Discrepancies in Infectivity of Flavivirus and SARS-CoV-2 Clinical Samples: An Improved Assay for Infectious Virus Shedding and Viremia Assessment
Abstract
:1. Introduction
2. Materials and Methods
2.1. Cell Lines
2.2. Virus
2.3. Patient Samples
2.4. Virus Overlay Medium
2.5. Infectious Virus Particle Titration from Laboratory-Established Strains and Clinical Samples
2.6. Data Analysis
3. Results
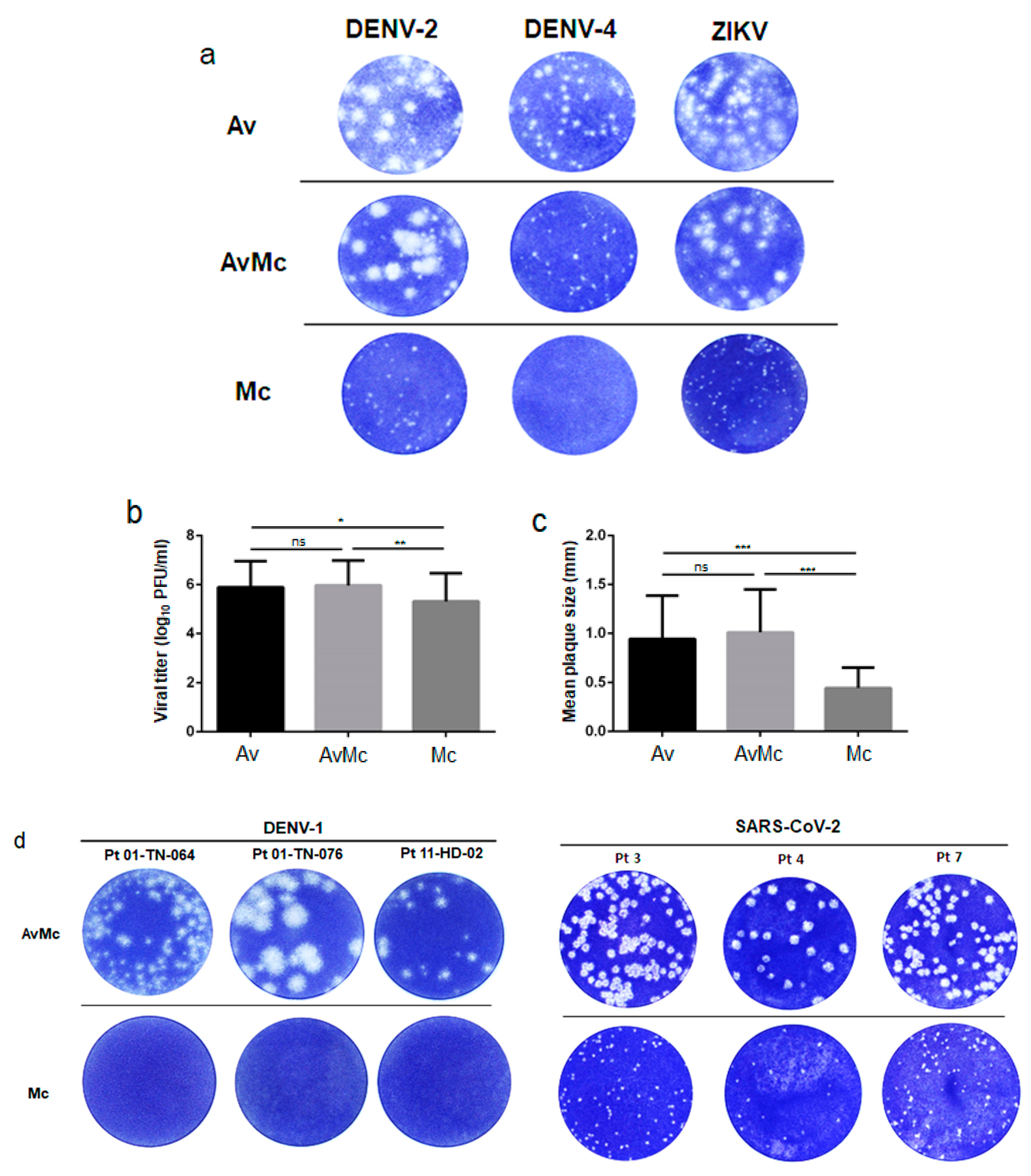
3.1. Infectious Virus Titration Using Laboratory-Established Flavivirus Strains and SARS-CoV-2 Strains
3.2. Dengue Virus Titers in Serum Samples from Dengue Patients and SARS-CoV-2 Titers in Nasopharyngeal Swab Samples from COVID-19 Patients
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Van Kampen, J.J.A.; van de Vijver, D.A.M.C.; Fraaij, P.L.A.; Haagmans, B.L.; Lamers, M.M.; Okba, N.; van den Akker, J.P.C.; Endeman, H.; Gommers, D.A.P.J.; Cornelissen, J.J.; et al. Duration and key determinants of infectious virus shedding in hospitalized patients with coronavirus disease-2019 (COVID-19). Nat. Commun. 2021, 12, 2675. [Google Scholar] [CrossRef]
- Vaughn, D.W.; Green, S.; Kalayanarooj, S.; Innis, B.L.; Nimmannitya, S.; Suntayakorn, S.; Endy, T.P.; Raengsakulrach, B.; Rothman, A.L.; Ennis, F.A.; et al. Dengue viremia titer, antibody response pattern, and virus serotype correlate with disease severity. J. Infect. Dis. 2000, 181, 2–9. [Google Scholar] [CrossRef]
- Murgue, B.; Roche, C.; Chungue, E.; Deparis, X. Prospective study of the duration of magnitude of viremia in children hospitalized during the 1996–1997 dengue-2 outbreak in French Polynesia. J. Med. Virol. 2000, 60, 432–438. [Google Scholar] [CrossRef]
- Libraty, D.H.; Endy, T.P.; Houng, H.S.; Green, S.; Kalayanarooj, S.; Suntayakorn, S.; Chansiriwongs, W.; Vaughn, D.W.; Nisalak, A.; Ennis, F.A.; et al. Differing influence of virus burden and immune activation on disease severity in secondary dengue-3 virus infection. J. Infect. Dis. 2002, 185, 1213–1221. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.K.; Chen, H.L.; Yang, C.H.; Hsieh, S.C.; Juan, C.C.; Chang, S.M.; Yu, C.C.; Lin, L.H.; Huang, J.H.; King, C.C. Slower Rates of Clearance of Viral Load and Virus-Containing Immune Complexes in Patients with Dengue Hemorrhagic Fever. Clin. Infect. Dis. 2006, 43, 1023–1030. [Google Scholar] [CrossRef] [PubMed]
