Detection of Free-Living Amoebae and Their Intracellular Bacteria in Borehole Water before and after a Ceramic Pot Filter Point-of-Use Intervention in Rural Communities in South Africa
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Consent
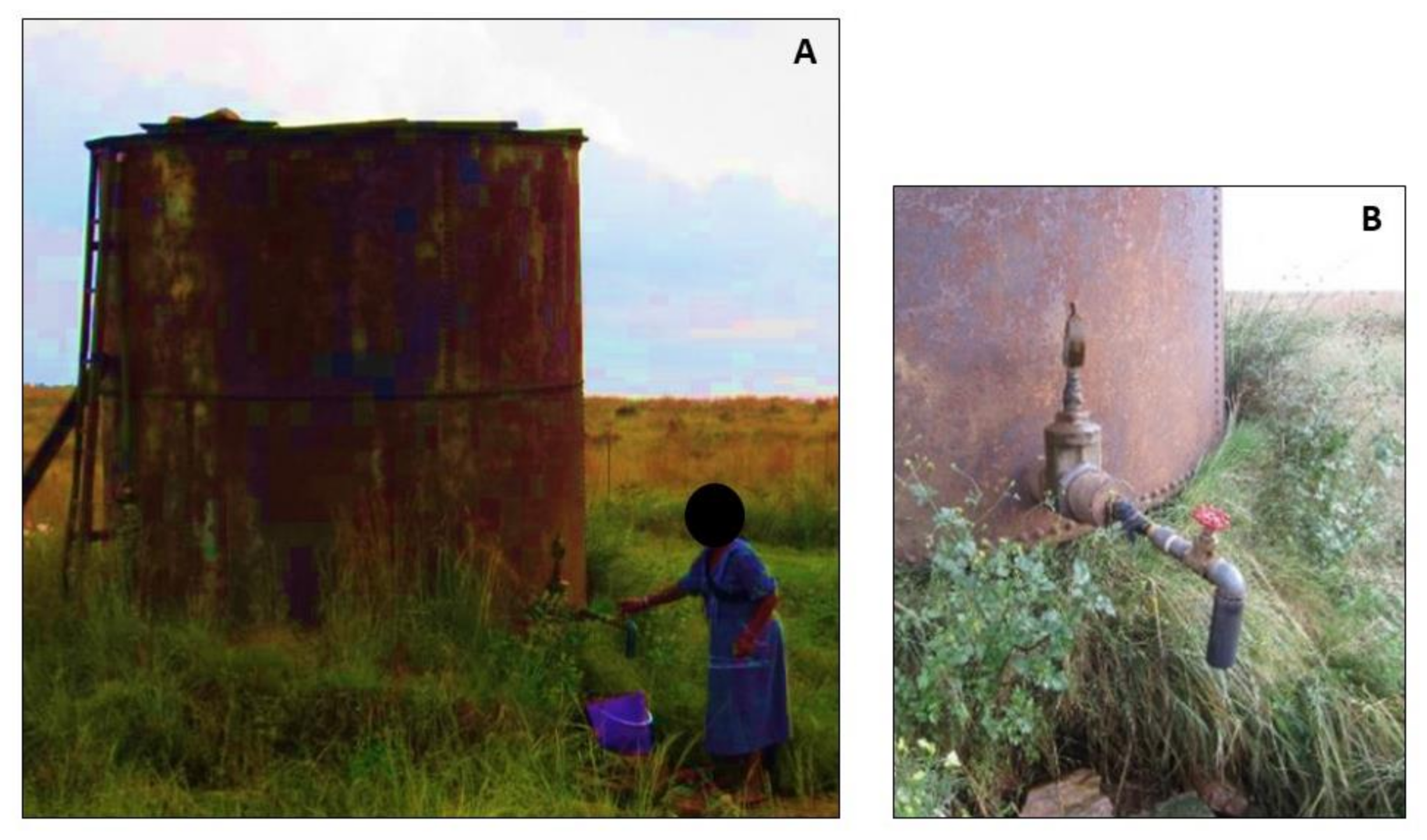
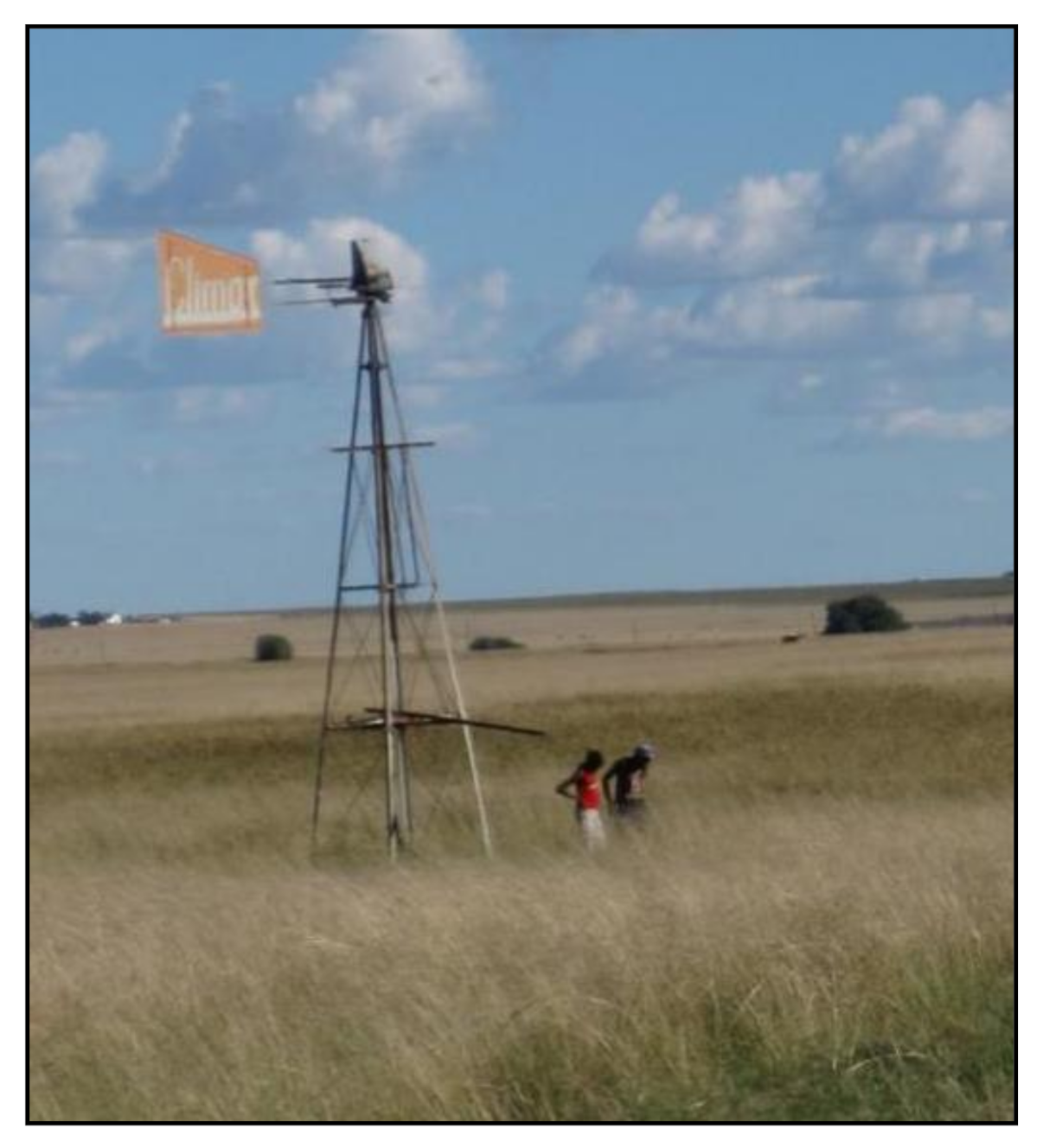
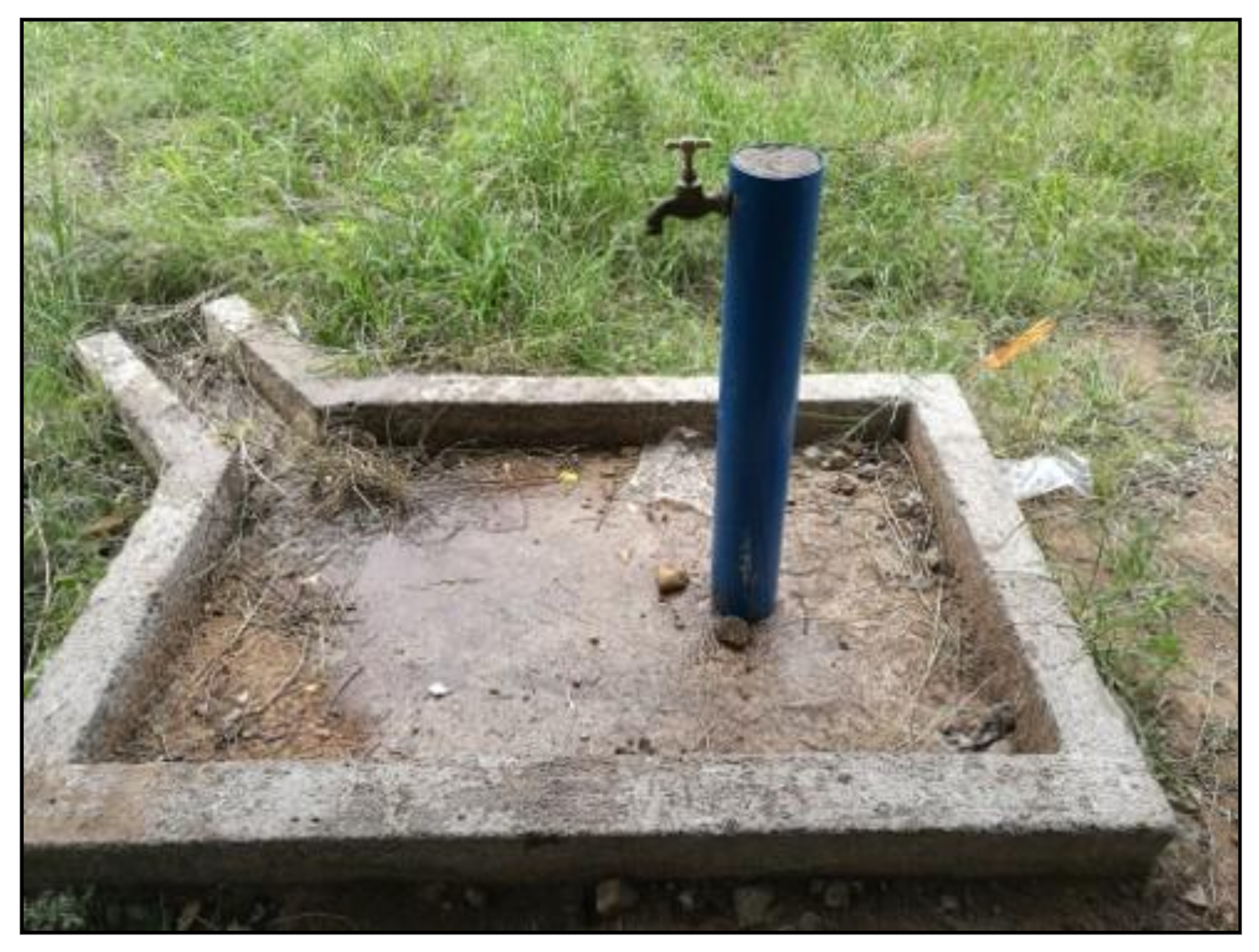
2.2. Study Site Description
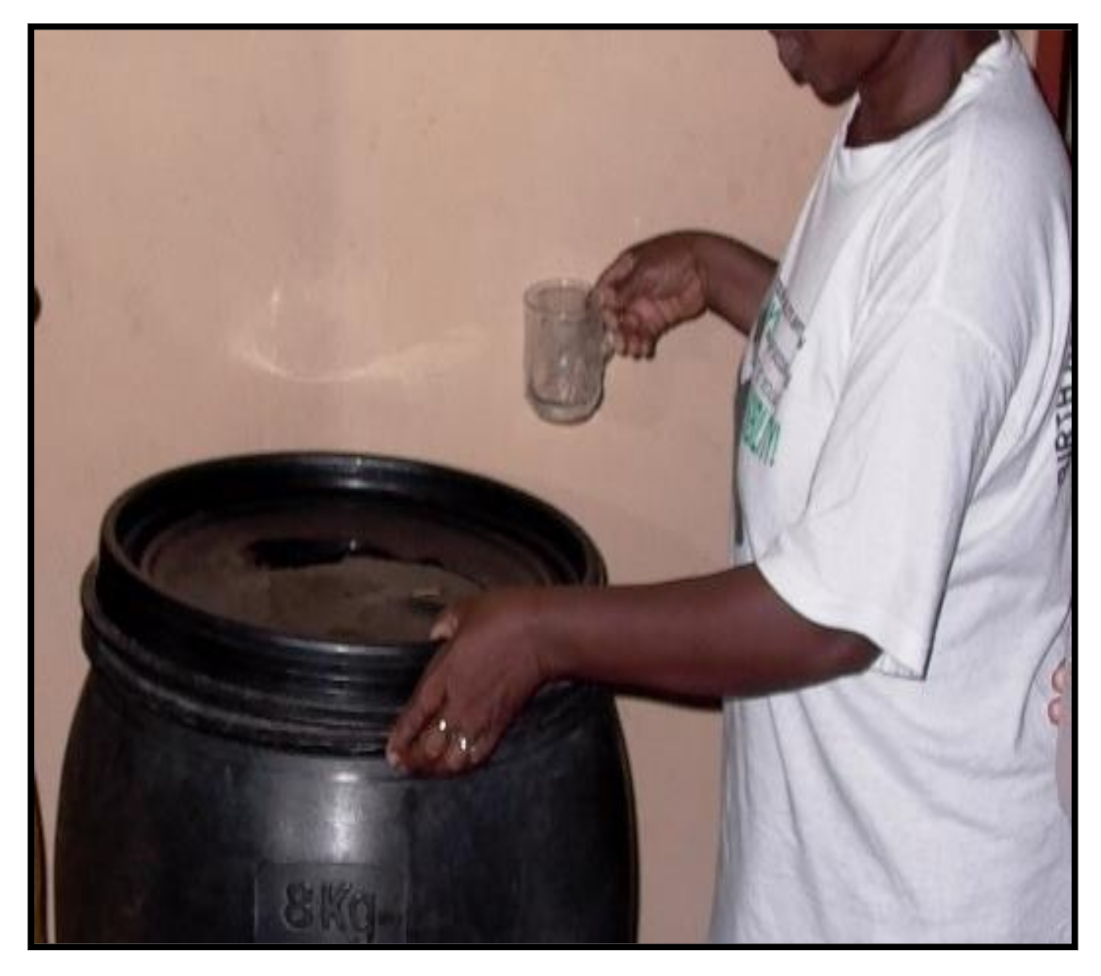
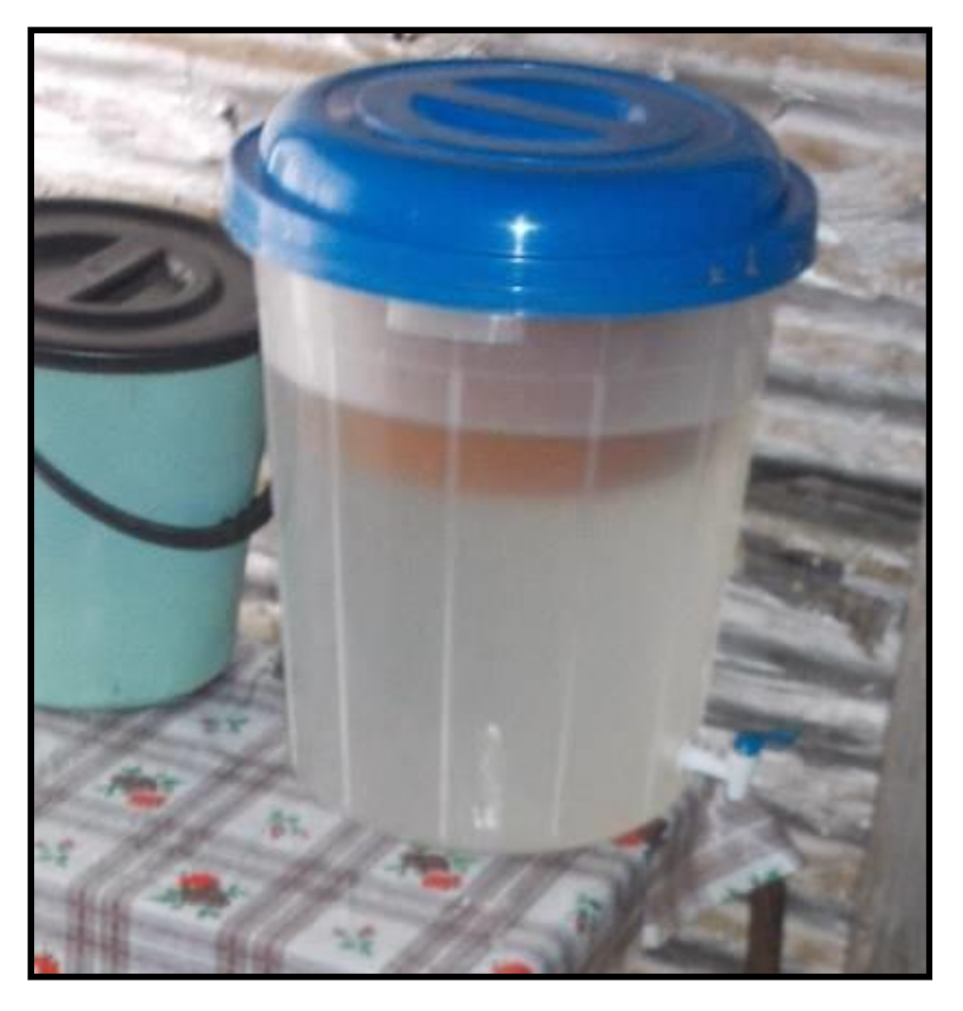
2.3. Point-of-Use Intervention
2.4. Water Quality Assessment Using E coli Bacteria Counts and Physical Parameters
2.5. Amoebal Enrichment of Water Samples
2.6. Isolation and Identification of FLA
2.7. Isolation and Identification of ARB
3. Results
3.1. Collection of Water Samples
3.2. Water Quality Assessment Using Indicators
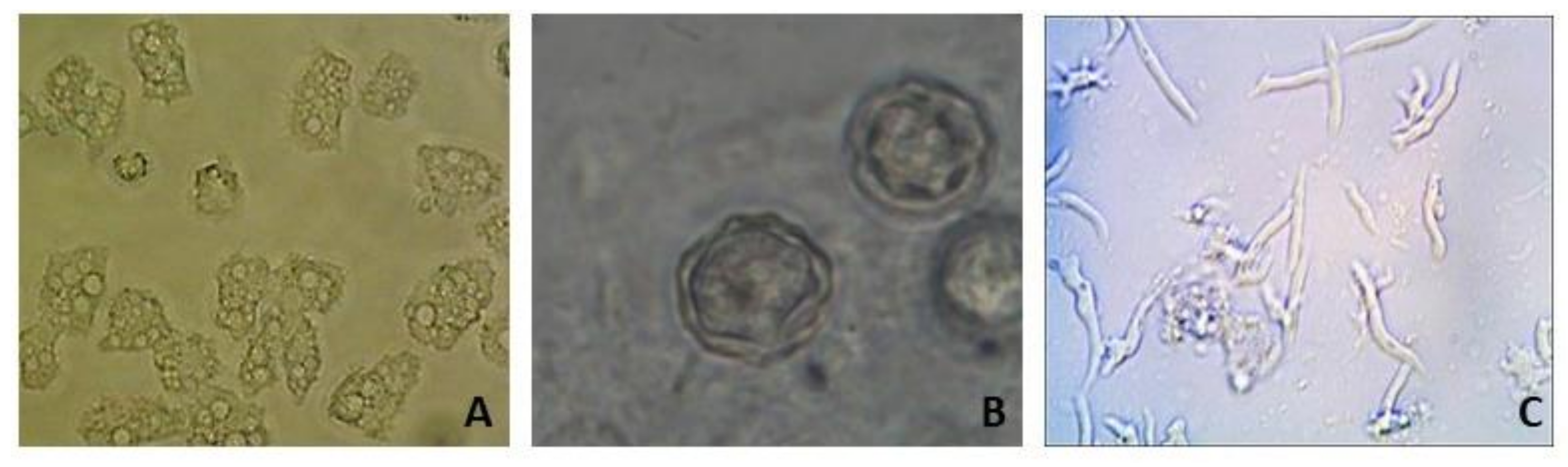
3.3. Isolation and Identification of FLA
3.4. Isolation and Identification of ARB
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Rodríguez-Zaragoza, S. Ecology of free-living amoebae. Crit. Rev. Microbiol. 1994, 20, 225–241. [Google Scholar] [CrossRef]
- Winiecka-Krusnell, J.; Linder, E. Bacterial infections of free-living amoebae. Res. Microbiol. 2001, 152, 613–619. [Google Scholar] [CrossRef]
- Rodriguez-Zaragoza, S.; Mayzlish, E.; Steinberger, Y. Vertical distribution of the free-living amoeba population in soil under desert shrubs in the Negev desert, Israel. Appl. Environ. Microbiol. 2005, 71, 2053–2060. [Google Scholar] [CrossRef] [PubMed]
- Schmitz-Esser, S.; Toenshoff, E.R.; Haider, S.; Heinz, E.; Hoenninger, V.M.; Wagner, M.; Horn, M. Diversity of Bacterial Endosymbionts of Environmental Acanthamoeba Isolates. Appl. Environ. Microbiol. 2008, 74, 5822–5831. [Google Scholar] [CrossRef]
- Hoffmann, R.; Michel, R. Distribution of free-living amoebae (FLA) during preparation and supply of drinking water. Int. J. Hyg. Environ. Health 2001, 203, 215–219. [Google Scholar] [CrossRef]
- Thomas, V.; Herrera-Rimann, K.; Blanc, D.S.; Greub, G. Biodiversity of amoebae and amoeba-resisting bacteria in a hospital water network. Appl. Environ. Microbiol. 2006, 72, 2428–2438. [Google Scholar] [CrossRef]
- Thomas, V.; McDonnel, G.; Denyer, S.P.; Maillard, J.-Y. Free-living amoebae and their intracellular pathogenic microorganisms: Risk for water quality. FEMS Microbiol. Rev. 2010, 34, 231–259. [Google Scholar] [CrossRef] [PubMed]
- Corsaro, D.; Feroldi, V.; Saucedo, G.; Ribas, F.; Loret, J.F.; Greub, G. Novel Chlamydiales strains isolated from a water treatment plant. Environ. Microbiol. 2009, 11, 188–200. [Google Scholar] [CrossRef]
- Muchesa, P.; Barnard, T.G.; Bartie, C. The prevalence of free-living amoebae in a South African hospital water distribution system. S. Afr. J. Sci. 2015, 111, 1–3. [Google Scholar] [CrossRef]
- Martinez, A.J.; Visvesvara, G.S. Free-living, amphizoic and opportunistic amebas. Brain Pathol. 1997, 7, 583–598. [Google Scholar] [CrossRef] [PubMed]
- Visvesvara, G.S.; Moura, H.; Schuster, F.L. Pathogenic and opportunistic free-living amoebae: Acanthamoeba spp., Balamuthia mandrillaris, Naegleria fowleri, and Sappinia diploidea. FEMS Immunol. Med. Microbiol. 2007, 50, 1–26. [Google Scholar] [CrossRef]
- Qvarnstrom, Y.; da Silva, A.J.; Schuster, F.L.; Gelman, B.B.; Visvesvara, G.S. Molecular confirmation of Sappinia pedata as a causative agent of amoebic encephalitis. J. Infect. Dis. 2009, 199, 1139–1142. [Google Scholar] [CrossRef]
- Schuster, F.L.; Visvesvara, G.S. Free-living amoebae as opportunistic and non-opportunistic pathogens of humans and animals. Int. J. Parasitol. 2004, 34, 1001–1027. [Google Scholar] [CrossRef]
- Gianinazzi, C.; Schild, M.; Wüthrich, F.; Müller, N.; Schürch, N.; Gottstein, B. Potentially human pathogenic Acanthamoeba isolated from a heated indoor swimming pool in Switzerland. Exp. Parasitol. 2009, 121, 180–186. [Google Scholar] [CrossRef] [PubMed]
- Rivera, F.; Rosas, I.; Castillo, M.; Chávez, M.; Gómez, R.; Chío, R.E.; Islas, J. Pathogenic and free-living protozoa cultured from the nasopharyngeal and oral regions of dental patients: II. Environ. Res. 1986, 39, 364–371. [Google Scholar] [CrossRef]
- Mergeryan, H. The prevalence of Acanthamoeba in the human environment. Rev. Infect. Dis. 1991, 13, 390–391. [Google Scholar] [CrossRef] [PubMed]
- Mendoza Cavazos, C.; Knoll, L.J. Entamoeba histolytica: Five facts about modeling a complex human disease in rodents. PLoS Pathogens 2020, 16, e1008950. [Google Scholar] [CrossRef] [PubMed]
- WHO: World Health Organization. Entamoeba taxonomy. Bull. World Health Organ. 1997, 75, 291–294. [Google Scholar]
- Carrero, J.C.; Reyes-López, M.; Serrano-Luna, J.; Shibayama, M.; Unzueta, J.; León-Sicairos, N.; De la Garza, M. Intestinal amoebiasis: 160 years of its first detection and remains as a health problem in developing countries. Int. J. Med. Microbiol. 2020, 310, 1513–1558. [Google Scholar] [CrossRef]
- Gelman, B.B.; Rauf, S.J.; Nader, R.; Popov, V.; Bokowski, J.; Chaljub, G.; Nauta, H.W.; Visvesvara, G.S. Amoebic encephalitis due to Sappinia diploidea. J. Am. Med Assoc. 2001, 285, 2450–2451. [Google Scholar] [CrossRef]
- Trabelsi, H.; Dendana, F.; Sellami, A.; Sellami, H.; Cheikhrouhou, F.; Neji, S.; Makni, F.; Ayadi, A. Pathogenic free-living amoebae: Epidemiology and clinical review. Pathol. Biol. 2012, 60, 399–405. [Google Scholar] [CrossRef]
- Stothard, D.R.; Schroeder-Diedrich, J.M.; Awwad, M.H.; Gast, R.J.; Ledee, D.R.; Rodriguez-Zaragoza, S.; Dean, C.L.; Fuerst, P.A.; Byers, T.J. The evolutionary history of the genus Acanthamoeba and the identification of eight new 18S rRNA gene sequence types. J. Eukaryot. Microbiol. 1998, 45, 45–54. [Google Scholar] [CrossRef] [PubMed]
- Horn, M.; Fritsche, T.R.; Gautom, R.K.; Schleifer, K.H.; Wagner, M. Novel bacterial endosymbionts of Acanthamoeba spp. related to the Paramecium caudatum symbiont Caedibacter caryophilus. Environ. Microbiol. 1999, 1, 357–367. [Google Scholar] [CrossRef] [PubMed]
- Gast, R.J. Development of an Acanthamoeba-specific reverse dot-blot and the discovery of a new ribotype. J. Eukaryot. Microbiol. 2001, 48, 609–615. [Google Scholar] [PubMed]
- Hewett, M.K.; Robinson, B.S.; Monis, P.T.; Saint, C.P. Identification of a new Acanthamoeba 18S rRNA Gene Sequence Type, Corresponding to the Species Acanthamoeba jacobsi Sawyer, Nerad and Visvesvara, 1992 (Lobosea: Acanthamoebidae). Acta Protozool. 2003, 42, 325–329. [Google Scholar]
- Corsaro, D.; Venditti, D. Detection of Chlamydiae from freshwater environments by PCR, amoeba coculture and mixed coculture. Res. Microbiol. 2009, 160, 547–552. [Google Scholar] [CrossRef] [PubMed]
