Wastewater-Based Epidemiology as an Early Warning System for the Spreading of SARS-CoV-2 and Its Mutations in the Population
Abstract
1. Introduction
2. Coronavirus SARS-CoV-2 and Wastewater
Wastewater—A Possible Source of Information
3. Detection of SARS-CoV-2
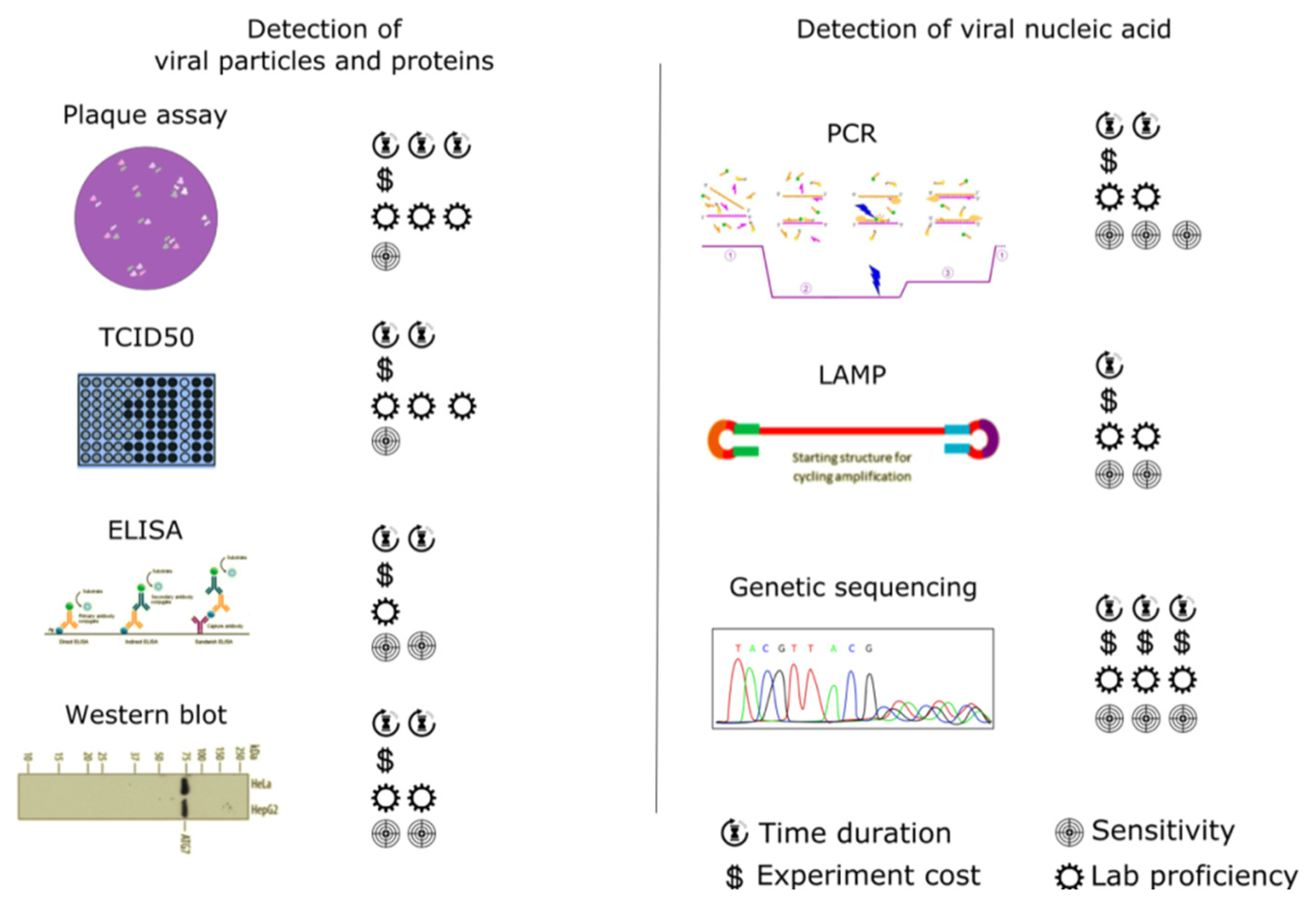
3.1. Traditional (Standard) Techniques for Virus Detection
3.1.1. Detection of Viral Particles and Proteins
3.1.2. Detection of Viral Nucleic Acids
3.1.3. Latest Research in Traditional Techniques for Virus Detection
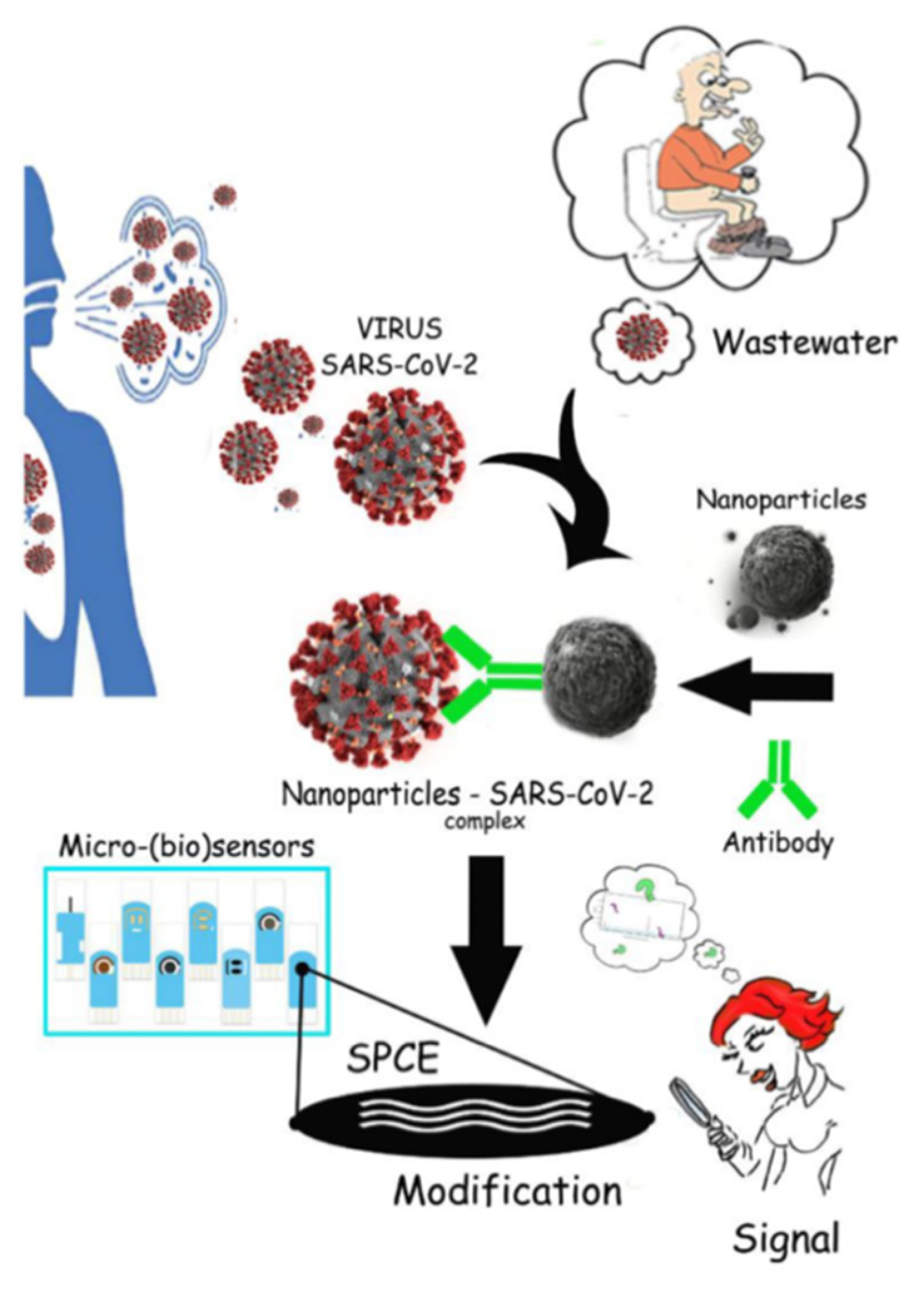
3.2. Biosensors—Early Warning System for Virus Detection in Wastewater
3.2.1. Biocatalytic Sensors
3.2.2. Affinity Biosensors
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Daughton, C.D.; Ternes, T.A. Pharmaceuticals and personal care products in the environment: Agents of subtle change? Environ. Health Perspect. 1999, 107, 907–938. [Google Scholar] [CrossRef] [PubMed]
- Zuccato, E.; Chiabrando, C.; Castiglioni, S.; Calamari, D.; Bagnati, R.; Schiarea, S.; Fanelli, R. Cocaine in surface waters: A new evidence-based tool to monitor community drug abuse. Environ. Health 2005, 4, 14–20. [Google Scholar] [CrossRef] [PubMed]
- Bijlsma, L.; Celma, A.; Lopez, F.J.; Hernandez, F. Monitoring new psychoactive substances use through wastewater analysis: Current situation, challenges and limitations. Curr. Opin. Environ. Sci. Health 2019, 9, 1–12. [Google Scholar] [CrossRef]
- Castrignanò, E.; Yang, Z.; Feil, E.J.; Bade, R.; Castiglioni, S.; Causanilles, A.; Gracia-Lor, E.; Hernandez, F.; Plosz, B.G.; Ramin, P.; et al. Enantiomeric profiling of quinolones and quinolones resistance gene qnrS in European wastewaters. Water Res. 2020, 175, 115653. [Google Scholar] [CrossRef] [PubMed]
- Gracia-Lor, E.; Zuccato, E.; Hernandez, F.; Castiglioni, S. Wastewater-based epidemiology for tracking human exposure to mycotoxins. J. Hazard. Mater. 2020, 382, 121108. [Google Scholar] [CrossRef]
- Rousis, I.N.; Gracia-Lor, E.; Reid, M.J.; Baz-Lomba, J.A.; Ryu, Y.; Zuccato, E.; Thomas, K.V.; Castiglioni, S. Assessment of human exposure to selected pesticides in Norway by wastewater analysis. Sci. Total Environ. 2020, 723, 138132. [Google Scholar] [CrossRef]
- Sims, N.; Kasprzyk-Hordern, B. Future perspectives of wastewater-based epidemiology: Monitoring infectious disease spread and resistance to the community level. Environ. Int. 2020, 139, 105689. [Google Scholar] [CrossRef]
- Castrignanò, E.; Mardal, M.; Rydevik, A.; Miserez, B.; Ramsey, J.; Shine, T.; Pantos, D.; Meyer, M.R.; Kasprzyk-Hordern, B. A newapproach towards biomarker selection in estimation of human exposure to chiral chemicals: A case study of mephedrone. Sci. Rep. 2017, 7, 13009. [Google Scholar] [CrossRef]
- Xagoraraki, I.; O’Brien, E. Wastewater-based epidemiology for early detection of viral outbreaks. In Women in Water Quality, Women in Engineering and Science; O’Bannon, D., Ed.; Springer Nature: Cham, Switzerland, 2020; pp. 75–97. [Google Scholar] [CrossRef]
- WHO. 2020. Available online: https://web.archive.org/web/20141214011751/ (accessed on 7 April 2020).
- WHO. 2020. Available online: https://www.who.int/en/news-room/fact-sheets/detail/rabies (accessed on 7 April 2020).
- WHO. 2020. Available online: https://www.who.int/csr/sars/country/table2004_04_21/en/ (accessed on 7 April 2020).
- Ozili, P.K.; Arun, T. Spillover of COVID-19: Impact on the global economy. SSRN Electron. J. 2020, 3562570. [Google Scholar] [CrossRef]
- Van Doremalen, N.; Bushmaker, T.; Morris, D.H.; Holbrook, M.G.; Gamble, A.; Williamson, B.N.; Tamin, A.; Harcourt, J.L.; Thornburg, N.J.; Gerber, S.I.; et al. Aerosol and surface stability of SARS-CoV-2 as compared with SARS-CoV-1. N. Engl. J. Med. 2020, 382, 1564–1567. [Google Scholar] [CrossRef]
- Tian, Y.; Rong, L.; Nian, W.; He, Y. Review article: Gastrointestinal features in COVID-19 and the possibility of faecal transmission. Aliment. Pharmacol. Ther. 2020, 51, 843–851. [Google Scholar] [CrossRef]
- Wang, D.; Hu, B.; Hu, C.; Zhu, F.; Liu, X.; Zhang, J.; Wang, B.; Xiang, H.; Cheng, Z.; Xiong, Y.; et al. Clinical characteristics of 138 hospitalized patients with 2019 novel coronavirus-infected pneumonia in Wuhan, China. J. Am. Med. Assoc. 2020, 323, 1061–1069. [Google Scholar] [CrossRef] [PubMed]
- WHO. Water, Sanitation, Hygiene and Waste Management for COVID-19. 2020. Available online: https://www.who.int/publications-detail/water-sanitation-hygiene-and-waste-management-for-covid-19 (accessed on 23 March 2020).
- Xiao, E.; Tang, M.; Zheng, X.; Liu, Y.; Li, C.; He, J.; Hong, Z.; Huang, S.; Zhang, Z.; Lin, X.; et al. Evidence for gastrointestinal infection of SARS-CoV. Gastroenterology 2020, 158, 1831–1833. [Google Scholar] [CrossRef]
- Yeo, C.; Kaushal, S.; Yeo, D. Enteric involvement of coronaviruses: Is faecal–oral transmission of SARS-CoV-2 possible? Lancet Gastroenterol. Hepatol. 2020, 5, 335–337. [Google Scholar] [CrossRef]