- Bullard, J.; Dust, K.; Funk, D.; Strong, J.E.; Alexander, D.; Garnett, L.; Boodman, C.; Bello, A.; Hedley, A.; Schiffman, Z.; et al. Predicting Infectious Severe Acute Respiratory Syndrome Coronavirus 2 from Diagnostic Samples. Clin. Infect. Dis. 2020, 71, 2663–2666. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.G.; Lee, K.M.; Hsiao, M.J.; Yang, S.L.; Huang, P.N.; Gong, Y.N.; Hsieh, T.H.; Huang, P.W.; Lin, Y.J.; Liu, Y.C.; et al. Culture-Based Virus Isolation To Evaluate Potential Infectivity of Clinical Specimens Tested for COVID-19. J. Clin. Microbiol. 2020, 58, e01068-20. [Google Scholar] [CrossRef] [PubMed]
- Bae, H.G.; Nitsche, A.; Teichmann, A.; Biel, S.S.; Niedrig, M. Detection of yellow fever virus: A comparison of quantitative real-time PCR and plaque assay. J. Virol. Methods 2003, 110, 185–191. [Google Scholar] [CrossRef]
- Yuill, T.M.; Sukhavachana, P.; Nisalak, A.; Russell, P.K. Dengue-virus recovery by direct and delayed plaques in LLC-MK2 cells. Am. J. Trop. Med. Hyg. 1968, 17, 441–448. [Google Scholar] [CrossRef] [PubMed]
- Storch, G.A. Diagnostic Virology. Clin. Infect. Dis. 2000, 31, 739–751. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, K.; Chen, Y.; Yuan, J.; Yi, P.; Ding, C.; Wu, W.; Li, Y.; Ni, Q.; Zou, R.; Li, X.; et al. Factors Associated with Prolonged Viral RNA Shedding in Patients with Coronavirus Disease 2019 (COVID-19). Clin. Infect. Dis. 2020, 71, 799–806. [Google Scholar] [CrossRef]
- Scola, B.L.; Bideau, M.L.; Andreani, J.; Hoang, V.T.; Grimaldier, C.; Colson, P.; Gautret, P.; Raoult, D. Viral RNA load as determined by cell culture as a management tool for discharge of SARS-CoV-2 patients from infectious disease wards. Eur. J. Clin. Microbiol. Infect. Dis. 2020, 39, 1059–1061. [Google Scholar] [CrossRef]
- Mendoza, E.J.; Manguiat, K.; Wood, H.; Drebot, M. Two detailed plaque assay protocols for the quantification of infectious SARS-CoV-2. Curr. Protoc. Microbiol. 2020, 57, e105. [Google Scholar] [CrossRef]
- Matrosovich, M.; Matrosovich, T.; Garten, W.; Klenk, H.D. New low-viscosity overlay medium for viral plaque assays. Virol. J. 2006, 3, 63. [Google Scholar] [CrossRef] [Green Version]
- Yin, Y.; Xu, Y.; Ou, Z.; Su, L.; Xia, H. A simple and highly repeatable viral plaque assay for enterovirus 71. J. Basic Microbiol. 2015, 55, 538–541. [Google Scholar] [CrossRef]
