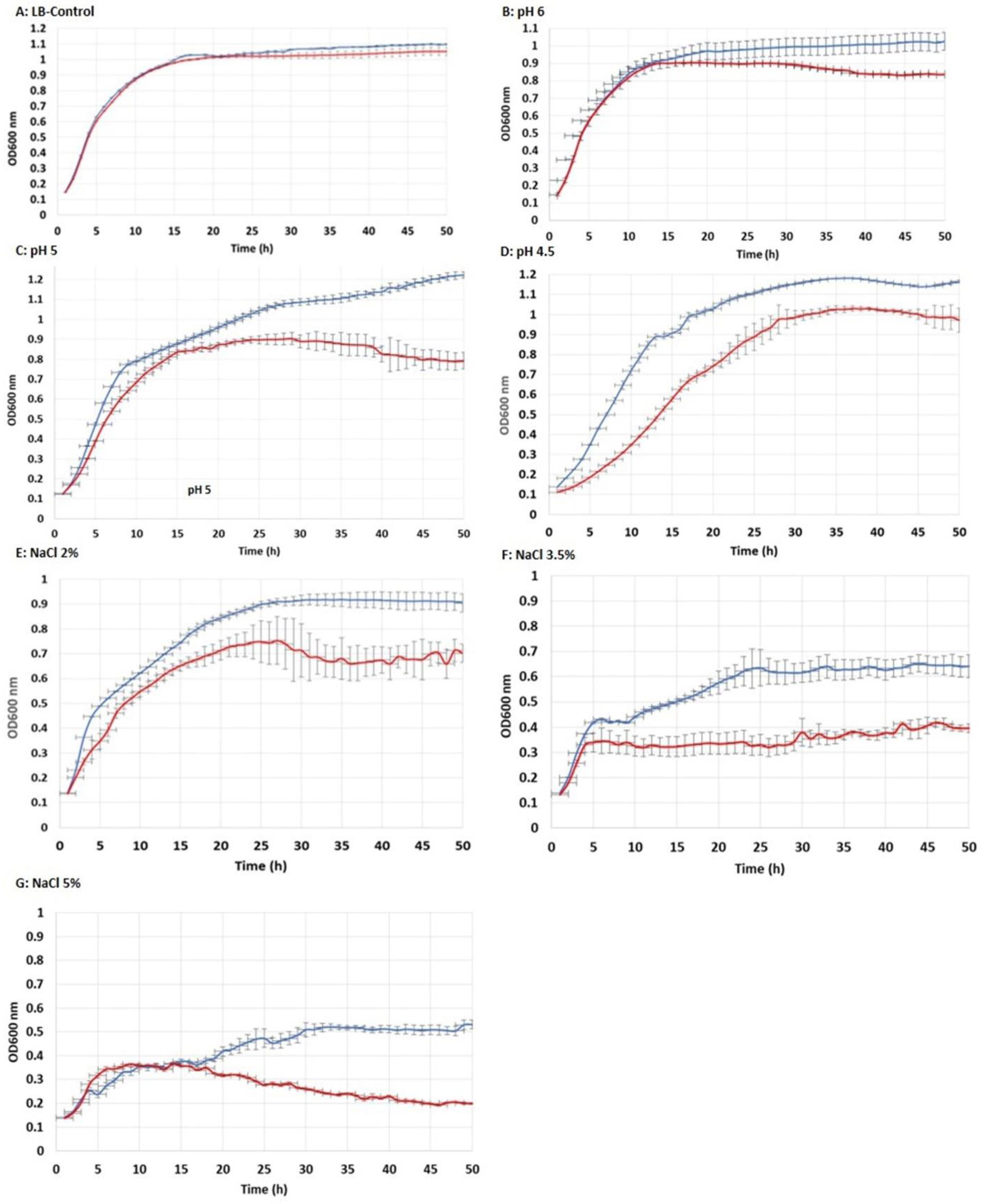
3.3. Tolerance to Acidic pH and Salt Exposure
3.4. Determination of Cellular Metal Concentration
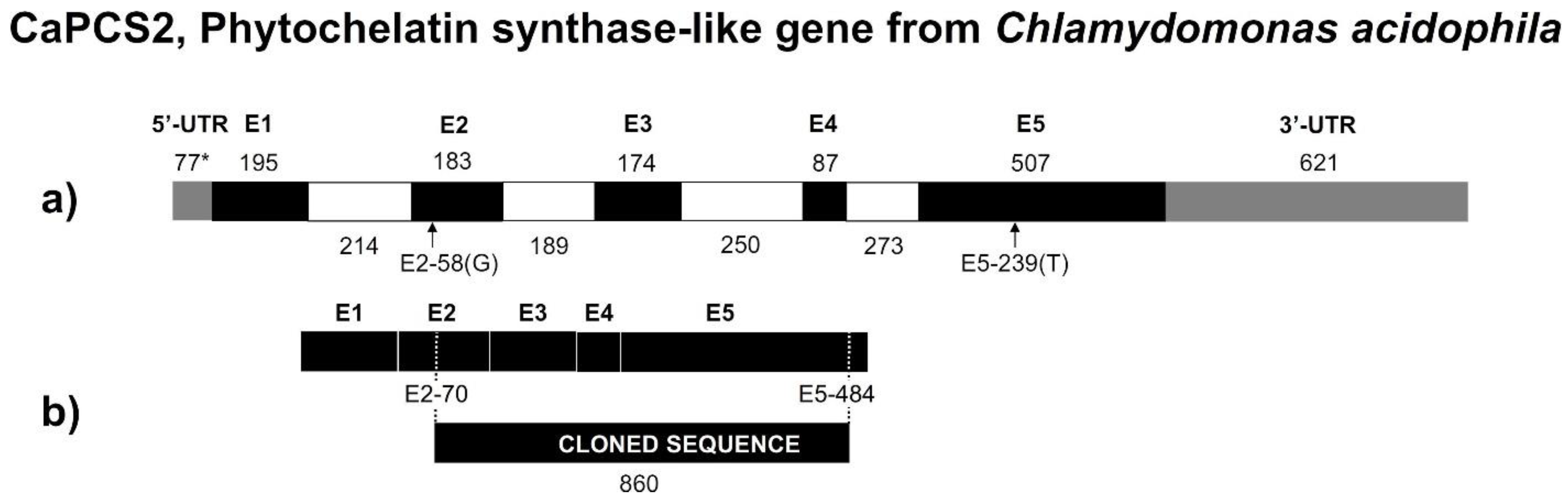
3.5. CaPCS2 Gene from Chlamydomonas Acidophila
4. Discussion
4.1. Role of Phytochelatin Sythases in Tolerance to Abiotic Stressors
4.2. Phytochelatin Sythetase and Heavy Metal(loid)s Bioaccumulation
5. Conclusions
- Heterologous expression of the PCS gene from C. acidophila (CaPCS2) confers resistance in E. coli to several metal(loid)s that directly or indirectly produce ROS. This PCS is not involved in the bioaccumulation of metal(loid)s in the cell.
- The E. coli strain transformed with the CaPCS2 gene showed increased resistance to exposure to various cellular stress factors, such as the presence of perchlorate and exposure to UVB radiation. This strain also exhibited greater tolerance to the presence of salt and acidic pH in the culture medium than the control strain.
