Infection Dynamics and Antimicrobial Resistance Profile of Salmonella Paratyphi B d-tartrate Positive (Java) in a Persistently Infected Broiler Barn
Abstract
:1. Introduction
2. Materials and Methods
2.1. Broiler Flocks and Farm Management
2.2. Salmonella DNA Detection after Cleaning and Disinfection
2.3. Salmonella Detection during Rearing Cycles
2.4. Quantitative PCR (qPCR) Set-up and Amplification
2.5. Biochemical and Serological Identification of Salmonella
2.6. Antimicrobial Susceptibility Testing of S. Java
2.7. Statistical Analysis
3. Results
3.1. Salmonella DNA Detection after Cleaning and Disinfection
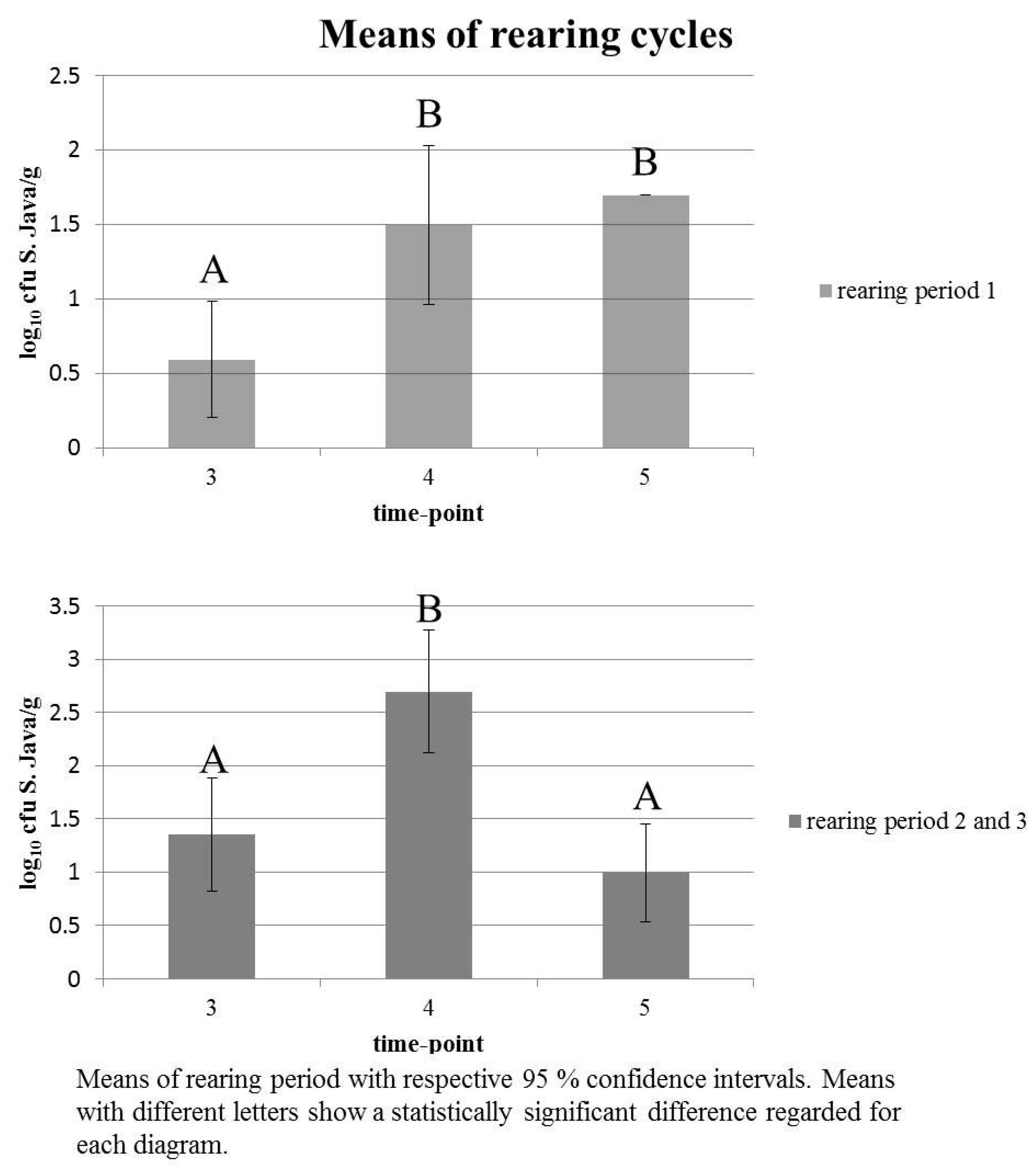
3.2. Salmonella Detection during Rearing Cycles
3.3. Biochemical and Serological Identification of Salmonella
3.4. Antimicrobial Susceptibility Testing of S. Java
4. Discussion
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Andino, A.; Hanning, I. Salmonella enterica: Survival, colonization, and virulence differences among serovars. Sci. World J. 2015, 2015, 520179. [Google Scholar] [CrossRef] [PubMed]
- Callaway, T.R.; Edrington, T.S.; Anderson, R.C.; Byrd, J.A.; Nisbet, D.J. Gastrointestinal microbial ecology and the safety of our food supply as related to Salmonella. J. Anim. Sci. 2008, 86, E163–E172. [Google Scholar] [CrossRef] [PubMed]
- Bolder, N.M. Salmonella Control in Broiler Flocks; International Society for Animal Hygiene: Saint-Malo, France, 2004; Available online: http://www.isah-soc.org/documents/2004/Bolder.pdf (accessed on 18 January 2015).