- Baer, A.; Kehn-Hall, K. Viral concentration determination through plaque assays: Using traditional and novel overlay systems. J. Vis. Exp. 2014, 93, e52065. [Google Scholar] [CrossRef]
- Zhao, G.H.; Kapur, N.; Carlin, B.; Selinger, E.; Guthrie, J.T. Characterisation of the interactive properties of microcrystalline cellulose-carboxymethyl cellulose hydrogels. Int. J. Pharm. 2011, 415, 95–101. [Google Scholar] [CrossRef]
- Moi, M.L.; Lim, C.K.; Kotaki, A.; Takasaki, T.; Kurane, I. Development of an antibody-dependent enhancement assay for dengue virus using stable BHK-21 cell lines expressing FcγRIIA. J. Virol. Methods 2010, 163, 205–209. [Google Scholar] [CrossRef]
- Balingit, J.C.; Phu, L.M.H.; Matsuda, M.; Suzuki, R.; Hasebe, F.; Morita, K.; Moi, M.M. A Simple and High-Throughput ELISA-Based Neutralization Assay for the Determination of Anti-Flavivirus Neutralizing Antibodies. Vaccines 2020, 8, 297. [Google Scholar] [CrossRef]
- Pabbaraju, K.; Wong, S.; Gill, K.; Fonseca, K.; Tipples, G.A.; Tellier, R. Simultaneous detection of Zika, Chikungunya and Dengue viruses by a multiplex real-time RT-PCR assay. J. Clin. Virol. 2016, 83, 66–71. [Google Scholar] [CrossRef]
- St. John, A.L.; Rathore, A.P.S. Adaptive immune responses to primary and secondary dengue virus infections. Nat. Rev. Immunol. 2019, 19, 218–230. [Google Scholar] [CrossRef]
- Falconar, A.K.; de Plata, E.; Romero-Vivas, C.M. Altered enzyme-linked immunosorbent assay immunoglobulin M (IgM)/IgG optical density ratios can correctly classify all primary or secondary dengue virus infections 1 day after the onset of symptoms, when all of the viruses can be isolated. Clin. Vaccine Immunol. 2006, 13, 1044–1051. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- WHO. Dengue Duidelines for Diagnosis, Treatment, Prevention and Control; WHO Press: Geneva, Switzerland, 2009. [Google Scholar]
- Yoshikawa, R.; Abe, H.; Igasaki, Y.; Negishi, S.; Goto, H.; Yasuda, J. Development and evaluation of a rapid and simple diagnostic assay for COVID-19 based on loop-mediated isothermal amplification. PLoS Negl. Trop. Dis. 2020, 14, e000855. [Google Scholar] [CrossRef]
- Cooper, P.D. The Plaque Assay of Animal Viruses. Adv. Virus Res. 1962, 8, 319–378. [Google Scholar] [CrossRef]
