Structure of a Sulfated Capsular Polysaccharide from the Marine Bacterium Cobetia marina KMM 1449 and a Genomic Insight into Its Biosynthesis
Abstract
1. Introduction
2. Results
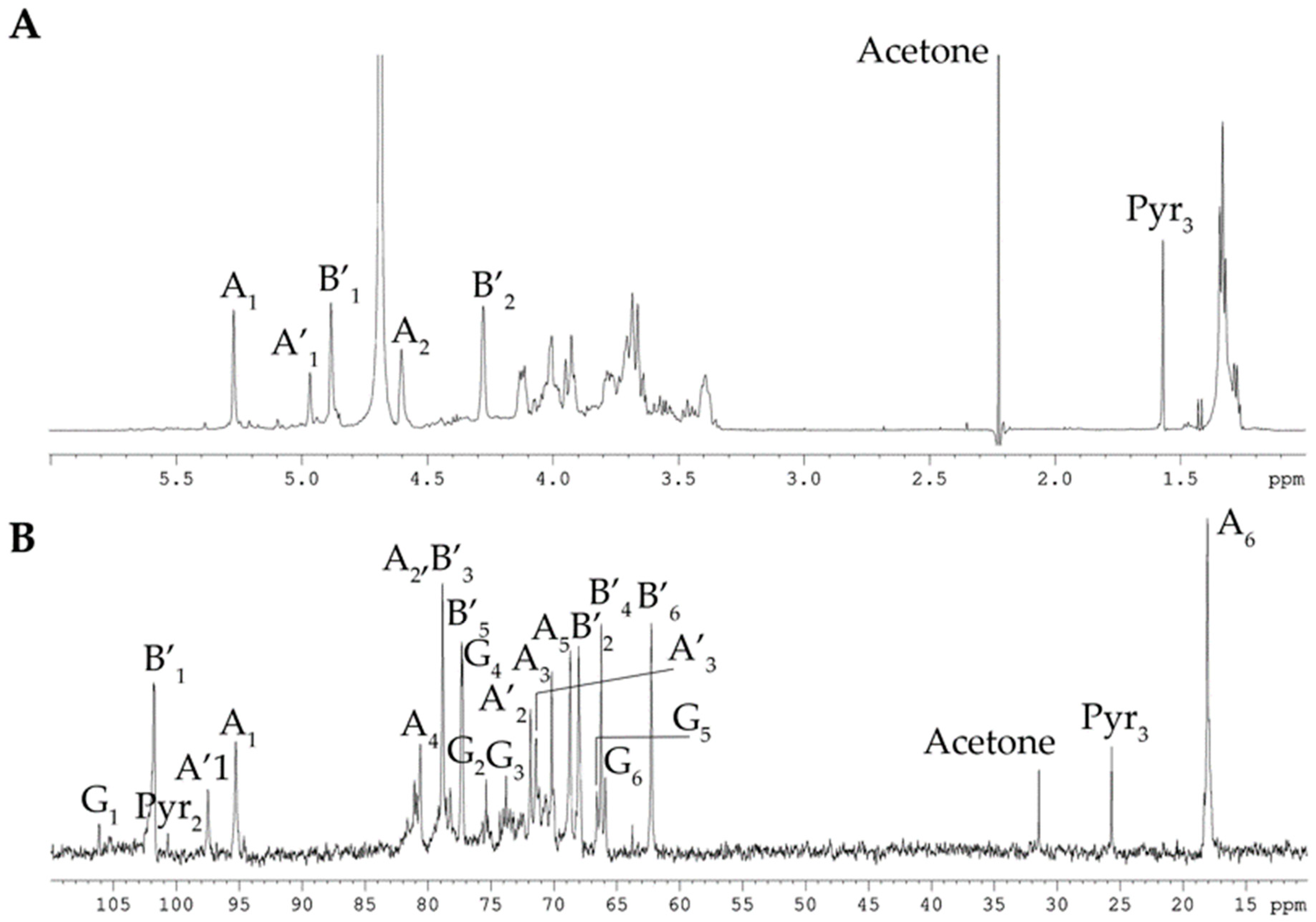
