Diversity of Bacterial Communities on Four Frequently Used Surfaces in a Large Brazilian Teaching Hospital
Abstract
:1. Introduction
2. Experimental Section
3. Results and Discussion
| Sample ID | Valid Reads | Number of OTU (>97% Identity) | Shannon Index | Goods Library Coverage |
|---|---|---|---|---|
| HC-EB | 195,814 | 78,236 | 9.065327 | 0.645633 |
| HC-EBTCS | 791,476 | 315,177 | 9.109842 | 0.61603 |
| HC-BMKB | 548,605 | 238,544 | 9.801774 | 0.594236 |
| Hc_Restrooms | 708,726 | 288,265 | 9.782897 | 0.615365 |
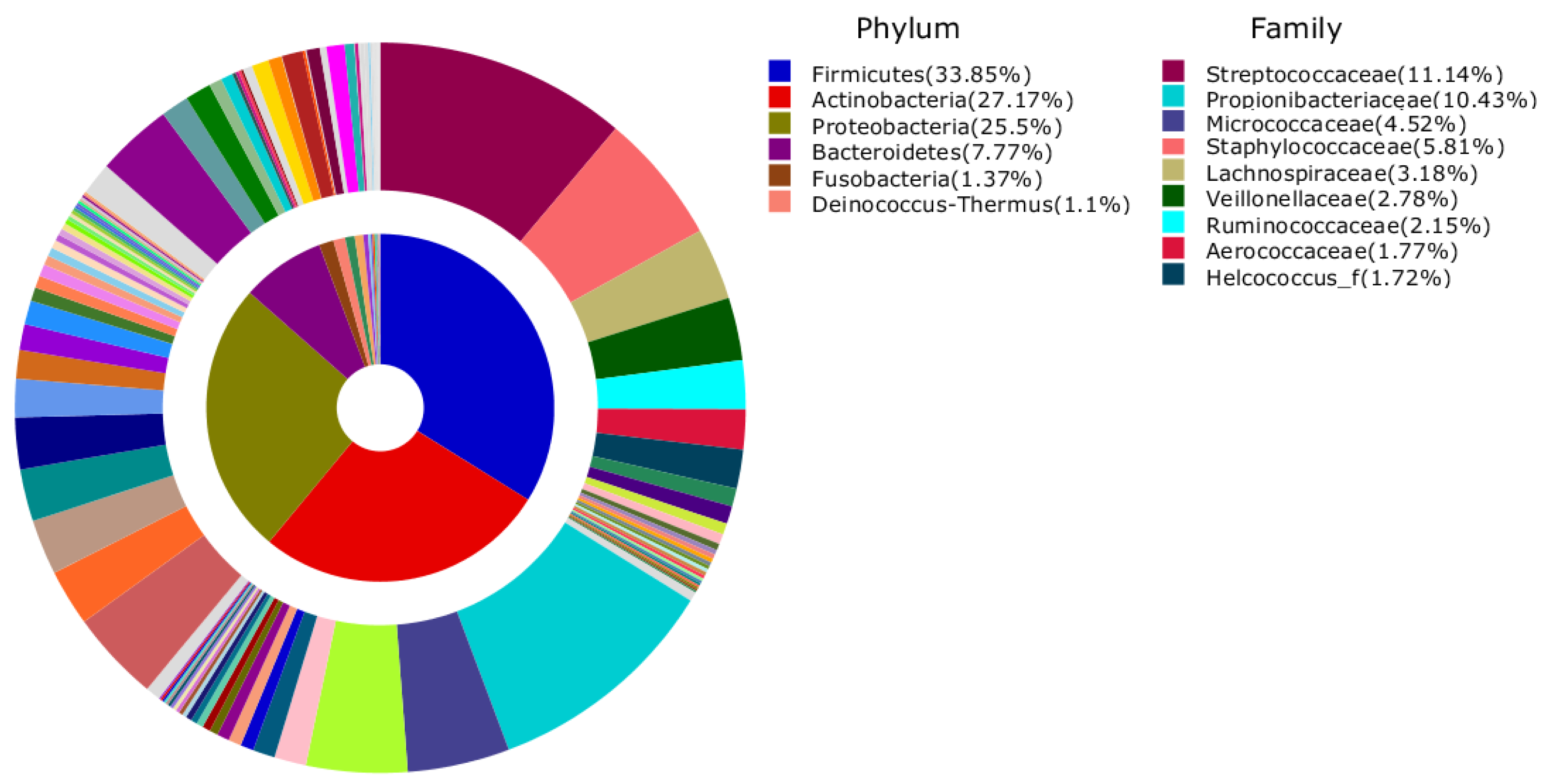
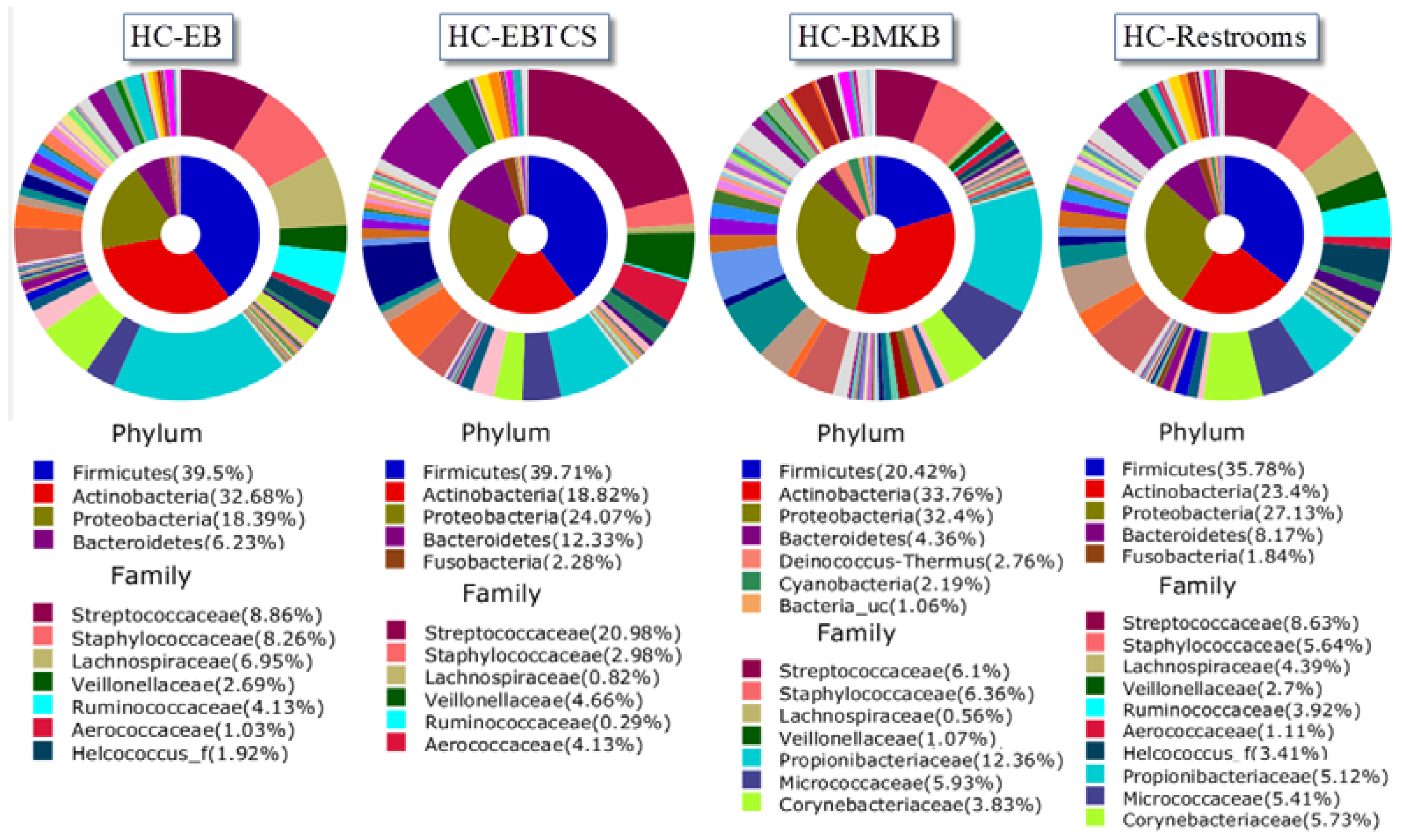
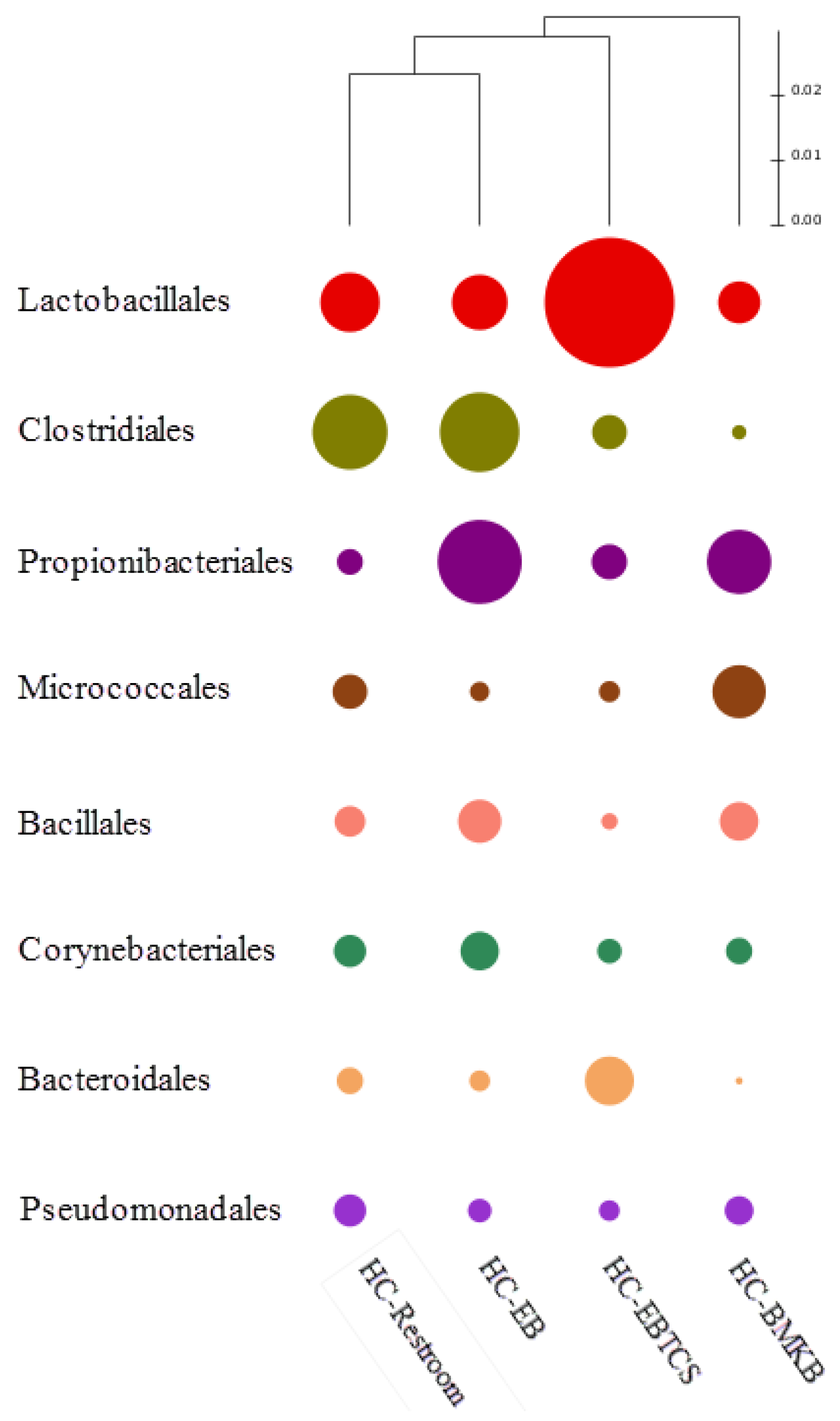
| HC-EB | HC-BMKB | HC-Restroom | HC-EBTCS | |
|---|---|---|---|---|
| Abundance Order | Taxa (Abundance) | |||
| 1 | Propionibacterium acnes (11.8%) | Propionibacterium acnes (7.46%) | Propionibacterium acnes (2.23%) | Streptococcus dentisani (8.45%) |
| 2 | Streptococcus dentisani (2.8%) | Propionibacteriaceae_uc_s (2.18%) | Rothia mucilaginosa (1.76%) | Propionibacterium acnes (4.72%) |
| 3 | Staphylococcus epidermidis (2.54%) | Streptococcus dentisani (1.85%) | Streptococcaceae_uc_s (1.57%) | Streptococcaceae_uc_s (3.84%) |
| 4 | Propionibacteriaceae_uc_s (2.46%) | Propionibacterium_uc (1.56%) | Streptococcus dentisani (1.54%) | Streptococcus_uc (3.08%) |
| 5 | Propionibacterium_uc (2.41%) | Staphylococcus epidermidis (1.31%) | HQ762034_s (1.34%) | Streptococcus salivarius (2.2%) |
