Screening and cDNA Cloning of Kv1 Potassium Channel Toxins in Sea Anemones
Abstract
:1. Introduction
2. Results and Discussion
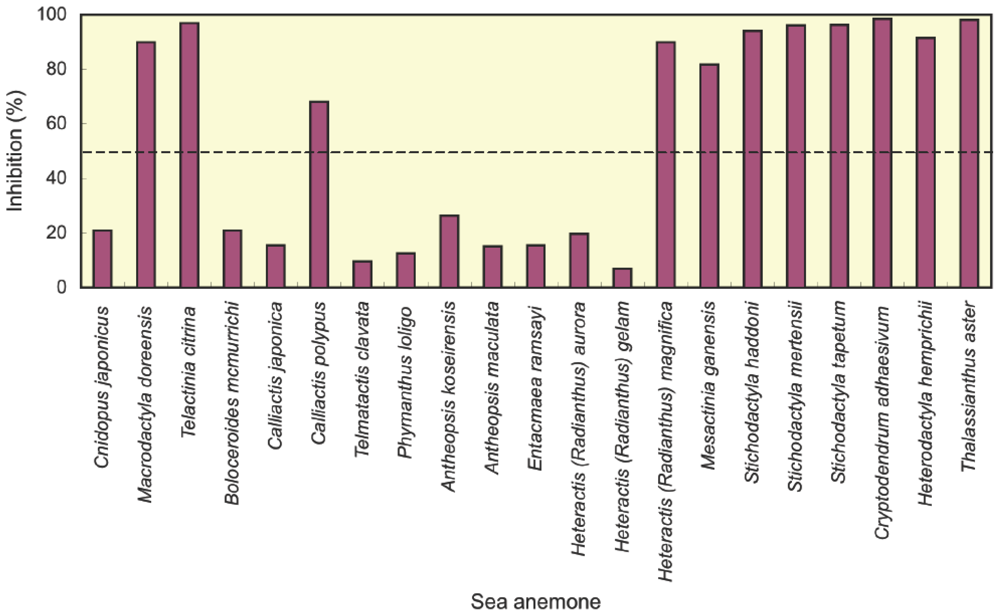
2.1. Screening of potassium channel toxins
2.2. Cloning of cDNAs encoding type 1 potassium channel toxins
2.3. Amino acid sequences of type 1 potassium channel toxins
3. Experimental Section
3.1. Sea anemone samples
3.2. Preparation of crude extracts
3.3. Assay of potassium channel toxicity
3.4. Cloning experiments
4. Conclusions
Supplementary Data
marinedrugs-08-02893s1.tifAcknowledgement
References
- Honma, T; Shiomi, K. Peptide toxins in sea anemones: structural and functional aspects. Mar Biotechnol 2006, 8, 1–10. [Google Scholar]
- Norton, RS. Structures of sea anemone toxins. Toxicon 2009, 54, 1075–1088. [Google Scholar]
- Shiomi, K. Novel peptide toxins recently isolated from sea anemones. Toxicon 2009, 54, 1112–1118. [Google Scholar]
- Norton, RS. Structure and structure-function relationships of sea anemone proteins that interact with the sodium channel. Toxicon 1991, 29, 1051–1084. [Google Scholar]
- Bosmans, F; Tytgat, J. Sea anemone venom as a source of insecticidal peptides acting on voltagegated Na+ channels. Toxicon 2007, 49, 550–560. [Google Scholar]
- Smith, JJ; Blumenthal, KM. Site-3 sea anemone toxins: molecular probes of gating mechanisms in voltage-dependent sodium channels. Toxicon 2007, 49, 159–170. [Google Scholar]
- Moran, Y; Gordon, D; Gurevitz, M. Sea anemone toxins affecting voltage-gated sodium channels – molecular and evolutionary features. Toxicon 2009, 54, 1089–1101. [Google Scholar]
- Wanke, E; Zaharenko, AJ; Redaelli, E; Schiavon, E. Actions of sea anemone type 1 neurotoxins on voltage-gated sodium channel isoforms. Toxicon 2009, 54, 1102–1111. [Google Scholar]
- Castaneda, O; Harvey, AL. Discovery and characterization of cnidarian peptide toxins that affect neuronal potassium ion channels. Toxicon 2009, 54, 1119–1124. [Google Scholar]
- King, GF; Gentz, MC; Escoubas, P; Nicholson, GM. A rational nomenclature for naming peptide toxins from spiders and other venomous animals. Toxicon 2008, 52, 264–276. [Google Scholar]
- Minagawa, S; Ishida, M; Nagashima, Y; Shiomi, K. Primary structure of a potassium channel toxin from the sea anemone Actinia equina. FEBS Lett 1998, 427, 149–151. [Google Scholar]
- Hasegawa, Y; Honma, T; Nagai, H; Ishida, M; Nagashima, Y; Shiomi, K. Isolation and cDNA cloning of a potassium channel peptide toxin from the sea anemone Anemonia erythraea. Toxicon 2006, 48, 536–542. [Google Scholar]
- Schweitz, H; Bruhn, T; Guillemare, E; Moinier, D; Lancelin, J-M; Béress, L; Lazdunski, M. Kalicludines and kaliseptine. Two different classes of sea anemone toxins for voltage-sensitive K+ channels. J Biol Chem 1995, 270, 25121–25126. [Google Scholar]