- Atkinson, B.; Petersen, E. SARS-CoV-2 shedding and infectivity. Lancet 2020, 395, 1339–1340. [Google Scholar] [CrossRef]
- Yamada, S.; Fukushi, S.; Kinoshita, H.; Ohnishi, M.; Suzuki, T.; Fujimoto, T.; Saijo, M.; Maeda, K. Assessment of SARS-CoV-2 infectivity of upper respiratory specimens from COVID-19 patients by virus isolation using VeroE6/TMPRSS2 cells. BMJ Open Respir. Res. 2021, 8, e000830. [Google Scholar] [CrossRef]
- Payne, A.F.; Binduga-Gajewska, I.; Kauffman, E.B.; Kramer, L.D. Quantitation of flaviviruses by fluorescent focus assay. J. Virol. Methods 2006, 134, 183–189. [Google Scholar] [CrossRef] [PubMed]
- Okuno, Y.; Fukunaga, T.; Tadano, M.; Okamoto, Y.; Ohnishi, T.; Takagi, M. Rapid focus reduction neutralization test of Japanese encephalitis virus in microtiter system. Brief report. Arch. Virol. 1985, 86, 129–135. [Google Scholar] [CrossRef]
- Kang, W.; Shin, E.C. Colorimetric Focus-Forming Assay with Automated Focus Counting by Image Analysis for Quantification of Infectious Hepatitis C Virions. PLoS ONE 2012, 7, e43960. [Google Scholar] [CrossRef]
- Zhao, H.; To, K.K.W.; Lam, H.; Zhou, X.; Chan, J.F.W.; Peng, Z.; Lee, A.C.Y.; Cai, J.; Chan, W.M.; lp, J.D.; et al. Cross-linking peptide and repurposed drugs inhibit both entry pathways of SARS-CoV-2. Nat. Commun. 2021, 12, 1517. [Google Scholar] [CrossRef]
- Park, J.G.; Oladunni, F.S.; Chiem, K.; Ye, C.; Pipenbrink, M.; Moran, T.; Walter, M.R.; Kobie, J.; Martinez-Sobrido, L. Rapid in vitro assays for screening neutralizing antibodies and antivirals against SARS-CoV-2. J. Virol. Methods 2021, 287, 113995. [Google Scholar] [CrossRef] [PubMed]
- Amarilla, A.A.; Modhiran, N.; Setoh, Y.X.; Peng, N.Y.G.; Sng, J.D.J.; Liang, B.; McMillan, C.L.D.; Freney, M.E.; Cheung, S.T.M.; Chappell, K.J. An Optimized High-Throughput Immuno-Plaque Assay for SARS-CoV-2. Front. Microbiol. 2021, 12, 625136. [Google Scholar] [CrossRef] [PubMed]
- Gwon, Y.D.; Strand, M.; Lindqvist, R.; Nilsson, E.; Saleeb, M.; Elfsson, M.; Overby, A.K.; Evander, M. Antiviral Activity of Benzavir-2 against Emerging Flaviviruses. Viruses 2020, 12, 351. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- WHO. Update on SARS-CoV-2 Variant Nomenclature, June 2021. Available online: https://www.who.int/docs/default-source/coronaviruse/risk-comms-updates/update60_nomenclature-variants.pdf?sfvrsn=27fc6fa_4 (accessed on 15 September 2021).