| 6 | EF188441_s (1.54%) | Streptococcaceae_uc_s (1.26%) | Staphylococcus epidermidis (1.33%) | Rothia mucilaginosa (1.67%) |
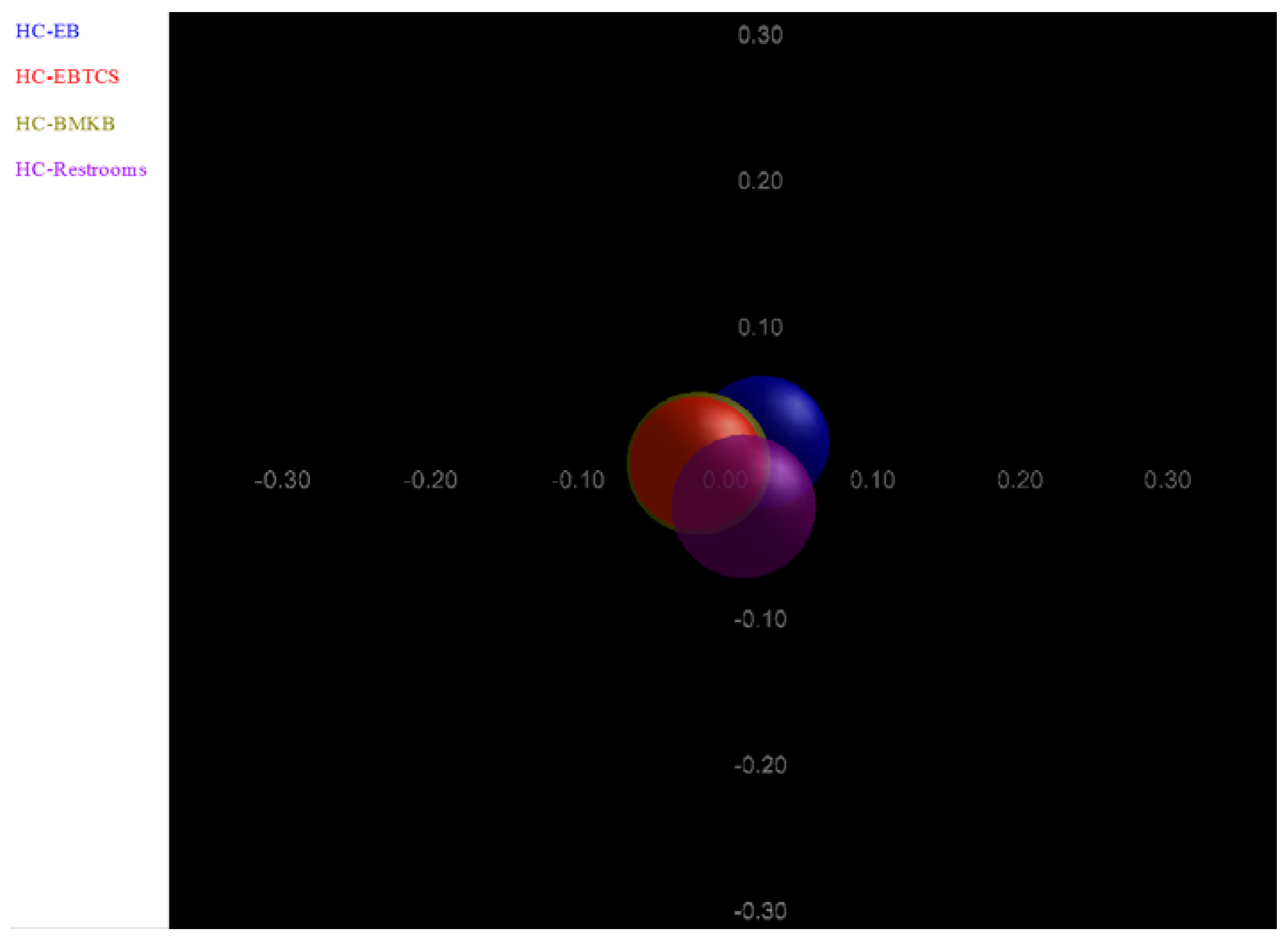
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Gravel, D.; Taylor, G.; Ofner, M.; Johnston, L.; Loeb, M.; Roth, V.R.; Stegenga, J.; Bryce, E.; Matlow, A. Point prevalence survey for healthcare-associated infections within canadian adult acute-care hospitals. J. Hosp. Infect. 2007, 66, 243–248. [Google Scholar] [CrossRef] [PubMed]
- Dancer, S.J. Importance of the environment in meticillin-resistant Staphylococcus aureus acquisition: The case for hospital cleaning. Lancet Infect. Dis. 2008, 8, 101–113. [Google Scholar] [CrossRef]
- Klevens, R.M.; Edwards, J.R.; Richards, C.L., Jr.; Horan, T.C.; Gaynes, R.P.; Pollock, D.A.; Cardo, D.M. Estimating health care-associated infections and deaths in U.S. Hospitals, 2002. Public Health Rep. 2007, 122, 160–166. [Google Scholar] [PubMed]
- Burke, J.P. Infection control—A problem for patient safety. N. Engl. J. Med. 2003, 348, 651–656. [Google Scholar] [CrossRef] [PubMed]
- Stone, P.W.; Braccia, D.; Larson, E. Systematic review of economic analyses of health care-associated infections. Am. J. Infect. Control 2005, 33, 501–509. [Google Scholar] [CrossRef] [PubMed]
- Sydnor, E.R.; Perl, T.M. Hospital epidemiology and infection control in acute-care settings. Clin. Microbiol. Rev. 2011, 24, 141–173. [Google Scholar] [CrossRef] [PubMed]
