Fungal Contaminants in Drinking Water Regulation? A Tale of Ecology, Exposure, Purification and Clinical Relevance
Abstract
:1. Introduction
2. Fungi and Water—Background Information
2.1. Regulations
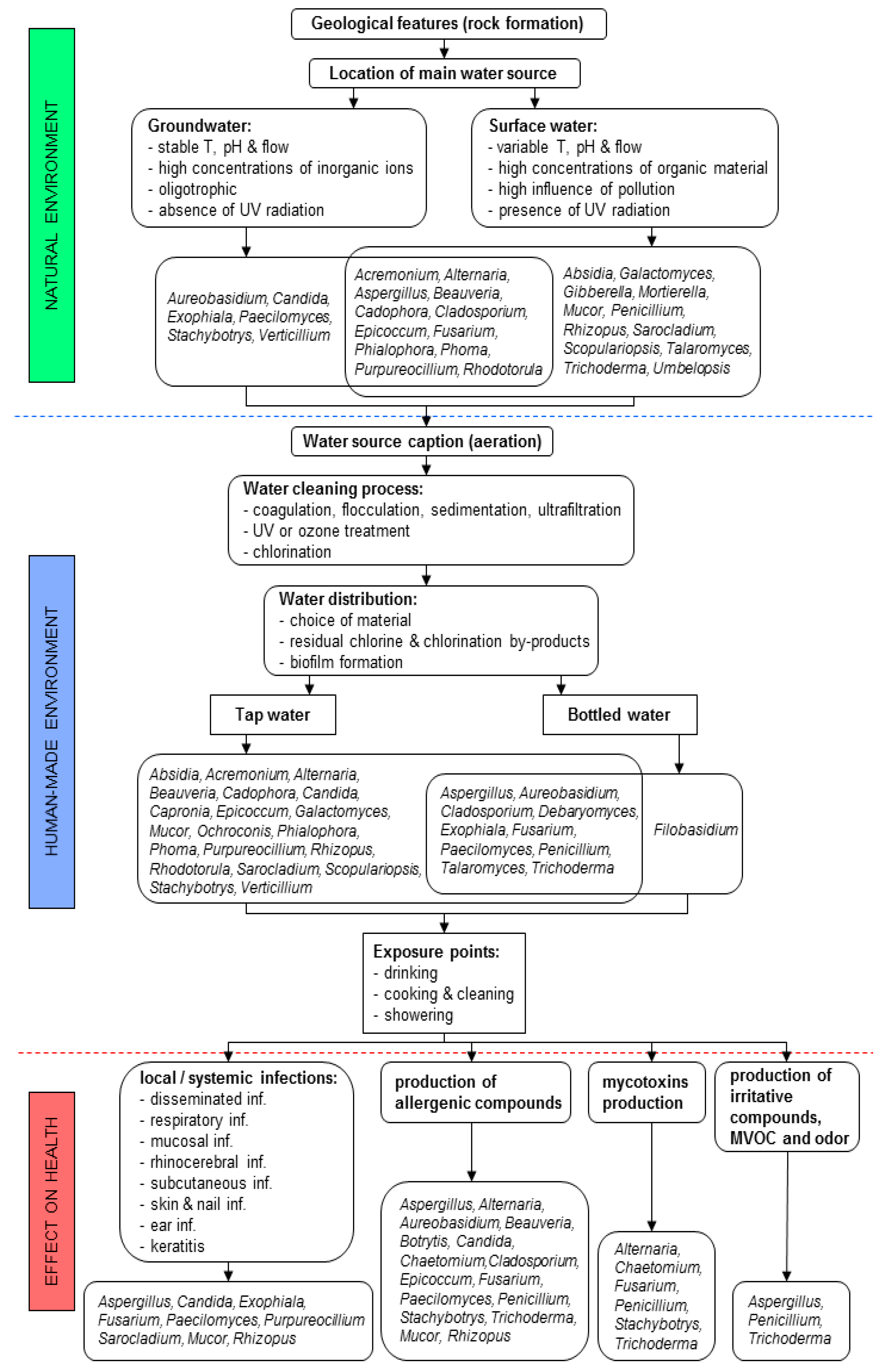
2.2. Ecology of Fungi in Water
2.3. Aqueous Geochemistry Processes Affect the Presence of Fungi in Water and Vice-Versa
2.4. Number and Diversity of Fungi Depends on Organic Matter Originating from Natural and Anthropogenic Sources
2.5. Effect of Sunlight and Water Temperature on Fungi in the Natural Environment
2.6. Effect of Drinking Water Treatment Processes on Fungal Contaminants
2.7. Materials Used for Building Water Supply Networks and Their Effect on Biofilm Formation
2.8. Commonly Used Methods for Isolation and Detection of Fungi in Water and Biofilms
3. Exposure to Fungi from Water in Indoor Environments and Their Medical Relevance
3.1. Direct Contact with Fungi
3.2. Indirect Contact with Fungi
3.3. Fungal Metabolites—Mycotoxins, Allergens, Microbial Volatile Organic Compounds (MVOCs)
4. Discussion
5. Conclusions
5.1. Future Scientific Research Needs
- Development of a consensus standard operating analytical procedure for the assessment of fungal contaminants in drinking water;
- Establishment of a geographically broad report on fungal contaminants in water (enumeration and variety) using a standardized analytical procedure.
- Development of sampling techniques necessary to detect sporadic particles released by biofilms.
- Large scale assessment of the presence and quantification of mycotoxins and MVOCs in drinking water.
- Generating agent specific epidemiological assessments of the health effects resulting from drinking-waterborne fungi.
5.2. Recommendations
- 1
- Surveillance of drinking water in relevant contexts.
- 2
- Adoption of the current Swedish legislation with an update of its fungal parameters to levels compatible with current knowledge.
- 3
- Special attention to be paid to hospitals and other open-to-public buildings, where immunocompromised people circulate or stay for a longer time and where molecular typing may be required in order to track sources or link infections together.
5.3. Afterword
- -
- Filtration: use of filters with a pore diameter of 0.45 μm and a filtration volume of 100 mL
- -
- Media: Rose Bengal Chloramphenicol and Chlortetracycline Agar (RBCC) for filamentous fungi and for yeasts
- -
- Incubation temperature: 25 °C.
- -
- Incubation time: 7 days
- -
- Results: maximum allowed number of moulds + yeasts = 100 CFU/100 mL [41]
- -
- Filtration: use of filters with a pore diameter of 0.45 μm and a filtration volume 100 mL
- -
- Media: Sabouraud agar for filamentous fungi and Dichloran Rose Bengal Chloramphenicol Agar (DRBC) for yeasts
- -
- Incubation temperature: 30 °C yields the highest diversity as reported by different authors
- -
- Incubation time: 7 days
- -
- Results: maximum allowed number (Unchanged due to the lack of epidemiological data that could support alterations) of moulds + yeasts = 100 CFU/100 mL
- -
- Detection and quantification of clinically relevant species/genera (culture-based + PCR-based in hospitals and other open-to-public buildings)
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Gostinčar, C.; Grube, M.; Gunde-Cimerman, N. Evolution of fungal pathogens in domestic environments? Fungal Biol. 2011, 115, 1008–1018. [Google Scholar] [CrossRef] [PubMed]
- DEFRA (Department for Environment, Food & Rural Affairs). A Review of Fungi in Drinking Water and the Implications for Human Health, 1st ed.; BIO Intelligence Service: Paris, France, 2011; p. 107.
- De Bruin, A.; Ibelings, B.; Kagami, M.; Mooij, W.M.; van Donk, E. Adaptation of the fungal parasite Zygorhizidium planktonicum during 200 generations of growth on homogeneous and heterogeneous populations of its host, the diatom Asterionella formosa. J. Eukaryot. Microbiol. 2008, 55, 69–74. [Google Scholar] [CrossRef] [PubMed]
- Wurzbacher, C.; Kerr, J.; Grossart, H.-P. Aquatic fungi. In The Dynamical Processes of Biodiversity: Case Studies of Evolution and Spatial Distribution, 1st ed.; Grillo, O., Venora, G., Eds.; InTech: Rijeka, Croatia, 2011; Volume 1, pp. 227–258. [Google Scholar]
- Hinzelin, F.; Block, J.C. Yeasts and filamentous fungi in drinking water. Environ. Technol. Lett. 1985, 6, 101–106. [Google Scholar] [CrossRef]
- Nyström, A.; Grimvall, A.; Krantz-Rülcker, C.; Sävenhed, R.; Åkerstrand, K. Drinking water off-flavour caused by 2,4,6-trichloroanisole. Water Sci. Technol. 1992, 25, 241–249. [Google Scholar]
- Frankova, E.; Horecka, M. Filamentous soil fungi and unidentified bacteria in drinking water from wells and water mains near Bratislava. Microbiol. Res. 1995, 150, 311–313. [Google Scholar] [CrossRef]
- Arvanitidou, M.; Kanellou, K.; Constantinides, T.C.; Katsouyannopoulos, V. The occurrence of fungi in hospital and community potable waters. Lett. Appl. Microbiol. 1999, 29, 81–84. [Google Scholar] [CrossRef] [PubMed]
- Kinsey, G.C.; Paterson, R.R.M.; Kelley, J. Methods for the determination of filamentous fungi in treated and untreated waters. J. Appl. Microbiol. 1999, 85, 214–224. [Google Scholar] [CrossRef] [PubMed]
- Warris, A.; Gaustad, P.; Meis, J.F.G.M.; Voss, A.; Verweij, P.E.; Abrahamsen, T.G. Recovery of filamentous fungi from water in a paediatric bone marrow transplantation. J. Hosp. Infect. 2001, 47, 143–148. [Google Scholar] [CrossRef] [PubMed]
- Göttlich, E.; van der Lubbe, W.; Lange, B.; Fiedler, S.; Melchert, I.; Reifenrath, M.; Flemming, H.-C.; de Hoog, S. Fungal flora in groundwater-derived public drinking water. Int. J. Hyg. Environ. Health 2002, 205, 269–279. [Google Scholar] [CrossRef] [PubMed]
- Rankovic, B. Five Serbian reservoirs contain different fungal propagules. Mycologia 2005, 97, 50–56. [Google Scholar] [CrossRef] [PubMed]
- Gonçalves, A.B.; Paterson, R.R.M.; Lima, N. Survey and significance of filamentous fungi from tap water. Int. J. Hyg. Environ. Health 2006, 209, 257–264. [Google Scholar] [CrossRef] [PubMed]
- Kanzler, D.; Buzina, W.; Paulitsch, A.; Haas, D.; Platzer, S.; Marth, E.; Mascher, F. Occurrence and hygienic relevance of fungi in drinking water. Mycoses 2008, 51, 165–169. [Google Scholar] [CrossRef] [PubMed]
- Pereira, V.J.; Basîlio, M.C.; Fernandes, D.; Domingues, M.; Paiva, J.M.; Benoliel, M.J.; Crespo, M.T.; San Romão, M.V. Occurrence of filamentous fungi and yeasts in three different drinking water sources. Water Res. 2009, 43, 3813–3819. [Google Scholar] [CrossRef] [PubMed]
- Hayette, M.-P.; Christiaens, G.; Mutsers, J.; Barbier, C.; Huynen, P.; Melin, P.; de Mol, P. Filamentous fungi recovered from the water distribution system of a Belgian university hospital. Med. Mycol. 2010, 48, 969–974. [Google Scholar] [CrossRef] [PubMed]
- Rudenko, A.V.; Savluk, O.S.; Saprykina, M.N.; Yastremskaya, A.V.; Goncharuk, V.V. Microscopic fungi in water of the Dnieper river. J. Water Chem. Technol. 2011, 33, 323–327. [Google Scholar] [CrossRef]
- Oliveira, B.R.; Crespo, M.T.; San Romão, M.V.; Benoliel, M.J.; Samson, R.A.; Pereira, V.J. New insights concerning the occurrence of fungi in water sources and their potential pathogenicity. Water Res. 2013, 47, 6338–6347. [Google Scholar] [CrossRef] [PubMed]
- Novak Babič, M.; Zalar, P.; Ženko, B.; Džeroski, S.; Gunde-Cimerman, N. Yeasts and yeast-like fungi in tap water and groundwater, and their transmission to household appliances. Fungal Ecol. 2016, 20, 30–39. [Google Scholar] [CrossRef]
- Hageskal, G.; Knutsen, A.K.; Gaustad, P.; de Hoog, G.S.; Skaar, I. Diversity and significance of mold species in Norwegian drinking water. Appl. Environ. Microbiol. 2006, 72, 7586–7593. [Google Scholar] [CrossRef] [PubMed]
- Kauffmann-Lacroix, C.; Costa, D.; Imbert, C. Fungi, water supply and biofilms. In Fungal Biofilms and Related Infections, 1st ed.; Imbert, C., Ed.; Springer International Publishing: Cham, Switzerland, 2016; Volume 3, pp. 49–61. [Google Scholar]
- World Health Organization (WHO). Guidelines for Drinking Water Quality, 3rd ed.; World Health Organization: Geneva, Switzerland, 2004; p. 515. [Google Scholar]
- Heinrichs, G.; Hübner, I.; Schmidt, K.C.; de Hoog, G.S.; Haase, G. Analysis of black fungal biofilms occurring at domestic water taps (I): Compositional analysis using Tag-encoded FLX amplicon pyrosequencing. Mycopathologia 2013, 175, 387–397. [Google Scholar] [CrossRef] [PubMed]
- U.S. EPA. Distribution Systems: A Best Practices Guide Introduction, 1st ed.; United States Environmental Protection Agency, Office of Water: Washington, DC, USA, 2006; p. 2.
- U.S. EPA. Best Management Practices for the Maintenance of Water Distribution Assets, 1st ed.; Water Research Foundation: Denver, CO, USA, 2015; p. 126.
