In Vitro Interactions between 17β-Estradiol and DNA Result in Formation of the Hormone-DNA Complexes
Abstract
:1. Introduction
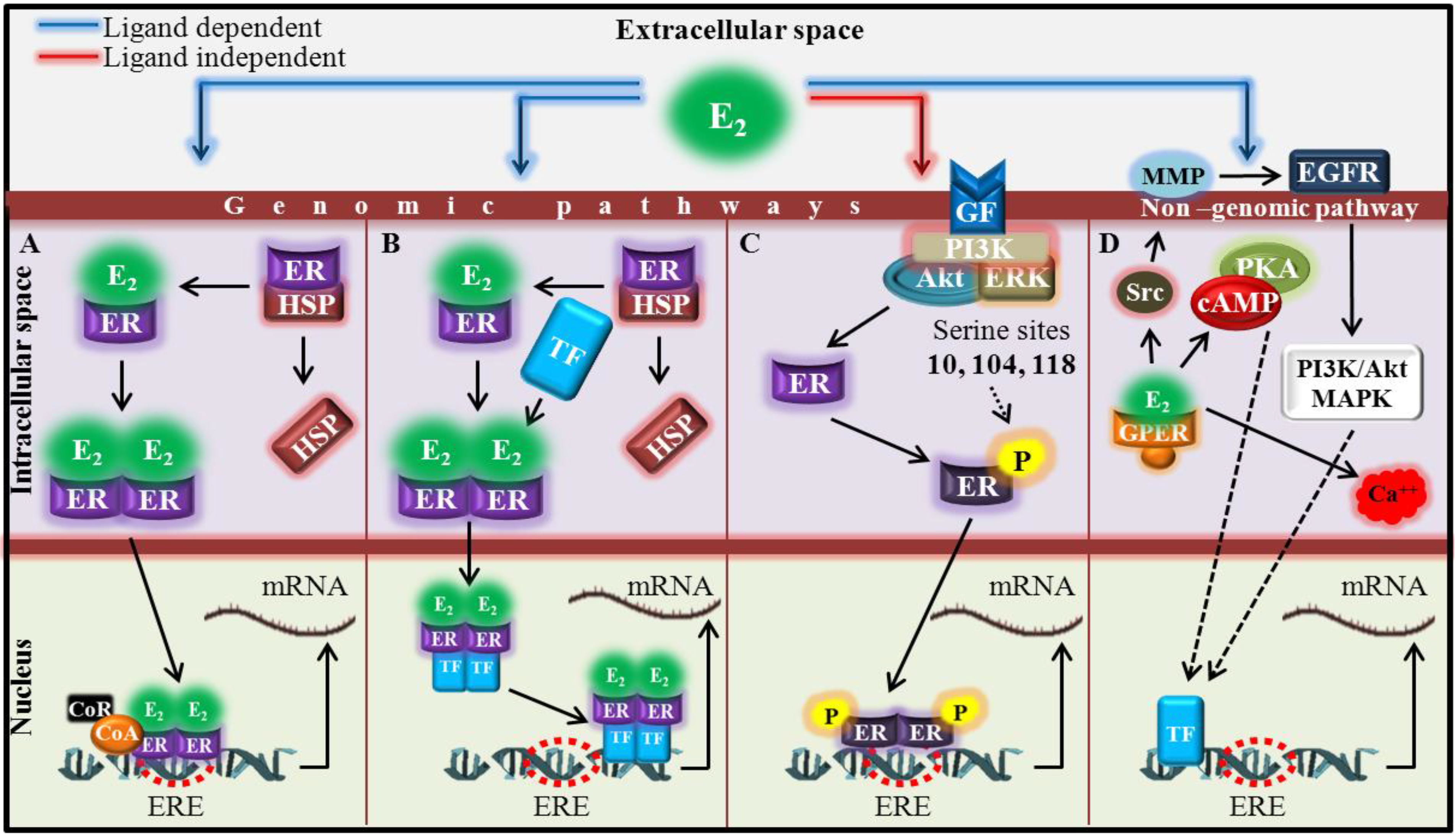
2. Experimental Section
2.1. Chemicals and pH Measurement
2.2. Oligonucleotides
| ERE1 | 5’-CTAATCACTCTGACCAT-3’ 3’-GATTAGTGAGACTGGTA-5’ |
| ERE2 | 5’-CCAGGTCAGAGTGACCTGAG-3’ 3’-GGTCCAGTCTCACTGGACTC-5’ |
| ERE3 | 5’-GCAGGTCAGAGTGACCTGAGCTAG-3’ 3’-CGTCCAGTCTCACTGGACTCGATC-5’ |
| ERE4 | 5’-CCAGGTCAGAGTGACCTGAGCTACGGTGACACAGGCAG-3’ 3’-GGTCCAGTCTCACTGGACTCGATGCCACTGTGTCCGTC-5’ |
2.3. The Hybridization Process of Single-Stranded DNA Fragments
2.4. Spectrophotometric Analysis of Interactions between E2 and DNA Fragments
2.5. MALDI-TOF MS Analysis of E2/DNA Fragments Interaction
2.6. Descriptive Statistics
3. Results and Discussion
3.1. An Optical Behavior of DNA Affected by 17β-Estradiol
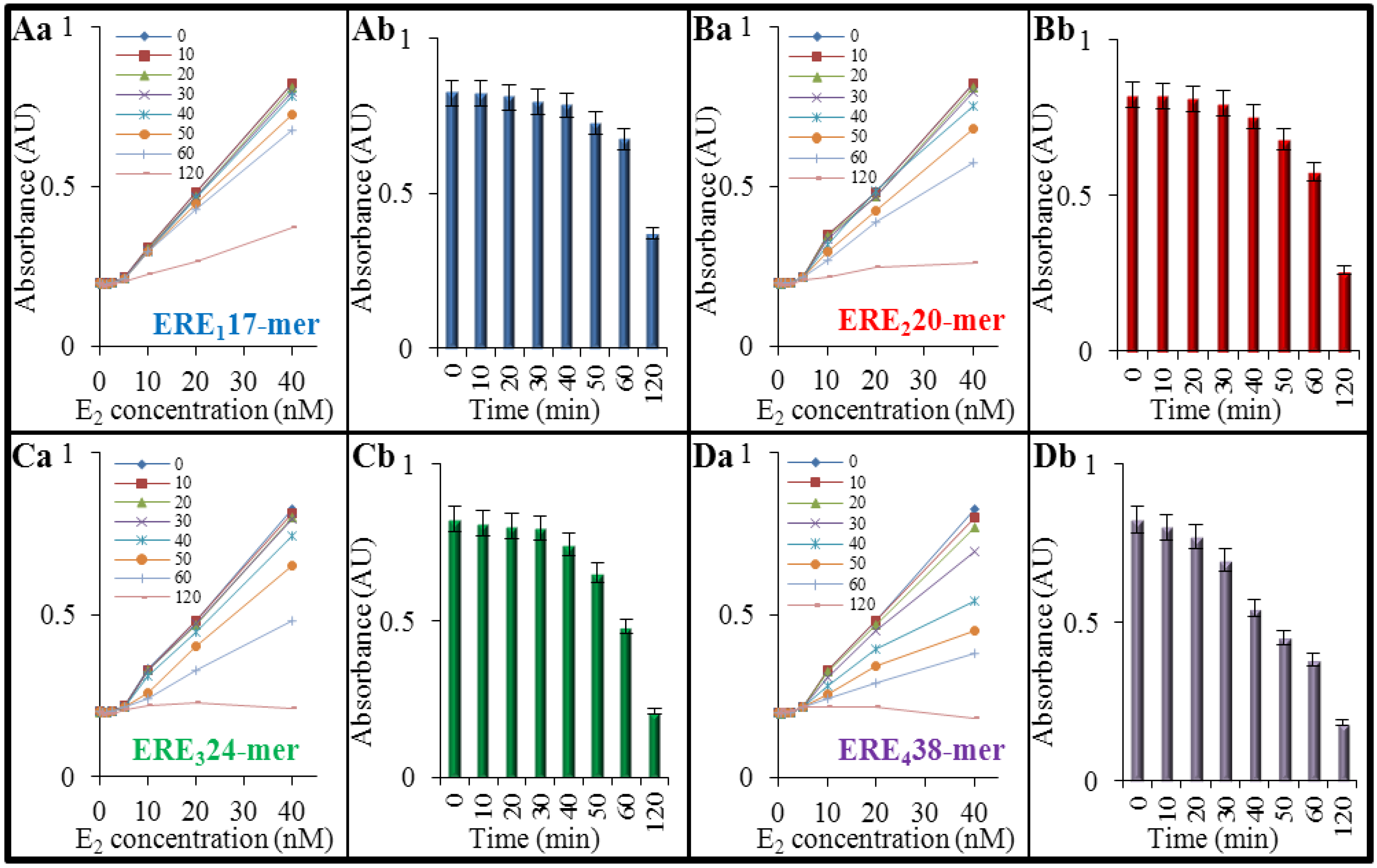
3.2. An Effect of 17β-Estradiol on Denaturation of Double-Stranded DNA
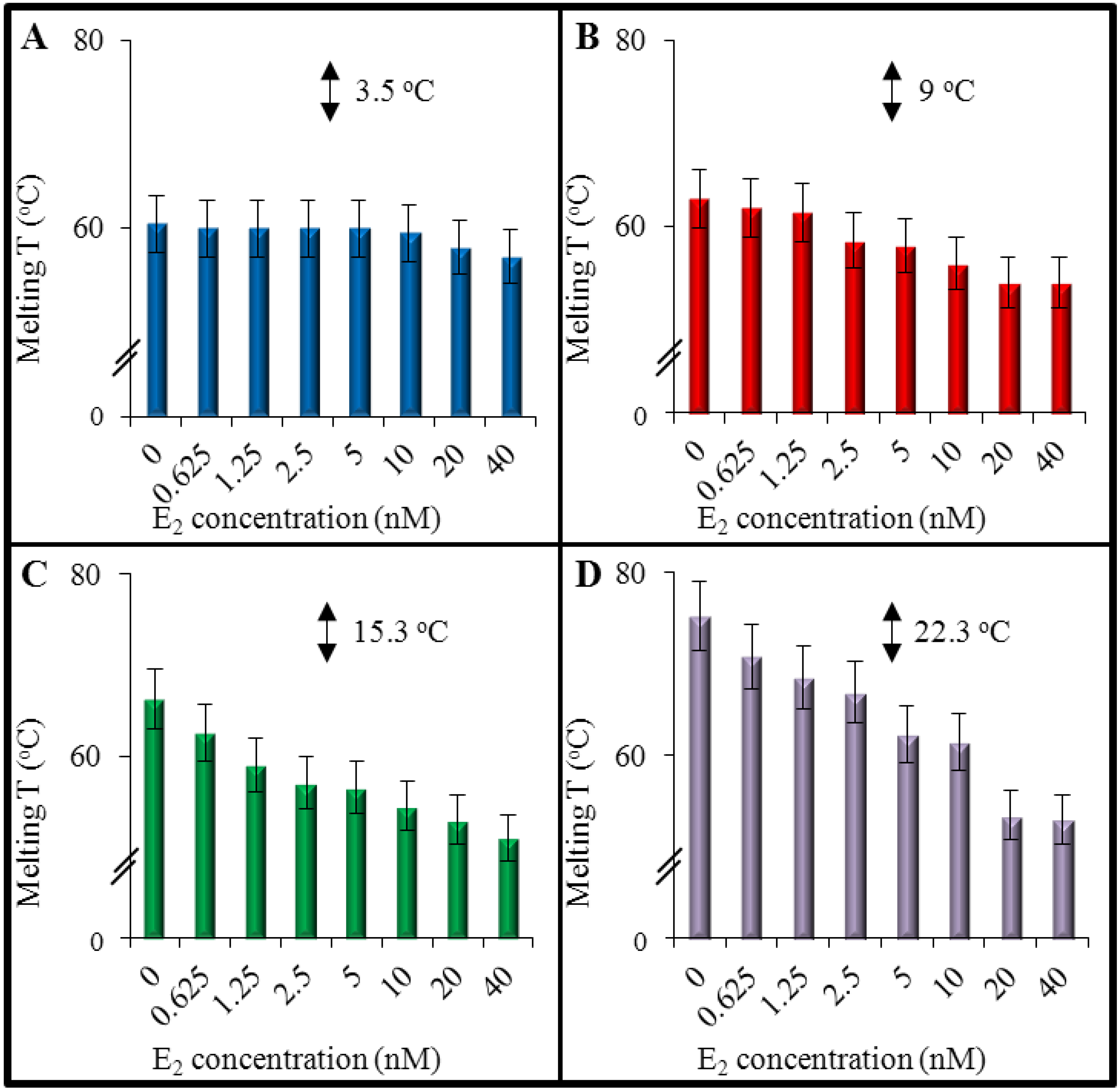
3.3.MALDI-TOF Mass Spectrometry for Evaluation of E2-DNA Interaction
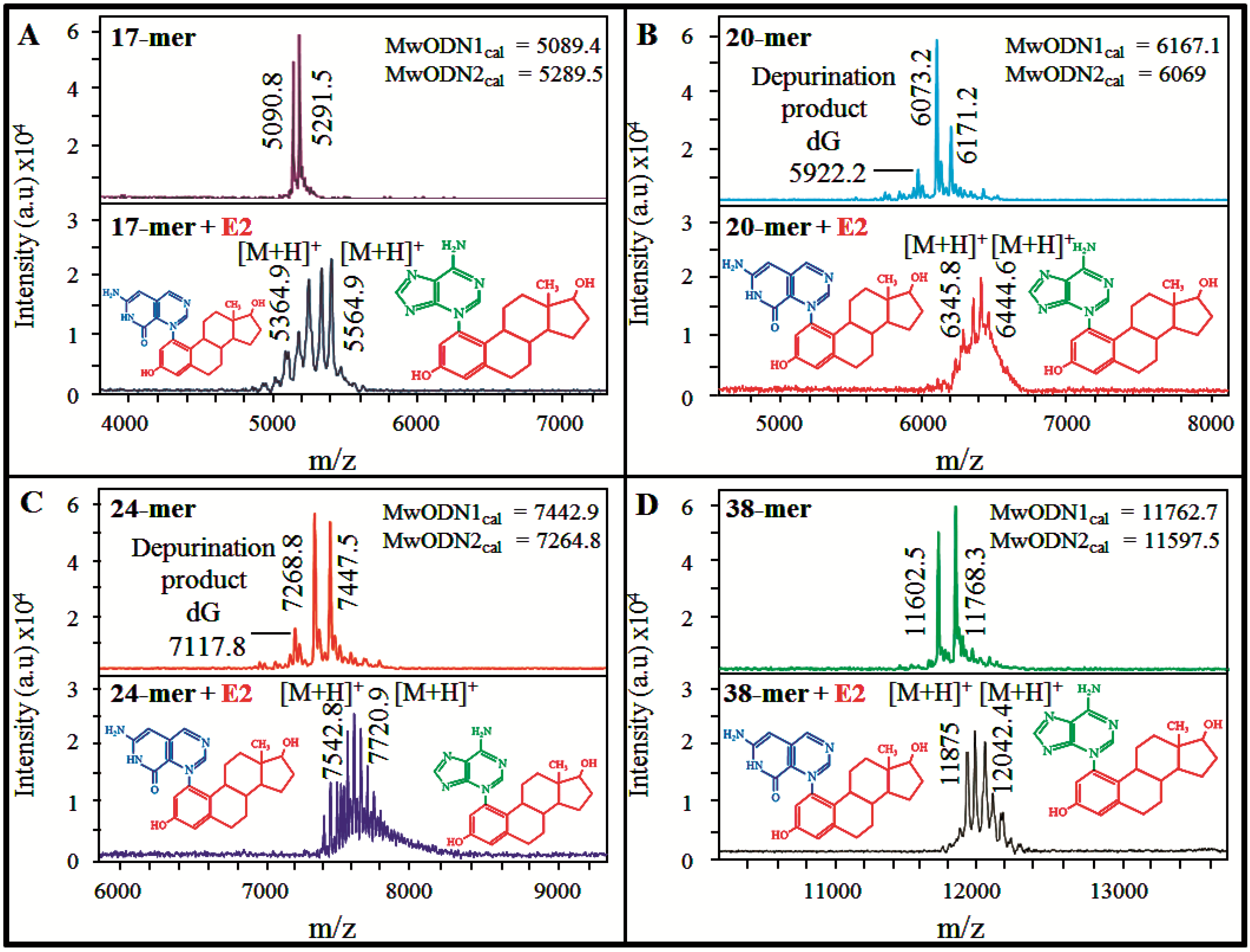
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Benotti, M.J.; Trenholm, R.A.; Vanderford, B.J.; Holady, J.C.; Stanford, B.D.; Snyder, S.A. Pharmaceuticals and endocrine disrupting compounds in US drinking water. Environ. Sci. Technol. 2009, 43, 597–603. [Google Scholar] [CrossRef]
- Boyd, G.R.; Reemtsma, H.; Grimm, D.A.; Mitra, S. Pharmaceuticals and personal care products (PPCPs) in surface and treated waters of Louisiana, USA and Ontario, Canada. Sci. Total Environ. 2003, 311, 135–149. [Google Scholar] [CrossRef]
- Kolpin, D.W.; Furlong, E.T.; Meyer, M.T.; Thurman, E.M.; Zaugg, S.D.; Barber, L.B.; Buxton, H.T. Pharmaceuticals, hormones, and other organic wastewater contaminants in US streams, 1999–2000: A national reconnaissance. Environ. Sci. Technol. 2002, 36, 1202–1211. [Google Scholar] [CrossRef]
