Antibiotic-Resistant Bacteria in Aquaculture and Climate Change: A Challenge for Health in the Mediterranean Area
Abstract
1. Introduction
2. Antibiotic Use in Aquaculture
3. Antibiotic Residues as Emerging Pollutants
4. Induction of Antibiotic Resistance
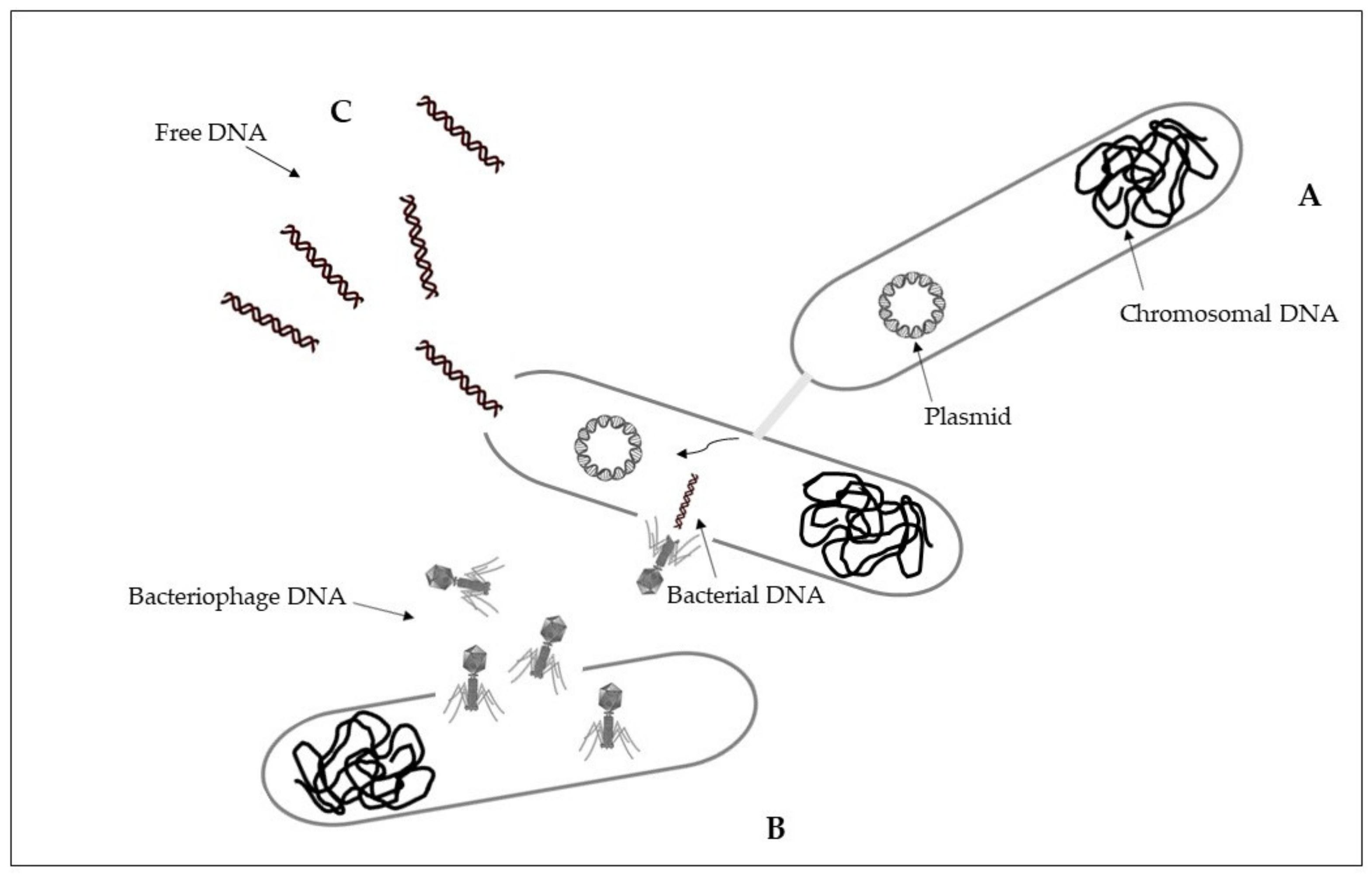
5. Transfer of Antibiotic Resistance between Bacteria
5.1. Conjugation
5.2. Transduction
5.3. Transformation
6. Transfer of Antibiotic Resistance from Aquaculture Bacteria to Human Pathogens
6.1. Quinolones
6.2. Tetracyclines
6.3. β-Lactams, Macrolides, Fosfomycin, Chloramphenicol, Colistin, Florfenicol
7. Antibiotic Resistance in the Mediterranean Basin
8. Antibiotic Resistance and Temperature
8.1. Effect Cross-Protection
8.2. Modifications Induced by Temperature
8.3. Epigenetic Modifications
9. Climate Change and Antibiotic Resistance
10. Climate Change in the Mediterranean Area
Mediterranean-Type Ecosystem
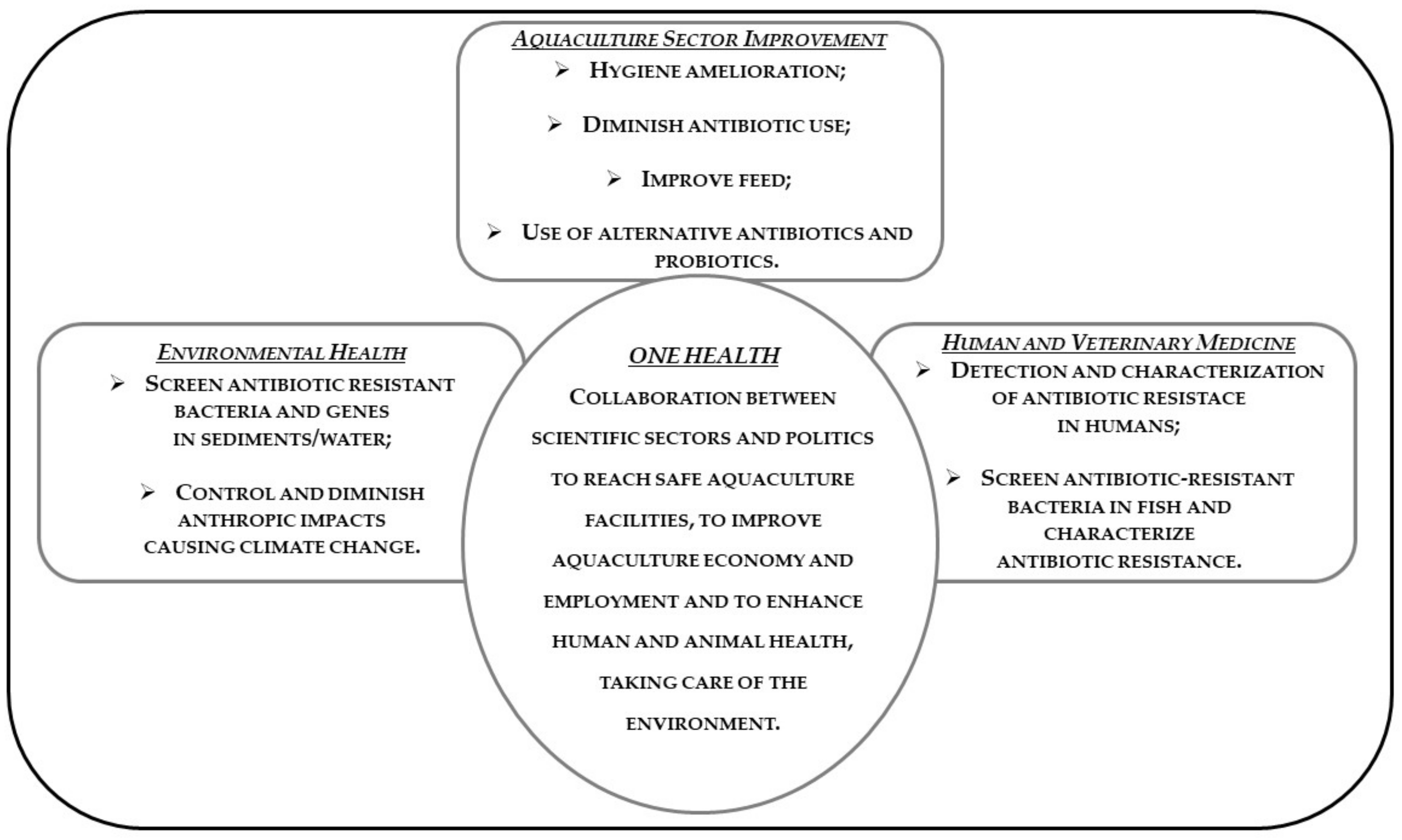
11. One Health and Antibiotic Resistance in Aquaculture
Good Practices to Be Adopted
12. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Maite, C.; Peral, I.P.; Ramos, S.; Basurco, B.; López-Francos, M.A.; Cavallo, M.; Perez, J.; Aguilera, C.; Fu-rones, D.; Reverté, C.; et al. Deliverable 1.2 of the Horizon 2020 Project MedAID (GA number 727315). Available online: https://archimer.ifremer.fr/doc/00515/62630 (accessed on 5 January 2021).
- Panagiotis, P. (Ed.) Challenges Facing Marine Aquaculture in the EU-Mediterranean. In Trends in Fisheries and Aquatic Animal Health; Bentham Science Publishers Ltd.: Sharjah, United Arab Emirates, 2017; pp. 42–84. [Google Scholar]
- UN. The Sustainable Development Goals Report; United Nations: New York, NY, USA, 2018. [Google Scholar]
- FAO. Definition of Aquaculture, Seventh Session of the IPFC Working Party of Expects on Aquaculture, IPFC/WPA/WPZ; RAPA/FAO: Bangkok, Thailand, 1988; pp. 1–3. [Google Scholar]
- Courtenay, W.R.; Bardach, J.E.; Ryther, J.H.; McLarney, W.O. Aquaculture, the Farming and Husbandry of Freshwater and Marine Organisms. Copeia 1973, 1973, 826. [Google Scholar] [CrossRef]
- Rabanal, H.R. History of Aquaculture; ASEAN/SF/88/Tech 7; FAO: Manila, Philippines, 1988. [Google Scholar]
- Mancuso, M. Farming Thunnus thynnus in Mediterranean Sea. Is an Illusion? Poult. Fish Wildl. Sci. 2015, 3, e110. [Google Scholar] [CrossRef]
- Barazi-Yeroulanos, L. Synthesis of Mediterranean Marine Finfish Aquaculture—A Marketing and Promotion Strategy. Studies and Reviews: General Fisheries Commission for the Mediterranean; FAO No. 88; FAO: Rome, Italy, 2010; p. 198. [Google Scholar]
- FAO. The State of World Fishery and Aquaculture 2020 (SOFIA); Food and Agriculture Organization of the United Nations: Rome, Italy, 2020. [Google Scholar] [CrossRef]
- Grigorakis, K.; Rigos, G. Aquaculture effects on environmental and public welfare—The case of Mediterranean mariculture. Chemosphere 2011, 85, 899–919. [Google Scholar] [CrossRef] [PubMed]
- FAO. The State of World Fisheries and Aquaculture 2018—Meeting the Sustainable Development Goals; FAO: Rome, Italy, 2018; Available online: http://www.fao.org/3/i9540en/I9540EN.pdf (accessed on 21 January 2021).
