Autism Spectrum Disorder Diagnostic System Using HOS Bispectrum with EEG Signals
Abstract
1. Introduction
2. Data Used
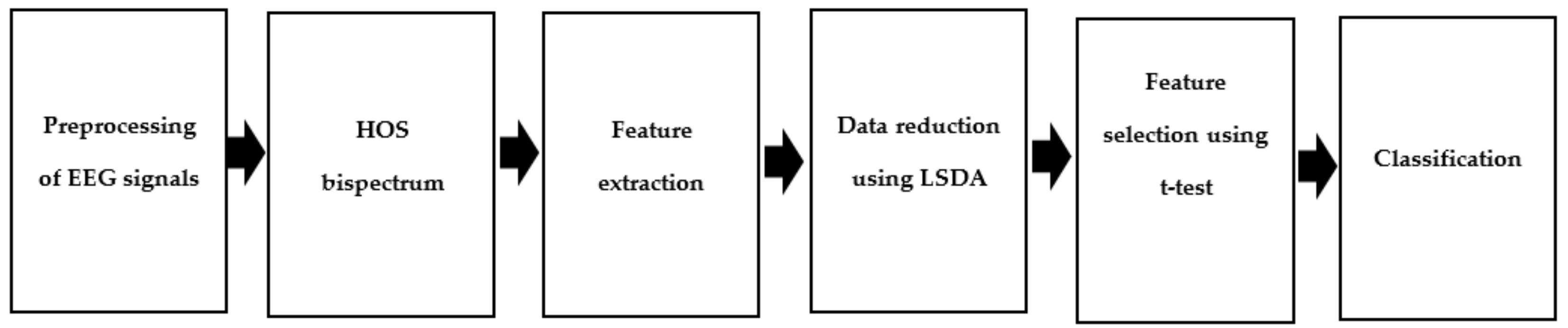
3. Methodology
3.1. Recording and Pre-Processing of Signals
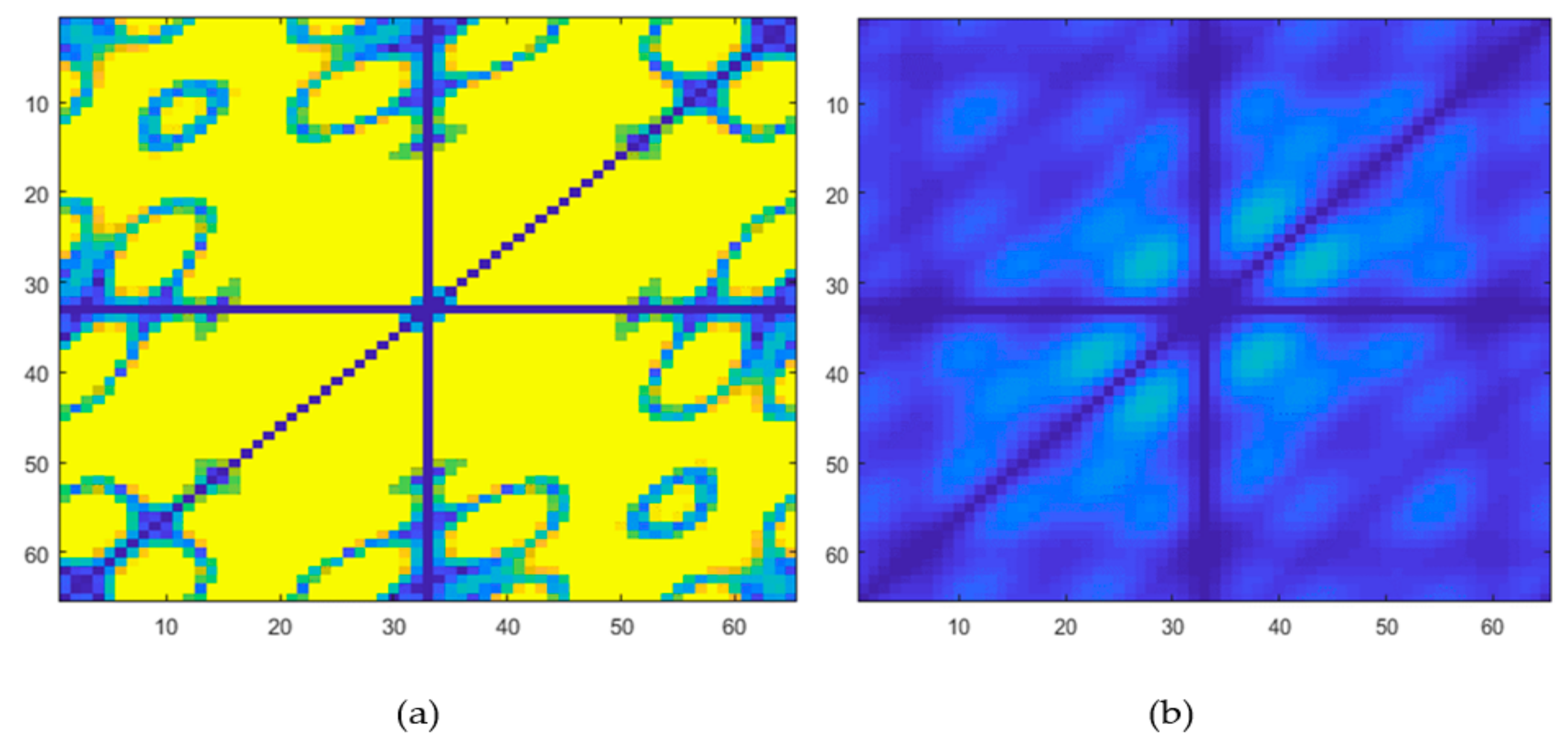
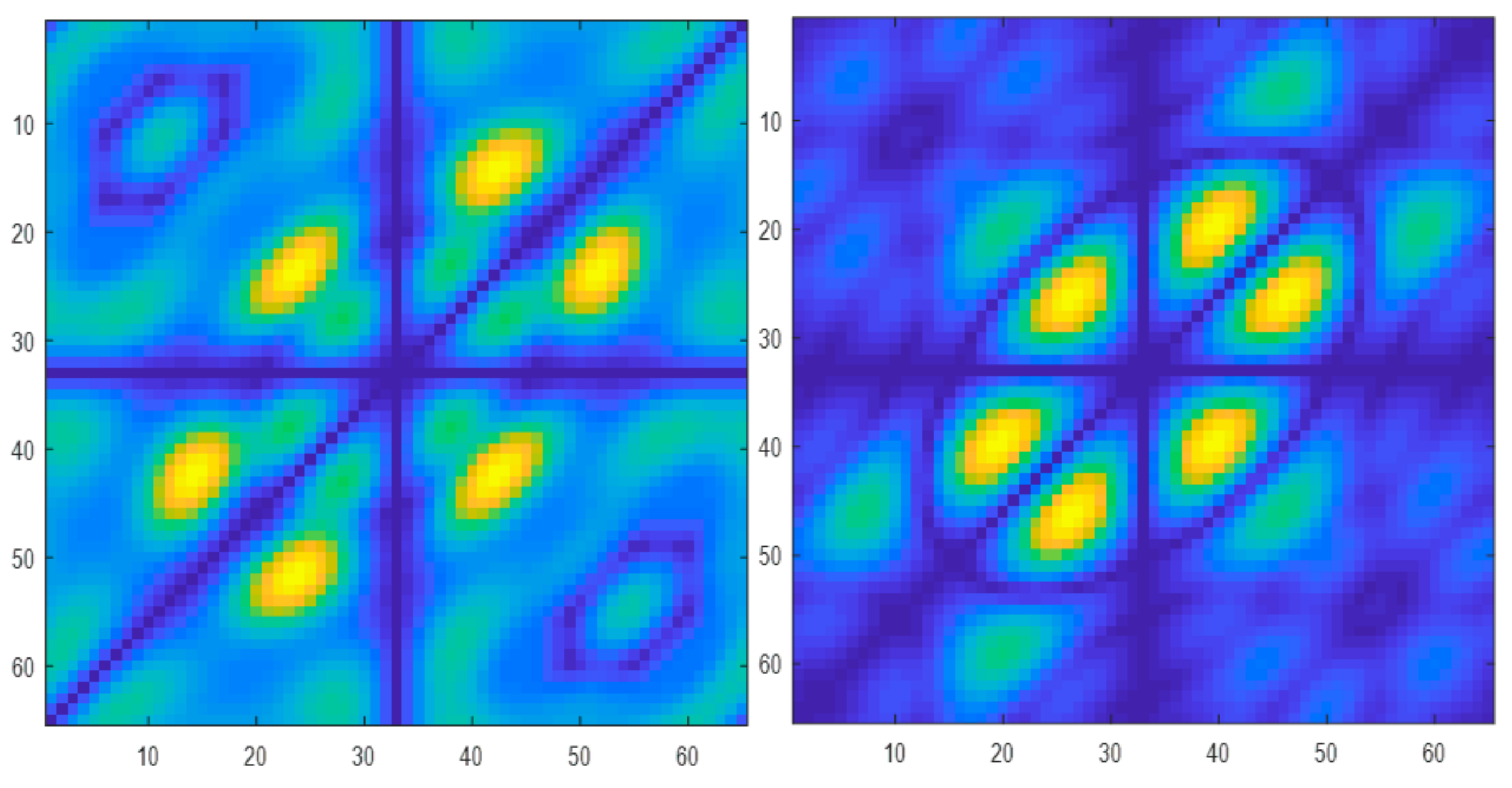
3.2. HOS Bispectrum
3.3. Feature Extraction
Description of Features
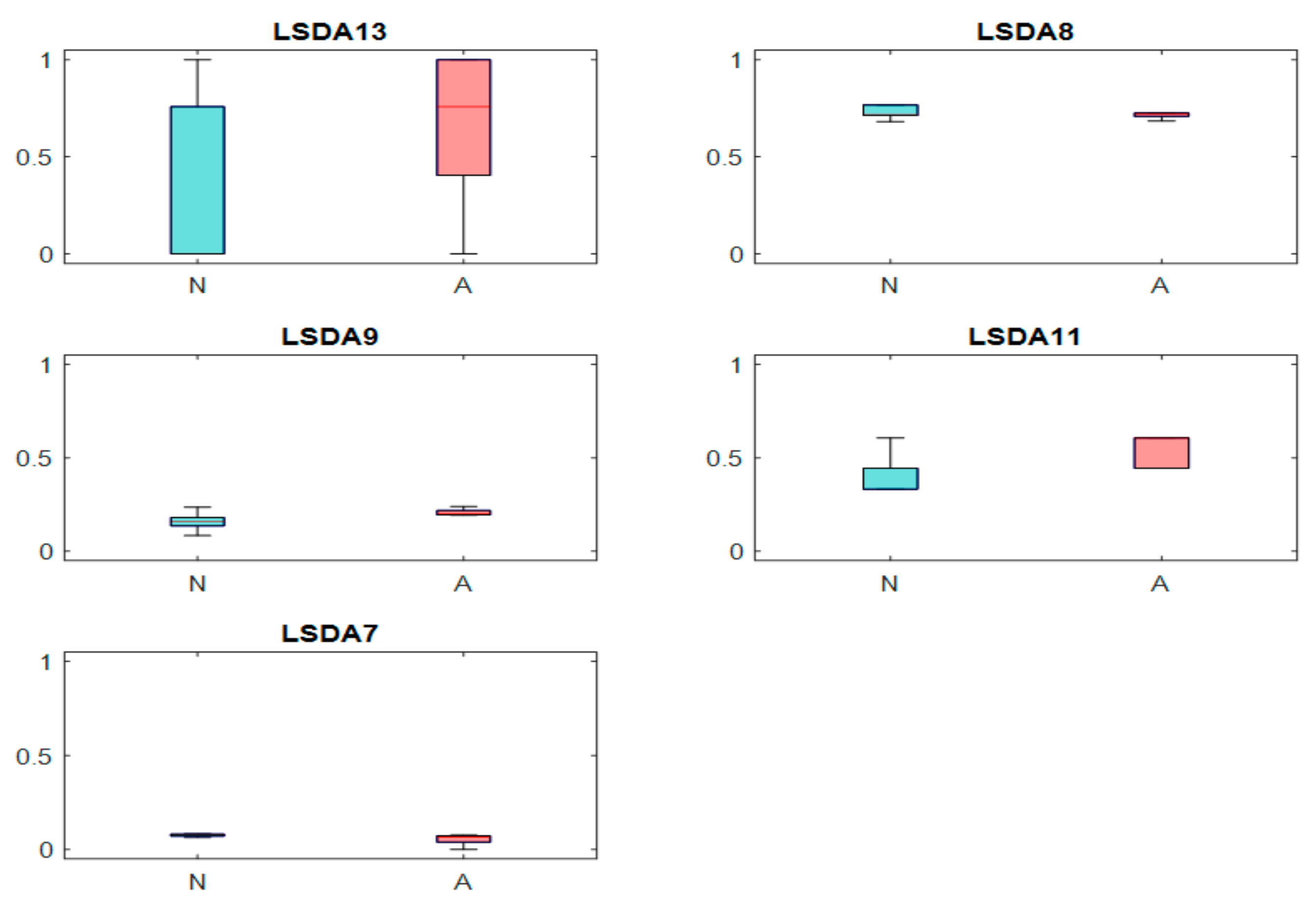
3.4. Feature Reduction and Selection
3.5. Classification
4. Results
5. Discussion
- Benefits:
- The recommended technique allows for rapid and accurate diagnosis of ASD.
- The diagnostic method is non-invasive.
- The method is promising, as the model used has been validated by 10-fold validation.
- Drawbacks:
- Feature extraction and selection processes are done manually.
- This technique only supports a small data size; thus, sizeable data cannot be studied for early detection.
6. Summary
7. Future work
Author Contributions
Funding
Conflicts of Interest
References
- Kim, D.G.; Park, H.R.; Lee, J.M.; Moon, H.E.; Lee, D.S.; Kim, B.N.; Kim, J.; Paek, S.H. A short review on the current understanding of autism spectrum disorders. Exp. Neurobiol. 2016, 25, 1–13. [Google Scholar]
- Gharani, N.; Benayed, R.; Mancuso, V.; Brzustowicz, L.M.; Millonig, J.H. Association of the homeobox transcription factor, ENGRAILED 2, 3, with autism spectrum disorder. Mol. Psychiatry 2004, 9, 474–484. [Google Scholar] [CrossRef] [PubMed]
- Miles, J.H. Autism spectrum disorders-A genetics review. Genet. Med. 2011, 13, 278–294. [Google Scholar] [CrossRef] [PubMed]
- Kern, J.K.; Jones, A.M. Evidence of toxicity, oxidative stress, and neuronal insult in autism. J. Toxicol. Environ. Health B Crit. Rev. 2007, 7404, 485–499. [Google Scholar] [CrossRef] [PubMed]
- Kolevzon, A.R.; Raz Gross, A. Prenatal and perinatal risk factors for autism. Arch. Pediatrics Adolesc. Med. 2007, 161, 326–333. [Google Scholar] [CrossRef]