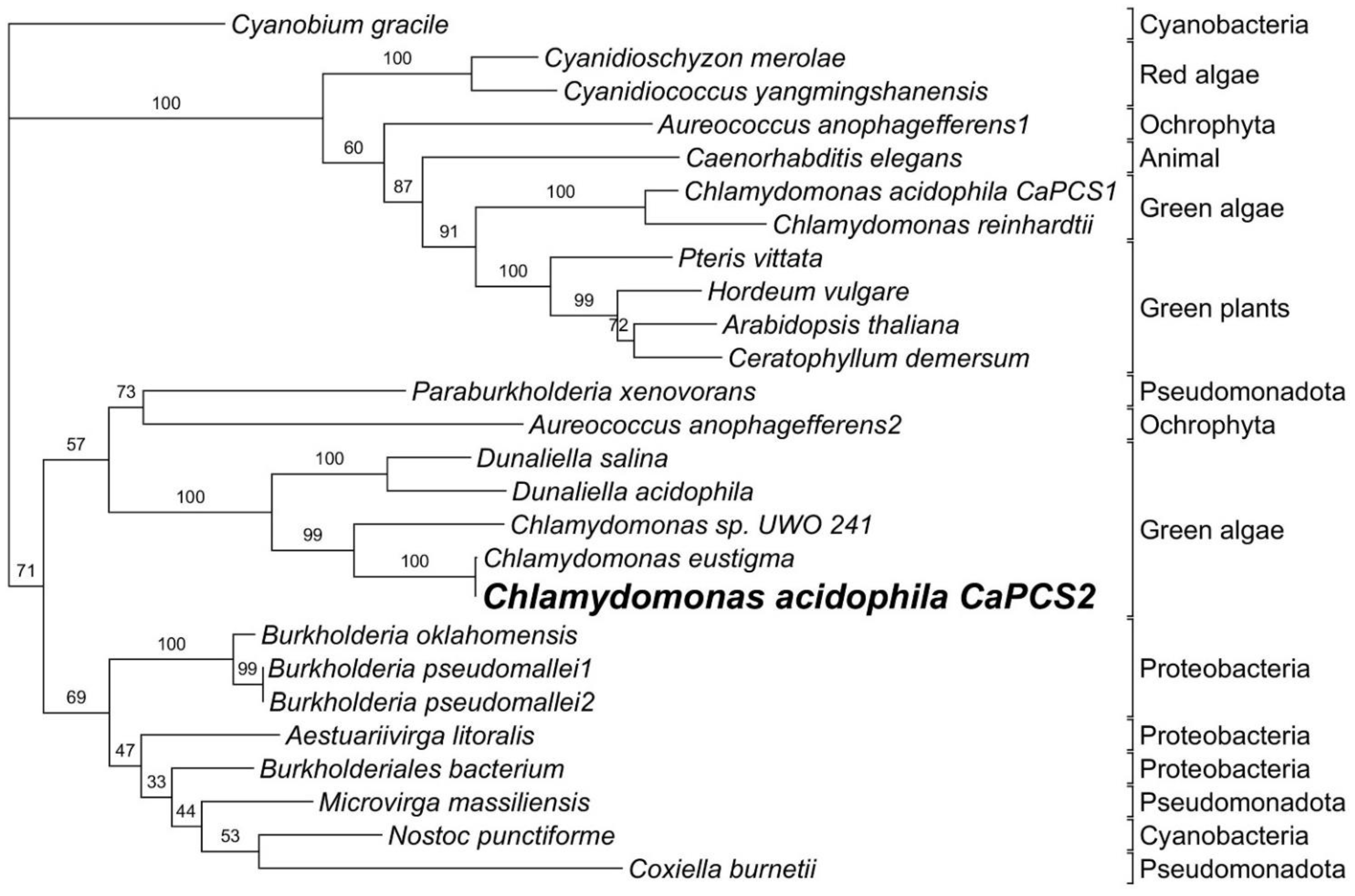
- The phylogenetic analysis of the CaPCS2 gene showed characteristics intermediate to eukaryotic and prokaryotic PCS genes. The amino acid sequence suggests closer relationships to bacterial PCS than to plant PCS. However, the complete gene sequence contains introns, such as eukaryotic genes.
- The strictly conserved Cys residues and catalytic triad (Cys56, His162, and Asp180) are also present in CaPCS2 (Cys70, His189, and Asp207). CaPCS2 lacks four of the five conserved Cys residues at the N-terminus, such as bacterial PCS.
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Cobbett, C.; Goldsbrough, P. Phytochelatins and metallothioneins: Roles in heavy metal detoxification and homeostasis. Annu. Rev. Plant Biol. 2002, 53, 159–182. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Grill, E.; Winnacker, E.L.; Zenk, M.H. Phytochelatins: The principal heavy-metal complexing peptides of higher plants. Science 1985, 230, 674–676. [Google Scholar] [CrossRef] [PubMed]
- Kondo, N.; Imai, K.; Isobe, M.; Goto, T.; Murasugi, A.; Wada-Nakagawa, C.; Hayashi, Y. Cadystin A and B, major unit peptides comprising cadmium binding peptides induced in a fission yeast—Separation, revision of structures and synthesis. Tetrahedron Lett. 1984, 25, 3869–3872. [Google Scholar] [CrossRef]
- Clemens, S.; Peršoh, D. Multi-tasking phytochelatin synthases. Plant Sci. 2009, 177, 266–271. [Google Scholar] [CrossRef]
- Petraglia, A.; De Benedictis, M.; Degola, F.; Pastore, G.; Calcagno, M.; Ruotolo, R.; Mengoni, A.; di Toppi, L.S. The capability to synthesize phytochelatins and the presence of constitutive and functional phytochelatin synthases are ancestral (plesiomorphic) characters for basal land plants. J. Exp. Bot. 2014, 65, 1153–1163. [Google Scholar] [CrossRef] [Green Version]
- Rea, P.A. Phytochelatin synthase: Of a protease a peptide polymerase made. Physiol. Plant 2012, 145, 154–164. [Google Scholar] [CrossRef]
- Fontanini, D.; Andreucci, A.; Castiglione, M.R.; Basile, A.; Sorbo, S.; Petraglia, A.; Degola, F.; Bellini, E.; Bruno, L.; Varotto, C.; et al. The phytochelatin synthase from Nitella mucronata (Charophyta) plays a role in the homeostatic control of iron(II)/(III). Plant Physiol. Biochem. 2018, 127, 88–96. [Google Scholar] [CrossRef]
- Gupton-Campolongo, T.; Damasceno, L.M.; Hay, A.G.; Ahner, B.A. Characterization of a high affinity phytochelatin synthase from the cd-utilizing marine diatom Thalassiosira pseudonana. J. Phycol. 2013, 49, 32–40. [Google Scholar] [CrossRef]
- Olsson, S.; Penacho, V.; Puente-Sánchez, F.; Díaz, S.; Gonzalez-Pastor, J.E.; Aguilera, A. Horizontal gene transfer of phytochelatin synthases from bacteria to extremophilic green algae. Microb. Ecol. 2017, 73, 50–60. [Google Scholar] [CrossRef] [Green Version]
- Osaki, Y.; Shirabe, T.; Nakanishi, H.; Wakagi, T.; Yoshimura, E. Characterization of phytochelatin synthase produced by the primitive red alga Cyanidioschyzon merolae. Metallomics 2009, 1, 353–358. [Google Scholar] [CrossRef]
- Perales-Vela, H.V.; Peña-Castro, J.M.; Cañizares-Villanueva, R.O. Heavy metal detoxification in eukaryotic microalgae. Chemosphere 2006, 64, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Bundy, J.G.; Kille, P. Metabolites and metals in Metazoa—What role do phytochelatins play in animals? Metallomics 2014, 6, 1576–1582. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- García-García, J.D.; Girard, L.; Hernández, G.; Saavedra, E.; Pardo, J.P.; Rodríguez-Zavala, J.S.; Encalada, R.; Reyes-Prieto, A.; Mendoza-Cózatl, D.G.; Moreno-Sánchez, R. Zn-bis-glutathionate is the best co-substrate of the monomeric phytochelatin synthase from the photosynthetic heavy metal-hyperaccumulator Euglena gracilis. Metallomics 2014, 6, 604–616. [Google Scholar] [CrossRef] [PubMed]
- Pawlik-Skowrońska, B. Correlations between toxic Pb effects and production of Pb-induced thiol peptides in the microalga Stichococcus bacillaris. Environ. Pollut. 2002, 119, 119–127. [Google Scholar] [CrossRef]
- Grill, E.; Loffler, S.; Winnacker, E.L.; Zenk, M.H. Phytochelatins, the heavy metal binding peptides of plants, are synthesized from glutathione by a specific gamma-glutamylcysteine depeptidyl transpeptidase (Phytochelatin synthase). Proc. Natl. Acad. Sci. USA 1989, 86, 6838–6842. [Google Scholar] [CrossRef] [Green Version]
- Vatamaniuk, O.K.; Mari, S.; Lang, A.; Chalasani, S.; Demkiv, L.O.; Rea, P.A. Phytochelatin synthase, a dipeptidyltransferase that undergoes multisite acylation with gamma-glutamylcysteine during catalysis: Stoichiometric and site-directed mutagenic analysis of Arabidopsis thaliana PCS1-catalyzed phytochelatin synthesis. J. Biol. Chem. 2004, 279, 22449–22460. [Google Scholar] [CrossRef] [Green Version]
- Romanyuk, N.D.; Rigden, D.J.; Vatamaniuk, O.K.; Lang, A.; Cahoon, R.E.; Jez, J.M.; Rea, P.A. Mutagenic definition of a papain-like catalytic triad, sufficiency of the N-terminal domain for single-site core catalytic enzyme acylation, and C-terminal domain for augmentative metal activation of a eukaryotic phytochelatin synthase. Plant Physiol. 2006, 141, 858–869. [Google Scholar] [CrossRef] [Green Version]
- Vivares, D.; Arnoux, P.; Pignol, D. A papain-like enzyme at work: Native and acyl-enzyme intermediate structures in phytochelatin synthesis. Proc. Natl. Acad. Sci. USA 2005, 102, 18848–18853. [Google Scholar] [CrossRef] [Green Version]