- Quach, C.; McArthur, M.; McGeer, A.; Li, L.; Simor, A.; Dionne, M.; Levesque, E.; Tremblay, L. Risk of infection following a visit to the emergency department: A cohort study. CMAJ 2012, 184, E232–E239. [Google Scholar] [CrossRef] [PubMed]
- Velasco, E.; Thuler, L.C.; Martins, C.A.; Dias, L.M.; Goncalves, V.M. Nosocomial infections in an oncology intensive care unit. Am. J. Infect. Control 1997, 25, 458–462. [Google Scholar] [CrossRef]
- Couto, R.C.; Carvalho, E.A.; Pedrosa, T.M.; Pedroso, E.R.; Neto, M.C.; Biscione, F.M. A 10-year prospective surveillance of nosocomial infections in neonatal intensive care units. Am. J. Infect. Control 2007, 35, 183–189. [Google Scholar] [CrossRef] [PubMed]
- Pessoa-Silva, C.L.; Richtmann, R.; Calil, R.; Santos, R.M.; Costa, M.L.; Frota, A.C.; Wey, S.B. Healthcare-associated infections among neonates in brazil. Infect. Control Hosp. Epidemiol. 2004, 25, 772–777. [Google Scholar] [CrossRef] [PubMed]
- Kawagoe, J.Y.; Segre, C.A.; Pereira, C.R.; Cardoso, M.F.; Silva, C.V.; Fukushima, J.T. Risk factors for nosocomial infections in critically ill newborns: A 5-year prospective cohort study. Am. J. Infect. Control 2001, 29, 109–114. [Google Scholar] [CrossRef] [PubMed]
- Nagata, E.; Brito, A.S.; Matsuo, T. Nosocomial infections in a neonatal intensive care unit: Incidence and risk factors. Am. J. Infect. Control 2002, 30, 26–31. [Google Scholar] [CrossRef] [PubMed]