- Heinrichs, G.; Hübner, I.; Schmidt, K.C.; de Hoog, G.S.; Haase, G. Analysis of black fungal biofilms occurring at domestic water taps (II): Potential routes of entry. Mycopathologia 2013, 175, 399–412. [Google Scholar] [CrossRef] [PubMed]
- Moat, J.; Rizoulis, A.; Fox, G.; Upton, M. Domestic shower hose biofilms contain fungal species capable of causing opportunistic infection. J. Water Health 2016, 14, 727–737. [Google Scholar] [CrossRef] [PubMed]
- Grabińska-Łoniewska, A.; Konillowicz-Kowalska, T.; Wardzynska, G.; Boryn, K. Occurrence of fungi in water distribution system. Pol. J. Environ. Stud. 2007, 16, 539–547. [Google Scholar]
- Paterson, R.R.M.; Lima, N. Fungal contamination of drinking water. In Water Encyclopedia, 1st ed.; Lehr, J., Keeley, J., Kingery, T.B., III, Eds.; John Wiley & Sons: New York, NY, USA, 2005; pp. 1–7. [Google Scholar]
- Schültze, N.; Lehmann, I.; Bönisch, U.; Simon, J.C.; Polte, T. Exposure to mycotoxins increases the allergic immune response in a murine asthma model. Am. J. Respir. Crit. Care Med. 2010, 181, 1188–1199. [Google Scholar] [CrossRef] [PubMed]
- Anaissie, E.J.; Stratton, S.L.; Dignani, M.C.; Summerbell, R.C.; Rex, J.H.; Monson, T.P.; Spencer, T.; Kasai, M.; Francesconi, A.; Walsh, J.T. Pathogenic Aspergillus species recovered from a hospital water system: A 3-year prospective study. Clin. Infect. Dis. 2002, 34, 780–789. [Google Scholar] [CrossRef] [PubMed]
- De Hoog, G.S.; Guarro, J.; Gené, J.; Figueras, M.J. Atlas of Clinical Fungi; Electronic Version 4.0; Centraalbureau voor Schimmelcultures: Utrecht, The Netherlands, 2014; Available online: http://www.clinicalfungi.org/ (accessed on 4 April 2017).
- National Health and Medical Research Council. National Water Quality Management Strategy, Australian Drinking Water Guidelines 6, 1st ed.; Commonwealth of Australia: Canberra, Australia, 2011; p. 1126. [Google Scholar]
- WHO. Guidelines for Drinking Water Quality, 4th ed.; World Health Organization: Geneva, Switzerland, 2011; p. 564. [Google Scholar]
- U.S. EPA. Regulatory Determinations Support. Document for Selected Contaminants from the Second Drinking Water Contaminant Candidate List (CCL 2), 1st ed.; United States Environmental Protection Agency, Office of Water: Washington, DC, USA, 2005; p. 497.
- US EPA. Drinking water contaminant candidate list CCL4. Fed. Regist. 2016, 81, 81099–81114. [Google Scholar]
- EEC. Council directive 98/83/EC on the quality of water intended for human consumption. Off. J. Eur. Commun. 1998, L330, 32–54. [Google Scholar]
- WHO. Heterotrophic Plate Counts and Drinking-Water Safety: The Significance of HPCs for Water Quality and Human Health, 1st ed.; IWA Publishing: London, UK, 2003; p. 271. [Google Scholar]
- Ministry of Health. Vyhláška Kterou se Stanoví Hygienické Požadavky na Pitnou a Teplou Vodu a Četnost a Rozsah Kontroly Pitné Vody, Decree 252/2004, 1st ed.; Ministry of Health: Prague, Czech Republic, 2004. [Google Scholar]
- Ministry of Health. Government Decree 201/2001 on the Quality and Monitoring Requirements of Drinking Water, 1st ed.; Ministry of Health: Budapest, Hungary, 2001.
- NFA. Livsmedelsverkets Föreskrifter om Dricksvatten, SLVFS 2001:30, 1st ed.; National Food Administration: Uppsala, Sweden, 2001; p. 33.
- Gray, F.N. Pathogen control in drinking water. In Microbiology of Waterborne Diseases, 2nd ed.; Percival, L.S., Yates, V.M., Eds.; Elsevier: Oxford, UK, 2014; Volume 1, pp. 537–570. [Google Scholar]
- Baldy, V.; Chauvet, E.; Charcosset, J.; Gessner, M.O. Microbial dynamics associated with leaves decomposing in the mainstem and floodplain pond of a large river. Aquat. Microb. Ecol. 2002, 28, 25–36. [Google Scholar] [CrossRef]
- Bärlocher, F.; Seena, S.; Wilson, K.P.; Williams, D.D. Raised water temperature lowers diversity of hyporheic aquatic hyphomycetes. Freshw. Biol. 2008, 53, 368–379. [Google Scholar] [CrossRef]
- Medeiros, A.O.; Pascoal, C.; Graça, M.A.S. Diversity and activity of aquatic fungi under low oxygen conditions. Freshw. Biol. 2009, 54, 142–149. [Google Scholar] [CrossRef]
- Sterflinger, K. Fungi: Their role in deterioration of cultural heritage. Fungal Biol. Rev. 2010, 24, 47–55. [Google Scholar] [CrossRef]
- Sohlberg, E.; Bomberg, M.; Miettinen, H.; Nyyssönen, M.; Salavirta, H.; Vikman, M.; Itävaara, M. Revealing the unexplored fungal communities in deep groundwater of crystalline bedrock fracture zones in Olkiluoto, Finland. Front. Microbiol. 2015, 6, 573. [Google Scholar] [CrossRef] [PubMed]
- Hou, W.; Dou, C.; Lian, B.; Dong, H. The interaction of fungus with calcite and the effect on aqueous geochemistry in karst systems. Carbonate Evaporite 2013, 28, 413–418. [Google Scholar] [CrossRef]
- Kumar, R.; Kumar, A.V. Biodeterioration of Stone in Tropical Environments: An Overview, 1st ed.; Getty Conservation Institute: Los Angeles, CA, USA, 1999; p. 96. [Google Scholar]
- Gadd, G.M. Fungi in Biogeochemical Cycles, 1st ed.; Cambridge University Press: New York, NY, USA, 2006; p. 469. [Google Scholar]
- Sterflinger, K. Fungi as geologic agents. Geomicrobiol. J. 2000, 17, 97–124. [Google Scholar] [CrossRef]
- Krauss, G.-J.; Solé, M.; Krauss, G.; Schlosser, D.; Wesenberg, D.; Bärlocher, F. Fungi in freshwaters: Ecology, physiology and biochemical potential. FEMS Microbiol. Rev. 2011, 35, 620–651. [Google Scholar] [CrossRef] [PubMed]
- Massaccesi, G.; Romero, M.C.; Cazau, M.C.; Bucsinszky, A.M. Cadmium removal capacities of filamentous soil fungi isolated from industrially polluted sediments, in La Plata (Argentina). World J. Microbiol. Biotechnol. 2002, 18, 817–820. [Google Scholar] [CrossRef]
- Steffen, K.T.; Hatakka, A.; Hofrichter, M. Removal and mineralization of polycyclic aromatic hydrocarbons by litter-decomposing basidiomycetous fungi. Appl. Microbiol. Biotechnol. 2002, 60, 212–217. [Google Scholar] [CrossRef] [PubMed]
- Ehrman, J.M.; Bärlocher, F.; Wennrich, R.; Krauss, G.-J.; Krauss, G. Fungi in a heavy metal precipitating stream in the Mansfeld mining district, Germany. Sci. Total Environ. 2008, 389, 486–496. [Google Scholar] [CrossRef] [PubMed]
- Karuppayil, S.M.; Szaniszlo, P.J. Importance of calcium to the regulation of polymorphism in Wangiella (Exophiala) dermatitidis. J. Med. Vet. Mycol. 1997, 35, 379–388. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Szaniszlo, J.P. Roles of the pH signalling transcription factor PacC in Wangiella (Exophiala) dermatitidis. Fungal Genet. Biol. 2009, 46, 657–666. [Google Scholar] [CrossRef] [PubMed]
- Zalar, P.; Novak, M.; de Hoog, G.S.; Gunde-Cimerman, N. Dishwashers-a man-made ecological niche accommodating human opportunistic fungal pathogens. Fungal Biol. 2011, 115, 997–1007. [Google Scholar] [CrossRef] [PubMed]
- Suberkropp, K. The influence of nutrients on fungal growth, productivity, and sporulation during leaf breakdown in streams. Can. J. Bot. 1995, 73, 1361–1369. [Google Scholar] [CrossRef]
- Sridhar, K.R.; Bärlocher, F. Water chemistry and sporulation by aquatic hyphomycetes. Mycol. Res. 1997, 101, 591–596. [Google Scholar] [CrossRef]
- Weber, S.D.; Hofmann, A.; Pilhofer, M.; Wanner, G.; Agerer, R.; Ludwig, W.; Schleifer, K.H.; Fried, J. The diversity of fungi in aerobic sewage granules assessed by 18S rRNA gene and ITS sequence analyses. FEMS Microbiol. Ecol. 2009, 68, 246–254. [Google Scholar] [CrossRef] [PubMed]
- Wurzbacher, C.M.; Bärlocher, F.; Grossart, H.-P. Fungi in lake ecosystems. Aquat. Microb. Ecol. 2010, 59, 125–149. [Google Scholar] [CrossRef]
- Tsui, C.K.M.; Baschien, C.; Goh, T.-K. Biology and ecology of freshwater fungi. In Biology of Microfungi, 1st ed.; Li, D.-W., Ed.; Springer International Publishing: Cham, Switzerland, 2016; pp. 285–313. [Google Scholar]
- Pereira, V.J.; Fernandes, D.; Carvalho, G.; Benoliel, M.J.; San Romão, M.V.; Barreto Crespo, M.T. Assessment of the presence and dynamics of fungi in drinking water sources using cultural and molecular methods. Water Res. 2010, 44, 4850–4859. [Google Scholar] [CrossRef] [PubMed]
- Madrid, H.; Hernández-Restrepo, M.; Gené, J.; Cano, J.; Guarro, J.; Silva, V. New and interesting chaetothyrialean fungi from Spain. Mycol. Progress 2016, 15, 1179–1201. [Google Scholar] [CrossRef]
- Biedunkiewicz, A.; Kowalska, K.; Schulz, Ł.; Stojek, K.; Dynowska, M.; Ejdys, E.; Sucharzewska, E.; Kubiak, D. Mycological monitoring of selected aquatic ecosystems in the context of epidemiological hazards. Drinking water. Ann. Parasitol. 2014, 60, 191–198. [Google Scholar] [PubMed]
- Zupančič, J.; Novak Babič, M.; Zalar, P.; Gunde-Cimerman, N. The black yeast Exophiala dermatitidis and other selected opportunistic human fungal pathogens spread from dishwashers to kitchens. PLoS ONE 2016, 11, e0148166. [Google Scholar] [CrossRef] [PubMed]
- Blasi, B.; Poyntner, C.; Rudavsky, T.; Prenafeta-Boldú, X.F.; de Hoog, S.; Tafer, H.; Sterflinger, K. Pathogenic yet environmentally friendly? Black fungal candidates for bioremediation of pollutants. Geomicrobiol. J. 2016, 33, 308–317. [Google Scholar] [CrossRef] [PubMed]
- Mehlman, M.A. Dangerous and cancer-causing properties of products and chemicals in the oil refining and petrochemical industry: VIII. Health effects of motor fuels: Carcinogenicity of gasoline: Scientific update. Environ. Res. 1992, 59, 238–249. [Google Scholar] [CrossRef]
- Taylor, R.T.; Hanna, M.L.; Shah, N.N.; Shonnard, D.R.; Duba, A.G.; Durham, W.B.; Jackson, K.J.; Knapp, R.B.; Wijesinghe, A.M.; Knezovich, J.P.; et al. In situ bioremediation of trichloroethylene-contaminated water by a resting-cell methanotrophic microbial filter. Hydrol. Sci. J. 1993, 38, 323–342. [Google Scholar] [CrossRef]
- Prenafeta-Boldú, X.F.; Summerbell, R.; de Hoog, G.S. Fungi growing on aromatic hydrocarbons: Biotechnology’s unexpected encounter with biohazard? FEMS Microbiol. Rev. 2006, 30, 109–130. [Google Scholar] [CrossRef] [PubMed]
- Gesell, M.; Hammer, E.; Specht, M.; Francke, W.; Schauer, F. Biotransformation of biphenyl by Paecilomyces lilacinus and characterization of ring cleavage products. Appl. Environ. Microb. 2001, 67, 1551–1557. [Google Scholar] [CrossRef] [PubMed]
- Verdin, A.; Sahraoui, A.L.; Durand, R. Degradation of benzo[a]pyrene by mitosporic fungi and extracellular oxidative enzymes. Int. Biodeterior. Biodegrad. 2004, 53, 65–70. [Google Scholar] [CrossRef]
- Mohamed, D.J.; Martiny, J.B.H. Patterns of fungal diversity and composition along a salinity gradient. ISME J. 2011, 5, 379–388. [Google Scholar] [CrossRef] [PubMed]
- Raghukumar, C. Biology of Marine Fungi, 1st ed.; Springer: Berlin, Germany, 2012; Volume 53, p. 328. [Google Scholar]
- Jebaraj, C.S.; Raghukumar, C.; Behnke, A.; Stoeck, T. Fungal diversity in oxygen-depleted regions of the Arabian Sea revealed by targeted environmental sequencing combined with cultivation. FEMS Microbiol. Ecol. 2010, 71, 399–412. [Google Scholar] [CrossRef] [PubMed]
- Singh, P.; Raghukumar, C.; Verma, P.; Shouche, Y. Assessment of fungal diversity in deep sea sediments by multiple primer approach. World J. Microbiol. Biotechnol. 2012, 28, 659–667. [Google Scholar] [CrossRef] [PubMed]