- Jiang, W.W.; Yan, Y.; Ma, M.; Wang, D.H.; Luo, Q.; Wang, Z.J.; Satyanarayanan, S.K. Assessment of source water contamination by estrogenic disrupting compounds in China. J. Environ. Sci. 2012, 24, 320–328. [Google Scholar] [CrossRef]
- Snyder, E.M.; Snyder, S.A.; Kelly, K.L.; Gross, T.S.; Villeneuve, D.L.; Fitzgerald, S.D.; Villalobos, S.A.; Giesy, J.P. Reproductive responses of common carp (Cyprinus carpio) exposed in cages to influent of the Las Vegas Wash in Lake Mead, Nevada, from late winter to early spring. Environ. Sci. Technol. 2004, 38, 6385–6395. [Google Scholar] [CrossRef]
- Isobe, T.; Serizawa, S.; Horiguchi, T.; Shibata, Y.; Managaki, S.; Takada, H.; Morita, M.; Shiraishi, H. Horizontal distribution of steroid estrogens in surface sediments in Tokyo Bay. Environ. Pollut. 2006, 144, 632–638. [Google Scholar] [CrossRef]
- David, A.; Fenet, H.; Gomez, E. Alkylphenols in marine environments: Distribution monitoring strategies and detection considerations. Mar. Pollut. Bull. 2009, 58, 953–960. [Google Scholar] [CrossRef]
- Boulay, F.; Perdiz, D. 17β-Estradiol modulates UVB-induced cellular responses in estrogen receptors positive human breast cancer cells. J. Photochem. Photobiol. B Biol. 2005, 81, 143–153. [Google Scholar] [CrossRef]
- Yan, Z.H.; Lu, G.H.; Wu, D.H.; Ye, Q.X.; Xie, Z.X. Interaction of 17 beta-estradiol and ketoconazole on endocrine function in goldfish (Carassius auratus). Aquat. Toxicol. 2013, 132, 19–25. [Google Scholar]
- Heldring, N.; Pike, A.; Andersson, S.; Matthews, J.; Cheng, G.; Hartman, J.; Tujague, M.; Strom, A.; Treuter, E.; Warner, M.; et al. Estrogen receptors: How do they signal and what are their targets. Physiol. Rev. 2007, 87, 905–931. [Google Scholar] [CrossRef]
- Lee, H.R.; Jeung, E.B.; Cho, M.H.; Kim, T.H.; Leung, P.C.K.; Choi, K.C. Molecular mechanism(s) of endocrine-disrupting chemicals and their potent oestrogenicity in diverse cells and tissues that express oestrogen receptors. J. Cell. Mol. Med. 2013, 17, 1–11. [Google Scholar] [CrossRef]
