Information Is Selection—A Review of Basics Shows Substantial Potential for Improvement of Digital Information Representation
Abstract
1. Introduction
2. Definition of Information
3. Global Definition of Information
3.1. Literature Research
3.2. Format of the Domain Vector (DV)
- The UL has a similar function to a link (i.e., a URL resp. “Uniform Resource Locator”) [6], but allows for maximal efficiency. It is a number sequence and typically has a hierarchical structure with a predefined meaning, where the first number represents the count of the subsequent numbers of the UL and the second number points to a global table of conventional internet addresses of online presences, where users can define DVs online in a standardized way. Subsequent numbers in the UL can provide detailed addresses within the chosen online presence;
- Numbers in the UL are self-elongating positive integers, starting with a half byte or byte, as shown in Figure 4 of [2];
- The number sequence after the UL is completely defined in the online definition of the online address given by the UL. This is also a metric (i.e., a distance function; see Section 4.4) for a similarity comparison of DVs when the UL is provided and the online definition is expandable. Necessary explanations and definitions are, at least, given in English, but should be language-independent, such that translation into other languages (i.e., multilingual definitions) is possible;
- Nesting and a posteriori combinations of DVs are possible and often efficient (e.g., date, time, and location, along with a sequence of measurement results);
- The binary format of the DV can be converted into a text-compatible form using, for example, the Base64 Data Encoding specification (RFC 4648) [7]. After this, it can be integrated into currently recommended approaches (e.g., into the FHIR resp. “Fast Health Interoperability Resources" standard [8]) as an extension [9].
4. Comparison of Information
4.1. Domain of Information: “Language Vocabulary”
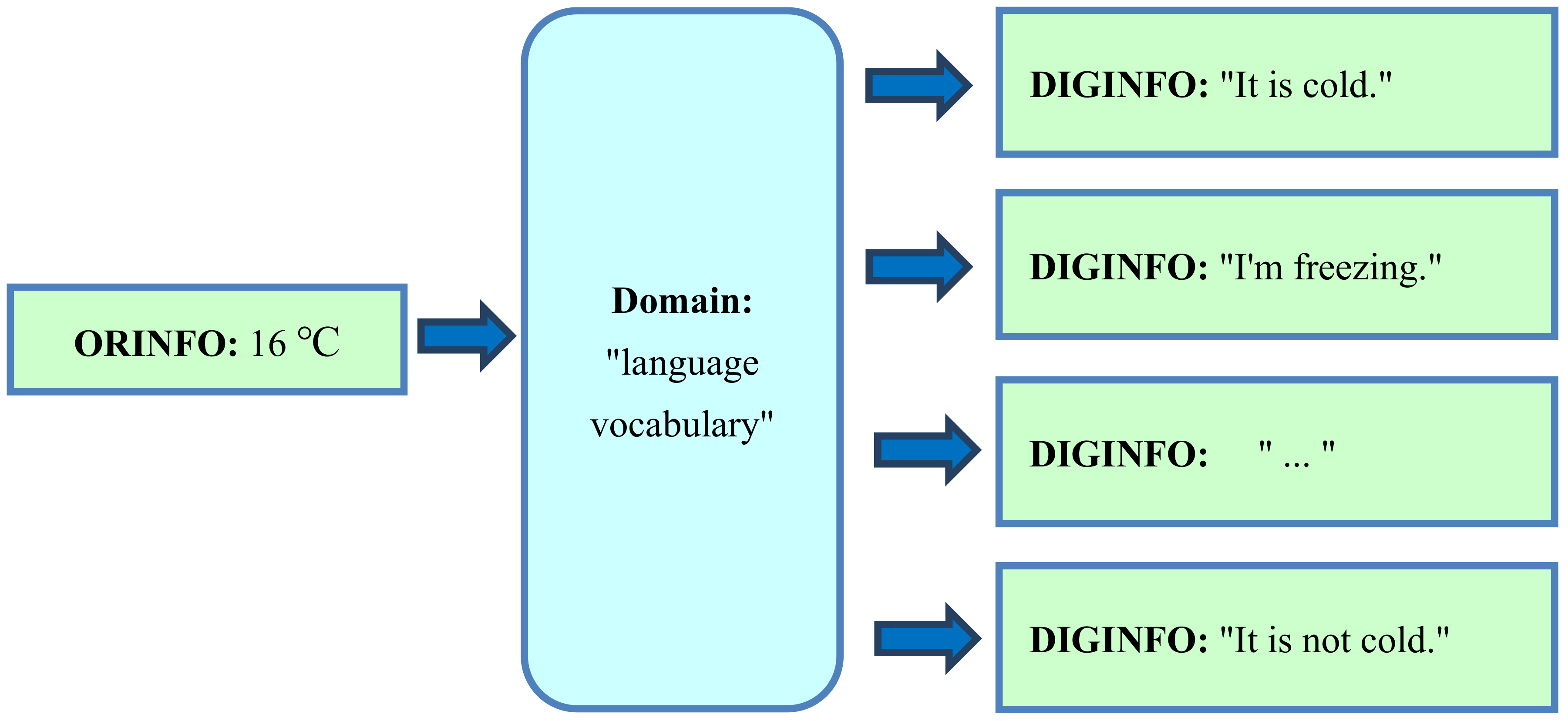
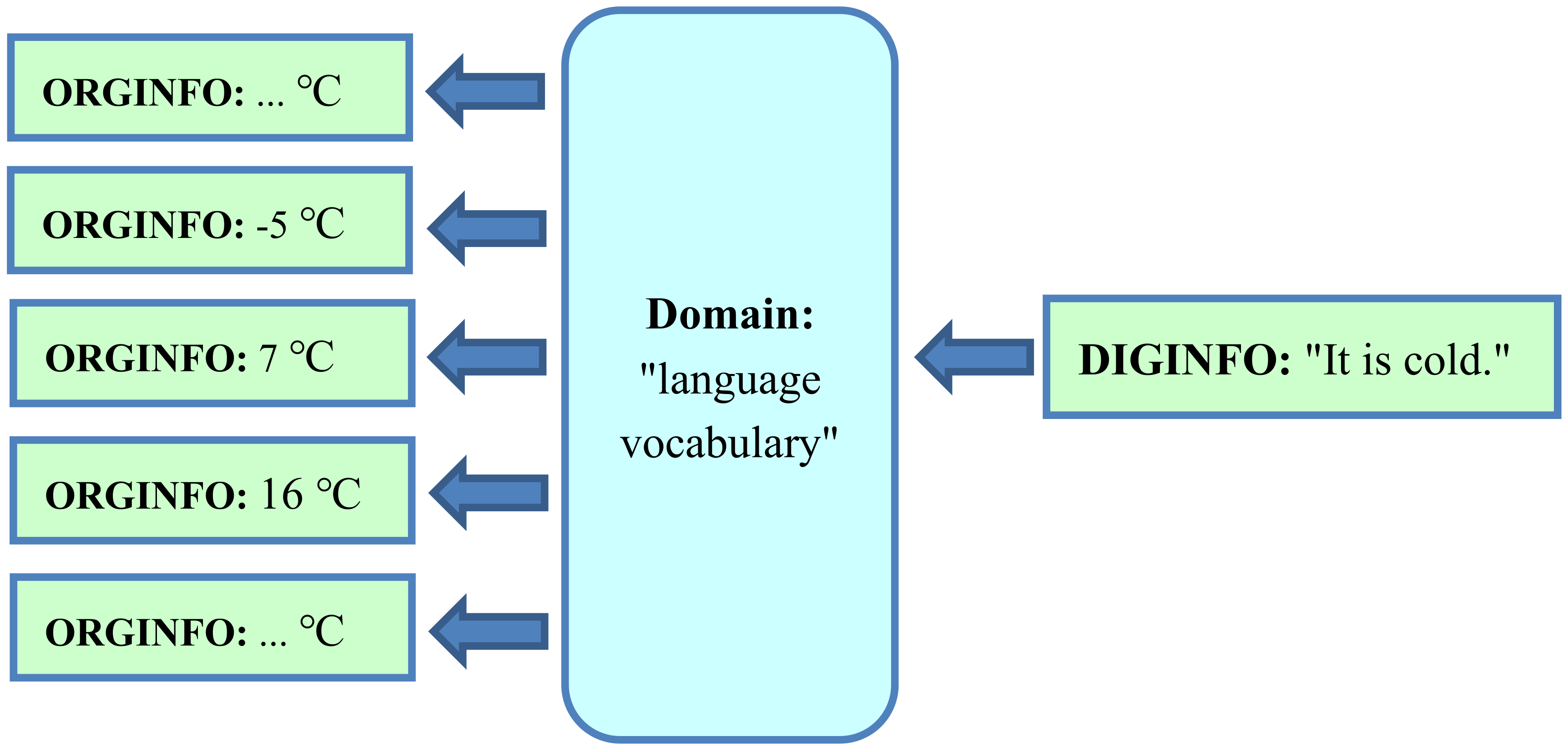
4.2. Translation of Original Information into Digital Representation using the Domain “Language Vocabulary”
4.3. Domain of Information: Adapted to the Topic
- (a)
- Which (additional) independent feature (parameter) is relevant within the chosen topic? If an appropriate quantification of this feature is available online, reuse it; otherwise, ask:
- (b)
- Which variants of the feature are possible? Quantify the feature, order its variants, and define a bijection to the numeric values of a parameter with the corresponding order.