- Orlowski, M. Mucor dimorphism. Microbiol. Rev. 1991, 55, 234–258. [Google Scholar] [PubMed]
- Kevei, J. New aspects in monitoring of drinking water: Mycolocicalé studies (in Hungarian). Budapesti Közegészségügy 1992, 2, 53–56. [Google Scholar]
- Novak Babič, M.; Zalar, P.; Ženko, B.; Schroers, H.-J.; Džeroski, S.; Gunde-Cimerman, N. Candida and Fusarium species known as opportunistic human pathogens from customer-accessible parts of residential washing machines. Fungal Biol. 2015, 119, 95–113. [Google Scholar] [CrossRef] [PubMed]
- Pedro-Botet, M.L.; Sanchez, I.; Sabria, M.; Sopena, N.; Mateu, L.; Garcia-Nunez, M.; Rey-Joly, C. Impact of copper and silver ionization on fungal colonization of the water supply in health care centers: Implications for immunocompromised patients. Clin. Infect. Dis. 2007, 45, 84–86. [Google Scholar] [CrossRef] [PubMed]
- Plutzer, J.; Törökné, A. Free-living microscopic organisms as indicators of changes in drinking-water quality. Water Pract. Technol. 2012, 7, 1–14. [Google Scholar] [CrossRef]
- Niemi, R.M.; Knuth, S.; Lundström, K. Actinomycetes and fungi in surface waters and in potable water. Appl. Environ. Microbiol. 1982, 43, 378–388. [Google Scholar] [PubMed]
- Pap, K.; Tornai-Lehoczki, J.; Syposs, Z. Mold challenge study in bottled natural mineral waters and spring waters. Acta Microbiol. Immunol. Hung. 2008, 55, 145–155. [Google Scholar] [CrossRef] [PubMed]
- Van der Wielen, P.W.; van der Kooij, D. Nontuberculous Mycobacteria, fungi, and opportunistic pathogens in unchlorinated drinking water in The Netherlands. Appl. Environ. Microbiol. 2013, 79, 825–834. [Google Scholar] [CrossRef] [PubMed]
- Otterholt, E.; Charnock, C. Microbial quality and nutritional aspects of Norwegian brand waters. Int. J. Food Microbiol. 2010, 144, 455–463. [Google Scholar] [CrossRef] [PubMed]
- Mata, A.T.; Ferreira, J.P.; Oliveira, B.R.; Batoréu, M.C.; Barreto Crespo, M.T.; Pereira, V.J.; Bronze, M.R. Bottled water: Analysis of mycotoxins by LC-MS/MS. Food Chem. 2015, 176, 455–464. [Google Scholar] [CrossRef] [PubMed]
- Nevarez, L.; Vasseur, V.; Le Madec, A.; Le Bras, M.A.; Coroller, L.; Leguérinel, I.; Barbier, G. Physiological traits of Penicillium glabrum strain LCP 08.5568, a filamentous fungus isolated from bottled aromatised mineral water. Int. J. Food Microbiol. 2009, 130, 166–171. [Google Scholar] [CrossRef]
- Onofri, S.; Anastasi, A.; Del Frate, G.; Di Piazza, S.; Garnero, N.; Guglielminetti, M.; Isola, D.; Panno, L.; Ripa, C.; Selbmann, L.; Varese, G.C.; Voyron, S.; Zotti, M.; Zucconi, L. Biodiversity of rock, beach and water fungi in Italy. Plant Biosyst. 2011, 145, 978–987. [Google Scholar] [CrossRef]
- Yin, R.; Dai, T.; Avci, P.; Serafim Jorge, A.E.; de Melo, C.M.A.W.; Vecchio, D.; Huang, Y.-Y.; Gupta, A.; Hamblin, R.M. Light based anti-infectives: Ultraviolet C irradiation, photodynamic therapy, blue light, and beyond. Curr. Opin. Pharmacol. 2013, 13, 731–762. [Google Scholar] [CrossRef] [PubMed]
- Joyce, T.M.; McGuigan, K.G.; Elmore-Meegan, M.; Conroy, R.M. Inactivation of fecal bacteria in drinking water by solar heating. Appl. Environ. Microbiol. 1996, 62, 399–402. [Google Scholar] [PubMed]
- Heaselgrave, W.; Kilvington, S. Antimicrobial activity of simulated solar disinfection against bacterial, fungal, and protozoan pathogens and its enhancement by riboflavin. Appl. Environ. Microbiol. 2010, 76, 6010–6012. [Google Scholar] [CrossRef] [PubMed]
- Mitakakis, Z.T.; O’Meara, J.T.; Tovey, R.E. The effect of sunlight on allergen release from spores of the fungus Alternaria. Grana 2003, 42, 43–46. [Google Scholar] [CrossRef]
- Lonnen, J.; Kilvington, S.; Kehoe, S.C.; Al-Touati, F.; McGuigan, K.G. Solar photocatalytic disinfection of protozoan, fungal and bacterial microbes in drinking water. Water Res. 2005, 39, 877–883. [Google Scholar] [CrossRef] [PubMed]
- Sichel, C.; de Cara, M.; Tello, J.; Blanco, J.; Fernández-Ibánez, P. Solar photocatalytic disinfection of agricultural pathogenic fungi: Fusarium species. Appl. Catal. B Environ. 2007, 74, 152–160. [Google Scholar] [CrossRef]
- Rainey, R.C.; Harding, A.K. Drinking water quality and solar disinfection: Effectiveness in peri-urban households in Nepal. J. Water Health 2005, 3, 239–248. [Google Scholar] [CrossRef] [PubMed]
- Dang, C.K.; Schindler, M.; Chauvet, E.; Gessner, M.O. Temperature oscillations coupled with fungal communities can modulate warming effects on litter decomposition. Ecology 2009, 90, 122–131. [Google Scholar] [CrossRef] [PubMed]
- Feller, G.; Gerday, C. Pyschrophilic enzymes: Hot topics in cold adaptation. Nat. Rev. Microbiol. 2003, 1, 200–208. [Google Scholar] [CrossRef] [PubMed]
- Margesin, R.; Gander, S.; Zacke, G.; Gounot, A.M.; Schinner, F. Hydrocarbon degradation and enzyme activities of cold-adapted bacteria and yeasts. Extremophiles 2008, 7, 451–458. [Google Scholar] [CrossRef] [PubMed]
- Percival, L.S.; Yates, V.M.; Williams, W.D.; Chalmers, R.M.; Gray, F.N. Microbiology of Waterborne Diseases, 2nd ed.; Elsevier: Oxford, UK, 2014; p. 590. [Google Scholar]
- Selecky, M.; White, B.; Grunenfelder, G. Guidance Document: Slow Sand Filtration and Diatomaceous Earth Filtration for Small Water Systems, 1st ed.; Washington State Department of Health: Washington, DC, USA, 2003; p. 118.
- MRWA. Coagulation and Flocculation Process. Fundamentals, 1st ed.; Minnesota Rural Water Association: Elbow Lake, MN, USA, 2003; p. 8. [Google Scholar]
- Alpatova, A.; Verbych, S.; Bryk, M.; Nigmatullin, R.; Hilal, N. Ultrafiltration of water containing natural organic matter: Heavy metal removing in the hybrid complexation-ultrafiltration process. Sep. Purif. Technol. 2004, 40, 155–162. [Google Scholar] [CrossRef]
- Kelley, J.; Kinsey, G.C.; Paterson, R.R.M.; Pitchers, R. Identification and Control of Fungi in Distribution Systems, 1st ed.; AWWA Research Foundation and American Water Works Association: Denver, CO, USA, 2001; p. 150. [Google Scholar]
- Stopar, P. Water Treatment of Spring Jama. Graduation Thesis, University of Ljubljana, Ljubljana, Slovenia, 9 October 2007. [Google Scholar]
- Ozcelik, B. Fungi/bactericidal and static effects of ultraviolet light in 254 and 354 nm wavelengths. Res. J. Microbiol. 2007, 2, 42–49. [Google Scholar] [CrossRef]
- Hageskal, G.; Lima, N.; Skaar, I. The study of fungi in drinking water. Mycol. Res. 2009, 113, 165–172. [Google Scholar] [CrossRef] [PubMed]
- Glaze, W.H. Drinking water treatment with ozone. Environ. Sci. Technol. 1987, 21, 224–230. [Google Scholar] [CrossRef] [PubMed]
- Coronel, B.; Duroselley, P.; Behry, H.; Moskovtchenko, J.F.; Frenyy, J. In situ decontamination of medical wastes using oxidative agents. J. Hosp. Infect. 2002, 50, 207–212. [Google Scholar] [CrossRef] [PubMed]
- Kottapalli, B.; Wolf-Hall, C.E.; Schwarz, P. Evaluation of gaseous ozone and hydrogen peroxide treatments for reducing Fusarium survival in malting barley. J. Food Prot. 2005, 68, 1236–1240. [Google Scholar] [CrossRef] [PubMed]
- Fujiwara, K.; Kadoya, M.; Hayashi, Y.; Kurata, K. Effects of ozonated water application on the population density of Fusarium oxysporum f.sp. lycopersici in soil columns. Ozone Sci. Eng. 2006, 28, 125–127. [Google Scholar] [CrossRef]
- Geweely, S.I.N. Antifungal activity of ozonized olive oil (oleozone). Int. J. Agric. Biol. 2006, 5, 670–675. [Google Scholar]
- Rojas-Valencia, M.N. Research on ozone application as disinfectant and action mechanisms on wastewater microorganisms. In Science Against Microbial Pathogens: Communicating Current Research and Technological Advances, 1st ed.; Mendez-Vilas, A., Ed.; Formatex Research Centre: Badajoz, Spain, 2011; Volume 1, pp. 263–271. [Google Scholar]
- Roushdy, M.M.; Abdel-Shakour, E.H.; Abdel-Ghany, T.M. Sporicidal effect of ozone on fungal and bacterial spores in water disinfection. J. Am. Sci. 2011, 7, 942–948. [Google Scholar]
- Kang, H.M.; Pengkit, A.; Choi, K.; Jeon, S.S.; Choi, W.H.; Shin, B.D.; Choi, H.E.; Uhm, S.H.; Park, G. Differential inactivation of fungal spores in water and on seeds by ozone and arc discharge plasma. PLoS ONE 2015, 10, e0139263. [Google Scholar] [CrossRef] [PubMed]
- Sharbaugh, R.J. Decontamination: Principles of disinfection. In Sterilization Technology for the Health Care Facility, 2nd ed.; Reichert, M., Young, J.H., Eds.; Aspen Publishers Inc.: Gaithersburg, MD, USA, 1997; pp. 21–28. [Google Scholar]
- Kelley, J.; Paterson, R.; Kinsey, G.; Pitchers, R.; Rossmoore, H. Identification, significance and control of fungi in water distribution systems. In Proceedings of the Water Technology Conference, Denver, CO, USA, 9–12 November 1997. [Google Scholar]
- Pereira, V.J.; Marques, R.; Marques, M.; Benoliel, M.J.; Barreto Crespo, M.T. Free chlorine inactivation of fungi in drinking water sources. Water Res. 2013, 47, 517–523. [Google Scholar] [CrossRef] [PubMed]
- 4MS. ACCEPTANCE of Metallic Materials Used for Products in Contact with Drinking Water, 1st ed.; 4MS Joint Management Comitee, Germany, France, The Netherlands and United Kingdom: Berlin, Germany, 2011; p. 19. [Google Scholar]
- NLZOH; ZAG; NIJZ. Priporočila za Ocenjevanje Primernosti Materialov in Proizvodov, ki Prihajajo v Stik s Pitno Vodo in so del Vodovodnega Omrežja in Interne Vodovodne Napeljave (P-MPPV), 1st ed.; Nacionalni Laboratorij za Zdravje, Okolje in Hrano, Zavod za Gradbeništvo Slovenije, Nacionalni Inštitut za Javno Zdravje: Ljubljana, Slovenija, 2016; p. 48. [Google Scholar]
- BELGAQUA. General Terms and Conditions for the Acceptance of Materials in Contact with Drinking Water and Water Intended for the Production of Drinking Water, 1st ed.; Belgian Federation for the Water Sector: Brussels, Belgium, 2012; p. 24. [Google Scholar]
- Donlan, R.M. Biofilms: Microbial life on surfaces. Emerg. Infect. Dis. 2002, 8, 881–890. [Google Scholar] [CrossRef] [PubMed]
- Le Chevallier, M.W. Biofilms in drinking water distribution systems: Significance and control. In Identifying Future Drinking Water Contaminants, 1st ed.; National Academy Press: Washington, DC, USA, 1999; Volume 1, pp. 206–219. [Google Scholar]
- Doggett, M.S. Characterisation of fungal biofilms within a municipal water distribution system. Appl. Environ. Microb. 2000, 66, 1249–1251. [Google Scholar] [CrossRef]
- Lehtola, M.; Miettinen, I.T.; Lampola, T.; Hirvonen, A.; Vartiainen, T.; Martikainen, P.J. Pipeline materials modify the effectiveness of disinfectants in drinking water distribution systems. Water Res. 2004, 39, 1962–1971. [Google Scholar] [CrossRef] [PubMed]
- Percival, S.; Knapp, J.S.; Wales, D.S.; Edyvean, R.G.J. The effect of turbulent flow and surface roughness on biofilm formation in drinking water. J. Ind. Microbiol. Biotechnol. 1999, 22, 152–159. [Google Scholar] [CrossRef]
- Cooper, R.I. Microbial biofilms: Case reviews of bacterial and fungal pathogens persisting on biomaterials and environmental substrata. In Current Research, Technology and Education Topics in Applied Microbiology and Microbial Biotechnology, 1st ed.; Mendez-Vilas, A., Ed.; Formatex Research Centre: Badajoz, Spain, 2010; Volume 2, pp. 807–817. [Google Scholar]