- Langone, M.; Petta, L.; Cellamare, C.M.; Ferraris, M.; Guzzinati, R.; Mattioli, D.; Sabia, G. SARS-CoV-2 in water services: Presence and impacts. Environ. Pollut. 2020, 268, 115806. [Google Scholar] [CrossRef] [PubMed]
- Medema, G.; Heijnen, L.; Elsinga, G.; Italiaander, R.; Brouwer, A. Presence of SARS-Coronavirus-2 RNA in Sewage and Correlation with Reported COVID-19 Prevalence in the Early Stage of the Epidemic in The Netherlands. Environ. Sci. Technol. Lett. 2020, 7, 511–516. [Google Scholar] [CrossRef]
- Orive, G.; Lertxundi, U.; Barcelo, D. Early SARS-CoV-2 outbreak detection by sewage-based epidemiology. Sci. Total Environ. 2020, 732, 139298. [Google Scholar] [CrossRef] [PubMed]
- Hokajärvi, A.M.; Rytkönen, A.; Tiwari, A.; Kauppinen, A.; Oikarinen, S.; Lehto, K.M.; Kankaanpää, A.; Gunnar, T.; Al-Hello, H.; Blomqvist, S.; et al. The detection and stability of the SARS-CoV-2 RNA biomarkers in wastewater influent in Helsinki, Finland. Sci. Total. Environ. 2021, 770, 145274. [Google Scholar] [CrossRef]
- Pan, Y.; Zhang, D.; Yang, P.; Poon, L.L.M.; Wang, Q. Viral load of SARS-CoV-2 in clinical samples. Lancet Infect. Dis. 2020, 20, 411–412. [Google Scholar] [CrossRef]
- Woelfel, R.; Corman, V.M.; Guggemos, W.; Seilmaier, M.; Zange, S.; Mueller, M.A.; Niemeyer, D.; Vollmar, P.; Rothe, C.; Hoelscher, M.; et al. Clinical presentation and virological assessment of hospitalized cases of coronavirus disease 2019 in a travel-associated transmission cluster. medRxiv 2020. [Google Scholar] [CrossRef]
- Chen, C.; Gao, G.; Xu, Y.; Pu, L.; Wang, Q.; Wang, L.; Wang, W.; Song, Y.; Chen, M.; Wang, L.; et al. SARS-CoV-2–positive sputum and feces after conversion of pharyngeal samples in patients with COVID-19. Ann. Intern. Med. 2020. [Google Scholar] [CrossRef]
- Lescure, F.X.; Bouadma, L.; Nguyen, D.; Parisey, M.; Wicky, P.H.; Behillil, S.; Gaymard, A.; Bouscambert-Duchamp, M.; Donati, F.; Le Hingrat, Q.; et al. Clinical and virological data of the first cases of COVID-19 in Europe: A case series. Lancet Infect. Dis. 2020, 20, 697–706. [Google Scholar] [CrossRef]
- Wang, W.; Xu, Y.; Gao, R.; Lu, R.; Han, K.; Wu, G.; Tan, W. Detection of SARS-CoV-2 in different types of clinical specimens. J. Am. Med. Assoc. 2020, 323, 1843–1844. [Google Scholar] [CrossRef]
- Wu, Y.; Guo, C.; Tang, L.; Hong, Z.; Zhou, J.; Dong, X.; Yin, H.; Xiao, Q.; Tang, Y.; Qu, X.; et al. Prolonged presence of SARS-CoV-2 viral RNA in faecal samples. Lancet Gastroenterol. Hepatol. 2020, 5, 434–435. [Google Scholar] [CrossRef]
- Xu, Y.; Li, X.; Zhu, B.; Liang, H.; Fang, C.; Gong, Y.; Guo, Q.; Sun, X.; Zhao, D.; Shen, J.; et al. Characteristics of pediatric SARS-CoV-2 infection and potential evidence for persistent fecal viral shedding. Nat. Med. 2020, 26, 502–505. [Google Scholar] [CrossRef]
- Randazzo, W.; Truchado, P.; Cuevas-Ferrando, E.; Simon, P.; Allende, A.; Sanchez, G. SARS-CoV-2 RNA in wastewater anticipated COVID-19 occurrence in a low prevalence area. Water Res. 2020, 181, 115942. [Google Scholar] [CrossRef]
- Haramoto, E.; Malla, B.; Thakali, O.; Kitajima, M. First environmental surveillance for the presence of SARS-CoV-2 RNA in wastewater and river water in Japan. Sci. Total Environ. 2020, 737, 140405. [Google Scholar] [CrossRef]
- Randazzo, W.; Cuevas-Ferrando, E.; Sanjuan, R.; Domingo-Calap, P.; Sanchez, G. Metropolitan wastewater analysis for COVID-19 epidemiological surveillance. Int. J. Hyg. Environ. Health. 2020, 230, 113621. [Google Scholar] [CrossRef] [PubMed]
- Fongaro, G.; Stoco, P.H.; Souza, D.S.M.; Grisard, E.C.; Magri, M.E.; Rogovski, P.; Schorner, M.A.; Barazzetti, F.H.; Christoff, A.P.; de Oliveira, L.F.V.; et al. SARS-CoV-2 in human sewage in Santa Catalina, Brazil, November 2019. Sci. Total Environ. 2021, 778, 146198. [Google Scholar] [CrossRef] [PubMed]
- La Rosa, G.; Mancini, P.; Ferraro, G.B.; Veneri, C.; Iaconelli, M.; Bonadonna, L.; Lucentini, L.; Suffredini, E. SARS-CoV-2 has been circulating in northern Italy since December 2019: Evidence from environmental monitoring. Sci. Total Environ. 2021, 750, 141711. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Pei, S.; Chen, B.; Song, Y.; Zhang, T.; Yang, W.; Shaman, J. Substantial undocumented infection facilitates the rapid dissemination of novel coronavirus (SARS-CoV-2). Science 2020, 368, 489–493. [Google Scholar] [CrossRef]
- Kitajima, M.; Ahmed, W.; Bibby, K.; Carducci, A.; Gerba, C.P.; Hamilton, K.A.; Harmoto, E.; Rose, J.B. SARS-CoV-2 in wastewater: State of the knowledge and research needs. Sci. Total Environ. 2020, 739, 139076. [Google Scholar] [CrossRef]
- La Rosa, G.; Bonadonna, L.; Lucentini, L.; Kenmore, S.; Suffredini, E. Coronavirus in water environments: Occurrence, persistence and concentration methods—A scoping review. Water Res. 2020, 179, 115899. [Google Scholar] [CrossRef]
- Bivins, A.; Greaves, J.; Fischer, R.; Yinda, K.C.; Ahmed, W.; Kitajima, M.; Munster, V.J.; Bibby, K. Persistence of SARS-CoV-2 in Water and Wastewater. Environ. Sci. Technol. Lett. 2020, 7, 937–942. [Google Scholar] [CrossRef]
