Soil Microbial Functional and Fungal Diversity as Influenced by Municipal Sewage Sludge Accumulation
Abstract
:1. Introduction
2. Materials and Methods
2.1. Site Description and Sample Collection
| Parameter | Values | |
|---|---|---|
| Soil | Sewage Sludge | |
| pH | 7.5 | 7.5 |
| EC (ms m−1) | 14.58 | 33.50 |
| TOC % | 0.845 | 30.00 |
| N (g 100 g−1) | 0.068 | 5.70 |
| P (mg kg−1) | 373.00 | 600.00 |
| K (mg kg−1) | 223.00 | 469.10 |
| C/N | 12.50 | 5.26 |
| Ca (g kg−1) | 6.38 | 2.52 |
| Mg (g kg−1) | 1.39 | 1.09 |
| Na (g kg−1) | 0.19 | - |
| Zn (mg kg−1) | 43.06 | 326.00 |
| Cu (mg kg−1) | 5.85 | 46.40 |
| Cd (mg kg−1) | 0.16 | - |
| Cr (mg kg−1) | 15.99 | 33.00 |
| Pb (mg kg−1) | 11.51 | 21.00 |
- a soil under the sewage sludge landfill (SS)—the landfill plot was a quadrat of 4 m by 4 m with 40 cm sludge layer on the soil surface,