- Wimpenny, J. An overview of biofilms as functional communities. In Community Structure and Co-Operation in Biofilms, 1st ed.; Allison, G.D., Ed.; Cambridge University Press: Cambridge, UK, 2000; Volume 59, pp. 1–24. [Google Scholar]
- Romani, A.M.; Fischer, H.; Mille-Lindblom, C.; Tranvik, L.J. Interactions of bacteria and fungi on decomposing litter: Differential extracellular enzyme activities. Ecology 2006, 87, 2559–2569. [Google Scholar] [CrossRef]
- Claus, H.; Gleixner, G.; Filip, Z. Formation of humic-like substances in mixed and pure cultures of aquatic microorganisms. Acta Hydrochim. Hydrobiol. 1999, 27, 200–207. [Google Scholar] [CrossRef]
- Harding, W.M.; Marques, L.L.R.; Howard, R.J.; Olson, M.E. Can filamentous fungi form biofilms? Trends Microbiol. 2009, 17, 475–480. [Google Scholar] [CrossRef] [PubMed]
- Simões, L.C.; Simões, M.; Lima, N. Kinetics of biofilm formation by drinking water isolated Penicillium expansum. Biofouling 2015, 31, 349–362. [Google Scholar] [CrossRef] [PubMed]
- Siqueira, V.M.; Oliveira, H.M.; Santos, C.; Paterson, R.R.M.; Gusmão, B.N.; Lima, N. Filamentous fungi in drinking water, particularly in relation to biofilm formation. Int. J. Environ. Res. Public Health 2011, 8, 456–469. [Google Scholar] [CrossRef] [PubMed]
- Biedunkiewicz, A.; Schulz, Ł. Fungi of the genus Exophiala in tap water—Potential etiological factors of phaeohyphomycoses. Mikol. Lek. 2012, 19, 23–26. [Google Scholar]
- Sammon, B.N.; Harrower, M.K.; Fabbro, D.L.; Reed, H.R. Incidence and distribution of microfungi in a treated municipal water supply system in sub-tropical Australia. Int. J. Environ. Res. Publ. Health 2010, 7, 1597–1611. [Google Scholar] [CrossRef] [PubMed]
- Kadaifciler, D.G.; Ökten, S.; Sen, B. Mycological contamination in dental unit waterlines in Istanbul, Turkey. Braz. J. Microbiol. 2013, 44, 977–981. [Google Scholar] [CrossRef] [PubMed]
- Schoch, C.L.; Seifert, K.A.; Huhndorf, S.; Robert, V.; Spouge, J.L.; Levesque, C.A.; Chen, W. Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for Fungi. Proc. Natl. Acad. Sci. USA 2012, 109, 6241–6246. [Google Scholar] [CrossRef] [PubMed]
- Irinyi, L.; Serena, C.; Garcia-Hermoso, D.; Arabatzis, M.; Desnos-Olivier, M.; Vu, D.; Cardinali, G.; Arthur, I.; Normand, A.C.; Giraldo, A.; et al. International Society of Human and Animal Mycology (ISHAM)-ITS reference DNA barcoding database—The quality controlled standard tool for routine identification of human and animal pathogenic fungi. Med. Mycol. 2015, 53, 313–337. [Google Scholar] [CrossRef] [PubMed]
- Stielow, J.B.; Lévesque, C.A.; Seifert, K.A.; Meyer, W.; Iriny, L.; Smits, D.; Renfurm, R.; Verkley, G.J.; Groenewald, M.; Chaduli, D.; et al. One fungus, which genes? Development and assessment of universal primers for potential secondary fungal DNA barcodes. Persoonia 2015, 35, 242–263. [Google Scholar] [CrossRef] [PubMed]
- Brinkman, E.N.; Haugland, A.R.; Wymer, J.L.; Byappanahalli, M.; Whitman, L.R.; Vesper, J.S. Evaluation of a rapid, Quantitative Real-Time PCR method for enumeration of pathogenic Candida cells in water. Appl. Environ. Microbiol. 2003, 69, 1775–1782. [Google Scholar] [CrossRef] [PubMed]
- Al-Gabr, H.M.; Zheng, T.; Yu, X. Occurrence and quantification of fungi and detection of mycotoxigenic fungi in drinking water in Xiamen City, China. Sci. Total Environ. 2013, 466–467, 1103–1111. [Google Scholar] [CrossRef] [PubMed]
- Mesquita-Rocha, S.; Godoy-Martinez, P.C.; Gonçalves, S.S.; Urrutia, M.D.; Carlesse, F.; Seber, A.; Silva, M.A.; Petrilli, A.S.; Colombo, A.L. The water supply system as a potential source of fungal infection in paediatric haematopoietic stem cell units. BMC Infect. Dis. 2013, 13, 289. [Google Scholar] [CrossRef] [PubMed]
- Lisboa, G.M.; Lisboa, Y.R.; Pinheiro, T.M.; Stegun, R.C.; da Silva-Filho, E.A. Microbial diversity in dental unit waterlines. Acta Odontol. Latinoam. 2014, 27, 110–114. [Google Scholar] [CrossRef] [PubMed]
- Brown, G.D.; Denning, D.W.; Gow, N.A.; Levitz, S.M.; Netea, M.G.; White, T.C. Hidden Killers: Human fungal infections. Sci. Transl. Med. 2012, 4, 165rv13. [Google Scholar] [CrossRef] [PubMed]
- Twaroch, T.E.; Focke, M.; Fleischmann, K.; Balic, N.; Lupinek, C.; Blatt, K.; Ferrara, R.; Mari, A.; Ebner, C.; Valent, P.; Spitzauer, S.; Swoboda, I.; Valenta, R. Carrier-bound Alt a 1 peptides without allergenic activity for vaccination against Alternaria alternata allergy. Clin. Exp. Allergy 2012, 42, 966–975. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.; Prakash, D.; Singh, A.B. Sensitization to different species of Aspergillus in bakery workers and general atopic population. Asian Pac. J. Allergy Immunol. 1998, 16, 5–15. [Google Scholar] [PubMed]
- Yu, C.J.; Chiou, S.H.; Lai, W.Y.; Chiang, B.L.; Chow, L.P. Characterization of a novel allergen, a major IgE-binding protein from Aspergillus flavus, as an alkaline serine protease. Biochem. Biophys. Res. Commun. 1999, 261, 669–675. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, K.F. Mycotoxin production by indoor molds. Fungal Genet. Biol. 2003, 39, 103–117. [Google Scholar] [CrossRef]
- Hedayati, M.T.; Pasqualotto, A.C.; Warn, P.A.; Bowyer, P.; Denning, D.W. Aspergillus flavus: Human pathogen, allergen and mycotoxin producer. Microbiology 2007, 153, 1677–1692. [Google Scholar] [CrossRef] [PubMed]
- Itabashi, T.; Hosoe, T.; Toyasaki, N.; Imai, T.; Adachi, M.; Kawai, K. Allergen activity of xerophilic fungus, Aspergillus restrictus. Arerugi 2007, 56, 101–108. [Google Scholar] [PubMed]
- Fiedler, K.; Schütz, E.; Geh, S. Detection of microbial volatile organic compounds (MVOCs) produced by moulds on various materials. Int. J. Hyg. Environ. Health 2001, 204, 111–121. [Google Scholar] [CrossRef] [PubMed]
- Stevens, D.A.; Moss, R.B.; Kurup, V.P.; Knutsen, A.P.; Greenberger, P.; Judson, M.A.; Denning, D.W.; Crameri, R.; Brody, A.S.; Light, M.; Skov, M.; Maish, W.; Mastella, G. Allergic bronchopulmonary aspergillosis in cystic fibrosis—State of the art: Cystic Fibrosis Foundation Consensus Conference. Clin. Infect. Dis. 2003, 1, S225–S264. [Google Scholar] [CrossRef] [PubMed]
- Gernez, Y.; Dunn, C.E.; Everson, C.; Mitsunaga, E.; Gudiputi, L.; Krasinska, K.; Davies, Z.A.; Herzenberg, L.A.; Tirouvanziam, R.; Moss, R.B. Blood basophils from cystic fibrosis patients with allergic bronchopulmonary aspergillosis are primed and hyper-responsive to stimulation by Aspergillus allergens. J. Cyst Fibros. 2012, 11, 502–510. [Google Scholar] [CrossRef] [PubMed]
- Zanjani, L.S.; Bakhtiari, A.; Sabokbar, A.; Khosravi, A.R.; Bahonar, A.; Memarnejadian, A. Sensibilisation of asthmatic patients to extracted antigens from strains of Aspergillus fumigatus, Aspergillus flavus and Aspergillus niger. J. Mycol. Med. 2012, 22, 58–63. [Google Scholar] [CrossRef] [PubMed]
- Taylor, P.E.; Esch, R.; Flagan, R.C.; House, J.; Tran, L.; Glovsky, M.M. Identification and possible disease mechanisms of an under-recognized fungus, Aureobasidium pullulans. Int. Arch. Allergy Immunol. 2006, 139, 45–52. [Google Scholar] [CrossRef] [PubMed]
- Westwood, G.S.; Huang, S.W.; Keyhani, N.O. Allergens of the entomopathogenic fungus Beauveria bassiana. Clin. Mol. Allergy 2005, 3, 1. [Google Scholar] [CrossRef] [PubMed]
- Jurgensen, C.W.; Madsen, A.M. Exposure to the airborne mould Botrytis and its health effects. Ann. Agric. Environ. Med. 2009, 16, 183–196. [Google Scholar] [PubMed]
- Koivikko, A.; Kalimo, K.; Nieminen, E.; Savolainen, J.; Viljanen, M.; Viander, M. Allergenic cross-reactivity of yeasts. Allergy 1988, 43, 192–200. [Google Scholar] [CrossRef] [PubMed]
- Khosravi, A.R.; Bandghorai, A.N.; Moazzeni, M.; Shokri, H.; Mansouri, P.; Mahmoudi, M. Evaluation of Candida albicans allergens reactive with specific IgE in asthma and atopic eczema patients. Mycoses 2009, 52, 326–333. [Google Scholar] [CrossRef] [PubMed]
- Beezhold, D.H.; Green, B.J.; Blachere, F.M.; Schmechel, D.; Weissman, D.N.; Velickoff, D.; Hogan, M.B.; Wilson, N.W. Prevalence of allergic sensitization to indoor fungi in West Virginia. Allergy Asthma Proc. 2008, 29, 29–34. [Google Scholar] [CrossRef] [PubMed]
- Barnes, C.; Pacheco, F.; Dhar, M.; Portnoy, J. Alternaria and Cladosporium fungal allergen epitopes are denatured by sodium hypochlorite. World Allergy Organ. J. 2009, 2, 296–302. [Google Scholar] [CrossRef] [PubMed]
- Dixit, A.; Kwilinski, K. 969 Cladosporium sphaerospermum-A new allergic species. J. Allergy Clin. Immunol. 2000, 105, S328. [Google Scholar] [CrossRef]
- Fukutomi, Y.; Taniguchi, M. Sensitization to fungal allergens: Resolved and unresolved issues. Allergol. Int. 2015, 64, 321–331. [Google Scholar] [CrossRef] [PubMed]
- Dixit, A.B.; Lewis, W.H.; Wedner, H.J. The allergens of Epicoccum nigrum link: I. Identification of the allergens by immunoblotting. J. Allergy Clin. Immunol. 1992, 90, 11–20. [Google Scholar] [CrossRef]
- Hoff, M.; Ballmer-Weber, B.K.; Niggemann, B.; Cistero-Bahima, A.; San Miguel-Moncín, M.; Conti, A.; Haustein, D.; Vieths, S. Molecular cloning and immunological characterisation of potential allergens from the mould Fusarium culmorum. Mol. Immunol. 2003, 39, 965–975. [Google Scholar] [CrossRef]
- Khosravi, A.; Fatahinia, M.; Shokri, H.; Yadegari, M. Allergens from Fusarium solani identified by immunoblotting in asthma patients in Iran. Arh. Hig. Rada Toksikol. 2012, 63, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Green, B.J.; Rittenour, W.R.; Hettick, J.M.; Janotka, E.; Beezhold, D.H. Characterization of Paecilomyces variotii allergens. J. Allergy Clin. Immunol. 2011, 127, AB264. [Google Scholar] [CrossRef]
- Park, H.S.; Jung, K.S.; Kim, S.O.; Kim, S.J. Hypersensitivity pneumonitis induced by Penicillium expansum in a home environment. Clin. Exp. Allergy 1994, 24, 383–385. [Google Scholar] [CrossRef] [PubMed]
- Lu-Ping, C.; Ning-Yuan, S.; Chia-Jung, Y.; Chiang, B.; Horng-Der, S. Identification and expression of Pen c2, a novel allergen from Penicillium citrinum. Biochem. J. 1999, 341, 51–59. [Google Scholar]
- Shen, H.D.; Chou, H.; Tam, M.F.; Chang, C.Y.; Lai, H.Y.; Wang, S.R. Molecular and immunological characterization of Pen ch 18, the vacuolar serine protease major allergen of Penicillium chrysogenum. Allergy 2003, 58, 993–1002. [Google Scholar] [CrossRef] [PubMed]
- Vijay, H.M.; Abebe, M.; Kumar, V.; DeVouge, M.; Schrader, T.; Thaker, A.; Comtois, P.; Escamilla-Garcia, B. Allergenic and mutagenic characterization of 14 Penicillium species. Aerobiologia 2005, 21, 95–103. [Google Scholar] [CrossRef]
- Sevinc, M.S.; Kumar, V.; Abebe, M.; Mohottalage, S.; Kumarathasan, P.; Vincent, R.; Vijay, H.M. Expression and characterization of Pen b 26 allergen of Penicillium brevicompactum in Escherichia coli. Protein Expr. Purif. 2009, 65, 8–14. [Google Scholar] [CrossRef] [PubMed]