- Nuprasert, W.; Putaporntip, C.; Pariyakanok, L.; Jongwutiwes, S. Identification of a novel T17 genotype of Acanthamoeba from environmental isolates and T10 genotype causing keratitis in Thailand. J. Clin. Microbiol. 2010, 48, 4636–4640. [Google Scholar] [CrossRef] [PubMed]
- Khan, N.A. Acanthamoeba: Biology and increasing importance in human health. Fems Microbiol. Rev. 2006, 30, 564–595. [Google Scholar] [CrossRef]
- Greub, G.; Raoult, D. Microorganisms resistant for free-living amoebae. Clin. Microbiol. Rev. 2004, 17, 413–433. [Google Scholar] [CrossRef]
- Strassmann, J.E.; Shu, L. Ancient bacteria–amoeba relationships and pathogenic animal bacteria. PLoS Biol. 2017, 15, e2002460. [Google Scholar] [CrossRef]
- Scheid, P.L. Vermamoeba vermiformis—A Free-Living Amoeba with Public Health and Environmental Health Significance. Open Parasitol. J. 2019, 7, 40–47. [Google Scholar] [CrossRef]
- Park, J.S. First Record of Potentially Pathogenic Amoeba Vermamoeba vermiformis (Lobosea: Gymnamoebia) Isolated from a Freshwater of Dokdo Island in the East Sea, Korea. Animal Systematics. Evol. Divers. 2016, 32, 1–8. [Google Scholar] [CrossRef]
- Molmeret, M.; Horn, M.; Wagner, M.; Santic, M.; Kwaik, Y.A. Amoebae as Training Grounds for Intracellular Bacterial Pathogens. Appl. Environ. Microbiol. 2005, 71, 20–28. [Google Scholar] [CrossRef]
- Swords, W.E.; Guenthner, P.C.; Birkness, K.A.; Lal, R.B.; Dezzutti, C.S.; Quinn, F.D. Mycobacterium xenopi multiplies within human macrophages and enhances HIV replication in vitro. Microb. Pathog. 2006, 40, 41–47. [Google Scholar] [CrossRef]
- Drancourt, M.; Adékambi, T.; Raoult, D. Interactions between Mycobacterium xenopi, amoeba and human cells. J. Hosp. Infect. 2007, 65, 138–142. [Google Scholar] [CrossRef]
- Greub, G.; La Scola, B.; Raoult, D. Amoebae-resisting bacteria isolated from human nasal swabs by amoebal coculture. Emerg. Infect. Dis. 2004, 10, 470–477. [Google Scholar] [CrossRef]
- Moliner, C.; Fournier, P.; Raoult, D. Genome analysis of microorganisms living in amoebae reveals a melting pot of evolution. Fems Microbiol. Rev. 2010, 34, 281–294. [Google Scholar] [CrossRef] [PubMed]
- Rowbotham, T.J. Current views on the relationships between amoebae, legionellae and man. Isr. J. Med Sci. 1986, 22, 678–689. [Google Scholar]
- Jadin, J.B. Amibes limax: Vecteurs possibles de Mycobactéries et de M. leprae. Acta Leprol. 1975, 59, 57–67. [Google Scholar]
- Thom, S.; Warhurst, D.; Drasar, B.S. Association of Vibrio cholerae with freshwater amoebae. J. Med Microbiol. 1992, 36, 303–306. [Google Scholar] [CrossRef] [PubMed]
- Olivier, K.N.; Weber, D.J.; Lee, J.H.; Handler, A.; Tudor, G.; Molina, P.L.; Tomashefski, J.; Knowles, M.R. Nontuberculous mycobacteria. II: Nested-cohort study of impact on cystic fibrosis lung disease. Am. J. Respir. Crit. Care Med. 2003, 167, 835–840. [Google Scholar] [CrossRef] [PubMed]
- Wallace, R.J., Jr.; Zhang, Y.; Wilson, R.W.; Mann, L.; Rossmoore, H. Presence of a single genotype of the newly described species Mycobacterium immunogenum in industrial metalworking fluids associated with hypersensitivity pneumonitis. Appl. Environ. Microbiol. 2002, 68, 5580–5584. [Google Scholar] [CrossRef] [PubMed]
- Meyers, H.; Brown-Elliott, B.A.; Moore, C.; Curry, J.; Truong, C.; Zhang, Y.; Wallace, R.J., Jr. An outbreak of Mycobacterium chelonae infection following liposuction. Clin. Infect. Dis. 2002, 34, 1500–1507. [Google Scholar] [CrossRef]
- Vugia, D.J.; Jang, Y.; Zizek, C.; Ely, J.; Winthrop, K.L.; Desmond, E. Mycobacteria in nail salon whirlpool footbaths, California. Emerg. Infect. Dis. 2005, 11, 616–618. [Google Scholar] [CrossRef]
- Trupiano, J.K.; Sebek, B.A.; Goldfarb, J.; Levy, L.R.; Hall, G.S.; Procop, G.W. Mastitis due to Mycobacterium abscessus after body piercing. Clin. Infect. Dis. 2001, 33, 131–134. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Galil, K.; Miller, L.A.; Yakrus, M.A.; Wallace, R.J., Jr.; Mosley, D.G.; England, B.; Huitt, G.; McNeil, M.M.; Perkins, B.A. Abscesses due to Mycobacterium abscessus linked to injection of unapproved alternative medication. Emerg. Infect. Dis. 1999, 5, 681–687. [Google Scholar] [CrossRef] [PubMed]
- Potgieter, N.; Becker, P.; Ehlers, M. Evaluation of the CDC safe water-storage intervention to improve the microbiological quality of point-of-use drinking water in rural communities in South Africa. Water SA 2009, 35, 505–516. [Google Scholar] [CrossRef]
- Traoré, A.; Mulaudzi, K.; Chari, G.; Foord, S.; Mudau, L.; Barnard, T.; Potgieter, N. The Impact of Human Activities on Microbial Quality of Rivers in the Vhembe District, South Africa. Int. J. Environ. Res. Public Health 2016, 13, 817. [Google Scholar] [CrossRef]
- Sobsey, M.D. Managing Water in the Home: Accelerated Health Gains from Improved Water Supply; WHO: Geneva, Switzerland, 2002. [Google Scholar]
- Potgieter, N.; Mudau, L.S.; Maluleke, F.R.S. The microbiological quality of ground water sources used by rural communities in the Limpopo Province, South Africa. Water Sci. Technol. 2006, 54, 371–377. [Google Scholar] [CrossRef]
- Taonameso, S.; Mudau, L.S.; Traoré, A.N.; Potgieter, N. Borehole water: A potential health risk to rural communities in South Africa. Water Sci. Technol. Water Supply 2019, 19, 128–136. [Google Scholar] [CrossRef]
- Lantagne, D. Investigation of the Potters for Peace Colloidal Silver Impregnated Ceramic Filter—Report 1, Intrinsic Effectiveness; Alethia Environmental: Alston, MA, USA, 2001. [Google Scholar]
- Omar, K.B.; Potgieter, N.; Barnard, T.G. Development of a rapid screening method for the detection of pathogenic Escherichia coli using a combination of Colilert® Quanti-Trays/2000 and PCR. Water Supply 2010, 10, 7–13. [Google Scholar] [CrossRef]
- Schroeder, J.M.; Booton, G.C.; Hay, J.; Niszl, I.A.; Seal, D.V.; Markus, M.B.; Fuerst, P.A.; Byers, T.J. Use of subgenic 18S ribosomal DNA PCR and sequencing for genus and genotype identification of acanthamoebae from humans with keratitis and from sewage sludge. J. Clin. Microbiol. 2001, 39, 1903–1911. [Google Scholar] [CrossRef]
- Corsaro, D.; Walochnik, J.; Köhsler, M.; Rott, M.B. Acanthamoeba misidentification and multiple labels: Redefining genotypes T16, T19, and T20 and proposal for Acanthamoeba micheli sp. nov. (genotype T19). Parasitol. Res. 2015, 114, 2481–2490. [Google Scholar] [CrossRef]
- International Union against Tuberculosis and Lung Disease. Technical guide for sputum examination for tuberculosis by direct microscopy. Bull. Int. Union Tuberc. Lung Dis. 1978, 2, 4–16. [Google Scholar]