| Log10 Virus Titer (PFU/mL) | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Cell Lines | BHK-21 | Vero | FcγR-Expressing BHK-21 | ||||||
| Virus Strain | Av a | AvMc b | Mc c | Av | AvMc | Mc | Av | AvMc | Mc |
| DENV-1 01-44-1 HuNIID | 6.4 | 6.6 | 4.1 | 3.3 | 4.4 | 3.8 | 6.5 | 6.1 | 5.7 |
| DENV-1 99St12A | 4.4 | 5.5 | 2.5 | 4.5 | 4.7 | 4.8 | 4.9 | 6.1 | 4.6 |
| DENV-2 DHF0663 | 5.4 | 5.1 | 4.4 | 5.4 | 5.6 | 5.0 | 5.9 | 5.8 | 5.2 |
| DENV-2 TLC-30 | 5.0 | 4.8 | 4.5 | 5.0 | 4.7 | 5.0 | 4.5 | 4.8 | 4.0 |
| DENV-2 08-77 | 4.9 | 4.3 | 5.2 | 5.6 | 5.4 | 5.6 | 6.1 | 5.8 | 5.2 |
| DENV-2 00St22A | 4.7 | 5.6 | 5.7 | 6.3 | 6.0 | 6.0 | 6.6 | 5.6 | 5.8 |
| DENV-3 CH53469 | 5.8 | 6.7 | 5.4 | 4.6 | 4.3 | 4.1 | 5.9 | 6.9 | 5.8 |
| DENV-4 SLMC318 | 5.7 | 5.6 | 4.9 | 5.4 | 4.1 | 4.4 | 6.1 | 6.0 | 4.3 |
| DENV-4 TVP360 | 4.7 | 5.1 | 3.4 | 4.8 | 4.2 | 4.6 | 4.9 | 5.0 | 3.9 |
| ZIKV MR766 | 6.2 | 6.8 | 6.9 | 8.2 | 7.6 | 7.1 | 6.9 | 7.4 | 6.1 |
| ZIKV PRVABC59 | 5.5 | 5.9 | 5.0 | 7.1 | 6.8 | 6.7 | 5.1 | 6.1 | 4.4 |
| ZIKV MRS_OPY_Martinique_PaRi_2015 | 5.8 | 6.1 | 5.6 | 7.1 | 6.9 | 6.8 | 5.9 | 6.2 | 4.9 |
| ZIKV H/PF/2013 | 6.5 | 6.7 | 5.9 | 7.4 | 7.2 | 6.2 | 6.8 | 6.7 | 5.7 |
| JEV OH0566 | 7.8 | 7.8 | 7.9 | 7.6 | 7.7 | 7.5 | 7.9 | 7.7 | 7.7 |
| YFV 17D | 6.6 | 6.5 | 5.2 | 5.8 | 6.3 | 6.5 | 6.9 | 6.6 | 4.9 |
| Cell Lines | BHK-21 | Vero | FcγR-Expressing BHK | ||||||
|---|---|---|---|---|---|---|---|---|---|
| Virus Strain | Av a | AvMc b | Mc c | Av | AvMc | Mc | Av | AvMc | Mc |
| DENV-1 01-44-1 HuNIID | 0.8 ± 0.2 | 0.8 ± 0.4 | 0.3 ± 0.1 | 0.3 ± 0.2 | 0.3 ± 0.2 | 0.3 ± 0.1 | 1.1 ± 0.3 | 0.4 ± 0.1 | 0.4 ± 0.1 |
| DENV-1 99St12A | 1.0 ± 0.3 | 1.5 ± 0.3 | 0.4 ± 0.1 | 0.5 ± 0.2 | 0.5 ± 0.2 | 0.3 ± 0.1 | 0.9 ± 0.2 | 1.0 ± 0.2 | 0.2 ± 0.0 |
| DENV-2 DHF0663 | 1.1 ± 0.3 | 1.1 ± 0.2 | 0.2 ± 0.1 | 1.1± 0.3 | 1.5 ± 0.3 | 0.7 ± 0.2 | 1.0 ± 0.4 | 1.1 ± 0.7 | 0.5 ± 0.1 |
| DENV-2 TLC-30 | 1.7 ± 0.5 | 1.8 ± 0.6 | 0.5 ± 0.1 | 1.2 ± 0.3 | 1.0 ± 0.5 | 0.6 ± 0.1 | 0.8 ± 0.1 | 1.3 ± 0.1 | 0.3 ± 0.0 |