- Iwen, P.C.; Schutte, S.D.; Florescu, D.F.; Noel-Hurst, R.K.; Sigler, L. Invasive Scopulariopsis brevicaulis infection in an immunocompromised patient and review of prior cases caused by Scopulariopsis and Microascus species. Med. Mycol. 2012, 50, 561–569. [Google Scholar] [CrossRef] [PubMed]
- Nayak, A.P.; Green, B.J.; Janotka, E.; Blachere, F.M.; Vesper, S.J.; Beezhold, D.H.; Schmechel, D. Production and characterization of IgM monoclonal antibodies against hyphal antigens of Stachybotrys species. Hybridoma 2011, 30, 29–36. [Google Scholar] [CrossRef] [PubMed]
- Das, S.; Gupta-Bhattacharya, S. Trichoderma harzianum: Occurrence in the air and clinical significance. Aerobiologia 2009, 25, 137–145. [Google Scholar] [CrossRef]
- Chou, H.; Tam, M.F.; Lee, S.S.; Tai, H.Y.; Chang, C.Y.; Chou, C.T.; Shen, H.D. A vacuolar serine protease (Rho m2) is a major allergen of Rhodotorula mucilaginosa and belongs to a class of highly conserved pan-fungal allergens. Int. Arch. Allergy Immunol. 2005, 138, 134–141. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Z.; Li, L.; Wan, Z.; Chen, W.; Liu, H.; Li, R. Simultaneous detection and identification of Aspergillus and mucorales species in tissues collected from patients with fungal rhinosinusitis. J. Clin. Microbiol. 2011, 49, 1501–1507. [Google Scholar] [CrossRef] [PubMed]
- Jerath, V.P.; Sood, M.; Nishchal, R. Prevalance of skin reactivity to fungal antigens in patients of nasobronchial allergy of Jalandhar and neighbouring area in Punjab. Indian J. Allergy Asthma Immunol. 2012, 26, 73–76. [Google Scholar] [CrossRef]
- Sridhara, S.; Gangal, S.V.; Joshi, A.P. Immunocheraical investigation of allergens from Rhizopus nigricans. Allergy 1990, 45, 577–586. [Google Scholar] [CrossRef] [PubMed]
- Sirkar, G.; Chakrabarti, H.S.; Saha, B.; Gupta-Bhattacharya, S. Identification of aero-allergens from Rhizopus oryzae: An immunoproteomic approach. J. Proteom. 2012, 77, 455–468. [Google Scholar] [CrossRef] [PubMed]
- Anaissie, E.J.; Stratton, S.L.; Dignani, M.C.; Lee, C.K.; Summerbell, R.C.; Rex, J.H.; Monson, T.P.; Walsh, T.J. Pathogenic molds (including Aspergillus species) in hospital water distribution systems: A 3-year prospective study and clinical implications for patients with hematologic malignancies. Blood 2003, 101, 2542–2546. [Google Scholar] [CrossRef] [PubMed]
- Hamada, N.; Abe, N. Physiological characteristics of 13 common fungal species in bathrooms. Mycoscience 2009, 50, 421–429. [Google Scholar] [CrossRef]
- Lotrakul, P.; Deenarn, P.; Prasongsuk, S.; Punnapayak, H. Isolation of Aureobasidium pullulans from bathroom surfaces and their antifungal activity against some Aspergilli. Afr. J. Microbiol. Res. 2009, 3, 253–257. [Google Scholar]
- Lian, X.; de Hoog, G.S. Indoor wet cells harbour melanized agents of cutaneous infection. Med. Mycol. 2010, 48, 622–628. [Google Scholar] [CrossRef] [PubMed]
- Abe, N.; Hamada, N. Molecular characterization and surfactant utilization of Scolecobasidium isolates from detergent-rich indoor environments. Biocontrol Sci. 2011, 16, 139–147. [Google Scholar] [CrossRef] [PubMed]
- Warris, A.; Klaassen, C.H.W.; Meis, J.F.G.M.; de Ruiter, M.T.; de Valk, H.A.; Abrahamsen, T.G.; Gaustad, P.; Verweij, P.E. Molecular epidemiology of Aspergillus fumigatus isolates recovered from water, air, and patients shows two clusters of genetically distinct strains. J. Clin. Microbiol. 2003, 41, 4101–4106. [Google Scholar] [CrossRef] [PubMed]
- Verweij, P.E.; Snelders, E.; Kema, G.J.H.; Mellado, E.; Melchers, W.J.G. Azole resistance in Aspergillus fumigatus: A side-effect of environmental fungicide use? Lancet Infect. Dis. 2009, 9, 789–795. [Google Scholar] [CrossRef]
- Nucci, M.; Anaissie, E. Fusarium infections in immunocompromised patients. Clin. Microbiol. Rev. 2007, 20, 695–704. [Google Scholar] [CrossRef] [PubMed]
- Guarro, J.; Gene, J. Opportunistic fusarial infections in humans. Eur. J. Clin. Microbiol. Infect. Dis. 1995, 14, 741–754. [Google Scholar] [CrossRef] [PubMed]
- Dóczi, I.; Gyetvai, T.; Kredics, L.; Nagy, E. Involvement of Fusarium spp. in fungal keratitis. Clin. Microbiol. Infect. 2004, 10, 773–776. [Google Scholar] [CrossRef] [PubMed]
- Bernal, M.D.; Acharya, N.R.; Lietman, T.M.; Strauss, E.C.; McLeod, S.D.; Hwang, D.G. Outbreak of Fusarium keratitis in soft contact lens wearers in San Francisco. Arch. Ophthalmol. 2006, 124, 1051–1053. [Google Scholar] [CrossRef] [PubMed]
- Muñoz, P.; Marín, M.; Tornero, P.; Martín Rabadán, P.; Rodríguez-Creixéms, M.; Bouza, E. Successful outcome of Scedosporium apiospermum disseminated infection treated with voriconazole in a patient receiving corticosteroid therapy. Clin. Infect. Dis. 2000, 31, 1499–1501. [Google Scholar] [CrossRef] [PubMed]
- Munk, S.; Johansen, C.; Stahnke, L.H.; Adler-Nissen, J. Microbial survival and odor in laundry. J. Surfactants Deterg. 2001, 4, 385–394. [Google Scholar] [CrossRef]
- Stapleton, K.; Hill, K.; Day, K.; Perry, D.J.; Dean, R.J. The potential impact of washing machines on laundry malodour generation. Lett. Appl. Microbiol. 2013, 56, 299–306. [Google Scholar] [CrossRef] [PubMed]
- Matos, T.; de Hoog, G.S.; de Boer, A.G.; de Crom, I.; Haase, G. High prevalence of the neurotrope Exophiala dermatitidis and related oligotrophic black yeasts in sauna facilities. Mycoses 2002, 45, 373–377. [Google Scholar] [CrossRef] [PubMed]
- Perlroth, J.; Choi, B.; Spellberg, B. Nosocomial fungal infections: Epidemiology, diagnosis and treatment. Med. Mycol. 2007, 45, 321–346. [Google Scholar] [CrossRef] [PubMed]
- Sabino, R.; Sampaio, P.; Rosado, L.; Videira, Z.; Grenouillet, F.; Pais, C. Analysis of clinical and environmental Candida parapsilosis isolates by microsatellite genotyping—A tool for hospital infections surveillance. Clin. Microbiol. Infect. 2015, 21. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.L.; Elfman, L.; Mi, Y.; Wieslander, G.; Smedje, G.; Norbäck, D. Indoor molds, bacteria, microbial volatile organic compounds and plasticizers in schools-associations with asthma and respiratory symptoms in pupils. Indoor Air 2007, 17, 153–163. [Google Scholar] [CrossRef] [PubMed]
- WHO. Guidelines for Indoor Air Quality: Dampness and Mould, 1st ed.; World Health Organization: Copenhagen, Denmark, 2009; p. 248. [Google Scholar]
- Brewer, J.H.; Thrasher, J.D.; Straus, D.C.; Madison, R.A.; Hooper, D. Detection of mycotoxins in patients with chronic fatigue syndrome. Toxins 2013, 5, 605–617. [Google Scholar] [CrossRef] [PubMed]
- Al-Gabr, H.M.; Zheng, T.; Yu, X. Fungi contamination of drinking water. Rev. Environ. Contam. Toxicol. 2014, 228, 121–139. [Google Scholar] [CrossRef] [PubMed]
- Miller, J.D.; Sun, M.; Gilyan, A.; Roy, J.; Rand, T.G. Inflammation associated gene transcription and expression in mouse lungs induced by low molecular weight compounds from fungi from the built environment. Chem. Biol. Interact. 2010, 183, 113–124. [Google Scholar] [CrossRef] [PubMed]
- Kirjavainen, P.V.; Täubel, M.; Karvonen, A.M.; Sulyok, M.; Tiittanen, P.; Krska, R.; Hyvärinen, A.; Pekkanen, J. Microbial secondary metabolites in homes in association with moisture damage and asthma. Indoor Air 2016, 26, 448–456. [Google Scholar] [CrossRef] [PubMed]
- Mayer, S.; Curtui, V.; Usleber, E.; Gareis, M. Airborne mycotoxins in dust from grain elevators. Mycotoxin Res. 2007, 23, 94–100. [Google Scholar] [CrossRef] [PubMed]
- Mayer, S.; Engelhart, S.; Kolk, A.; Blome, H. The significance of mycotoxins in the framework of assessing workplace related risks. Mycotoxin Res. 2008, 24, 151–164. [Google Scholar] [CrossRef] [PubMed]
- Mayer, S.; Vishwanath, V.; Sulyok, M. Airborne workplace exposure to microbial metabolites in waste sorting plants. In Bioaerosols, 1st ed.; Johanning, E., Morrey, P.R., Auger, P., Eds.; Fungal Research Group Foundation Inc.: Albany, NY, USA, 2012. [Google Scholar]
- Viegas, S.; Veiga, L.; Almeida, A.; dos Santos, M.; Carolino, E.; Viegas, C. Occupational exposure to aflatoxin B1 in a Portuguese poultry slaughterhouse. Ann. Occup. Hyg. 2015, 60, 176–183. [Google Scholar] [CrossRef] [PubMed]
- Viegas, S.; Veiga, L.; Figueiredo, P.; Almeida, A.; Carolino, E.; Sabino, R.; Veríssimo, C.; Viegas, C. Occupational exposure to aflatoxin B1 in swine production and possible contamination sources. J. Toxicol. Environ. Health A 2013, 76, 944–951. [Google Scholar] [CrossRef] [PubMed]
- Boonen, J.; Malysheva, S.V.; Taevernier, L.; Di Mavungu, J.D.; De Saeger, S.; De Spiegeleer, B. Human skin penetration of selected model mycotoxins. Toxicology 2012, 301, 21–32. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, T. Airborne fungal colony-forming units in outdoor and indoor environments in Yokohama, Japan. Mycopathologia 1997, 139, 23–33. [Google Scholar] [CrossRef] [PubMed]
- Kauffman, H.F.; van der Heide, S. Exposure, sensitization, and mechanisms of fungus-induced asthma. Curr. Allergy Asthma Rep. 2003, 3, 430–437. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, N.; Shendell, D.G.; Peccia, J. Assessing allergenic fungi in house dust by floor wipe sampling and quantitative PCR. Indoor Air 2011, 21, 521–530. [Google Scholar] [CrossRef] [PubMed]
- Amend, A.S.; Seifert, K.A.; Samson, R.; Bruns, T.D. Indoor fungal composition is geographically patterned and more diverse in temperate zones than in the tropics. Proc. Natl. Acad. Sci. USA 2010, 107, 13748–13753. [Google Scholar] [CrossRef] [PubMed]
- Dannemiller, K.; Lang-Yona, N.; Yamamoto, N.; Rudich, Y.; Peccia, J. Combining real-time PCR and next-generation DNA sequencing to provide quantitative comparisons of fungal aerosol populations. Atmos. Environ. 2014, 84, 113–121. [Google Scholar] [CrossRef]
- Kredics, L.; Antal, Z.; Dóczi, I.; Manczinger, L.; Kevei, F.; Nagy, E. Clinical importance of the genus Trichoderma. A review. Acta Microbiol. Immunol. Hung. 2003, 50, 105–117. [Google Scholar] [CrossRef] [PubMed]
- Low, C.-Y.; Rotstein, C. Emerging fungal infections in immunocompromised patients. F1000 Med. Rep. 2011, 3, 14. [Google Scholar] [CrossRef] [PubMed]
- Mershon-Shier, K.L.; Deville, J.G.; Delair, S.; Fothergill, A.W.; Wickes, B.; de Hoog, G.S.; Sutton, D.A.; Lewinski, M.A. Aureobasidium pullulans var. melanogenum fungemia in a pediatric patient. Med. Mycol. 2011, 49, 80–83. [Google Scholar] [CrossRef] [PubMed]
- Chowdhary, A.; Kathuria, S.; Agarwal, K.; Sachdeva, N.; Singh, P.K.; Jain, S.; Meis, J.F. Voriconazole-resistant Penicillium oxalicum: An emerging pathogen in immunocompromised hosts. Open Forum Infect. Dis. 2014, 1, ofu029. [Google Scholar] [CrossRef] [PubMed]
- Denning, W.D.; Pashley, C.; Hartl, D.; Wardlaw, A.; Godet, C.; Del Giacco, S.; Delhaes, L.; Sergejeva, S. Fungal allergy in asthma-state of the art and research needs. Clin. Transl. Allergy 2014, 4, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Vinh, D.C.; Sugui, J.A.; Hsu, A.P.; Freeman, A.F.; Holland, S.M. Invasive fungal disease in autosomal-dominant hyper-IgE syndrome. J. Allergy Clin. Immunol. 2010, 125, 1389–1390. [Google Scholar] [CrossRef] [PubMed]