- a soil located 3 m from a sewage sludge landfill (SS3), and
- a control soil (C) consisting of an undisturbed soil located 200 m from the sewage sludge landfill.
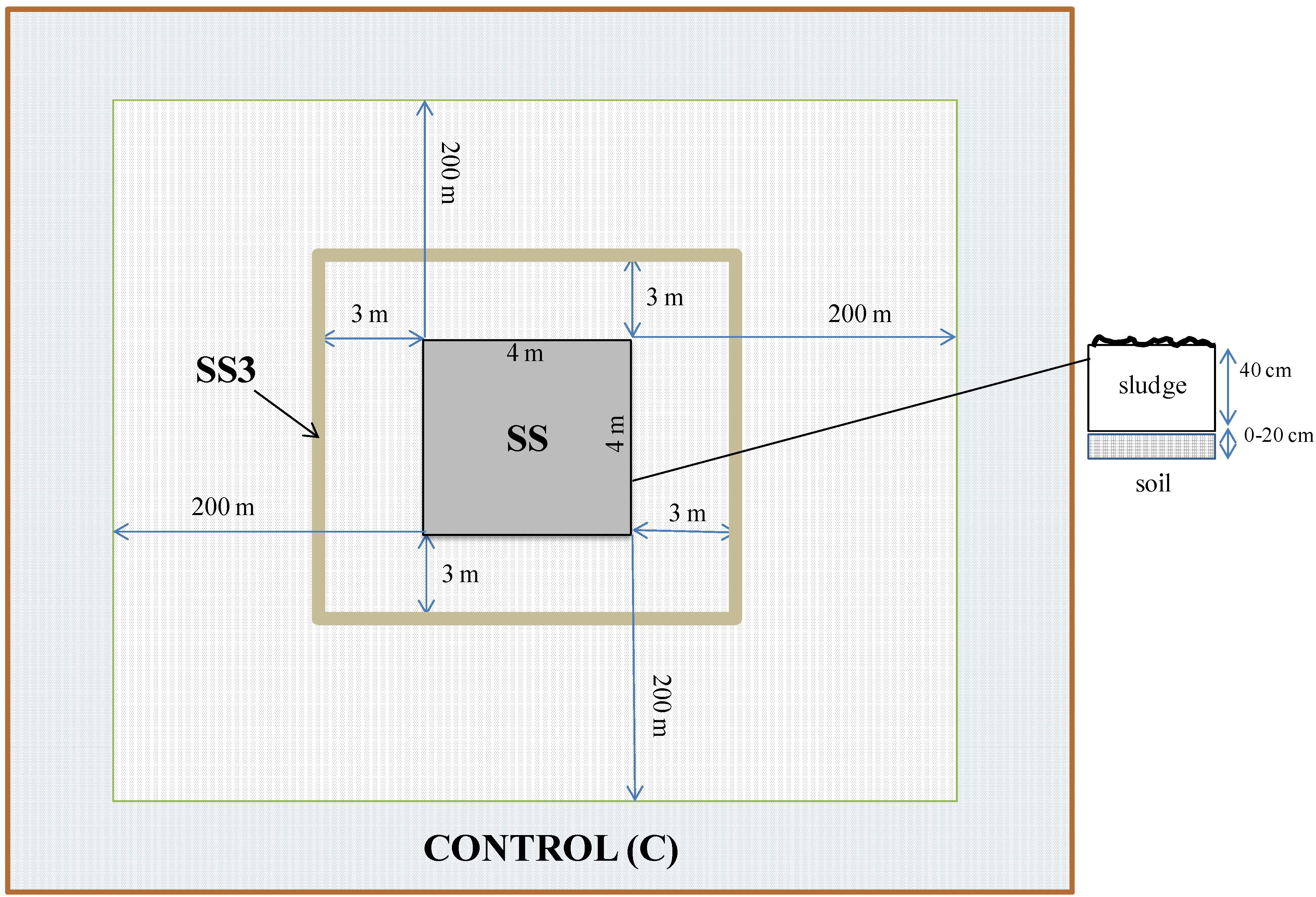
2.2. Chemical Analysis
2.3. Community Level Physiological Profiling (CLPP) Analysis
2.4. Fungal Enumeration and Morphotypic Identification of Fungal Strains
2.5. Molecular Identification of Fungi Isolated from Sewage Sludge
| Initial Step | Each of 30 Cycles | 1 Cycle | Final Step | |
|---|---|---|---|---|
| Melt | Anneal | Final Extension | ||
| HOLD | CYCLE | CYCLE | HOLD | |
| 95 °C | 95 °C | 64 °C | 72 °C | 4 °C |
| 10 s | 0 s | 15 s | 1 min | ∞ |
2.6. Statistical Analysis
3. Results
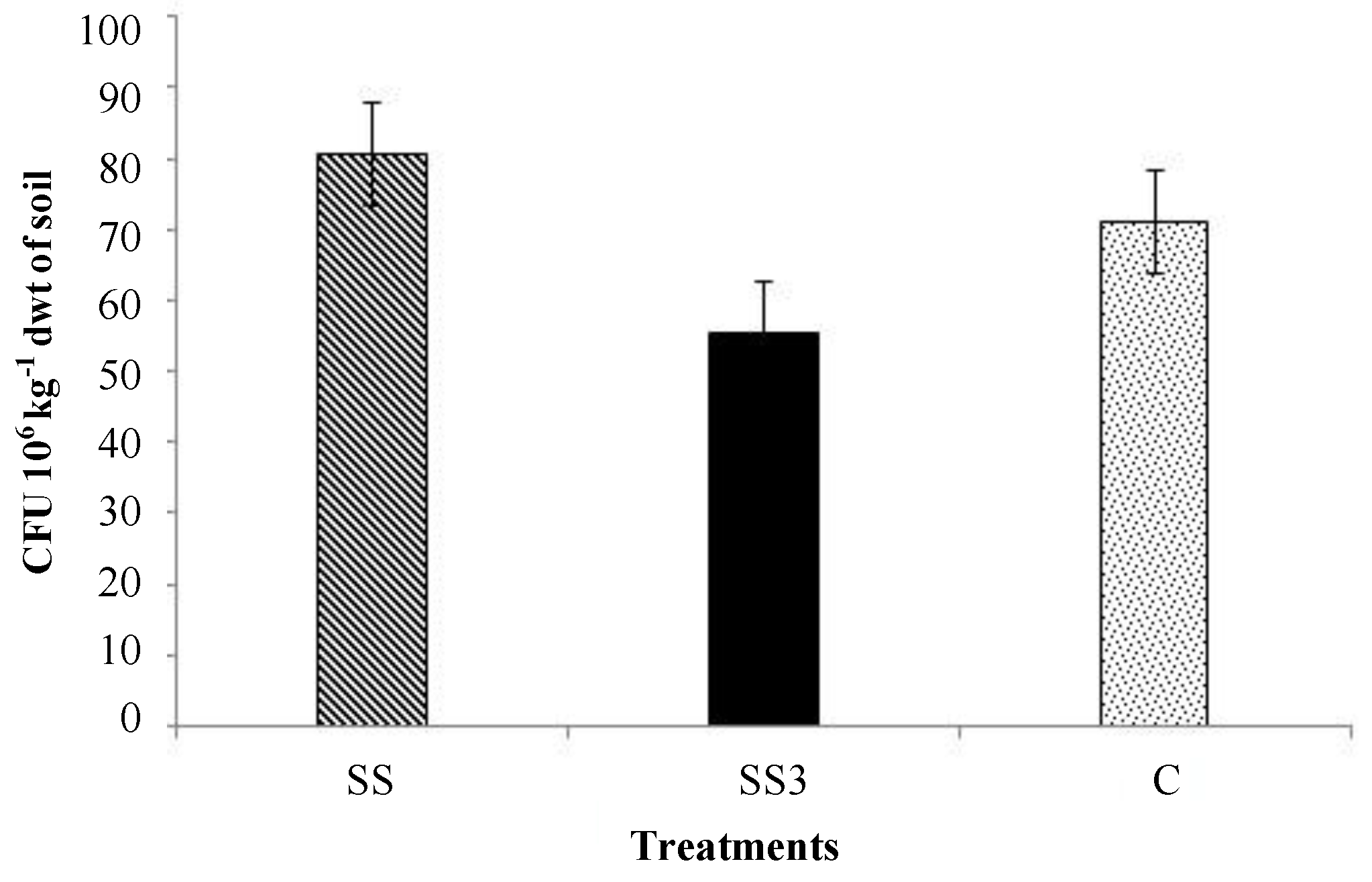
3.1. Microbial Activity
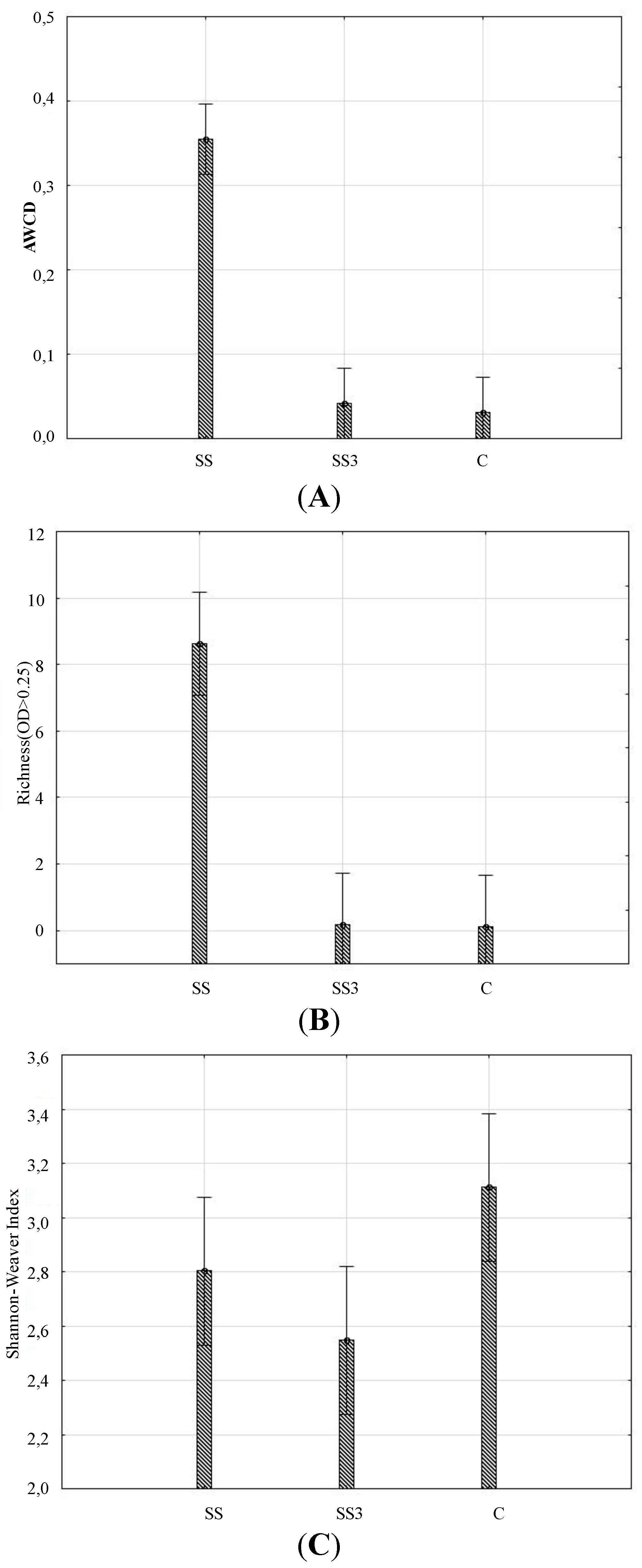
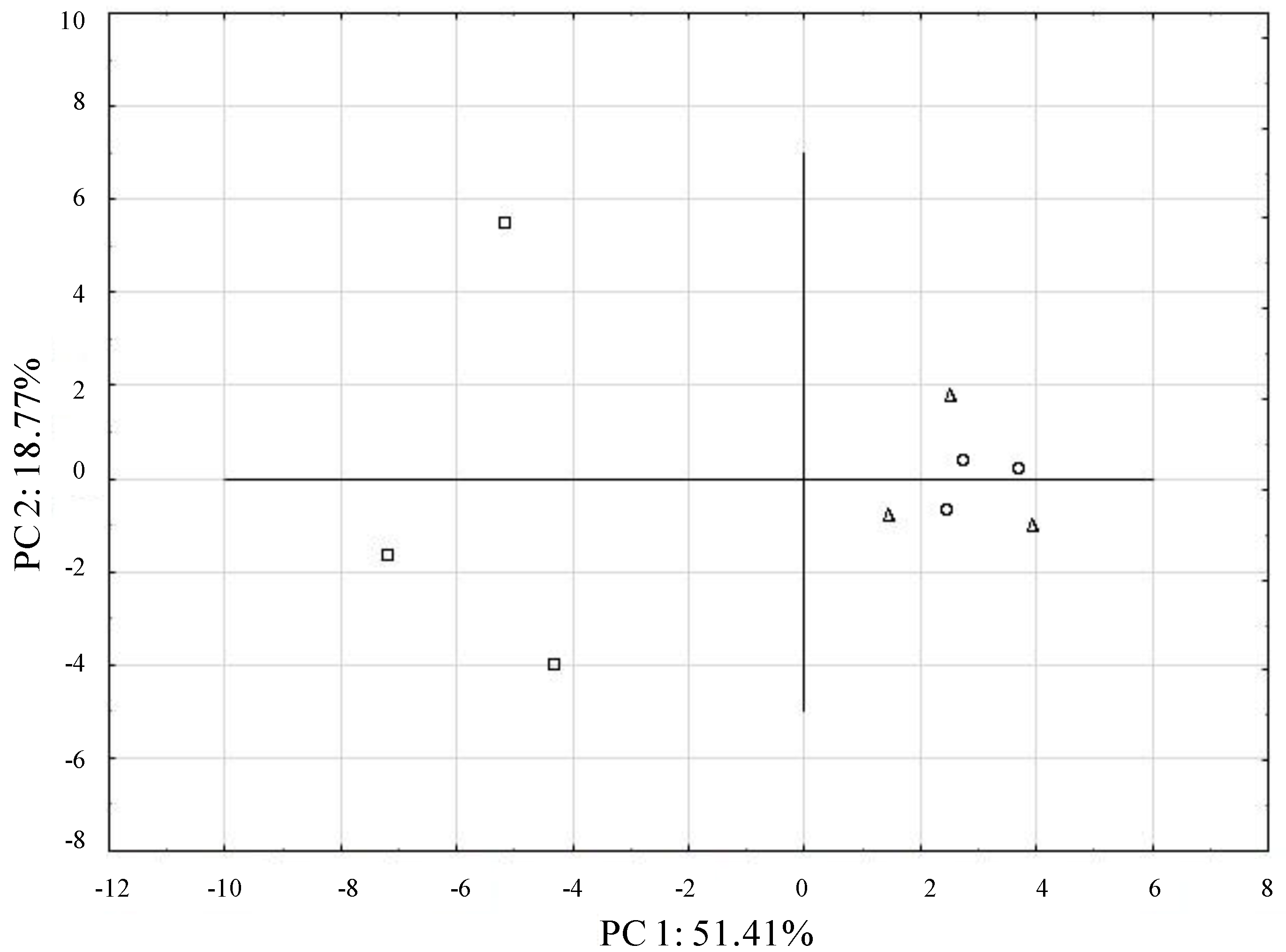
| Principal Components | ||
|---|---|---|
| PC1 | PC2 | |