- Jones, D.L.; Baluja, M.Q.; Graham, D.W.; Corbishley, A.; McDonald, J.E.; Malham, S.K.; Hillary, L.S.; Connor, T.R.; Gaze, W.H.; Moura, I.B.; et al. Fecal Shedding of SARS-CoV-2 and its Potential Role in Person-To-Person Transmission and the Environment- Based Spread of COVID-19. Sci. Total Environ. 2020, 749, 141364. [Google Scholar] [CrossRef] [PubMed]
- Westhouse, S.; Weber, F.A.; Schiwy, S.; Linnemann, V.; Brinkmann, M.; Widera, M.; Greve, C.; Janke, A.; Hollert, H.; Wintgens, T.; et al. Detection of SARS-CoV-2 in raw and treated wastewater in Germany—Suitability for COVID-19 surveillance and potential transmission risks. Sci. Total Environ. 2021, 751, 141750. [Google Scholar] [CrossRef]
- Hart, O.E.; Halden, R.U. Computational analysis of SARS-CoV-2/COVID-19 surveillance by wastewater-based epidemiology locally and globally: Feasibility, economy, opportunities and challenges. Sci. Total Environ. 2020, 730, 138875. [Google Scholar] [CrossRef] [PubMed]
- Katayama, H.; Haramoto, E.; Oguma, K.; Yamashita, H.; Tajima, A.; Nakajima, H.; Ohgaki, S. One-year monthly quantitative survey of noroviruses, enteroviruses, and adenoviruses in wastewater collected from six plants in Japan. Water Res. 2008, 42, 1441–1448. [Google Scholar] [CrossRef] [PubMed]
- Skraber, S.; Ogorzaly, L.; Helmi, K.; Maul, A.; Hoffmann, L.; Cauchie, H.M.; Gantzer, C. Occurrence and persistence of enteroviruses, noroviruses and F-specific RNA phages in natural wastewater biofilms. Water Res. 2009, 43, 4780–4789. [Google Scholar] [CrossRef]
- Bashir, M.F.; Ma, B.; Bilal; Komal, B.; Bashir, M.A.; Tan, D.; Bashir, M. Correlation between climate indicators and COVID-19 pandemic in New York, USA. Sci. Total Environ. 2020, 728, 138835. [Google Scholar] [CrossRef]
- Gupta, S.; Parker, J.; Smits, S.; Underwood, J.; Dolwani, S. Persistent viral shedding of SARS-CoV-2 in faeces—A rapid review. Colorectal Dis. 2020, 22, 611–622. [Google Scholar] [CrossRef] [PubMed]
- Núñez-Delgado, A. What do we know about the SARS-CoV-2 coronavirus in the environment? Sci. Total Environ. 2020, 727, 138647. [Google Scholar] [CrossRef] [PubMed]
- Wu, F.Q.; Xiao, A.; Zhang, J.B.; Gu, X.Q.; Lee, W.L.; Kauffman, K.; Hanage, W.P.; Matus, M.; Ghaeli, N.; Endo, N.; et al. SARS-CoV-2 titers in wastewater are higher than expected from clinically confirmed cases. mSystems 2020, 5, e00614-20. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, W.; Angel, N.; Edson, J.; Bibby, K.; Bivins, A.; O’Brien, J.W.; Choi, P.M.; Kitajima, M.; Simpson, S.L.; Li, J.; et al. First confirmed detection of SARS-CoV-2 in untreated wastewater in Australia: A proof of concept for the wastewater surveillance of COVID-19 in the community. Sci. Total Environ. 2020, 728, 138764. [Google Scholar] [CrossRef]
- Wurtzer, S.; Marechal, V.; Mouchel, J.M.; Moulin, L. Time course quantitative detection of SARS-CoV-2 in Parisian wastewaters correlates with COVID-19 confirmed cases. medRxiv 2020. [Google Scholar] [CrossRef]
- Kocamemi, B.A.; Kurt, H.; Hacioglu, S.; Yarali, C.; Saatci, A.M.; Pakdemirli, B. First Data-Set on SARS-CoV-2 Detection for Istanbul Wastewaters in Turkey. medRxiv 2020. [Google Scholar] [CrossRef]
- Rimoldi, S.G.; Stefani, F.; Gigantiello, A.; Polesello, S.; Comandatore, F.; Mileto, D.; Maresca, M.; Longobardi, C.; Mancon, A.; Romeri, F.; et al. Presence and infectivity of SARS-CoV-2 virus in wastewaters and rivers. Sci. Total Environ. 2020, 744, 140911. [Google Scholar] [CrossRef]
- Iglesias, N.G.; Gebhard, L.G.; Carballeda, J.M.; Aiello, I.; Recalde, E.; Terny, G.; Ambrosolio, S.; L’Arco, G.; Jonatan, K.; Brardinelli, J.I. SARS-CoV-2 surveillance in untreated wastewater: First detection in a low-resource community in Buenos Aires, Argentina. medRxiv 2020. [Google Scholar] [CrossRef]
- Sherchan, S.P.; Shahin, S.; Ward, L.M.; Tandukar, S.; Aw, T.G.; Schmitz, B.; Ahmed, W.; Kitajima, M. First detection of SARS-CoV-2 RNA in wastewater in North America: A study in Louisiana, USA. Sci. Total Environ. 2020, 743, 140621. [Google Scholar] [CrossRef] [PubMed]
- Kocamemi, B.A.; Kurt, H.; Sait, A.; Sarac, F.; Saatci, A.M.; Pakdemirli, B. SARS-CoV-2 Detection in Istanbul Wastewater Treatment Plant Sludges. medRxiv 2020. [Google Scholar] [CrossRef]
- Miyani, B.; Fonoll, X.; Norton, J.; Mehrotra, A.; Xagoraraki, I. SARS-CoV-2 in Detroit Wastewater. J. Environ. Eng. 2020, 146, 06020004. [Google Scholar] [CrossRef]
- Mlejnková, H.; Sovova, K.; Vasickova, P.; Ocenaskova, V.; Jasikova, L.; Juranova, E. Preliminary Study of Sars-Cov-2 Occurrence in Wastewater in the Czech Republic. Int. J. Environ. Res. Public Health 2020, 17, 5508. [Google Scholar] [CrossRef] [PubMed]
- Agrawal, S.; Orschler, L.; Lackner, S. Long-term monitoring of SARS-CoV-2 in wastewater of the Frankfurt metropolitan area in Southern Germany. Sci. Rep. 2021, 11, 5372. [Google Scholar] [CrossRef]