- Tang, X.J.; Naveedullah; Hashmi, M.Z.; Zhang, H.; Qian, M.R.; Yu, C.N.; Shen, C.F.; Qin, Z.H.; Huang, R.L.; Qiao, J.N.; et al. A preliminary study on the occurrence and dissipation of estrogen in livestock wastewater. Bull. Environ. Contam. Toxicol. 2013, 90, 391–396. [Google Scholar] [CrossRef]
- Macon, M.B.; Fenton, S.E. Endocrine disruptors and the breast: Early life effects and later life disease. J. Mammary Gland Biol. Neoplasia 2013, 18, 43–61. [Google Scholar] [CrossRef]
- Jenkins, S.; Betancourt, A.M.; Wang, J.; Lamartiniere, C.A. Endocrine-active chemicals in mammary cancer causation and prevention. J. Steroid Biochem. Mol. Biol. 2012, 129, 191–200. [Google Scholar] [CrossRef]
- Ombra, M.N.; di Santi, A.; Abbondanza, C.; Migliaccio, A.; Avvedimento, E.V.; Perillo, B. Retinoic acid impairs estrogen signaling in breast cancer cells by interfering with activation of LSD1 via PKA. Biochim. Biophys. Acta 2013, 1829, 480–486. [Google Scholar] [CrossRef]
- Lacroix, M.; Leclercq, G. Relevance of breast cancer cell lines as models for breast tumours: An update. Breast Cancer Res. Treat. 2004, 83, 249–289. [Google Scholar] [CrossRef]
- Resende, F.A.; de Oliveira, A.P.S.; de Camargo, M.S.; Vilegas, W.; Varanda, E.A. Evaluation of estrogenic potential of flavonoids using a recombinant yeast strain and MCF7/BUS cell proliferation assay. PLoS One 2013, 8. [Google Scholar] [CrossRef]
- Cavalieri, E.L.; Rogan, E.G. Depurinating estrogen-DNA adducts in the etiology and prevention of breast and other human cancers. Future Oncol. 2010, 6, 75–91. [Google Scholar] [CrossRef]
- Cavalieri, E.L.; Stack, D.E.; Devanesan, P.D.; Todorovic, R.; Dwivedy, I.; Higginbotham, S.; Johansson, S.L.; Patil, K.D.; Gross, M.L.; Gooden, J.K.; et al. Molecular origin of cancer: Catechol estrogen-3,4-quinones as endogenous tumor initiators. Proc. Natl. Acad. Sci. USA 1997, 94, 10937–10942. [Google Scholar] [CrossRef]