- Ha, S.B.; Smith, A.P.; Howden, R.; Dietrich, W.M.; Bugg, S.; O’Connell, M.J.; Goldsbrough, P.B.; Cobbett, C.S. Phytochelatin synthase genes from Arabidopsis and the yeast Schizosaccharomyces pombe. Plant Cell 1999, 11, 1153–1164. [Google Scholar] [CrossRef]
- Tsuji, N.; Nishikori, S.; Iwabe, O.; Matsumoto, S.; Shiraki, K.; Miyasaka, H.; Takagi, M.; Miyamoto, K.; Hirata, K. Comparative analysis of the two-step reaction catalyzed by prokaryotic and eukaryotic phytochelatin synthase by an ion-pair liquid chromatography assay. Planta 2005, 222, 181–191. [Google Scholar] [CrossRef]
- Bellini, E.; Varotto, C.; Borsò, M.; Rugnini, L.; Bruno, L.; Sanità di Toppi, L. Eukaryotic and prokaryotic phytochelatin synthases differ less in functional terms than previously thought: A comparative analysis of Marchantia polymorpha and Geitlerinema sp. PCC 7407. Plants 2020, 9, 914. [Google Scholar] [CrossRef] [PubMed]
- Foyer, C.H.; Noctor, G. Redox homeostasis and antioxidant signaling: A metabolic interface between stress perception and physiological responses. Plant Cell 2005, 17, 1866–1875. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mullineaux, P.M.; Rausch, T. Glutathione, photosynthesis and the redox regulation of stress-responsive gene expression. Photosynth. Res. 2005, 86, 459–474. [Google Scholar] [CrossRef] [PubMed]
- Freeman, J.L.; Persans, M.W.; Nieman, K.; Albrecht, C.; Peer, W.; Pickering, I.J.; Salt, D.E. Increased glutathione biosynthesis plays a role in nickel tolerance in Thlaspi nickel hyperaccumulators. Plant Cell 2004, 16, 2176–2191. [Google Scholar] [CrossRef] [Green Version]
- Dixon, D.P.; Cummins, L.; Cole, D.J.; Edwards, R. Glutathione-mediated detoxification systems in plants. Curr. Opin. Plant Biol. 1998, 1, 258–266. [Google Scholar] [CrossRef]
- Tsuji, N.; Nishikori, S.; Iwabe, O.; Shiraki, K.; Miyasaka, H.; Takagi, M.; Hirata, K.; Miyamoto, K. Characterization of phytochelatin synthase-like protein encoded by alr0975 from a prokaryote, Nostoc sp. PCC 7120. Biochem. Biophys. Res. Commun. 2004, 315, 751–755. [Google Scholar] [CrossRef]
- Beck, A.; Lendzian, K.; Oven, M.; Christmann, A.; Grill, E. Phytochelatin synthase catalyzes key step in turnover of glutathione conjugates. Phytochemistry 2003, 62, 423–431. [Google Scholar] [CrossRef]
- Harada, E.; Vonroepenacklahaye, E.; Clemens, S. A cyanobacterial protein with similarity to phytochelatin synthases catalyzes the conversion of glutathione to -glutamylcysteine and lacks phytochelatin synthase activity. Phytochemistry 2004, 65, 3179–3185. [Google Scholar] [CrossRef]
- Hirata, K.; Tsuji, N.; Miyamoto, K. Biosynthetic regulation of phytochelatins, heavy metal-binding peptides. J. Biosci. Bioeng. 2005, 100, 593–599. [Google Scholar] [CrossRef]
- Mehra, R.K.; Kodati, V.R.; Abdullah, R. Chain lengthdependent Pb (II)-coordination in phytochelatins. Biochem. Biophys. Res. Commun. 1995, 215, 730–736. [Google Scholar] [CrossRef]
- Dennis, K.K.; Uppal, K.; Liu, K.H.; Ma, C.; Liang, B.; Go, Y.M.; Jones, D.P. Phytochelatin database: A resource for phytochelatin complexes of nutritional and environmental metals. Database 2019, 2019, baz083. [Google Scholar] [CrossRef] [PubMed]
- Chekmeneva, E.; Prohens, R.; Díaz-Cruz, J.M.; Ariño, C.; Esteban, M. Competitive binding of Cd and Zn with the phytochelatin (gamma-Glu-Cys)4-Gly: Comparative study by mass spectrometry, voltammetry-multivariate curve resolution, and isothermal titration calorimetry. Environ. Sci. Technol. 2008, 42, 2860–2866. [Google Scholar] [CrossRef] [PubMed]
- May, M.J.; Leaver, C.J. oxidative stimulation of glutathione synthesis in Arabidopsis thaliana suspension cultures. Plant Physiol. 1993, 103, 621–627. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tsuji, N.; Hirayanagi, N.; Okada, M.; Miyasaka, H.; Hirata, K.; Zenk, M.H.; Miyamoto, K. Enhancement of tolerance to heavy metals and oxidative stress in Dunaliella tertiolecta by Zn-induced phytochelatin synthesis. Biochem. Biophys. Res. Commun. 2002, 293, 653–659. [Google Scholar] [CrossRef]
- Filiz, E.; Saracoglu, I.A.; Ozyigit, I.I.; Yalcin, B. Comparative analyses of phytochelatin synthase (PCS) genes in higher plants. Biotechnol. Biotechnol. Equip. 2019, 33, 178–194. [Google Scholar] [CrossRef] [Green Version]
- Mendoza-Cozatl, D.G.; Moreno-Sanchez, R. Cd2+ transport and storage in the chloroplast of Euglena gracilis. Biochim. Biophys. Acta Bioenerg. 2005, 1706, 88–97. [Google Scholar] [CrossRef] [Green Version]
- Noctor, G.; Mhamdi, A.; Chaouch, S.; Han, Y.; Neukermans, J.; Marquez-Garcia, B.; Queval, G.; Foyer, C.H. Glutathione in plants: An integrated overview. Plant Cell Environ. 2012, 35, 454–484. [Google Scholar] [CrossRef]
- Pal, R.; Rai, J.P.N. Phytochelatins: Peptides involved in heavy metal detoxification. Appl. Biochem. Biotechnol. 2010, 160, 945–963. [Google Scholar] [CrossRef]
- Ahner, B.A.; Morel, F.M.M. Phytochelatin production in marine algae. 2. Induction by various metals. Limnol. Oceanogr. 1995, 40, 658–665. [Google Scholar] [CrossRef] [Green Version]
- Kawakami, S.K.; Gledhill, M.; Achterberg, E.P. Production of phytochelatins and glutathione by marine phytoplankton in response to metal stress. J. Phycol. 2006, 42, 975–989. [Google Scholar] [CrossRef]