- NIH. Autism Spectrum Disorder Fact Sheet. Available online: https://www.ninds.nih.gov/Disorders/Patient-Caregiver-Education/Fact-Sheets/Autism-Spectrum-Disorder-Fact-Sheet (accessed on 31 January 2020).
- Fisch, G.S. Erratum to In the article by Gene S. Fisch, entitled “Nosology and Epidemiology in Autism: Classification Counts” in. American Journal of Medical Genetics Part C. Am. J. Med. Genet. Part A 2013, 161, 2399. [Google Scholar] [CrossRef]
- Robinson, E.B.; Lichtenstein, P.; Anckarsäter, H.; Happé, F.; Ronald, A. Examining and interpreting the female protective effect against autistic behavior. Proc. Natl. Acad. Sci. USA 2013, 110, 5258–5262. [Google Scholar] [CrossRef]
- Mattila, M.L.; Kielinen, M.; Linna, S.L.; Jussila, K.; Ebeling, H.; Bloigu, R.; Joseph, R.M.; Moilanen, I. Autism spectrum disorders according to DSM-IV-TR and comparison with DSM-5 draft criteria: An epidemiological study. J. Am. Acad. Child Adolesc. Psychiatry 2011, 50, 583–592. [Google Scholar] [CrossRef]
- Leventhal, B.L.; Kim, Y.S.; Koh, Y.J.; Fombonne, E.; Laska, E.; Lim, E.C.; Cheon, K.A.; Kim, S.J.; Kim, Y.K.; Lee, H. Prevalence of autism spectrum disorder in a total population sample. Am. J. Psychiatry 2011, 168, 904–912. [Google Scholar]
- Falkmer, T.; Anderson, K.; Falkmer, M.; Horlin, C. Diagnostic procedures in autism spectrum disorders: A systematic literature review. Eur. Child Adolesc. Psychiatry 2013, 22, 329–340. [Google Scholar] [CrossRef]
- Zalla, T.; Sperduti, M. The Amygdala and the Relevance Detection Theory of Autism: An Evolutionary Perspective. Front. Hum. Neurosci. 2013, 7, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Howard, M.A.; Cowell, P.E.; Boucher, J.; Broks, P.; Mayes, A.; Farrant, A.; Roberts, N. Convergent neuroanatomical and behavioural evidence of an amygdala hypothesis of autism. Neuroreport 2000, 11, 2931–2935. [Google Scholar] [CrossRef] [PubMed]