| DENV-2 08-77 | 0.4 ± 0.1 | 0.6 ± 0.1 | 0.5 ± 0.1 | 1.0 ± 0.5 | 0.6 ± 0.2 | 0.4 ± 0.1 | 0.5 ± 0.1 | 1.4 ± 0.2 | 0.3 ± 0.0 |
| DENV-2 00St22A | 0.6 ± 0.1 | 0.8 ± 0.2 | 0.3 ± 0.1 | 1.2 ± 0.4 | 0.9 ± 0.4 | 0.6 ± 0.2 | 0.7 ± 0.2 | 1.5 ± 0.3 | 0.3 ± 0.1 |
| DENV-3 CH53469 | 1.5 ± 0.5 | 1.8 ± 0.7 | 0.5 ± 0.1 | 1.3 ± 0.5 | 0.6 ± 0.2 | 0.2± 0.0 | 1.9 ± 0.8 | 1.7 ± 0.8 | 0.4 ± 0.1 |
| DENV-4 SLMC318 | 0.9 ± 0.2 | 0.7 ± 0.4 | 0.5 ± 0.1 | 0.6 ± 0.1 | 0.7 ± 0.3 | 0.4 ± 0.1 | 0.7 ± 0.1 | 0.4 ± 0.3 | 0.4 ± 0.1 |
| DENV-4 TVP360 | 0.9 ± 0.2 | 1.8 ± 0.6 | 0.4 ± 0.1 | 0.9 ± 0.6 | 0.8 ± 0.2 | 0.8 ± 0.3 | 1.0 ± 0.3 | 1.6 ± 0.3 | 0.3 ± 0.1 |
| ZIKV MR766 | 0.8 ± 0.2 | 1.1 ± 0.6 | 0.4 ± 0.1 | 2.2 ± 0.0 | 1.7 ± 0.4 | 1.1 ± 0.2 | 1.2 ± 0.4 | 1.5 ± 0.6 | 0.3 ± 0.0 |
| ZIKV PRVABC59 | 0.4 ± 0.1 | 0.6 ± 0.2 | 0.5 ± 0.1 | 1.2 ± 0.5 | 1.3 ± 0.3 | 0.5 ± 0.1 | 0.4 ± 0.1 | 0.5 ± 0.1 | 0.4 ± 0.2 |
| ZIKV MRS_OPY_Martinique_PaRi_2015 | 0.6 ± 0.1 | 0.8 ± 0.2 | 0.4 ± 0.1 | 1.3 ± 0.6 | 1.2 ± 0.4 | 0.5 ± 0.2 | 0.5 ± 0.2 | 0.6 ± 0.1 | 0.2 ± 0.1 |
| ZIKV H/PF/2013 | 0.4 ± 0.1 | 0.8 ± 0.3 | 0.4 ± 0.1 | 1.4± 0.9 | 1.4 ± 0.6 | 0.9 ± 0.3 | 0.6 ± 0.2 | 0.8 ± 0.3 | 0.2 ± 0.0 |
| JEV OH0566 | 0.2 ±0.0 | 0.6 ± 0.1 | 1.0 ± 0.3 | 1.4 ± 0.4 | 1.5 ± 0.4 | 0.8 ± 0.1 | 1.3 ± 0.9 | 0.7 ± 0.3 | 0.3 ± 0.1 |
| YFV 17D | 1.2 ± 0.5 | 0.6 ± 0.1 | 0.3 ± 0.1 | 0.3 ± 0.1 | 0.7 ± 0.2 | 0.3 ± 0.1 | 1.4 ± 0.2 | 0.8 ± 0.3 | 0.4 ± 0.1 |
| Virus Titer (Log10 PFU/mL) Vero | Plaque Size | |||
|---|---|---|---|---|
| SARS-CoV-2 Strain | AvMc a | Mc b | AvMc | Mc |
| 2019-nCoV/Japan/TY/WK-521/2020 | 4.9 | 4.8 | 1.4 ± 0.5 | 0.6 ± 0.1 |
| NGS1B | 5.5 | 4.6 | 1.3 ± 0.5 | 0.6 ± 0.1 |
| hCoV-19/Japan/QK002/2020 (Alpha) d | 3.7 | ND c | 0.7 ± 0.2 | ND |
| hCoV-19/Japan/TY7-501/2021(Gamma) | 3.6 | ND | 0.8 ± 0.1 | ND |
| hCoV-19/Japan/TY8-612/2021 (Beta) | 3.5 | 3.2 | 0.7 ± 0.1 | 0.3 ± 0.1 |
| Sample Code | Days a | Log10 Genome Copies/mL | Dengue ELISA | Virus Titer (Log10 PFU/mL) | |||
|---|---|---|---|---|---|---|---|
| OD e-IgG | OD-IgM | IgM/IgG OD Ratio | AvMc c | Mc d | |||
| Primary Infection | |||||||