- Been, F.; Rossi, L.; Ort, C.; Rudaz, S.; Delemont, O.; Esseiva, P. Population normalization with ammonium in wastewater-based epidemiology: Application to illicit drug monitoring. Environ. Sci. Technol. 2014, 48, 8162–8169. [Google Scholar] [CrossRef]
- Chen, C.; Kostakis, C.; Gerber, J.P.; Tscharke, B.J.; Irvine, R.J.; White, J.M. Towards finding apopulation biomarker for wastewater epidemiology studies. Sci. Total Environ. 2014, 487, 621–628. [Google Scholar] [CrossRef]
- Choi, M.P.; Tscharke, B.J.; Donner, E.; O’Brien, J.W.; Grant, S.C.; Kaserzon, S.L.; Mackie, R.; O’Malley, E.; Crosbie, N.D.; Thomas, K.V.; et al. Wastewater-based epidemiology biomarkers: Past, present and future. Trends Anal. Chem. 2018, 105, 453–469. [Google Scholar] [CrossRef]
- Lomba, B.A.J.; Di Ruscio, F.; Amador, A.; Reid, M.; Thomas, K.V. Assessing alternative population size proxies in a wastewater catchment area using mobile device data. Environ. Sci. Technol. 2019, 53, 1994–2001. [Google Scholar] [CrossRef] [PubMed]
- Larson, R.C.; Berman, O.; Nourinejad, M. Sampling manholes to home in on SARS-CoV-2 infections. PLoS ONE 2020, 15, e0240007. [Google Scholar] [CrossRef]
- Daughton, C. The international imperative to rapidly and inexpensively monitor community-wide Covid-19 infection status and trends. Sci. Total Environ. 2020, 726, 138149. [Google Scholar] [CrossRef]
- Farkas, K.; Walker, D.I.; Adriaenssens, E.M.; McDonald, J.E.; Hillary, L.S.; Malham, S.K.; Jones, D.L. Viral indicators for tracking domestic wastewater contamination in the aquatic environment. Water Res. 2020, 181, 115926. [Google Scholar] [CrossRef]
- Thomas, K.V.; Bijilsma, L.; Castiglioni, S.; Covaci, A.; Emke, E.; Grabic, R.; Hernandez, F.; Karolak, S.; Kasprzyk-Hordern, B.; Lindeberg, R.H.; et al. Comparing illicit drug use in 19 European cities through sewage analysis. Sci. Total Environ. 2012, 432, 432–439. [Google Scholar] [CrossRef] [PubMed]
- Ort, C.; van Nuijs, A.L.N.; Berset, J.D.; Bijlsma, L.; Castiglioni, S.; Covaci, A.; de Voogt, P.; Emke, E.; Fatta-Kassinos, D.; Griffiths, P.; et al. Spatial differences and temporal changes in illicit drugs use in Europe quantified by wastewater analysis. Addiction 2014, 109, 1338–1352. [Google Scholar] [CrossRef] [PubMed]
- Krizman-Matasic, I.; Senta, I.; Kostanjevecki, P.; Ahel, M.; Terzic, S. Long-term monitoring of drug consumption patterns in a large-sized European city using wastewater-based epidemiology: Comparison of two sampling schemes for the assessment of multiannual trends. Sci. Total Environ. 2019, 647, 474–485. [Google Scholar] [CrossRef]
- COVID19 Wbe Collaborative. Available online: https://www.covid19wbec.org/ (accessed on 7 April 2020).
- Polo, D.; Quintela-Baluja, M.; Corbishley, A.; Jones, D.L.; Singer, A.C.; Graham, D.W.; Romalde, J.L. Making waves: Wastewater-based epidemiology for COVID-19—Approaches and challenges for surveillance and prediction. Water Res. 2020, 186, 116404. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, F.; Islam, A.; Kumar, M.; Hossain, M.; Bhattacharya, P.; Islam, T.; Hossen, F.; Hossain, S.; Islam, S.; Uddin, M.; et al. First detection of SARS-CoV-2 genetic material in the vicinity of COVID-19 isolation centre through wastewater surveillance in Bangladesh. medRxiv 2020. [Google Scholar] [CrossRef]
- Albastaki, A.; Naji, M.; Lootah, R.; Almeheiri, R.; Almulla, H.; Almarri, I.; Alreyami, A.; Aden, A.; Alghafri, R. First confirmed detection of SARS-COV-2 in untreated municipal and aircraft wastewater in Dubai, UAE: The use of wastewater based epidemiology as an early warning tool to monitor the prevalence of COVID-19. Sci. Total Environ. 2021, 760, 143350. [Google Scholar] [CrossRef]
- Kumar, M.; Patel, A.K.; Shah, A.V.; Raval, J.; Rajpara, N.; Joshi, M.; Joshi, C.G. First proof of the capability of wastewater surveillance for COVID-19 in India through detection of genetic material of SARS-CoV-2. Sci. Total Environ. 2020, 746, 141326. [Google Scholar] [CrossRef]
- La Rosa, G.; Iaconelli, M.; Mancini, P.; Bonanno, F.; Veneri, C.; Bonadonna, L.; Lucentini, L.; Suffredini, E. First detection of SARS-CoV-2 in untreated wastewaters in Italy. Sci. Total Environ. 2020, 736, 139652. [Google Scholar] [CrossRef]
- Bar Or, I.; Yaniv, K.; Shagan, M.; Ozer, E.; Erster, O.; Mendelson, E.; Mannasse, B.; Shirazi, R.; Kramarsky-Winter, E.; Nir, O.; et al. Regressing SARS-CoV-2 sewage measurements onto COVID-19 burden in the population: A proof-of-concept for quantitative environmental surveillance. medRxiv 2020. [Google Scholar] [CrossRef]
- Guerrero-Latorre, L.; Ballesteros, I.; Villacrés-Granda, I.; Grandam, M.G.; Freire-Paspuel, B.; Ríos Touma, B. SARS-CoV-2 in river water: Implications in low sanitation countries. Sci. Total Environ. 2020, 743, 140832. [Google Scholar] [CrossRef]
- Available online: https://www.cdc.gov/coronavirus/2019-ncov/lab/rt-pcr-panel-primer-probes.html (accessed on 24 May 2021).
