Do Rural Second Homes Shape Commensal Microbiota of Urban Dwellers? A Pilot Study among Urban Elderly in Finland
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Area and Participants
2.2. Sample Collection, DNA Extraction, Amplification, and Sequencing
2.3. Bioinformatics
2.4. Statistical Analyses
3. Results
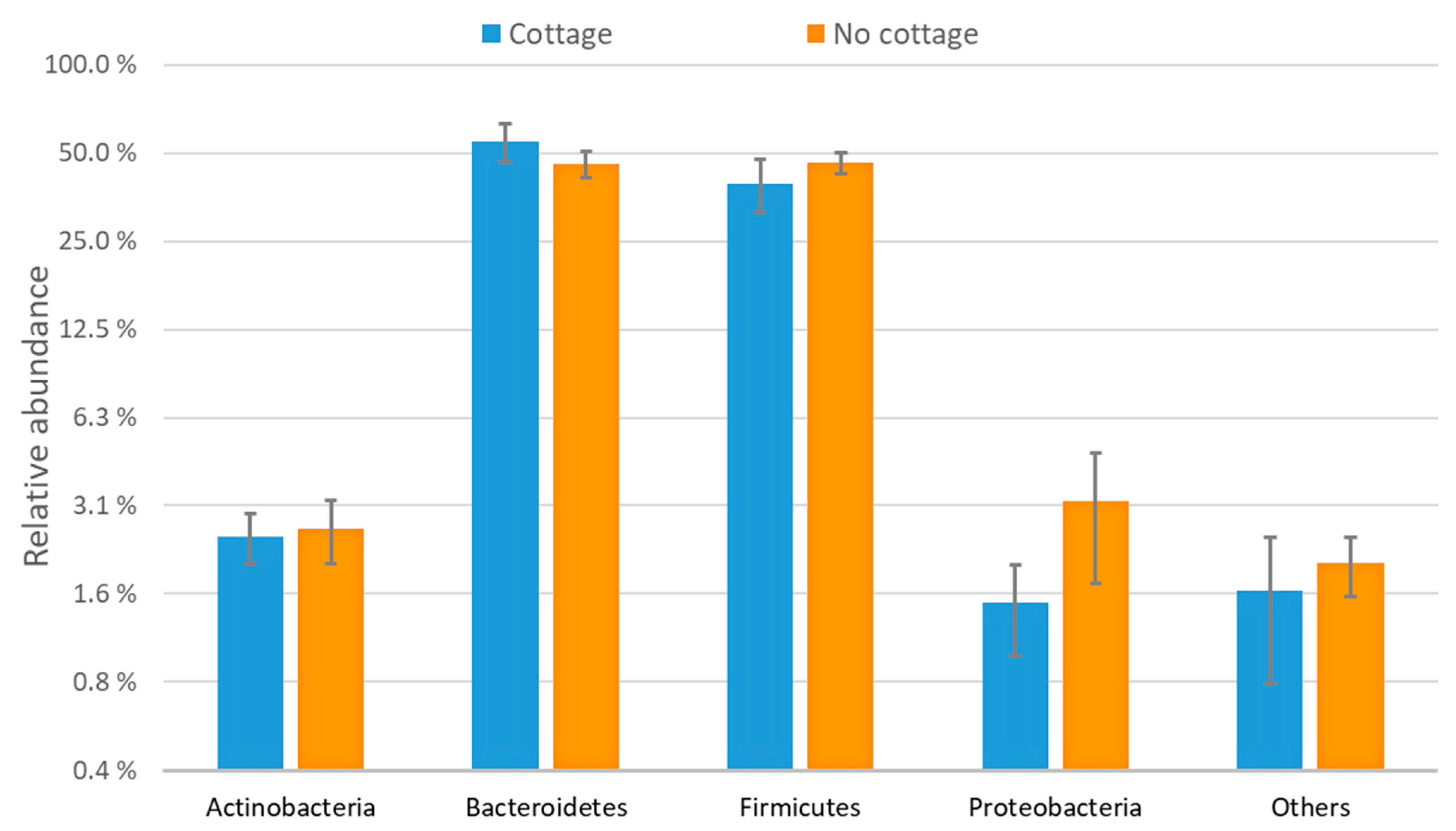
3.1. Characterization of the Bacterial Communities
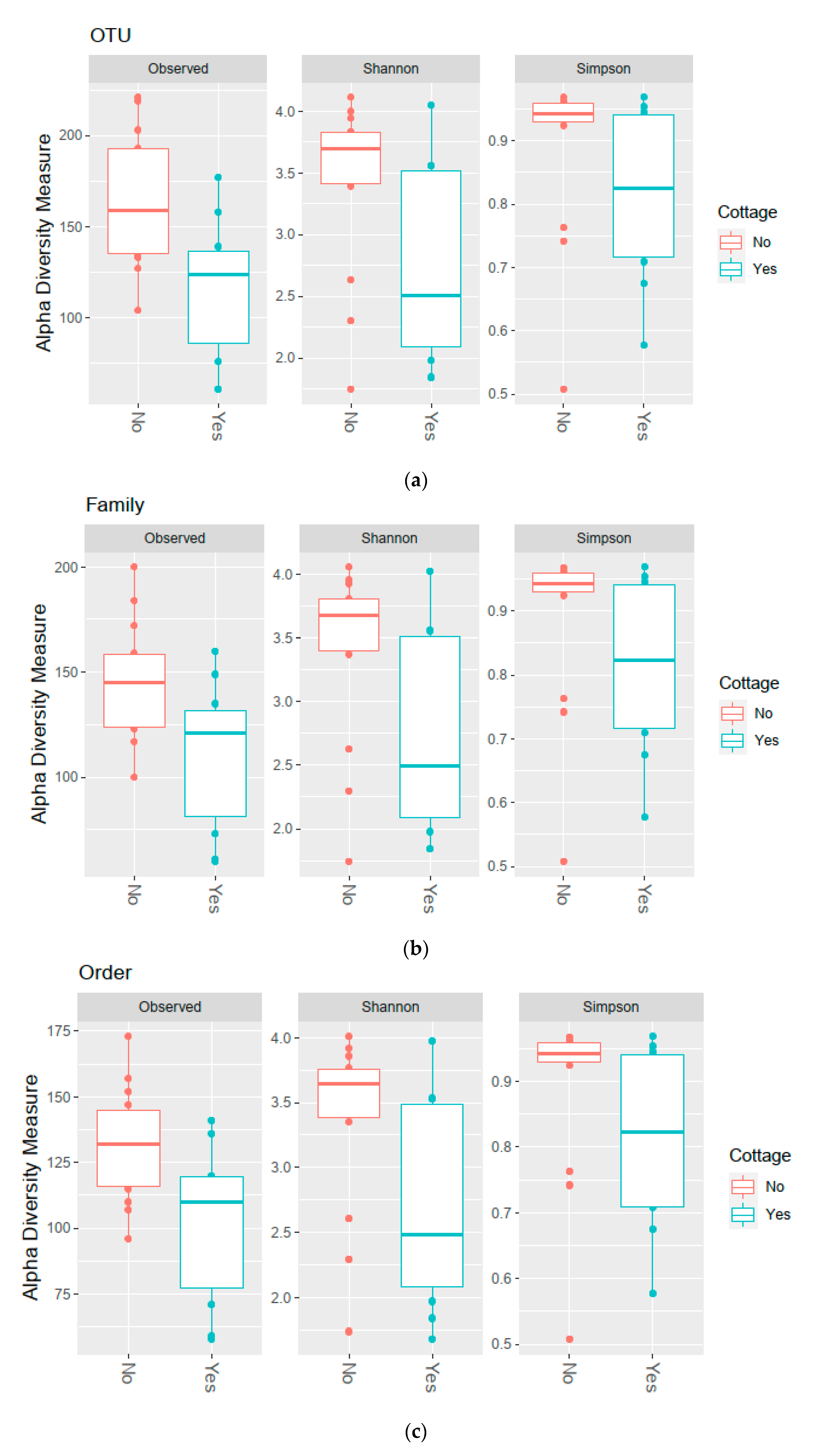
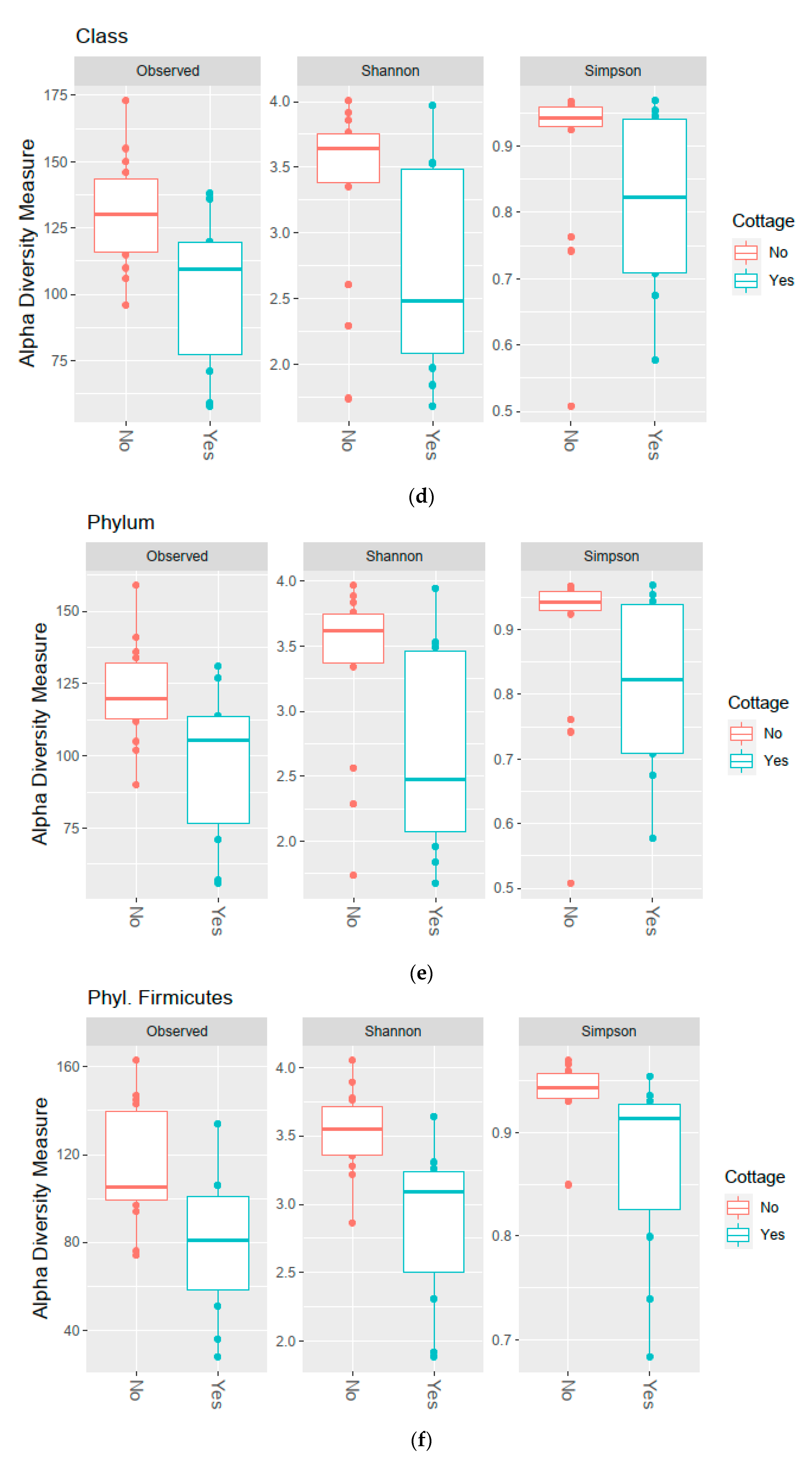
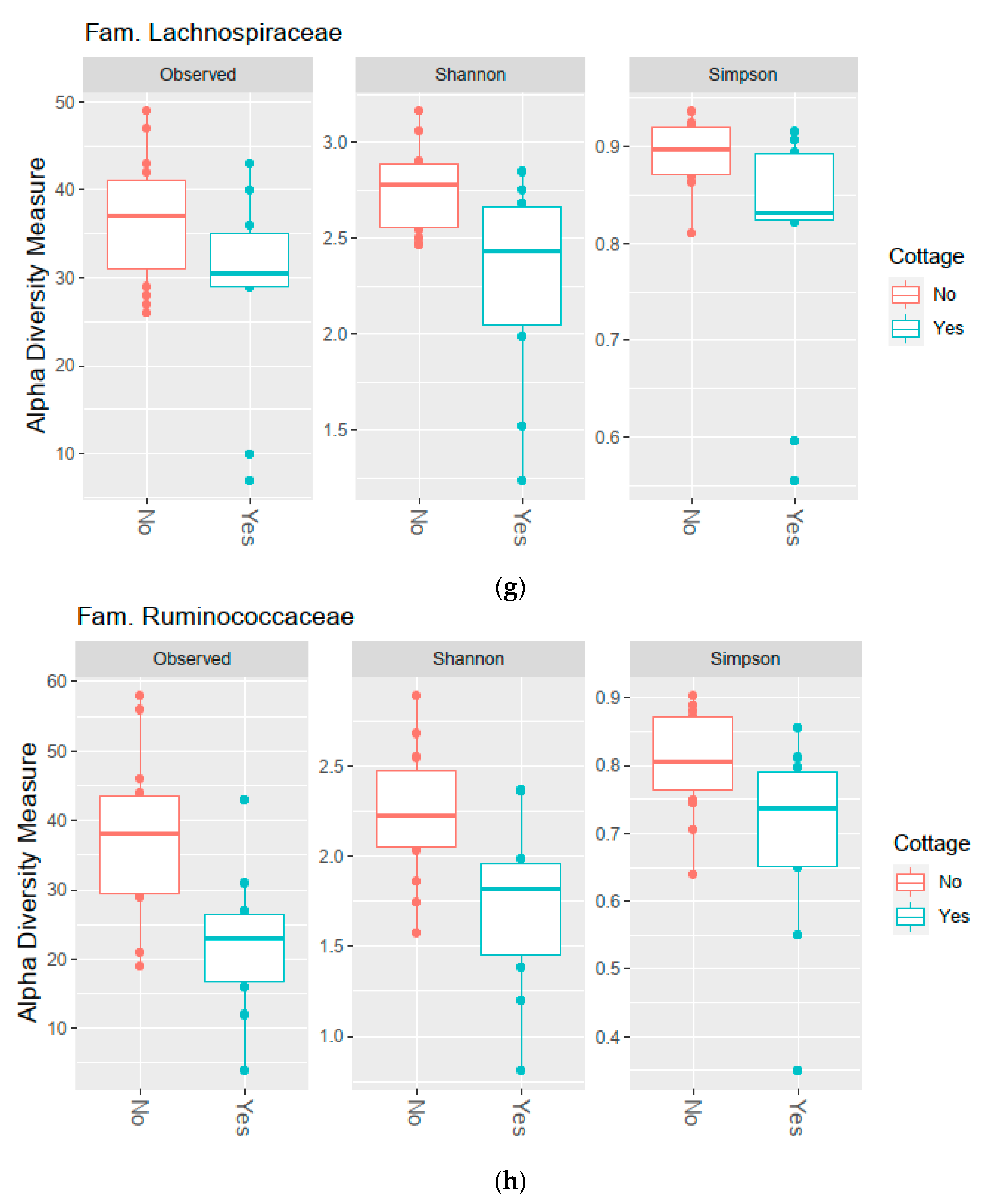
3.2. Diversity of the Bacterial Communities
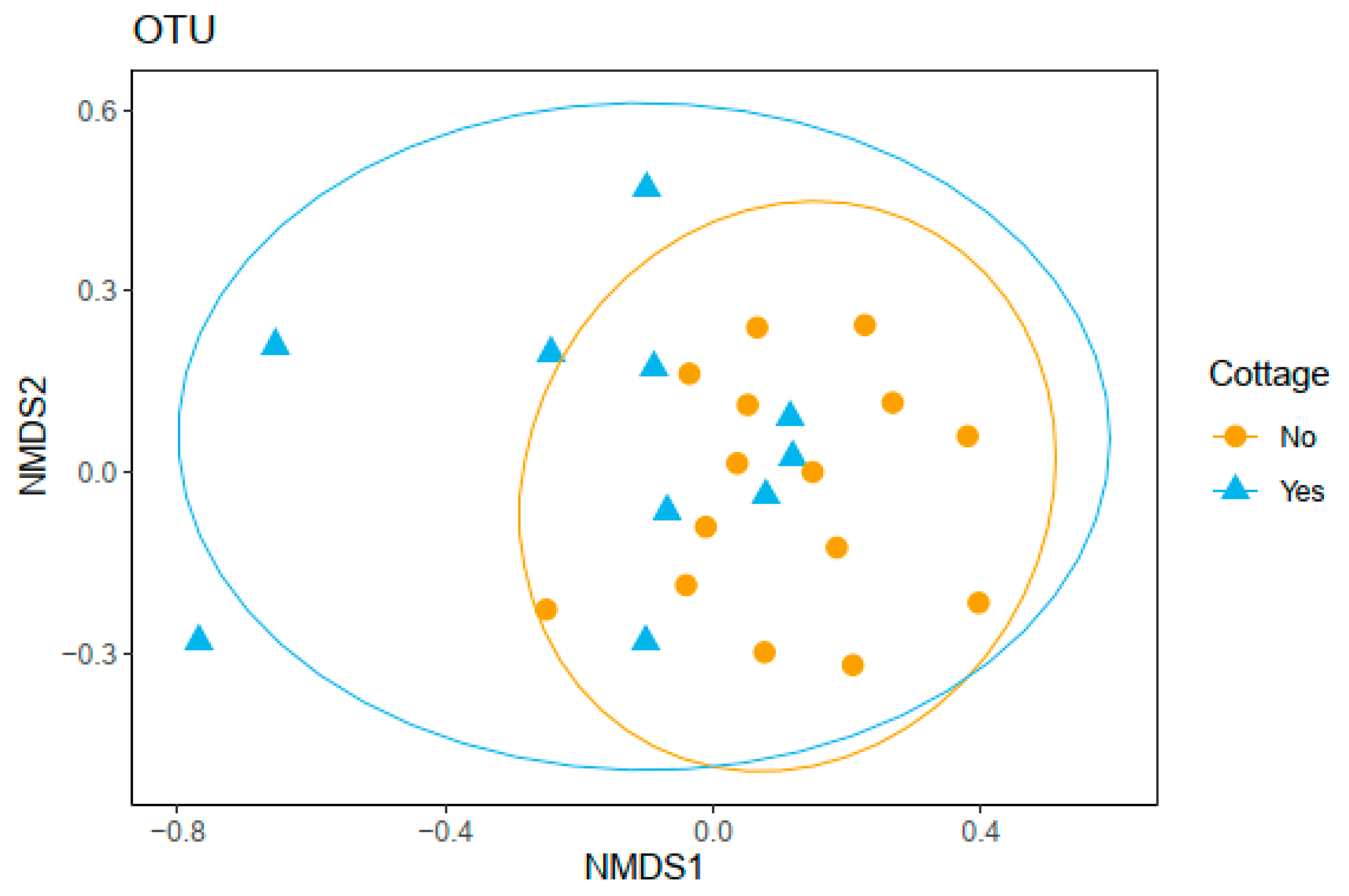
3.3. Bacterial Community Composition and Functional Profile
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lerner, A.; Jeremias, P.; Matthias, T. The World Incidence and Prevalence of Autoimmune Diseases Is Increasing. IJCD 2016, 3, 151–155. [Google Scholar] [CrossRef]
- To, T.; Stanojevic, S.; Moores, G.; Gershon, A.S.; Bateman, E.D.; Cruz, A.A.; Boulet, L.-P. Global Asthma Prevalence in Adults: Findings from the Cross-Sectional World Health Survey. BMC Public Health 2012, 12, 204. [Google Scholar] [CrossRef]
- Seto, K.C.; Güneralp, B.; Hutyra, L.R. Global Forecasts of Urban Expansion to 2030 and Direct Impacts on Biodiversity and Carbon Pools. Proc. Natl. Acad. Sci. USA 2012, 109, 16083–16088. [Google Scholar] [CrossRef]
- United Nations, Department of Economic and Social Affairs, Population Division. World Urbanization Prospects: The 2018 Revision (ST/ESA/SER.A/420); United Nations: New York, NY, USA, 2019. [Google Scholar]
- Strachan, D.P. Hay Fever, Hygiene, and Household Size. BMJ 1989, 299, 1259–1260. [Google Scholar] [CrossRef]
- Stiemsma, L.T.; Reynolds, L.A.; Turvey, S.E.; Finlay, B.B. The Hygiene Hypothesis: Current Perspectives and Future Therapies. Immunotargets Ther. 2015, 4, 143–157. [Google Scholar] [CrossRef]
- Hanski, I.; von Hertzen, L.; Fyhrquist, N.; Koskinen, K.; Torppa, K.; Laatikainen, T.; Karisola, P.; Auvinen, P.; Paulin, L.; Mäkelä, M.J.; et al. Environmental Biodiversity, Human Microbiota, and Allergy Are Interrelated. Proc. Natl. Acad. Sci. USA 2012, 109, 8334–8339. [Google Scholar] [CrossRef] [PubMed]
- Rook, G.A.W.; Adams, V.; Hunt, J.; Palmer, R.; Martinelli, R.; Brunet, L.R. Mycobacteria and Other Environmental Organisms as Immunomodulators for Immunoregulatory Disorders. Springer Semin. Immun. 2004, 25, 237–255. [Google Scholar] [CrossRef]
- Noverr, M.C.; Huffnagle, G.B. The “microflora Hypothesis” of Allergic Diseases. Clin. Exp. Allergy 2005, 35, 1511–1520. [Google Scholar] [CrossRef] [PubMed]
- Ege, M.J.; Mayer, M.; Schwaiger, K.; Mattes, J.; Pershagen, G.; van Hage, M.; Scheynius, A.; Bauer, J.; von Mutius, E. Environmental Bacteria and Childhood Asthma. Allergy 2012, 67, 1565–1571. [Google Scholar] [CrossRef] [PubMed]