- Cotton, J; Crest, M; Bouet, F; Alessandri, N; Gola, M; Forest, E; Karlsson, E; Castaneda, O; Harvey, AL; Vita, C; Ménez, A. A potassium-channel toxin from the sea anemone Bunodosoma granulifera, an inhibitor for Kv1 channels. Revision of the amino acid sequence, disulfide-bridge assignment, chemical synthesis, and biological activity. Eur J Biochem 1997, 244, 192–202. [Google Scholar]
- Gendeh, GS; Young, LC; de Medeiros, LC; Jeyaseelan, K; Harvey, AL; Chung, MCM. A new potassium channel toxin from the sea anemone Heteractis magnifica: isolation, cDNA cloning, and functional expression. Biochemistry 1997, 36, 11461–11471. [Google Scholar]
- Castaneda, O; Sotolongo, V; Amor, AM; Stöklin, R; Anderson, AJ; Harvey, AL; Engström, Å; Wernstedt, C; Karlsson, E. Characterization of a potassium channel toxin from the Caribbean sea anemone Stichodactyla helianthus. Toxicon 1995, 33, 603–613. [Google Scholar]
- Honma, T; Kawahata, S; Ishida, M; Nagai, H; Nagashima, Y; Shiomi, K. Novel peptide toxins from the sea anemone Stichodactyla haddoni. Peptides 2008, 29, 536–544. [Google Scholar]
- Diochot, S; Schweitz, H; Béress, L; Lazdunski, M. Sea anemone peptides with a specific blocking activity against the fast inactivating potassium channel Kv3.4. J Biol Chem 1998, 273, 6744–6749. [Google Scholar]
- Diochot, S; Loret, E; Bruhn, T; Béress, L; Lazdunski, M. APETx1, a new toxin from the sea anemone Anthopleura elegantissima, blocks voltage-gated human ether-a-go-go-related gene potassium channels. Mol Pharmacol 2003, 64, 59–69. [Google Scholar]
- Zhang, M; Liu, XS; Diochot, S; Lazdunski, M; Tseng, GN. APETx1 from sea anemone Anthopleura elegantissima is a gating modifier peptide toxin of the human ether-a-go-go-related (hERG) potassium channel. Mol Pharmacol 2007, 72, 259–268. [Google Scholar]
- Harvey, AL; Rowan, EG; Vatanpour, H; Young, L; Castaneda, O; Mebs, D; Cervenansky, C; Karlsson, E. Potassium channel neurotoxins from sea anemones. In Biochemical Aspects of Marine Pharmacology; Lazarovici, P, Spira, ME, Zlotkin, E, Eds.; Alaken Inc: Fort Collins, CO, USA, 1996; pp. 121–131. [Google Scholar]
- Shiomi, K; Minagawa, S; Lin, X-Y; Yokoyama, A; Shimakura, K; Nagashima, Y. Screening of toxins and protease inhibitors in sea anemones. Fish Sci 1998, 64, 172–173. [Google Scholar]
- Harvey, AL. Twenty years of dendrotoxins. Toxicon 2001, 39, 15–26. [Google Scholar]
- Bendtsen, JD; Nielsen, H; von Heijne, G; Brunak, S. Improved prediction of signal peptides: SignalP 3.0. J Mol Biol 2004, 340, 783–795. [Google Scholar]
- Anderluh, G; Podlesek, Z; Macek, P. A common motif in proparts of Cnidarian toxins and nematocyst collagens and its putative role. Biochim Biophys Acta 2000, 1476, 372–376. [Google Scholar]
- Honma, T; Nagai, H; Nagashima, Y; Shiomi, K. Molecular cloning of an epidermal growth factor-like toxin and two sodium channel toxins from the sea anemone Stichodactyla gigantea. Biochim Biophys Acta 2003, 1652, 103–106. [Google Scholar]
- Honma, T; Hasegawa, Y; Ishida, M; Nagai, H; Nagashima, Y; Shiomi, K. Isolation and molecular cloning of novel peptide toxins from the sea anemone Antheopsis maculata. Toxicon 2005, 45, 33–41. [Google Scholar]