- Leland, D.S.; Ginocchio, C.C. Role of cell culture for virus detection in the age of technology. Clin. Microbiol. Rev. 2017, 20, 49–78. [Google Scholar] [CrossRef]
- Modrow, S.; Falke, D.; Truyen, U.; Schätzl, H. Laboratory methods for detecting viral infections. In Molecular Virology; Springer: Berlin, Germany, 2013; pp. 163–181. [Google Scholar] [CrossRef]
- Schauflinger, M.; Villinger, C.; Walther, P. Three-dimensional visualization of virus-infected cells by serial sectioning: An electron microscopic study using resin embedded cells. In Virus-Host Interactions: Methods and Protocols; Bailer, S.M., Lieber, D., Eds.; Humana Press: Stuttgart, Germany, 2013; pp. 227–237. [Google Scholar] [CrossRef]
- Dilnessa, T.; Zeleke, H. Cell Culture, Cytopathic effect and immunofluorescence diagnosis of viral infection. J. Microbiol. Modern. Tech. 2017, 2, 1–8. [Google Scholar] [CrossRef]
- Stephenson, J.R.; Warnes, A. Diagnostic Virology Protocols; Humana Press: Totowa, NJ, USA, 2011; p. 470. [Google Scholar] [CrossRef]
- Mattison, K.; Bidawid, S. Analytical methods for food and environmental viruses. Food Environ. Virol. 2009, 1, 107–122. [Google Scholar] [CrossRef]
- Barardi, C.R.M.; Viancelli, A.; Rigotto, C.; Correa, A.A.; Moresco, V.; Souza, D.S.M.; ElMahdy, M.E.I.; Fongaro, G.; Pilotto, M.R.; Nascimento, M.A. Monitoring viruses in environmental samples. Int. J. Environ. Sci. Eng. Res. 2012, 3, 62–79. [Google Scholar]
- Calgua, B.; Barardi, C.R.M.; Bofill-Mas, S.; Rodriguez-Manzano, J.; Girones, R. Detection and quantitation of infectious human adenoviruses and JC polyomaviruses in water by immunofluorescence assay. J. Virol. Methods 2011, 171, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Greening, G.E.; Hewitt, J.; Lewis, G.D. Evaluation of integrated cell culture-PCR (C-PCR) for virological analysis of environmental samples. J. Appl. Microbiol. 2002, 93, 745–750. [Google Scholar] [CrossRef] [PubMed]
- Haramoto, E.; Kitajima, M.; Hata, A.; Torrey, J.R.; Masago, Y.; Sano, D.; Katayama, H. A review on recent progress in the detection methods and prevalence of human enteric viruses in water. Water Res. 2018, 135, 68–186. [Google Scholar] [CrossRef]
- Engvall, E.; Perlmann, P. Enzyme-linked immunosorbent assay (ELISA). Quantitative assay of immunoglobulin G. Immunochemestry 1971, 8, 871–874. [Google Scholar] [CrossRef]
- Crowther, J.R. Methods in Molecular Biology; The ELISA Guidebook; Humana Press: Vienna, Austria, 2009; p. 566. [Google Scholar] [CrossRef]
- Maier, R.M.; Pepper, I.L.; Gerba, C.P. Enviromental Microbiology, 2nd ed.; Academic Press: Oxford, UK, 2009; p. 598. [Google Scholar] [CrossRef]
- He, J. Practical guide to ELISA development. In The Immunoassay Handbook; Wild, D., Ed.; Elsevier Science: Oxford, UK, 2013; pp. 381–393. [Google Scholar] [CrossRef]
- Sakamoto, S.; Putalun, W.; Vimolmangkang, S.; Phoolcharoen, W.; Shoyama, Y.; Tanaka, H.; Morimoto, S. Enzyme-linked immunosorbent assay for the quantitative/qualitative analysis of plant secondary metabolites. J. Nat. Med. 2018, 72, 32–42. [Google Scholar] [CrossRef]
- McFeters, G.A. Drinking Water Microbiology Progress and Recent Developements; Springer: Bozeman, MT, USA, 1990; p. 502. [Google Scholar] [CrossRef]
- Towbin, H.; Staehelin, T.; Gordon, J. Electrophoretic transfer of proteins from polyacrylamide gels to nitrocellulose sheets: Procedure and some applications. Proc. Natl. Acad. Sci. USA 1979, 76, 4350–4354. [Google Scholar] [CrossRef]
- Bass, J.J.; Wilkinson, D.J.; Rankin, D.; Phillips, B.E.; Szewczyk, N.J.; Smith, K.; Atherton, P.J. An overview of technical considerations for Western blotting applications to physiological research. Scand. J. Med. Sci. Sports 2016, 27, 4–25. [Google Scholar] [CrossRef] [PubMed]
- Jensen, E.C. The basics of western blotting. Anat. Rec. 2012, 295, 369–371. [Google Scholar] [CrossRef] [PubMed]
- Eslami, A.; Lujan, J. Western blotting: Sample preparation to detection. J. Vis. Exp. 2010, 44, 2359. [Google Scholar] [CrossRef] [PubMed]
- Kurien, B.; Scofield, R. Western blotting. Methods 2006, 38, 283–293. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Sun, H.; Huang, G.; Liu, G.; Li, Z.; Yang, H.; Jin, L.; Cui, X.; Shi, L.; Ma, T.; et al. A fixation method for the optimisation of western blotting. Sci. Rep. 2019, 9, 6649. [Google Scholar] [CrossRef]
- Meads, M.B.; Medveczky, P.G. Application of western blotting to diagnosis of viral infections. In Clinical Virology Manual, 4th ed.; Specter, S., Hodinka, R., Young, S., Wiedbrauk, D., Eds.; ASM Press: Washington, DC, USA, 2009; pp. 150–155. [Google Scholar] [CrossRef]
- Mahmood, T.; Yang, P.C. Western blot: Technique, theory, and trouble shooting. N. Am. J. Med. Sci. 2012, 4, 429–434. [Google Scholar] [CrossRef]
- Taylor, S.C.; Berkelman, T.; Yadav, G.; Hammond, M. A defined methodology for reliable quantification of western blot data. Mol. Biotechnol. 2013, 55, 217–226. [Google Scholar] [CrossRef]
- La Rosa, G.; Muscillo, M. Molecular detection of viruses in water and sewage. In Viruses in Food and Water: Risks, Surveillance and Control; Cook, N., Ed.; Woodhead Publishing: Cambridge, UK, 2013; pp. 97–125. [Google Scholar] [CrossRef]
- Buzdin, A.A. Nucleic acids hybridization: Potentials and limitations. In Nucleic Acids Hybridization Modern Applications; Buzdin, A.A., Lukyanov, S.A., Eds.; Springer: Dordrecht, The Netherlands, 2007; pp. 1–28. [Google Scholar] [CrossRef]
- Mays Hoopes, L.L. Nucleic acid blotting: Southern and northern. In Current Protocols Essential Laboratory Techniques; Gallagher, S.R., Wiley, E.A., Eds.; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2008; pp. 821–824. [Google Scholar] [CrossRef]
- Yeh, H.Y.; Yates, M.V.; Chen, W.; Mulchandani, A. Real-time molecular methods to detect infectious viruses. Semin. Cell. Dev. Biol. 2009, 20, 49–54. [Google Scholar] [CrossRef]
- Girones, R.; Ferrús, M.A.; Alonso, J.L.; Rodriguez-Manzano, J.; Calgua, B.; Corrêa, A.A.; Hundesa, A.; Carratala, A.; Bofill-Mas, S. Molecular detection of pathogens in water-the pros and cons of molecular techniques. Water Res. 2010, 44, 4325–4339. [Google Scholar] [CrossRef]
- Van Pelt-Verkuil, E.; van Belkum, A.; Hays, J.P. The polymerase chain reaction. In Principles and Technical Aspects of PCR Amplification; van Pelt-Verkuil, E., van Belkum, A., Hays, J.P., Eds.; Springer: Dordrecht, The Netherlands, 2008; pp. 1–7. [Google Scholar] [CrossRef]