- Valkonen, M.; Wouters, I.M.; Täubel, M.; Rintala, H.; Lenters, V.; Vasara, R.; Genuneit, J.; Braun-Fahrländer, C.; Piarroux, R.; von Mutius, E.; et al. Bacterial Exposures and Associations with Atopy and Asthma in Children. PLoS ONE 2015, 10, e0131594. [Google Scholar] [CrossRef] [PubMed]
- Stein, M.M.; Hrusch, C.L.; Gozdz, J.; Igartua, C.; Pivniouk, V.; Murray, S.E.; Ledford, J.G.; Marques dos Santos, M.; Anderson, R.L.; Metwali, N.; et al. Innate Immunity and Asthma Risk in Amish and Hutterite Farm Children. N. Engl. J. Med. 2016, 375, 411–421. [Google Scholar] [CrossRef] [PubMed]
- Kondrashova, A.; Seiskari, T.; Ilonen, J.; Knip, M.; Hyöty, H. The ‘Hygiene Hypothesis’ and the Sharp Gradient in the Incidence of Autoimmune and Allergic Diseases between Russian Karelia and Finland. APMIS 2013, 121, 478–493. [Google Scholar] [CrossRef]
- Sala, O.E.; Chapin, F.S., III; Armesto, J.J.; Berlow, E.; Bloomfield, J.; Dirzo, R.; Huber-Sanwald, E.; Huenneke, L.F.; Jackson, R.B. Global Biodiversity Scenarios for the Year 2100. Science 2000, 287, 1770–1774. [Google Scholar] [CrossRef] [PubMed]
- Parajuli, A.; Grönroos, M.; Kauppi, S.; Płociniczak, T.; Roslund, M.I.; Galitskaya, P.; Laitinen, O.H.; Hyöty, H.; Jumpponen, A.; Strömmer, R.; et al. The Abundance of Health-Associated Bacteria Is Altered in PAH Polluted Soils—Implications for Health in Urban Areas? PLoS ONE 2017, 12, e0187852. [Google Scholar] [CrossRef]
- Roslund, M.I.; Rantala, S.; Oikarinen, S.; Puhakka, R.; Hui, N.; Parajuli, A.; Laitinen, O.H.; Hyöty, H.; Rantalainen, A.-L.; Sinkkonen, A.; et al. Endocrine Disruption and Commensal Bacteria Alteration Associated with Gaseous and Soil PAH Contamination among Daycare Children. Environ. Int. 2019, 130, 104894. [Google Scholar] [CrossRef] [PubMed]
- Vari, H.K.; Roslund, M.I.; Oikarinen, S.; Nurminen, N.; Puhakka, R.; Parajuli, A.; Grönroos, M.; Siter, N.; Laitinen, O.H.; Hyöty, H.; et al. Associations between Land Cover Categories, Gaseous PAH Levels in Ambient Air and Endocrine Signaling Predicted from Gut Bacterial Metagenome of the Elderly. Chemosphere 2020, 128965. [Google Scholar] [CrossRef]
- von Hertzen, L.; Hanski, I.; Haahtela, T. Natural Immunity. EMBO Rep. 2011, 12, 1089–1093. [Google Scholar] [CrossRef] [PubMed]
- Haahtela, T.; Laatikainen, T.; Alenius, H.; Auvinen, P.; Fyhrquist, N.; Hanski, I.; von Hertzen, L.; Jousilahti, P.; Kosunen, T.U.; Markelova, O.; et al. Hunt for the Origin of Allergy—Comparing the Finnish and Russian Karelia. Clin. Exp. Allergy 2015, 45, 891–901. [Google Scholar] [CrossRef] [PubMed]
- Haahtela, T. A Biodiversity Hypothesis. Allergy 2019, 74, 1445–1456. [Google Scholar] [CrossRef]
- von Hertzen, L.; Haahtela, T. Disconnection of Man and the Soil: Reason for the Asthma and Atopy Epidemic? J. Allergy Clin. Immunol. 2006, 117, 334–344. [Google Scholar] [CrossRef] [PubMed]
- Ruokolainen, L.; von Hertzen, L.; Fyhrquist, N.; Laatikainen, T.; Lehtomäki, J.; Auvinen, P.; Karvonen, A.M.; Hyvärinen, A.; Tillmann, V.; Niemelä, O.; et al. Green Areas around Homes Reduce Atopic Sensitization in Children. Allergy 2015, 70, 195–202. [Google Scholar] [CrossRef]