4.4. Comparability of Information
- F(DV1, DV2) ≥ 0,
- F(DV1, DV2) = 0 if and only if DV1 = DV2,
- F(DV1, DV2) + F(DV2, DV3) ≥ F(DV1, DV3), and
- F(DV1, DV2) = F(DV2, DV1).
4.5. Domains of Information in Databases
5. Search of Information
5.1. Text Search of Information
5.2. Search of Information in Databases
5.3. Search of Information in General
5.4. Similarity Search of Information
5.5. User-Defined Global Similarity Search of Information
- (a)
- Ask for relevant features within the chosen topic;
- (b)
- Quantify them, reusing already existing online definitions.
5.6. Medical Example
6. Discussion
6.1. Comparison with Current Approaches
6.2. User Defined Similarity Search of Medical Information
6.3. Urgent Questions in Information Science and Informatics
- Why has the exact definition of information as a selection from an ordered set (or domain) (1)not been consequently emphasized and technically utilized from the beginning? This is far-reaching, as adapted domains can be defined online for all possible applications (Figure 3). If it is unclear how to define an ordered set (i.e., domain) and the numbers that select from this set, advanced training (e.g., study of the medical example in Section 5.6) is necessary—information experts (by definition) need to know about this. A “language vocabulary” is only one example of a domain. Semantic concepts and other a posteriori combinations of information are derived applications and also need a basis.
- Why can users not (especially professionals, experts, and specialists) define adapted domains (Section 4.3) online for precise language-independent global communication in their areas of expertise?
- Digital information consists of number sequences. Why have these, up to now, been defined in variable and complex ways by context? Why have globally defined, identified, and searchable information carriers (such as the domain vectors detailed above (2), up to now, not been introduced (as selections from an online defined and adapted domain), decades after the introduction of the internet?
- Why are global information searches still essentially restricted to text searching?
7. Conclusions
Conflicts of Interest
References
- Dirac, P.A.M. The Principles of Quantum Mechanics (No. 27); Oxford University Press: Oxford, UK, 1981. [Google Scholar]
- Orthuber, W. Global predefinition of digital information. Digit. Med. 2018, 4, 148. [Google Scholar] [CrossRef]
- Orthuber, W. Online definition of comparable and searchable medical information. Digit. Med. 2018, 4, 77. [Google Scholar] [CrossRef]
- Orthuber, W.; Hasselbring, W. Proposal for a New Basic Information Carrier on the Internet: URL Plus Number Sequence. In Proceedings of the 15th International Conference WWW/Internet, Mannheim, Germany, 28–30 October 2016; pp. 279–284. [Google Scholar]
- Harzing, A.W.K.; Van der Wal, R. Google Scholar as a new source for citation analysis. Ethics Sci. Environ. Polit. 2008, 8, 61–73. [Google Scholar] [CrossRef]
- Fielding, R.; Gettys, J.; Mogul, J.; Frystyk, H.; Masinter, L.; Leach, P.; Berners-Lee, T. Hypertext transfer protocol–HTTP/1.1. 1999. Available online: http://www.hjp.at/doc/rfc/rfc2616.html (accessed on 23 April 2020).
- Josefsson, S. The base16, base32, and base64 data encodings. RFC 4648. October 2006, pp. 1–18. Available online: https://www.hjp.at/doc/rfc/rfc4648.html (accessed on 23 April 2020).