- European Food Safety Authority and European Center for Disease Control. The European Union Summary Report on Trends and Sources of Zoonoses, Zoonotic Agents and Food-borne Outbreaks in 2010. EFSA J. 2012, 10, 2597. [Google Scholar] [CrossRef]
- Al-Nakhli, H.M.; Al-Ogaily, Z.H.; Nassar, T.J. Representative Salmonella serovars isolated from poultry and poultry environments in Saudi Arabia. Rev. Sci. Tech. 1999, 18, 700–709. [Google Scholar] [CrossRef] [PubMed]
- Barua, H.; Biswas, P.K.; Talukder, K.A.; Olsen, K.E.P.; Christensen, J.P. Poultry as a possible source of non-typhoidal Salmonella enterica serovars in humans in Bangladesh. Vet. Microbiol. 2014, 168, 372–380. [Google Scholar] [CrossRef] [PubMed]
- Wollin, R. A study of invasiveness of different Salmonella serovars based on analysis of the Enter-net database. Euro Surveill. 2007, 12, E070927-3. [Google Scholar] [PubMed]
- European Food Safety Authority and European Center for Disease Control. The European Union summary report on trends and sources of zoonoses, zoonotic agents and food-borne outbreaks in 2014 European Food Safety Authority European Centre for Disease Prevention and Control. EFSA J. 2015, 13, 4329. [Google Scholar]
- Desenclos, J.C.; Bouvet, P.; Benz-Lemoine, E.; Grimont, F.; Desqueyroux, H.; Rebière, I.; Grimont, P.A. Large outbreak of Salmonella enterica serotype paratyphi B infection caused by a goats’ milk cheese, France, 1993: A case finding and epidemiological study. BMJ 1996, 312, 91–94. [Google Scholar] [CrossRef] [PubMed]
- Gaulin, C.; Vincent, C.; Alain, L.; Ismaïl, J. Outbreak of Salmonella paratyphi B linked to aquariums in the province of Quebec, 2000. Can. Commun. Dis. Rep. 2002, 28, 89–93. [Google Scholar] [PubMed]
- Stratton, J.; Stefaniw, L.; Grimsrud, K.; Werker, D.H.; Ellis, A.; Ashton, E.; Chui, L.; Blewett, E.; Ahmed, R.; Clark, C.; et al. Outbreak of Salmonella paratyphi B var java due to contaminated alfalfa sprouts in Alberta, British Columbia and Saskatchewan. Can. Commun. Dis. Rep. 2001, 27, 133–137. [Google Scholar] [PubMed]
- Toboldt, A.; Tietze, E.; Helmuth, R.; Fruth, A.; Junker, E.; Malorny, B. Human infections attributable to the D-tartrate-fermenting variant of Salmonella enterica serovar Paratyphi B in Germany originate in reptiles and, on rare occasions, poultry. Appl. Environ. Microbiol. 2012, 78, 7347–7357. [Google Scholar] [CrossRef] [PubMed]
- Van Pelt, W.; Van der Zee, H.; Wannet, W.J.B.; Van de Giessen, A.W.; Mevius, D.J.; Bolder, N.M.; Komijn, R.E.; Van Duynhoven, Y.T.H.P. Explosive increase of Salmonella Java in poultry in the Netherlands: Consequences for public health. Euro Surveill. 2003, 8, 31–35. [Google Scholar] [PubMed]
- Brown, D.J.; Mather, H.; Browning, L.M.; Coia, J.E. Investigation of human infections with Salmonella enterica serovar Java in Scotland and possible association with imported poultry. Euro Surveill. 2003, 8, 35–40. [Google Scholar] [PubMed]
- FAO; WHO; OIE. Joint FAO/WHO/OIE Expert Meeting on Critically Important Antimicrobials. 2008. Available online: http://www.fao.org/3/a-i0204e.pdf (accessed on 14 February 2016).