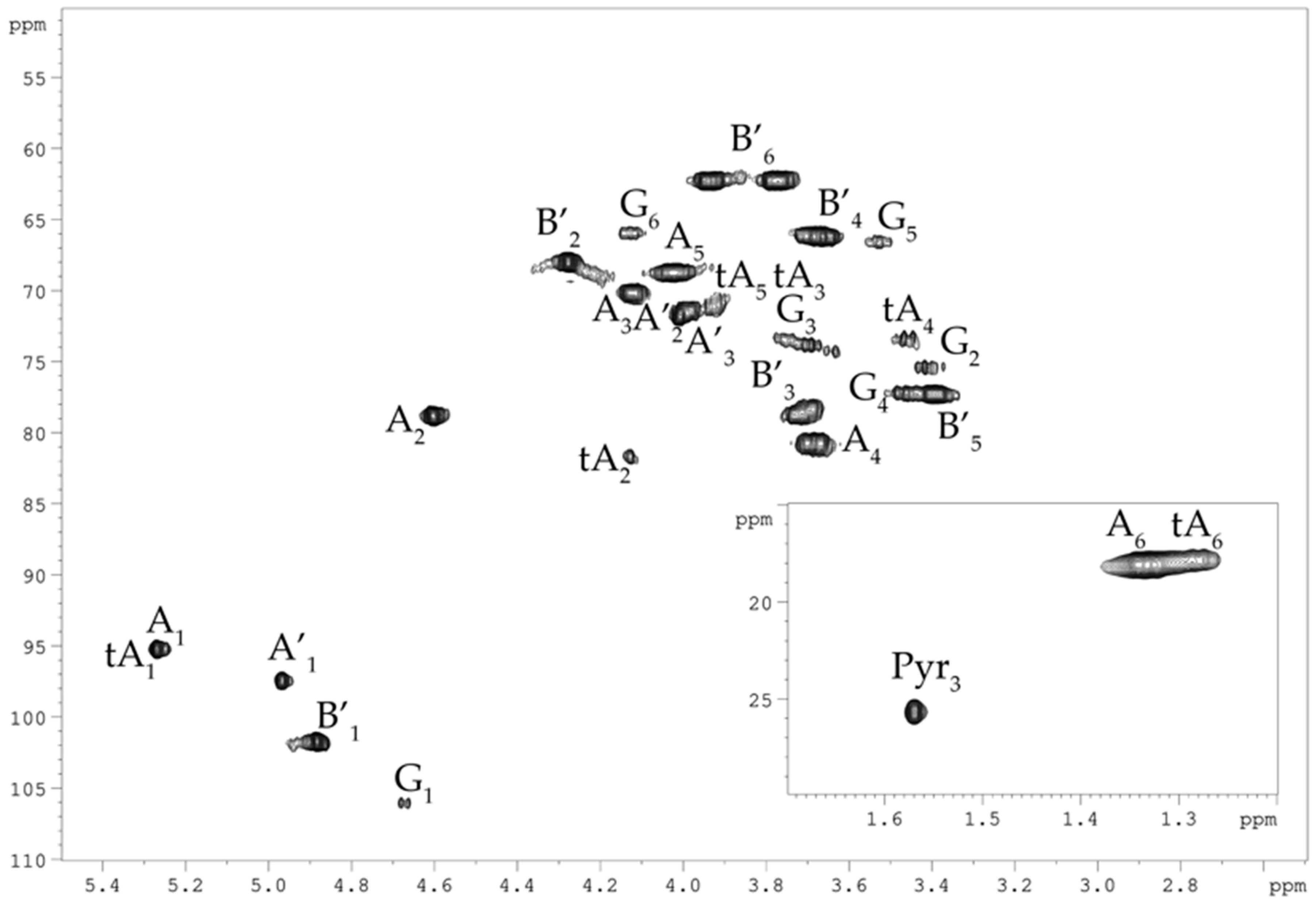
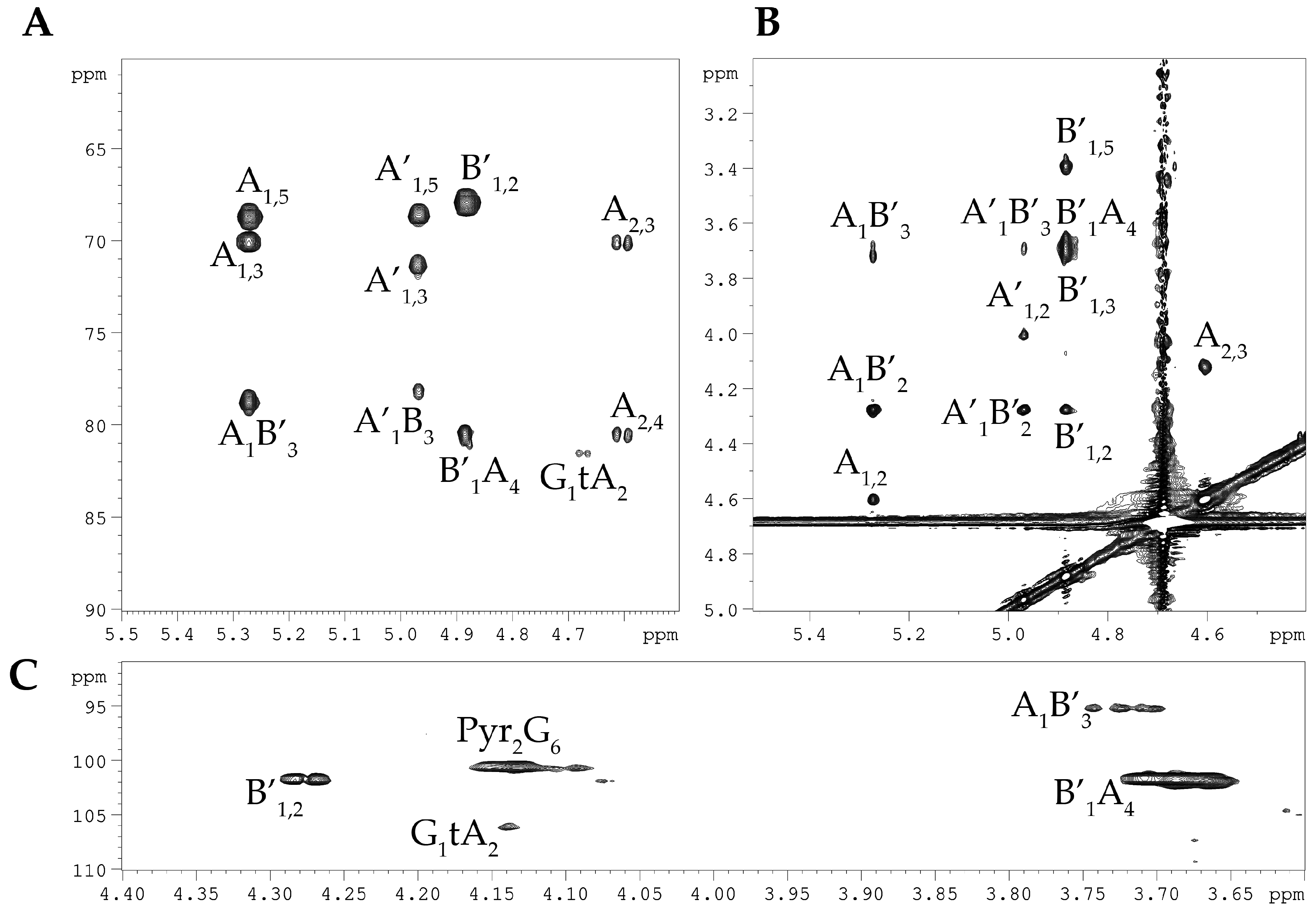