- Balzano, S.; Sardo, A.; Blasio, M.; Chahine, T.B.; Dell’Anno, F.; Sansone, C.; Brunet, C. microalgal metallothioneins and phytochelatins and their potential use in bioremediation. Front Microbiol. 2020, 11, 517. [Google Scholar] [CrossRef] [PubMed]
- Blaby-Haas, C.E.; Merchant, S.S. Comparative and functional algal genomics. Annu. Rev. Plant Biol. 2019, 70, 605–638. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Keeling, P.J.; Burki, F.; Wilcox, H.M.; Allam, B.; Allen, E.E.; Amaral-Zettler, L.A. The marine microbial eukaryote transcriptome sequencing project (MMETSP): Illuminating the functional diversity of eukaryotic life in the oceans through transcriptome sequencing. PLoS Biol. 2014, 12, e1001889. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Olsson, S.; Puente-Sánchez, F.; Gómez, M.J.; Aguilera, A. Transcriptional response to copper excess and identification of genes involved in heavy metal tolerance in the extremophilic microalga Chlamydomonas acidophila. Extremophiles 2015, 19, 657–672. [Google Scholar] [CrossRef] [PubMed]
- Hirooka, S.; Hirose, Y.; Kanesaki, Y.; Higuchi, S.; Fujiwara, T.; Onuma, R.; Era, A.; Ohbayashi, R.; Uzuka, A.; Nozaki, H.; et al. Acidophilic green algal genome provides insights into adaptation to an acidic environment. Proc. Natl. Acad. Sci. USA 2017, 114, E8304–E8313. [Google Scholar] [CrossRef] [Green Version]
- Aguilera, A.; Manrubia, S.C.; Gómez, F.; Rodríguez, N.; Amils, R. Eukaryotic community distribution and its relationship to water physicochemical parameters in an extreme acidic environment, Rio Tinto (Southwestern Spain). Appl. Environ. Microbiol. 2006, 72, 5325–5330. [Google Scholar] [CrossRef] [Green Version]
- Altschul, S.F.; Madden, T.L.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [Green Version]
- Katoh, K.; Misawa, K.; Kuma, K.; Miyata, T. MAFFT: A novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 2002, 30, 3059–3066. [Google Scholar] [CrossRef] [Green Version]
- Kalyaanamoorthy, S.; Minh, B.Q.; Wong, T.; von Haeseler, A.; Jermiin, L.S. ModelFinder: Fast model selection for accurate phylogenetic estimates. Nat. Methods 2017, 14, 587–589. [Google Scholar] [CrossRef] [Green Version]
- Nguyen, L.T.; Schmidt, H.A.; von Haeseler, A.; Minh, B.Q. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef]
- Hoang, D.T.; Chernomor, O.; von Haeseler, A.; Minh, B.Q.; Vinh, L.S. UFBoot2: Improving the Ultrafast Bootstrap Approximation. Mol. Biol. Evol. 2018, 35, 518–522. [Google Scholar] [CrossRef] [PubMed]
- Stöver, B.C.; Müller, K.F. TreeGraph 2: Combining and visualizing evidence from different phylogenetic analyses. BMC Bioinform. 2010, 11, 7. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rothschild, L.J.; Mancinelli, R.L. Life in extreme environments. Nature 2001, 409, 1092–1101. [Google Scholar] [CrossRef] [PubMed]
- Gisbert, C.; Ros, R.; De Haro, A.; Walker, D.J.; Pilar Bernal, M.; Serrano, R.; Navarro-Aviñó, J. A plant genetically modified that accumulates Pb is especially promising for phytoremediation. Biochem. Biophys. Res. Commun. 2003, 303, 440–445. [Google Scholar] [CrossRef]
- Bae, W.; Mehra, R.K.; Mulchandani, A.; Chen, W. Genetic engineering of Escherichia coli for enhanced uptake and bioaccumulation of mercury. Appl. Environ. Microbiol. 2001, 67, 5335–5338. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mehra, R.K.; Winge, D.R. Metal ion resistance in fungi: Molecular mechanisms and their regulated expression. J. Cell Biochem. 1991, 45, 30–40. [Google Scholar] [CrossRef]
- Vatamaniuk, O.K.; Mari, S.; Lu, Y.P.; Rea, P.A. AtPCS1, a phytochelatin synthase from Arabidopsis: Isolation and in vitro reconstitution. Proc. Natl. Acad. Sci. USA 1999, 96, 7110–7115. [Google Scholar] [CrossRef] [Green Version]
- Zenk, M.H. Heavy metal detoxification in higher plants—A review. Gene 1996, 179, 21–30. [Google Scholar] [CrossRef]
- Andrews, G.K. Regulation of metallothionein gene expression by oxidative stress and metal ions. Biochem. Pharmacol. 2000, 59, 95–104. [Google Scholar] [CrossRef]
- Sato, M.; Kondoh, M. Recent studies on metallothionein: Protection against toxicity of heavy metals and oxygen free radicals. Tohoku J. Exp. Med. 2002, 196, 9–22. [Google Scholar] [CrossRef] [Green Version]
- Brunetti, P.; Zanella, L.; Proia, A.; De Paolis, A.; Falasca, G.; Altamura, M.M.; Sanità di Toppi, L.; Costantino, P.; Cardarelli, M. Cadmium tolerance and phytochelatin content of Arabidopsis seedlings over-expressing the phytochelatin synthase gene AtPCS1. J. Exp. Bot. 2011, 62, 5509–5519. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lee, S.; Moon, J.S.; Ko, T.S.; Petros, D.; Goldsbrough, P.B.; Korban, S.S. Overexpression of Arabidopsis phytochelatin synthase paradoxically leads to hypersensitivity to cadmium stress. Plant Physiol. 2003, 131, 656–663. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, Y.; Dhankher, O.P.; Carreira, L.; Lee, D.; Chen, A.; Schroeder, J.I.; Balish, R.S.; Meagher, R.B. Overexpression of phytochelatin synthase in Arabidopsis leads to enhanced arsenic tolerance and cadmium hypersensitivity. Plant Cell Physiol. 2004, 45, 1787–1797. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, G.Y.; Zhang, Y.X.; Chai, T.Y. Phytochelatin synthase of Thlaspi caerulescens enhanced tolerance and accumulation of heavy metals when expressed in yeast and tobacco. Plant Cell Rep. 2011, 30, 1067–1076. [Google Scholar] [CrossRef] [PubMed]