- Parajuli, A.; Hui, N.; Puhakka, R.; Oikarinen, S.; Grönroos, M.; Selonen, V.A.O.; Siter, N.; Kramna, L.; Roslund, M.I.; Vari, H.K.; et al. Yard Vegetation Is Associated with Gut Microbiota Composition. Sci. Total Environ. 2020, 713, 136707. [Google Scholar] [CrossRef] [PubMed]
- Lehtimäki, J.; Karkman, A.; Laatikainen, T.; Paalanen, L.; von Hertzen, L.; Haahtela, T.; Hanski, I.; Ruokolainen, L. Patterns in the Skin Microbiota Differ in Children and Teenagers between Rural and Urban Environments. Sci. Rep. 2017, 7, 45651. [Google Scholar] [CrossRef] [PubMed]
- Parajuli, A.; Grönroos, M.; Siter, N.; Puhakka, R.; Vari, H.K.; Roslund, M.I.; Jumpponen, A.; Nurminen, N.; Laitinen, O.H.; Hyöty, H.; et al. Urbanization Reduces Transfer of Diverse Environmental Microbiota Indoors. Front. Microbiol. 2018, 9. [Google Scholar] [CrossRef]
- Macpherson, A.J.; Harris, N.L. Interactions between Commensal Intestinal Bacteria and the Immune System. Nat. Rev. Immunol. 2004, 4, 478–485. [Google Scholar] [CrossRef]
- Frank, D.N.; Amand, A.L.S.; Feldman, R.A.; Boedeker, E.C.; Harpaz, N.; Pace, N.R. Molecular-Phylogenetic Characterization of Microbial Community Imbalances in Human Inflammatory Bowel Diseases. Proc. Natl. Acad. Sci. USA 2007, 104, 13780–13785. [Google Scholar] [CrossRef]
- Round, J.L.; Mazmanian, S.K. The Gut Microbiota Shapes Intestinal Immune Responses during Health and Disease. Nat. Rev. Immunol. 2009, 9, 313–323. [Google Scholar] [CrossRef]
- Sekirov, I.; Russell, S.L.; Antunes, L.C.M.; Finlay, B.B. Gut Microbiota in Health and Disease. Physiol. Rev. 2010, 90, 859–904. [Google Scholar] [CrossRef]
- Clarke, G.; Stilling, R.M.; Kennedy, P.J.; Stanton, C.; Cryan, J.F.; Dinan, T.G. Minireview: Gut Microbiota: The Neglected Endocrine Organ. Mol. Endocrinol. 2014, 28, 1221–1238. [Google Scholar] [CrossRef] [PubMed]
- Meng, X.; Zhang, G.; Cao, H.; Yu, D.; Fang, X.; de Vos, W.M.; Wu, H. Gut Dysbacteriosis and Intestinal Disease: Mechanism and Treatment. J. Appl. Microbiol. 2020, 129, 787–805. [Google Scholar] [CrossRef]
- Ott, S.J.; Musfeldt, M.; Wenderoth, D.F.; Hampe, J.; Brant, O.; Fölsch, U.R.; Timmis, K.N.; Schreiber, S. Reduction in Diversity of the Colonic Mucosa Associated Bacterial Microflora in Patients with Active Inflammatory Bowel Disease. Gut 2004, 53, 685–693. [Google Scholar] [CrossRef] [PubMed]
- Scher, J.U.; Ubeda, C.; Artacho, A.; Attur, M.; Isaac, S.; Reddy, S.M.; Marmon, S.; Neimann, A.; Brusca, S.; Patel, T.; et al. Decreased Bacterial Diversity Characterizes the Altered Gut Microbiota in Patients with Psoriatic Arthritis, Resembling Dysbiosis in Inflammatory Bowel Disease. Arthritis Rheumatol. 2015, 67, 128–139. [Google Scholar] [CrossRef] [PubMed]