- Abbondanza, C.; de Rosa, C.; D’Arcangelo, A.; Pacifico, M.; Spizuoco, C.; Piluso, G.; Di Zazzo, E.; Gazzerro, P.; Medici, N.; Moncharmont, B.; et al. Identification of a functional estrogen-responsive enhancer element in the promoter 2 of PRDM2 gene in breast cancer cell lines. J. Cell. Physiol. 2012, 227, 964–975. [Google Scholar] [CrossRef]
- Hung, Y.C.; Chang, W.C.; Chen, L.M.; Chang, Y.Y.; Wu, L.Y.; Chung, W.M.; Lin, T.Y.; Chen, L.C.; Ma, W.L. Non-genomic estrogen/estrogen receptor alpha promotes cellular malignancy of immature ovarian teratoma in vitro. J. Cell. Physiol. 2014, 229, 752–761. [Google Scholar] [CrossRef]
- Cirillo, F.; Nassa, G.; Taralla, R.; Stellato, C.; De Filippo, M.R.; Arribrosino, C.; Baumann, M.; Nyman, T.A.; Weisz, A. Molecular mechanisms of selective estrogen receptor modulator activity in human breast cancer cells: Identification of novel nuclear cofactors of antiestrogen-ER alpha complexes by interaction proteomics. J. Proteome Res. 2013, 12, 421–431. [Google Scholar] [CrossRef]
- Heldring, N.; Isaacs, G.D.; Diehl, A.G.; Sun, M.; Cheung, E.; Ranish, J.A.; Kraus, W.L. Multiple sequence-specific DNA-binding proteins mediate estrogen receptor signaling through a tethering pathway. Mol. Endocrinol. 2011, 25, 564–574. [Google Scholar] [CrossRef]
- Acconcia, F.; Kumar, R. Signaling regulation of genomic and nongenomic functions of estrogen receptors. Cancer Lett. 2006, 238, 1–14. [Google Scholar] [CrossRef]
- McGlynn, L.M.; Tovey, S.; Bartlett, J.M.S.; Doughty, J.; Cooke, T.G.; Edwards, J. Interactions between MAP kinase and oestrogen receptor in human breast cancer. Eur. J. Cancer 2013, 49, 1176–1186. [Google Scholar] [CrossRef]