- Pomponi, M.; Censi, V.; Di Girolamo, V.; De Paolis, A.; di Toppi, L.S.; Aromolo, R.; Costantino, P.; Cardarelli, M. Overexpression of Arabidopsis phytochelatin synthase in tobacco plants enhances Cd(2+) tolerance and accumulation but not translocation to the shoot. Planta 2006, 223, 180–190. [Google Scholar] [CrossRef]
- Wojas, S.; Clemens, S.; Hennig, J.; Sklodowska, A.; Kopera, E.; Schat, H.; Bal, W.; Antosiewicz, D.M. Overexpression of phytochelatin synthase in tobacco: Distinctive effects of AtPCS1 and CePCS genes on plant response to cadmium. J. Exp. Bot. 2008, 59, 2205–2219. [Google Scholar] [CrossRef] [Green Version]
- Clemens, S.; Kim, E.J.; Neumann, D.; Schroeder, J.I. Tolerance to toxic metals by a gene family of phytochelatin synthases from plants and yeast. EMBO J. 1999, 18, 3325–3333. [Google Scholar] [CrossRef] [Green Version]
- Clemens, S.; Schroeder, J.I.; Degenkolb, T. Caenorhabditis elegans expresses a functional phytochelatin synthase. Eur. J. Biochem. 2001, 268, 3640–3643. [Google Scholar] [CrossRef]
- Dong, R. Molecular cloning and characterization of a phytochelatin synthase gene, PvPCS1, from Pteris vittata L. J. Ind. Microbiol. Biotechnol. 2005, 32, 527–533. [Google Scholar] [CrossRef]
- Gasic, K.; Korban, S.S. Expression of Arabidopsis phytochelatin synthase in Indian mustard (Brassica juncea) plants enhances tolerance for Cd and Zn. Planta 2007, 225, 1277–1285. [Google Scholar] [CrossRef]
- Heiss, S.; Wachter, A.; Bogs, J.; Cobbett, C.; Rausch, T. Phytochelatin synthase (PCS) protein is induced in Brassica juncea leaves after prolonged Cd exposure. J. Exp. Bot. 2003, 54, 1833–1839. [Google Scholar] [CrossRef] [PubMed]
- Ramos, J.; Naya, L.; Gay, M.; Abián, J.; Becana, M. Functional characterization of an unusual phytochelatin synthase, LjPCS3, of Lotus japonicus. Plant Physiol. 2008, 148, 536–545. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shri, M.; Dave, R.; Diwedi, S.; Shukla, D.; Kesari, R.; Tripathi, R.D.; Trivedi, P.K.; Chakrabarty, D. Heterologous expression of Ceratophyllum demersum phytochelatin synthase, CdPCS1, in rice leads to lower arsenic accumulation in grain. Sci. Rep. 2014, 4, 5784. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Vatamaniuk, O.K.; Bucher, E.A.; Ward, J.T.; Rea, P.A. A new pathway for heavy metal detoxification in animals. Phytochelatin synthase is required for cadmium tolerance in Caenorhabditis elegans. J. Biol. Chem. 2001, 276, 20817–20820. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, M.; Stragliati, L.; Bellini, E.; Ricci, A.; Saba, A.; Sanità di Toppi, L.; Varotto, C. Evolution and functional differentiation of recently diverged phytochelatin synthase genes from Arundo donax L. J. Exp. Bot. 2019, 70, 5391–5405. [Google Scholar] [CrossRef] [Green Version]
- Shukla, D.; Kesari, R.; Mishra, S.; Dwivedi, S.; Tripathi, R.D.; Nath, P.; Trivedi, P.K. Expression of phytochelatin synthase from aquatic macrophyte Ceratophyllum demersum L. enhances cadmium and arsenic accumulation in tobacco. Plant Cell Rep. 2012, 31, 1687–1699. [Google Scholar] [CrossRef]
- Zhang, X.; Rui, H.; Zhang, F.; Hu, Z.; Xia, Y.; Shen, Z. Overexpression of a functional Vicia sativa PCS1 homolog increases cadmium tolerance and phytochelatins synthesis in Arabidopsis. Front. Plant Sci. 2018, 9, 107. [Google Scholar] [CrossRef] [Green Version]
- Chaurasia, N.; Mishra, Y.; Rai, L.C. Cloning expression and analysis of phytochelatin synthase (pcs) gene from Anabaena sp. PCC 7120 offering multiple stress tolerance in Escherichia coli. Biochem. Biophys. Res. Commun. 2008, 376, 225–230. [Google Scholar] [CrossRef]
- Kang, S.H.; Singh, S.; Kim, J.Y.; Lee, W.; Mulchandani, A.; Chen, W. Bacteria metabolically engineered for enhanced phytochelatin production and cadmium accumulation. Appl. Environ. Microbiol. 2007, 73, 6317–6320. [Google Scholar] [CrossRef] [Green Version]
- Li, Y.; Dankher, O.P.; Carreira, L.; Smith, A.P.; Meagher, R.B. The shoot-specific expression of gamma-glutamylcysteine synthetase directs the long-distance transport of thiol-peptides to roots conferring tolerance to mercury and arsenic. Plant Physiol. 2006, 141, 288–298. [Google Scholar] [CrossRef] [Green Version]
- Sauge-Merle, S.; Cuiné, S.; Carrier, P.; Lecomte-Pradines, C.; Luu, D.T.; Peltier, G. Enhanced toxic metal accumulation in engineered bacterial cells expressing Arabidopsis thaliana phytochelatin synthase. Appl. Environ. Microbiol. 2003, 69, 490–494. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Singh, S.; Lee, W.; Dasilva, N.A.; Mulchandani, A.; Chen, W. Enhanced arsenic accumulation by engineered yeast cells expressing Arabidopsis thaliana phytochelatin synthase. Biotechnol. Bioeng. 2008, 99, 333–340. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.; Kang, S.H.; Lee, W.; Mulchandani, A.; Chen, W. Systematic engineering of phytochelatin synthesis and arsenic transport for enhanced arsenic accumulation in E. coli. Biotechnol. Bioeng. 2010, 105, 780–785. [Google Scholar] [CrossRef] [PubMed]
- Tsai, S.L.; Singh, S.; Dasilva, N.A.; Chen, W. Co-expression of Arabidopsis thaliana phytochelatin synthase and Treponema denticola cysteine desulfhydrase for enhanced arsenic accumulation. Biotechnol. Bioeng. 2012, 109, 605–608. [Google Scholar] [CrossRef]