| d-Galactonic Acid | N-Acetyl-d-Glucosamine | d-Xylose |
| l-Arginine | d-Glucosaminic Acid | Cyclodextrin |
| Pyruvic Acid Methyl Ester | Itaconic Acid | α-A-d Lactose |
| i-Erythritol | Glycyl-l-Glutamic Acid | Putrescine |
| 2-Hydroxybenzoic Acid | d-Glucose-1-Phosphate | |
| Tween 80 | α-Ketobutyric Acid | |
| d-Mannitol | Phenylethylamine | |
| 4-Hydroxybenzoic Acid | dl-α-Glycerol Phosphate | |
| l-Serine | d-Malic Acid | |
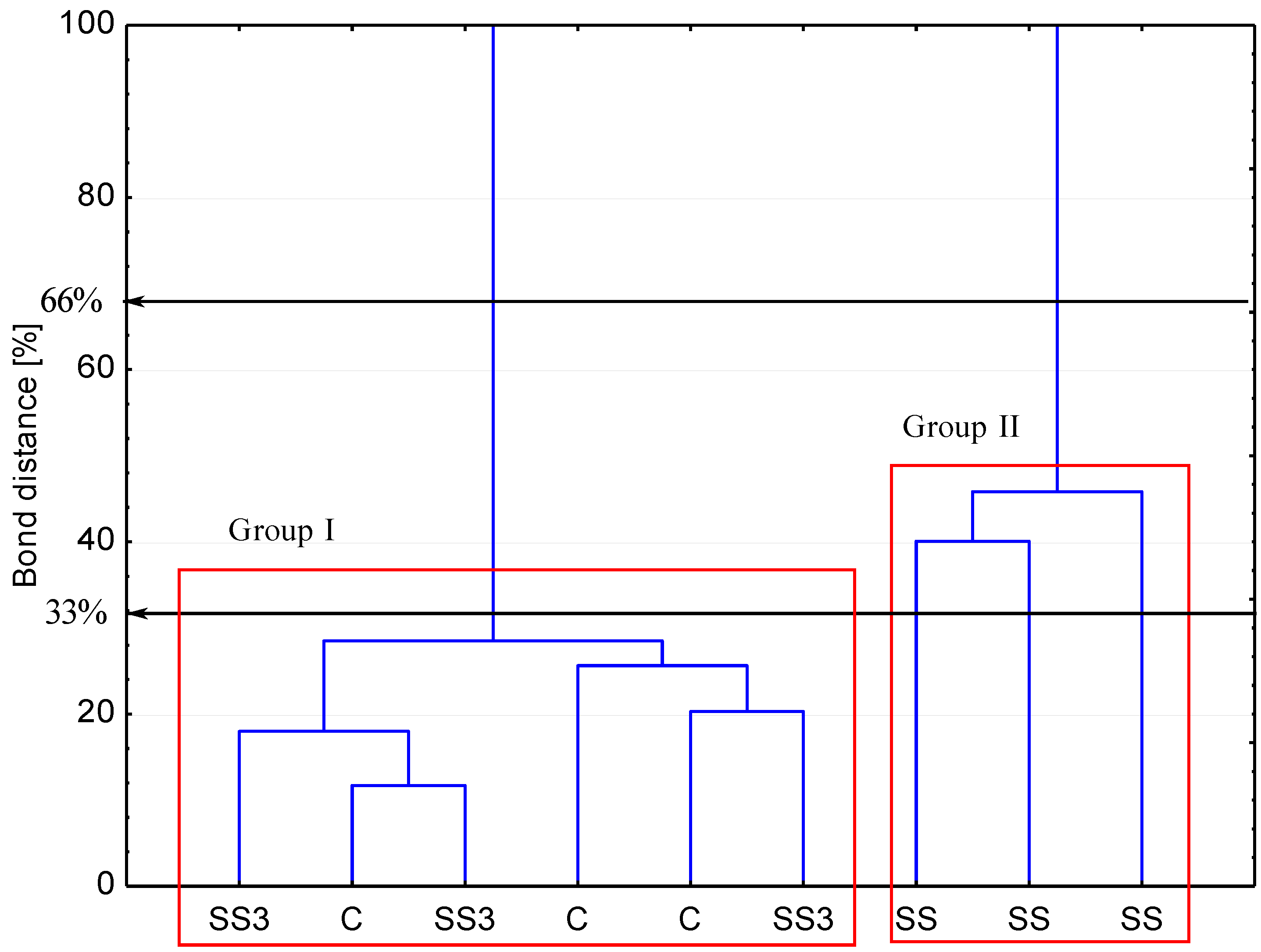
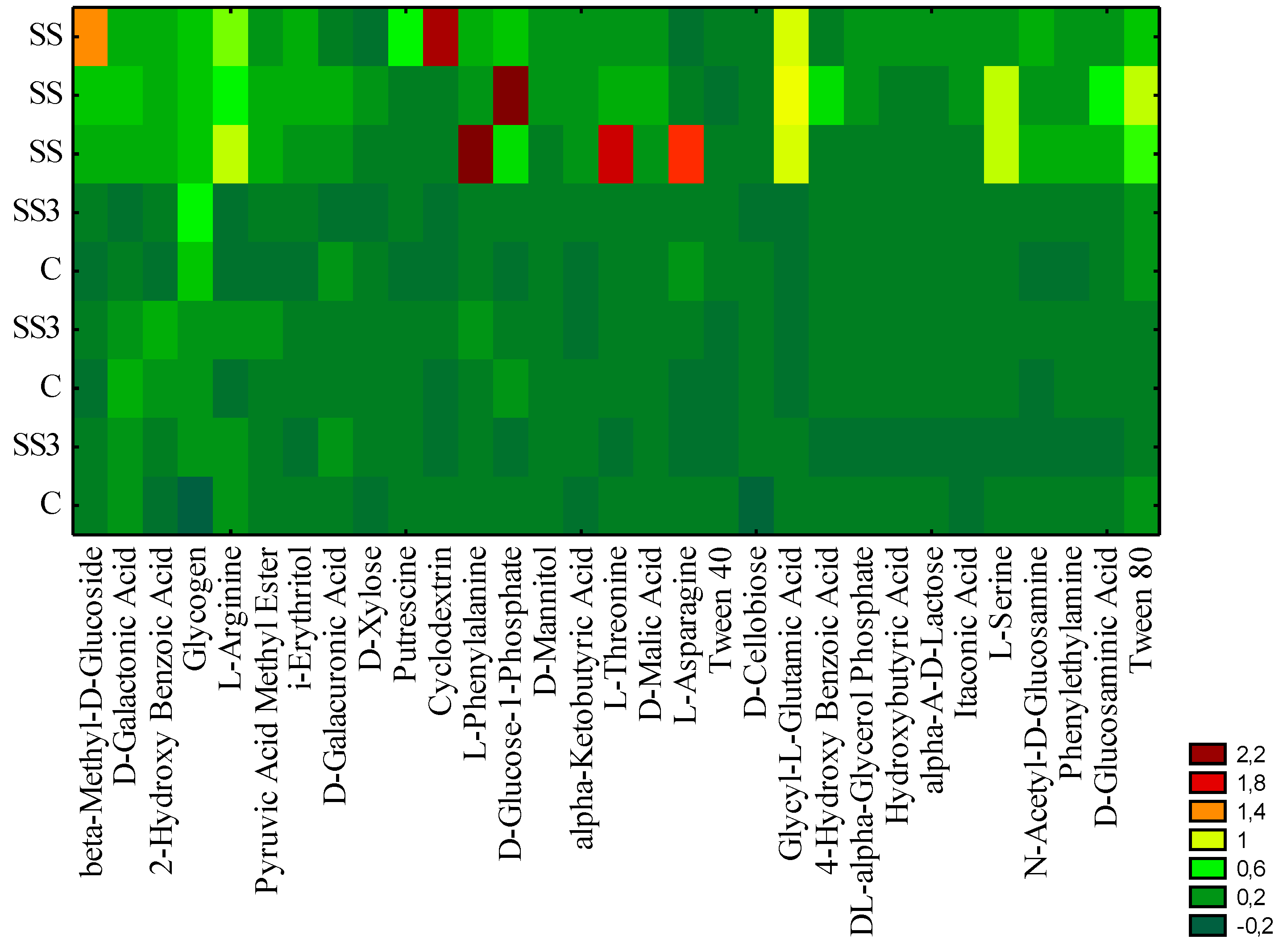

3.2. Pathogenic Fungi and Yeast
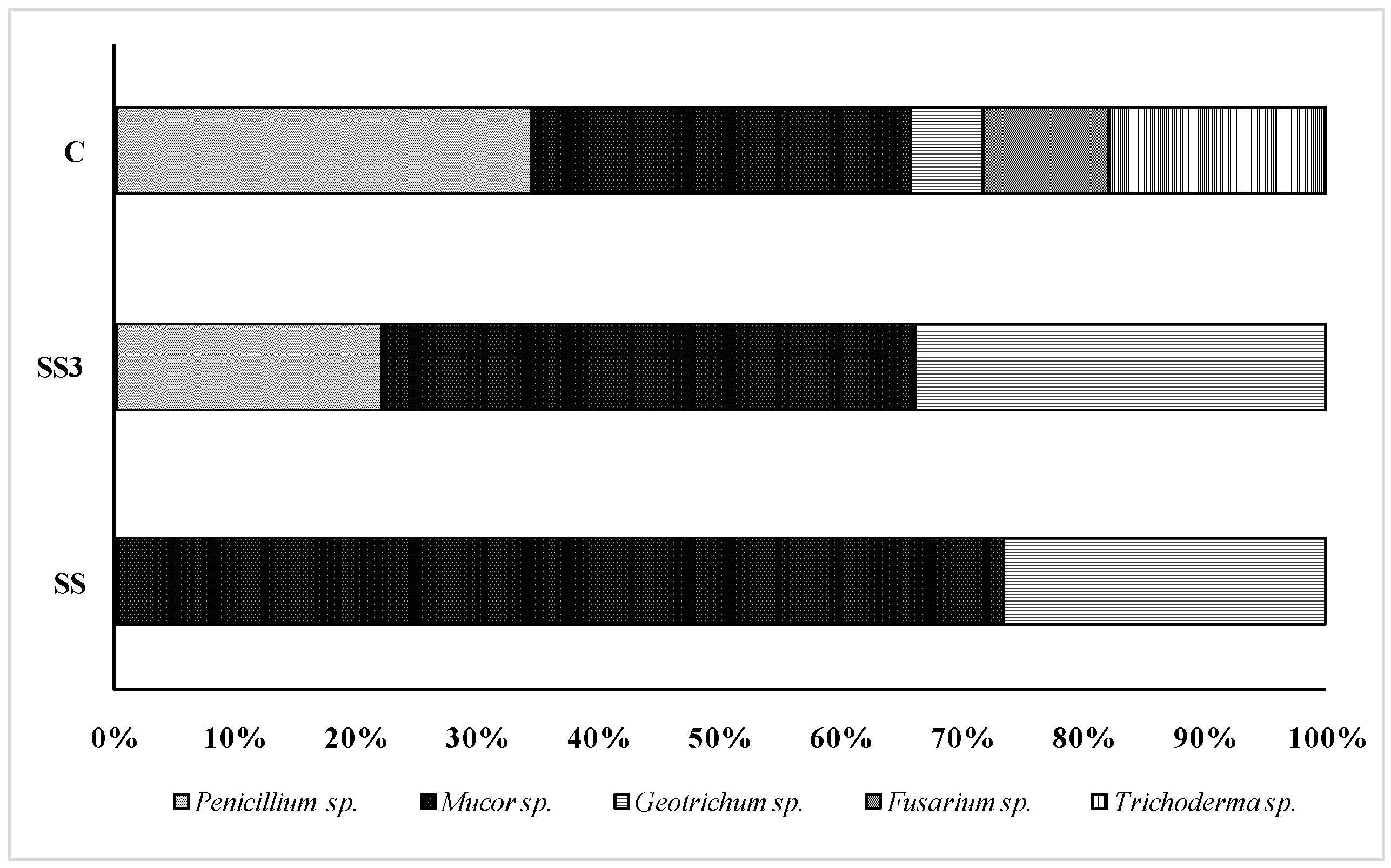
| Sample Name | Specimen Score | Top Match | % Match | Consensus Length | Library Entry Length |
|---|---|---|---|---|---|
| GNO_1_M_circinelloides | 44 | Mucor circinelloides (CBS = 195.68) | 99.62 | 370 | 384 |
| GNO_2_M_circinelloides | 40 | Mucor circinelloides (CBS = 195.68) | 99.30 | 384 | 384 |