- Frank, J.A.; Reich, C.I.; Sharma, S.; Weisbaum, J.S.; Wilson, B.A.; Olsen, G.J. Critical evaluation of two primers commonly used for amplification of bacterial 16S rRNA genes. Appl. Environ. Microbiol. 2008, 74, 2461–2470. [Google Scholar] [CrossRef]
- Fredriksson, N.J.; Hermansson, M.; Wilén, B.M. The Choice of PCR Primers Has Great Impact on Assessments of Bacterial Community Diversity and Dynamics in a Wastewater Treatment Plant. PLoS ONE 2013, 8, e76431. [Google Scholar] [CrossRef] [PubMed]
- Hughes, M.S.; Ball, N.W.; Beck, L.A.; de Lisle, G.W.; Skuce, R.A.; Neill, S.D. Determination of the etiology of presumptive feline leprosy by 16S rRNA gene analysis. J. Clin. Microbiol. 1997, 35, 2464–2467. [Google Scholar] [CrossRef] [PubMed]
- Miyamoto, H.; Yamamoto, H.; Arima, K.; Fujii, J.; Maruta, K.; Izu, K.; Shiomori, T.; Yoshida, S. Development of a new semi-nested PCR method for detection of Legionella species and its application to surveillance of legionellae in hospital cooling tower water. Appl. Environ. Microbiol. 1997, 63, 2489–2494. [Google Scholar] [CrossRef]
- Altschul, S.F.; Madden, T.L.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [PubMed]
- SANS 241–1, 2015. South African National Standards (SANS) Ed 2 for Drinking Water. Available online: http://www.sabs.co.za (accessed on 3 October 2016).
- Department of Water Affairs Forestry (DWAF). South African Water Quality Guidelines, 2nd ed.; Domestic Use; The Government Printer: Pretoria, South Africa, 1996; Volume 2.
- Pawlowski, J.; Adl, S.M.; Audic, S.; Bass, D.; Belbahri, L.; Berney, C.; Bowser, S.; Cepicka, I.; Decelle, J.; Dunthorn, M.; et al. CBOL protist working group: Barcoding eukaryotic richness beyond the animal, plant and fungal kingdoms. PLoS Biol. 2012, 10, e1001419. [Google Scholar] [CrossRef] [PubMed]
- Risse-Buhl, U.; Herrmann, M.; Lange, P.; Akob, D.M.; Pizani, N.; Schönborn, W.; Totsche, K.U.; Küsel, K. Phagotrophic Protist Diversity in the Groundwater of a Karstified Aquifer—Morphological and Molecular Analysis. J. Eukaryot. Microbiol. 2013, 60, 467–479. [Google Scholar] [CrossRef]
- Neff, R.J. Purification, Axenic Cultivation, and Description of a Soil Amoeba, Acanthamoeba sp. J. Protozool. 1957, 4, 176–182. [Google Scholar] [CrossRef]
- Arnalich-Montiel, F.; Lumbreras-Fernández, B.; Martín-Navarro, C.M.; Valladares, B.; Lopez-Velez, R.; Morcillo-Laiz, R.; Lorenzo-Morales, J. Influence of Acanthamoeba Genotype on Clinical Course and Outcomes for Patients with Acanthamoeba keratitis in Spain. J. Clin. Microbiol. 2014, 52, 1213–1216. [Google Scholar] [CrossRef] [PubMed]
- Corsaro, D.; Köhsler, M.; Montalbano Di Filippo, M.; Venditti, D.; Monno, R.; Di Cave, D.; Berrilli, F.; Walochnik, J. Update on Acanthamoeba jacobsi genotype T15, including full-length 18S rDNA molecular phylogeny. Parasitol. Res. 2017, 116, 1273–1284. [Google Scholar] [CrossRef]
- Baumgartner, M.; Yapi, A.; Gröbner-Ferreira, R.; Stetter, K.O. Cultivation and properties of Echinamoeba thermarum n. sp., an extremely thermophilic amoeba thriving in hot springs. Extremophiles 2003, 7, 267–274. [Google Scholar] [CrossRef] [PubMed]
- Page, F.C. Taxonomic criteria for limax amoebae, with descriptions of 3 new species of Hartmannella and 3 of Vahlkampfia. J. Protozool. 1967, 14, 499–521. [Google Scholar] [CrossRef]
- Tachibana, H.; Yanagi, T.; Lama, C.; Pandey, K.; Feng, M.; Kobayashi, S.; Sherchand, J.B. Prevalence of Entamoeba nuttalli infection in wild rhesus macaques in Nepal and characterization of the parasite isolates. Parasitol. Int. 2013, 62, 230–235. [Google Scholar] [CrossRef] [PubMed]
- Michel, R.; Smirnov, A. The genus Flamella Schaeffer, 1926 (lobosea, gymnamoebia), with description of two new species. Eur. J. Protistol. 1999, 35, 403–410. [Google Scholar] [CrossRef]
- Shmakova, L.; Bondarenko, N.; Smirnov, A. Viable Species of Flamella (Amoebozoa: Variosea) Isolated from Ancient Arctic Permafrost Sediments. Protist 2016, 167, 13–30. [Google Scholar] [CrossRef]
- Kudryavtsev, A.; Wylezich, C.; Schlegel, M.; Walochnik, J.; Michel, R. Ultrastructure, SSU rRNA gene sequences and phylogenetic relationships of Flamella Schaeffer, 1926 (Amoebozoa), with description of three new species. Protist 2009, 160, 21–40. [Google Scholar] [CrossRef]
- Fields, B.S. Legionella and protozoa: Interaction of a pathogen and its natural environment. In Legionella: Current Status and Emerging Perspectives; Barbaree, J.M., Breiman, R.F., Dufour, A.P., Eds.; American Society for Microbiology: Washington, DC, USA, 1993; pp. 129–136. [Google Scholar]
- Page, F.C. Platyamoeba stenopodia. ng, n. sp., a Freshwater Amoeba. J. Protozool. 1969, 16, 437–441. [Google Scholar] [CrossRef] [PubMed]
- Pagnier, I.; Valles, C.; Raoult, D.; La Scola, B. Isolation of Vermamoeba vermiformis and associated bacteria in hospital water. Microb. Pathog. 2015, 80, 14–20. [Google Scholar] [CrossRef]
- Javanmard, E.; Niyyati, M.; Lorenzo-Morales, J.; Lasjerdi, Z.; Behniafar, H.; Mirjalali, H. Molecular identification of waterborne free living amoebae (Acanthamoeba, Naegleria and Vermamoeba) isolated from municipal drinking water and environmental sources, Semnan province, north half of Iran. Exp. Parasitol. 2017, 183, 240–244. [Google Scholar] [CrossRef] [PubMed]
- Mulec, J.; Dietersdorfer, E.; Üstüntürk-Onan, M.; Walochnik, J. Acanthamoeba and other free-living amoebae in bat guano, an extreme habitat. Parasitol. Res. 2016, 115, 1375–1383. [Google Scholar] [CrossRef]
- Van Wichelen, J.; D’Hondt, S.; Claeys, M.; Vyverman, W.; Berney, C.; Bass, D.; Vanormelingen, P. A Hotspot of Amoebae Diversity: 8 New Naked Amoebae Associated with the Planktonic Bloom-forming Cyanobacterium Microcystis. Acta Protozool. 2015, 55, 61–87. [Google Scholar]
- Gray, J.S.; Birmingham, J.M.; Fenton, J.I. Got black swimming dots in your cell culture? Identification of Achromobacter as a novel cell culture contaminant. Biologicals 2010, 38, 273–277. [Google Scholar] [CrossRef] [PubMed]
- Coenye, T.; Vancanneyt, M.; Falsen, E.; Swings, J.; Vandamme, P. Achromobacter insolitus sp. nov., Achromobacter spanius sp. nov., from human clinical samples. Int. J. Syst. Evol. Microbiol. 2003, 53, 1819–1824. [Google Scholar] [CrossRef][Green Version]
- Gomila, M.; Tvrzová, L.; Teshim, A.; Sedlácek, I.; González-Escalona, N.; Zdráhal, Z.; Sedo, O.; González, J.F.; Bennasar, A.; Moore, E.R.; et al. Achromobacter marplatensis sp. nov., isolated from a pentachlorophenol-contaminated soil. Int. J. Syst. Evol. Microbiol. 2011, 61, 2231–2237. [Google Scholar] [CrossRef] [PubMed]