- Abell, F.; Krams, M.; Ashburner, J.; Passingham, R.; Friston, K.; Frackowiak, R.; Happe, F.; Frith, C.; Frith, U. The neuroanatomy of autism: A voxel-based whole brain analysis of structural scans. Sci. Ment. Heal. Vol. 2 Autism 2013, 10, 123–125. [Google Scholar] [CrossRef] [PubMed]
- Nam, D.; Barrack, R.L.; Potter, H.G. What are the advantages and disadvantages of imaging modalities to diagnose wear-related corrosion problems? Clin. Orthop. Relat. Res. 2014, 472, 3665–3673. [Google Scholar] [CrossRef]
- Salmon, E.; Bernard Ir, C.; Hustinx, C. Pitfalls and limitations of PET/CT in brain imaging. Semin. Nucl. Med. 2015, 45, 541–551. [Google Scholar] [CrossRef]
- Acharya, U.R.; Oh, S.L.; Hagiwara, Y.; Tan, J.H.; Adeli, H.; Subha, D.P. Automated EEG-based screening of depression using deep convolutional neural network. Comput. Methods Programs Biomed. 2018, 161, 103–113. [Google Scholar] [CrossRef]
- Acharya, U.R. Characterization of focal EEG signals: A review. Futur. Gener. Comput. Syst. 2019, 91, 290–299. [Google Scholar] [CrossRef]
- Jahmunah, V.; Oh, S.L.; Rajinikanth, V.; Ciaccio, E.J.; Cheong, K.H.; Arunkumar, N.; Acharya, U.R. Automated detection of schizophrenia using nonlinear signal processing methods. Artif. Intell. Med. 2019, 100, 101698. [Google Scholar] [CrossRef]
- Bhat, S.; Acharya, U.R.; Adeli, H.; Bairy, G.M.; Adeli, A. Autism: Cause factors, early diagnosis and therapies. Rev. Neurosci. 2014, 25, 841–850. [Google Scholar] [CrossRef]
- Hadoush, H.; Alafeef, M.; Abdulhay, E. Automated identification for autism severity level: EEG analysis using empirical mode decomposition and second order difference plot. Behav. Brain Res. 2019, 362, 240–248. [Google Scholar] [CrossRef]
- Oh, S.L.; Hagiwara, Y.; Raghavendra, U.; Yuvaraj, R.; Arunkumar, N.; Murugappan, M.; Acharya, U.R. A deep learning approach for Parkinson’s disease diagnosis from EEG signals. Neural Comput. Appl. 2018, 5, 1–7. [Google Scholar] [CrossRef]