- Wang, F.; Wang, Z.; Zhu, C. Heteroexpression of the wheat phytochelatin synthase gene (TaPCS1) in rice enhances cadmium sensitivity. Acta Biochim. Biophys. Sin. 2012, 44, 886–893. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wawrzyńska, A.; Wawrzyński, A.; Gaganidze, D.; Kopera, E.; Piatek, K.; Bal, W.; Sirko, A. Overexpression of genes involved in phytochelatin biosynthesis in Escherichia coli: Effects on growth, cadmium accumulation and thiol level. Acta Biochim. Pol. 2005, 52, 109–116. [Google Scholar] [CrossRef] [Green Version]
- Wojas, S.; Clemens, S.; Skłodowska, A.; Antosiewicz, D.M. Arsenic response of AtPCS1- and CePCS-expressing plants—Effects of external As(V) concentration on As-accumulation pattern and NPT metabolism. J. Plant Physiol. 2010, 167, 169–175. [Google Scholar] [CrossRef]
- Wojas, S.; Ruszczyńska, A.; Bulska, E.; Clemens, S.; Antosiewicz, D.M. The role of subcellular distribution of cadmium and phytochelatins in the generation of distinct phenotypes of AtPCS1- and CePCS3-expressing tobacco. J. Plant Physiol. 2010, 167, 981–988. [Google Scholar] [CrossRef]
- Lee, B.D.; Hwang, S. Tobacco phytochelatin synthase (NtPCS1) plays important roles in cadmium and arsenic tolerance and in early plant development in tobacco. Plant Biotechnol. Rep. 2015, 9, 107–114. [Google Scholar] [CrossRef]
- Kühnlenz, T.; Schmidt, H.; Uraguchi, S.; Clemens, S. Arabidopsis thaliana phytochelatin synthase 2 is constitutively active in vivo and can rescue the growth defect of the PCS1-deficient cad1-3 mutant on Cd-contaminated soil. J. Exp. Bot. 2014, 65, 4241–4253. [Google Scholar] [CrossRef] [Green Version]
- Cazalé, A.C.; Clemens, S. Arabidopsis thaliana expresses a second functional phytochelatin synthase. FEBS Lett. 2001, 507, 215–219. [Google Scholar] [CrossRef] [Green Version]
- Uraguchi, S.; Tanaka, N.; Hofmann, C.; Abiko, K.; Ohkama-Ohtsu, N.; Weber, M.; Kamiya, T.; Sone, Y.; Nakamura, R.; Takanezawa, Y.; et al. Phytochelatin synthase has contrasting effects on cadmium and arsenic accumulation in rice grains. Plant Cell Physiol. 2017, 58, 1730–1742. [Google Scholar] [CrossRef] [PubMed]
- Meyer, C.L.; Peisker, D.; Courbot, M.; Craciun, A.R.; Cazalé, A.C.; Desgain, D.; Schat, H.; Clemens, S.; Verbruggen, N. Isolation and characterization of Arabidopsis halleri and Thlaspi caerulescens phytochelatin synthases. Planta 2011, 234, 83–95. [Google Scholar] [CrossRef] [PubMed]
- Mallik, N.; Rai, L.C. Characterization of Cd-induced low molecular weight protein in N2 fixing cyanobacterium Anabaena doliolum with special reference to co-/multiple stress tolerance. Biometals 1998, 11, 55–61. [Google Scholar] [CrossRef]
- Seki, M.; Narusaka, M.; Ishida, J.; Nanjo, T.; Fujita, M.; Oono, Y.; Kamiya, A.; Nakajima, M.; Enju, A.; Sakurai, T.; et al. Monitoring the expression profiles of 7000 Arabidopsis genes under drought, cold and high-salinity stresses using a full-length cDNA microarray. Plant J. 2002, 31, 279–292. [Google Scholar] [CrossRef]
- Stroiński, A.; Chadzinikolau, T.; Gizewska, K.; Zielezińska, M. ABA or cadmium induced phytochelatin synthesis in potato tubers. Biol. Plant. 2010, 54, 117–120. [Google Scholar] [CrossRef]
- Zhang, H.; Xu, W.; Guo, J.; He, Z.; Ma, M. Coordinated responses of phytochelatins and metallothioneins to heavy metals in garlic seedlings. Plant Sci. 2005, 169, 1059–1065. [Google Scholar] [CrossRef]
- Bhargava, P.; Srivastava, A.K.; Urmil, S.; Rai, L.C. Phytochelatin plays a role in UV-B tolerance in N2-fixing cyanobacterium Anabaena doliolum. J. Plant Physiol. 2005, 162, 1220–1225. [Google Scholar] [CrossRef]
- Chaurasia, N.; Mishra, Y.; Chatterjee, A.; Rai, R.; Yadav, S.; Rai, L.C. Overexpression of phytochelatin synthase (pcs) enhances abiotic stress tolerance by altering the proteome of transformed Anabaena sp. PCC 7120. Protoplasma 2017, 254, 1715–1724. [Google Scholar] [CrossRef]
- Hall, J.L. Cellular mechanism for heavy metal detoxification and tolerance. J. Exp. Bot. 2001, 53, 1–11. [Google Scholar] [CrossRef]
- Gill, S.S.; Tuteja, N. Reactive oxygen species and antioxidant machinery in abiotic stress tolerance in crop plants. Plant Physiol. Biochem. 2010, 48, 909–930. [Google Scholar] [CrossRef] [PubMed]
- Rabenstein, D.L. Metal complexes of glutathione and their biologicalsignificance. In Glutathione: Chemical, Biochemical and Medical Aspects; Dolphin, D., Poulson, R., Avramovic, O., Eds.; John Wiley and Sons: New York, NY, USA, 1989; pp. 147–186. [Google Scholar]
- Catling, D.C.; Claire, M.W.; Zahnle, K.J.; Quinn, R.C.; Clark, B.C.; Hecht, M.H.; Kounaves, S. Atmospheric origins of perchlorate on Mars and in the Atacama. J. Geophys. Res. Planets 2010, 115, E00E11. [Google Scholar] [CrossRef] [Green Version]
- Díaz-Rullo, J.; Rodríguez-Valdecantos, G.; Torres-Rojas, F.; Cid, L.; Vargas, I.T.; González, B.; González-Pastor, J.E. mining for perchlorate resistance genes in microorganisms from sediments of a hypersaline pond in Atacama desert, Chile. Front. Microbiol. 2021, 12, 723874. [Google Scholar] [CrossRef] [PubMed]
- Kumarathilaka, P.; Oze, C.; Indraratne, S.P.; Vithanage, M. Perchlorate as an emerging contaminant in soil, water and food. Chemosphere 2016, 150, 667–677. [Google Scholar] [CrossRef]