- Ramírez-Castillo, F.Y.; Loera-Muro, A.; Jacques, M.; Garneau, P.; Avelar-González, F.; Harel, J.; Guerrero-Barrera, A.L. Waterborne pathogens: Detection methods and challenges. Pathogens 2015, 4, 307–334. [Google Scholar] [CrossRef]
- Kadri, K. Polymerase chain reaction (PCR): Principle and applications. In Synthetic Biology—New Interdisciplinary Science; Nagpal, M.L., Boldura, O.M., Balta, C., Enany, S., Eds.; IntechOpen: London, UK, 2020. [Google Scholar] [CrossRef]
- Hryniszyn, A.; Skonieczna, M.; Wiszniowski, J. Methods for detection of viruses in water and wastewater. Adv. Microbiol. 2013, 3, 442–449. [Google Scholar] [CrossRef]
- Elnifro, E.M.; Ashshi, A.M.; Cooper, R.J.; Klapper, P.E. Multiplex PCR: Optimization and application in diagnostic virology. Clin. Microbiol. Rev. 2000, 13, 559–570. [Google Scholar] [CrossRef]
- Rodríguez, R.A.; Pepper, I.L.; Gerba, C.P. Application of PCR-based methods to assess the infectivity of enteric viruses in environmental samples. Appl. Environ. Microbiol. 2009, 75, 297–307. [Google Scholar] [CrossRef]
- Chen, H. Nucleic acid detection of major foodborne viral pathogens: Human noroviruses and hepatitis A virus. In Nucleic Acids—From Basic Aspects to Laboratory Tools; Larramendy, M., Soloneski, S., Eds.; IntechOpen: London, UK, 2016; pp. 36–57. [Google Scholar] [CrossRef]
- Kralik, P.; Ricchi, M. A basic guide to real time PCR in microbial diagnostics: Definitions, parameters, and everything. Front. Microbiol. 2017, 8, 108. [Google Scholar] [CrossRef]
- Srivastava, K.R.; Awasthi, S.; Mishra, P.K.; Srivastava, P.K. Biosensors/molecular tools for detection of waterborne pathogens. In Waterborne Pathogens—Detection and Treatment; Prasad, M.N.V., Grobelak, A., Eds.; Butterworth-Heinemann: Oxford, UK, 2020; pp. 237–277. [Google Scholar] [CrossRef]
- Watzinger, F.; Ebner, K.; Lion, T. Detection and monitoring of virus infections by real-time PCR. Mol. Asp. Med. 2006, 27, 254–298. [Google Scholar] [CrossRef]
- Singh, J.; Birbian, N.; Sinha, S.; Goswami, A. A critical review on PCR, its types and applications. Int. J. Adv. Res. Biol. Sci. 2014, 1, 65–80. [Google Scholar]
- Wagner, E.M. Monitoring gene expression: Quantitative real-time rt-PCR. Methods Mol. Biol. 2013, 1027, 19–45. [Google Scholar] [CrossRef]
- Hawkins, S.F.; Guest, P.C. Multiplex analyses using real-time quantitative PCR. Methods Mol. Biol. 2017, 1546, 125–133. [Google Scholar] [CrossRef]
- Lievens, A.; Jacchia, S.; Kagkli, D.; Savini, C.; Querci, M. Measuring digital PCR quality: Performance parameters and their optimization. PLoS ONE 2016, 11, e0153317. [Google Scholar] [CrossRef]
- Demeke, T.; Dobnik, D. Critical assessment of digital PCR for the detection and quantification of genetically modified organisms. Anal. Bioanal. Chem. 2018, 410, 4039–4050. [Google Scholar] [CrossRef]
- Neault, N.; Baig, A.T.; Graber, T.E.; D’Aoust, P.M.; Mercier, E.; Alexandrov, I.; Crosby, D.; Baird, S.; Mayne, J.; Pounds, T.; et al. SARS-CoV-2 Protein in Wastewater Mirrors COVID-19 Prevalence. medRxiv 2020. [Google Scholar] [CrossRef]
- Ongerth, J.E. RT qLAMP-Direct Detection of SARS-CoV-2 in Raw Sewage. medRxiv 2020. [Google Scholar] [CrossRef]
- Johnston, J.; Behrens, S. Seasonal Dynamics of the Activated Sludge Microbiome in Sequencing Batch Reactors, Assessed Using 16S rRNA Transcript Amplicon Sequencing. Appl. Environ. Microbiol. 2020, 86. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Li, Y.; Li, S.; Hu, N.; He, Y.; Pong, R.; Lin, D.; Lu, L.; Law, M. Comparison of next-generation sequencing systems. J. Biomed. Biotechnol. 2012, 2012, 251364. [Google Scholar] [CrossRef] [PubMed]
- Chan, A.W.; Naphtali, J.; Schellhorn, H.E. High-throughput DNA sequencing technologies for water and wastewater analysis. Sci. Prog. 2019, 102, 351–376. [Google Scholar] [CrossRef] [PubMed]
- Urban, L.; Holzer, A.; Baronas, J.J.; Hall, M.B.; Braeuninger-Weimer, P.; Scherm, M.J.; Kunz, D.J.; Perera, S.N.; Martin-Herranz, D.E.; Tipper, E.T.; et al. Freshwater monitoring by nanopore sequencing. eLife 2021, 10, e61504. [Google Scholar] [CrossRef] [PubMed]
- Acharya, K.; Blackburn, A.; Mohammed, J.; Haile, A.T.; Hiruy, A.M.; Werner, D. Metagenomic water quality monitoring with a portable laboratory. Water Res. 2020, 184, 116112. [Google Scholar] [CrossRef]
- Kemp, S.; Collier, D.; Datir, R.; Ferreira, I.; Gayed, S.; Jahun, A.; Hosmillo, M.; Rees-Spear, C.; Mlcochova, P.; Lumb, I.U.; et al. Neutralising antibodies in Spike mediated SARS-CoV-2 adaptation. Nature 2021, 592, 277–282. [Google Scholar] [CrossRef]
- Crits-Christoph, A.; Kantor, R.S.; Olm, M.R.; Whitney, O.N.; Al-Shayeb, B.; Lou, Y.C.; Flamholz, A.; Kennedy, L.C.; Greenwald, H.; Hinkle, A.; et al. Genome sequencing of sewage detects regionally prevalent SARS-CoV-2 variants. mBio 2021, 12, e02703–e02720. [Google Scholar] [CrossRef]
- Izquierdo-Lara, R.; Elsinga, G.; Heijnen, L.; Oude Munnink, B.B.; Schapendonk, C.M.E.; Nieuwenhuijse, D.; Kon, M.; Lu, L.; Aarestrup, F.M.; Lycett, S.; et al. Monitoring SdumkARS-CoV-2 circulation and diversity through community wastewater sequencing. medRxiv 2020. [Google Scholar] [CrossRef]
- Jahn, K.; Dreifuss, D.; Topolsky, I.; Kull, A.; Ganesanandamoorthy, P.; Fernandez-Cassi, X.; Bänziger, C.; Stachler, E.; Fuhrmann, L.; Jablonski, K.P.; et al. Detection of SARS-CoV-2 variants in Switzerland by genomic analysis of wastewater samples. Infect. Dis. 2021. preprint. [Google Scholar] [CrossRef]
- Dumke, R.; de la Cruz Barron, M.; Oertel, R.; Helm, B.; Kallies, R.; Berendonk, T.U.; Dalpke, A. Evaluation of two methods to concentrate SARS-CoV-2 from untreated wastewater. Pathogens 2021, 10, 195. [Google Scholar] [CrossRef]
- Bhalla, N.; Jolly, P.; Formisano, N.; Estrela, P. Introduction to biosensors. Essays Biochem. 2016, 60, 1. [Google Scholar] [CrossRef] [PubMed]
- Men, D.; Zhou, J.; Li, W.; Leng, Y.; Chen, X.; Tao, S.; Zhang, X.E. Fluorescent Protein Nanowire-Mediated Protein Microarrays for Multiplexed and Highly Sensitive Pathogen Detection. ACS Appl. Mater. Interfaces 2016, 8, 17472. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Anglès d‘Auriac, M.; Goggins, S.; Kasprzyk-Hordern, B.; Thomas, K.V.; Frost, C.G.; Estrela, P. A novel DNA biosensor using a ferrocenyl intercalator applied to the potential detection of human population biomarkers in wastewater. Environ. Sci. Technol. 2015, 49, 5609. [Google Scholar] [CrossRef] [PubMed]