- Bender, D.; Sartipi, K. HL7 FHIR: An Agile and RESTful approach to healthcare information exchange. In Proceedings of the 26th IEEE international symposium on computer-based medical systems, Porto, Portugal, 20–22 June 2013; pp. 326–331. [Google Scholar]
- HL7 FHIR Release 4, Extensibility. Available online: https://www.hl7.org/fhir/extensibility.html (accessed on 27 January 2020).
- Zezula, P.; Amato, G.; Dohnal, V.; Batko, M. Similarity Search: The Metric Space Approach; Springer Science & Business Media, Inc.: New York, NY, USA, 2006; Volume 32. [Google Scholar]
- Hamet, P.; Tremblay, J. Artificial intelligence in medicine. Metabolism 2017, 69, 36–40. [Google Scholar] [CrossRef] [PubMed]
- Henderson, T.; Shepheard, J.; Sundararajan, V. Quality of diagnosis and procedure coding in ICD-10 administrative data. Med Care 2006, 44, 1011–1019. [Google Scholar] [CrossRef] [PubMed]
- Southern, D.; Eastwood, C.; Quan, H.; Ghali, W. Enhancing description of hospital-conditions with ICD-11 cluster coding: Better codes for monitoring and prevention. Int. J. Popul. Data Sci. 2018, 3. [Google Scholar] [CrossRef]
- Mazurek, M. Indexing the NoSQL Repository of Medical Records with Ontology Concepts. Coll. Econ. Anal. Ann. 2018, 52, 71–82. [Google Scholar]
- Ryan, A. Towards semantic interoperability in healthcare: Ontology mapping from SNOMED-CT to HL7 version 3. In Proceedings of the Second Australasian Workshop on Advances in Ontologies, Hobart, Australia, 5 December 2006; Australian Computer Society, Inc.: Darlinghurst, Australian, 2006; Volume 72, pp. 69–74. [Google Scholar]
- Deshpande, P.; Rasin, A.; Brown, E.T.; Furst, J.; Montner, S.M.; Armato III, S.G.; Raicu, D.S. Augmenting medical decision making with text-based search of teaching file repositories and medical ontologies: Text-based search of radiology teaching files. Int. J. Knowl. Discov. Bioinform. (Ijkdb) 2018, 8, 18–43. [Google Scholar] [CrossRef]
- Lu, W.; Hou, J.; Yan, Y.; Zhang, M.; Du, X.; Moscibroda, T. MSQL: Efficient similarity search in metric spaces using SQL. Vldb J. 2017, 26, 829–854. [Google Scholar] [CrossRef]
- Keim, D.A. Efficient geometry-based similarity search of 3D spatial databases. ACM Sigmod Rec. 1999, 28, 419–430. [Google Scholar] [CrossRef]
- Negrel, R.; Picard, D.; Gosselin, P.H. Compact tensor based image representation for similarity search. In Proceedings of the 19th IEEE International Conference on Image Processing, Orlando, FL, USA, 30 September–3 October 2012; pp. 2425–2428. [Google Scholar]
- Moise, D.; Shestakov, D.; Gudmundsson, G.; Amsaleg, L. Terabyte-scale image similarity search: Experience and best practice. In Proceedings of the 2013 IEEE International Conference on Big Data, Silicon Valley, CA, USA, 6–9 October 2013; pp. 674–682. [Google Scholar]
- Stanchev, P.; Amato, G.; Falchi, F.; Gennaro, C.; Rabitti, F.; Savino, P. Selection of MPEG-7 image features for improving image similarity search on specific data sets. In Proceedings of the 7-th IASTED International Conference on Computer Graphics and Imaging, CGIM 2004, Kauai, HI, USA, 17–19 August 2004; pp. 395–400. [Google Scholar]
- Uysal, M.S.; Beecks, C.; Schmücking, J.; Seidl, T. Efficient similarity search in scientific databases with feature signatures. In Proceedings of the 27th International Conference on Scientific and Statistical Database Management, La Jolla, CA, USA, 29 June–1 July 2015; pp. 1–12. [Google Scholar]
- Harmsen, D.; Rothgänger, J.; Frosch, M.; Albert, J. RIDOM: Ribosomal differentiation of medical micro-organisms database. Nucleic Acids Res. 2002, 30, 416–417. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Petrakis, E.G.M.; Faloutsos, A. Similarity searching in medical image databases. IEEE Trans. Knowl. Data Eng. 1997, 9, 435–447. [Google Scholar] [CrossRef]
- Korn, P.; Sidiropoulos, N.D.; Faloutsos, C.; Siegel, E.L.; Protopapas, Z. Fast and effective similarity search in medical tumor databases using morphology. In Multimedia Storage and Archiving Systems; International Society for Optics and Photonics: Bellingham, WA, USA, 1996; Volume 2916, pp. 116–129. [Google Scholar]
- Wichterich, M.; Kranen, P.; Assent, I.; Seidl, T. Efficient EMD-based Similarity Search in Medical Image Databases. Sci. Eng. Biol. Inform. 2010, 6, 175–201. [Google Scholar]
- NumericSearch. Available online: http://numericsearch.com (accessed on 29 October 2019).