- Backus, S.M.; Klawuun, P.; Brown, S.; D’sa, I.; Sharp, S.; Surette, C.; Williams, D.J. Determination of perchlorate in selected surface waters in the Great Lakes Basin by HPLC/MS/MS. Chemosphere 2005, 61, 834–843. [Google Scholar] [CrossRef]
- Motzer, W.E. Perchlorate: Problems, detection, and solutions. Environ. Forensics 2001, 2, 301–311. [Google Scholar] [CrossRef]
- Lamprecht-Grandío, M.; Cortesão, M.; Mirete, S.; de la Cámara, M.B.; de Figueras, C.G.; Pérez-Pantoja, D.; White, J.J.; Farías, M.E.; Rosselló-Móra, R.; González-Pastor, J.E. Novel genes involved in resistance to both ultraviolet radiation and perchlorate from the metagenomes of hypersaline environments. Front. Microbiol. 2020, 11, 453. [Google Scholar] [CrossRef]
- Kim, Y.O.; Kang, H.; Ahn, S.J. Overexpression of phytochelatin synthase AtPCS2 enhances salt tolerance in Arabidopsis thaliana. J. Plant Physiol. 2019, 240, 153011. [Google Scholar] [CrossRef]
- Huang, B. MetaPocket: A meta approach to improve protein ligand binding site prediction. OMICS J. Integr. Biol. 2009, 13, 325–330. [Google Scholar] [CrossRef]
- Jin, Y.; Jing, W.; Zhang, Q.; Zhang, W. Cyclic nucleotide gated channel 10 negatively regulates salt tolerance by mediating Na+ transport in Arabidopsis. J. Plant Res. 2015, 128, 211–220. [Google Scholar] [CrossRef]
- Zhang, M.; Zhang, G.Q.; Kang, H.H.; Zhou, S.M.; Wang, W. TaPUB1, a putative E3 ligase gene from wheat, enhances salt stress tolerance in transgenic Nicotiana benthamiana. Plant Cell Physiol. 2017, 58, 1673–1688. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- del Río, L.A.; Palma, J.M.; Sandalio, L.M.; Corpas, F.J.; Pastori, G.M.; Bueno, P.; López-Huertas, E. Peroxisomes as a source of superoxide and hydrogen peroxide in stressed plants. Biochem. Soc. Trans. 1996, 24, 434–438. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Quartacci, M.F.; Cosi, E.; Navari-Izzo, F. Lipids and NADPH-dependent superoxide production in plasma membrane vesicles from roots of wheat grown under copper deficiency or excess. J. Exp. Bot. 2001, 52, 77–84. [Google Scholar] [PubMed]
- Li, M.; Barbaro, E.; Bellini, E.; Saba, A.; Sanità di Toppi, L.; Varotto, C. Ancestral function of the phytochelatin synthase C-terminal domain in inhibition of heavy metal-mediated enzyme overactivation. J. Exp. Bot. 2020, 71, 6655–6669. [Google Scholar] [CrossRef] [PubMed]
- Shukla, D.; Kesari, R.; Tiwari, M.; Dwivedi, S.; Tripathi, R.D.; Nath, P.; Trivedi, P.K. Expression of Ceratophyllum demersum phytochelatin synthase, CdPCS1, in Escherichia coli and Arabidopsis enhances heavy metal(loid)s accumulation. Protoplasma 2013, 250, 1263–1272. [Google Scholar] [CrossRef] [PubMed]
- Díaz, S.; Francisco, P.; Olsson, S.; Aguilera, Á.; González-Toril, E.; Martín-González, A. Toxicity, physiological, and ultrastructural effects of arsenic and cadmium on the extremophilic microalga Chlamydomonas acidophila. Int. J. Environ. Res. Public Health 2020, 17, 1650. [Google Scholar] [CrossRef] [Green Version]
- Adams, M.S.; Dillon, C.T.; Vogt, S.; Lai, B.; Stauber, J.; Jolley, D.F. Copper uptake, intracellular localization, and speciation in marine microalgae measured by synchrotron radiation X-ray fluorescence and absorption microspectroscopy. Environ. Sci. Technol. 2016, 50, 8827–8839. [Google Scholar] [CrossRef]
- Wang, Y.; Zhang, C.; Zheng, Y.; Ge, Y. Bioaccumulation kinetics of arsenite and arsenate in Dunaliella salina under different phosphate regimes. Environ. Sci. Pollut. Res. Int. 2017, 24, 21213–21221. [Google Scholar] [CrossRef]
- Wang, Y.; Zhang, C.; Yu, X.; Ge, Y. Arsenite Oxidation by Dunaliella salina is affected by external phosphate concentration. Bull Environ. Contam. Toxicol. 2020, 105, 868–873. [Google Scholar] [CrossRef]
- Hayashi, S.; Tanikawa, H.; Kuramata, M.; Abe, T.; Ishikawa, S. Domain exchange between Oryza sativa phytochelatin synthases reveals a region that determines responsiveness to arsenic and heavy metals. Biochem. Biophys. Res. Commun. 2020, 523, 548–553. [Google Scholar] [CrossRef]
- Ruotolo, R.; Peracchi, A.; Bolchi, A.; Infusini, G.; Amoresano, A.; Ottonello, S. Domain organization of phytochelatin synthase: Functional properties of truncated enzyme species identified by limited proteolysis. J. Biol. Chem. 2004, 279, 14686–14693. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Loscos, J.; Naya, L.; Ramos, J.; Clemente, M.R.; Matamoros, M.A.; Becana, M. A reassessment of substrate specificity and activation of phytochelatin synthases from model plants by physiologically relevant metals. Plant Physiol. 2006, 140, 1213–1221. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kumar, K.; Dahms, H.U.; Won, E.J.; Lee, J.S.; Shin, K.H. Microalgae—A promising tool for heavy metal remediation. Ecotoxicol. Environ. Saf. 2015, 113, 329–352. [Google Scholar] [CrossRef] [PubMed]
- Bräutigam, A.; Schaumlöffel, D.; Krauss, G.J.; Wesenberg, D. Analytical approach for characterization of cadmium-induced thiol peptides--a case study using Chlamydomonas reinhardtii. Anal. Bioanal. Chem. 2009, 395, 1737–1747. [Google Scholar] [CrossRef]
- Bräutigam, A.; Bomke, S.; Pfeifer, T.; Karst, U.; Krauss, G.J.; Wesenberg, D. Quantification of phytochelatins in Chlamydomonas reinhardtii using ferrocene-based derivatization. Metallomics 2010, 2, 565–570. [Google Scholar] [CrossRef]