- Fleming, A. On the Antibacterial Action of Cultures of a Penicillium, with Special Reference to their Use in the Isolation of B. influenzæ. Br. J. Exp. Pathol. 1929, 10, 226–236. [Google Scholar] [CrossRef]
- EMA/AMEG. Preliminary Risk Profiling for New Antimicrobial Veterinary Medicinal Products. Available online: https://www.ema.europa.eu/en/documents/regulatory-procedural-guideline/answer-request-european-commission-updating-scientific-advice-impact-public-health-animal-health-use_en-0.pdf2019 (accessed on 2 February 2021).
- Kümmerer, K. Antibiotics in the aquatic environment: A review: Part I. Chemosphere 2009, 75, 417–434. [Google Scholar] [CrossRef] [PubMed]
- Done, H.Y.; Venkatesan, A.K.; Halden, R.U. Does the Recent Growth of Aquaculture Create Antibiotic Resistance Threats Different from those Associated with Land Animal Production in Agriculture? AAPS J. 2015, 17, 513–524. [Google Scholar] [CrossRef]
- Quesada, S.P.; Paschoal, J.A.R.; Reyes, F.G.R. Considerations on the Aquaculture Development and on the Use of Veterinary Drugs: Special Issue for Fluoroquinolones-A Review. J. Food Sci. 2013, 78, R1321–R1333. [Google Scholar] [CrossRef] [PubMed]
- Santos, L.; Ramos, F. Antimicrobial resistance in aquaculture: Current knowledge and alternatives to tackle the problem. Int. J. Antimicrob. Agents 2018, 52, 135–143. [Google Scholar] [CrossRef]
- Rigos, G.; Troisi, G.M. Antibacterial Agents in Mediterranean Finfish Farming: A Synopsis of Drug Pharmacoki-netics in Important Euryhaline Fish Species and Possible Environmental Implications. Rev. Fish Biol. Fish. 2005, 15, 53–73. [Google Scholar] [CrossRef]
- Fang, H.; Huang, K.; Yu, J.; Ding, C.; Wang, Z.; Zhao, C.; Yuan, H.; Wang, Z.; Wang, S.; Hu, J.; et al. Meta-genomic analysis of bacterial communities and antibiotic resistance genes in the Eriocheir sinensis freshwater aqua-culture environment. Chemosphere 2019, 224, 202–211. [Google Scholar] [CrossRef] [PubMed]
- Sarmah, A.K.; Meyer, M.T.; Boxall, A.B. A global perspective on the use, sales, exposure pathways, occurrence, fate and effects of veterinary antibiotics (VAs) in the environment. Chemosphere 2006, 65, 725–759. [Google Scholar] [CrossRef] [PubMed]
- Sørum, H. Antimicrobial drug resistance in fish pathogens. In Antimicrobial Resistance in Bacteria of Animal Origin; Aarestrup, F.M., Ed.; ASM Press: Washington, DC, USA, 2006; pp. 213–238. [Google Scholar]
- Sapkota, A.; Sapkota, A.R.; Kucharski, M.; Burke, J.; McKenzie, S.; Walker, P.; Lawrence, R. Aquaculture practices and potential human health risks: Current knowledge and future priorities. Environ. Int. 2008, 34, 1215–1226. [Google Scholar] [CrossRef]
- Cabello, F.C.; Godfrey, H.P.; Tomova, A.; Ivanova, L.; Dölz, H.; Millanao, A.; Buschmann, A.H. Antimicrobial use in aquaculture re-examined: Its relevance to antimicrobial resistance and to animal and human health. Environ. Microbiol. 2013, 15, 1917–1942. [Google Scholar] [CrossRef] [PubMed]
- Armstrong, S.M.; Armstrong, S.M.; Hargrave, B.T.; Hargrave, B.T.; Haya, K.; Haya, K. Antibiotic Use in Finfish Aquaculture: Modes of Action, Environmental Fate, and Microbial Resistance. In Handbook of Environmental Chemistry; Springer: Berlin/Heidelberg, Germany, 2005; Volume 5M, pp. 341–357. [Google Scholar] [CrossRef]
- Buschmann, A.H.; Tomova, A.; López, A.; Maldonado, M.A.; Henríquez, L.A.; Ivanova, L.; Moy, F.; Godfrey, H.P.; Cabello, F.C. Salmon aqua-culture and antimicrobial resistance in the marine environment. PLoS ONE 2012, 7, e42724. [Google Scholar] [CrossRef] [PubMed]
- Capone, D.G.; Weston, D.P.; Miller, V.; Shoemaker, C. Antibacterial residues in marine sediments and inverte-brates following chemotherapy in aquaculture. Aquaculture 1996, 145, 55–75. [Google Scholar] [CrossRef]
- Boxall, A.B.; Fogg, L.A.; Blackwell, P.A.; Kay, P.; Pemberton, E.J.; Croxford, A. Veterinary medicines in the envi-ronment. Rev. Environ. Contam. Toxicol. 2004, 180, 1–91. [Google Scholar] [PubMed]
- Hektoen, H.; Berge, J.A.; Hormazabal, V.; Yndestad, M. Persistence of antibacterial agents in marine sediments. Aquaculture 1995, 133, 175–184. [Google Scholar] [CrossRef]
- Serwecińska, L. Antimicrobials and Antibiotic-Resistant Bacteria: A Risk to the Environment and to Public Health. Water 2020, 12, 3313. [Google Scholar] [CrossRef]
- Koeypudsa, W.; Yakupitiyage, A.; Tangtrongpiros, J. The fate of chlortetracycline residues in a simulated chick-en–fish integrated farming systems. Aquac. Res. 2005, 36, 570–577. [Google Scholar] [CrossRef]
- Lai, H.-T.; Lin, J.-J. Degradation of oxolinic acid and flumequine in aquaculture pond waters and sediments. Chemosphere 2009, 75, 462–468. [Google Scholar] [CrossRef] [PubMed]
- Hoa, P.T.P.; Nonaka, L.; Viet, P.H.; Suzuki, S. Detection of the sul1, sul2, and sul3 genes in sulfonamide-resistant bacteria from wastewater and shrimp ponds of north Vietnam. Sci. Total Environ. 2008, 405, 377–384. [Google Scholar] [CrossRef]
- Smith, P. Antimicrobial resistance in aquaculture. Rev. Sci. Tech. OIE 2008, 27, 243–264. [Google Scholar] [CrossRef]
- Miranda, C. Antimicrobial resistance associated with salmonid farming. In Antimicrobial Resistance in the Environment; Keen, P.L., Montforts, M.H.H.M., Eds.; Wiley-Blackwell: Hoboken, NJ, USA, 2012; pp. 423–451. [Google Scholar]
- Kümmerer, K. Resistance in the environment. J. Antimicrob. Chemother. 2004, 54, 311–320. [Google Scholar] [CrossRef]
- Kalantzi, I.; Rico, A.; Mylon, K.; Pergantis, S.A.; Tsapakis, M. Fish farming, metals and antibiotics in the eastern Mediterranean Sea: Is there a threat to sediment wild life? Sci. Total Environ. 2021, 764, 142843. [Google Scholar] [CrossRef] [PubMed]
- Andersson, D.I.; Hughes, D. Evolution of antibiotic resistance at non-lethal drug concentrations. Drug Resist. Updat. 2012, 15, 162–172. [Google Scholar] [CrossRef] [PubMed]
- Gullberg, E.; Cao, S.; Berg, O.G.; Ilbäck, C.; Sandegren, L.; Hughes, D.; Andersson, D.I. Selection of Resistant Bacteria at Very Low Antibiotic Concentrations. PLoS Pathog. 2011, 7, e1002158. [Google Scholar] [CrossRef]
- Bengtsson-Palme, J.; Larsson, D.J. Concentrations of antibiotics predicted to select for resistant bacteria: Pro-posed limits for environmental regulation. Environ. Int. 2016, 86, 140–149. [Google Scholar] [CrossRef] [PubMed]
- Miranda, C.D.; Tello, A.; Keen, P.L. Mechanisms of antimicrobial resistance in finfish aquaculture environments. Front. Microbiol. 2013, 4, 233. [Google Scholar] [CrossRef]
- Zhu, Y.-G.; Zhao, Y.; Li, B.; Huang, C.-L.; Zhang, S.-Y.; Si-Yu, Z.; Chen, Y.-S.; Zhang, T.; Gillings, M.R.; Su, J.-Q. Continental-scale pollution of estuaries with antibiotic resistance genes. Nat. Microbiol. 2017, 2, 16270. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.; Nishino, K.; Roberts, M.C.; Tolmasky, M.; Aminov, R.I.; Zhang, L. Mechanisms of antibiotic resistance. Front. Microbiol. 2015, 6, 34. [Google Scholar] [CrossRef] [PubMed]
- Romero, J.; Gloria, C.; Navarrete, P. Antibiotics in Aquaculture—Use, Abuse and Alternatives. In Health and Environment in Aquaculture; IntechOpen: Rijeka, Croatia, 2012. [Google Scholar] [CrossRef]
- Rigos, G.; Kogiannou, D.; Padrós, F.; Cristòfol, C.; Florio, D.; Fioravanti, M.; Zarza, C. Best therapeutic practices for the use of antibacterial agents in finfish aquaculture: A particular view on European seabass (Dicentrarchus labrax) and gilthead seabream (Sparus aurata) in Mediterranean aquaculture. Rev. Aquac. 2020, 1, 39. [Google Scholar] [CrossRef]
- Watts, J.E.M.; Schreier, H.J.; Lanska, L.; Hale, M.S. The Rising Tide of Antimicrobial Resistance in Aquaculture: Sources, Sinks and Solutions. Mar. Drugs 2017, 15, 158. [Google Scholar] [CrossRef]
- Zago, V.; Veschetti, L.; Patuzzo, C.; Malerba, G.; Lleo, M.M. Shewanella algae and Vibrio spp. strains isolated in Italian aquaculture farms are reservoirs of antibiotic resistant genes that might constitute a risk for human health. Mar. Pollut. Bull. 2020, 154, 111057. [Google Scholar] [CrossRef]