- Busse, H.J.; Auling, G. Genus II Achromobacter. Yabuuchi and Yano 1981, 477VP emend. Yabuuchi, Kawamura, Kosako and Ezaki 1998a, 1083. In Bergey’s Manual of Systematic Bacteriology, 2nd ed.; Brenner, D.J., Krieg, N.R., Garrity, G.M., Staley, J.T., Boone, D.R., de Vos, P., Goodfellow, M., Rainey, F.A., Schleifer, K.H., Eds.; Springer: New York, NY, USA, 2005; Volume 2C, pp. 658–662. [Google Scholar]
- Duggan, J.M.; Goldstein, S.J.; Chenoweth, C.E.; Kauffman, C.A.; Bradley, S.F. Achromobacter xylosoxidans bacteremia: Report of four cases and review of the literature. Clin. Infect. Dis. 1996, 23, 569–576. [Google Scholar] [CrossRef]
- Sandu, C.; Chiribau, C.; Sachelaru, P.; Brandsch, R. Plasmids for Nicotine-Dependent and Independent Gene Expression in Arthrobacter nicotinovorans and Other Arthrobacter Species. Appl. Environ. Microbiol. 2005, 71, 8920–8924. [Google Scholar] [CrossRef]
- Urakami, T.; Oyanag, H.; Arak, H.; Suzuki, K.; Komagata, K. Recharacterization and Emended Description Mycoplana and Description of Two New of the Genus Species, Mycoplana ramosa and Mycoplana segnis. Int. J. Syst. Bacteriol. 1990, 40, 434–442. [Google Scholar] [CrossRef]
- Murugaiyan, J.; Krueger, K.; Roesler, U.; Weinreich, J.; Schierack, P. Assessment of species and antimicrobial resistance among Enterobacteriaceae isolated from mallard duck faeces. Environ. Monit. Assess. 2015, 187, 127. [Google Scholar] [CrossRef] [PubMed]
- Nyangacha, R.M.; Odongo, D.; Oyieke, F.; Ochwoto, M.; Korir, R.; Ngetich, R.K.; Nginya, G.; Makwaga, O.; Bii, C.; Mwitari, P.; et al. Secondary bacterial infections and antibiotic resistance among tungiasis patients in Western, Kenya. PLoS Negl. Trop. Diseases 2017, 11, e0005901. [Google Scholar] [CrossRef] [PubMed]
- Brenner, D.J.; McWhorter, A.C.; Kai, A.; Steigerwalt, A.G.; Farmer, J.J. Enterobacter asburiae sp. nov., a new species found in clinical specimens, and reassignment of Erwinia dissolvens and Erwinia nimipressuralis to the genus Enterobacter as Enterobacter dissolvens comb. nov., Enterobacter nimipressuralis comb. nov. J. Clin. Microbiol. 1986, 23, 1114–1120. [Google Scholar] [CrossRef]
- Stock, I.; Wiedemann, B. Natural antibiotic susceptibility of Enterobacter amnigenus, Enterobacter cancerogenus, Enterobacter gergoviae and Enterobacter sakazakii strains. Clin. Microbiol. Infect. 2002, 8, 564–578. [Google Scholar] [CrossRef][Green Version]
- Hoffmann, H.; Stindl, S.; Ludwig, W.; Stumpf, A.; Mehlen, A.; Heesemann, J.; Monget, D.; Schleifer, K.H.; Roggenkamp, A. Reassignment of Enterobacter dissolvens to Enterobacter cloacae as E. cloacae subspecies dissolvens comb. nov., emended description of Enterobacter asburiae and Enterobacter kobei. Syst. Appl. Microbiol. 2005, 28, 196–205. [Google Scholar] [CrossRef] [PubMed]
- Garazzino, S.; Aprato, A.; Maiello, A.; Massé, A.; Biasibetti, A.; De Rosa, F.G.; Di Perri, G. Osteomyelitis Caused by Enterobacter cancerogenus Infection following a Traumatic Injury: Case Report and Review of the Literature. J. Clin. Microbiol. 2005, 43, 1459–1461. [Google Scholar] [CrossRef]
- Podschun, R.; Ullmann, U. Klebsiella spp. as nosocomial pathogens: Epidemiology, taxonomy, typing methods, and pathogenicity factors. Clin. Microbiol. Rev. 1998, 11, 589–603. [Google Scholar] [CrossRef] [PubMed]
- Lowe, C.; Willey, B.; O’Shaughnessy, A.; Lee, W.; Lum, M.; Pike, K.; Larocque, C.; Dedier, H.; Dales, L.; Moore, C.; et al. Outbreak of Extended-Spectrum β-Lactamase–producing Klebsiella oxytoca Infections Associated with Contaminated Handwashing Sinks. Emerg. Infect. Dis. 2012, 18, 1242–1247. [Google Scholar] [CrossRef]
- Yang, B.; Feng, Y.; McNally, A.; Zong, Z. Occurrence of Enterobacter hormaechei carrying blaNDM-1 and blaKPC-2 in China. Diagn. Microbiol. Infect. Dis. 2018, 90, 139–142. [Google Scholar] [CrossRef] [PubMed]
- Podder, M.P.; Rogers, L.; Daley, P.K.; Keefe, G.P.; Whitney, H.G.; Tahlan, K. Klebsiella Species Associated with Bovine Mastitis in Newfoundland. PLoS ONE 2014, 9, e106518. [Google Scholar] [CrossRef]
- Lu, M.G.; Jiang, J.; Liu, L.; Ma, A.P.; Leung, F.C. Complete Genome Sequence of Klebsiella pneumoniae Strain HKUOPLC, a Cellulose-Degrading Bacterium Isolated from Giant Panda Feces. Genome Announc. 2015, 3, e01318-15. [Google Scholar] [CrossRef]
- Reyna-Flores, F.; Barrios-Camacho, H.; Dantán-González, E.; Ramírez-Trujillo, J.A.; Beltrán, L.F.L.A.; Rodríguez-Medina, N.; Garza-Ramos, U.; Suárez-Rodríguez, R. Draft Genome Sequences of Endophytic Isolates of Klebsiella variicola and Klebsiella pneumoniae Obtained from the Same Sugarcane Plant. Genome Announc. 2018, 6. [Google Scholar] [CrossRef]
- Mukherjee, N.; Bartelli, D.; Patra, C.; Chauhan, B.V.; Dowd, S.E.; Banerjee, P. Microbial Diversity of Source and Point-of-Use Water in Rural Haiti—A Pyrosequencing-Based Metagenomic Survey. PLoS ONE 2016, 11, e0167353. [Google Scholar] [CrossRef]
- Gneiding, K.; Frodl, R.; Funke, G. Identities of Microbacterium spp. Encountered in Human Clinical Specimens. J. Clin. Microbiol. 2008, 46, 3646–3652. [Google Scholar] [CrossRef] [PubMed]
- Padda, K.P.; Puri, A.; Zeng, Q.; Chanway, C.P.; Wu, X. Effect of GFP-tagging on nitrogen fixation and plant growth promotion of an endophytic diazotrophic strain of Paenibacillus polymyxa. Botany 2017, 95, 933–942. [Google Scholar] [CrossRef]
- McSpadden Gardener, B.B. Ecology of Bacillus and Paenibacillus spp. in agricultural systems. Phytopathology 2004, 94, 1252–1258. [Google Scholar] [CrossRef] [PubMed]
- Snopková, K.; Sedlár, K.; Bosák, J.; Chaloupková, E.; Provazník, I.; Šmajsa, D. Complete Genome Sequence of Pragia fontium 24613, an Environmental Bacterium from the Family Enterobacteriaceae. Genome Announc. 2015, 3, 1. [Google Scholar] [CrossRef]
- Gilardi, G.L. Infrequently Encountered Pseudomonas Species Causing Infection in Humans. Ann. Intern. Med. 1972, 77, 211–215. [Google Scholar] [CrossRef]
- Cho, J.C.; Tiedje, J.M. Biogeography and degree of endemicity of fluorescent Pseudomonas strains in soil. Appl. Environ. Microbiol. 2000, 66, 5448–5456. [Google Scholar] [CrossRef]
- Alouache, S.; Kada, M.; Messai, Y.; Estepa, V.; Torres, C.; Bakour, R. Antibiotic resistance and extended-spectrum β-Lactamases in isolated bacteria from seawater of Algiers beaches (Algeria). Microbes Environ. 2012, 27, 80–86. [Google Scholar] [CrossRef] [PubMed]