2.1. Isolation and Structural Analyses of the CPS
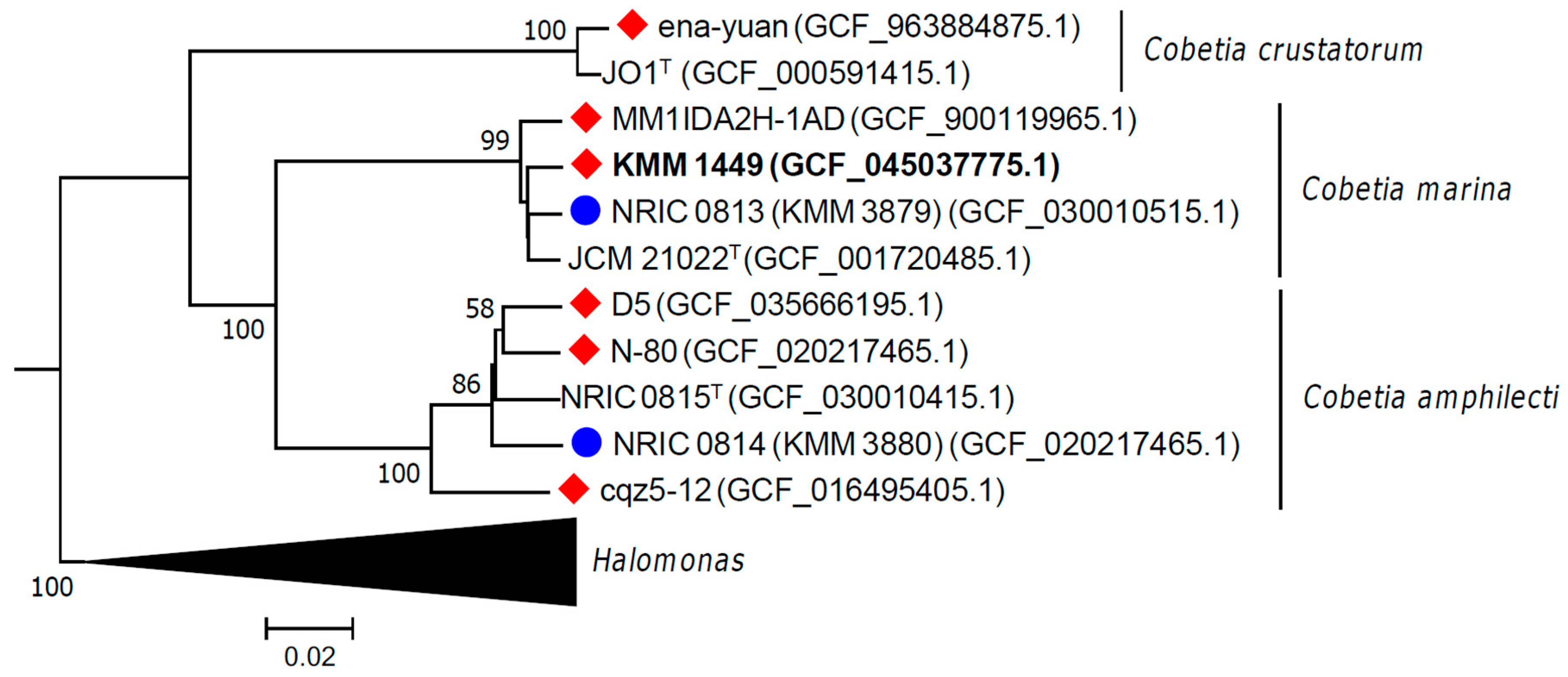
2.2. Phylogenetic and Genomic Analyses