- de Paiva, C.S.; Jones, D.B.; Stern, M.E.; Bian, F.; Moore, Q.L.; Corbiere, S.; Streckfus, C.F.; Hutchinson, D.S.; Ajami, N.J.; Petrosino, J.F.; et al. Altered Mucosal Microbiome Diversity and Disease Severity in Sjögren Syndrome. Sci. Rep. 2016, 6, 23561. [Google Scholar] [CrossRef]
- Manichanh, C.; Rigottier-Gois, L.; Bonnaud, E.; Gloux, K.; Pelletier, E.; Frangeul, L.; Nalin, R.; Jarrin, C.; Chardon, P.; Marteau, P.; et al. Reduced Diversity of Faecal Microbiota in Crohn’s Disease Revealed by a Metagenomic Approach. Gut 2006, 55, 205–211. [Google Scholar] [CrossRef]
- Rajilić–Stojanović, M.; Biagi, E.; Heilig, H.G.H.J.; Kajander, K.; Kekkonen, R.A.; Tims, S.; de Vos, W.M. Global and Deep Molecular Analysis of Microbiota Signatures in Fecal Samples From Patients With Irritable Bowel Syndrome. Gastroenterology 2011, 141, 1792–1801. [Google Scholar] [CrossRef] [PubMed]
- Louis, P.; Flint, H.J. Diversity, Metabolism and Microbial Ecology of Butyrate-Producing Bacteria from the Human Large Intestine. Fems. Microbiol. Lett. 2009, 294, 1–8. [Google Scholar] [CrossRef]
- Plöger, S.; Stumpff, F.; Penner, G.; Schulzke, J.; Gäbel, G.; Martens, H.; Shen, Z.; Günzel, D.; Aschenbach, J. Microbial Butyrate and Its Role for Barrier Function in the Gastrointestinal Tract. Ann. N. Y. Acad. Sci. 2012, 1258, 52–59. [Google Scholar] [CrossRef]
- Geirnaert, A.; Calatayud, M.; Grootaert, C.; Laukens, D.; Devriese, S.; Smagghe, G.; De Vos, M.; Boon, N.; Van de Wiele, T. Butyrate-Producing Bacteria Supplemented in Vitro to Crohn’s Disease Patient Microbiota Increased Butyrate Production and Enhanced Intestinal Epithelial Barrier Integrity. Sci. Rep. 2017, 7, 11450. [Google Scholar] [CrossRef]
- Greenhalgh, K.; Meyer, K.M.; Aagaard, K.M.; Wilmes, P. The Human Gut Microbiome in Health: Establishment and Resilience of Microbiota over a Lifetime. Environ. Microbiol. 2016, 18, 2103–2116. [Google Scholar] [CrossRef]
- Rinninella, E.; Raoul, P.; Cintoni, M.; Franceschi, F.; Miggiano, G.A.D.; Gasbarrini, A.; Mele, M.C. What Is the Healthy Gut Microbiota Composition? A Changing Ecosystem across Age, Environment, Diet, and Diseases. Microorganisms 2019, 7, 14. [Google Scholar] [CrossRef] [PubMed]
- O’Toole, P.W.; Claesson, M.J. Gut Microbiota: Changes throughout the Lifespan from Infancy to Elderly. Int. Dairy J. 2010, 20, 281–291. [Google Scholar] [CrossRef]
- Nagpal, R.; Mainali, R.; Ahmadi, S.; Wang, S.; Singh, R.; Kavanagh, K.; Kitzman, D.W.; Kushugulova, A.; Marotta, F.; Yadav, H. Gut Microbiome and Aging: Physiological and Mechanistic Insights. Nutr. Healthy Aging 2018, 4, 267–285. [Google Scholar] [CrossRef] [PubMed]
- Nurminen, N.; Lin, J.; Grönroos, M.; Puhakka, R.; Kramna, L.; Vari, H.K.; Viskari, H.; Oikarinen, S.; Roslund, M.; Parajuli, A.; et al. Nature-Derived Microbiota Exposure as a Novel Immunomodulatory Approach. Future Microbiol. 2018, 13, 737–744. [Google Scholar] [CrossRef] [PubMed]