- Heger, Z.; Zitka, O.; Krizkova, S.; Beklova, M.; Kizek, R.; Adam, V. Molecular biology of beta-estradiol-estrogen receptor complex binding to estrogen response element and the effect on cell proliferation. Neuroendocrinol. Lett. 2013, 34, 123–129. [Google Scholar]
- Losel, R.M.; Falkenstein, E.; Feuring, M.; Schultz, A.; Tillmann, H.C.; Rossol-Haseroth, K.; Wehling, M. Nongenomic steroid action: Controversies, questions, and answers. Physiol. Rev. 2003, 83, 965–1016. [Google Scholar]
- Suman, M.; Giacomello, M.; Corain, L.; Ballarin, C.; Montelli, S.; Cozzi, B.; Peruffo, A. Estradiol effects on intracellular Ca2+ homeostasis in bovine brain-derived endothelial cells. Cell Tissue Res. 2012, 350, 109–118. [Google Scholar] [CrossRef]
- Candelaria, N.R.; Liu, K.; Lin, C.Y. Estrogen receptor alpha: Molecular mechanisms and emerging insights. J. Cell. Biochem. 2013, 114, 2203–2208. [Google Scholar] [CrossRef]
- Imaoka, T.; Nishimura, M.; Doi, K.; Tani, S.; Ishikawa, K.; Yamashita, S.; Ushijima, T.; Imai, T.; Shimada, Y. Molecular characterization of cancer reveals interactions between ionizing radiation and chemicals on rat mammary carcinogenesis. Int. J. Cancer 2014, 134, 1529–1538. [Google Scholar] [CrossRef]
- Wang, Y.C.; Lin, W.L.; Lin, Y.J.; Tang, F.Y.; Chen, Y.M.; Chiang, E.P.I. A novel role of the tumor suppressor GNMT in cellular defense against DNA damage. Int. J. Cancer 2014, 134, 799–810. [Google Scholar] [CrossRef]
- Saeed, M.; Zahid, M.; Gunselman, S.J.; Rogan, E.; Cavalieri, E. Slow loss of deoxyribose from the N7deoxygruanosine adducts of estradiol-3,4-quinone and hexestrol-3,4’-quinone. Implications for mutagenic activity. Steroids 2005, 70, 29–35. [Google Scholar] [CrossRef]
- Peng, B.; Chen, X.; Du, K.J.; Yu, B.L.; Chao, H.; Ji, L.N. Synthesis, characterization and DNA-binding studies of ruthenium(II) mixed-ligand complexes containing dipyrido 1,2,5 oxadiazolo 3,4-b quinoxaline. Spectroc. Acta A Mol. Biomol. Spectr. 2009, 74, 896–901. [Google Scholar] [CrossRef]