- Collis, W.B.; White, P.R.; Hammond, J.K. Higher-Order Spectra: The Bispectrum and Trispectrum. Mech. Syst. Signal Process. 1998, 12, 375–394. [Google Scholar] [CrossRef]
- Acharya, U.R.; Vidya, K.S.; Koh, J.E.W.; Martis, R.J.; Tan, J.H.; Oh, S.L.; Adam, M.; Hagiwara, Y.; Mookiah, M.R.K.; Chua, K.P.; et al. Application of higher-order spectra for the characterization of coronary artery disease using electrocardiogram signals. Biomed. Signal Process. Control 2017, 31, 31–43. [Google Scholar] [CrossRef]
- Tan, J.H.; Ng, E.Y.K.; Acharya, U.R.; Chee, C. Study of normal ocular thermogram using textural parameters. Infrared Phys. Technol. 2010, 53, 120–126. [Google Scholar]
- Savare, G.; Toscani, G. The concavity of rényi entropy power. IEEE Trans. Inf. Theory 2014, 60, 2687–2693. [Google Scholar] [CrossRef]
- Shannon, C.E. A mathematical theory of communication. Bell Syst. Tech. J. 1948, 27, 623–656. [Google Scholar] [CrossRef]
- Darbellay, G.A.; Vajda, I. Entropy expressions for multivariate continuous distributions. IEEE Trans. Inf. Theory 2000, 46, 709–712. [Google Scholar] [CrossRef]
- Hu, Q.; Yu, D. Entropies of fuzzy indiscernibility relation and its operations. Int. J. Uncertain. Fuzziness Knowlege-Based Syst. 2004, 12, 575–589. [Google Scholar] [CrossRef]
- Tang, X. Automated diagnosis of glaucoma using texture and higher order spectra features. IEEE Trans. Image Process. 1998, 7, 1602–1609. [Google Scholar]
- Galloway, M.M. Texture analysis using gray level run lengths. Comput. Graph. Image Process. 1975, 2, 172–179. [Google Scholar] [CrossRef]
- Xu, D.H.; Kurani, A.S.; Furst, J.D.; Raicu, D.S. Run-length encoding for volumetric texture. Proc. Fourth IASTED Int. Conf. Vis. Imaging Image Process. 2004, pp. 534–539. Available online: http://facweb.cs.depaul.edu/research/vc/Publications/final_submission_paper_452_131_last.pdf (accessed on 31 January 2020).