| GNO_3_G_citri-aurantii | 42 | Geotrichum citri-aurantii (CBS = 175.89) | 93.12 | 271 | 271 |
| GNO_4_G_citri-aurantii | 42 | Geotrichum citri-aurantii (CBS = 175.89) | 92.42 | 271 | 271 |
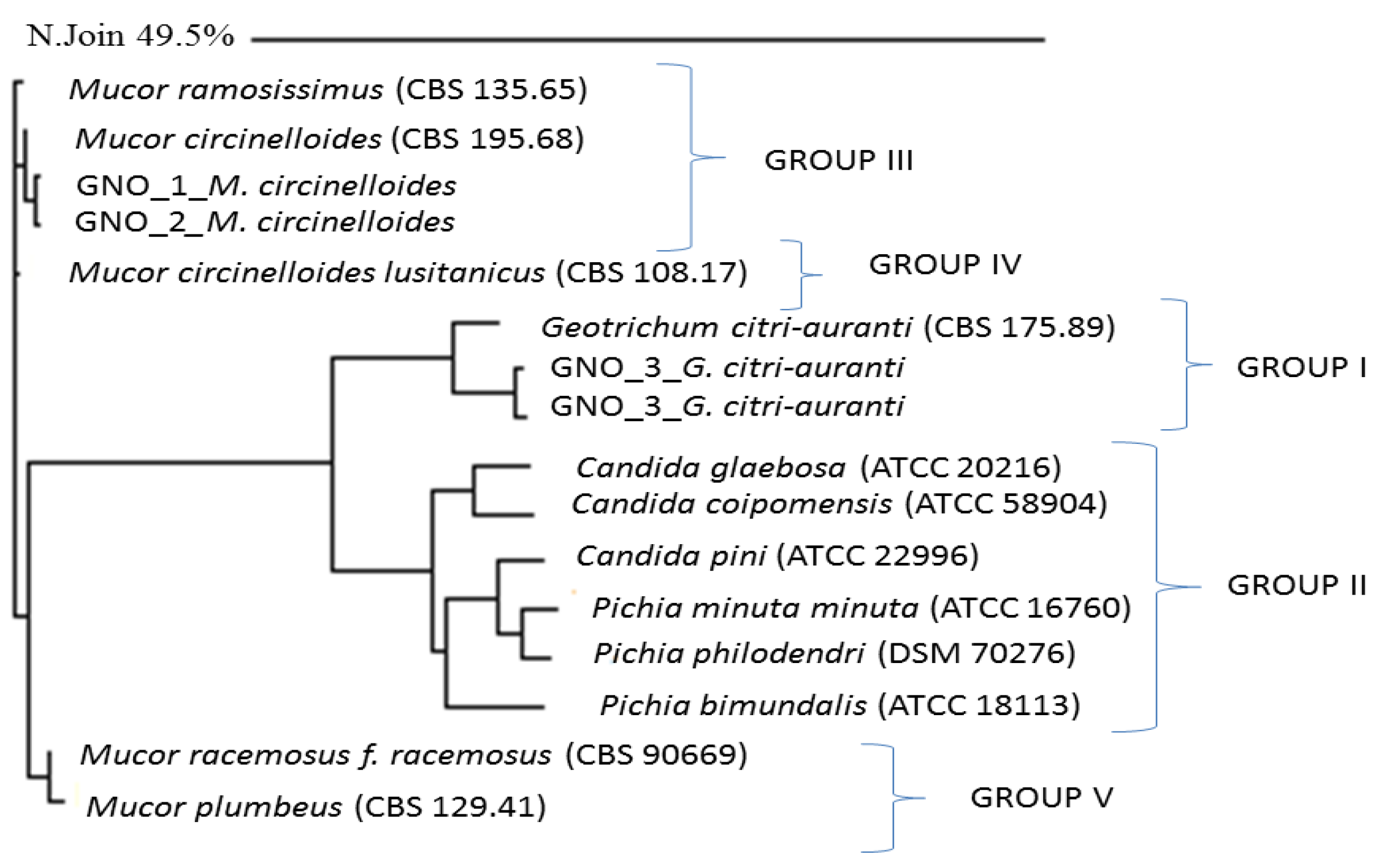
4. Discussion
4.1. Microbial Activity
4.2. Pathogenic Fungi and Yeast
5. Conclusions
- This study showed that community-level physiological profiling using Biolog EcoPlatesTM can be used as a sensitive and effective indicator for evaluating microbial functional diversity and microbial communities under sewage sludge accumulation. These parameters can be useful for monitoring the soil microbial state following sewage sludge application.
- Municipal sewage sludge is rich in molds and yeasts, which are known as fungal plant pathogens. Most fungi that were recovered in this investigation can be considered as potential pathogens. Therefore, all