- Treakle, A.M.; Thom, K.A.; Furuno, J.P.; Strauss, S.M.; Harris, A.D.; Perencevich, E.N. Bacterial contamination of health care workers’ white coats. Am. J. Infect. Control 2009, 37, 101–105. [Google Scholar] [CrossRef] [PubMed]
- Tang, P.H.; Worster, A.; Srigley, J.A.; Main, C.L. Examination of staphylococcal stethoscope contamination in the emergency department (pilot) study (EXSSCITED pilot study). CJEM 2011, 13, 239–244. [Google Scholar] [PubMed]
- Redelmeier, D.A.; Livesley, N.J. Adhesive tape and intravascular-catheter-associated infections. J. Gen. Intern. Med. 1999, 14, 373–375. [Google Scholar] [CrossRef] [PubMed]
- Schultz, M.; Gill, J.; Zubairi, S.; Huber, R.; Gordin, F. Bacterial contamination of computer keyboards in a teaching hospital. Infect. Control Hosp. Epidemiol. 2003, 24, 302–303. [Google Scholar] [CrossRef] [PubMed]
- Kandel, C.E.; Simor, A.E.; Redelmeier, D.A. Elevator buttons as unrecognized sources of bacterial colonization in hospitals. Open Med. 2014, 8, e81–e86. [Google Scholar] [PubMed]
- Brady, R.R.; Verran, J.; Damani, N.N.; Gibb, A.P. Review of mobile communication devices as potential reservoirs of nosocomial pathogens. J. Hosp. Infect. 2009, 71, 295–300. [Google Scholar] [CrossRef] [PubMed]
- Mullaney, P.J.; Munthali, P.; Vlachou, P.; Jenkins, D.; Rathod, A.; Entwisle, J. How clean is your probe? Microbiological assessment of ultrasound transducers in routine clinical use, and cost-effective ways to reduce contamination. Clin. Radiol. 2007, 62, 694–698. [Google Scholar] [CrossRef] [PubMed]