- Hornero, R.; Abasolo, D.; Jimeno, N.; Sanchez, C.I.; Poza, J.; Aboy, M. Variability, regularity, and complexity of time series generated by schizophrenic patients and control subjects. IEEE Trans. Biomed. Eng. 2006, 53, 210–218. [Google Scholar] [CrossRef] [PubMed]
- Acharya, U.R.; Sudarshan, V.K.; Adeli, H.; Santhosh, J.; Koh, J.E.W.; Adeli, A. Computer-aided diagnosis of depression using EEG signals. Eur. Neurol. 2015, 73, 329–336. [Google Scholar] [CrossRef] [PubMed]
- Haidekker, M. Advanced Biomedical Image Analysis, 1st ed.; John Wiley & Sons: Hoboken, NJ, USA, 2011. [Google Scholar]
- Cai, D.; He, X.; Zhou, K.; Han, J.; Bao, H. Locality Sensitive Discriminant Analysis. In Proceedings of the 20th International Joint Conference on Artificial Intelligence, Hyderabad, India, 6–12 January 2007; IJCAI: San Francisco, CA, USA, 2007; pp. 708–713. [Google Scholar]
- Kim, T.K. T test as a parametric statistic. Korean J. Anesthesiol. 2015, 68, 540–546. [Google Scholar] [CrossRef] [PubMed]
- Hasan, M.R.; Ibrahimy, M.I.; Motakabber, S.M.A.; Shahid, S. Classification of multichannel EEG signal by linear discriminant analysis. In Progress in Systems Engineering; Springer: Cham, Switzerland, 2015; Volume 366, pp. 279–282. [Google Scholar]
- Ghojogh, B.; Crowley, M. Introduction to machine learning: K-nearest neighbors. Ann. Transl. Med. 2019, 4, 1–16. [Google Scholar]
- Zhang, Z. Support vector machines. Ann. Transl. Med. 2016, 4, 1–7. [Google Scholar]
- Bakheet, S. An SVM framework for malignant melanoma detection based on optimized HOG features. Computation 2017, 5, 1–13. [Google Scholar] [CrossRef]
- Men, S.; Yan, L.; Liu, J.; Qian, H.; Luo, Q. A classification method for seed viability assessment with infrared thermography. Sensors 2017, 17, 845. [Google Scholar] [CrossRef]
- Apostolidis-Afentoulis, V.; Lioufi, K.I. SVM Classification with Linear and RBF Kernels. Available online: https://www.researchgate.net/publication/279913074_SVM_Classification_with_Linear_and_RBF_kernels (accessed on 31 January 2020).
- Karatzoglou, A.; Meyer, D.; Hornik, K. Support vector machines in R. J. Stat. Softw. 2006, 15, 1–28. [Google Scholar] [CrossRef]
- Duda, D.G.S.R.O.; Hart, P.E. Pattern Classification. John Wiley and Sons: New York, NY, USA, 2001. [Google Scholar]
- Jadhav, P.N.; Shanamugan, D.; Chourasia, A.; Ghole, A.R.; Acharyya, A.; Naik, G. Automated detection and correction of eye blink and muscular artefacts in EEG signal for analysis of Autism Spectrum Disorder. In Proceedings of the 36th Annual International Conference of the IEEE Engineering in Medicine and Biology Society, Chicago, IL, USA, 26–30 August 2014; IEEE: Chicago, IL, USA; pp. 1881–1884. [Google Scholar]
- Djemal, R.; Al Sharabi, K.; Ibrahim, S.; Alsuwailem, A. EEG-Based computer aided diagnosis of autism spectrum disorder using wavelet, entropy, and ANN. Biomed. Res. Int. 2017, 2017, 1–9. [Google Scholar] [CrossRef]
- Elhabashy, H.; Raafat, O.; Afifi, L.; Raafat, H.; Abdullah, K. Quantitative EEG in autistic children. Egypt. J. Neurol. Psychiatry Neurosurg. 2015, 52, 176–182. [Google Scholar]
- Bosl, W.J.; Tager-Flusberg, H.; Nelson, C.A. EEG analytics for early detection of autism spectrum disorder: A data-driven approach. Sci. Rep. 2018, 8, 1–20. [Google Scholar] [CrossRef] [PubMed]
- Bhat, S.; Acharya, U.R.; Adeli, H.; Bairy, G.M.; Adeli, A. Automated diagnosis of autism: In search of a mathematical marker. Rev. Neurosci. 2014, 25. [Google Scholar] [CrossRef] [PubMed]
- Lushchekina, E.A.; Podreznaya, E.D.; Lushchekin, V.S.; Novototskii-Vlasov, V.Y.; Strelets, V.B. Comparative studies of EEG theta and gamma rhythms in normal children and children with early childhood autism. Neurosci. Behav. Physiol. 2014, 44, 902–908. [Google Scholar] [CrossRef]