- Wu, H.; Jiang, D.; Cai, P.; Rong, X.; Dai, K.; Liang, W.; Huang, Q. Adsorption of Pseudomonas putida on soil particle size fractions: Effects of solution chemistry and organic matter. J. Soils Sediments 2012, 12, 143–149. [Google Scholar] [CrossRef]
- Molina, L.; Udaondo, Z.; Duque, E.; Fernández, M.; Molina-Santiago, C.; Roca, A.; Porcel, M.; de la Torre, J.; Segura, A.; Plesiat, P.; et al. Antibiotic Resistance Determinants in a Pseudomonas putida Strain Isolated from a Hospital. PLoS ONE 2014, 9, e81604. [Google Scholar] [CrossRef] [PubMed]
- Ayers, J.C. The relationship of organisms of the genus Pseudomonas to the spoilage of meat, poultry and eggs. J. Appl. Bacteriol. 1960, 23, 471–486. [Google Scholar] [CrossRef]
- Thomas, S.B. Psychrophilic micro-organisms in milk and dairy products. Dairy Sci. Abstr. 1958, 20, 356–369, 468. [Google Scholar]
- Willits, C.O.; Frank, H.A.; Bell, R.A. Maple sirup. XIX. Flavor and color through controlled fermentation of maple sap. Food Technol. 1961, 15, 473–474. [Google Scholar]
- Sikorski, J.; Stackebrandt, E.; Wackernagel, W. Pseudomonas kilonensis sp. nov., a bacterium isolated from agricultural soil. Int. J. Syst. Evol. Microbiol. 2001, 51, 1549–1555. [Google Scholar] [CrossRef]
- Kwon, S.W.; Kim, J.S.; Park, I.C.; Yoon, S.H.; Park, D.H.; Lim, C.K.; Go, S.J. Pseudomonas koreensis sp. nov., Pseudomonas umsongensis sp. nov., Pseudomonas jinjuensis sp. nov., novel species from farm soils in Korea. Int. J. Syst. Evol. Microbiol. 2003, 53, 21–27. [Google Scholar] [CrossRef] [PubMed]
- Elomari, M.; Coroler, L.; Verhille, S.; Izard, D.; Leclerc, H. Pseudomonas monteilii sp. nov., isolated from clinical specimens. Int. J. Syst. Bacteriol. 1997, 47, 846–852. [Google Scholar] [CrossRef]
- Tvrzová, L.; Schumann, P.; Sproer, C.; Sedlacek, I.; Pacova, Z.; Sedo, O.; Zdrahal, Z.; Steffen, M.; Lang, E. Pseudomonas moraviensis sp. nov., Pseudomonas vranovensis sp. nov., soil bacteria isolated on nitroaromatic compounds, and emended description of Pseudomonas asplenii. Int. J. Syst. Evol. Microbiol. 2006, 56, 2657–2663. [Google Scholar] [CrossRef]
- Clark, L.L.; Dajcs, J.J.; McLean, C.H.; Bartell, J.G.; Stroman, D.W. Pseudomonas otitidis sp. nov., isolated from patients with otic infections. Int. J. Syst. Evol. Microbiol. 2006, 56, 709–714. [Google Scholar] [CrossRef]
- Behrendt, U.; Ulrich, A.; Schumann, P. Fluorescent pseudomonads associated with the phyllosphere of grasses; Pseudomonas trivialis sp. nov., Pseudomonas poae sp. nov., Pseudomonas congelans sp. nov. Int. J. Syst. Evol. Microbiol. 2003, 53, 1461–1469. [Google Scholar] [CrossRef]
- Elomari, M.; Coroler, L.; Izard, D.; Leclerc, H. A numerical taxonomic study of fluorescent Pseudomonas strains isolated from natural mineral waters. J. Appl. Bacteriol. 1995, 78, 71–81. [Google Scholar] [CrossRef] [PubMed]
- Gardan, L.; Shafik, H.; Belouin, S.; Broch, R.; Grimont, F.; Grimont, P.A.D. DNA relatedness among the pathovars of Pseudomonas syringae and description of Pseudomonas tremae sp. nov., Pseudomonas cannabina sp. nov. (ex Sutic and Dowson 1959). Int. J. Syst. Evol. Microbiol. 1999, 49, 469–478. [Google Scholar] [CrossRef]
- Mohn, W.W.; Wilson, A.E.; Bicho, P.; Moore, E.R. Physiological and phylogenetic diversity of bacteria growing on resin acids. Syst. Appl. Microbiol. 1999, 22, 68–78. [Google Scholar] [CrossRef]
- Park, S.D.; Uh, Y.; Jang, I.H.; Yoon, K.J.; Kim, H.M.; Bae, Y.J. Rhodococcus erythropolis septicaemia in a patient with acute lymphocytic leukaemia. J. Med. Microbiol. 2011, 60, 252–255. [Google Scholar] [CrossRef]
- Bhadra, B.; Roy, P.; Chakraborty, R. Serratia ureilytica sp. nov., a novel urea-utilizing species. Int. J. Syst. Evol. Microbiol. 2005, 55, 2155–2158. [Google Scholar] [CrossRef] [PubMed]
- Parkins, M.D.; Floto, R.A. Emerging bacterial pathogens and changing concepts of bacterial pathogenesis in cystic fibrosis. J. Cyst. Fibros. 2015, 14, 293–304. [Google Scholar] [CrossRef]
- Chang, Y.; Lin, C.; Chen, Y.; Hsueh, P. Update on infections caused by Stenotrophomonas maltophilia with particular attention to resistance mechanisms and therapeutic options. Front. Microbiol. 2015, 6, 893. [Google Scholar] [CrossRef] [PubMed]
- Brooke, J.S. Stenotrophomonas maltophilia: And Emerging Global Opportunistic Pathogen. Clin. Microbiol. Rev. 2012, 25, 2–41. [Google Scholar] [CrossRef] [PubMed]
- Wolf, A.; Fritze, A.; Hagemann, M.; Berg, G. Stenotrophomonas rhizophila sp. nov., a novel plant-associated bacterium with antifungal properties. Int. J. Syst. Evol. Microbiol. 2002, 52, 1937–1944. [Google Scholar]
- Mishra, S.; Singh, B.R.; Naqvi, A.H.; Singh, H.B. Potential of biosynthesized silver nanoparticles using Stenotrophomonas sp. BHU-S7 (MTCC 5978) for management of soil-borne and foliar phytopathogens. Sci. Rep. 2017, 7, 45154. [Google Scholar] [CrossRef]
- Phung, L.T.; Rimble, W.L.; Meyer, F.; Gilbert, J.A.; Silver, S. Draft Genome Sequence of Alcaligenes faecalis subsp. faecalis NCIB 8687 (CCUG 2071). J. Bacteriol. 2012, 194, 5153. [Google Scholar] [CrossRef]
- Sarkar, J.K.; Choudhury, B.; Tribedi, B.P. Alcaligenes faecalis; its systematic study. Indian J. Med Res. 1959, 47, 1–12. [Google Scholar]
- Rehfuss, M.; Urban, J. Alcaligenes faecalis subsp. phenolicus subsp. nov.—A phenol-degrading, denitrifying bacterium isolated from a graywater bioprocessor. Syst. Appl. Microbiol. 2005, 28, 421–429. [Google Scholar]
- Wong, W.C.; Preece, T.F. Pseudomonas tolaasi in Mushroom Crops: A Note on Primary and Secondary Sources of the Bacterium on a Commercial Farm in England. J. Appl. Microbiol. 1980, 49, 305–314. [Google Scholar]
- Stackebrandt, E.; Fowler, V.J.; Fiedler, F.; Seiler, H. Taxonomic studies on Arthrobacter nicotianae and related taxa: Description of Arthrobacter uratoxydans sp. nov., Arthrobacter sulfureus sp. nov., reclassification of Brevibacterium protophormiae as Arthrobacter protophormiae comb. nov. Syst. Appl. Microbiol. 1983, 4, 470–486. [Google Scholar] [CrossRef]
- Häggblom, M.M.; Nohynek, L.J.; Palleroni, N.J.; Kronqvist, K.; Nurmiaho-Lassila, E.L.; Salkinoja-Salonen, M.S.; Klatte, S.; Kroppenstedt, R.M.; Hägglblom, M.M. Transfer of polychlorophenol-degrading Rhodococcus chlorophenolicus (Apajalahti et al. 1986) to the genus Mycobacterium as Mycobacterium chlorophenolicum comb. nov. Int. J. Syst. Evol. Microbiol. 1994, 44, 485–493. [Google Scholar] [CrossRef]
- Kempker, R.R.; Kipiani, M.; Mirtskhulava, V.; Tukvadze, N.; Magee, M.J.; Blumberg, H.M. Acquired Drug Resistance in Mycobacterium tuberculosis and Poor Outcomes among Patients with Multidrug-Resistant Tuberculosis. Emerg. Infect. Dis. 2015, 21, 992–1001. [Google Scholar] [CrossRef]