sewage sludge research and accumulation workers should be careful to avoid mycotic infections, and their production must be adapted to control the spread of pathogenic fungi in the environment.
- Fungi in sewage sludges are not only pathogens but also have practical uses for the development of biopreparations for environmental technologies and agricultural biotechnology, especially for the disposal and recycling of organic wastes. Future research studies are needed to assess the properties and use of fungal strains that are isolated from sludges and soils during organic waste degradation.
Acknowledgments
Author Contributions
Conflicts of Interest
References and Notes
- Pascual, I.; Avilés, M.; Aguirreolea, J.; Sánchez-Diaz, M. Effect of sanitized and non-sanitized sewage sludge on soil microbial community and the physiology of pepper plants. Plant. Soil 2008, 310, 41–53. [Google Scholar] [CrossRef]
- Kot, A.; Frąc, M. Methods used in the evaluation of the organic wastes influence on soil microbial activity. Post. Microb. 2014, 53, 183–193. [Google Scholar]
- Frąc, M.; Jezierska-Tys, S. Agricultural utilisation of dairy sewage sludge: Its effect on enzymatic activity and microorganisms of the soil environment. Afr. J. Microb. Res. 2011, 5, 1755–1762. [Google Scholar]
- Zaidi, M.K. Wastewater reuse-risk assessment. Dec. Mak. Environ. Sec. 2007, 1, 357–366. [Google Scholar]
- Łagód, G.; Montusiewicz, A.; Chomczyńska, M.; Bieganowski, A.; Ryżak, M.; Sochan, A. Measurement of activated sludge particle diameters using laser diffraction method. Ecol. Chem. Eng. 2012, 19, 597–608. [Google Scholar]
- National Plan of Wastes Management. 2010. Available online: http://www.mos.gov.pl/kategoria/197_odpady (Accessed on 4 April 2014).