- Kramer, A.; Schwebke, I.; Kampf, G. How long do nosocomial pathogens persist on inanimate surfaces? A systematic review. BMC Infect. Dis. 2006, 6. [Google Scholar] [CrossRef] [PubMed]
- Kei, J.; Richards, J.R. The prevalence of methicillin-resistant Staphylococcus aureus on inanimate objects in an urban emergency department. J. Emerg. Med. 2011, 41, 124–127. [Google Scholar] [CrossRef] [PubMed]
- Flores, G.E.; Bates, S.T.; Caporaso, J.G.; Lauber, C.L.; Leff, J.W.; Knight, R.; Fierer, N. Diversity, distribution and sources of bacteria in residential kitchens. Environ. Microbiol. 2013, 15, 588–596. [Google Scholar] [CrossRef] [PubMed]
- Flores, G.E.; Bates, S.T.; Knights, D.; Lauber, C.L.; Stombaugh, J.; Knight, R.; Fierer, N. Microbial biogeography of public restroom surfaces. PLoS ONE 2011, 6, e28132. [Google Scholar] [CrossRef] [PubMed]
- Rintala, H.; Pitkaranta, M.; Toivola, M.; Paulin, L.; Nevalainen, A. Diversity and seasonal dynamics of bacterial community in indoor environment. BMC Microbiol. 2008, 8. [Google Scholar] [CrossRef] [PubMed]
- Paulino, L.C.; Tseng, C.H.; Strober, B.E.; Blaser, M.J. Molecular analysis of fungal microbiota in samples from healthy human skin and psoriatic lesions. J. Clin. Microbiol. 2006, 44, 2933–2941. [Google Scholar] [CrossRef] [PubMed]
- Herlemann, D.P.; Labrenz, M.; Jurgens, K.; Bertilsson, S.; Waniek, J.J.; Andersson, A.F. Transitions in bacterial communities along the 2000 km salinity gradient of the baltic sea. ISME J. 2011, 5, 1571–1579. [Google Scholar] [CrossRef] [PubMed]