- Miko, A.; Pries, K.; Schroeter, A.; Helmuth, R. Multiple-drug resistance in D-tartrate-positive Salmonella enterica serovar paratyphi B isolates from poultry is mediated by class 2 integrons inserted into the bacterial chromosome. Antimicrob. Agents Chemother. 2003, 47, 3640–3643. [Google Scholar] [CrossRef] [PubMed]
- Doublet, B.; Praud, K.; Nguyen-Ho-Bao, T.; Argudín, M.A.; Bertrand, S.; Butaye, P.; Cloeckaert, A. Extended-spectrum β-lactamase- and AmpC β-lactamase-producing D-tartrate-positive Salmonella enterica serovar Paratyphi B from broilers and human patients in Belgium, 2008–10. J. Antimicrob. Chemother. 2014, 69, 1257–1264. [Google Scholar] [CrossRef] [PubMed]
- Namata, H.; Welby, S.; Aerts, M.; Faes, C.; Abrahantes, J.C.; Imberechts, H.; Vermeersch, K.; Hooyberghs, J.; Méroc, E.; Mintiens, K. Identification of risk factors for the prevalence and persistence of Salmonella in Belgian broiler chicken flocks. Prev. Vet. Med. 2009, 90, 211–222. [Google Scholar] [CrossRef] [PubMed]
- Van Immerseel, F.; Meulemans, L.; De Buck, J.; Pasmans, F.; Velge, P.; Bottreau, E.; Haesebrouck, F.; Ducatelle, R. Bacteria-host interactions of Salmonella B d+ Tartrate in poultry. Epidemiol. Infect. 2004, 132, 239–243. [Google Scholar] [CrossRef] [PubMed]
- Kloska, F.; Casteel, M.; Wilms-Schulze Kump, F.; Klein, G. Implementation of a risk-orientated hygiene analysis for the control of S almonella Java in broiler production. Curr. Microbiol. 2017, in press. [Google Scholar]
- Bundesministerium für Justiz und für Verbraucherschutz Tierschutzgesetz in der Fassung der Bekanntmachung vom 18. Mai 2006 (BGBl. I S. 1206, 1313), das zuletzt durch Artikel 4 Absatz 87 des Gesetzes vom 18. Juli 2016 (BGBl. I S. 1666) geändert worden ist. Available online: https://www.gesetze-im-internet.de/tierschg/BJNR012770972.html (accessed on 30 December 2016).
- Malorny, B.; Paccassoni, E.; Fach, P.; Bunge, C.; Martin, A.; Helmuth, R. Diagnostic real-time PCR for detection of Salmonella in food. Appl. Environ. Microbiol. 2004, 70, 7046–7052. [Google Scholar] [CrossRef] [PubMed]
- Deutsches Institut Für Normung DIN 58940: Medical Microbiology—Susceptibility Testing of Pathogens to Antimicrobial Agents; Beuth Verlag: Berlin, Germany, 2007.
- Anonymous. Antimicrobial Wild Type Distributions of Microorganisms. Available online: http://mic.eucast.org/Eucast2/SearchController/search.jsp?action=init (accessed on 12 March 2016).