- Grönroos, M.; Parajuli, A.; Laitinen, O.H.; Roslund, M.I.; Vari, H.K.; Hyöty, H.; Puhakka, R.; Sinkkonen, A. Short-Term Direct Contact with Soil and Plant Materials Leads to an Immediate Increase in Diversity of Skin Microbiota. MicrobiologyOpen 2019, 8, e00645. [Google Scholar] [CrossRef]
- Hui, N.; Grönroos, M.; Roslund, M.; Parajuli, A.; Vari, H.K.; Soininen, L.; Laitinen, O.H.; Sinkkonen, A. Diverse Environmental Microbiota as a Tool to Augment Biodiversity in Urban Landscaping Materials. Front. Microbiol. 2019, 10, 536. [Google Scholar] [CrossRef] [PubMed]
- Roslund, M.I.; Puhakka, R.; Grönroos, M.; Nurminen, N.; Oikarinen, S.; Gazali, A.M.; Cinek, O.; Kramná, L.; Siter, N.; Vari, H.K.; et al. Biodiversity Intervention Enhances Immune Regulation and Health-Associated Commensal Microbiota among Daycare Children. Sci. Adv. 2020, 6, eaba2578. [Google Scholar] [CrossRef]
- Lehtimäki, J.; Sinkko, H.; Hielm-Björkman, A.; Salmela, E.; Tiira, K.; Laatikainen, T.; Mäkeläinen, S.; Kaukonen, M.; Uusitalo, L.; Hanski, I.; et al. Skin Microbiota and Allergic Symptoms Associate with Exposure to Environmental Microbes. Proc. Natl. Acad. Sci. USA 2018, 115, 4897–4902. [Google Scholar] [CrossRef] [PubMed]
- Sievänen, T.; Neuvonen, M. Luonnon virkistyskäytön kysyntä 2010 ja kysynnän muutos. In Luonnon Virkistyskäyttö 2010; Sievänen, T., Neuvonen, M., Eds.; Working Papers of the Finnish Forest Research Institute: Vantaa, Finland, 2011; Volume 212, pp. 37–79. [Google Scholar]
- Pitkänen, K.; Lehtimäki, J.; Puhakka, R. How Do Rural Second Homes Affect Human Health and Well-Being? Review of Potential Impacts. Int. J. Environ. Res. Public Health 2020, 17, 6748. [Google Scholar] [CrossRef]
- Adamiak, C.; Vepsäläinen, M.; Strandell, A.; Hiltunen, M.; Pitkänen, K.; Hall, M.; Rinne, J.; Hannonen, O.; Paloniemi, R.; Åkerlund, U. Second Home Tourism in Finland—Perceptions of Citizens and Municipalities on the State and Development of Second Home Tourism; Suomen Ympäristökeskus: Helsinki, Finland, 2015. [Google Scholar]
- Statistics Finland. Free-Time Residences 2019. Helsinki. 2020. Available online: http://www.stat.fi/til/rakke/2019/rakke_2019_2020-05-27_kat_001_en.html (accessed on 2 December 2020).
- Pitkänen, K.; Hannonen, O.; Toso, S.; Gallent, N.; Hamiduddin, I.; Halseth, G.; Hall, M.C.; Müller, D.K.; Treivish, A.; Nefedova, T. Second Homes during Corona—Safe or Unsafe Haven and for Whom? Reflections from Researchers around the World. Matkailututkimus 2020, 16, 20–39. [Google Scholar] [CrossRef]
- Sievänen, T.; Pouta, E.; Neuvonen, M. Recreational Home Users—Potential Clients for Countryside Tourism? Scand. J. Hosp. Tour. 2007, 7, 223–242. [Google Scholar] [CrossRef]
- FCG Finnish Consulting Group Oy. Mökkibarometri 2016; Saaristoasiain Neuvottelukunta, Maa-ja Metsätalousministeriö: Helsinki, Finland, 2016; Available online: https://valtioneuvosto.fi/documents/1410837/1880296/Mokkibarometri+2016/7b69ab48-5859-4b55-8dc2-5514cdfa6000 (accessed on 22 December 2020).