- Bustos, B.; Keim, D.A.; Saupe, D.; Schreck, T.; Vranić, D.V. Feature-based similarity search in 3D object databases. Acm Comput. Surv. 2005, 37, 345–387. [Google Scholar] [CrossRef]
- Yan, Z.; Dijkman, R.; Grefen, P. Fast business process similarity search with feature-based similarity estimation. In OTM Confederated International Conferences “On the Move to Meaningful Internet Systems”; Springer: Berlin/Heidelberg, Germany, 2010; pp. 60–77. [Google Scholar]
- Kriegel, H.P.; Brecheisen, S.; Kröger, P.; Pfeifle, M.; Schubert, M. Using sets of feature vectors for similarity search on voxelized CAD objects. In Proceedings of the 2003 ACM SIGMOD International Conference on Management of Data, San Diego, CA, USA, 9–12 June 2003; pp. 587–598. [Google Scholar]
- Yan, X.; Zhu, F.; Yu, P.S.; Han, J. Feature-based similarity search in graph structures. Acm Trans. Database Syst. 2006, 31, 1418–1453. [Google Scholar] [CrossRef]
- Liu, L.; Yu, M.; Shao, L. Unsupervised local feature hashing for image similarity search. IEEE Trans. Cybern. 2015, 46, 2548–2558. [Google Scholar] [CrossRef]
- Jurado, E.; Barrena, M. Efficient similarity search in feature spaces with the Q-tree. In Proceedings of the East European Conference on Advances in Databases and Information Systems, Bratislava, Slovakia, 8–11 September 2002; pp. 177–190. [Google Scholar]
- Abolmaali, S.B.; Ostermann, C.; Zell, A. The Compressed Feature Matrix—A novel descriptor for adaptive similarity search. J. Mol. Model. 2003, 9, 66–75. [Google Scholar] [CrossRef]
- Precioso, F.; Cord, M.; Gorisse, D.; Thome, N. Efficient bag-of-feature kernel representation for image similarity search. In Proceedings of the 2011 18th IEEE International Conference on Image Processing, Brussels, Belgium, 11–14 September 2011; pp. 109–112. [Google Scholar]
- Gennaro, C.; Mordacchini, M.; Orlando, S.; Rabitti, F. A Scalable Distributed Data Structure for Multi-Feature Similarity Search. In Proceedings of the Sixteenth Italian Symposium on Advanced Database Systems, SEBD 2008, Mondello, Italy, 22–25 June 2008; pp. 302–309. [Google Scholar]
- Song, K.T.; Nam, H.J.; Chang, J.W. A cell-based index structure for similarity search in high-dimensional feature spaces. In Proceedings of the 2001 ACM symposium on Applied computing, Las Vegas, NV, USA, 11–14 March 2001; pp. 264–268. [Google Scholar]
- Lv, Q.; Josephson, W.; Wang, Z.; Charikar, M.; Li, K. Ferret: A toolkit for content-based similarity search of feature-rich data. Acm Sigops Oper. Syst. Rev. 2006, 40, 317–330. [Google Scholar] [CrossRef]
- Benson, T.; Grieve, G. Principles of Health Interoperability: SNOMED CT, HL7 and FHIR; Springer: Berlin/Heidelberg, Germany, 2016. [Google Scholar]
- Bhargava, A.; Kim, T.; Quine, D.B.; Hauser, R.G. A 20-Year Evaluation of LOINC in the United States’ Largest Integrated Health System. Arch. Pathol. Lab. Med. 2020, 144, 478–484. [Google Scholar] [CrossRef]
- Saripalle, R.; Runyan, C.; Russell, M. Using Hl7 FHIR to achieve interoperability in patient health record. J. Biomed. Inform. 2019, 94, 103188. [Google Scholar] [CrossRef] [PubMed]
- HL7 FHIR Release 4, Observation-example-f001-glucose.xml. Available online: https://www.hl7.org/fhir/observation-example-f001-glucose.xml.html (accessed on 29 October 2019).