- 136. Whitman, W.; Goodfellow, M.; Kämpfer, P.; Busse, H.; Trujillo, M.; Ludwig, W.; Suzuki, K.; Parte, A. Bergey’s Manual of Systematic Bacteriology; Springer: New York, NY, USA, 2012; ISBN 978-0-387-68233-4. Available online: https://www.springer.com/series/4157 (accessed on 13 March 2019).
- Satsuma, K.; Masuda, M. Reductive dechlorination of methoxychlor by bacterial species of environmental origin: Evidence for primary biodegradation of methoxychlor in submerged environments. J. Agric. Food Chem. 2012, 60, 2018–2023. [Google Scholar] [CrossRef] [PubMed]
- Cooksey, R.C.; de Waard, J.H.; Yakrus, M.A.; Rivera, I.; Chopite, M.; Toney, S.R.; Morlock, G.P.; Butler, W.R. Mycobacterium cosmeticum sp. nov., a novel rapidly growing species isolated from a cosmetic infection and from a nail salon. Int. J. Syst. Evol. Microbiol. 2004, 54, 2385–2391. [Google Scholar] [CrossRef] [PubMed]
- Shojaei, H.; Magee, J.G.; Freeman, R.; Yates, M.; Horadagoda, N.U.; Goodfellow, M. Mycobacterium elephantis sp. nov., a rapidly growing non-chromogenic Mycobacterium isolated from an elephant. Int. J. Syst. Evol. Microbiol. 2000, 50, 1817–1820. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Lévy-Frébault, V.; Daffé, M.; Goh, K.S.; Lanéelle, M.A.; Asselineau, C.; David, H.L. Identification of Mycobacterium fortuitum and Mycobacterium chelonei. J. Clin. Microbiol. 1983, 17, 744–752. [Google Scholar] [CrossRef] [PubMed]
- Chamoiseau, G. Etiology of farcy in African bovines: Nomenclature of the causal organisms Mycobacterium farcinogenes Chamoiseau and Mycobacterium senegalense (Chamoiseau) comb.nov. Int. J. Syst. Bacteriol. 1979, 29, 407–410. [Google Scholar] [CrossRef][Green Version]
- Tortoli, E.; Rindi, L.; Goh, K.S.; Katila, M.L.; Mariottini, A.; Mattei, R.; Mazzarelli, G.; Suomalainen, S.; Torkko, P.; Rastogi, N. Mycobacterium florentinum sp. nov., isolated from humans. Int. J. Syst. Evol. Microbiol. 2005, 55, 1101–1106. [Google Scholar] [CrossRef]
- Lambaret, O.; Drancourt, M. Mycobacterium gilvum illustrates size-correlated relationships between mycobacteria and Acanthamoeba polyphaga. Appl. Environ. Microbiol. 2013, 79, 1606–1611. [Google Scholar]
- Meier, A.; Kirschner, P.; Schröder, K.H.; Wolters, J.; Kroppenstedt, R.M.; Böttger, E.C. Mycobacterium intermedium sp. nov. Int. J. Syst. Evol. Microbiol. 1993, 43, 204–209. [Google Scholar] [CrossRef]
- Gomila, M.; Ramirez, A.; Gasco, J.; Lalucat, J. Mycobacterium llatzerense sp. nov., a facultatively autotrophic, hydrogen-oxidizing bacterium, isolated from haemodialysis water. Int. J. Syst. Evol. Microbiol. 2008, 58, 2769–2773. [Google Scholar] [CrossRef]
- Van Ingen, J.; Boeree, M.J.; De Lange, W.C.; De Haas, P.E.; Van der Zanden, A.G.; Mijs, W.; Rigouts, L.; Dekhuijzen, P.N.; Van Soolingen, D. Mycobacterium noviomagense sp. nov., clinical relevance evaluated in 17 patients. Int. J. Syst. Evol. Microbiol. 2009, 59, 845–849. [Google Scholar] [CrossRef]
- 147. Hennessee, C.T.; Seo, J.-S.; Alvarez, A.M.; Li, Q.X. Polycyclic aromatic hydrocarbon-degrading species isolated from Hawaiian soils: Mycobacterium crocinum sp. nov., Mycobacterium pallens sp. nov., Mycobacterium rutilum sp. nov., Mycobacterium rufum sp. nov., Mycobacterium aromaticivorans sp. nov. Int. J. Syst. Evol. Microbiol. 2009, 59, 378–387. [Google Scholar] [CrossRef] [PubMed]
- Padgitt, P.J.; Moshier, S.E. Mycobacterium poriferae sp. nov., a scotochromogenic, rapidly growing species isolated from a marine sponge. Int. J. Syst. Bacteriol. 1987, 37, 186–191. [Google Scholar] [CrossRef]
- Whipps, C.M.; Butler, W.R.; Pourahmad, F.; Watral, V.G.; Kent, M.L. Molecular systematics support the revival of Mycobacterium salmoniphilum (ex Ross 1960) sp. nov., nom. rev.; a species closely related to Mycobacterium chelonae. Int. J. Syst. Evol. Microbiol. 2007, 57, 2525–2531. [Google Scholar] [CrossRef]
- Trujillo, M.E.; Velazquez, E.; Kroppenstedt, R.M.; Schumann, P.; Rivas, R.; Mateos, P.F.; Martinez-Molina, E. Mycobacterium psychrotolerans sp. nov., isolated from pond water near a uranium mine. Int. J. Syst. Evol. Microbiol. 2004, 54, 1459–1463. [Google Scholar] [CrossRef] [PubMed]
- Tsukamura, M. Proposal of a new genus, Gordona, for slightly acid-fast organisms occurring in sputa of patients with pulmonary disease and in soil. J. Gen. Microbiol. 1971, 68, 15–26. [Google Scholar] [CrossRef]
- Ross, A.J. Mycobacterium salmoniphilum sp. nov. from salmonoid fishes. Am. Rev. Respir. Dis. 1960, 81, 241–250.–250. [Google Scholar]
- Wallace, R.J., Jr.; Nash, D.R.; Tsukamura, M.; Blacklock, Z.M.; Silcox, V.A. Human Disease Due to Mycobacterium smegmatis. J. Infect. Dis. 1988, 158, 52–59. [Google Scholar] [CrossRef] [PubMed]
- Wallace, R.J., Jr.; Musser, J.M.; Hull, S.I.; Silcox, V.A.; Steele, L.C.; Forrester, G.D.; Labidi, A.; Selander, R.K. Diversity and sources of rapidly growing mycobacteria associated with infections following cardiac bypass surgery. J. Infect. Dis. 1989, 159, 708–716. [Google Scholar] [CrossRef] [PubMed]
- Tsukamura, M. New species of rapidly growing, scotochromogenic mycobacteria, Mycobacterium chubuense Tsukamura, Mycobacterium aichiense Tsukamura, and Mycobacterium tokaiense Tsukamura. Med. Biol. 1973, 86, 13–17. [Google Scholar]
- Suomalainen, S.; Koukila-Kähkölä, P.; Brander, E.; Katila, M.L.; Piilonen, A.; Paulin, L.; Mattson, K. Pulmonary infection caused by an unusual, slowly growing nontuberculous mycobacterium. J. Clin. Microbiol. 2001, 39, 2668–2671. [Google Scholar] [CrossRef]
- Cabello-Vílchez, A.; Mena, R.; Zuñiga, J.; Reyes-Batlle, M.; Piñero, J.; Valladares, B.; Lorenzo-Morales, J. Endosymbiotic Mycobacterium chelonae in a Vermamoeba vermiformis strain isolated from the nasal mucosa of an HIV patient in Lima, Peru. Exp. Parasitol. 2014, 145, S127–S130. [Google Scholar] [CrossRef]
- Wheat, W.H.; Casali, A.L.; Thomas, V.; Spencer, J.S.; Lahiri, R.; Williams, D.L.; McDonnell, G.E.; Gonzalez-Juarrero, M.; Brennan, P.J.; Jackson, M. Long-term survival and virulence of Mycobacterium leprae in amoebal cysts. Plos Negl. Trop. Dis. 2014, 18, 3405. [Google Scholar] [CrossRef] [PubMed]
- Bajrai, L.H.; Benamar, S.; Azhar, E.I.; Robert, C.; Levasseur, A.; Raoult, D.; La Scola, B. Kaumoebavirus, a new virus that clusters with Faustoviruses and Asfarviridae. Viruses 2016, 8, 278. [Google Scholar] [CrossRef] [PubMed]
- Delafont, V.; Mougari, F.; Cambau, E.; Joyeux, M.; Bouchon, D.; Héchard, Y.; Moulin, L. First evidence of amoebae-mycobacteria association in drinking water network. Environ. Sci. Technol. 2014, 48, 11872–11882. [Google Scholar] [CrossRef]