- Pohl, J; Hubalek, F; Byrnes, ME; Nielsen, KR; Woods, A; Pennington, MW. Assignment of the three disulfide bonds in ShK toxin: A potent potassium channel inhibitor from the sea anemone Stichodactyla helianthus. Lett Pept Sci 1995, 1, 291–297. [Google Scholar]
- Pennington, MW; Mahnir, VM; Krafte, DS; Zaydenberg, I; Byrnes, ME; Khaytin, I; Crowley, K; Kem, WR. Identification of three separate binding sites on SHK toxin, a potent inhibitor of voltage-dependent potassium channels in human T-lymphocytes and rat brain. Biochem Biophys Res Commun 1996, 219, 696–701. [Google Scholar]
- Dauplais, M; Lecoq, A; Song, J; Cotton, J; Jamin, N; Gilquin, B; Roumestand, C; Vita, C; de Medeiros, CL; Rowan, EG; Harvey, AL; Ménez, A. On the convergent evolution of animal toxins. Conservation of a dyad of functional residues in potassium channel-blocking toxins with unrelated structures. J Biol Chem 1997, 272, 4302–4309. [Google Scholar]
- Alessandri-Haber, N; Lecoq, A; Gasparini, S; Grangier-Macmath, G; Jacquet, G; Harvey, AL; de Medeiros, C; Rowan, EG; Gola, M; Ménez, A; Crest, M. Mapping the functional anatomy of BgK on Kv1.1, Kv1.2, and Kv1.3. Clues to design analogs with enhanced selectivity. J Biol Chem 1999, 274, 35653–35661. [Google Scholar]
- Gilquin, B; Racape, J; Wrisch, A; Visan, V; Lecoq, A; Grissmer, S; Ménez, A; Gasparini, S. Structure of the BgK-Kv1.1 complex based on distance restraints identified by double mutant cycles. Molecular basis for convergent evolution of Kv1 channel blockers. J Biol Chem 2002, 277, 37406–37413. [Google Scholar]
- Harvey, AL; Marshall, DL; De-Allie, FA; Strong, PN. Interactions between dendrotoxin, a blocker of voltage-dependent potassium channels, and charybdotoxin, a blocker of calciumactivated potassium channels, at binding sites on neuronal membranes. Biochem Biophys Res Commun 1989, 163, 394–397. [Google Scholar]


| Sea anemone | Potassium channel toxin | Reference | ||
|---|---|---|---|---|
| Suborder | Family | Species | ||
| Endocoelantheae | Halcuriidae | Halcurias sp. | − | [22] |
| Nynantheae | Actiniidae | Actinia bermudensis | + | [21] |
| Actinia equina | + | [11,22] | ||
| Adamsia maculata | − | [21] | ||
| Alcyonium digitatum | − | [21] | ||
| Anemonia erythraea | + | [12,22] | ||
| Anemonia sulcata (viridis) | + | [13,21] | ||
| Anthopleura asiatica | − | [22] | ||
| Anthopleura japonica | − | [22] | ||
| Anthopleura pacifica | − | [22] | ||
| Anthopleura xanthogrammica | − | [22] | ||
| Bunodosoma cangicum | + | [21] | ||
| Bunodosoma granulifera | + | [14,21] | ||
| Cnidopus japonicus | − | This study | ||
| Condylactis passiflora | − | [22] | ||
| Dofleinia armata | − | [22] | ||
| Macrodactyla doreensis | + | This study | ||
| Tealia coriacea | − | [22] | ||
| Telactinia citrina | + | This study | ||
| Actinodendronidae | Actinodendron plumosum (arboreum) | − | [22] | |
| Boloceroididae | Boloceroides mcmurrichi | − | This study | |