- Pilevar, M.; Kim, K.T.; Lee, W.H. Recent advances in biosensors for detecting viruses in water and wastewater. J. Hazard. Mater. 2021, 410, 124656. [Google Scholar] [CrossRef]
- Velusamy, V.; Arshak, K.; Korostynska, O.; Oliwa, K.; Adley, C. An overview of foodborne pathogen detection: In the perspective of biosensors. Biotechnol. Adv. 2010, 28, 232–254. [Google Scholar] [CrossRef]
- Ahmed, A.; Rushworth, J.V.; Hirst, N.A.; Millner, P.A. Biosensors for whole-cell bacterial detection. Clin. Microbiol. Rev. 2014, 27, 631. [Google Scholar] [CrossRef]
- Mustafa, F.; Andreescu, S. Chemical and biological sensors for food-quality monitoring and smart packaging. Foods 2018, 7, 168. [Google Scholar] [CrossRef]
- Ashrafi, A.M.; Koudelkova, Z.; Sedlackova, E.; Richtera, L.; Adam, V. Electrochemical sensors and biosensors for determination of mercury ions. J. Electrochem. Soc. 2018, 165, B824–B834. [Google Scholar] [CrossRef]
- Peixoto, A.C.; Silva, A.F. 11—Smart devices: Micro- and nanosensors. In Bioinspired Materials for Medical Applications; Rodrigues, L., Mota, M., Eds.; Woodhead Publishing: Cambridge, UK, 2017; pp. 297–329. [Google Scholar]
- Thevenot, D.R.; Toth, K.; Durst, R.A.; Wilson, G.S. Electrochemical biosensors: Recommended definitions and classification. Pure Appl. Chem. 1999, 71, 2333–2348. [Google Scholar] [CrossRef]
- Ilkhani, H.; Farhad, S. A novel electrochemical DNA biosensor for Ebola virus detection. Anal. Biochem. 2018, 557, 151–155. [Google Scholar] [CrossRef] [PubMed]
- Saylan, Y.; Erdem, O.; Unal, S.; Denizli, A. An Alternative Medical Diagnosis Method: Biosensors for Virus Detection. Biosensors 2019, 9, 65. [Google Scholar] [CrossRef]
- Freitas, T.A.; Proenca, C.A.; Baldo, T.A.; Materon, E.M.; Wong, A.; Magnani, R.F.; Faria, R.C. Ultrasensitive immunoassay for detection of Citrus tristeza virus in citrus sample using disposable microfluidic electrochemical device. Talanta 2019, 205, 120110. [Google Scholar] [CrossRef]
- Ozer, T.; Geiss, B.J.; Henry, C.S. Chemical and Biological Sensors for Viral Detection. J. Electrochem. Soc. 2020, 167, 037523. [Google Scholar] [CrossRef]
- Han, J.H.; Lee, D.; Chew, C.H.C.; Kim, T.; Pak, J.J. A multi-virus detectable microfluidic electrochemical immunosensor for simultaneous detection of H1N1, H5N1, and H7N9 virus using ZnO nanorods for sensitivity enhancement. Sens. Actuators B Chem. 2016, 228, 36–42. [Google Scholar] [CrossRef]
- Kaya, S.I.; Karadurmus, L.; Ozcelikay, G.; Bakirhan, N.K.; Ozkan, S.A. Electrochemical virus detections with nanobiosensors. In Nanosensors for Smart Cities; Han, B., Tomer, V.K., Nguyen, T.A., Farmani, A., Kumar Singh, P., Eds.; Elsevier: Amsterdam, The Netherlands, 2020; pp. 303–326. [Google Scholar]
- Siuzdak, K.; Niedziałkowski, P.; Sobaszek, M.; Łęga, T.; Sawczak, M.; Czaczyk, E.; Dziabowska, K.; Ossowski, T.; Nidzworski, D.; Bogdanowicz, R. Biomolecular influenza virus detection based on the electrochemical impedance spectroscopy using the nanocrystalline boron-doped diamond electrodes with covalently bound antibodies. Sens. Actuators B Chem. 2019, 280, 263–271. [Google Scholar] [CrossRef]
- Anik, Ü.; Tepeli, Y.; Sayhi, M.; Nsiri, J.; Diouani, M.F. Towards the electrochemical diagnostic of influenza virus: Development of a graphene–Au hybrid nanocomposite modified influenza virus biosensor based on neuraminidase activity. Analyst 2018, 143, 150–156. [Google Scholar] [CrossRef] [PubMed]
- Shariati, M.; Ghorbani, M.; Sasanpour, P.; Karimizefreh, A. An ultrasensitive label free human papilloma virus DNA biosensor using gold nanotubes based on nanoporous polycarbonate in electrical alignment. Anal. Chim. Acta 2019, 1048, 31–41. [Google Scholar] [CrossRef]
- Cabral-Miranda, G.; Cardoso, A.R.; Ferreira, L.C.S.; Sales, M.G.F.; Bachmann, M.F. Biosensor-based selective detection of Zika virus specific antibodies in infected individuals. Biosens. Bioelectron. 2018, 113, 101–107. [Google Scholar] [CrossRef]
- Palomar, Q.; Gondran, C.; Marks, R.; Cosnier, S.; Holzinger, M. Impedimetric quantification of anti-dengue antibodies using functional carbon nanotube deposits validated with blood plasma assays. Electrochim. Acta 2018, 274, 84–90. [Google Scholar] [CrossRef]
- Lai, H.C.; Chin, S.F.; Pang, S.C.; Sum, H.; Sia, M.; Perera, D. Carbon nanoparticles based electrochemical biosensor strip for detection of Japanese Encephalitis Virus. J. Nanomat. 2017, 2017, 7. [Google Scholar] [CrossRef]
- Lee, T.; Park, S.Y.; Jang, H.; Kim, G.H.; Lee, Y.; Park, C.; Mohsen, M.; Lee, M.H.; Min, J. Fabrication of electrochemical biosensor consisted of multi-functional DNA structure/porous au nanoparticle for avian influenza virus (H5N1) in chicken serum. Mater. Sci. Eng. C 2019, 99, 511–519. [Google Scholar] [CrossRef]
- Zhang, Y.; Gao, Y.; Zhang, X.; Wang, H.; Xia, T.; Bian, C.; Liang, S.; Tng, X.; Wang, X. Electrochemical immunosensor for HBe antigen detection based on a signal amplification strategy: The co-catalysis of horseradish peroxidase and nanoporous gold. Sens. Actuators B Chem. 2019, 284, 296–304. [Google Scholar] [CrossRef]
- Hou, Y.H.; Wang, J.J.; Jiang, Y.Z.; Lv, C.; Xia, L.; Hong, S.L.; Lin, M.; Lin, Y.; Zhang, Z.L.; Pang, D.W. A colorimetric and electrochemical immunosensor for point-of-care detection of enterovirus 71. Biosens. Bioelectron. 2018, 99, 186–192. [Google Scholar] [CrossRef] [PubMed]
- Layqah, L.A.; Eissa, S. An electrochemical immunosensor for the corona virus associated with the Middle East respiratory syndrome using an array of gold nanoparticle-modified carbon electrodes. Microchim. Acta 2019, 186, 224–233. [Google Scholar] [CrossRef] [PubMed]
- Sayhi, M.; Ouerghi, O.; Belgacem, K.; Arbi, M.; Tepeli, Y.; Ghram, A.; Anik, U.; Osterlunf, L.; Laouini, D.; Diouani, M.F. Electrochemical detection of influenza virus H9N2 based on both immunomagnetic extraction and gold catalysis using an immobilization-free screen printed carbon microelectrode. Biosens. Bioelectron. 2018, 107, 170–177. [Google Scholar] [CrossRef]
- Gao, Z.; Li, Y.; Zhang, X.; Feng, J.; Kong, L.; Wang, P.; Chen, Z.; Dong, Y.; Wei, Q. Ultrasensitive electrochemical immunosensor for quantitative detection of HBeAg using Au@Pd/MoS2@MWCNTs nanocomposite as enzyme-mimetic labels. Biosens. Bioelectron. 2018, 102, 189–195. [Google Scholar] [CrossRef] [PubMed]
- Tu, H.; Lin, K.; Lun, Y.; Yu, L. Magnetic bead/capture DNA/glucose-loaded nanoliposomes for amplifying the glucometer signal in the rapid screening of hepatitis C virus RNA. Anal. Bioanal. Chem. 2018, 410, 3661–3669. [Google Scholar] [CrossRef]