- Anastasiou, T.I.; Mandalakis, M.; Krigas, N.; Vézignol, T.; Lazari, D.; Katharios, P.; Dailianis, T.; Antonopoulou, E. Comparative Evaluation of Essential Oils from Medicinal-Aromatic Plants of Greece: Chemical Composition, Antioxidant Capacity and Antimicrobial Activity against Bacterial Fish Pathogens. Molecules 2019, 25, 148. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.; Li, J.; Chen, H.; Bond, P.L.; Yuan, Z. Metagenomic analysis reveals wastewater treatment plants as hotspots of antibiotic resistance genes and mobile genetic elements. Water Res. 2017, 123, 468–478. [Google Scholar] [CrossRef]
- Erauso, G.; Lakhal, F.; Bidault-Toffin, A.; Le Chevalier, P.; Bouloc, P.; Paillard, C.; Jacq, A. Evidence for the Role of Horizontal Transfer in Generating pVT1, a Large Mosaic Conjugative Plasmid from the Clam Pathogen, Vibrio tapetis. PLoS ONE 2011, 6, e16759. [Google Scholar] [CrossRef] [PubMed]
- Abe, K.; Nomura, N.; Suzuki, S. Biofilms: Hot spots of horizontal gene transfer (HGT) in aquatic environments, with a focus on a new HGT mechanism. FEMS Microbiol. Ecol. 2020, 96, 031. [Google Scholar] [CrossRef]
- Barnes, M.A.; Turner, C.R.; Jerde, C.L.; Renshaw, M.A.; Chadderton, W.L.; Lodge, D.M. Environmental Conditions Influence eDNA Persistence in Aquatic Systems. Environ. Sci. Technol. 2014, 48, 1819–1827. [Google Scholar] [CrossRef]
- Dong, P.; Wang, H.; Fang, T.; Wang, Y.; Ye, Q. Assessment of extracellular antibiotic resistance genes (eARGs) in typical environmental samples and the transforming ability of eARG. Environ. Int. 2019, 125, 90–96. [Google Scholar] [CrossRef] [PubMed]
- Yuan, K.; Wang, X.; Chen, X.; Zhao, Z.; Fang, L.; Chen, B.; Jiang, J.; Luan, T.; Chen, B. Occurrence of antibiotic re-sistance genes in extracellular and intracellular DNA from sediments collected from two types of aquaculture farms. Chemosphere 2019, 234, 520–527. [Google Scholar] [CrossRef]
- Nielsen, K.M.; Johnsen, P.J.; Bensasson, D.; Daffonchio, D. Release and persistence of extracellular DNA in the environment. Environ. Biosaf. Res. 2007, 6, 37–53. [Google Scholar] [CrossRef] [PubMed]
- Poly, F.; Chenu, C.; Simonet, P.; Rouiller, J.; Monrozier, L.J. Differences between Linear Chromosomal and Supercoiled Plasmid DNA in Their Mechanisms and Extent of Adsorption on Clay Minerals. Langmuir 2000, 16, 1233–1238. [Google Scholar] [CrossRef]
- Zarei-Baygi, A.; Smith, A.L. Intracellular versus extracellular antibiotic resistance genes in the environment: Prevalence, horizontal transfer, and mitigation strategies. Bioresour. Technol. 2021, 319, 124181. [Google Scholar] [CrossRef]
- Liu, S.-S.; Qu, H.-M.; Yang, D.; Hu, H.; Liu, W.-L.; Qiu, Z.-G.; Hou, A.-M.; Guo, J.; Li, J.-W.; Shen, Z.-Q.; et al. Chlorine disinfection increases both intracellular and extracellular antibiotic resistance genes in a full-scale wastewater treatment plant. Water Res. 2018, 136, 131–136. [Google Scholar] [CrossRef] [PubMed]
- Thomas, C.M.; Nielsen, K.M. Mechanisms of, and Barriers to, Horizontal Gene Transfer between Bacteria. Nat. Rev. Genet. 2005, 3, 711–721. [Google Scholar] [CrossRef] [PubMed]
- Von Wintersdorff, C.J.H.; Penders, J.; van Niekerk, J.M.; Mills, N.D.; Majumder, S.; van Alphen, L.B.; Savelkoul, P.H.M.; Wolffs, P.F.G. Dissemination of Antimicrobial Resistance in Microbial Ecosystems through Horizontal Gene Transfer. Front. Microbiol. 2016, 7, 173. [Google Scholar] [CrossRef]
- Vrancianu, C.O.; Popa, L.I.; Bleotu, C.; Chifiriuc, M.C. Targeting Plasmids to Limit Acquisition and Transmission of Antimicrobial Resistance. Front. Microbiol. 2020, 11, 761. [Google Scholar] [CrossRef] [PubMed]
- Lopatkin, A.J.; Meredith, H.R.; Srimani, J.K.; Pfeiffer, C.; Durrett, R.; You, L. Persistence and reversal of plas-mid-mediated antibiotic resistance. Nat. Commun. 2017, 8, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Burrus, V.; Pavlovic, G.; Decaris, B.; Guédon, G. Conjugative transposons: The tip of the iceberg. Mol. Microbiol. 2002, 46, 601–610. [Google Scholar] [CrossRef] [PubMed]
- Chandrasekaran, S.; Venkatesh, B.; Lalithakumari, D. Transfer and expression of a multiple antibiotic resistance plasmid in marine bacteria. Curr. Microbiol. 1998, 37, 347–351. [Google Scholar] [CrossRef] [PubMed]
- Bergman, J.; Fineran, P.; Petty, N.; Salmond, G. Transduction: The transfer of host DNA by bacteriophages. In Encyclopedia of Microbiology, 4th ed.; Elsevier: Cambridge, MA, USA, 2019; pp. 458–473. [Google Scholar]
- Jiang, S.; Paul, J.H. Gene Transfer by Transduction in the Marine Environment. Appl. Environ. Microbiol. 1998, 64, 2780–2787. [Google Scholar] [CrossRef]
- Brown-Jaque, M.; Calero-Cáceres, W.; Muniesa, M. Transfer of antibiotic-resistance genes via phage-related mobile elements. Plasmid 2015, 79, 1–7. [Google Scholar] [CrossRef]
- Modi, S.R.; Lee, H.H.; Spina, C.S.; Collins, J.J. Antibiotic treatment expands the resistance reservoir and ecologi-cal network of the phage metagenome. Nature 2013, 499, 219–222. [Google Scholar] [CrossRef] [PubMed]
- Landén, R.; Heierson, A.; Boman, H.G. A phage for generalized transduction in Bacillus thuringiensis and map-ping of four genes for antibiotic resistance. Microbiology 1981, 123, 49–59. [Google Scholar] [CrossRef][Green Version]
- Varga, M.; Kuntová, L.; Pantůček, R.; Mašlaňová, I.; Růžičková, V.; Doškař, J. Efficient transfer of antibiotic re-sistance plasmids by transduction within methicillin-resistant Staphylococcus aureus USA300 clone. FEMS Microbiol. Lett. 2012, 332, 146–152. [Google Scholar] [CrossRef]
- Volkova, V.V.; Lu, Z.; Besser, T.; Gröhn, Y.T. Modeling the infection dynamics of bacteriophages in enteric Escherichia coli: Estimating the contribution of transduction to antimicrobial gene spread. Appl. Environ. Microbiol. 2014, 80, 4350–4362. [Google Scholar] [CrossRef]
- Kakasis, A.; Panitsa, G. Bacteriophage therapy as an alternative treatment for human infections. A comprehensive review. Int. J. Antimicrob. Agents 2019, 53, 16–21. [Google Scholar] [CrossRef] [PubMed]
- Vlassov, V.V.; Laktionov, P.P.; Rykova, E.Y. Extracellular nucleic acids. BioEssays 2007, 29, 654–667. [Google Scholar] [CrossRef] [PubMed]
- Pietramellara, G.; Ascher-Jenull, J.; Borgogni, F.; Ceccherini, M.T.; Guerri, G.; Nannipieri, P. Extracellular DNA in soil and sediment: Fate and ecological relevance. Biol. Fertil. Soils 2009, 45, 219–235. [Google Scholar] [CrossRef]
- Stewart, G.J.; Sinigalliano, C.D. Detection of Horizontal Gene Transfer by Natural Transformation in Native and Introduced Species of Bacteria in Marine and Synthetic Sediments. Appl. Environ. Microbiol. 1990, 56, 1818–1824. [Google Scholar] [CrossRef]
- Dubnau, D. DNA Uptake in Bacteria. Annu. Rev. Microbiol. 1999, 53, 217–244. [Google Scholar] [CrossRef] [PubMed]
- Johnsborg, O.; Eldholm, V.; Håvarstein, L.S. Natural genetic transformation: Prevalence, mechanisms and function. Res. Microbiol. 2007, 158, 767–778. [Google Scholar] [CrossRef] [PubMed]
- Palmen, R.; Hellingwerf, K.J. Uptake and processing of DNA by Acinetobacter calcoaceticus—A review. Gene 1997, 192, 179–190. [Google Scholar] [CrossRef]
- Zawadzki, P.; Cohan, F.M. The size and continuity of DNA segments integrated in Bacillus transformation. Genetics 1995, 141, 1231–1243. [Google Scholar] [CrossRef]
- Tello, A.; Austin, B.; Telfer, T. Selective Pressure of Antibiotic Pollution on Bacteria of Importance to Public Health. Environ. Health Perspect. 2012, 120, 1100–1106. [Google Scholar] [CrossRef] [PubMed]
- Pham, T.D.M.; Ziora, Z.M.; Blaskovich, M.A.T. Quinolone antibiotics. MedChemComm 2019, 10, 1719–1739. [Google Scholar] [CrossRef]
- Cattoir, V.; Nordmann, P. Plasmid-mediated quinolone resistance in gram-negative bacterial species: An update. Curr. Med. Chem. 2009, 16, 1028–1046. [Google Scholar] [CrossRef]
- Cattoir, V.; Poirel, L.; Aubert, C.; Soussy, C.-J.; Nordmann, P. Unexpected Occurrence of Plasmid-mediated Quinolone Resistance Determinants in Environmental Aeromonas spp. Emerg. Infect. Dis. 2008, 14, 231. [Google Scholar] [CrossRef]
- Tomova, A.; Ivanova, L.; Buschmann, A.H.; Godfrey, H.P.; Cabello, F.C. Plasmid-Mediated Quinolone Resistance (PMQR) Genes and Class 1 Integrons in Quinolone-Resistant Marine Bacteria and Clinical Isolates of Escherichia coli from an Aquacultural Area. Microb. Ecol. 2017, 75, 104–112. [Google Scholar] [CrossRef]