- Eldridge, J.; Lane, A.E.; Belkin, M.; Dennis, S. Robust features for the automatic identification of autism spectrum disorder in children. J. Neurodev. Disord. 2014, 6, 1–12. [Google Scholar] [CrossRef]
- Yousef, A.; Youssef, U.; El-Shabrawy, A.; Abdel Fattah, N.A.; Khedr, H.; Khedr, H. EEG abnormalities and severity of symptoms in non-epileptic autistic children. Egypt. J. Psychiatry 2017, 38, 59. [Google Scholar] [CrossRef]
- Heunis, T.; Aldrich, C.; Peters, J.M.; Jeste, S.S.; Sahin, M.; Scheffer, C.; Vries, P.J. Recurrence quantification analysis of resting state EEG signals in autism spectrum disorder—A systematic methodological exploration of technical and demographic confounders in the search for biomarkers. BMC Med. 2018, 16, 1–17. [Google Scholar] [CrossRef]
- Hadoush, H.; Alafeef, M.; Abdulhay, E. Brain complexity in children with mild and severe autism spectrum disorders: Analysis of multiscale entropy in EEG. Brain Topogr. 2019, 32, 914–921. [Google Scholar] [CrossRef]
- Hashemian, H.P.M. Decision-level fusion-based structure of autism diagnosis uisng interpretation of EEG signals related to facial expression modes. Neurophysiology 2017, 49, 59–71. [Google Scholar] [CrossRef]
- Grossi, E.; Olivieri, C.; Buscema, M. Diagnosis of autism through EEG processed by advanced computational algorithms: A pilot study. Comput. Methods Programs Biomed. 2017, 142, 73–79. [Google Scholar] [CrossRef]
- Thapaliya, S.; Jayarathna, S.; Jaime, M. Evaluating the EEG and eye movements for autism spectrum disorder. In Proceedings of the 2018 IEEE International Conference on Big Data, Seattle, WA, USA, 10–13 December 2018; IEEE: Chicago, IL, USA, 2018; pp. 2328–2336. [Google Scholar]
- Haputhanthri, D.; Brihadiswaran, G.; Gunathilaka, S.; Meedeniya, D.; Jayawardena, Y.; Jayarathna, S.; Jaime, M. An EEG based channel optimized classification approach for autism spectrum disorder. Proceedings of 2019 Moratuwa Engineering Research Conference (MERCon), Moratuwa, Sri Lanka, 3–5 July 2019; IEEE: Chicago, IL, USA, 2019; pp. 123–128. [Google Scholar]
- Harun, N.H.; Hamzah, N.; Zaini, N.; Sani, M.M.; Norhazman, H.; Yassin, I.M. EEG classification analysis for diagnosing autism spectrum disorder based on emotions. J. Telecommun. Electron. Comput. Eng. 2018, 10, 87–93. [Google Scholar]
- Haartsen, R.; Jones, E.J.H.; Orekhova, E.V.; Charman, T.; Johnson, M.H. Functional EEG connectivity in infants associates with later restricted and repetitive behaviours in autism: A replication study. Transl. Psychiatry 2019, 9. [Google Scholar] [CrossRef] [PubMed]
- Acharya, U.R.; Fujita, H.; Oh, S.L.; Raghavendra, U.; Tan, J.H.; Adam, M.; Gertych, A.; Hagiwara, Y. Automated identification of shockable and non-shockable life-threatening ventricular arrhythmias using convolutional neural network. Future Gener. Comp. Syst. 2018, 79, 952–959. [Google Scholar] [CrossRef]
- Oh, S.L.; Ng, E.Y.K.; San Tan, R.; Acharya, U.R. Automated diagnosis of arrhythmia using combination of CNN and LSTM techniques with variable length heart beats. Comput. Biol. Med. 2018, 102, 278–287. [Google Scholar] [CrossRef]
- Raghavendra, U.; Fujita, H.; Bhandary, S.V.; Gudigar, A.; Tan, J.H.; Acharya, U.R. Deep convolution neural network for accurate diagnosis of glaucoma using digital fundus images. Inf. Sci. 2018, 441, 41–49. [Google Scholar] [CrossRef]
- Tan, J.H.; Hagiwara, Y.; Pang, W.; Lim, I.; Oh, S.L.; Adam, M.; Tan, R.S.; Chen, M.; Acharya, U.R. Application of stacked convolutional and long short-term memory network for accurate identification of CAD ECG signals. Comput. Biol. Med. 2018, 94, 19–26. [Google Scholar] [CrossRef]

| Classifier | Number of Features | Accuracy (%) | Sensitivity (%) | Specificity (%) | Positive Predictive Value (%) |
|---|---|---|---|---|---|