- Da Pereira Fonseca, T.A.; Pessoa, R.; Sanabani, S.S. Molecular analysis of bacterial microbiota on brazilian currency note surfaces. Int. J. Environ. Res. Public Health 2015, 12, 13276–13288. [Google Scholar] [CrossRef] [PubMed]
- Masella, A.P.; Bartram, A.K.; Truszkowski, J.M.; Brown, D.G.; Neufeld, J.D. Pandaseq: Paired-end assembler for illumina sequences. BMC Bioinform. 2012, 13. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C.; Haas, B.J.; Clemente, J.C.; Quince, C.; Knight, R. Uchime improves sensitivity and speed of chimera detection. Bioinformatics 2011, 27, 2194–2200. [Google Scholar] [CrossRef] [PubMed]
- Kim, O.S.; Cho, Y.J.; Lee, K.; Yoon, S.H.; Kim, M.; Na, H.; Park, S.C.; Jeon, Y.S.; Lee, J.H.; Yi, H.; et al. Introducing EzTaxon-e: A prokaryotic 16s rRNA gene sequence database with phylotypes that represent uncultured species. Int. J. Syst. Evol. Microbiol. 2012, 62, 716–721. [Google Scholar] [CrossRef] [PubMed]
- Hamady, M.; Lozupone, C.; Knight, R. Fast unifrac: Facilitating high-throughput phylogenetic analyses of microbial communities including analysis of pyrosequencing and phylochip data. ISME J. 2010, 4, 17–27. [Google Scholar] [CrossRef] [PubMed]
- Tairacan, A.P.F.; Sabri, S.S. Diversity of Bacterial Communities on Four Frequently Used Surfaces in a Large Brazilian Teaching Hospital. Available online: http://dx.doi.org/10.5281/zenodo.35584 (accessed on 21 December 2015).