- Yang, H.; Duan, G.; Zhu, J.; Zhang, W.; Xi, Y.; Fan, Q. Prevalence and characterisation of plasmid-mediated quinolone resistance and mutations in the gyrase and topoisomerase IV genes among Shigella isolates from Henan, China, between 2001 and 2008. Int. J. Antimicrob. Agents 2013, 42, 173–177. [Google Scholar] [CrossRef]
- Poirel, L.; Cattoir, V.; Nordmann, P. Plasmid-Mediated Quinolone Resistance; Interactions between Human, Animal, and Environmental Ecologies. Front. Microbiol. 2012, 3, 24. [Google Scholar] [CrossRef] [PubMed]
- Eftekhar, F.; Seyedpour, S.M. Prevalence of qnr and aac(6′)-Ib-cr Genes in Clinical Isolates of Klebsiella pneumoni-ae from Imam Hussein Hospital in Tehran. Iran. J. Med. Sci. 2015, 40, 6. [Google Scholar]
- Khajanchi, B.K.; Fadl, A.A.; Borchardt, M.A.; Berg, R.L.; Horneman, A.J.; Stemper, M.E.; Joseph, S.W.; Moyer, N.P.; Sha, J.; Chopra, A.K. Distribution of Virulence Factors and Molecular Fingerprinting of Aeromonas Species Isolates from Water and Clinical Samples: Suggestive Evidence of Water-to-Human Transmission. Appl. Environ. Microbiol. 2010, 76, 2313–2325. [Google Scholar] [CrossRef] [PubMed]
- Sánchez-Céspedes, J.; Blasco, M.D.; Marti, S.; Alcalde, E.; Vila, J.; Alba, V.; Esteve, C. Plasmid-Mediated QnrS2 Determinant from a Clinical Aeromonas veronii Isolate. Antimicrob. Agents Chemother. 2008, 52, 2990–2991. [Google Scholar] [CrossRef][Green Version]
- Lavilla, S.; Gonzalez-Lopez, J.J.; Sabate, M.; Garcia-Fernandez, A.; Larrosa, M.N.; Bartolome, R.M.; Carattoli, A.; Prats, G. Prevalence of qnr genes among extended-spectrum β-lactamase-producing enterobacterial isolates in Barcelona, Spain. J. Antimicrob. Chemother. 2008, 61, 291–295. [Google Scholar] [CrossRef]
- Park, Y.J.; Yu, J.K.; Lee, S.; Oh, E.J.; Woo, G.J. Prevalence and diversity of qnr alleles in AmpC-producing Enterobacter cloacae, Enterobacter aerogenes, Citrobacter freundii and Serratia marcescens: A multicentre study from Korea. J. Antimicrob. Chemother. 2007, 60, 868–871. [Google Scholar] [CrossRef] [PubMed]
- Heuer, O.E.; Kruse, H.; Grave, K.; Collignon, P.; Karunasagar, I.; Angulo, F.J. Human Health Consequences of Use of Antimicrobial Agents in Aquaculture. Clin. Infect. Dis. 2009, 49, 1248–1253. [Google Scholar] [CrossRef]
- Wen, Y.; Pu, X.; Zheng, W.; Hu, G. High Prevalence of Plasmid-Mediated Quinolone Resistance and IncQ Plasmids Carrying qnrS2 Gene in Bacteria from Rivers near Hospitals and Aquaculture in China. PLoS ONE 2016, 11, e0159418. [Google Scholar] [CrossRef]
- Poirel, L.; Rodriguez-Martinez, J.M.; Mammeri, H.; Liard, A.; Nordmann, P. Origin of plasmid-mediated quino-lone resistance determinant QnrA. Antimicrob. Agents Chemother. 2005, 49, 3523–3525. [Google Scholar] [CrossRef] [PubMed]
- Cattoir, V.; Poirel, L.; Mazel, D.; Soussy, C.J.; Nordmann, P. Vibrio splendidus as the source of plasmid-mediated QnrS-Like quinolone resistance determinants. Antimicrob. Agents Chemother. 2007, 51, 2650–2651. [Google Scholar] [CrossRef]
- World Health Organization. Antimicrobial Resistance: Global Report on Surveillance 2014; WHO: Geneva, Switzerland, 2014; Available online: http://apps.who.int/iris/bitstream/10665/112642/1/9789241564748_eng.pdf (accessed on 11 February 2021).
- The Center for Disease, Dynamics Economics and Policy. Resistance Map: Antibiotic Resistance. 2021. Available online: https://resistancemap.cddep.org/AntibioticResistance.php (accessed on 11 February 2021).
- Chopra, I.; Roberts, M. Tetracycline Antibiotics: Mode of Action, Applications, Molecular Biology, and Epidemiology of Bacterial Resistance. Microbiol. Mol. Biol. Rev. 2001, 65, 232–260. [Google Scholar] [CrossRef]
- Speer, B.S.; Shoemaker, N.B.; Salyers, A.A. Bacterial Resistance to Tetracycline: Mechanisms, Transfer, and Clinical Significance. Clin. Microbiol. Rev. 1992, 5, 387–399. [Google Scholar] [CrossRef]
- Furushita, M.; Shiba, T.; Maeda, T.; Yahata, M.; Kaneoka, A.; Takahashi, Y.; Torii, K.; Hasegawa, T.; Ohta, M. Similarity of tetracycline resistance genes isolated from fish farm bacteria to those from clinical isolates. Appl. Environ. Microbiol. 2003, 69, 5336–5342. [Google Scholar] [CrossRef] [PubMed]
- Rhodes, G.; Huys, G.; Swings, J.; McGann, P.; Hiney, M.; Smith, P. Distribution of oxytetracycline re-sistance plasmids between Aeromonads in hospital and aquaculture environments: Implication of Tn1721 in dis-semination of the tetracycline resistance determinant Tet A. Appl. Environ. Microbiol. 2000, 66, 3883–3890. [Google Scholar] [CrossRef] [PubMed]
- Briggs, C.E.; Fratamico, P.M. Molecular Characterization of an Antibiotic Resistance Gene Cluster of Salmonella typhimuriumDT104. Antimicrob. Agents Chemother. 1999, 43, 846–849. [Google Scholar] [CrossRef]
- Angulo, F.J.; Johnson, K.R.; Tauxe, R.V.; Cohen, M.L. Origins and consequences of antimicrobial-resistant non-typhoidal Salmonella: Implications for the use of fluoroquinolones in food animals. Microb. Drug Resist. 2000, 6, 77–183. [Google Scholar] [CrossRef]
- Angulo, F.J.; Griffin, P.M. Changes in Antimicrobial Resistance in Salmonella enterica Serovar Typhimurium. Emerg. Infect. Dis. 2000, 6, 436–437. [Google Scholar] [CrossRef] [PubMed]
- Cabello, F.C.; Godfrey, H.P.; Buschmann, A.H.; Dölz, H.J. Aquaculture as yet another environmental gateway to the development and globalisation of antimicrobial resistance. Lancet Infect. Dis. 2016, 16, e127–e133. [Google Scholar] [CrossRef]
- Sousa, M.; Torres, C.; Barros, J.; Somalo, S.; Igrejas, G.; Poeta, P. Gilthead Seabream (Sparus aurata) as Carriers of SHV-12 and TEM-52 Extended-Spectrum Beta-Lactamases-Containing Escherichia coli Isolates. Foodborne Pathog. Dis. 2011, 8, 1139–1141. [Google Scholar] [CrossRef]
- Eckburg, P.B.; Lister, T.; Walpole, S.; Keutzer, T.; Utley, L.; Tomayko, J.; Kopp, E.; Farinola, N.; Coleman, S. Safety, Tolerability, Pharmacokinetics, and Drug Interaction Potential of SPR741, an Intravenous Potentiator, after Single and Multiple Ascending Doses and When Combined with β-Lactam Antibiotics in Healthy Subjects. Antimicrob. Agents Chemother. 2019, 63, e00892-19. [Google Scholar] [CrossRef]
- Ibrahim, M.E.; Abbas, M.; Al-Shahrai, A.M.; Elamin, B.K. Phenotypic Characterization and Antibiotic Re-sistance Patterns of Extended-Spectrum β-Lactamase- and AmpC β-Lactamase-Producing Gram-Negative Bacteria in a Referral Hospital, Saudi Arabia. Can. J. Infect. Dis. Med. Microbiol. 2019, 6054694. [Google Scholar] [CrossRef]
- Liu, Y.-Y.; Wang, Y.; Walsh, T.; Yi, L.-X.; Zhang, R.; Spencer, J.; Doi, Y.; Tian, G.; Dong, B.; Huang, X.; et al. Emergence of plasmid-mediated colistin resistance mechanism MCR-1 in animals and human beings in China: A microbiological and molecular biological study. Lancet Infect. Dis. 2016, 16, 161–168. [Google Scholar] [CrossRef]
- Telke, A.A.; Rolain, J.-M. Functional genomics to discover antibiotic resistance genes: The paradigm of resistance to colistin mediated by ethanolamine phosphotransferase in Shewanella algae MARS 14. Int. J. Antimicrob. Agents 2015, 46, 648–652. [Google Scholar] [CrossRef] [PubMed]
- Tomova, A.; Ivanova, L.; Buschmann, A.H.; Rioseco, M.L.; Kalsi, R.K.; Godfrey, H.P.; Cabello, F.C. Antimicrobial resistance genes in marine bacteria and human uropathogenic Escherichia coli from a region of intensive aquaculture. Environ. Microbiol. Rep. 2015, 7, 803–809. [Google Scholar] [CrossRef]
- Bills, G.F.; Gloer, J.B. Biologically active secondary metabolites from the fungi. Fungal Kingd. 2017, 1087–1119. [Google Scholar] [CrossRef]
- Bolton, L.F.; Kelley, L.C.; Lee, M.D.; Fedorka-Cray, P.J.; Maurer, J.J. Detection of multidrug-resistant Salmonella enterica serotype Typhimurium DT104 based on a gene which confers cross-resistance to florfenicol and chloram-phenicol. J. Clin. Microbiol. 1999, 37, 1348–1351. [Google Scholar] [CrossRef]
- Boyd, D.; Peters, G.A.; Cloeckaert, A.; Boumedine, K.S.; Chaslus-Dancla, E.; Imberechts, H.; Mulvey, M.R. Complete nucleotide sequence of a 43-kilobase genomic island associated with the multidrug resistance region of Salmonella enterica serovar Typhimurium DT104 and its identification in phage type DT120 and serovar Agona. J. Bacteriol. 2001, 183, 5725–5732. [Google Scholar] [CrossRef]