- Cateau, E.; Imbert, C.; Rodier, M.H. Hartmanella vermiformis can be permissive for Pseudomonas aeruginosa. Lett. Appl. Microbiol. 2008, 47, 475–477. [Google Scholar] [CrossRef]
- Maciver, S.K.; Asif, M.; Simmen, M.W.; Lorenzo-Morales, J. A systematic analysis of Acanthamoeba genotype frequency correlated with source and pathogenicity: T4 is confirmed as a pathogen-rich genotype. Eur. J. Protistol. 2013, 49, 217–221. [Google Scholar] [CrossRef]
- Siddiqui, R.; Khan, N.A. War of the microbial worlds: Who is the beneficiary in Acanthamoeba–bacterial interactions? Exp. Parasitol. 2012, 130, 311–313. [Google Scholar] [CrossRef] [PubMed]
- Omaña-Molina, M.; Vanzzini-Zago, V.; Hernandez-Martinez, D.; Gonzalez-Robles, A.; Salazar-Villatoro, L.; Ramirez-Flores, E.; Oregon-Miranda, E.; Lorenzo-Morales, J.; Martinez-Palomo, A. Acanthamoeba genotypes T3 and T4 as causative agents of amoebic keratitis in Mexico. Parasitol. Res. 2016, 115, 873–878. [Google Scholar] [CrossRef]
- Di Cave, D.; Monno, R.; Bottalico, P.; Guerriero, S.; D’Amelio, S.; D’Orazi, C.; Berrilli, F. Acanthamoeba T4 and T15 genotypes associated with keratitis infections in Italy. Eur. J. Clin. Microbiol. Infect. Dis. 2008, 28, 607–612. [Google Scholar] [CrossRef]
- Landell, M.F.; Salton, J.; Caumo, K.; Broetto, L.; Rott, M.B. Isolation and genotyping of free-living environmental isolates of Acanthamoeba spp. from bromeliads in Southern Brazil. Exp. Parasitol. 2013, 134, 290–294. [Google Scholar] [CrossRef]
- Tachibana, H.; Yanagi, T.; Pandey, K.; Cheng, X.J.; Kobayashi, S.; Sherchand, J.B.; Kanbara, H. An Entamoeba sp. strain isolated from rhesus monkey is virulent but genetically different from Entamoeba histolytica. Mol. Biochem. Parasitol. 2007, 153, 107–114. [Google Scholar] [CrossRef] [PubMed]
- Debenham, J.J.; Tysnes, K.; Khunger, S.; Robertson, L.J. Occurrence of Giardia, Cryptosporidium, and Entamoeba in wild rhesus macaques (Macaca mulatta) living in urban and semi-rural North-West India. Int. J. Parasitol. Parasites Wildl. 2017, 6, 29–34. [Google Scholar] [CrossRef] [PubMed]
- Schilde, C.; Schaap, P. The Amoebozoa. Methods Mol. Biol. 2013, 983, 1–15. [Google Scholar]
- Denoncourt, A.M.; Paquet, V.E.; Charette, S.J. Potential role of bacteria packaging by protozoa in the persistence and transmission of pathogenic bacteria. Front. Microbiol. 2014, 5, 240. [Google Scholar] [CrossRef]
- Buse, H.Y.; Donohue, M.J.; Ashbolt, N.J. Hartmannella vermiformis inhibition of Legionella pneumophila cultivability. Microbiol. Ecol. 2013. [Google Scholar] [CrossRef] [PubMed]
- Potgieter, N.; Bolton, M.; Barnard, T.G. Evaluation of Ceramic Filters for Its Effectiveness as a Household Treatment System; WRC Report 1653; Water Research Commission: Pretoria, South Africa, 2011; ISBN 978-1-4312-0191-4. [Google Scholar]
- Lantagne, D. Investigation of the Potters for Peace Colloidal Silver Impregnated Ceramic Filter—Report 2, Field Investigations; Alethia Environmental: Alston, MA, USA, 2001. [Google Scholar]
- WHO. Results of Round II of the WHO International Scheme to Evaluate Household Water Treatment Technologies; World Health Organization: Geneva, Switzerland, 2019; ISBN 978-92-4-151603-7. [Google Scholar]
- Van Halem, D. Ceramic Silver Impregnated Pot Filters for Household Drinking Water Treatment in Developing Countries. Master’s Thesis, Faculty of Civil Engineering, Delft University of Technology, Delft, The Netherlands, 2006. [Google Scholar]
- Van Halem, D.; Heijman, S.; Soppe, A.; van Dijk, J.; Amy, G. Ceramic silver-impregnated pot filters for household drinking water treatment in developing countries: Material characterization and performance study. Water Sci. Technol. 2007, 7, 9–17. [Google Scholar] [CrossRef]
- Visvesvara, G.S. Infections with free-living amebae. Handb. Clin. Neurol. 2013, 114, 153–168. [Google Scholar]
- Yu, H.S.; Jeong, H.J.; Hong, Y.C.; Seol, S.Y.; Chung, D.I.; Kong, H.H. Natural occurrence of Mycobacterium as an endosymbiont of Acanthamoeba isolated from a contact lens storage case. Korean J. Parasitol. 2007, 45, 11–18. [Google Scholar] [CrossRef]
- Niyyati, M.; Lasgerdi, Z.; Lorenzo-Morales, J. Detection and Molecular Characterization of Potentially Pathogenic Free-living Amoebae from Water Sources in Kish Island, Southern Iran. Microbiol. Insights 2015, 8, 1–6. [Google Scholar] [CrossRef]
- Kuchta, J.M.; Navratil, J.S.; Shepherd, M.E.; Wadowsky, R.M.; Dowling, J.N.; States, S.J.; Yee, R.B. Impact of Chlorine and heat on the survival of Hartmannella vermiformis and subsequent growth of Legionella pneumophila. Appl. Environ. Microbiol. 1993, 59, 4096–4100. [Google Scholar] [CrossRef]
- Critchley, M.; Bentham, R. The efficacy of biocides and other chemical additives in cooling water systems in the control of amoebae. J. Appl. Microbiol. 2009, 106, 784–789. [Google Scholar] [CrossRef] [PubMed]
- Fouque, E.; Yefimova, M.; Trouilhé, M.C.; Quellard, N.; Fernandez, B.; Rodier, M.H.; Thomas, V.; Humeau, P.; Héchard, Y. Morphological Study of the Encystment and Excystment of Vermamoeba vermiformis Revealed Original Traits. J. Eukaryot. Microbiol. 2015, 62, 327–373. [Google Scholar] [CrossRef] [PubMed]
- Anwar, A.; Abdalla, S.A.O.; Aslam, Z.; Shah, M.R.; Siddiqui, R.; Khan, N.A. Oleic acid-conjugated silver nanoparticles as efficient anti-amoebic agent against Acanthamoeba castellanii. Parasitol. Res. 2019, 118, 2295–2304. [Google Scholar] [CrossRef]
- Rajendran, K.; Anwar, A.; Khan, N.A.; Aslam, Z.; Shah, M.R.; Siddiqui, R. Oleic Acid Coated Silver Nanoparticles Showed Better in Vitro Amoebicidal Effects against Naegleria fowleri than Amphotericin B. ACS Chem. Neurosci. 2020, 11, 2431–2437. [Google Scholar] [CrossRef] [PubMed]
- Squire, S.A.; Yang, R.; Robertson, I.; Ayi, I.; Squire, D.S.; Ryan, U. Gastrointestinal helminths in farmers and their ruminant livestock from the Coastal Savannah zone of Ghana. Parasitol. Res. 2018, 117, 3183–3194. [Google Scholar] [CrossRef] [PubMed]
- Ngcamphalala, P.; Lamb, J.; Mukaratirwa, S. Molecular identification of hookworm isolates from stray dogs, humans and selected wildlife from South Africa. J. Helminthol. 2020, 94, E39. [Google Scholar] [CrossRef]
- Ward, J.V.; Voelz, N.J.; Harvey, J.H. Groundwater Faunas as Indicators of Groundwater Quality: The South Platte River System; Colorado Water Resources Research Institute Completion Report no. 150; Colorado Water: Denver, CO, USA, 1989; Available online: https://mountainscholar.org/bitstream/handle/10217/3034/CR_150.pdf?sequence=1 (accessed on 3 October 2016).