- Hui, N.; Parajuli, A.; Puhakka, R.; Grönroos, M.; Roslund, M.I.; Vari, H.K.; Selonen, V.A.O.; Yan, G.; Siter, N.; Nurminen, N.; et al. Temporal Variation in Indoor Transfer of Dirt-Associated Environmental Bacteria in Agricultural and Urban Areas. Environ. Int. 2019, 132, 105069. [Google Scholar] [CrossRef]
- Kondrashova, A.; Reunanen, A.; Romanov, A.; Karvonen, A.; Viskari, H.; Vesikari, T.; Ilonen, J.; Knip, M.; Hyöty, H. A Six-fold Gradient in the Incidence of Type 1 Diabetes at the Eastern Border of Finland. Ann. Med. 2005, 37, 67–72. [Google Scholar] [CrossRef] [PubMed]
- Fogelholm, M.; Valve, R.; Absetz, P.; Heinonen, H.; Uutela, A.; Patja, K.; Karisto, A.; Konttinen, R.; Mäkelä, T.; Nissinen, A.; et al. Rural—Urban Differences in Health and Health Behaviour: A Baseline Description of a Community Health-Promotion Programme for the Elderly. Scand. J. Public Health 2006, 34, 632–640. [Google Scholar] [CrossRef] [PubMed]
- Schloss, P.D.; Westcott, S.L.; Ryabin, T.; Hall, J.R.; Hartmann, M.; Hollister, E.B.; Lesniewski, R.A.; Oakley, B.B.; Parks, D.H.; Robinson, C.J.; et al. Introducing Mothur: Open-Source, Platform-Independent, Community-Supported Software for Describing and Comparing Microbial Communities. Appl. Environ. Microbiol. 2009, 75, 7537–7541. [Google Scholar] [CrossRef] [PubMed]
- Schloss, P.D.; Westcott, S.L. Assessing and Improving Methods Used in Operational Taxonomic Unit-Based Approaches for 16S RRNA Gene Sequence Analysis. Appl. Environ. Microbiol. 2011, 77, 3219–3226. [Google Scholar] [CrossRef]
- Kozich, J.J.; Westcott, S.L.; Baxter, N.T.; Highlander, S.K.; Schloss, P.D. Development of a Dual-Index Sequencing Strategy and Curation Pipeline for Analyzing Amplicon Sequence Data on the MiSeq Illumina Sequencing Platform. Appl. Environ. Microbiol. 2013, 79, 5112–5120. [Google Scholar] [CrossRef]
- Pruesse, E.; Quast, C.; Knittel, K.; Fuchs, B.M.; Ludwig, W.; Peplies, J.; Glöckner, F.O. SILVA: A Comprehensive Online Resource for Quality Checked and Aligned Ribosomal RNA Sequence Data Compatible with ARB. Nucleic Acids Res. 2007, 35, 7188–7196. [Google Scholar] [CrossRef]
- Oliver, A.K.; Brown, S.P.; Callaham, M.A.; Jumpponen, A. Polymerase Matters: Non-Proofreading Enzymes Inflate Fungal Community Richness Estimates by up to 15%. Fungal Ecol. 2015, 15, 86–89. [Google Scholar] [CrossRef]
- DeSantis, T.Z.; Hugenholtz, P.; Larsen, N.; Rojas, M.; Brodie, E.L.; Keller, K.; Huber, T.; Dalevi, D.; Hu, P.; Andersen, G.L. Greengenes, a Chimera-Checked 16S RRNA Gene Database and Workbench Compatible with ARB. Appl. Environ. Microbiol. 2006, 72, 5069–5072. [Google Scholar] [CrossRef]
- Langille, M.G.I.; Zaneveld, J.; Caporaso, J.G.; McDonald, D.; Knights, D.; Reyes, J.A.; Clemente, J.C.; Burkepile, D.E.; Vega Thurber, R.L.; Knight, R.; et al. Predictive Functional Profiling of Microbial Communities Using 16S RRNA Marker Gene Sequences. Nat. Biotechnol. 2013, 31, 814–821. [Google Scholar] [CrossRef] [PubMed]
- Benjamini, Y.; Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J. R. Stat. Soc. Ser. B 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Donaldson, G.P.; Lee, S.M.; Mazmanian, S.K. Gut Biogeography of the Bacterial Microbiota. Nat. Rev. Microbiol. 2016, 14, 20–32. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2020; Available online: https://www.R-project.org/ (accessed on 22 December 2020).
- McMurdie, P.J.; Holmes, S. Phyloseq: An R Package for Reproducible Interactive Analysis and Graphics of Microbiome Census Data. PLoS ONE 2013, 8, e61217. [Google Scholar] [CrossRef]
- Wickham, H. Ggplot2: Elegant Graphics for Data Analysis; Springer: New York, NY, USA, 2016; ISBN 978-0-387-98141-3. [Google Scholar]
- Oksanen, J.; Blanchet, F.G.; Friendly, M.; Kindt, R.; Legendre, P.; McGlinn, D.; Minchin, P.R.; O’Hara, R.B.; Simpson, G.L.; Solymos, P.; et al. Vegan: Community Ecology Package. R Package Version 2.5-6. 2019. Available online: https://CRAN.R-project.org/package=vegan (accessed on 22 December 2020).
- Bartosch, S.; Fite, A.; Macfarlane, G.T.; McMurdo, M.E.T. Characterization of Bacterial Communities in Feces from Healthy Elderly Volunteers and Hospitalized Elderly Patients by Using Real-Time PCR and Effects of Antibiotic Treatment on the Fecal Microbiota. Appl. Environ. Microbiol. 2004, 70, 3575–3581. [Google Scholar] [CrossRef]
- Woodmansey, E.J.; McMurdo, M.E.T.; Macfarlane, G.T.; Macfarlane, S. Comparison of Compositions and Metabolic Activities of Fecal Microbiotas in Young Adults and in Antibiotic-Treated and Non-Antibiotic-Treated Elderly Subjects. Appl. Environ. Microbiol. 2004, 70, 6113–6122. [Google Scholar] [CrossRef]
- Zwielehner, J.; Liszt, K.; Handschur, M.; Lassl, C.; Lapin, A.; Haslberger, A.G. Combined PCR-DGGE Fingerprinting and Quantitative-PCR Indicates Shifts in Fecal Population Sizes and Diversity of Bacteroides, Bifidobacteria and Clostridium Cluster IV in Institutionalized Elderly. Exp. Gerontol. 2009, 44, 440–446. [Google Scholar] [CrossRef] [PubMed]
- Mäkivuokko, H.; Tiihonen, K.; Tynkkynen, S.; Paulin, L.; Rautonen, N. The Effect of Age and Non-Steroidal Anti-Inflammatory Drugs on Human Intestinal Microbiota Composition. Br. J. Nutr. 2010, 103, 227–234. [Google Scholar] [CrossRef]
- Kostic, A.D.; Gevers, D.; Pedamallu, C.S.; Michaud, M.; Duke, F.; Earl, A.M.; Ojesina, A.I.; Jung, J.; Bass, A.J.; Tabernero, J.; et al. Genomic Analysis Identifies Association of Fusobacterium with Colorectal Carcinoma. Genome Res. 2012, 22, 292–298. [Google Scholar] [CrossRef] [PubMed]
- Larsen, N.; Vogensen, F.K.; van den Berg, F.W.J.; Nielsen, D.S.; Andreasen, A.S.; Pedersen, B.K.; Al-Soud, W.A.; Sørensen, S.J.; Hansen, L.H.; Jakobsen, M. Gut Microbiota in Human Adults with Type 2 Diabetes Differs from Non-Diabetic Adults. PLoS ONE 2010, 5, e9085. [Google Scholar] [CrossRef]