| Linear discriminant analysis | 6 | 93.51 | 97.50 | 89.10 | 90.70 |
| Quadratic discriminant analysis | 5 | 85.71 | 87.50 | 83.78 | 85.37 |
| SVM polynomial 1 | 6 | 93.51 | 97.50 | 89.19 | 90.70 |
| SVM polynomial 2 | 5 | 97.40 | 97.50 | 97.30 | 97.50 |
| SVM polynomial 3 | 4 | 96.10 | 95.00 | 97.30 | 97.44 |
| KNN | 3 | 92.21 | 92.50 | 91.90 | 92.50 |
| SVMRBF | 2 | 97.40 | 100.00 | 94.60 | 95.24 |
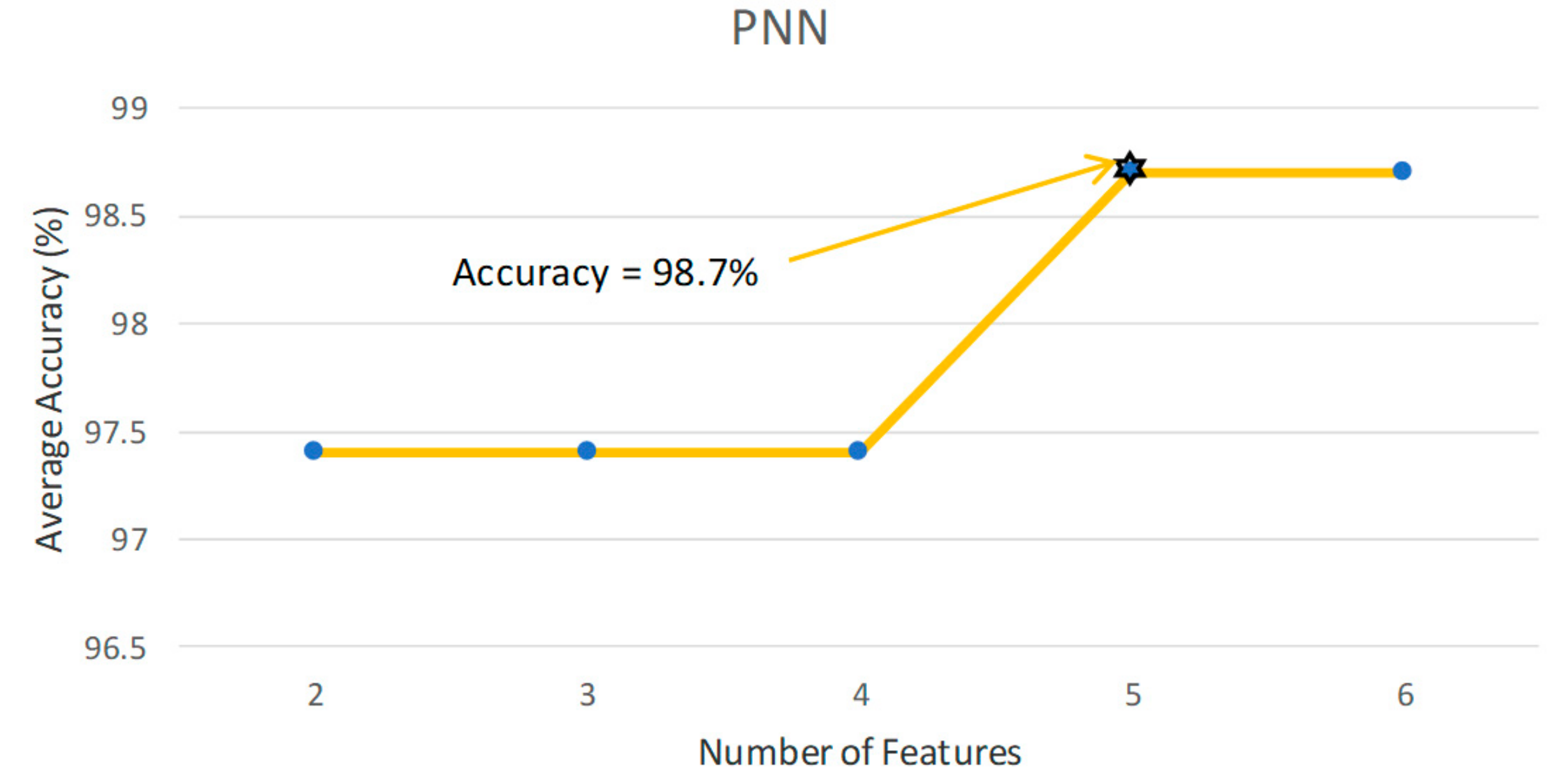
| PNN | 5 | 98.70 | 100.00 | 97.30 | 97.56 |
| Normal | ASD | |||||
|---|---|---|---|---|---|---|
| Features | Mean | SD | Mean | SD | p-Value | t-Value |
| LSDA13 | −1756.04 | 1126.778 | −801.964 | 1080.377 | 0.000309 | 3.786288 |
| LSDA8 | −1402.45 | 544.1245 | −2004.56 | 909.222 | 0.000711 | 3.55602 |
| LSDA9 | −886.62 | 264.4797 | −314.428 | 1157.47 | 0.003981 | 3.041854 |
| LSDA11 | 1918.153 | 1133.604 | 2545.265 | 1297.72 | 0.026577 | 2.262406 |
| LSDA7 | −583.943 | 600.9221 | −805.991 | 116.416 | 0.033149 | 2.209627 |
| LSDA2 | 133.0712 | 364.5094 | 291.3328 | 311.3471 | 0.044995 | 2.040697 |
| LSDA6 | −833.493 | 651.3617 | −998.316 | 145.0319 | 0.140299 | 1.505079 |
| LSDA1 | −385.252 | 98.16647 | −548.472 | 803.8656 | 0.209993 | 1.273933 |
| LSDA4 | −531.886 | 140.8786 | −567.485 | 125.164 | 0.246415 | 1.168582 |
| LSDA5 | −680.707 | 70.31738 | −691.059 | 23.01104 | 0.397739 | 0.854162 |
| LSDA14 | −657.845 | 501.4798 | −545.09 | 1308.884 | 0.614934 | 0.50615 |
| LSDA21 | −592.889 | 3.035538 | −590.386 | 44.26157 | 0.723211 | 0.356711 |
| LSDA10 | 796.1476 | 2058.705 | 922.609 | 867.6855 | 0.730657 | 0.346282 |
| LSDA12 | −5132.27 | 4467.789 | −4754.77 | 5277.353 | 0.735127 | 0.339583 |
| LSDA24 | −1464.89 | 71.78779 | −1461.35 | 7.848605 | 0.767 | 0.298501 |
| LSDA23 | −801.367 | 2047.65 | −706.917 | 504.8907 | 0.786254 | 0.273003 |
| LSDA28 | 1383.901 | 772.3631 | 1413.86 | 61.62772 | 0.815334 | 0.235248 |
| LSDA27 | 1029.853 | 696.2088 | 1005.313 | 73.09999 | 0.832258 | 0.213319 |
| LSDA29 | 585.8519 | 1.168018 | 585.4515 | 12.84125 | 0.845341 | 0.196346 |
| LSDA22 | −295.577 | 1400.244 | −339.624 | 97.81121 | 0.849659 | 0.19091 |
| LSDA17 | 445.7471 | 353.7109 | 485.0725 | 1609.695 | 0.880972 | 0.150629 |
| LSDA15 | 460.2031 | 37.12207 | 463.1301 | 119.2549 | 0.883222 | 0.147686 |
| LSDA19 | −592.541 | 1998.428 | −546.308 | 461.9218 | 0.891436 | 0.137369 |
| LSDA20 | −1035.72 | 1877.321 | −993.439 | 381.1509 | 0.893739 | 0.134455 |
| LSDA25 | −588.542 | 1679.106 | −621.33 | 116.2695 | 0.906315 | 0.118513 |
| LSDA16 | −1775.64 | 457.8857 | −1799.15 | 1321.607 | 0.91614 | 0.105843 |
| LSDA18 | −1565.5 | 2122.529 | −1523.15 | 1425.022 | 0.919109 | 0.101969 |
| LSDA26 | −663.813 | 14.99397 | −664.267 | 26.89385 | 0.926769 | 0.092285 |
| LSDA30 | −653.938 | 158.0741 | −653.208 | 37.69053 | 0.978273 | 0.027406 |
| LSDA3 | 346.7321 | 107.2744 | 338.2296 | 2320.293 | 0.981649 | 0.023149 |
| Year Published | Techniques | Number of Participants/Database/Demographics | Results |
|---|---|---|---|
| [46] 2014 |
| Caltech, PhysioNet, and Swartz Center for Computational Neuroscience: 20 subjects | Average correlation coefficient: 0.7574 Regression: 0.6992 |
| [50] 2014 |
| N: 1 subject A: 1 patient | Nonlinear features can be used as pointers to diagnose at early stages of ASD. |
| [51] 2014 |
| N: 24 subjects (boys; mean age of 6.05 ± 0.86 years) A: 27 patients (5.79 ± 1.42 years) | Spectral power of theta rhythm was lower in autistic children than in healthy children, whereas gamma power was larger. |
| [52] 2014 |
| N: 30 subjects A : 19 patients | Naïve Bayes: Ay: 79% |
| [48] 2015 |
| Child Psychiatry Outpatient Clinic: N: 21 subjects (aged between 4 and 12) A: 21 patients (aged between 4 and 12) | Statistically large differences in EEG power between the two groups; larger EEG power in delta and theta bands were found in the frontal and posterior regions. |