- Khanna, N.; Stuehler, C.; Lünemann, A.; Wójtowicz, A.; Bochud, P.Y.; Leibundgut-Landmann, S. Host response to fungal infections—How immunology and host genetics could help to identify and treat patients at risk. Swiss Med. Wkly. 2016, 146, w14350. [Google Scholar] [CrossRef] [PubMed]
- Pana, Z.D.; Farmaki, E.; Roilides, E. Host genetics and opportunistic fungal infections. Clin. Microbiol. Infect. 2014, 12, 1254–1264. [Google Scholar] [CrossRef] [PubMed]
- Johnson, D.C. Chronic candidal bronchitis: A consecutive series. Open Respir. Med. J. 2012, 6, 145–149. [Google Scholar] [CrossRef] [PubMed]
- Denning, W.D. Global burden of human fungal diseases and their underlying diseases. In Proceedings of the International Union of Microbiological Societies, Montreal, QC, Canada, 27 July–1 August 2014; Abstract Number BR-01.02. p. 15. [Google Scholar]
- HSE/HPSC. Guidelines for the Prevention and Control of Infection from Water Systems in Healthcare Facilities, 1st ed.; Health Service Executive; Health Protection Surveillance Center: Dublin, Ireland, 2015; p. 100. [Google Scholar]
- SIS. SS 028192: Vattenundersökningar—Mikrosvampar i vatten—Kvantitativ Bestämning med Membranfiltermetod (English: Microfungi in Water—Quantitative Determination with the Membrane Filter Method.), 1st ed.; Swedish Standard Institute (SIS): Stockholm, Sweden, 1989; p. 4. [Google Scholar]
- AIHA. Field Guide for the Determination of Biological Contaminants in Environmental Samples, 1st ed.; American Industrial Hygiene Association: Fairfax, VA, USA, 1996; p. 174. [Google Scholar]
| Fungal Species | BSL * | Water Type | Country | Reference | |||
|---|---|---|---|---|---|---|---|
| Ground Water | Surface Water | Tap Water | Non-Mineral Bottled Water | ||||
| Ascomycota (phylum) | |||||||
| Acremonium psammosporum | 1 | + | − | + | − | Germany | [11] |
| Acremonium spp. | 1/2 | + | + | + | − | Germany, Greece, Slovakia, France, Austria, Portugal, Norway, Belgium, Serbia, UK, Sweden, Hungary | [5,6,7,8,9,10,11,12,13,14,16,18,79] |
| Acrostalagmus luteoalbus | 1 | + | + | + | − | Germany, Serbia | [11,12] |
| Alternaria alternata | 1 | + | + | + | − | Austria, Portugal, Ukraine, Serbia, Slovenia, UK, Hungary | [9,12,14,15,17,79,80] |
| Alternaria atra | 1 | − | + | − | − | UK | [9] |
| Alternaria botrytis | 1 | − | − | + | − | UK | [9] |
| Alternaria infectoria | 1 | − | + | − | − | Portugal, UK | [9,15] |
| Alternaria spp. | 1 | + | − | + | − | Greece, Slovakia, Portugal, Norway, Hungary, Belgium, Spain, Germany, UK | [7,8,9,10,13,16,18,23,81,82] |
| Alternaria tenuissima | 1 | − | − | + | − | Hungary | [79] |
| Arthrinium phaeospermum | 1 | − | + | + | − | Norway, UK | [9,20] |
| Arthrobotrys spp. | 1 | + | − | − | − | Slovakia | [7] |
| Arthrographis spp. | 1/2 | − | − | + | − | Poland, Norway, UK | [9,10,66] |
| Ascochyta spp. | 1 | − | − | + | − | UK | [9] |
| Aspergillus aculeatus | 1 | − | + | − | − | UK | [9] |
| Aspergillus alliaceus | 1 | − | + | − | − | Portugal | [15] |
| Aspergillus brasiliensis | 1 | − | + | − | − | Portugal | [15] |
| Aspergillus calidoustus | 1 | − | + | + | − | Portugal, Norway | [18,20] |
| Aspergillus candidus | 1 | − | + | − | − | Serbia | [12] |
| Aspergillus carbonarius | 1 | − | − | + | − | Greece | [8] |
| Aspergillus chevalieri | 1 | − | + | − | − | Portugal | [15] |
| Aspergillus clavatus | 1 | + | + | + | − | Norway, UK | [9,20] |
| Aspergillus fischeri | 1 | − | − | + | − | Slovenia | [80] |
| Aspergillus flavus | 2 | + | + | + | − | Germany, Greece, Belgium, Serbia, UK | [8,9,11,12,16] |
| Aspergillus fumigatus | 2 | + | + | + | + | Germany, Greece, Poland, Hungary, Norway, Portugal, The Netherlands, Finland, Belgium, Serbia, UK | [8,9,10,11,12,15,16,18,20,28,66,83,84,85] |
| Aspergillus glaucus | 1 | − | − | + | − | Greece | [8] |
| Aspergillus inflatus | 1 | − | + | − | − | Norway | [20] |
| Aspergillus insuetus | 1 | + | − | − | − | Portugal | [18] |
| Aspergillus japonicus | 1 | − | + | − | − | UK | [9] |
| Aspergillus nidulans | 1 | − | − | + | − | Greece, Belgium | [8,16] |
| Aspergillus niger | 1 | + | + | + | − | Germany, Greece, Poland, Norway, Belgium, Ukraine, Serbia, UK, Portugal | [8,9,10,11,12,16,17,18,20,28] |
| Aspergillus ochraceus | 1 | − | − | + | − | Greece | [8] |
| Aspergillus ostianus | 1 | − | − | + | − | Greece | [8] |
| Aspergillus parasiticus | 1 | − | − | + | − | Greece, Poland | [8,28] |
| Aspergillus parvulus | 1 | − | − | + | − | UK | [9] |
| Aspergillus repens | 1 | + | − | − | − | Portugal | [18] |
| Aspergillus restrictus | 1 | + | − | + | − | Greece, The Netherlands | [8,85] |
| Aspergillus sydowii | 1 | − | + | + | − | Norway, Belgium | [16,20] |
| Aspergillus terreus | 1 | + | + | + | − | Greece, Austria, Portugal, Norway, UK | [8,9,10,14,15,18] |
| Aspergillus tubingensis | 1 | − | + | − | − | Portugal | [15] |
| Aspergillus ustus | 1 | + | + | + | − | Poland, Norway, Portugal, Serbia | [12,15,20,28] |
| Aspergillus versicolor | 1 | + | + | + | + | Germany, Poland, Serbia, Slovenia, UK | [9,11,12,28,80] |
| Aspergillus viridinutans | 1 | − | + | − | − | Portugal | [18] |
| Aspergillus spp. | 1/2 | + | − | + | − | Slovakia, France, Austria, Portugal, Norway, Spain, Slovenia, Hungary | [5,7,10,13,14,19,79,81] |
| Asteroma sp. | 1 | − | + | − | − | UK | [9] |
| Asteromella sp. | 1 | − | − | + | − | UK | [9] |
| Aureobasidium melanogenum | 1 | + | − | + | − | Slovenia | [19,67,80] |
| Aureobasidium pullulans | 1 | + | + | + | + | Greece, Norway, Austria, Ukraine, Serbia | [8,12,14,17,20,86] |
| Aureobasidium spp. | 1 | + | + | + | − | Slovakia, UK, Portugal, Hungary | [7,9,18,79] |
| Beauveria bassiana | 1 | + | + | + | − | Norway, Austria, UK, Portugal | [9,14,18,20] |
| Beauveria brongniartii | 1 | − | + | − | − | Norway, UK | [9,20] |
| Beauveria spp. | 1 | + | − | − | − | Slovakia | [7] |
| Bionectria ochroleuca | 1 | + | − | − | − | Portugal | [18] |
| Bionectria sp. | No data | + | − | − | − | Portugal | [18] |
| Bipolaris spp. | 1/2 | − | − | + | − | Greece | [8] |
| Biscogniauxia sp. | No data | − | + | − | − | Portugal | [18] |
| Bisifusarium dimerum | 1 | + | − | + | − | Norway, Slovenia | [19,20,67] |
| Boeremia exigua | 1 | − | − | + | − | UK | [9] |
| Botryotrichum spp. | 1 | + | − | − | − | Slovakia | [7] |
| Botrytis cinerea | 1 | − | + | + | − | Norway, Portugal, Serbia, UK | [9,12,15,20] |
| Botrytis elliptica | 1 | − | + | − | − | Norway | [20] |
| Byssochlamys lagunculariae | 1 | − | + | − | − | Norway | [20] |
| Cadophora luteo-olivacea | 1 | + | − | − | − | Germany | [23] |
| Cadophora malorum | 1 | + | + | + | − | Germany, Poland, Norway, Austria | [14,20,23,28] |
| Cadophora melinii | 1 | − | + | − | − | Norway | [20] |
| Candida albicans | 2 | − | + | − | − | Ukraine | [17] |
| Candida glaebosa | 1 | − | − | + | − | Slovenia | [67] |
| Candida intermedia | 1 | − | − | + | − | Poland, Slovenia | [66,67] |
| Candida orthopsilosis | 2 | − | − | + | − | Slovenia | [19] |
| Candida parapsilosis | 2 | + | − | + | − | Poland, Slovenia | [19,66,67] |
| Candida pararugosa | 1 | − | − | + | − | Slovenia | [19,67] |
| Candida pseudointermedia | 1 | − | − | + | − | Slovenia | [19] |
| Candida saitoana | 1 | − | − | + | − | Slovenia | [19] |
| Candida sake | 1 | − | + | − | − | Portugal | [15] |
| Candida sp. | No data | + | − | + | − | Portugal, Greece | [8,15] |
| Candida tropicalis | 2 | − | − | + | − | Greece | [8] |
| Candida versatilis | 1 | − | − | + | − | Poland | [66] |
| Capronia munkii | 1 | − | + | − | − | Portugal | [18] |
| Capronia pilosella | 1 | − | − | + | − | Germany | [23] |
| Capronia sp. | No data | − | − | + | − | Slovenia | [67] |
| Cephalosporium spp. | 1/2 | + | + | + | − | Slovakia, Portugal | [7,18] |
| Ceratocystis fimbriata | 1 | − | + | − | − | Norway | [20] |
| Chaetomium globosum | 1 | − | + | − | − | Norway, Serbia, UK | [9,12,20] |
| Chaetomium spp. | 1 | + | − | + | − | Greece, Norway, Portugal | [8,13,20] |
| Chalara sp. | No data | + | − | + | − | Germany | [11] |
| Chalaropsis spp. | 1 | + | − | − | − | Slovakia | [7] |
| Chrysosporium spp. | 1 | − | − | + | − | Greece | [8] |
| Chrysonilia sp. | No data | + | + | − | − | Norway | [20] |
| Cistella acuum | 1 | + | − | + | − | Austria | [14] |
| Cladosporium cladosporioides | 1 | + | + | + | − | Germany, Greece, Poland, Norway, Portugal, The Netherlands, Serbia, Slovenia, UK, Hungary | [8,9,11,12,15,18,20,23,28,79,80,85] |
| Cladosporium cucumerinum | 1 | − | + | − | − | Serbia | [12] |
| Cladosporium diaphanum | 1 | − | + | − | − | Serbia | [12] |
| Cladosporium halotolerans | 1 | + | + | + | − | Portugal, Germany | [15,18,23] |
| Cladosporium herbarum | 1 | + | + | + | − | Germany, Norway, Portugal, Serbia, UK | [9,11,12,15,20] |
| Cladosporium macrocarpum | 1 | − | + | − | − | Portugal | [18] |
| Cladosporium oxysporum | 1 | − | + | − | − | Serbia | [12] |
| Cladosporium pseudocladosporioides | 1 | − | − | + | − | Slovenia | [80] |
| Cladosporium sphaerospermum | 1 | − | + | + | − | Poland, Norway, UK | [9,20,28] |
| Cladosporium spp. | 1 | + | + | + | + | Greece, Slovakia, France, Austria, Portugal, Norway, Hungary, Belgium, Ukraine, Spain, UK | [5,7,8,9,10,13,14,15,16,17,18,81,82,83,84,87] |
| Cladosporium tenuissimum | 1 | − | + | − | − | Portugal | [15] |
| Cladosporium variabile | 1 | − | + | − | − | Serbia | [12] |
| Clavispora lusitaniae | 1 | − | − | + | − | Slovenia | [19] |
| Clethridium corticola | 1 | − | − | + | − | UK | [9] |
| Clonostachys candelabrum | 1 | − | + | + | − | Poland | [28] |
| Coniochaeta hoffmannii | 1 | + | + | + | − | Norway, Austria, Portugal | [14,18,20] |
| Coniochaeta velutina | 1 | − | + | − | − | Portugal | [18] |
| Coniothyrium olivaceum | 1 | − | + | + | − | UK | [9] |
| Cordyceps bassiana | 1 | + | − | + | − | Austria | [14] |
| Cosmospora arxii | 1 | + | − | + | − | Germany | [11] |
| Cosmospora berkeleyana | 1 | + | − | + | − | Germany | [11] |
| Cosmospora butyri | 1 | + | + | − | − | Norway | [20] |
| Cosmospora sp. | No data | − | + | − | − | Portugal | [15] |
| Curvularia spp. | 1/2 | + | − | + | − | Greece, Slovakia | [7,8] |
| Cyclothyrium sp. | No data | − | − | + | − | UK | [9] |
| Cylindrocarpon spp. | 1/2 | + | + | − | − | Slovakia, UK | [7,9] |
| Cyphellophora europaea | 2 | − | − | + | − | Germany | [23] |
| Cyphellophora reptans | 1 | + | − | + | − | Germany | [23] |
| Cyphellophora sessilis | 1 | + | − | + | − | Germany | [11,23] |
| Cytospora sp. | No data | − | + | + | − | UK | [9] |
| Dactylaria spp. | 1/2 | + | − | + | − | Slovakia, Austria | [7,14] |
| Dactylella spp. | 1 | + | − | + | − | Slovakia | [7] |
| Debaryomyces hansenii | 1 | − | − | + | + | Poland, Slovenia, France | [5,19,66] |
| Didymella molleriana | 1 | + | + | + | − | Norway, Austria, Portugal | [14,15,18,20] |
| Didymella musae | 1 | − | + | + | − | UK | [9] |
| Diplocladium spp. | No data | + | − | − | − | Slovakia | [7] |
| Discosporium sp. | No data | − | − | + | − | UK | [9] |
| Doratomyces spp. | 1 | − | − | + | − | Greece | [8] |
| Embellisia sp. | No data | − | + | − | − | UK | [9] |