- Clark, G.M.; Kaufmann, A.F.; Gangarosa, E.J. Epidemiology of an international outbreak of Salmonella Agona. Lancet 1973, 2, 490–493. [Google Scholar] [CrossRef]
- Weber, J.T.; Mintz, E.D.; Canizares, R.; Semiglia, A.; Gomez, I.; Sempertegui, R.; Davila, A.; Greene, K.D.; Puhr, N.D.; Cameron, D.N.; et al. Epidemic cholera in Ecuador: Multidrug-resistance and transmission by water and seafood. Epidemiol. Infect. 1994, 112, 1–11. [Google Scholar] [CrossRef][Green Version]
- Nogales, B.; Lanfranconi, M.P.; Piña-Villalonga, J.M.; Bosch, R. Anthropogenic perturbations in marine microbial communities. FEMS Microbiol. Rev. 2011, 35, 275–298. [Google Scholar] [CrossRef] [PubMed]
- Pridgeon, J.W.; Klesius, P.H. Major bacterial diseases in aquaculture and their vaccine development. Cab Rev. 2012, 7, 1–16. [Google Scholar] [CrossRef]
- Chelossi, E.; Vezzulli, L.; Milano, A.; Branzoni, M.; Fabiano, M.; Riccardi, G.; Banat, I.M. Antibiotic resistance of benthic bacteria in fish-farm and control sediments of the Western Mediterranean. Aquaculture 2003, 219, 83–97. [Google Scholar] [CrossRef]
- Bakopoulos, V.; Adams, A.; Richards, R.H. Some biochemical properties and antibiotic sensitivities of Pasteurel-la piscicida isolated in Greece and comparison with strains from Japan, France and Italy. J. Fish Dis. 1995, 18, 1–7. [Google Scholar] [CrossRef]
- Zorilla, I.; Chabrillon, M.; Arijo, S.; Diaz-Rosales, P.; Martinez-Manzanares, E.; Babelona, M.; Morinigo, M. Bacteria recovered from diseased cultured gilthead seabream (Sparus aurata) in southwestern Spain. Aquaculture 2003, 218, 11–20. [Google Scholar] [CrossRef]
- Smith, P.; Christofilogiannis, P. Application of Normalised Resistance Interpretation to the detection of multiple low-level resistance in strains of Vibrio anguillarum obtained from Greek fish farms. Aquaculture 2007, 272, 223–230. [Google Scholar] [CrossRef]
- Labella, A.; Gennari, M.; Ghidini, V.; Trento, I.; Manfrin, A.; Borrego, J.; Lleo, M. High incidence of antibiotic multi-resistant bacteria in coastal areas dedicated to fish farming. Mar. Pollut. Bull. 2013, 70, 197–203. [Google Scholar] [CrossRef] [PubMed]
- Elmahdi, S.; DaSilva, L.V.; Parveen, S. Antibiotic resistance of Vibrio parahaemolyticus and Vibrio vulnificus in various countries: A review. Food Microbiol. 2016, 57, 128–134. [Google Scholar] [CrossRef]
- Ottaviani, D.; Bacchiocchi, I.; Masini, L.; Francesca, L.; Carraturo, A.; Giammarioli, M.; Sbaraglia, G. Antimicrobial susceptibility of potentially pathogenic halophilic vibrios isolated from seafood. Int. J. Antimicrob. Agents 2001, 18, 135–140. [Google Scholar] [CrossRef]
- Scarano, C.; Piras, F.; Virdis, S.; Ziino, G.; Nuvoloni, R.; Dalmasso, A.; De Santis, E.; Spanu, C. Antibiotic resistance of Aeromonas ssp. strains isolated from Sparus aurata reared in Italian mariculture farms. Int. J. Food Microbiol. 2018, 284, 91–97. [Google Scholar] [CrossRef]
- Salgueiro, V.; Manageiro, V.; Bandarra, N.M.; Reis, L.; Ferreira, E.; Caniça, M. Bacterial Diversity and Antibiotic Susceptibility of Sparus aurata from Aquaculture. Microorganisms 2020, 8, 1343. [Google Scholar] [CrossRef]
- Nikolakopoulou, T.L.; Giannoutsou, E.P.; Karabatsou, A.A.; Karagouni, A.D. Prevalence of tetracycline re-sistance genes in Greek seawater habitats. J. Microbiol. 2008, 46, 633. [Google Scholar] [CrossRef] [PubMed]
- Allen, H.K.; Donato, J.; Wang, H.H.; Cloud-Hansen, K.A.; Davies, J.; Handelsman, J. Call of the wild: Antibiotic resistance genes in natural environments. Nat. Rev. Genet. 2010, 8, 251–259. [Google Scholar] [CrossRef]
- Sutili, F.J.; Gressler, L.T. Chapter 3—Antimicrobial agents. In Aquaculture Pharmacology, 1st ed.; Kibenge, F.S.B., Baldisserotto, B., Chong, R.S.-M., Eds.; Elsevier: Amsterdam, The Netherlands, 2020; pp. 131–196. [Google Scholar]
- Cassini, A. Attributable deaths and disability-adjusted life-years caused by infections with antibiotic-resistant bacteria in the EU and the European Economic Area in 2015: A population-level modelling analysis. Lancet Infect. Dis. 2019, 19, 56–66. [Google Scholar] [CrossRef]
- Reverter, M.; Sarter, S.; Caruso, D.; Avarre, J.-C.; Combe, M.; Pepey, E.; Pouyaud, L.; Vega-Heredía, S.; De Verdal, H.; Gozlan, R.E. Aquaculture at the crossroads of global warming and antimicrobial resistance. Nat. Commun. 2020, 11, 1–8. [Google Scholar] [CrossRef]
- Van Boeckel, T.P.; Glennon, E.E.; Chen, D.; Gilbert, M.; Robinson, T.P.; Grenfell, B.B.; Levin, S.A.S.; Bonhoeffer, S.; Laxminarayan, R.R. Reducing antimicrobial use in food animals. Science 2017, 357, 1350–1352. [Google Scholar] [CrossRef]
- Arab, S.; Nalbone, L.; Giarratana, F.; Berbar, A. Occurrence of Vibrio spp. along the Algerian Mediterranean coast in wild and farmed Sparus aurata and Dicentrarchus labrax. Vet. World 2020, 13, 1199–1208. [Google Scholar] [CrossRef]
- Kolda, A.; Mujakić, I.; Perić, L.; Vardić Smrzlić, I.; Kapetanović, D. Microbiological Quality Assessment of Water and Fish from Karst Rivers of the Southeast Black Sea Basin (Croatia), and Antimicrobial Susceptibility of Aeromonas Isolates. Curr. Microbiol. 2020, 77, 2322–2332. [Google Scholar] [CrossRef]
- Kapetanović, D.; Vardić, S.; Valić, D.; Teskeredžić Emin, E. Occurrence, characterization and antimicrobial susceptibility of Vibrio alginolyticus in the Eastern Adriatic Sea. Mar. Pollut. Bull. 2013, 75, 46–52. [Google Scholar]
- El-Gohary, F.A.; Zahran, E.; El-Gawad, E.A.A.; El-Gohary, A.H.; Abdelhamid, F.M.; El-Mleeh, A.; Elmahallawy, E.K.; Elsayed, M.M. Investigation of the Prevalence, Virulence Genes, and Antibiogram of Motile Aeromonads Isolated from Nile Tilapia Fish Farms in Egypt and Assessment of their Water Quality. Animals 2020, 10, 1432. [Google Scholar] [CrossRef] [PubMed]
- Fadel, A.; Mabrok, M.; Aly, S. Epizootics of Pseudomonas anguilliseptica among cultured seabream (Sparus aurata) populations: Control and treatment strategies. Microb. Pathog. 2018, 121, 1–8. [Google Scholar] [CrossRef]
- Hamouda, A.H.; Moustafa, E.M.; Zayed, M.M. Overview on the most prevailing bacterial diseases infecting Oreochromis niloticus at Aswan fish hatchery, Egypt. Adv. Anim. Vet. Sci. 2019, 7, 950–961. [Google Scholar] [CrossRef]
- Hamza, D.; Dorgham, S.; Ismael, E.; El-Moez, S.I.A.; ElHariri, M.; Elhelw, R.; Hamza, E. Emergence of β-lactamase- and carbapenemase- producing Enterobacteriaceae at integrated fish farms. Antimicrob. Resist. Infect. Control. 2020, 9, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Calvez, S.; Navarro-Gonzalez, N.; Siekoula-Nguedia, C.; Fournel, C.; Duchaud, E. High Genetic Diversity in Flavobacterium psychrophilum Isolates from Healthy Rainbow Trout (Oncorhynchus mykiss) Farmed in the Same Watershed, Revealed by Two Typing Methods. Appl. Environ. Microbiol. 2021, 87. [Google Scholar] [CrossRef]
- Moreau, E.; Thomas, T.; Brevet, M.; Thorin, C.; Fournel, C.; Calvez, S. Mutations involved in the emergence of Yersinia ruckeri biotype 2 in France. Transbound. Emerg. Dis. 2019, 66, 1387–1394. [Google Scholar] [CrossRef] [PubMed]
- Di Cesare, A.; Luna, G.M.; Vignaroli, C.; Pasquaroli, S.; Tota, S.; Paroncini, P.; Biavasco, F. Aquaculture Can Promote the Presence and Spread of Antibiotic-Resistant Enterococci in Marine Sediments. PLoS ONE 2013, 8, e62838. [Google Scholar] [CrossRef]
- Laganà, P.; Caruso, G.; Minutoli, E.; Zaccone, R.; Santi, D. Susceptibility to antibiotics of Vibrio spp. and Photobacterium damsela ssp. piscicida strains isolated from Italian aquaculture farms. New Microbiol. 2011, 34, 53–63. [Google Scholar] [PubMed]
- Harakeh, S.; Yassine, H.; El-Fadel, M. Antimicrobial-resistance of Streptococcus pneumoniae isolated from the Lebanese environment. Mar. Environ. Res. 2006, 62, 181–193. [Google Scholar] [CrossRef]
- Alcaide, E.; Blasco, M.-D.; Esteve, C. Occurrence of Drug-Resistant Bacteria in Two European Eel Farms. Appl. Environ. Microbiol. 2005, 71, 3348–3350. [Google Scholar] [CrossRef] [PubMed]
- Del Cerro, A.; Márquez, I.; Prieto, J.M. Genetic diversity and antimicrobial resistance of Flavobacterium psychrophilum isolated from cultured rainbow trout, Onchorynchus mykiss (Walbaum), in Spain. J. Fish Dis. 2010, 33, 285–291. [Google Scholar] [CrossRef]