- Costello, E.K.; Lauber, C.L.; Hamady, M.; Fierer, N.; Gordon, J.I.; Knight, R. Bacterial community variation in human body habitats across space and time. Science 2009, 326, 1694–1697. [Google Scholar] [CrossRef] [PubMed]
- Fierer, N.; Hamady, M.; Lauber, C.L.; Knight, R. The influence of sex, handedness, and washing on the diversity of hand surface bacteria. Proc. Natl. Acad. Sci. USA 2008, 105, 17994–17999. [Google Scholar] [CrossRef] [PubMed]
- Grice, E.A.; Kong, H.H.; Conlan, S.; Deming, C.B.; Davis, J.; Young, A.C.; Bouffard, G.G.; Blakesley, R.W.; Murray, P.R.; Green, E.D.; et al. Topographical and temporal diversity of the human skin microbiome. Science 2009, 324, 1190–1192. [Google Scholar] [CrossRef] [PubMed]
- Reed, D.; Kemmerly, S.A. Infection control and prevention: A review of hospital-acquired infections and the economic implications. Ochsner J. 2009, 9, 27–31. [Google Scholar] [PubMed]
- Anderson, R.N.; Smith, B.L. Deaths: Leading causes for 2002. Natl. Vital. Stat. Rep. 2005, 53, 1–89. [Google Scholar] [PubMed]
- Gao, Z.; Tseng, C.H.; Pei, Z.; Blaser, M.J. Molecular analysis of human forearm superficial skin bacterial biota. Proc. Natl. Acad. Sci. USA 2007, 104, 2927–2932. [Google Scholar] [CrossRef] [PubMed]
- Lee, L.; Tin, S.; Kelley, S.T. Culture-independent analysis of bacterial diversity in a child-care facility. BMC Microbiol. 2007, 7. [Google Scholar] [CrossRef] [PubMed]
- McManus, C.J.; Kelley, S.T. Molecular survey of aeroplane bacterial contamination. J. Appl. Microbiol. 2005, 99, 502–508. [Google Scholar] [CrossRef] [PubMed]
- Claesson, M.J.; O’Sullivan, O.; Wang, Q.; Nikkila, J.; Marchesi, J.R.; Smidt, H.; de Vos, W.M.; Ross, R.P.; O’Toole, P.W. Comparative analysis of pyrosequencing and a phylogenetic microarray for exploring microbial community structures in the human distal intestine. PLoS ONE 2009, 4, e6669. [Google Scholar] [CrossRef] [PubMed]
- Ley, R.E.; Turnbaugh, P.J.; Klein, S.; Gordon, J.I. Microbial ecology: Human gut microbes associated with obesity. Nature 2006, 444, 1022–1023. [Google Scholar] [CrossRef] [PubMed]
- Mukherjee, N.; Dowd, S.E.; Wise, A.; Kedia, S.; Vohra, V.; Banerjee, P. Diversity of bacterial communities of fitness center surfaces in a U.S. Metropolitan area. Int. J. Environ. Res. Public Health 2014, 11, 12544–12561. [Google Scholar] [CrossRef] [PubMed]
- Spellerberg, B.; Brandt, C. Manual of Clinical Microbiology, 10th ed.; ASM Press: Washington, DC, USA, 2011; pp. 331–349. [Google Scholar]
- Kloos, W.E.; Musselwhite, M.S. Distribution and persistence of Staphylococcus and micrococcus species and other aerobic bacteria on human skin. Appl. Microbiol. 1975, 30, 381–385. [Google Scholar] [PubMed]
- National Nosocomial Infections Surveillance System. National nosocomial infections surveillance (NNIS) system report, data summary from January 1992 through June 2004, issued October 2004. Am. J. Infect. Control 2004, 32, 470–485.