| [53] 2017 |
| Psychiatric Outpatients Clinics, Faculty of Medicine N: 40 subjects (aged between 4 and 12) A : 40 patients (aged between 2 to 12 years, 28 boys) | Abnormal EEG signals and brainwave regions were found to correlate with ASD severity. |
| [47] 2017 |
| King Abdulaziz University Brain Computer Interface Group: N: 10 subjects (males; aged 9 to 16) A: 9 patients (6 males, 3 females; aged 10–16) | Discrete wavelet transform (DWT)+ Shannon entropy: Ay: 99.71% |
| [56] 2017 |
| N: 6 boys (aged 7 to 9 years) A: 6 children (4 boys, 2 girls; aged 7 to 9 years) | The method proposed is able to differentiate normal and ASD classes. |
| [57] 2017 |
| Villa Santa Maria Institute N: 10 subjects (4 males, 6 females; aged 7 to 12 years) A: 15 patients (13 males, 2 females; aged 7 to 14 years) | Random forest classifier: Ay: 92.8% |
| [49] 2018 |
| Boston Children’s Hospital/Harvard Medical School N: 89 infants (with low risk of ASD) A: 99 infants (with older siblings having ASD diagnosis) | Sp, se: close to 100% Prediction scores correlated with actual scores. |
| [54] 2018 |
| N: 7 subjects (aged 2–6 years) A: 7 patients (aged 2-6 years) | SVM classifier: Ay: 92.9% Se: 100% Sp: 85.7% |
| [55] 2018 |
| Mild A: 18 patients Severe A: 18 patients | Mean multiscale entropy (MSE) values were found to be higher in children with mild A as compared to those with severe A. Increased sample entropy values in children with mild A. |
| [60] 2018 |
| - | Classification of ASD versus normal without emotions: Artificial neural network: Ay: 90.5% Classification of ASD versus normal with emotions: Artificial neural network: Ay: 92.5% Autistic children express a more complexed emotion than normal children. |
| [58] 2019 |
| 34 participants | Eye + EEG data: Naïve Bayes: Ay: 100% Logistic: Ay: 100% Only eye data: Logistic: Ay: 100% Deep neural network: Ay: 100% |
| [59] 2019 |
| N: 5 subjectsA: 10 patients (9 males, 6 females; between 5 and 17 years) | Random forest classifier: Ay: 93% |
| [61] 2019 |
| N (low risk infants): 20 subjects A (high-risk infants): 81 patients | Insignificant increase in global functional connectivity and networks in the alpha range between high-risk (HR) and low-risk (LR) groups and other groups being compared. |
| Present study |
| N: 37 healthy A: 40 patients | Probabilistic neural network classifier: Ay: 98.7% |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pham, T.-H.; Vicnesh, J.; Wei, J.K.E.; Oh, S.L.; Arunkumar, N.; Abdulhay, E.W.; Ciaccio, E.J.; Acharya, U.R. Autism Spectrum Disorder Diagnostic System Using HOS Bispectrum with EEG Signals. Int. J. Environ. Res. Public Health 2020, 17, 971. https://doi.org/10.3390/ijerph17030971
Pham T-H, Vicnesh J, Wei JKE, Oh SL, Arunkumar N, Abdulhay EW, Ciaccio EJ, Acharya UR. Autism Spectrum Disorder Diagnostic System Using HOS Bispectrum with EEG Signals. International Journal of Environmental Research and Public Health. 2020; 17(3):971. https://doi.org/10.3390/ijerph17030971
Chicago/Turabian StylePham, The-Hanh, Jahmunah Vicnesh, Joel Koh En Wei, Shu Lih Oh, N. Arunkumar, Enas. W. Abdulhay, Edward J. Ciaccio, and U. Rajendra Acharya. 2020. "Autism Spectrum Disorder Diagnostic System Using HOS Bispectrum with EEG Signals" International Journal of Environmental Research and Public Health 17, no. 3: 971. https://doi.org/10.3390/ijerph17030971
APA StylePham, T.-H., Vicnesh, J., Wei, J. K. E., Oh, S. L., Arunkumar, N., Abdulhay, E. W., Ciaccio, E. J., & Acharya, U. R. (2020). Autism Spectrum Disorder Diagnostic System Using HOS Bispectrum with EEG Signals. International Journal of Environmental Research and Public Health, 17(3), 971. https://doi.org/10.3390/ijerph17030971