| Emmonsia spp. | 1/2 | − | − | + | − | Greece | [8] |
| Epicoccum nigrum | 1 | + | + | + | − | Norway, Austria, UK, Serbia | [9,12,14,20] |
| Epicoccum spp. | 1 | − | − | + | − | Greece | [8] |
| Eupenicillium sp. | No data | − | − | + | − | UK | [9] |
| Eurotium spp. | 1 | − | − | + | − | Greece | [8] |
| Exophiala alcalophila | 1 | − | − | + | − | Slovenia, Germany | [19,23,67] |
| Exophiala angulospora | 1 | + | − | + | − | Germany | [11,23] |
| Exophiala cancerae | 1 | − | − | + | − | Germany | [23] |
| Exophiala castellanii | 2 | + | − | + | − | Germany, Poland | [11,23,66] |
| Exophiala dermatitidis | 2 | + | − | + | − | Slovenia | [19,67] |
| Exophiala equina | 1 | + | − | + | − | Germany | [23] |
| Exophiala jeanselmei | 2 | − | − | + | − | Poland, UK | [9,66] |
| Exophiala lecanii-corni | 1 | − | − | + | − | Slovenia, Germany | [19,23,67] |
| Exophiala mesophila | 1 | + | − | + | − | Slovenia, Germany | [19,23] |
| Exophiala oligosperma | 2 | + | − | + | − | Slovenia, Germany | [19,23] |
| Exophiala opportunistica | 1 | − | − | + | − | Germany | [23] |
| Exophiala phaeomuriformis | 2 | − | − | + | − | Slovenia, Germany | [19,23,67] |
| Exophiala pisciphila | 1 | + | − | + | − | Germany | [11] |
| Exophiala psychrophila | 1 | + | − | + | − | Germany | [23] |
| Exophiala salmonis | 1 | + | − | + | − | Germany | [23] |
| Exophiala spinifera | 2 | − | − | + | + | Poland | [66] |
| Exophiala spp. | 1/2 | + | − | + | − | Germany, Greece | [8,11] |
| Exophiala xenobiotica | 1 | + | − | + | − | Slovenia, Germany | [19,23] |
| Fusarium begoniae | 1 | − | + | − | − | Portugal | [15] |
| Fusarium culmorum | 1 | − | + | + | − | Serbia, UK | [9,12] |
| Fusarium flocciferum | 1 | − | + | − | − | UK | [9] |
| Fusarium foetens | 1 | − | + | − | − | Portugal | [18] |
| Fusarium incarnatum | 1 | − | + | − | − | Serbia | [12] |
| Fusarium oxysporum | 2 | + | + | + | − | Norway, Serbia, UK | [9,12,20] |
| Fusarium solani | 2 | + | + | + | − | Germany, Greece, Poland, Serbia, UK | [8,9,11,12,28] |
| Fusarium sporotrichioides | 1 | − | + | − | − | Serbia | [12] |
| Fusarium spp. | 1/2 | + | + | + | + | Germany, Slovakia, Austria, Portugal, Norway, Belgium, Ukraine, Spain, Hungary, UK | [7,9,10,11,14,15,16,17,18,79,81,84,87] |
| Fusarium torulosum | 1 | − | + | − | − | UK | [9] |
| Fusicolla aquaeductuum | 1 | − | − | + | − | UK | [9] |
| Fusicolla merismoides | 1 | + | − | + | − | Germany | [11] |
| Galactomyces geotrichum | 1 | − | + | + | − | Slovenia, Portugal, Poland, Serbia, UK | [9,12,18,19,28,67] |
| Geomyces sp. | No data | + | − | + | − | Germany | [11] |
| Geotrichum spp. | 1/2 | + | + | + | − | Slovakia, Norway, Hungary | [7,20,79] |
| Gibberella avenacea | 1 | − | + | + | − | UK | [9] |
| Gibberella fujikuroi | 1 | − | + | − | − | UK | [9] |
| Gibberella gordonii | 1 | − | + | − | − | Serbia | [12] |
| Gibberella intricans | 1 | − | + | − | − | UK | [9] |
| Gliocladium spp. | 1 | + | + | + | − | Greece, Slovakia, UK, Hungary | [7,8,9,79] |
| Graphium silanum | 1 | + | − | + | − | Austria | [14] |
| Hormiscium spp. | 1/2 | + | − | + | − | Slovakia | [7] |
| Hyphopichia burtonii | 1 | − | + | − | − | Portugal | [15] |
| Humicola grisea | 1 | − | − | + | − | Hungary | [79] |
| Isaria farinosa | 1 | + | + | + | − | Germany, Norway, Serbia | [11,12,20] |
| Issatchenkia orientalis | 1 | − | − | + | − | Poland | [66] |
| Kloeckera spp. | 1 | + | − | + | − | Greece, Portugal | [8,15] |
| Kluyveromyces lactis | 1 | − | − | + | − | Poland | [66] |
| Kluyveromyces marxianus | 1 | − | − | + | − | Poland | [66] |
| Lecanicillium lecanii | 1 | + | + | + | − | Germany, Poland, Norway | [11,20,28] |
| Leptodontidium sp. | No data | − | − | + | − | UK | [9] |
| Leptosphaeria sp. | No data | + | + | + | − | Austria, UK | [9,14] |
| Leucostoma persoonii | 1 | − | + | − | − | Norway | [20] |
| Mauginiella sp. | No data | − | − | + | − | UK | [9] |
| Melanospora simplex | 1 | − | + | + | − | Poland | [28] |
| Metarhizium carneum | 1 | + | + | − | − | Norway | [20] |
| Meyerozyma caribbica | 1 | − | − | + | − | Slovenia | [19,67] |
| Meyerozyma guilliermondii | 1 | − | − | + | − | Slovenia | [19] |
| Microdochium sp. | No data | + | − | + | − | Austria | [14] |
| Microsphaeropsis sp. | No data | − | + | − | − | UK | [9] |
| Microsporum spp. | 1/2 | − | − | + | − | Slovakia | [7] |
| Monascus ruber | 1 | − | + | − | − | Norway | [20] |
| Monilia spp. | 1/2 | + | − | + | − | Slovakia, Belgium | [7,16] |
| Nakazawaea holstii | 1 | − | + | − | − | Portugal | [15] |
| Neurospora sp. | No data | − | + | − | − | UK | [9] |
| Ochroconis musae | 1 | − | − | + | − | Germany | [23] |
| Ochroconis sp. | 1 | + | − | + | − | Germany | [11] |
| Oosporidium margaritiferum | 1 | − | − | + | − | Poland | [66] |
| Paecilomyces spp. | 1 | + | − | + | + | Slovakia, Austria, Norway, Belgium, Spain, Poland | [7,10,14,16,66,81] |
| Paecilomyces variotii | 1 | + | + | + | − | Norway, Austria, Greece | [8,14,20] |
| Papulaspora sp. | No data | + | − | + | − | Slovakia | [7] |
| Paraconiothyrium sp. | No data | − | + | − | − | Portugal | [15] |
| Paraphaeosphaeria minitans | 1 | − | + | − | − | Potugal | [18] |
| Paraphaeosphaeria sporulosa | 1 | − | + | − | − | Portugal | [15] |
| Paraphoma fimeti | 1 | + | − | + | − | Germany | [23] |
| Paspalomyces sp. | No data | + | − | + | − | Slovakia | [7] |
| Penicillium atrofulvum | 1 | − | + | − | − | Portugal | [18] |
| Penicillium aurantiogriseum | 1 | − | + | + | − | UK, Portugal | [9,15] |
| Penicillium brevicompactum | 1 | + | + | + | − | Germany, Norway, Portugal, UK | [9,11,13,18,20] |
| Penicillium canescens | 1 | − | + | − | − | Norway, Portugal, Serbia | [12,15,18,20] |
| Penicillium chrysogenum | 1 | + | + | + | + | Germany, Norway, Serbia, Slovenia, UK, Hungary | [9,11,12,20,80,84] |
| Penicillium citrinum | 1 | − | + | + | − | Norway, Portugal, UK | [9,15,18,20] |
| Penicillium corylophilum | 1 | + | + | + | − | Portugal, UK | [9,13,18] |
| Penicillium dierckxii | 1 | − | + | − | − | Portugal, Norway | [15,18,20] |
| Penicillium digitatum | 1 | − | + | − | − | Portugal | [18] |
| Penicillium echinulatum | 1 | − | + | − | − | UK | [9] |
| Penicillium expansum | 1 | − | + | + | − | Norway, Portugal, UK | [9,13,18,20] |
| Penicillium glabrum | 1 | + | + | + | + | Germany, Norway, Portugal, UK, France, Poland | [9,11,13,15,18,20,28,88] |
| Penicillium griseofulvum | 1 | − | + | + | − | Portugal, Serbia, UK | [9,12,13,15,18] |
| Penicillium hirsutum | 1 | − | − | + | − | UK | [9] |
| Penicillium implicatum | 1 | − | + | − | − | Norway, Portugal | [15,20] |
| Penicillium janczewskii | 1 | − | + | + | − | Norway, UK | [9,20] |
| Penicillium jensenii | 1 | − | + | − | − | Norway | [20] |
| Penicillium lanosum | 1 | − | + | − | − | Norway | [20] |
| Penicillium megasporum | 1 | − | + | − | − | Norway | [20] |
| Penicillium melanoconidium | 1 | − | + | − | − | Portugal | [15] |
| Penicillium melinii | 1 | − | + | − | − | Norway | [20] |
| Penicillium miczynskii | 1 | − | + | − | − | Norway | [20] |
| Penicillium montanense | 1 | + | + | − | − | Norway | [20] |
| Penicillium novae-zeelandiae | 1 | − | + | − | − | Portugal | [18] |
| Penicillium ochrochloron | 1 | − | + | − | − | Portugal | [15] |
| Penicillium ochrosalmoneum | 1 | − | − | + | − | UK | [9] |
| Penicillium olsonii | 1 | − | + | − | − | Norway, Portugal | [18,20] |
| Penicillium oxalicum | 1 | − | + | − | − | Norway | [20] |
| Penicillium pancosmium | 1 | − | + | − | − | Portugal | [18] |
| Penicillium paxilli | 1 | − | + | − | − | Norway | [20] |
| Penicillium phoeniceum | 1 | − | + | − | − | Norway | [20] |
| Penicillium purpurogenum | 1 | − | + | + | − | Norway, UK | [9,20] |
| Penicillium raistrickii | 1 | − | + | + | − | Norway, Portugal, UK | [9,13,15,20] |
| Penicillium resedanum | 1 | − | + | − | − | Serbia | [12] |
| Penicillium restrictum | 1 | − | + | − | − | Norway, Portugal | [15,18,20] |
| Penicillium roseopurpureum | 1 | − | + | − | − | Norway | [20] |
| Penicillium sanguifluum | 1 | − | + | − | − | Portugal | [18] |
| Penicillium scabrosum | 1 | − | + | − | − | Portugal | [15] |
| Penicillium simplicissimum | 1 | − | + | − | − | Norway, UK, Portugal | [9,18,20] |
| Penicillium solitum | 1 | − | + | + | − | Norway, UK, Portugal | [9,13,15,18,20] |
| Penicillium spinulosum | 1 | + | + | + | − | Norway, UK | [9,20] |
| Penicillium spp. | 1/2 | + | + | + | + | Germany, Greece, Slovakia, France, Austria, Norway, Belgium, Ukraine, Spain, Sweden, Portugal, Hungary | [5,6,7,8,10,11,14,16,17,79,81,87] |
| Penicillium thomii | 1 | − | + | − | − | Norway, Portugal, Serbia | [12,15,20] |
| Penicillium verrucosum | 1 | + | + | − | − | Norway, Serbia | [12,20] |
| Penicillium virgatum | 1 | − | + | − | − | Portugal | [18] |
| Penicillium waksmanii | 1 | − | + | + | − | Portugal, UK | [9,13] |
| Penicillium westlingii | 1 | − | + | − | − | Norway | [20] |
| Phaeosphaeria juncophila | 1 | + | − | + | − | Austria | [14] |
| Phialemonium sp. | No data | − | + | − | − | Portugal | [18] |
| Phialocephala dimorphospora | 1 | − | − | + | − | Germany | [23] |
| Phialophora cyclaminis | 1 | − | + | − | − | Norway | [20] |
| Phialophora fastigiata | 1 | + | + | + | − | Italy, Germany, Norway, UK | [9,20,23,89] |
| Phialophora spp. | 1/2 | + | − | + | − | Germany, Greece, Slovakia, Austria, Portugal, Sweden | [6,7,8,11,13,14] |
| Phialophora verrucosa | 2 | − | + | − | − | Norway | [20] |
| Phoma herbarum | 1 | + | + | + | − | Germany, Serbia | [11,12] |
| Phoma leveillei | 1 | + | + | + | − | Germany, Italy, UK | [9,11,89] |
| Phoma macrostoma | 1 | − | + | + | − | UK | [9] |
| Phoma medicaginis | 1 | − | + | + | − | Serbia, UK | [9,12] |
| Phoma sp. | No data | + | + | + | − | Poland, Norway, Portugal, Serbia | [10,12,15,20,28] |
| Phomatodes nebulosa | 1 | − | − | + | − | UK | [9] |
| Phomopsis spp. | 1 | + | − | + | − | Austria, UK | [9,14] |
| Pichia fermentans | 1 | − | − | + | − | Slovenia | [19] |
| Pichia membranifaciens | 1 | − | − | + | − | France, Greece | [5,8] |
| Pilidium concavum | 1 | + | + | + | − | UK, Portugal | [9,18] |
| Priceomyces carsonii | 1 | − | − | + | − | Poland | [66] |
| Prosthecium pyriforme | 1 | + | − | − | − | Portugal | [18] |
| Pseudeurotium hygrophilum | 1 | − | + | − | − | UK | [9] |
| Pseudogymnoascus pannorum | 1 | − | + | − | − | Norway | [20] |
| Pseudogymnoascus roseus | 1 | − | + | − | − | Norway | [20] |
| Pseudopithomyces sacchari | 1 | − | + | − | − | UK | [9] |
| Purpureocillium lilacinum | 1 | + | + | + | − | UK, Portugal, Poland, Norway, Italy | [9,18,20,28,89] |
| Pyrenochaeta spp. | 1/2 | − | + | + | − | Greece, Italy, UK | [8,9,89] |
| Pyrenochaeta unguis-hominis | 2 | − | − | + | − | Germany | [23] |
| Rhinocladiella similis | 2 | + | − | + | − | Slovenia, Germany | [19,23,67] |
| Saccharomycopsis capsularis | 1 | − | − | + | − | Poland | [66] |
| Saprochaete suaveolens | 1 | − | − | + | − | Poland | [66] |
| Sarocladium kiliense | 2 | − | + | + | − | Poland, UK | [9,66] |