- Syed, S.N.; Schulze, H.; Macdonald, D.; Crain, J.; Mount, A.R.; Bachmann, T.T. Cyclic denaturation and renaturation of double-stranded DNA by redox-state switching of DNA intercalators. J. Am. Chem. Soc. 2013, 135, 5399–5407. [Google Scholar] [CrossRef]
- Vizard, D.L.; White, R.A.; Ansevin, A.T. Comparison of theory to experiment for DNA thermal-denaturation. Nature 1978, 275, 250–251. [Google Scholar] [CrossRef]
- Breslauer, K.J.; Frank, R.; Blocker, H.; Marky, L.A. Predicting DNA duplex stability from the base sequence. Proc. Natl. Acad. Sci. USA 1986, 83, 3746–3750. [Google Scholar] [CrossRef]
- Melchior, W.B.; Vonhippe, P.H. Alteration of relative stability of dA-dT and dG-dC base pairs in DNA. Proc. Natl. Acad. Sci. USA 1973, 70, 298–302. [Google Scholar] [CrossRef]
- Srinivasan, J.R.; Liu, Y.H.; Venta, P.J.; Siemieniak, D.; Killeen, A.A.; Zhu, Y.D.; Lubman, D.M. Matrix-assisted laser desorption/ionization time-of-flight mass spectrometry as a rapid screening method to detect mutations causing Tay-Sachs disease. Rapid Commun. Mass Spectrom. 1997, 11, 1144–1150. [Google Scholar] [CrossRef]
- Zhang, L.K.; Gross, M.L. Matrix-assisted laser desorption/ionization mass spectrometry methods for oligodeoxynucleotides: Improvements in matrix, detection limits, quantification, and sequencing. J. Am. Soc. Mass Spectrom. 2000, 11, 854–865. [Google Scholar] [CrossRef]
- Yu, F.; Xu, S.Y.; Pan, C.S.; Ye, M.L.; Zou, H.F.; Guo, B.C. A matrix of 3,4-diaminobenzophenone for the analysis of oligonucleotides by matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. Nucleic Acids Res. 2006, 34, 1–7. [Google Scholar] [CrossRef]
- Ohara, K.; Smietana, M.; Vasseur, J.J. Characterization of specific noncovalent complexes between guanidinium derivatives and single-stranded DNA by MALDI. J. Am. Soc. Mass Spectrom. 2006, 17, 283–291. [Google Scholar] [CrossRef] [Green Version]