- Ortega, C.; Gimeno, O.; Blanc, V.; Cortés, P.; Anía, S.; Llagostera, M. Antibiotic susceptibility of strains of Aeromonas salmonicida isolated from Spanish salmonids. Rev. Med. Vet. 2006, 157, 410–414. [Google Scholar]
- Lajnef, R.; Snoussi, M.; López Romalde, J.; Nozha, C.; Hassen, A. Comparative study on the antibiotic susceptibility and plasmid profiles of Vibrio alginolyticus strains isolated from four Tunisian marine biotipes. World J. Microbiol. Biotechnol. 2012, 28, 3345–3363. [Google Scholar] [CrossRef] [PubMed]
- Ben Said, L.; Hamdaoui, M.; Jouini, A.; Boudabous, A.; Ben Slama, K.; Torres, C.; Klibi, N. First Detection of CTX-M-1 in Extended-Spectrum β-Lactamase–Producing Escherichia coli in Seafood from Tunisia. J. Food Prot. 2017, 80, 1877–1881. [Google Scholar] [CrossRef]
- Capkin, E.; Terzi, E.; Altinok, I. Occurrence of antibiotic resistance genes in culturable bacteria isolated from Turkish trout farms and their local aquatic environment. Dis. Aquat. Org. 2015, 114, 127–137. [Google Scholar] [CrossRef] [PubMed]
- Duman, M.; Altun, S.; Cengiz, M.; Saticioglu, I.B.; Buyukekiz, A.G.; Sahinturk, P. Genotyping and antimicrobial resistance genes of Yersinia ruckeri isolates from rainbow trout farms. Dis. Aquat. Org. 2017, 125, 31–44. [Google Scholar] [CrossRef] [PubMed]
- Onuk, E.E.; Didinen, B.I.; Giftci, A.; Yardimci, B.; Pekmezci, G.Z. Molecular characterisation of antibiotic resistance in Yersinia ruckeri isolates from Turkey. Bull. Eur. Assoc. Fish Pathol. 2019, 39, 145–155. [Google Scholar]
- Povraz, N.; Flu, M.B.M. Antibiotic resistant bacteria profile and microbial community structure of trout farming ponds in Bielecki, Turkey. Fresenius. Environ. Bull. 2020, 29, 9164–9174. [Google Scholar]
- McGough, S.F.; MacFadden, D.R.; Hattab, M.W.; Mølbak, K.; Santillana, M. Rates of increase of antibiotic resistance and ambient temperature in Europe: A cross-national analysis of 28 countries between 2000 and 2016. Eurosurveillance 2000, 25, 1900414. [Google Scholar] [CrossRef]
- Greulich, P.; Scott, M.; Evans, M.R.; Allen, R.J. Growth-dependent bacterial susceptibility to ribosome-targeting antibiotics. Mol. Syst. Biol. 2015, 11, 796. [Google Scholar] [CrossRef] [PubMed]
- Ramirez, M.S.; Tolmasky, M.E. Aminoglycoside modifying enzymes. Drug Resist. Updat. 2010, 13, 151–171. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.T.; Connelly, M.B.; Amolo, C.; Otani, S.; Yaver, D.S. Global Transcriptional Response of Bacillus subtilis to Treatment with Subinhibitory Concentrations of Antibiotics That Inhibit Protein Synthesis. Antimicrob. Agents Chemother. 2005, 49, 1915–1926. [Google Scholar] [CrossRef] [PubMed]
- Kindrachuk, K.N.; Fernández, L.; Bains, M.; Hancock, R.E.W. Involvement of an ATP-Dependent Protease, PA0779/AsrA, in Inducing Heat Shock in Response to Tobramycin in Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 2011, 55, 1874–1882. [Google Scholar] [CrossRef] [PubMed]
- Kohanski, M.A.; Dwyer, D.J.; Wierzbowski, J.; Cottarel, G.; Collins, J.J. Mistranslation of Membrane Proteins and Two-Component System Activation Trigger Antibiotic-Mediated Cell Death. Cell 2008, 135, 679–690. [Google Scholar] [CrossRef] [PubMed]
- Van Bogelen, R.A.; Neidhardt, F.C. Ribosomes as sensors of heat and cold shock in Escherichia coli. Proc. Natl. Acad. Sci. USA 1990, 87, 5589–5593. [Google Scholar] [CrossRef] [PubMed]
- Cruz-Loya, M.; Kang, T.M.; Lozano, N.A.; Watanabe, R.; Tekin, E.; Damoiseaux, R.; Savage, V.M.; Yeh, P.J. Stressor interaction networks suggest antibiotic resistance co-opted from stress responses to temperature. ISME J. 2019, 13, 12–23. [Google Scholar] [CrossRef] [PubMed]
- Cardoso, K.; Gandra, R.F.; Wisniewski, E.S.; Osaku, C.A.; Kadowaki, M.K.; Felipach-Neto, V.; Haus, L.F.A.-A.; Simão, R.D.C.G. DnaK and GroEL are induced in response to antibiotic and heat shock in Acinetobacter baumannii. J. Med. Microbiol. 2010, 59, 1061–1068. [Google Scholar] [CrossRef]
- Poole, K. Stress responses as determinants of antimicrobial resistance in Gram-negative bacteria. Trends Microbiol. 2012, 20, 227–234. [Google Scholar] [CrossRef] [PubMed]
- Monem, S.; Furmanek-Blaszk, B.; Łupkowska, A.; Kuczyńska-Wiśnik, D.; Stojowska-Swędrzyńska, K.; Laskowska, E. Mechanisms Protecting Acinetobacter baumannii against Multiple Stresses Triggered by the Host Immune Response, Antibiotics, and outside Host Environment. Int. J. Mol. Sci. 2020, 21, 5498. [Google Scholar] [CrossRef]
- Goltermann, L.; Good, L.; Bentin, T. Chaperonins fight aminoglycoside-induced protein misfolding and pro-mote short term tolerance in Escherichia coli. J. Biol. Chem. 2013, 288, 10483–10489. [Google Scholar] [CrossRef] [PubMed]
- Murata, M.; Fujimoto, H.; Nishimura, K.; Charoensuk, K.; Nagamitsu, H.; Raina, S.; Kosaka, T.; Oshima, T.; Ogasawara, N.; Yamada, M. Molecular Strategy for Survival at a Critical High Temperature in Eschierichia coli. PLoS ONE 2011, 6, e20063. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Verdugo, A.; Lozano-Huntelman, N.; Cruz-Loya, M.; Savage, V.; Yeh, P. Compounding Effects of Climate Warming and Antibiotic Resistance. iScience 2020, 23, 101024. [Google Scholar] [CrossRef]
- Rodríguez-Verdugo, A.; Gaut, B.S.; Tenaillon, O. Evolution of Escherichia coli rifampicin resistance in an antibiotic-free environment during thermal stress. BMC Evol. Biol. 2013, 13, 50. [Google Scholar] [CrossRef] [PubMed]
- Dragosits, M.; Mozhayskiy, V.; Quinones-Soto, S.; Park, J.; Tagkopoulos, I. Evolutionary potential, cross-stress behavior and the genetic basis of acquired stress resistance in Escherichia coli. Mol. Syst. Biol. 2013, 9, 643. [Google Scholar] [CrossRef] [PubMed]
- Machielsen, R.; Van Alen-Boerrigter, I.J.; Koole, L.A.; Bongers, R.S.; Kleerebezem, M.; Vlieg, J.E.T.V.H. Indigenous and Environmental Modulation of Frequencies of Mutation in Lactobacillus plantarum. Appl. Environ. Microbiol. 2009, 76, 1587–1595. [Google Scholar] [CrossRef]
- Rodríguez-Verdugo, A.; Tenaillon, O.; Gaut, B.S. First-Step Mutations during Adaptation Restore the Expression of Hundreds of Genes. Mol. Biol. Evol. 2016, 33, 25–39. [Google Scholar] [CrossRef] [PubMed]
- Walsh, T.R.; Weeks, J.; Livermore, D.M.; Toleman, M.A. Dissemination of NDM-1 positive bacteria in the New Delhi environment and its implications for human health: An environmental point prevalence study. Lancet Infect. Dis. 2011, 11, 355–362. [Google Scholar] [CrossRef]
- Kent, A.G.; Garcia, C.A.; Martiny, A.C. Increased biofilm formation due to high temperature adaptation in marine Roseobacter. Nat. Microbiol. 2018, 3, 989–995. [Google Scholar] [CrossRef] [PubMed]
- Payne, J.L.; Wagner, A. The causes of evolvability and their evolution. Nat. Rev. Genet. 2019, 20, 24–38. [Google Scholar] [CrossRef]
- Foster, P.L. Stress-Induced Mutagenesis in Bacteria. Crit. Rev. Biochem. Mol. Biol. 2007, 42, 373–397. [Google Scholar] [CrossRef] [PubMed]
- MacLean, R.C.; Bell, G.; Rainey, P.B. The evolution of a pleiotropic fitness tradeoff in Pseudomonas fluorescens. Proc. Natl. Acad. Sci. USA 2004, 101, 8072–8077. [Google Scholar] [CrossRef] [PubMed]
- Andersson, D.I.; Hughes, D. Antibiotic resistance and its cost: Is it possible to reverse resistance? Nat. Rev. Genet. 2010, 8, 260–271. [Google Scholar] [CrossRef]
- Knies, J.L.; Cai, F.; Weinreich, D.M. Enzyme efficiency but not thermostability drives cefotaxime resistance evolution in TEM-1 ßlactamase. Mol. Biol. Evol. 2017, 34, 1040–1054. [Google Scholar] [PubMed]
- Trindade, S.; Sousa, A.; Gordo, I. Antibiotic resistance and stress in the light of fisher’s model. Evolution 2012, 66, 3815–3824. [Google Scholar] [CrossRef]
- Chu, X.; Zhang, D.; Buckling, A.; Zhang, Q. Warmer temperatures enhance beneficial mutation effects. J. Evol. Biol. 2020, 33, 1020–1027. [Google Scholar] [CrossRef] [PubMed]
- Schwartzman, D.; Lineweaver, C. The hyperthermophilic origin of life revisited. Biochem. Soc. Trans. 2004, 32, 168–171. [Google Scholar] [CrossRef] [PubMed]
- Åkerfelt, M.; Morimoto, R.I.; Sistonen, L. Heat shock factors: Integrators of cell stress, development and lifespan. Nat. Rev. Mol. Cell Biol. 2010, 11, 545–555. [Google Scholar] [CrossRef]