| 01-TN-145 | 0 | 9.5 | 0.8 | 0.1 | 0.2 | 6.1 ± 0.03 | ND |
| 01-TN-087 | 1 | 10.2 | 0.5 | 0.1 | 0.2 | 5.4 ± 0.04 | ND |
| 29-HD-011 | 1 | 9.5 | 0.1 | 0.1 | 1.0 | 5.1 ± 0.04 | ND |
| 01-TN-067 | 2 | 7.6 | 0.5 | 0.1 | 0.2 | 6.4 ± 0.1 | 4.49 ± 0.03 |
| 01-TN-076 | 2 | 10.3 | 0.8 | 0.07 | 0.09 | 7.3 ± 0.1 | ND |
| 01-TN-078 | 2 | 9.2 | 0.9 | 0.06 | 0.06 | 6.2 ± 0.2 | ND |
| 01-TN-091 | 2 | 10.5 | 0.4 | 0.2 | 0.4 | 6.6 ± 0.02 | ND |
| 01-TN-040 | 3 | 8.1 | 0.5 | 0.2 | 0.3 | 6.0 ± 0.2 | ND |
| 01-TN-063 | 3 | 6.9 | 0.8 | 0.2 | 0.2 | 6.1 ± 0.07 | ND |
| 04-HD-010 | 7 | 5.2 | 1.8 | 3.5 | 2.0 | ND b | ND |
| 08-HD-001 | 8 | 7.6 | 1.0 | 2.6 | 2.5 | ND | ND |
| 09-HD-014 | 8 | 5.4 | 1.6 | 3.3 | 2.0 | ND | ND |
| Secondary Infection | |||||||
| 01-TN-088 | 1 | 10.3 | 1.1 | 0.13 | 8.5 | 6.3 ± 0.2 | ND |
| 01-TN-077 | 1 | 10.4 | 1.0 | 0.05 | 22.1 | 5.0 ± 0.1 | ND |
| 01-TN-106 | 3 | 10.0 | 1.5 | 0.16 | 9.9 | 5.3 ± 0.06 | ND |
| 11-HD-006 | 3 | 7.7 | 2.3 | 0.45 | 5.2 | ND | ND |
| 11-HD-012 | 3 | 8.9 | 1.0 | 0.14 | 7.2 | 5.8 ± 0.1 | ND |
| 29-HD-001 | 3 | 8.9 | 1.4 | 0.30 | 4.7 | ND | ND |
| 01-TN-056 | 3 | 7.6 | 1.5 | 0.13 | 11.5 | 6.1 ± 0.1 | ND |
| 01-TN-061 | 3 | 6.8 | 1.3 | 0.13 | 10.0 | 6.7 ± 0.03 | ND |
| 01-TN-064 | 3 | 7.5 | 1.0 | 0.14 | 6.7 | 6.4 ± 0.08 | ND |
| 11-HD-002 | 4 | 8.3 | 1.1 | 0.16 | 0.2 | 6.0 ± 0.05 | 5.5 ± 0.03 |
| 01-TN-062 | 5 | 7.2 | 1.6 | 0.13 | 12.8 | 6.1 ± 0.1 | ND |
| 01-TN-068 | 5 | 6.5 | 2.0 | 0.39 | 5.1 | 5.8 ± 0.2 | ND |
| 01-PK-119 | 6 | 5.0 | 1.4 | 0.20 | 7.0 | ND | ND |
| 11-HD-004 | 6 | 6.1 | 1.9 | 2.0 | 1.0 | ND | ND |
| 03-HD-014 | 8 | 9.5 | 1.9 | 0.79 | 2.4 | 6.5 ± 0.01 | ND |
| Sample No | Virus Load (log10 Genome Copies/mL) | Virus Titer (Log10 PFU/mL) | Plaque Size (Mean Diameters mm ± SD) | ||
|---|---|---|---|---|---|
| AvMc | Mc | AvMc | Mc | ||
| 1 | 10.9 | 4.3 | 4.2 | 0.87 ± 0.34 | 0.29 ± 0.1 |
| 2 | 10.2 | 3.4 | ND a | NT | NT |
| 3 | 9.9 | 5.1 | 4.9 | 1.0 ± 0.2 | 0.4 ± 0.07 |
| 4 | 9.7 | 3.8 | 3.6 | 1.3 ± 0.2 | 0.3 ± 0.09 |
| 5 | 9.7 | 3.2 | 3.1 | 1.4 ± 0.2 | 0.38 ± 0.1 |