- Sanchez Monedero, M.A.; Mondini, C.; De Nobili, M.; Leita, L.; Roig, A. Land applications of biosolids. Soil response to different stabilization degree or treated organic matter. Waste Manag. 2004, 24, 325–332. [Google Scholar]
- More, T.T.; Yan, S.; Tyagi, R.D.; Surampalli, R.Y. Potential use of filamentous fungi for wastewater sludge treatment. Biores. Technol. 2010, 101, 7691–7700. [Google Scholar] [CrossRef]
- Martins, A.M.; Pagilla, K.; Heijnen, J.J.; van Loosdrecht, M.C. Filamentous bulking sludge—A critical review. Water Res. 2004, 38, 793–817. [Google Scholar] [CrossRef] [PubMed]
- Jezierska-Tys, S.; Frąc, M.; Tys, J. Microbiological hazards resulting from application of dairy sewage sludge: effects on occurrence of pathogenic microorganisms in soil. J. Toxicol. Environ. Health Part. A 2010, 73, 1194–1201. [Google Scholar]
- Singh, R.P.; Agrawal, M. Potential benefits and risks of land application of sewage sludge. Waste Manag. 2008, 28, 347–358. [Google Scholar]
- Ros, M.; Klammer, S.; Knapp, B.; Aichberger, K.; Insam, H. Long-term effects of compost amendment of soil on functional and structural diversity and microbial activity. Soil Use Manag. 2006, 22, 209–218. [Google Scholar] [CrossRef]
- Pankhurst, C.E.; Hawke, B.G.; McDonald, H.J.; Kirby, C.A.; Buckerfield, J.C.; Michelsen, P.; O’Brien, K.A.; Gupta, K.A.; Doube, B.M. Evaluation of soil biological properties as potential bioindicators of soil health. Austr. J. Exp. Agric. 1995, 35, 1015–1028. [Google Scholar] [CrossRef]
- Nannipieri, P.; Ascher, J.; Ceccherini, M.T.; Landi, L.; Pietramellara, G.; Renella, G. Microbial diversity and soil functions. Eur. J. Soil Sci. 2003, 54, 655–670. [Google Scholar] [CrossRef]
- Garland, J.L.; Mills, A.L. Classification and characterization of heterotrophic microbial communities on the basis of patterns community level sole-carbon-source utilization. App. Environ. Microbiol. 1991, 57, 2351–2359. [Google Scholar]
- Frąc, M. Mycological evaluation of dairy sewage sludge and its influence on functional diversity of soil microorganisms. Acta Agrophys. Monograph. 2012, 1, 1–142. [Google Scholar]
- Nielsen, U.N.; Ayres, E.; Wall, D.H.; Bardgett, R.D. Soil biodiversity and carbon cycling: A review and synthesis of studies examining diversity-function relationships. Eur. J. Soil Sci. 2011, 62, 105–116. [Google Scholar] [CrossRef]
- Stefanowicz, A. The Biolog plates technique as a tool in ecological studies of microbial communities. Pol. J. Environ. Stud. 2006, 15, 669–676. [Google Scholar]
- Garland, J.L. Analysis and interpretation of community-level physiological profiles in microbial ecology. FEMS Microbiol. Ecol. 1997, 24, 289–300. [Google Scholar] [CrossRef]
- Gomez, E.; Ferreras, L.; Toresani, S. Soil bacterial functional diversity as influenced by organic amendment application. Biores. Technol. 2006, 97, 1484–1489. [Google Scholar] [CrossRef]
- Stenberg, B. Monitoring soil quality of arable land: Microbiological indicators. Acta Agric. Scand. Sec. B Soil Plant. Sci. 1999, 49, 1–24. [Google Scholar]
- Gomez, E.; Garland, J.; Conti, M. Reproducibility in the response of soil bacterial community-level physiological profiles from a land use intensification gradient. App. Soil Ecol. 2004, 26, 21–30. [Google Scholar] [CrossRef]
- Frąc, M.; Oszust, K.; Lipiec, J. Community level physiological profiles (CLPP), characterization and microbial activity of soil amended with dairy sewage sludge. Sensors 2012, 12, 3253–3268. [Google Scholar]
- Hageskal, G.; Leema, N.; Skaar, I. The study of fungi in drinking water. Microbiol. Res. 2009, 113, 165–172. [Google Scholar]
- Awad, M.F.; Kraume, M. The occurrence of fungi in activated sludge from MBRs. Int. J. Chem. Biol. Eng. 2010, 3, 180–184. [Google Scholar]
- Theodoratos, P.; Moirou, A.; Xenidis, A.; Paspaliaris, I. The use of municipal sewage sludge for the stabilization of soil contaminated by mining activities. J. Haz. Mat. 2000, B 77, 177–191. [Google Scholar]
- Song, U.; Lee, E.J. Environmental and economic assessment of sewage sludge compost application on soil and plants in a landfill. Res. Conserv. Recycl. 2010, 54, 1109–1116. [Google Scholar] [CrossRef]
- Islam, M.R.; Chauhan, P.S.; Kim, Y.; Kim, M.; Sa, T. Community level functional diversity and enzyme activities in paddy soils under different long-term fertilizer management practices. Biol. Fertil. Soils 2011, 47, 599–604. [Google Scholar] [CrossRef]
- Domsch, K.H.; Gams, W.; Anderson, T.H. Compendium of Soil Fungi; Academic Press: Waltham, MA, USA, 1980. [Google Scholar]
- Burnett, H.L.; Hunter, B.B. Illustrated Genera of Imperfect Fungi; Burgess Publishing Company: Minneapolis, MN, USA, 1987. [Google Scholar]
- Watanabe, T. Pictorial Atlas of Soil and Seed Fungi: Morphologies of Cultured Fungi and Key to Species, 3rd ed.; CRC Press: Boca Raton, FL, USA, 2010. [Google Scholar]