- Somani, V.; Goldin, Q.; Madhuri, K.P. Occurrence of Rotifers and its Relation to the Water Quality during the Bioremediation process in Lake Kacharali, Thane, MS, India. Int. Res. J. Biol. Sci. 2012, 1, 54–58. [Google Scholar]





| Indicator | Household Storage Water | Household Filtered Water | South African Water Quality Guideline [62] |
|---|---|---|---|
| E. coli count (MPN/100 mL) | 173 (<1–>2419.6) | 23 (<1–>2419.6) | 0 |
| pH | 8.25 (5.65–9.29) | 8.11 (6.87–9.18) | ≥5.0–≤9.7 |
| Turbidity (NTU) | 0.66 (0.11–18.4) | 0.46 (0.17–2.99) | 1 |
| Conductivity (mS/cm) | 91.73 (18.5–185.3) | 74.11 (17.2–221.9) | 170 |
| FLA Species | n | Stored Water | Filtered Water | Published Studies Indicating Where the Organism Was Previously Isolated with References |
|---|---|---|---|---|
| Amoebozoa spp. | 5 | 3 | 2 | Marine-/fresh water and soil [64]; Groundwater [65] |
| Acanthamoeba spp. | 2 | 2 | 0 | Soil [66] |
| Genotype T3 | 1 | 1 | 0 | Corneal biopsy of contact lens wearer [67] |
| Genotype T4 | 2 | 2 | 0 | Corneal scrape of contact lens wearer [67] |
| Genotype T15 | 1 | 1 | 0 | Corresponding to only A. jacobsi [25,68] |
| Genotype T16 | 2 | 2 | 0 | Freshwater pond, Italy [26] |
| A. castellanii str. Neff | 3 | 2 | 1 | Soil [66] |
| Echinamoeba spp. | 1 | 0 | 1 | Hot water springs [69] |
| E. exudans | 2 | 0 | 2 | Leaf litter [70] |
| Entamoeba spp. | ||||
| E. nuttalli | 3 | 2 | 1 | Rhesus macaque [71] |
| Flamella spp. | 2 | 2 | 0 | Depends on species [72] |
| F. beringiania | 1 | 1 | 1 | Siberian permafrost [73] |
| F. fluviatalis | 1 | 1 | 0 | Fresh water [74] |
| * Hartmanella spp. | 2 | 1 | 1 | Potable water supplies [75] |
| Stenamoeba spp. | 2 | 2 | 0 | Freshwater lake sediment [76] |
| Vermamoeba spp. | ||||
| V. vermiformis | 34 | 17 | 17 | Variety of water sources [77,78]; bat guano [79] |
| Vexillifera spp. | ||||
| V. westveldii | 2 | 1 | 1 | Pond water [80] |
| ARB Species | n | Stored Water | Filtered Water | Published Studies Indicating Where the Organism Was Previously Isolated with References |
|---|---|---|---|---|
| Achromobacter spp. | 4 | 3 | 1 | Freshwater; sea water, soil [81] |
| A. insolitus | 1 | 1 | 0 | Laboratory sink, urine, wounds [82] |
| A. marplatensis | 1 | 1 | 0 | Pentachlorophenol-contaminated soil [83] |
| A. spanius | 2 | 1 | 1 | Human clinical samples [82] |
| A. xylosoxidans | 1 | 0 | 1 | Oligotrophic aquatic niches; opportunistic pathogen; human bacteremia; pneumonia [84,85] |
| Arthrobacter spp. | ||||
| A. nicotinovorans | 1 | 1 | 0 | Soil [86] |
| Caulobacter spp. | ||||
| C. segnis | 1 | 1 | 0 | Soil [87] |
| Enterobacter spp. | ||||
| E. amnigenus | 1 | 1 | 0 | Mallard duck intestines; parasitic disease caused by Kenyan sand fly [88,89] |
| E. asburiae | 6 | 6 | 0 | Human clinical samples [90] |
| E. cancerogenus | 2 | 2 | 0 | Human clinical samples [91] |
| E. kobei | 2 | 2 | 0 | Implanted medical devices; nosocomial bacteria in ICU patients [92,93] |
| E. ludwigii | 8 | 6 | 2 | Human clinical samples [92] |
| Klebsiella spp. | ||||
| K. oxytoca | 1 | 1 | 0 | Mammal mucosal surfaces; hospital-acquired pathogen; various human infections [94,95,96] |
| K. variicola | 1 | 1 | 0 | Environmental samples [97,98,99,100] |
| Microbacterium spp. | 1 | 1 | 0 | Human clinical samples [101] |
| M. oxydans | 4 | 3 | 1 | Human clinical samples [101] |
| M. paraoxydans | 12 | 6 | 6 | Human clinical samples [101] |
| Paenibacillus spp. | ||||
| P. validus | 1 | 1 | 0 | Soil, water, rhizosphere, vegetable matters, fargae, insect larvae and human clinical samples [102,103] |
| Pragia spp. | ||||
| P. fontium | 1 | 1 | 0 | Water wells; pipes [104] |
| Pseudomonas spp. | 4 | 1 | 3 | Saprophytes; contaminated human clinical samples [105] |
| P. fluorescens | 7 | 3 | 4 | Plants, soil, rhizosphere, human clinical samples [106,107,108,109] |
| P. geniculata | 1 | 1 | 0 | Refrigerated meat; dairy products; maple tree sap [110,111,112] |
| P. kilonensis | 1 | 1 | 0 | Agricultural soil [113] |
| P. koreensis | 2 | 1 | 1 | Agricultural soil [114] |
| P. monteilii | 1 | 0 | 1 | Human clinical samples [115] |
| P. moraviensis | 1 | 0 | 1 | Soil [116] |
| P. otitidis | 1 | 0 | 1 | Clinical specimen—human ears [117] |
| P. poae | 1 | 1 | 0 | Phyllosphere of grasses [118] |
| P. rhodesiae | 1 | 0 | 1 | Natural mineral water [119] |
| P. tremae | 1 | 1 | 0 | Plants [120] |
| P. vancouverensis | 1 | 1 | 0 | Soil [121] |
| P. putida | 3 | 1 | 2 | Plants, soil, rhizosphere, human clinical samples [106,107,108,109] |
| Rhodococcus spp. | ||||
| R. erythropolis | 1 | 1 | 0 | Soil, rocks, groundwater, seawater, plants, animals, gut of insects [122] |
| Serratia spp. | ||||
| S. ureilytica | 1 | 1 | 0 | River water [123] |
| Stenotrophomonas spp. | ||||
| S. maltophilia | 8 | 4 | 4 | Human pathogen; humid surfaces; medical devices [124,125,126] |
| S. rhizophila | 1 | 0 | 1 | Plant-associated [127,128] |
| Alcaligenes faecalis and Achromobacter marplatensis | 1 | 1 | 0 | A. faecalis—faeces, soil, water, other environments [129,130,131]; |
| A. marplatensis—Pentachlorophenol-contaminated soil [83] | ||||
| Pseudomonas spp. | ||||
| P. poae/P. tolaasii | 1 | 1 | 0 | Phyllosphere of grasses [118]; compost; casting soil in mushroom production [132] |
| P. fluorescens/P. rhodesiae | 1 | 0 | 1 | Plants, soil, rhizosphere; human clinical samples [106,107,108,109]; Natural mineral water [120] |
| Glutamicibacter uratoxydans and Stenotrophomonas maltophilia | 2 | 0 | 2 | G. uratoxydans (previously known as Arthrobacter uratoxydans) in humus soil [133]; S. maltophilia—Human pathogen; humid surfaces; medical devices [124,125,126] |
| Mycobacterium spp. | ||||
| M. chlorophenolicum | 1 | 1 | 0 | Soil; sludge; Pneumonia [134,135] |
| M. chubuense | 4 | 3 | 1 | Water; soil [136,137] |
| M. cosmeticum | 1 | 0 | 1 | Nail salons [138] |
| M. elephantis | 1 | 1 | 0 | Elephant carcass [139] |
| M. fallax | 1 | 1 | 0 | Water [140] |
| M. farcinogenes | 1 | 1 | 0 | Bovine farcy [141] |
| M. florentinum | 10 | 8 | 2 | Sputum; lymph node [142] |
| M. gilvum | 1 | 1 | 0 | Water; soil [143] |
| M. intermedium | 5 | 4 | 1 | Sputum [144] |
| M. llatzerense | 7 | 4 | 3 | Hemodialysis water [145] |
| M. noviomagense | 1 | 1 | 0 | Respiratory samples [146] |
| M. pallens | 3 | 3 | 0 | Soil [147] |
| M. poriferae | 7 | 6 | 1 | Treatment plants [148,149] |
| M. psychrotolerans | 1 | 1 | 0 | Water near uranium mine [150] |
| M. rhodesiae | 1 | 1 | 0 | Human clinical samples; soil [151] |
| M. salmoniphilum | 15 | 15 | 0 | Salmonid fishes [152] |
| M. smegmatis | 1 | 1 | 0 | Environmental samples; rare human pathogen [151,153,154] |
| M. tokaiense | 1 | 1 | 0 | Soil [155] |
| M. triplex | 3 | 2 | 1 | Human pulmonary mycobacteriosis [156] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
van der Loo, C.; Bartie, C.; Barnard, T.G.; Potgieter, N. Detection of Free-Living Amoebae and Their Intracellular Bacteria in Borehole Water before and after a Ceramic Pot Filter Point-of-Use Intervention in Rural Communities in South Africa. Int. J. Environ. Res. Public Health 2021, 18, 3912. https://doi.org/10.3390/ijerph18083912
van der Loo C, Bartie C, Barnard TG, Potgieter N. Detection of Free-Living Amoebae and Their Intracellular Bacteria in Borehole Water before and after a Ceramic Pot Filter Point-of-Use Intervention in Rural Communities in South Africa. International Journal of Environmental Research and Public Health. 2021; 18(8):3912. https://doi.org/10.3390/ijerph18083912
Chicago/Turabian Stylevan der Loo, Clarissa, Catheleen Bartie, Tobias George Barnard, and Natasha Potgieter. 2021. "Detection of Free-Living Amoebae and Their Intracellular Bacteria in Borehole Water before and after a Ceramic Pot Filter Point-of-Use Intervention in Rural Communities in South Africa" International Journal of Environmental Research and Public Health 18, no. 8: 3912. https://doi.org/10.3390/ijerph18083912
APA Stylevan der Loo, C., Bartie, C., Barnard, T. G., & Potgieter, N. (2021). Detection of Free-Living Amoebae and Their Intracellular Bacteria in Borehole Water before and after a Ceramic Pot Filter Point-of-Use Intervention in Rural Communities in South Africa. International Journal of Environmental Research and Public Health, 18(8), 3912. https://doi.org/10.3390/ijerph18083912