| 6 | 9.5 | 2.6 | ND | NT b | NT |
| 7 | 9.4 | 4.1 | 3.94 | 0.96 ± 0.2 | 0.37 ± 0.1 |
| 8 | 9.21 | 4.0 | 3.81 | 1.1 ± 0.3 | 0.37 ± 0.1 |
| 9 | 9.03 | 3.7 | 4.11 | 1.7 ± 0.3 | 0.4 ± 0.1 |
| 10 | 8.51 | 3.3 | 3.20 | 0.7 ± 0.1 | 0.4 ± 0.1 |
| 11 | 8.44 | 2.9 | 3.23 | 1.4 ± 0.2 | 0.5 ± 0.1 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fukuta, M.; Nguyen, C.T.; Nguyen, T.T.T.; Nguyen, T.T.N.; Vu, T.B.H.; Takemura, T.; Nguyen, L.K.H.; Inoue, S.; Morita, K.; Le, T.Q.M.; et al. Discrepancies in Infectivity of Flavivirus and SARS-CoV-2 Clinical Samples: An Improved Assay for Infectious Virus Shedding and Viremia Assessment. Int. J. Environ. Res. Public Health 2021, 18, 9845. https://doi.org/10.3390/ijerph18189845
Fukuta M, Nguyen CT, Nguyen TTT, Nguyen TTN, Vu TBH, Takemura T, Nguyen LKH, Inoue S, Morita K, Le TQM, et al. Discrepancies in Infectivity of Flavivirus and SARS-CoV-2 Clinical Samples: An Improved Assay for Infectious Virus Shedding and Viremia Assessment. International Journal of Environmental Research and Public Health. 2021; 18(18):9845. https://doi.org/10.3390/ijerph18189845
Chicago/Turabian StyleFukuta, Mizuki, Co Thach Nguyen, Thi Thu Thuy Nguyen, Thi Thanh Ngan Nguyen, Thi Bich Hau Vu, Taichiro Takemura, Le Khanh Hang Nguyen, Shingo Inoue, Kouichi Morita, Thi Quynh Mai Le, and et al. 2021. "Discrepancies in Infectivity of Flavivirus and SARS-CoV-2 Clinical Samples: An Improved Assay for Infectious Virus Shedding and Viremia Assessment" International Journal of Environmental Research and Public Health 18, no. 18: 9845. https://doi.org/10.3390/ijerph18189845
APA StyleFukuta, M., Nguyen, C. T., Nguyen, T. T. T., Nguyen, T. T. N., Vu, T. B. H., Takemura, T., Nguyen, L. K. H., Inoue, S., Morita, K., Le, T. Q. M., Hasebe, F., & Moi, M. L. (2021). Discrepancies in Infectivity of Flavivirus and SARS-CoV-2 Clinical Samples: An Improved Assay for Infectious Virus Shedding and Viremia Assessment. International Journal of Environmental Research and Public Health, 18(18), 9845. https://doi.org/10.3390/ijerph18189845