- Hansson, I.; Pudas, N.; Harbom, B.; Engvall, E.O. Within-flock variations of Campylobacter loads in caeca and on carcasses from broilers. Int. J. Food Microbiol. 2010, 141, 51–55. [Google Scholar] [CrossRef] [PubMed]
- European Food Safety Authority. Opinion of the Scientific Panel on Biological Hazards on the request from the Commission related to the use of antimicrobials for the control of Salmonella in poultry. EFSA J. 2004, 115, 1–76. [Google Scholar]
- Bronzwaer, S. Harmonised monitoring of antimicrobial resistance in Salmonella and Campylobacter isolates from food animals in the European Union. Clin. Microbiol. Infect. 2008, 14, 522–533. [Google Scholar] [CrossRef] [PubMed]
- Miko, A.; Guerra, B.; Schroeter, A.; Dorn, C.; Helmuth, R. Molecular characterization of multiresistant d-tartrate-positive Salmonella enterica serovar paratyphi B isolates. J. Clin. Microbiol. 2002, 40, 3184–3191. [Google Scholar] [CrossRef] [PubMed]
- Hasman, H.; Mevius, D.; Veldman, K.; Olesen, I.; Aarestrup, F.M. Beta-Lactamases among extended-spectrum beta-lactamase (ESBL)-resistant Salmonella from poultry, poultry products and human patients in The Netherlands. J. Antimicrob. Chemother. 2005, 56, 115–121. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez, I.; Barownick, W.; Helmuth, R.; Mendoza, M.C.; Rodicio, M.R.; Schroeter, A.; Guerra, B. Extended-spectrum {beta}-lactamases and AmpC {beta}-lactamases in ceftiofur-resistant Salmonella enterica isolates from food and livestock obtained in Germany during 2003–07. J. Antimicrob. Chemother. 2009, 64, 301–309. [Google Scholar] [CrossRef] [PubMed]
- Angulo, F.J.; Johnson, K.R.; Tauxe, R.V.; Cohen, M.L. Origins and consequences of antimicrobial-resistant nontyphoidal Salmonella: Implications for the use of fluoroquinolones in food animals. Microb. Drug Resist. 2000, 6, 77–83. [Google Scholar] [CrossRef] [PubMed]
- Schroeter, A.; Hoog, B.; Helmuth, R. Resistance of Salmonella isolates in Germany. J. Vet. Med. B Infect. Dis. Vet. Public Health 2004, 51, 389–392. [Google Scholar] [CrossRef] [PubMed]
- Weill, F.-X.; Fabre, L.; Grandry, B.; Grimont, P.A.D.; Casin, I. Multiple-antibiotic resistance in Salmonella enterica serotype Paratyphi B isolates collected in France between 2000 and 2003 is due mainly to strains harboring Salmonella genomic islands 1, 1-B, and 1-C. Antimicrob. Agents Chemother. 2005, 49, 2793–2801. [Google Scholar] [CrossRef] [PubMed]
- Huehn, S.; Helmuth, R.; Bunge, C.; Guerra, B.; Junker, E.; Davies, R.H.; Wattiau, P.; Van Pelt, W.; Malorny, B. Characterization of pathogenic and resistant genome repertoire reveals two clonal lines in Salmonella enterica subsp. enterica serovar Paratyphi B (+)-tartrate positive. Foodborne Pathog. Dis. 2009, 6, 431–443. [Google Scholar] [CrossRef] [PubMed]
- Boumart, Z.; Roche, S.M.; Lalande, F.; Virlogeux-Payant, I.; Hennequet-Antier, C.; Menanteau, P.; Gabriel, I.; Weill, F.-X.; Velge, P.; Chemaly, M. Heterogeneity of persistence of Salmonella enterica serotype Senftenberg strains could explain the emergence of this serotype in poultry flocks. PLoS ONE 2012, 7, e35782. [Google Scholar] [CrossRef] [PubMed]
- Allen, C.A.; Fedorka-Cray, P.J.; Vazquez-Torres, A.; Suyemoto, M.; Altier, C.; Ryder, L.R.; Fang, F.C.; Libby, S.J. In vitro and in vivo assessment of Salmonella enterica serovar Typhimurium DT104 virulence. Infect. Immun. 2001, 69, 4673–4677. [Google Scholar] [CrossRef] [PubMed]
- Foley, S.L.; Johnson, T.J.; Ricke, S.C.; Nayak, R.; Danzeisen, J. Salmonella pathogenicity and host adaptation in chicken-associated serovars. Microbiol. Mol. Biol. Rev. 2013, 7, 582–607. [Google Scholar] [CrossRef] [PubMed]