- Rogers, K.L.; Fey, P.D.; Rupp, M.E. Coagulase-negative staphylococcal infections. Infect. Dis. Clin. North Am. 2009, 23, 73–98. [Google Scholar] [CrossRef] [PubMed]
- Otto, M. Staphylococcus epidermidis—The “accidental” pathogen. Nat. Rev. Microbiol. 2009, 7, 555–567. [Google Scholar] [CrossRef] [PubMed]
- Mack, D.; Fischer, W.; Krokotsch, A.; Leopold, K.; Hartmann, R.; Egge, H.; Laufs, R. The intercellular adhesin involved in biofilm accumulation of Staphylococcus epidermidis is a linear beta-1,6-linked glucosaminoglycan: Purification and structural analysis. J. Bacteriol. 1996, 178, 175–183. [Google Scholar] [PubMed]
- Lowy, F.D. Staphylococcus aureus infections. N. Engl. J. Med. 1998, 339, 520–532. [Google Scholar] [CrossRef] [PubMed]
- Otto, M. Mrsa virulence and spread. Cell Microbiol. 2012, 14, 1513–1521. [Google Scholar] [CrossRef] [PubMed]
- Sanford, M.D.; Widmer, A.F.; Bale, M.J.; Jones, R.N.; Wenzel, R.P. Efficient detection and long-term persistence of the carriage of methicillin-resistant Staphylococcus aureus. Clin. Infect. Dis. 1994, 19, 1123–1128. [Google Scholar] [CrossRef] [PubMed]
- Miller, L.G.; Diep, B.A. Clinical practice: Colonization, fomites, and virulence: Rethinking the pathogenesis of community-associated methicillin-resistant Staphylococcus aureus infection. Clin. Infect. Dis. 2008, 46, 752–760. [Google Scholar] [CrossRef] [PubMed]
- David, M.Z.; Mennella, C.; Mansour, M.; Boyle-Vavra, S.; Daum, R.S. Predominance of methicillin-resistant Staphylococcus aureus among pathogens causing skin and soft tissue infections in a large urban jail: Risk factors and recurrence rates. J. Clin. Microbiol. 2008, 46, 3222–3227. [Google Scholar] [CrossRef] [PubMed]
- Rackham, D.M.; Ray, S.M.; Franks, A.S.; Bielak, K.M.; Pinn, T.M. Community-associated methicillin-resistant Staphylococcus aureus nasal carriage in a college student athlete population. Clin. J. Sport Med. 2010, 20, 185–188. [Google Scholar] [CrossRef] [PubMed]
- Montgomery, K.; Ryan, T.J.; Krause, A.; Starkey, C. Assessment of athletic health care facility surfaces for MRSA in the secondary school setting. J. Environ. Health 2010, 72, 8–11. [Google Scholar] [PubMed]
- Stanforth, B.; Krause, A.; Starkey, C.; Ryan, T.J. Prevalence of community-associated methicillin-resistant Staphylococcus aureus in high school wrestling environments. J. Environ. Health 2010, 72, 12–16. [Google Scholar] [PubMed]
- Gebel, J.; Exner, M.; French, G.; Chartier, Y.; Christiansen, B.; Gemein, S.; Goroncy-Bermes, P.; Hartemann, P.; Heudorf, U.; Kramer, A.; et al. The role of surface disinfection in infection prevention. GMS Hyg. Infect. Control 2013, 8. [Google Scholar] [CrossRef]
- Weber, D.J.; Rutala, W.A. Self-disinfecting surfaces: Review of current methodologies and future prospects. Am. J. Infect. Control 2013, 41, S31–S35. [Google Scholar] [CrossRef] [PubMed]
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons by Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pereira da Fonseca, T.A.; Pessôa, R.; Felix, A.C.; Sanabani, S.S. Diversity of Bacterial Communities on Four Frequently Used Surfaces in a Large Brazilian Teaching Hospital. Int. J. Environ. Res. Public Health 2016, 13, 152. https://doi.org/10.3390/ijerph13020152
Pereira da Fonseca TA, Pessôa R, Felix AC, Sanabani SS. Diversity of Bacterial Communities on Four Frequently Used Surfaces in a Large Brazilian Teaching Hospital. International Journal of Environmental Research and Public Health. 2016; 13(2):152. https://doi.org/10.3390/ijerph13020152
Chicago/Turabian StylePereira da Fonseca, Tairacan Augusto, Rodrigo Pessôa, Alvina Clara Felix, and Sabri Saeed Sanabani. 2016. "Diversity of Bacterial Communities on Four Frequently Used Surfaces in a Large Brazilian Teaching Hospital" International Journal of Environmental Research and Public Health 13, no. 2: 152. https://doi.org/10.3390/ijerph13020152
APA StylePereira da Fonseca, T. A., Pessôa, R., Felix, A. C., & Sanabani, S. S. (2016). Diversity of Bacterial Communities on Four Frequently Used Surfaces in a Large Brazilian Teaching Hospital. International Journal of Environmental Research and Public Health, 13(2), 152. https://doi.org/10.3390/ijerph13020152