- Krogius-Kurikka, L.; Lyra, A.; Malinen, E.; Aarnikunnas, J.; Tuimala, J.; Paulin, L.; Mäkivuokko, H.; Kajander, K.; Palva, A. Microbial Community Analysis Reveals High Level Phylogenetic Alterations in the Overall Gastrointestinal Microbiota of Diarrhoea-Predominant Irritable Bowel Syndrome Sufferers. BMC Gastroenterol. 2009, 9, 95. [Google Scholar] [CrossRef] [PubMed]
- Matsuoka, K.; Kanai, T. The Gut Microbiota and Inflammatory Bowel Disease. Semin. Immunopathol. 2015, 37, 47–55. [Google Scholar] [CrossRef]
- Zeng, X.; Gao, X.; Peng, Y.; Wu, Q.; Zhu, J.; Tan, C.; Xia, G.; You, C.; Xu, R.; Pan, S.; et al. Higher Risk of Stroke Is Correlated With Increased Opportunistic Pathogen Load and Reduced Levels of Butyrate-Producing Bacteria in the Gut. Front. Cell. Infect. Microbiol. 2019, 9. [Google Scholar] [CrossRef] [PubMed]
- Chiu, Y.-H.; Macmillan, J.B.; Chen, Z.J. RNA Polymerase III Detects Cytosolic DNA and Induces Type I Interferons through the RIG-I Pathway. Cell 2009, 138, 576–591. [Google Scholar] [CrossRef]
- Martínez, I.; Oliveros, J.C.; Cuesta, I.; de la Barrera, J.; Ausina, V.; Casals, C.; de Lorenzo, A.; García, E.; García-Fojeda, B.; Garmendia, J.; et al. Apoptosis, Toll-like, RIG-I-like and NOD-like Receptors Are Pathways Jointly Induced by Diverse Respiratory Bacterial and Viral Pathogens. Front. Microbiol. 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Laitinen, O.H.; Svedin, E.; Kapell, S.; Nurminen, A.; Hytönen, V.P.; Flodström-Tullberg, M. Enteroviral Proteases: Structure, Host Interactions and Pathogenicity. Rev. Med. Virol. 2016, 26, 251–267. [Google Scholar] [CrossRef] [PubMed]
- Oshiumi, H. Recent Advances and Contradictions in the Study of the Individual Roles of Ubiquitin Ligases That Regulate RIG-I-Like Receptor-Mediated Antiviral Innate Immune Responses. Front. Immunol. 2020, 11, 1296. [Google Scholar] [CrossRef] [PubMed]
- Ren, Z.; Ding, T.; Zuo, Z.; Xu, Z.; Deng, J.; Wei, Z. Regulation of MAVS Expression and Signaling Function in the Antiviral Innate Immune Response. Front. Immunol. 2020, 11, 1030. [Google Scholar] [CrossRef]
- Sharma, S.; Fitzgerald, K.A.; Cancro, M.P.; Marshak-Rothstein, A. Nucleic Acid-Sensing Receptors: Rheostats of Autoimmunity and Autoinflammation. J. Immunol. 2015, 195, 3507–3512. [Google Scholar] [CrossRef]
- Baechler, E.C.; Batliwalla, F.M.; Karypis, G.; Gaffney, P.M.; Ortmann, W.A.; Espe, K.J.; Shark, K.B.; Grande, W.J.; Hughes, K.M.; Kapur, V.; et al. Interferon-Inducible Gene Expression Signature in Peripheral Blood Cells of Patients with Severe Lupus. Proc. Natl. Acad. Sci. USA 2003, 100, 2610–2615. [Google Scholar] [CrossRef]
- Bennett, L.; Palucka, A.K.; Arce, E.; Cantrell, V.; Borvak, J.; Banchereau, J.; Pascual, V. Interferon and Granulopoiesis Signatures in Systemic Lupus Erythematosus Blood. J. Exp. Med. 2003, 197, 711–723. [Google Scholar] [CrossRef]
- Lübbers, J.; Brink, M.; van de Stadt, L.A.; Vosslamber, S.; Wesseling, J.G.; van Schaardenburg, D.; Rantapää-Dahlqvist, S.; Verweij, C.L. The Type I IFN Signature as a Biomarker of Preclinical Rheumatoid Arthritis. Ann. Rheum. Dis. 2013, 72, 776–780. [Google Scholar] [CrossRef] [PubMed]
- Postal, M.; Vivaldo, J.F.; Fernandez-Ruiz, R.; Paredes, J.L.; Appenzeller, S.; Niewold, T.B. Type I Interferon in the Pathogenesis of Systemic Lupus Erythematosus. Curr. Opin. Immunol. 2020, 67, 87–94. [Google Scholar] [CrossRef] [PubMed]
- Tossberg, J.T.; Heinrich, R.M.; Farley, V.M.; Crooke, P.S.; Aune, T.M. Adenosine-to-Inosine RNA Editing of Alu Double-Stranded (Ds)RNAs Is Markedly Decreased in Multiple Sclerosis and Unedited Alu DsRNAs Are Potent Activators of Proinflammatory Transcriptional Responses. J. Immunol. 2020, 205, 2606–2617. [Google Scholar] [CrossRef]
- Claesson, M.J.; Jeffery, I.B.; Conde, S.; Power, S.E.; O’Connor, E.M.; Cusack, S.; Harris, H.M.B.; Coakley, M.; Lakshminarayanan, B.; O’Sullivan, O.; et al. Gut Microbiota Composition Correlates with Diet and Health in the Elderly. Nature 2012, 488, 178–184. [Google Scholar] [CrossRef] [PubMed]
- Mueller, S.; Saunier, K.; Hanisch, C.; Norin, E.; Alm, L.; Midtvedt, T.; Cresci, A.; Silvi, S.; Orpianesi, C.; Verdenelli, M.C.; et al. Differences in Fecal Microbiota in Different European Study Populations in Relation to Age, Gender, and Country: A Cross-Sectional Study. Appl. Environ. Microbiol. 2006, 72, 1027–1033. [Google Scholar] [CrossRef]
- Hertzen, L.V.; Laatikainen, T.; Pitkänen, T.; Vlasoff, T.; Mäkelä, M.J.; Vartiainen, E.; Haahtela, T. Microbial Content of Drinking Water in Finnish and Russian Karelia—Implications for Atopy Prevalence. Allergy 2007, 62, 288–292. [Google Scholar] [CrossRef]
- Hesselmar, B.; Hicke-Roberts, A.; Wennergren, G. Allergy in Children in Hand Versus Machine Dishwashing. Pediatrics 2015, 135, e590–e597. [Google Scholar] [CrossRef]
- Turkki, P. Lähiruokaa Mökille: Etelä-Savon Vapaa-Ajan Asukkaiden Ruokaostoskäyttäytyminen ja Lähiruoan Saatavuus; Tutkimuksia ja Raportteja—Research Reports 106; Mikkelin Ammattikorkeakoulu: Mikkeli, Finland, 2016; ISBN 978-951-588-553-1. [Google Scholar]
- Martínez, I.; Kim, J.; Duffy, P.R.; Schlegel, V.L.; Walter, J. Resistant Starches Types 2 and 4 Have Differential Effects on the Composition of the Fecal Microbiota in Human Subjects. PLoS ONE 2010, 5, e15046. [Google Scholar] [CrossRef]
- David, L.A.; Maurice, C.F.; Carmody, R.N.; Gootenberg, D.B.; Button, J.E.; Wolfe, B.E.; Ling, A.V.; Devlin, A.S.; Varma, Y.; Fischbach, M.A.; et al. Diet Rapidly and Reproducibly Alters the Human Gut Microbiome. Nature 2014, 505, 559–563. [Google Scholar] [CrossRef] [PubMed]
- Venkataraman, A.; Sieber, J.R.; Schmidt, A.W.; Waldron, C.; Theis, K.R.; Schmidt, T.M. Variable Responses of Human Microbiomes to Dietary Supplementation with Resistant Starch. Microbiome 2016, 4, 33. [Google Scholar] [CrossRef]
- Lin, D.; Peters, B.A.; Friedlander, C.; Freiman, H.J.; Goedert, J.J.; Sinha, R.; Miller, G.; Bernstein, M.A.; Hayes, R.B.; Ahn, J. Association of Dietary Fibre Intake and Gut Microbiota in Adults. Br. J. Nutr. 2018, 120, 1014–1022. [Google Scholar] [CrossRef] [PubMed]