- Bailey, J.S. Integrated colonization control of Salmonella in poultry. Poult. Sci. 1988, 67, 928–932. [Google Scholar] [CrossRef] [PubMed]
| Type of Condition | Field Survey 1 | Field Survey 2 | Field Survey 3 |
|---|---|---|---|
| flock information | |||
| flock size | 33,000 | 33,000 | 33,000 |
| breed | Ross 308 | Ross 308 | Ross 308 |
| season | June–August | August–September | October–November |
| biosecurity | |||
| rubber boots | changed | Changed | changed |
| clothes | not changed | not changed | not changed |
| prethinning | yes | Yes | yes |
| vaccination (first 15 days of life) | Gumboro disease, Newcastle disease, Infectious bronchitis | Gumboro disease, Newcastle disease, Infectious bronchitis | Gumboro disease, Newcastle disease, Infectious bronchitis |
| feed additives | |||
| first 18 days of life | vitamins A, D3, E 1 | vitamins A, D3, E 1 | vitamins A, D3, E 1 |
| whole rearing period | buffered acids * | buffered acids * | buffered acids * |
| antibiotic therapy | amoxicillin/colistin on days 33–35 | lincomycin-spectinomycin on days 1–4 | lincomycin-spectinomycin on days 1–4 |
| Sampling Point | qPCR Result |
|---|---|
| water lines | negative |
| feed lines | negative |
| feed suppliers | negative |
| Wall | positive |
| supply air system | positive |
| exhaust air system | positive |
| barn gateway | positive |
| heating system | negative |
| windows and window frames | negative |
| Ceiling | negative |
| cable duct at the ceiling | positive |
| ligament suspension of lines | negative |
| floor, joints, and crannies | positive |
| anteroom | positive |
| housing forecourt | negative |
| Time Point | Statistic | ||
|---|---|---|---|
| 3 | 4 | 5 | |
| 1.349 | 2.694 | 0.990 | mean |
| 0 | 0 | 0 | min |
| 5.322 | 5.602 | 5.204 | max |
| 0.818; 1.880 | 2.118; 3.270 | 0.528; 1.451 | 95% CI |
| A * | B * | A * | LSD test |
| Time Point | Statistic | ||
|---|---|---|---|
| 3 | 4 | 5 1 | |
| 0.594 | 1.497 | 1.698 | mean |
| 0 | 0 | 1.698 | min |
| 1.698 | 4.954 | 1.698 | max |
| 0.205; 0.983 | 0.962; 2.031 | - | 95% CI |
| A * | B * | B * | LSD test |
| Time-Point | Positive | Negative | p-Value |
| 3 | 34 | 26 | |
| 4 | 12 | 48 | |
| 5 | 24 | 36 | |
| Time-Point | Odds Ratio | 95% CI | |
| 4 vs. 3 | 5.231 | 2.320; 11.793 | <0.0001 |
| 5 vs. 3 | 1.962 | 0.949; 4.055 | >0.05 |
| 5 vs. 4 | 0.375 | 0.166; 0.849 | <0.05 |
© 2017 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kloska, F.; Beyerbach, M.; Klein, G. Infection Dynamics and Antimicrobial Resistance Profile of Salmonella Paratyphi B d-tartrate Positive (Java) in a Persistently Infected Broiler Barn. Int. J. Environ. Res. Public Health 2017, 14, 101. https://doi.org/10.3390/ijerph14010101
Kloska F, Beyerbach M, Klein G. Infection Dynamics and Antimicrobial Resistance Profile of Salmonella Paratyphi B d-tartrate Positive (Java) in a Persistently Infected Broiler Barn. International Journal of Environmental Research and Public Health. 2017; 14(1):101. https://doi.org/10.3390/ijerph14010101
Chicago/Turabian StyleKloska, Franziska, Martin Beyerbach, and Günter Klein. 2017. "Infection Dynamics and Antimicrobial Resistance Profile of Salmonella Paratyphi B d-tartrate Positive (Java) in a Persistently Infected Broiler Barn" International Journal of Environmental Research and Public Health 14, no. 1: 101. https://doi.org/10.3390/ijerph14010101
APA StyleKloska, F., Beyerbach, M., & Klein, G. (2017). Infection Dynamics and Antimicrobial Resistance Profile of Salmonella Paratyphi B d-tartrate Positive (Java) in a Persistently Infected Broiler Barn. International Journal of Environmental Research and Public Health, 14(1), 101. https://doi.org/10.3390/ijerph14010101