- Tang, K.; Fu, D.J.; Kotter, S.; Cotter, R.J.; Cantor, C.R.; Koster, H. Matrix-assisted laser desorption/ionization mass-spectrometry of immobilized duplex DNA probes. Nucleic Acids Res. 1995, 23, 3126–3131. [Google Scholar] [CrossRef]
- Hop, C.; Bakhtiar, R. An introduction to electrospray ionization and matrix-assisted laser desorption/ionization mass spectrometry: Essential tools in a modern biotechnology environment. Biospectroscopy 1997, 3, 259–280. [Google Scholar] [CrossRef]
- Gilar, M.; Belenky, A.; Wang, B.H. High-throughput biopolymer desalting by solid-phase extraction prior to mass spectrometric analysis. J. Chromatogr. A 2001, 921, 3–13. [Google Scholar] [CrossRef]
- Chifotides, H.T.; Koomen, J.M.; Kang, M.J.; Tichy, S.E.; Dunbar, K.R.; Russell, D.H. Binding of DNA purine sites to dirhodium compounds probed by mass spectrometry. Inorg. Chem. 2004, 43, 6177–6187. [Google Scholar] [CrossRef]
- Tsuchiya, Y.; Nakajima, M.; Yokoi, T. Cytochrome P450-mediated metabolism of estrogens and its regulation in human. Cancer Lett. 2005, 227, 115–124. [Google Scholar] [CrossRef]
- Nussinov, R.; Tsai, C.J.; Mattos, C. “Pathway drug cocktail”: Targeting Ras signaling based on structural pathways. Trends Mol. Med. 2013, 19, 695–704. [Google Scholar] [CrossRef]
- Farmer, P.B.; Sweetman, G.M.A. Mass-spectrometric detection of carcinogen adducts. J. Mass Spectrom. 1995, 30, 1369–1379. [Google Scholar] [CrossRef]
- Saeed, M.; Rogan, E.; Cavalieri, E. Mechanism of metabolic activation and DNA adduct formation by the human carcinogen diethylstilbestrol: The defining link to natural estrogens. Int. J. Cancer 2009, 124, 1276–1284. [Google Scholar] [CrossRef]
- Hendry, L.B.; Roach, L.W.; Mahesh, V.B. Multidimensional screening and design of pharmaceuticals by using endocrine pharmacophores. Steroids 1999, 64, 570–575. [Google Scholar] [CrossRef]