- Orthuber, W.; Fiedler, G.; Kattan, M.; Sommer, T.; Fischer-Brandies, H. Design of a global medical database which is searchable by human diagnostic patterns. Open Med. Inform. J. 2008, 2, 21–31. [Google Scholar] [CrossRef] [PubMed]
- Orthuber, W.; Sommer, T. A searchable patient record database for decision support. Medical Informatics in a United and Healthy Europe. In Proceedings of the MIE 2009, The XXIInd International Congress of the European Federation for Medical Informatics, Sarajevo, Bosnia and Herzegovina, 30 August–2 September 2009; pp. 584–588. [Google Scholar]
- Orthuber, W.; Dietze, S. Towards Standardized Vectorial Resource Descriptors on the Web. In Proceedings of the Informatik 2010: Service Science—Neue Perspektiven für die Informatik, Beiträge der 40, Jahrestagung der Gesellschaft für Informatik e.V. (GI), Band 2, Leipzig, Germany, 27 September–1 October 2010; pp. 453–458. [Google Scholar]
- Orthuber, W.; Papavramidis, E. Standardized vectorial representation of medical data in patient records. Med. Care Compunetics 2010, 6, 153–166. [Google Scholar]




| ORGINFO | DIGINFO | F = |DIGINFO-16| | RANK |
|---|---|---|---|
| 16 °C | 16 | 0 | 1 |
| 15 °C | 15 | 1 | 2 |
| 14 °C | 14 | 2 | 3 |
| 18 °C | 18 | 2 | 4 |
| 0 °C | 0 | 16 | 5 |
| −5 °C | −5 | 21 | 6 |
| 45 °C | 45 | 29 | 7 |
Line Code 01 <?xml version="1.0" encoding="UTF-8"?> 02 <Observation xmlns="http://hl7.org/fhir"> 03 <code> 04 <coding> 05 <system value="http://loinc.org"/> 06 <code value="15074-8"/> 07 <display value="Glucose [Moles/volume] in Blood"/> 08 </coding> 09 </code> 10 <issued value="2013-04-03T15:30:10+01:00"/> 11 <valueQuantity> 12 <value value="6.3"/> 13 <unit value="mmol/l"/> 14 <system value="http://unitsofmeasure.org"/> 15 <code value="mmol/L"/> 16 </valueQuantity> 17 <interpretation> 18 <coding> 19 <system value="http://terminology.hl7.org/CodeSystem/ 20 v3-ObservationInterpretation"/> 21 <code value="H"/> 22 <display value="High"/> 23 </coding> 24 </interpretation> 25 <referenceRange> 26 <low> 27 <value value="3.1"/> 28 <unit value="mmol/l"/> 29 <system value="http://unitsofmeasure.org"/> 30 <code value="mmol/L"/> 31 </low> 32 <high> 33 <value value="6.2"/> 34 <unit value="mmol/l"/> 35 <system value="http://unitsofmeasure.org"/> 36 <code value="mmol/L"/> 37 </high> 38 </referenceRange> 39 </Observation> |
© 2020 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Orthuber, W. Information Is Selection—A Review of Basics Shows Substantial Potential for Improvement of Digital Information Representation. Int. J. Environ. Res. Public Health 2020, 17, 2975. https://doi.org/10.3390/ijerph17082975
Orthuber W. Information Is Selection—A Review of Basics Shows Substantial Potential for Improvement of Digital Information Representation. International Journal of Environmental Research and Public Health. 2020; 17(8):2975. https://doi.org/10.3390/ijerph17082975
Chicago/Turabian StyleOrthuber, Wolfgang. 2020. "Information Is Selection—A Review of Basics Shows Substantial Potential for Improvement of Digital Information Representation" International Journal of Environmental Research and Public Health 17, no. 8: 2975. https://doi.org/10.3390/ijerph17082975
APA StyleOrthuber, W. (2020). Information Is Selection—A Review of Basics Shows Substantial Potential for Improvement of Digital Information Representation. International Journal of Environmental Research and Public Health, 17(8), 2975. https://doi.org/10.3390/ijerph17082975