- Ganser, L.R.; Kelly, M.L.; Herschlag, D.; Al-Hashimi, H.M. The roles of structural dynamics in the cellular functions of RNAs. Nat. Rev. Mol. Cell Biol. 2019, 20, 474–489. [Google Scholar] [CrossRef]
- Zhang, H.; Miller, B.L. Immunosensor-based label-free and multiplex detection of influenza viruses: State of the art. Biosens. Bioelectron. 2019, 141, 111476. [Google Scholar] [CrossRef] [PubMed]
- Xiao, T.; Huang, J.; Wang, D.; Meng, T.; Yang, X. Au and Au-Based nanomaterials: Synthesis and recent progress in electrochemical sensor applications. Talanta 2020, 206, 120210. [Google Scholar] [CrossRef] [PubMed]
- Faria, H.A.M.; Zucolotto, V. Label-free electrochemical DNA biosensor for zika virus identification. Biosens. Bioelectron. 2019, 131, 149–155. [Google Scholar] [CrossRef]
- Khater, M.; de la Escosura-Muñiz, A.; Quesada-González, D.; Merkoçi, A. Electrochemical detection of plant virus using gold nanoparticle-modified electrodes. Anal. Chim. Acta 2019, 1046, 123–131. [Google Scholar] [CrossRef] [PubMed]
- Malik, A.A.; Nantasenamat, C.; Piacham, T. Molecularly imprinted polymer for human viral pathogen detection. Mater. Sci. Eng. C 2017, 77, 1341–1348. [Google Scholar] [CrossRef] [PubMed]
- Babamiri, B.; Salimi, A.; Hallaj, R. A molecularly imprinted electrochemiluminescence sensor for ultrasensitive HIV-1 gene detection using EuS nanocrystals as luminophore. Biosens. Bioelectron. 2018, 117, 332–339. [Google Scholar] [CrossRef]
- Wangchareansak, T.; Thitithanyanont, A.; Chuakheaw, D.; Gleeson, M.P.; Lieberzeit, P.A.; Sangma, C. Influenza A virus molecularly imprinted polymers and their application in virus sub-type classification. J. Mater. Chem. B 2013, 1, 2190–2197. [Google Scholar] [CrossRef] [PubMed]
- Wangchareansak, T.; Thitithanyanont, A.; Chuakheaw, D.; Gleeson, M.P.; Lieberzeit, P.A.; Sangma, C. A novel approach to identify molecular binding to the influenza virus H5N1: Screening using molecularly imprinted polymers (MIPs). MedChemComm 2014, 5, 617–621. [Google Scholar] [CrossRef]
- Tai, D.F.; Lin, C.Y.; Wu, T.Z.; Chen, L.K. Recognition of dengue virus protein using epitope-mediated molecularly imprinted film. Anal. Chem. 2005, 77, 5140–5143. [Google Scholar] [CrossRef]
- Altintas, Z.; Pocock, J.; Thompson, K.A.; Tothill, I.E. Comparative investigations for adenovirus recognition and quantification: Plastic or natural antibodies? Biosens. Bioelectron. 2015, 74, 996–1004. [Google Scholar] [CrossRef] [PubMed]
- Jenik, M.; Schirhagl, R.; Schirk, C.; Hayden, O.; Lieberzeit, P.; Blaas, D.; Paul, G. Sensing picornaviruses using molecular imprinting techniques on a quartz crystal microbalance. Anal. Chem. 2009, 81, 5320–5326. [Google Scholar] [CrossRef]
- Lodder, W.; de Roda Husman, A.M. SARS-CoV-2 in wastewater: Potential health risk, but also data source. Lancet Gastroenterol. Hepatol. 2020, 5, 533–534. [Google Scholar] [CrossRef]
- Mao, K.; Zhang, H.; Yang, Z. Can a Paper-Based Device Trace COVID-19 Sources with Wastewater-Based Epidemiology? Environ. Sci. Technol. 2020, 54, 3733–3735. [Google Scholar] [CrossRef] [PubMed]
- Klug, K.E.; Reynolds, K.A.; Yoon, J.Y. A Capillary Flow Dynamics-Based Sensing Modality for Direct Environmental Pathogen Monitoring. Chem. Eur. J. 2018, 24, 6025–6029. [Google Scholar] [CrossRef] [PubMed]
- McCracken, K.E.; Tat, T.; Paz, V.; Reynolds, K.A.; Yoon, J.Y. Immunoagglutinated particle rheology sensing on a microfluidic paper-based analytical device for pathogen detection. In ASABE Annual International Meeting; American Society of Agricultural and Biological Engineers: St. Joseph, MI, USA, 2017. [Google Scholar] [CrossRef]
| SARS-CoV-2 Gene | Source | Date of Sample Collection | Ref. |
|---|---|---|---|
| N1, N2, N3, E | Netherlands | 5 February–25 March 2020 | [21] |
| E, N2 | Helsinki, Finland | 19–20 April and 24–25 May 2020 | [23] |
| N1, N2, N3 | Region of Murcia, Spain | 12 March–14 April 2020 | [31] |
| ORF1a, S, N1, N2 | Yamanashi Prefecture, Japan | 17 March–7 May 2020 | [32] |
| RdRp, E, N, M | Germany | 8–9 May 2020 | [41] |
| N1, N2, N3 | Massachusetts, USA | 18–25 March 2020 | [48] |
| N | Brisbane, Australia | 27 March–1 April 2020 | [49] |
| RdRp, E | Paris, France | 5 March–23 April 2020 | [50] |
| ORF1ab, E, N | Milan and Monza, Italy | 14 and 22 April 2020 | [52] |
| RdRp | Istanbul, Turkey | 7 May 2020 | [55] |
| ORF1ab, N | Bangladesh | 10 July–29 August 2020 | [71] |
| ORF1ab, N, S | Dubai, UAE | 22 and 28 April 2020 | |
| 4 May 2020 | [72] | ||
| 7 May–7 July 2020 | |||
| ORF1ab, N, S | Ahmedabad, India | 8 and 27 May 2020 | [73] |
| ORF1ab, S, RdRp | Milan and Rome, Italy | 3 February–2 April | [74] |
| E | Israel | 10 March–21 April 2020 | [75] |
| N1, N2 | Quito, Ecuador | 5 June 2020 | [76] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mackuľak, T.; Gál, M.; Špalková, V.; Fehér, M.; Briestenská, K.; Mikušová, M.; Tomčíková, K.; Tamáš, M.; Butor Škulcová, A. Wastewater-Based Epidemiology as an Early Warning System for the Spreading of SARS-CoV-2 and Its Mutations in the Population. Int. J. Environ. Res. Public Health 2021, 18, 5629. https://doi.org/10.3390/ijerph18115629
Mackuľak T, Gál M, Špalková V, Fehér M, Briestenská K, Mikušová M, Tomčíková K, Tamáš M, Butor Škulcová A. Wastewater-Based Epidemiology as an Early Warning System for the Spreading of SARS-CoV-2 and Its Mutations in the Population. International Journal of Environmental Research and Public Health. 2021; 18(11):5629. https://doi.org/10.3390/ijerph18115629
Chicago/Turabian StyleMackuľak, Tomáš, Miroslav Gál, Viera Špalková, Miroslav Fehér, Katarína Briestenská, Miriam Mikušová, Karolína Tomčíková, Michal Tamáš, and Andrea Butor Škulcová. 2021. "Wastewater-Based Epidemiology as an Early Warning System for the Spreading of SARS-CoV-2 and Its Mutations in the Population" International Journal of Environmental Research and Public Health 18, no. 11: 5629. https://doi.org/10.3390/ijerph18115629
APA StyleMackuľak, T., Gál, M., Špalková, V., Fehér, M., Briestenská, K., Mikušová, M., Tomčíková, K., Tamáš, M., & Butor Škulcová, A. (2021). Wastewater-Based Epidemiology as an Early Warning System for the Spreading of SARS-CoV-2 and Its Mutations in the Population. International Journal of Environmental Research and Public Health, 18(11), 5629. https://doi.org/10.3390/ijerph18115629