- D’Costa, V.M.; King, C.E.; Kalan, L.; Morar, M.; Sung, W.; Schwarz, C.; Froese, D.G.; Zazula, G.D.; Calmels, F.; Debruyne, R.; et al. Antibiotic resistance is ancient. Nature 2011, 477, 457–461. [Google Scholar] [CrossRef]
- Mondal, S.; Pathak, B.K.; Ray, S.; Barat, C. Impact of P-Site tRNA and Antibiotics on Ribosome Mediated Protein Folding: Studies Using the Escherichia coli Ribosome. PLoS ONE 2014, 9, e101293. [Google Scholar] [CrossRef]
- Vabulas, R.M.; Raychaudhuri, S.; Hayer-Hartl, M.; Hartl, F.U. Protein Folding in the Cytoplasm and the Heat Shock Response. Cold Spring Harb. Perspect. Biol. 2010, 2, a004390. [Google Scholar] [CrossRef] [PubMed]
- Völker, U.; Mach, H.; Schmid, R.; Hecker, M. Stress proteins and cross-protection by heat shock and salt stress in Bacillus subtilis. J. Gen. Microbiol. 1992, 138, 2125–2135. [Google Scholar] [CrossRef]
- Richter, K.; Haslbeck, M.; Buchner, J. The Heat Shock Response: Life on the Verge of Death. Mol. Cell 2010, 40, 253–266. [Google Scholar] [CrossRef]
- Gunasekera, T.S.; Csonka, L.N.; Paliy, O. Genome-wide transcriptional responses of Escherichia coli K-12 to continuous osmotic and heat stresses. J. Bacteriol. 2008, 190, 3712–3720. [Google Scholar] [CrossRef] [PubMed]
- Blount, Z.D.; Barrick, J.E.; Davidson, C.J.; Lenski, R.E. Genomic analysis of a key innovation in an experimental Escherichia coli population. Nature 2012, 489, 513–518. [Google Scholar] [CrossRef]
- Quandt, E.M.; Deatherage, D.E.; Ellington, A.D.; Georgiou, G.; Barrick, J.E. Recursive genome wide recombina-tion and sequencing reveals a key refinement step in the evolution of a metabolic innovation in Escherichia coli. Proc. Natl. Acad. Sci. USA 2014, 111, 2217–2222. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, D.; Veeraraghavan, B.; Elangovan, R.; Perumal, V. Antibiotic Resistance and Epigenetics: More to It than Meets the Eye. Antimicrob. Agents Chemother. 2020, 64. [Google Scholar] [CrossRef]
- Adam, M.; Murali, B.; Glenn, N.O.; Potter, S.S. Epigenetic inheritance based evolution of antibiotic resistance in bacteria. BMC Evol. Biol. 2008, 8, 52. [Google Scholar] [CrossRef] [PubMed]
- Herrel, A.; Joly, D.; Danchin, E. Epigenetics in ecology and evolution. Funct. Ecol. 2019, 34, 381–384. [Google Scholar] [CrossRef]
- O’Dea, R.; Noble, D.W.A.; Johnson, S.L.; Hesselson, D.; Nakagawa, S. The role of non-genetic inheritance in evolutionary rescue: Epigenetic buffering, heritable bet hedging and epigenetic traps. Environ. Epigenetics 2016, 2, dvv014. [Google Scholar] [CrossRef]
- Jaenisch, R.; Bird, A. Epigenetic regulation of gene expression: How the genome integrates intrinsic and environmental signals. Nat. Genet. 2003, 33, 245–254. [Google Scholar] [CrossRef]
- Moore, L.D.; Le, T.; Fan, G. DNA Methylation and Its Basic Function. Neuropsychopharmacology 2013, 38, 23–38. [Google Scholar] [CrossRef]
- Huss, R.A.; Davidson, S.; Militello, K.T. DNA cytosine methyltransferase increases stationary phase fitness un-der temperature stress in Escherichia coli. FASEB J. 2016, 30, 581.2. [Google Scholar]
- Blum, P.; Payne, S. Evidence of an Epigenetics System in Archaea. Epigenetics Insights 2019, 12, 251686571986528. [Google Scholar] [CrossRef] [PubMed]
- McCaw, B.A.; Stevenson, T.J.; Lancaster, L.T. Epigenetic Responses to Temperature and Climate. Integr. Comp. Biol. 2020, 60, 1469–1480. [Google Scholar] [CrossRef] [PubMed]
- MacFadden, D.R.; McGough, S.F.; Fisman, D.; Santillana, M.; Brownstein, J.S. Antibiotic resistance increases with local temperature. Nat. Clim. Chang. 2018, 8, 510–514. [Google Scholar] [CrossRef] [PubMed]
- WHO, World Health Organization. 68th World Health Assembly: WHA Resolution 68.7; WHO: Geneva, Switzerland, 2015. [Google Scholar]
- ECDC (European Centre for Disease Prevention and Control). Surveillance of Antimicrobial Resistance in Europe 2016; Annual Report of the European Antimicrobial Resistance Surveillance Network (EARS-Net); ECDC: Stockholm, Sweden, 2017. [Google Scholar]
- Pärnänen, K.M.M.; Narciso-Da-Rocha, C.; Kneis, D.; Berendonk, T.U.; Cacace, D.; Do, T.T.; Elpers, C.; Fatta-Kassinos, D.; Henriques, I.; Jaeger, T.; et al. Antibiotic resistance in European wastewater treatment plants mirrors the pattern of clinical antibiotic resistance prevalence. Sci. Adv. 2019, 5, eaau9124. [Google Scholar] [CrossRef] [PubMed]
- Köppen, W. Versuch einer Klassifikation der Klimate, vorzugsweise nach ihren Beziehungen zur Pflanzenwelt (Attempted climate classification in relation to plant distributions). Geogr. Z. 1900, 6, 593–611. [Google Scholar]
- Lionello, P.; Malanotte-Rizzoli, P.; Boscolo, R.; Alpert, P.; Artale, V.; Li, L.; Luterbacher, J.; May, W.; Trigo, R.; Tsimplis, M.; et al. The Mediterranean climate: An overview of the main characteristics and issues. In Mediterranean Climate Variability; Lionello, P., Malanotte-Rizzoli, P., Boscolo, R., Eds.; Elsevier: Amsterdam, The Netherlands, 2006; pp. 1–26. [Google Scholar]
- Ulbrich, U. The Mediterranean climate change under global warming. In Mediterranean Climate Variability; Lionello, P., Malanotte-Rizzoli, P., Boscolo, R., Eds.; Elsevier: Amsterdam, The Netherlands, 2006; pp. 398–415. [Google Scholar]
- Giorgi, F. Climate change hot-spots. Geophys. Res. Lett. 2006, 33. [Google Scholar] [CrossRef]
- Luterbacher, J. Mediterranean climate variability over the last centuries. A review. In Mediterranean Climate Variability; Lionello, P., Malanotte-Rizzoli, P., Boscolo, R., Eds.; Elsevier: Amsterdam, The Netherlands, 2006; pp. 27–148. [Google Scholar]
- Hurrell, J.W. Decadal Trends in the North Atlantic Oscillation: Regional Temperatures and Precipitation. Science 1995, 269, 676–679. [Google Scholar] [CrossRef] [PubMed]
- Trigo, R. Relations between variability in the Mediterranean region and mid-latitude variability. In Mediterranean Climate Variability; Lionello, P., Malanotte-Rizzoli, P., Boscolo, R., Eds.; Elsevier: Amsterdam, The Netherlands, 2006; pp. 179–226. [Google Scholar]
- Alpert, P. Relations between climate variability in the Mediterranean region and the Tropics: ENSO, South Asian and African monsoons, hurricanes and Saharan dust. In Mediterranean Climate Variability; Lionello, P., Malanotte-Rizzoli, P., Boscolo, R., Eds.; Elsevier: Amsterdam, The Netherlands, 2006; pp. 149–177. [Google Scholar]
- Lionello, P.; Bhend, J.; Buzzi, A.; Della-Marta, P.M.; Krichak, S.O.; Jansa, A.; Maheras, P.; Sanna, A.; Trigo, I.F.; Trigo, R. Cyclones in the Mediterranean region: Climatology and effects on the environment. In Mediterranean Climate Variability; Lionello, P., Malanotte-Rizzoli, P., Boscolo, R., Eds.; Elsevier: Amsterdam, The Netherlands, 2006; pp. 325–372. [Google Scholar]
- Tuel, A.; Eltahir, E.A.B. Why Is the Mediterranean a Climate Change Hot Spot? J. Clim. 2020, 33, 5829–5843. [Google Scholar] [CrossRef]
- Contribution of Working Group I to the Fourth Assessment Report of the Intergovernmental Panel on Climate Change. In IPCC, Climate Change 2007: The Physical Science Basis; Solomon, S., Qin, D., Manning, M., Chen, Z., Marquis, M., Averyt, K.B., Tignor, M., Miller, H.L., Eds.; Cambridge University Press: Cambridge, UK; New York, NY, USA, 2007; p. 996. [Google Scholar]
- Giorgi, F.; Lionello, P. Climate change projections for the Mediterranean region. Glob. Planet. Chang. 2008, 63, 90–104. [Google Scholar] [CrossRef]
- Vautard, R.; Gobiet, A.; Sobolowski, S.; Kjellström, E.; Stegehuis, A.I.; Watkiss, P.; Mendlik, T.; Landgren, O.; Nikulin, G.; Teichmann, C.; et al. The European climate under a 2 °C global warming. Environ. Res. Lett. 2014, 9, 034006. [Google Scholar] [CrossRef]
- Forzieri, G.; Feyen, L.; Rojas, R.; Flörke, M.; Wimmer, F.; Bianchi, A. Ensemble projections of future streamfow droughts in Europe. Hydrol. Earth Syst. Sci. 2014, 18, 85–108. [Google Scholar] [CrossRef]
- IPCC, Intergovernmental Panel on Climate Change. Climate Change 2014: Synthesis Report; Contribution of Working Groups I, II and III to the Fifth Assessment Report of the Intergovernmental Panel on Climate Change; IPCC: Geneva, Switzerland, 2014; Available online: https://www.ipcc.ch/report/ar5/syr/ (accessed on 17 March 2021).