- Olmsted, S.L.; Tongcharoensirikul, P.; McCaskill, E.; Gandiaga, K.; Labaree, D.; Hochberg, R.B.; Hanson, R.N. Synthesis and evaluation of 17 alpha-E-20-(heteroaryl)norpregn-1,3,5(10),20 tetraene-3,17 beta-diols 17 alpha-(heteroaryl)vinyl estradiols as ligands for the estrogen receptor-alpha ligand binding domain (ER alpha-LBD). Bioorg. Med. Chem. Lett. 2012, 22, 977–979. [Google Scholar] [CrossRef]
- Izzotti, A.; Larghero, P.; Longobardi, M.; Cartiglia, C.; Camoirano, A.; Steele, V.E.; de Flora, S. Dose-responsiveness and persistence of microRNA expression alterations induced by cigarette smoke in mouse lung. Mutat. Res. Fundam. Mol. Mech. Mutagen. 2011, 717, 9–16. [Google Scholar] [CrossRef]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Heger, Z.; Guran, R.; Zitka, O.; Beklova, M.; Adam, V.; Kizek, R. In Vitro Interactions between 17β-Estradiol and DNA Result in Formation of the Hormone-DNA Complexes. Int. J. Environ. Res. Public Health 2014, 11, 7725-7739. https://doi.org/10.3390/ijerph110807725
Heger Z, Guran R, Zitka O, Beklova M, Adam V, Kizek R. In Vitro Interactions between 17β-Estradiol and DNA Result in Formation of the Hormone-DNA Complexes. International Journal of Environmental Research and Public Health. 2014; 11(8):7725-7739. https://doi.org/10.3390/ijerph110807725
Chicago/Turabian StyleHeger, Zbynek, Roman Guran, Ondrej Zitka, Miroslava Beklova, Vojtech Adam, and Rene Kizek. 2014. "In Vitro Interactions between 17β-Estradiol and DNA Result in Formation of the Hormone-DNA Complexes" International Journal of Environmental Research and Public Health 11, no. 8: 7725-7739. https://doi.org/10.3390/ijerph110807725
APA StyleHeger, Z., Guran, R., Zitka, O., Beklova, M., Adam, V., & Kizek, R. (2014). In Vitro Interactions between 17β-Estradiol and DNA Result in Formation of the Hormone-DNA Complexes. International Journal of Environmental Research and Public Health, 11(8), 7725-7739. https://doi.org/10.3390/ijerph110807725