- Ellis, J.T.; Amoyal, G.; Ryce, C.; Harper, P.A.; Clough, K.A.; Homan, W.L.; Brindley, P.J. Comparison of the large subunit ribosomal DNA of Neospora and Toxoplasma and development of a new genetic marker for their differentiation based on the D2 domain. Mol. Cell. Prob. 1998, 12, 1–13. [Google Scholar] [CrossRef]
- Rozynek, P.; Gilges, S.; Brüning, T.; Wilhelm, M. Quality test of the MicroSeq D2 LSU Fungal Sequencing Kit for the identification of fungi. Int. J. Hyg. Environ. Health 2003, 206, 297–299. [Google Scholar]
- Bossio, D.A.; Scow, K.M. Impact of carbon and flooding on the metabolic diversity of microbial communities in soil. Appl. Environ. Microb. 1995, 61, 4043–4050. [Google Scholar]
- Sort, X.; Alcaniz, J.M. Modification of soil porosity after application of sewage sludge. Soil Till. Res. 1999, 49, 337–345. [Google Scholar] [CrossRef]
- Sort, X.; Alcan, Ä.J. Contribution of sewage sludge to erosion control in the rehabilitation of limestones quarries. Land Degrad. Develop. 1996, 7, 69–76. [Google Scholar]
- Leitão, A.L. Potential of Penicillium species in the bioremediation field. Int. J. Environ. Res. Public Health 2009, 6, 1393–1417. [Google Scholar]
- Rejman, J.; Brodowski, R. Rill characteristics and sediment transport as a function of slope length during a storm event on loess soil. Earth Sur. Proc. Land. 2005, 30, 231–239. [Google Scholar] [CrossRef]
- Kacprzak, M.; Stańczyk-Mazanek, E. Changes in the structure of fungal communities of soil treated with sewage sludge. Biol. Fertil. Soils 2003, 38, 89–95. [Google Scholar] [CrossRef]
- Brendecke, J.W.; Axelson, R.D.; Pepper, I.L. Soil microbial activity as an indicator of soil fertility: long-term effects of municipal sewage sludge on an arid soil. Soil Biol. Biochem. 1993, 25, 751–758. [Google Scholar] [CrossRef]
- Nishijima, K.A.; Wall, M.M.; Wei, Y.; Wong, D.K. First report of association of Mucor circinelloides on Noni (Morinda citrifolia) in Hawaii. Plant. Disease 2011, 95, 360. [Google Scholar] [CrossRef]
- McKay, A.H.; Förster, H.; Adaskaveg, J.E. Toxicity and resistance potential of selected fungicides to Galactomyces and Penicillium spp. causing postharvest fruit decays of citrus and other crops. Plant. Disease 2012, 96, 1, 87–96. [Google Scholar]
- Farr, D.F.; Bills, G.F.; Chamuris, G.P.; Rossman, A.Y. Fungi on Plants and Plant. Products in the United States; APS Press: St. Paul, MN, USA, 1995. [Google Scholar]
- Andrade, V.S.; Sarubbo, L.A.; Fukushima, K.; Miyaji, M.; Nishimura, A.; de Campos-Takaki, G.M. Production of extracellular proteases by Mucor circinelloides using D-glucose as carbon source/substrate. Braz. J. Microbiol. 2002, 33, 106–110. [Google Scholar]
- Martin, J.F.; Gudiña, E.; Barredo, J.L. Conversion of β-carotene into astaxanthin: Two separate enzymes or a bifunctional hydroxylase-ketolase protein? Microb. Cell. Factor. 2008, 7, 1–10. [Google Scholar] [CrossRef]
- Rodríguez-Frómeta, R.A.; Gutiérrez, A.; Torres-Martínez, S.; Garre, V. Malic enzyme activity is not the only bottleneck for lipid accumulation in the oleaginous fungus Mucor circinelloides. Appl. Microbiol. Biotechnol. 2013, 97, 3063–3072. [Google Scholar]
- Vicente, G.; Bautista, L.F.; Gutierrez, F.J.; Rodriguez, R.; Martinez, V.; Rodriguez-Frometa, R.A.; Ruiz-Vazquez, R.M.; Torres-Martinez, S.; Garre, V. Direct transformation of fungal biomass from submerged cultures into biodiesel. Energy Fuel 2010, 24, 3173–3178. [Google Scholar] [CrossRef]
- Frąc, M.; Ziemiński, K. Methane fermentation process for utilisation of organic waste. Int. Agrophys. 2012, 26, 3, 317–330. [Google Scholar]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Frąc, M.; Oszust, K.; Lipiec, J.; Jezierska-Tys, S.; Nwaichi, E.O. Soil Microbial Functional and Fungal Diversity as Influenced by Municipal Sewage Sludge Accumulation. Int. J. Environ. Res. Public Health 2014, 11, 8891-8908. https://doi.org/10.3390/ijerph110908891
Frąc M, Oszust K, Lipiec J, Jezierska-Tys S, Nwaichi EO. Soil Microbial Functional and Fungal Diversity as Influenced by Municipal Sewage Sludge Accumulation. International Journal of Environmental Research and Public Health. 2014; 11(9):8891-8908. https://doi.org/10.3390/ijerph110908891
Chicago/Turabian StyleFrąc, Magdalena, Karolina Oszust, Jerzy Lipiec, Stefania Jezierska-Tys, and Eucharia Oluchi Nwaichi. 2014. "Soil Microbial Functional and Fungal Diversity as Influenced by Municipal Sewage Sludge Accumulation" International Journal of Environmental Research and Public Health 11, no. 9: 8891-8908. https://doi.org/10.3390/ijerph110908891
APA StyleFrąc, M., Oszust, K., Lipiec, J., Jezierska-Tys, S., & Nwaichi, E. O. (2014). Soil Microbial Functional and Fungal Diversity as Influenced by Municipal Sewage Sludge Accumulation. International Journal of Environmental Research and Public Health, 11(9), 8891-8908. https://doi.org/10.3390/ijerph110908891