- Vilas-Boas, J.A.; Arenas-Sánchez, A.; Vighi, M.; Romo, S.; van den Brink, P.J.; Pedroso Dias, R.J.; Rico, A. Multi-ple stressors in Mediterranean coastal wetland ecosystems: Influence of salinity and an insecticide on zooplankton communities under different temperature conditions. Chemosphere 2021, 269, 129381. [Google Scholar] [CrossRef] [PubMed]
- Cramer, W.; Guiot, J.; Fader, M.; Garrabou, J.; Gattuso, J.-P.; Iglesias, A.; Lange, M.A.; Lionello, P.; Llasat, M.C.; Paz, S.; et al. Climate change and interconnected risks to sustainable development in the Mediterranean. Nat. Clim. Chang. 2018, 8, 972–980. [Google Scholar] [CrossRef]
- Macias, D.; Garcia-Gorriz, E.; Stips, A. Understanding the Causes of Recent Warming of Mediterranean Waters. How Much Could Be Attributed to Climate Change? PLoS ONE 2013, 8, e81591. [Google Scholar] [CrossRef]
- Seneviratne, S.I.; Donat, M.G.; Pitman, A.J.; Knutti, R.; Wilby, R.L. Allowable CO2 emissions based on regional and impact-related climate targets. Nature 2016, 529, 477–483. [Google Scholar] [CrossRef] [PubMed]
- Di Castri, F.; Mooney, H.A. Mediterranean Type Ecosystems: Origin and Structure; Mooney, H.A., di Cas-tri, F., Eds.; Springer: Berlin/Heidelberg, Germany, 1973. [Google Scholar]
- NCEZID; Centers for Disease Control and Prevention. National Center for Emerging and Zoonotic Infectious Diseases; Centers for Disease Control and Prevention: Atlanta, GA, USA, 2018. [Google Scholar]
- EU. Guidelines for the Prudent Use of Antimicrobials in Veterinary Medicine; Official Journal of the European Union: Brussels, Belgium, 2015. [Google Scholar]
- EU. Council Conclusions on the Next Steps under a One Health Approach to Combat Antimicrobial Resistance; Official Journal of the European Union: Brussels, Belgium, 2016. [Google Scholar]
- Zrnčić, S. European Union’s Action Plan on Antimicrobial Resistance and Implications for Trading Partners with Example of National Action Plan for Croatia. Asian Fish Sci. 2020, 33S, 75–82. [Google Scholar] [CrossRef]
- EU SWD Commission Staff Working Document. Evaluation of the Action Plan against the Rising Threats from Antimicrobial Resistance; Official Journal of the European Union: Brussels, Belgium, 2016. [Google Scholar]
- EU. A European One Health Action Plan against Antimicrobial Resistance (AMR); Official Journal of the European Union: Brussels, Belgium, 2017. [Google Scholar]
- Zinsstag, J.; Schelling, E.; Waltner-Toews, D. One Health: The Theory and Practice of Integrated Health Approaches; CABI: Wallingford, UK, 2015. [Google Scholar]
- Zinsstag, J.; Schelling, E.; Wyss, K.; Mahamat, M.B. Potential of cooperation between human and animal health to strengthen health systems. Lancet 2005, 366, 2142–2145. [Google Scholar] [CrossRef]
- Blaha, T. One World-One Health: The Threat of Emerging Diseases. A European Perspective. Transbound. Emerg. Dis. 2012, 59, 3–8. [Google Scholar] [CrossRef] [PubMed]
- Patz, J.A.; Hahn, M.B. Climate change and human health: A One Health approach. Curr. Top. Microbiol. Immunol. 2013, 366, 141–171. [Google Scholar] [PubMed]
- Watts, N.; Adger, W.N.; Ayeb-Karlsson, S.; Belesova, K.; Bouley, T.; Boykoff, M.; Byass, P.; Cai, W.; Campbell-Lendrum, D.; Chambers, J.; et al. The Lancet Countdown: Tracking progress on health and cli-mate change. Lancet 2017, 389, 1151–1164. [Google Scholar] [CrossRef]
- Zinsstag, J.; Crump, L.; Schelling, E.; Hattendorf, J.; Maidane, Y.O.; Ali, K.O.; Muhummed, A.; Umer, A.A.; Aliyi, F.; Nooh, F.; et al. Climate change and One Health. FEMS Microbiol. Lett. 2018, 365, fny085. [Google Scholar] [CrossRef]
- Shomaker, T.S.; Green, E.M.; Yandow, S.M. Perspective: One Health: A compelling convergence. Acad. Med. 2013, 88, 49–55. [Google Scholar] [CrossRef] [PubMed]
- Rabinowitz, P.M.; Natterson-Horowitz, B.J.; Kahn, L.H.; Kock, R.; Pappaioanou, M. Incorporating one health into medical education. BMC Med. Educ. 2017, 17, 45. [Google Scholar] [CrossRef]
- Videla, O.E.; Urzua, J.O. The benefits of incorporating the One Health concept into the organisation of veterinary services. Rev. Sci. Tech. 2014, 33, 401–406. [Google Scholar] [PubMed]
- Nyatanyi, T.; Wilkes, M.; McDermott, H.; Nzietchueng, S.; Gafarasi, I.; Mudakikwa, A.; Kinani, J.F.; Rukelibuga, J.; Omolo, J.; Mupfasoni, D.; et al. Implementing One Health as an integrated approach to health in Rwanda. BMJ Glob. Health 2017, 2, e000121. [Google Scholar] [CrossRef] [PubMed]
| Class | Antibiotic |
|---|---|
| Worldwide [15,16] | |
| aminoglycosides | streptomycin |
| amphenicols | florfenicol |
| β-lactams | amoxicillin, ampicillin |
| diaminopyrimidines | ormethoprim |
| macrolides | erythromycin |
| quinolones | oxolinic acid, flumequine, sarafloxacin, enrofloxacin |
| sulfonamides | sulfadimethoxine |
| tetracyclines | tetracycline, oxytetracycline |
| Most European Countries [17] | |
| amphenicols | florfenicol |
| macrolides | erythromycin |
| quinolones | sarafloxacin |
| sulphonamides | boosted with trimethoprim or ormethoprim |
| tetracyclines | oxytetracycline |
| In Mediterranean Aquaculture [18] | |
| quinolones | oxolinic acid, flumequine |
| tetracyclines | oxytetracycline |
| Country | Mean Percentage (%) | Range Percentage (%) | Year | Isolated Strains | Source |
|---|---|---|---|---|---|
| Bosnia and Herzegovina | 29 | 23–35 | 2016 | 215 | CAESAR |
| Croatia | 29 | 26–32 | 2017 | 1150 | EARS-Net |
| France | 17 | 16–18 | 2017 | 13,328 | EARS-Net |
| Greece | 34 | 32–36 | 2017 | 1464 | EARS-Net |
| Italy | 47 | 46–48 | 2017 | 6945 | EARS-Net |
| Lebanon | 45 | 33–57 | 2016 | 65 | GLASS |
| Portugal | 30 | 29–31 | 2017 | 6424 | EARS-Net |
| Spain | 33 | 32–34 | 2017 | 5557 | EARS-Net |
| Tunisia | 19 | 13–29 | 2017 | 78 | GLASS |
| Turkey | 55 | 53–57 | 2016 | 3670 | CAESAR |
| Country/Area | Isolated Bacteria Species/Genus/Family/Order/Class | Antibiotic-Resistance | Antibiotic-Resistant Genes | References |
|---|---|---|---|---|
| Algeria | Vibrio alginolyticus, V. cholerae, V. fluvialis, V. hollisae | [133] | ||
| Croatia | Aeromonas spp. | [134] | ||
| Eastern Adriatic | Vibrio spp. | flumequinone, chloramphenicol, oxytetracycline | [135] | |
| Egypt | Aeromonas spp. | chloramphenicol, kanamycin, azithromycin | [136] | |
| Egypt | Pseudomonas anguilliseptica | [137] | ||
| Egypt | Aeromonas hydrophila | penicillin, erythromycin | [138] | |
| Egypt | Enterobacteriaceae | cephalosporins, carbapenem | blaKPC (blaCTX-M-15, blaSHV, blaTEM, blaPER-1) | [139] |
| France | Flavobacterium psychrophilum | [140] | ||
| France | Yersinia ruckeri | [141] | ||
| Greece | Acinetobacter spp., Bacillus spp., Pseudomonas spp., Staphylococcus spp., Stenotrophomonas spp. | tetracycline | tetA, tetK, tetC, tetE, tetM | [127] |
| Italy | Enterococci | ampicillin, gentamicin | tetM, tetL, tetO, ermB, mef | [142] |
| Italy | Aeromonas spp., Photobacterium spp., Shewanella spp., Vibrio spp. | tetracycline, flumequine, trimethoprim | [122] | |
| Italy | Photobacterium damselae ssp. piscicida, Vibrio fluvialis, V. alginolyticus, V. parahaemolyticus, V. metschnikovii | ampicillin, carbenicillin, kanamycin, cefalothin | [143] | |
| Italy | Aeromonas spp. | ampicillin, amoxicillin, cephalothin, erythromycin, streptomycin, sulfadiazine, trimethoprim | [125] | |
| Italy | Shewanella algae, Vibrio spp. | beta-lactams, quinolones, tetracyclines, macrolides, polymyxins, chloramphenicol, fosfomycin, erythromycin | blaOXA-55-like, blaAmpC, mexB-OprM, mdtG, mdlB,tet34, tet35, tetR, eptA, cat, mdtL | [46] |
| Lebanon | Streptococcus pneumoniae | polymyxins, chloramphenicol, fosfomycin, erythromycin | [144] | |
| Spain | Aeromonas spp., Salmonella spp., Vibrio mimicus, V. furnissii. | oxytetracycline, nitrofurantoin, oxacillin, sulfomethoxazole/trimethoprim | [145] | |
| Spain | Flavobacterium psychrophilum | oxytetracycline, florfenicol | [146] | |
| Spain | Aeromonas salmonicida | nalidixic acid, oxytetracycline | [147] | |
| Tunisia | Vibrio alginolyticus | ampicillin, erythromycin, kanamycin, cefataxime, streptomycin, trimetoprim | [148] | |
| Tunisia | Escherichia coli | tertracycline, streptomycin, ampicillin, trimethoprim, sulfamethoxazole | tetA-tetB | [149] |
| Turkey | E. coli, coliforms, fish pathogens | sulfamethoxazole, ampicillin, sulfamethoxazole, imipenem, aztreonam | ampC, blaCTX-M1, tetA, sul2, blaTEM | [150] |
| Turkey | Y. ruckeri | floR, sulI, tetC, tetD, tetE | [151] | |
| Turkey | Y. ruckeri | erythromycin, florfenicol, sulfonamide, tetracycline, trimetophrin | ermB, ermY, floR, su/I, suffll, tetA-tetG | [152] |
| Turkey | Aeromonas media, A. rivipollensis, A. salmonicida, Bacillus pumilus, B. zhangzhouensis, Hafnia alvei, Kluyvera intermedia, Pantoea spp., Pseudomonas spp., P. protegens, Staphylococcus spp., Gammaproteobacteria, Betaproteobacteria, Enterobacteriales, Burkholderiales | sulfamethacin, sulfamerazine, erythromycin, tetracycline | [153] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pepi, M.; Focardi, S. Antibiotic-Resistant Bacteria in Aquaculture and Climate Change: A Challenge for Health in the Mediterranean Area. Int. J. Environ. Res. Public Health 2021, 18, 5723. https://doi.org/10.3390/ijerph18115723
Pepi M, Focardi S. Antibiotic-Resistant Bacteria in Aquaculture and Climate Change: A Challenge for Health in the Mediterranean Area. International Journal of Environmental Research and Public Health. 2021; 18(11):5723. https://doi.org/10.3390/ijerph18115723
Chicago/Turabian StylePepi, Milva, and Silvano Focardi. 2021. "Antibiotic-Resistant Bacteria in Aquaculture and Climate Change: A Challenge for Health in the Mediterranean Area" International Journal of Environmental Research and Public Health 18, no. 11: 5723. https://doi.org/10.3390/ijerph18115723
APA StylePepi, M., & Focardi, S. (2021). Antibiotic-Resistant Bacteria in Aquaculture and Climate Change: A Challenge for Health in the Mediterranean Area. International Journal of Environmental Research and Public Health, 18(11), 5723. https://doi.org/10.3390/ijerph18115723