| Sarocladium strictum | 1 | + | + | + | − | Germany, Italy, Norway, Serbia | [11,12,20,89] |
| Sarocladium terricola | 1 | − | + | + | − | Serbia, Poland | [12,28] |
| Sclerotinia sclerotiorum | 1 | − | − | + | − | Poland | [28] |
| Scopulariopsis acremonium | 1 | − | + | − | − | UK | [9] |
| Scopulariopsis brevicaulis | 2 | − | + | + | − | Greece, Norway, UK | [8,9,20] |
| Scopulariopsis fusca | 1 | − | + | + | − | Poland | [20,66] |
| Scopulariopsis spp. | 1/2 | − | − | + | − | Greece | [8] |
| Sepedonium spp. | 1 | − | − | + | − | Greece, Norway | [8,10] |
| Sporothrix spp. | 1/2 | − | + | + | − | UK | [9] |
| Stachybotrys chartarum | 1 | + | + | + | − | Poland, Portugal | [18,28] |
| Stachybotrys spp. | 1 | + | − | + | − | Greece, Slovakia | [7,8] |
| Staphylotrichum sp. | No data | − | + | − | − | Norway | [20] |
| Stemphylium sp. | No data | + | − | + | − | Slovakia | [7] |
| Stephanoma strigosum | 1 | − | − | + | − | Hungary | [79] |
| Sydowia polyspora | 1 | − | − | + | − | UK | [9] |
| Talaromyces funiculosus | 1 | − | + | − | − | Serbia | [12] |
| Talaromyces minioluteus | 1 | − | − | + | − | UK | [9] |
| Talaromyces pinophilus | 1 | − | − | + | − | UK | [9] |
| Talaromyces ruber | 1 | − | + | + | − | Poland | [28] |
| Talaromyces rugulosus | 1 | − | − | − | + | Poland | [66] |
| Talaromyces verruculosus | 1 | − | − | + | − | Slovenia | [67] |
| Trichoderma asperellum | 1 | − | + | − | − | Portugal | [18] |
| Trichoderma citrinoviride | 1 | − | + | + | − | Slovenia, Portugal | [18,80] |
| Trichoderma harzianum | 1 | + | + | + | − | Portugal, UK | [9,15,18] |
| Trichoderma koningii | 1 | − | + | + | − | Serbia, UK, Portugal | [9,12,18] |
| Trichoderma longibrachiatum | 1 | − | − | − | + | Poland | [66] |
| Trichoderma pleuroticola | 1 | − | + | − | − | Portugal | [18] |
| Trichoderma polysporum | 1 | − | + | + | − | UK | [9] |
| Trichoderma pseudokoningii | 1 | − | + | + | − | UK | [9] |
| Trichoderma spp. | 1 | + | + | + | − | Greece, Slovakia, Norway, France, Austria, Belgium, Spain, Serbia, Hungary | [5,7,8,10,12,14,16,20,79,81] |
| Trichoderma viride | 1 | + | + | + | − | Poland, Austria, Ukraine, Serbia | [12,14,17,28] |
| Trichomonascus ciferrii | 1 | − | − | + | − | Greece | [8] |
| Trichothecium sp. | No data | + | − | + | − | Greece, Slovakia, Hungary | [7,8,79] |
| Trichophyton sp. | No data | + | − | + | − | Slovakia | [7] |
| Tritirachium sp. | No data | + | − | + | − | Slovakia | [7] |
| Truncatella angustata | 1 | − | + | − | − | UK | [9] |
| Varicosporium spp. | 1 | + | − | − | − | Slovakia | [7] |
| Verticillium spp. | 1 | + | − | + | − | Greece, Slovakia, UK, Hungary | [7,8,9,79] |
| Volutella sp. | No data | + | − | + | − | Germany | [11] |
| Westerdykella dispersa | 1 | − | + | − | − | UK | [9] |
| Wickerhamomyces anomalus | 1 | − | − | + | − | Poland | [66] |
| Yarrowia lipolytica | 1 | − | − | + | − | Slovenia | [19] |
| Basidiomycota (phylum) | |||||||
| Apiotrichum montevideense | 1 | − | − | + | − | Slovenia | [19,67] |
| Cryptococcus sp. | No data | − | + | − | − | Portugal | [15] |
| Cystobasidiopsis lactophilus | 1 | − | − | + | − | Poland | [66] |
| Cystobasidium minuta | 1 | − | + | + | − | France, Portugal | [5,15] |
| Cystobasidium slooffiae | 1 | − | − | + | − | Slovenia | [19,67] |
| Cystofilobasidium lari-marini | 1 | − | − | + | − | Poland | [66] |
| Filobasidium magnum | 1 | − | − | − | + | Norway | [86] |
| Naganishia albida | 1 | − | + | − | − | Portugal | [15] |
| Rhizoctonia spp. | 1 | + | − | − | − | Slovakia | [7] |
| Rhodotorula glutinis | 1 | − | + | + | − | France, Ukraine | [5,17] |
| Rhodotorula mucilaginosa | 1 | + | − | + | − | Slovenia | [19,67] |
| Rhodotorula spp. | 1 | + | + | + | − | Germany, Greece, Poland, Austria, Portugal | [8,11,14,15,66] |
| Schizophyllum commune | 1 | − | − | + | − | Slovenia | [67] |
| Sporidiobolus salmonicolor | 1 | + | − | − | − | Slovenia | [19] |
| Sporobolomyces japonicus | 1 | − | − | + | − | Poland | [66] |
| Sporobolomyces ruberrimus | 1 | − | − | + | − | Slovenia | [80] |
| Sporotrichum spp. | 1/2 | + | + | − | − | Slovakia, UK | [7,9] |
| Stereum sp. | No data | − | − | + | − | UK | [9] |
| Tilletiopsis sp. | No data | + | − | + | − | Germany | [11] |
| Trametes versicolor | 1 | + | − | + | − | Austria | [14] |
| Trichosporon coremiiforme | 1 | + | − | − | − | Slovenia | [19] |
| Triodiomyces crassus | 1 | − | − | + | − | Slovenia | [19,67] |
| Mucoromycotina (subphylum) | |||||||
| Absidia cylindrospora | 1 | − | + | + | − | Norway, UK | [9,20] |
| Absidia glauca | 1 | − | + | + | − | Norway, UK | [9,20] |
| Absidia spp. | 1/2 | + | − | + | − | Slovakia, Spain | [7,81] |
| Chaetocladium brefeldii | 1 | − | + | − | − | UK | [9] |
| Cunninghamella elegans | 1 | − | + | − | − | Portugal | [18] |
| Gongronella butleri | 1 | − | + | − | − | UK | [9] |
| Lichtheimia corymbifera | 2 | − | + | − | − | Norway | [20] |
| Mortierella alpina | 1 | − | + | − | − | UK | [9] |
| Mortierella elongata | 1 | − | + | − | − | UK | [9] |
| Mortierella zychae | 1 | − | − | + | − | UK | [9] |
| Mucor azygosporus | 1 | − | + | − | − | Norway | [20] |
| Mucor circinelloides | 1 | − | + | + | − | Norway, UK | [9,20] |
| Mucor fuscus | 1 | − | + | − | − | UK | [9] |
| Mucor hiemalis | 1 | − | + | + | − | Norway, Serbia, UK | [9,12,20] |
| Mucor moelleri | 1 | − | + | − | − | UK, Portugal | [9,18] |
| Mucor mucedo | 1 | − | − | + | − | Greece | [8] |
| Mucor plumbeus | 1 | − | + | + | − | Norway, UK | [9,20] |
| Mucor racemosus | 1 | − | + | + | − | Portugal, UK | [9,15,18] |
| Mucor spp. | 1/2 | + | + | + | − | Germany, Slovakia, France, Norway, Spain, Serbia, Hungary | [5,7,10,11,12,18,79,81] |
| Mucor strictus | 1 | − | + | + | − | UK | [9] |
| Rhizomucor spp. | 1/2 | − | − | + | − | Norway | [10] |
| Rhizopus arrhizus | 1 | − | + | − | − | Ukraine | [17] |
| Rhizopus spp. | 1/2 | − | − | + | − | Greece, Slovakia, France, Norway, Spain | [5,7,8,10,81] |
| Rhizopus stolonifer | 1 | − | + | + | − | Portugal, UK, Serbia | [9,12,13] |
| Syncephalastrum racemosum | 1 | − | − | + | − | UK | [9] |
| Umbelopsis isabellina | 1 | − | + | − | − | UK | [9] |
| Umbelopsis ramanniana | 1 | − | + | + | − | UK | [9] |
| Fungal Species | Local or Systemic Infections | Allergenic Compounds | Mycotoxins Production | Irritative Compounds, MVOC, Odor | References |
|---|---|---|---|---|---|
| Alternaria: A. alternata | respiratory infections, skin and nail infections, keratitis | X | X | No data | [32,145] |
| Aspergillus: A. flavus A. fumigatus A. niger A. terreus A. ustus A. versicolor | disseminated infections, respiratory infections, subcutaneous infections, rhinocerebral infections, skin and nail infections, ear infections, keratitis | X | X | X | [32,146,147,148,149,150,151,152,153,154] |
| Aureobasidium: A. pullulans A. melanogenum | skin and nail infections, keratitis | X | No data | No data | [32,155] |
| Beauveria: B. bassiana | disseminated infections, keratitis | X | No data | No data | [32,156] |
| Botrytis: B. cinerea | No data | X | No data | No data | [157] |
| Candida: C. albicans C. parapsilosis species complex | disseminated infections, mucosal infections | X | No data | No data | [32,158,159] |
| Chaetomium: C. globosum | respiratory infections, rhinocerebral infections, skin and nail infections | X | X | No data | [32,160] |
| Cladosporium: C. cladosporioides C. herbarum C. sphaerospermum | respiratory infections, skin and nail infections, keratitis | X | No data | No data | [32,161,162,163] |
| Epicoccum: E. nigrum | No data | X | No data | No data | [164] |
| Exophiala: E. dermatitidis E. jeanselmei | disseminated infections, respiratory infections, skin and nail infections | No data | No data | No data | [32] |
| Fusarium: F. oxysporum F. solani | disseminated infections, keratitis, skin and nail infections | X | X | No data | [32,165,166] |
| Paecilomyces: P. variotii | disseminated infections, respiratory infections, keratitis, skin and nail infections | X | No data | No data | [32,167] |
| Penicillium: P. brevicompactum P. chrysogenum P. citrinum P. expansum P. glabrum P. simplicissimum | respiratory infections, endocarditis, rhinocerebral infections, keratitis | X | X | X | [32,151,168,169,170,171,172] |
| Purpureocillium: P. lilacinum | disseminated infections, respiratory infections, keratitis, subcutaneous infections, skin and nail infections | No data | No data | No data | [32] |
| Sarocladium: S. kiliense S. strictum | disseminated infections, respiratory infections, keratitis, subcutaneous infections, skin and nail infections | No data | No data | No data | [32] |
| Scopulariopsis: S. brevicaulis | skin and nail infections, keratitis, endocarditis | X | No data | No data | [32,173] |
| Stachybotrys: S. chartarum | respiratory infections | X | X | No data | [174] |
| Trichoderma: T. harzianum T. viride | disseminated infections, respiratory infections | X | X | X | [32,151,160,175] |
| Rhodotorula: R. mucilaginosa | catheter-related fungemia | X | No data | No data | [32,176] |
| Mucor: M. circinelloides M. hiemalis M. racemosus | disseminated infections, keratitis, rhinocerebral infections, skin and nail infections, subcutaneous infections | X | No data | No data | [32,177,178] |
| Rhizopus: R. arrhizus R. stolonifer | disseminated infections, keratitis, subcutaneous infections, skin and nail infections | X | No data | No data | [32,179,180] |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Babič, M.N.; Gunde-Cimerman, N.; Vargha, M.; Tischner, Z.; Magyar, D.; Veríssimo, C.; Sabino, R.; Viegas, C.; Meyer, W.; Brandão, J. Fungal Contaminants in Drinking Water Regulation? A Tale of Ecology, Exposure, Purification and Clinical Relevance. Int. J. Environ. Res. Public Health 2017, 14, 636. https://doi.org/10.3390/ijerph14060636
Babič MN, Gunde-Cimerman N, Vargha M, Tischner Z, Magyar D, Veríssimo C, Sabino R, Viegas C, Meyer W, Brandão J. Fungal Contaminants in Drinking Water Regulation? A Tale of Ecology, Exposure, Purification and Clinical Relevance. International Journal of Environmental Research and Public Health. 2017; 14(6):636. https://doi.org/10.3390/ijerph14060636
Chicago/Turabian StyleBabič, Monika Novak, Nina Gunde-Cimerman, Márta Vargha, Zsófia Tischner, Donát Magyar, Cristina Veríssimo, Raquel Sabino, Carla Viegas, Wieland Meyer, and João Brandão. 2017. "Fungal Contaminants in Drinking Water Regulation? A Tale of Ecology, Exposure, Purification and Clinical Relevance" International Journal of Environmental Research and Public Health 14, no. 6: 636. https://doi.org/10.3390/ijerph14060636
APA StyleBabič, M. N., Gunde-Cimerman, N., Vargha, M., Tischner, Z., Magyar, D., Veríssimo, C., Sabino, R., Viegas, C., Meyer, W., & Brandão, J. (2017). Fungal Contaminants in Drinking Water Regulation? A Tale of Ecology, Exposure, Purification and Clinical Relevance. International Journal of Environmental Research and Public Health, 14(6), 636. https://doi.org/10.3390/ijerph14060636