- Bräutigam, A.; Wesenberg, D.; Preud’homme, H.; Schaumlöffel, D. Rapid and simple UPLC-MS/MS method for precise phytochelatin quantification in alga extracts. Anal. Bioanal. Chem. 2010, 398, 877–883. [Google Scholar] [CrossRef]
- Bräutigam, A.; Schaumlöffel, D.; Preud’homme, H.; Thondorf, I.; Wesenberg, D. Physiological characterization of cadmium-exposed Chlamydomonas reinhardtii. Plant Cell Environ. 2011, 34, 2071–2082. [Google Scholar] [CrossRef]
- Hirata, K.; Tsujimoto, Y.; Namba, T.; Ohta, T.; Hirayanagi, N.; Miyasaka, H.; Zenk, M.H.; Miyamoto, K. Strong induction of phytochelatin synthesis by zinc in marine green alga, Dunaliella tertiolecta. J. Biosci. Bioeng. 2001, 92, 24–29. [Google Scholar] [CrossRef]
- Jiang, Y.; Purchase, D.; Jones, H.; Garelick, H. Effects of arsenate (AS5+) on growth and production of glutathione (GSH) and phytochelatins (PCS) in Chlorella vulgaris. Int. J. Phytoremediation 2011, 13, 834–844. [Google Scholar] [CrossRef] [Green Version]
- Kobayashi, I.; Fujiwara, S.; Saegusa, H.; Inouhe, M.; Matsumoto, H.; Tsuzuki, M. Relief of arsenate toxicity by Cd-stimulated phytochelatin synthesis in the green alga Chlamydomonas reinhardtii. Mar. Biotechnol. 2006, 8, 94–101. [Google Scholar] [CrossRef]
- Le Faucheur, S.; Behra, R.; Sigg, L. Phytochelatin induction, cadmium accumulation, and algal sensitivity to free cadmium ion in Scenedesmus vacuolatus. Environ. Toxicol. Chem. 2005, 24, 1731–1737. [Google Scholar] [CrossRef] [PubMed]
- Pawlik-Skowrónska, B.; Pirszel, J.; Kalinowska, R.; Skowrónski, T. Arsenic availability, toxicity and direct role of GSH and phytochelatins in As detoxification in the green alga Stichococcus bacillaris. Aquat. Toxicol. 2004, 70, 201–212. [Google Scholar] [CrossRef] [PubMed]
- Simmons, D.B.; Hayward, A.R.; Hutchinson, T.C.; Emery, R.J. Identification and quantification of glutathione and phytochelatins from Chlorella vulgaris by RP-HPLC ESI-MS/MS and oxygen-free extraction. Anal. Bioanal. Chem. 2009, 395, 809–817. [Google Scholar] [CrossRef] [PubMed]
- Volland, S.; Schaumlöffel, D.; Dobritzsch, D.; Krauss, G.J.; Lütz-Meindl, U. Identification of phytochelatins in the cadmium-stressed conjugating green alga Micrasterias denticulata. Chemosphere 2013, 91, 448–454. [Google Scholar] [CrossRef]
- Wang, Y.; Zhang, C.; Zheng, Y.; Ge, Y. Phytochelatin synthesis in Dunaliella salina induced by arsenite and arsenate under various phosphate regimes. Ecotoxicol. Environ. Saf. 2017, 139, 150–160. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.J.; Wang, W.X. Cadmium sensitivity, uptake, subcellular distribution and thiol induction in a marine diatom: Exposure to cadmium. Aquat. Toxicol. 2011, 101, 377–386. [Google Scholar] [CrossRef]
- Wu, Y.; Wang, W.X. Thiol compounds induction kinetics in marine phytoplankton during and after mercury exposure. J. Hazard. Mater. 2012, 217–218, 271–278. [Google Scholar] [CrossRef]
- Wu, Y.; Wang, W.X. Intracellular speciation and transformation of inorganic mercury in marine phytoplankton. Aquat. Toxicol. 2014, 148, 122–129. [Google Scholar] [CrossRef]
| Metal/ | Concentration (mM) | Bioaccumulation | Bioaccumulation | t-Test | ||
|---|---|---|---|---|---|---|
| Metalloid | (mgg−1d.w) | (mgg−1d.w) | For Equality ofMeans | |||
| NC | SD | CaPCS2 | SD | Sig(2-tailed) | ||
| 1 | 6299 | 1814 | 6624 | 2734 | 0.902 | |
| Cu | 2 | 1006 | 116 | 1111 | 59 | 0.232 |
| 4 | 3822 | 1192 | 4204 | 1536 | 0.504 | |
| Cd | 0.3 | 4653 | 893 | 4640 | 1294 | 0.982 |
| 1 | 13,377 | 3461 | 14,547 | 2773 | 0.533 | |
| As | 1 | 246 | 63 | 272 | 70 | 0.513 |
| 0.5 | 381 | 25 | 338 | 148 | 0.642 | |
| Ni | 1 | 1375 | 102 | 1017 | 200 | 0.051 |
| 4 | 8486 | 1099 | 6234 | 1365 | 0.011 * | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Díaz, S.; Aguilera, Á.; de Figueras, C.G.; de Francisco, P.; Olsson, S.; Puente-Sánchez, F.; González-Pastor, J.E. Heterologous Expression of the Phytochelatin Synthase CaPCS2 from Chlamydomonas acidophila and Its Effect on Different Stress Factors in Escherichia coli. Int. J. Environ. Res. Public Health 2022, 19, 7692. https://doi.org/10.3390/ijerph19137692
Díaz S, Aguilera Á, de Figueras CG, de Francisco P, Olsson S, Puente-Sánchez F, González-Pastor JE. Heterologous Expression of the Phytochelatin Synthase CaPCS2 from Chlamydomonas acidophila and Its Effect on Different Stress Factors in Escherichia coli. International Journal of Environmental Research and Public Health. 2022; 19(13):7692. https://doi.org/10.3390/ijerph19137692
Chicago/Turabian StyleDíaz, Silvia, Ángeles Aguilera, Carolina G. de Figueras, Patricia de Francisco, Sanna Olsson, Fernando Puente-Sánchez, and José Eduardo González-Pastor. 2022. "Heterologous Expression of the Phytochelatin Synthase CaPCS2 from Chlamydomonas acidophila and Its Effect on Different Stress Factors in Escherichia coli" International Journal of Environmental Research and Public Health 19, no. 13: 7692. https://doi.org/10.3390/ijerph19137692
APA StyleDíaz, S., Aguilera, Á., de Figueras, C. G., de Francisco, P., Olsson, S., Puente-Sánchez, F., & González-Pastor, J. E. (2022). Heterologous Expression of the Phytochelatin Synthase CaPCS2 from Chlamydomonas acidophila and Its Effect on Different Stress Factors in Escherichia coli. International Journal of Environmental Research and Public Health, 19(13), 7692. https://doi.org/10.3390/ijerph19137692