| Diadumenidae | Haliplanella lineata | − | [22] | |
| Hormathiidae | Calliactis japonica | − | This study | |
| Calliactis polypus | + | This study | ||
| Isophellidae | Telmatactis clavata | − | This study | |
| Metridiidae | Metridium senile | − | [22] | |
| Nemanthidae | Nemanthus nitidus | − | [22] | |
| Phymanthidae | Phymanthus loligo | − | This study | |
| Stichodactylidae | Antheopsis koseirensis | − | This study | |
| Antheopsis maculata | − | This study | ||
| Entacmaea actinostoloides | − | [22] | ||
| Entacmaea ramsayi | − | This study | ||
| Heteractis (Radianthus) aurora | − | This study | ||
| Heteractis (Radianthus) crispus | − | [22] | ||
| Heteractis (Radianthus) gelam | − | This study | ||
| Heteractis (Radianthus) magnifica | + | [15], This study | ||
| Mesactinia ganensis | + | This study | ||
| Stichodactyla gigantea | + | This study a | ||
| Stichodactyla haddoni | + | [17], This study | ||
| Stichodactyla helianthus | + | [16,21] | ||
| Stichodactyla mertensii | + | [21], This study | ||
| Stichodactyla tapetum | + | This study | ||
| Thalassianthidae | Cryptodendrum adhaesivum | + | This study | |
| Heterodactyla hemprichii | + | This study | ||
| Thalassianthus aster | + | This study | ||
| Experiment | Designation of primer | Nucleotide sequence of primer |
|---|---|---|
| RT-PCR | RT-f | 5′-GATCGCAYTTTGTMTGTSTGT-3′ |
| RT-r | 5′-TGTATTTCATTGACGTYCTACA-3′ | |
| 3′RACE | 3′-Sha-f | 5′-ACGGTTTCCAAAAGAGGAGG-3′ |
| 3′-Ca-f | 5′-GAACTGCAGGATGTCGAGGA-3′ | |
| 3′-Hh-f | 5′-GAATTGCAGGATGACGAGGA-3′ | |
| AUAP | 5′-GGCCACGCGTCGACTAGTAC-3′ | |
| 5′RACE | 5′-Syn-SgSha-r | 5′-ACAAAAGCTCAGTCTGTATTTC-3′ |
| 5′-Syn-Sm-r | 5′-ACAGAAGCTAAGTCTGTATTTC-3′ | |
| 5′-Syn-CaHhTa-r | 5′-TTGAAATGCTGTGCATCGGC-3′ | |
| 5′-Sg-r | 5′-TCCTGCAGTTCCATCCTGG-3′ | |
| 5′-ShaSm-r | 5′-CCTCCTCTTTTGGAAACCGT-3′ | |
| 5′-Ca-r | 5′-GTCCTCCTCTTTTGGAAAGA-3′ | |
| 5′-HhTa-r | 5′-GTGCATCGGCTCTTCGGTA-3′ | |
| AUAP | 5′-GGCCACGCGTCGACTAGTAC-3′ | |
| AAP | 5′-GGCCACGCGTCGACTAGTACGGGGGGGGGGGGGGGG-3′ |
© 2010 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Yamaguchi, Y.; Hasegawa, Y.; Honma, T.; Nagashima, Y.; Shiomi, K. Screening and cDNA Cloning of Kv1 Potassium Channel Toxins in Sea Anemones. Mar. Drugs 2010, 8, 2893-2905. https://doi.org/10.3390/md8122893
Yamaguchi Y, Hasegawa Y, Honma T, Nagashima Y, Shiomi K. Screening and cDNA Cloning of Kv1 Potassium Channel Toxins in Sea Anemones. Marine Drugs. 2010; 8(12):2893-2905. https://doi.org/10.3390/md8122893
Chicago/Turabian StyleYamaguchi, Yoshikazu, Yuichi Hasegawa, Tomohiro Honma, Yuji Nagashima, and Kazuo Shiomi. 2010. "Screening and cDNA Cloning of Kv1 Potassium Channel Toxins in Sea Anemones" Marine Drugs 8, no. 12: 2893-2905. https://doi.org/10.3390/md8122893
APA StyleYamaguchi, Y., Hasegawa, Y., Honma, T., Nagashima, Y., & Shiomi, K. (2010). Screening and cDNA Cloning of Kv1 Potassium Channel Toxins in Sea Anemones. Marine Drugs, 8(12), 2893-2905. https://doi.org/10.3390/md8122893