- Bjørkhaug, S.T.; Aanes, H.; Neupane, S.P.; Bramness, J.G.; Malvik, S.; Henriksen, C.; Skar, V.; Medhus, A.W.; Valeur, J. Characterization of Gut Microbiota Composition and Functions in Patients with Chronic Alcohol Overconsumption. Gut Microbes 2019, 10, 663–675. [Google Scholar] [CrossRef] [PubMed]
- Wan, Y.; Wang, F.; Yuan, J.; Li, J.; Jiang, D.; Zhang, J.; Li, H.; Wang, R.; Tang, J.; Huang, T.; et al. Effects of Dietary Fat on Gut Microbiota and Faecal Metabolites, and Their Relationship with Cardiometabolic Risk Factors: A 6-Month Randomised Controlled-Feeding Trial. Gut 2019, 68, 1417–1429. [Google Scholar] [CrossRef] [PubMed]
- Flint, H.J.; Duncan, S.H.; Scott, K.P.; Louis, P. Links between Diet, Gut Microbiota Composition and Gut Metabolism. Proc. Nutr. Soc. 2015, 74, 13–22. [Google Scholar] [CrossRef]
| Years Used | Visiting Days per Year | Time Spent Outdoors vs. Indoors |
|---|---|---|
| 46 | 40 | Outdoors |
| 23 | 10 | Outdoors |
| 30 | 35 | Outdoors |
| 27 | 90 | Equally |
| - | 100 | Outdoors |
| 44 | 30 | Outdoors |
| 29 | 150 | Equally |
| 53 | 30 | Outdoors |
| 61 | - | Outdoors |
| 9 | 120 | Equally |
| Taxa | Role and Geography | Note |
|---|---|---|
| Bacteroidetes | Dominant phylum in the gut | |
| Firmicutes | Dominant phylum in the gut | |
| Actinobacteria | Dominant phylum in the gut | |
| Proteobacteria | Dominant phylum in the gut | |
| Verrucomicrobia | Dominant phylum in the gut | Not analyzed due to frequent zero-valued observations |
| Ruminococcaceae | Predominant family in the inter-fold regions of the lumen | |
| Lachnospiraceae | Predominant family in the inter-fold regions of the lumen | |
| Bacteroidaceae | Predominant family in the digesta | |
| Prevotellaceae | Predominant family in the digesta | Not analyzed due to frequent zero-valued observations |
| Rikenellaceae | Predominant family in the digesta | Not analyzed due to frequent zero-valued observations |
| Diversity Measure | Level | p-Value | Benjamini–Hochberg Critical Value [66] | Significant with FDR 0.1 | Test |
|---|---|---|---|---|---|
| Shannon | Phyl. Firmicutes | 0.001 | 0.008 | Yes | U test |
| Fam. Ruminococcaceae | 0.014 | 0.017 | Yes | t-test | |
| Fam. Lachnospiraceae | 0.017 | 0.025 | Yes | U test | |
| Phylum | 0.031 | 0.033 | Yes | U test | |
| Class | 0.031 | 0.033 | Yes | U test | |
| Order | 0.031 | 0.033 | Yes | U test | |
| OTU | 0.035 | 0.042 | Yes | U test | |
| Family | 0.035 | 0.042 | Yes | U test | |
| Phyl. Proteobacteria | 0.154 | 0.050 | No | t-test | |
| Phyl. Bacteroidetes | 0.229 | 0.058 | No | t-test | |
| Fam. Bacteroidaceae | 0.542 | 0.067 | No | U test | |
| Phyl. Actinobacteria | 0.868 | 0.075 | No | t-test | |
| Simpson | Phyl. Firmicutes | 0.004 | 0.008 | Yes | U test |
| Fam. Ruminococcaceae | 0.027 | 0.017 | No | U test | |
| Fam. Lachnospiraceae | 0.031 | 0.025 | No | U test | |
| OTU | 0.120 | 0.033 | No | U test | |
| Phylum | 0.120 | 0.033 | No | U test | |
| Class | 0.120 | 0.033 | No | U test | |
| Order | 0.120 | 0.033 | No | U test | |
| Family | 0.120 | 0.033 | No | U test | |
| Phyl. Bacteroidetes | 0.267 | 0.042 | No | U test | |
| Fam. Bacteroidaceae | 0.579 | 0.050 | No | U test | |
| Phyl. Proteobacteria | 0.680 | 0.058 | No | t-test | |
| Phyl. Actinobacteria | 0.868 | 0.067 | No | U test | |
| Observed richness | Fam. Ruminococcaceae | 0.003 | 0.008 | Yes | t-test |
| OTU | 0.008 | 0.017 | Yes | t-test | |
| Phyl. Firmicutes | 0.009 | 0.025 | Yes | t-test | |
| Family | 0.018 | 0.033 | Yes | t-test | |
| Class | 0.019 | 0.042 | Yes | t-test | |
| Order | 0.019 | 0.05 | Yes | t-test | |
| Phylum | 0.024 | 0.058 | Yes | t-test | |
| Phyl. Proteobacteria | 0.053 | 0.067 | No | t-test | |
| Fam. Lachnospiraceae | 0.126 | 0.075 | No | U test | |
| Phyl. Actinobacteria | 0.548 | 0.083 | No | t-test | |
| Phyl. Bacteroidetes | 0.760 | 0.092 | No | U test | |
| Fam. Bacteroidaceae | 0.889 | 0.1 | No | t-test |
| Level | p-Value | F | R2 |
|---|---|---|---|
| Phyl. Proteobacteria | 0.068 | 1.441 | 0.059 |
| Phyl. Actinobacteria | 0.080 | 1.661 | 0.067 |
| OTU | 0.139 | 1.327 | 0.055 |
| Fam. Bacteroidaceae | 0.172 | 1.376 | 0.056 |
| Phyl. Bacteroidetes | 0.254 | 1.205 | 0.050 |
| Phylum | 0.300 | 1.117 | 0.046 |
| Order | 0.318 | 1.088 | 0.045 |
| Class | 0.324 | 1.089 | 0.045 |
| Fam. Lachnospiraceae | 0.328 | 1.104 | 0.046 |
| Family | 0.343 | 1.071 | 0.044 |
| Fam. Ruminococcaceae | 0.408 | 0.990 | 0.041 |
| Phyl. Firmicutes | 0.427 | 1.024 | 0.043 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Saarenpää, M.; Roslund, M.I.; Puhakka, R.; Grönroos, M.; Parajuli, A.; Hui, N.; Nurminen, N.; Laitinen, O.H.; Hyöty, H.; Cinek, O.; et al. Do Rural Second Homes Shape Commensal Microbiota of Urban Dwellers? A Pilot Study among Urban Elderly in Finland. Int. J. Environ. Res. Public Health 2021, 18, 3742. https://doi.org/10.3390/ijerph18073742
Saarenpää M, Roslund MI, Puhakka R, Grönroos M, Parajuli A, Hui N, Nurminen N, Laitinen OH, Hyöty H, Cinek O, et al. Do Rural Second Homes Shape Commensal Microbiota of Urban Dwellers? A Pilot Study among Urban Elderly in Finland. International Journal of Environmental Research and Public Health. 2021; 18(7):3742. https://doi.org/10.3390/ijerph18073742
Chicago/Turabian StyleSaarenpää, Mika, Marja I. Roslund, Riikka Puhakka, Mira Grönroos, Anirudra Parajuli, Nan Hui, Noora Nurminen, Olli H. Laitinen, Heikki Hyöty, Ondrej Cinek, and et al. 2021. "Do Rural Second Homes Shape Commensal Microbiota of Urban Dwellers? A Pilot Study among Urban Elderly in Finland" International Journal of Environmental Research and Public Health 18, no. 7: 3742. https://doi.org/10.3390/ijerph18073742
APA StyleSaarenpää, M., Roslund, M. I., Puhakka, R., Grönroos, M., Parajuli, A., Hui, N., Nurminen, N., Laitinen, O. H., Hyöty, H., Cinek, O., Sinkkonen, A., & the ADELE Research Group. (2021). Do Rural Second Homes Shape Commensal Microbiota of Urban Dwellers? A Pilot Study among Urban Elderly in Finland. International Journal of Environmental Research and Public Health, 18(7), 3742. https://doi.org/10.3390/ijerph18073742