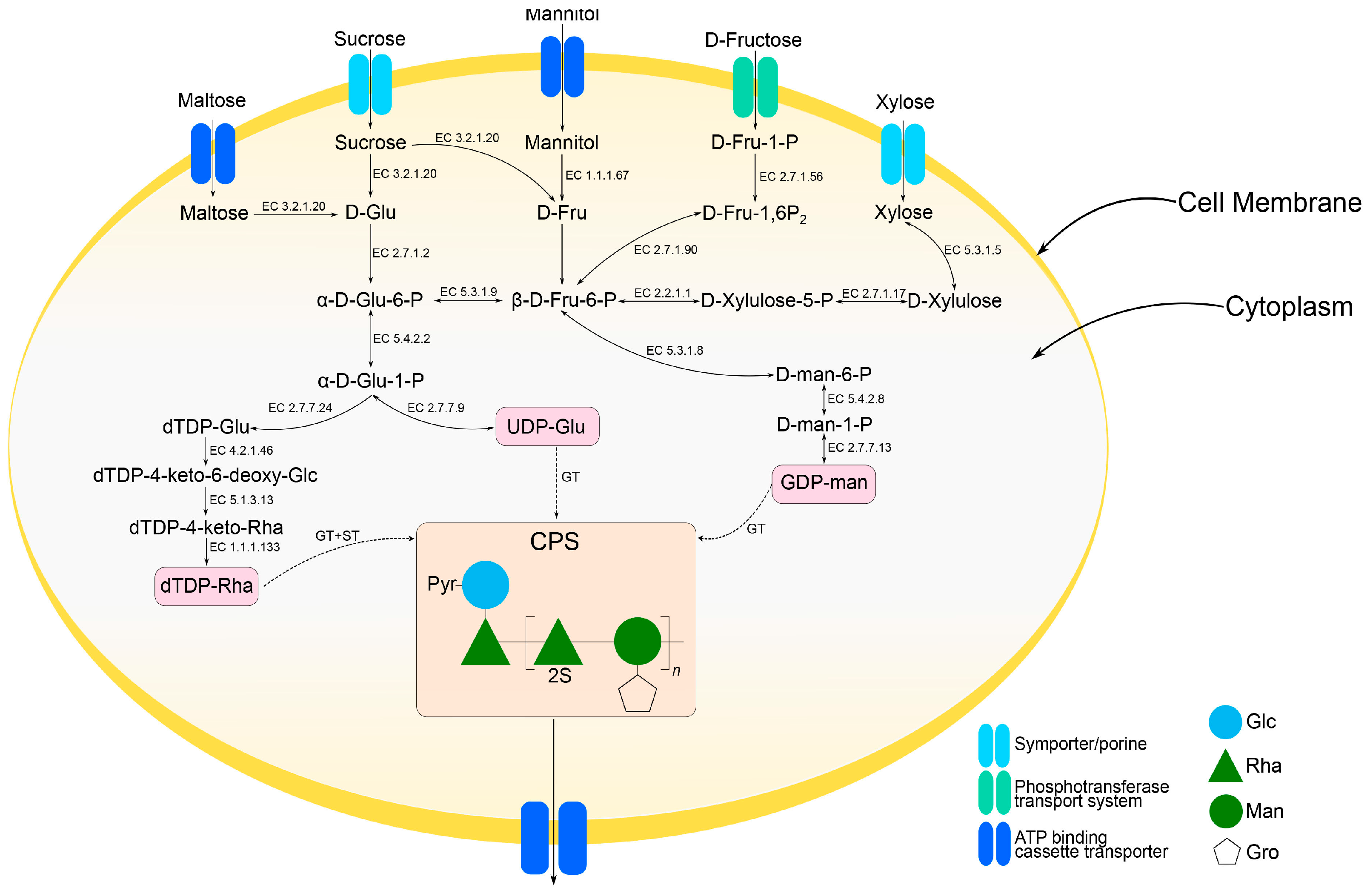
2.3. Predicted CPS Gene Cluster and Biosynthesis
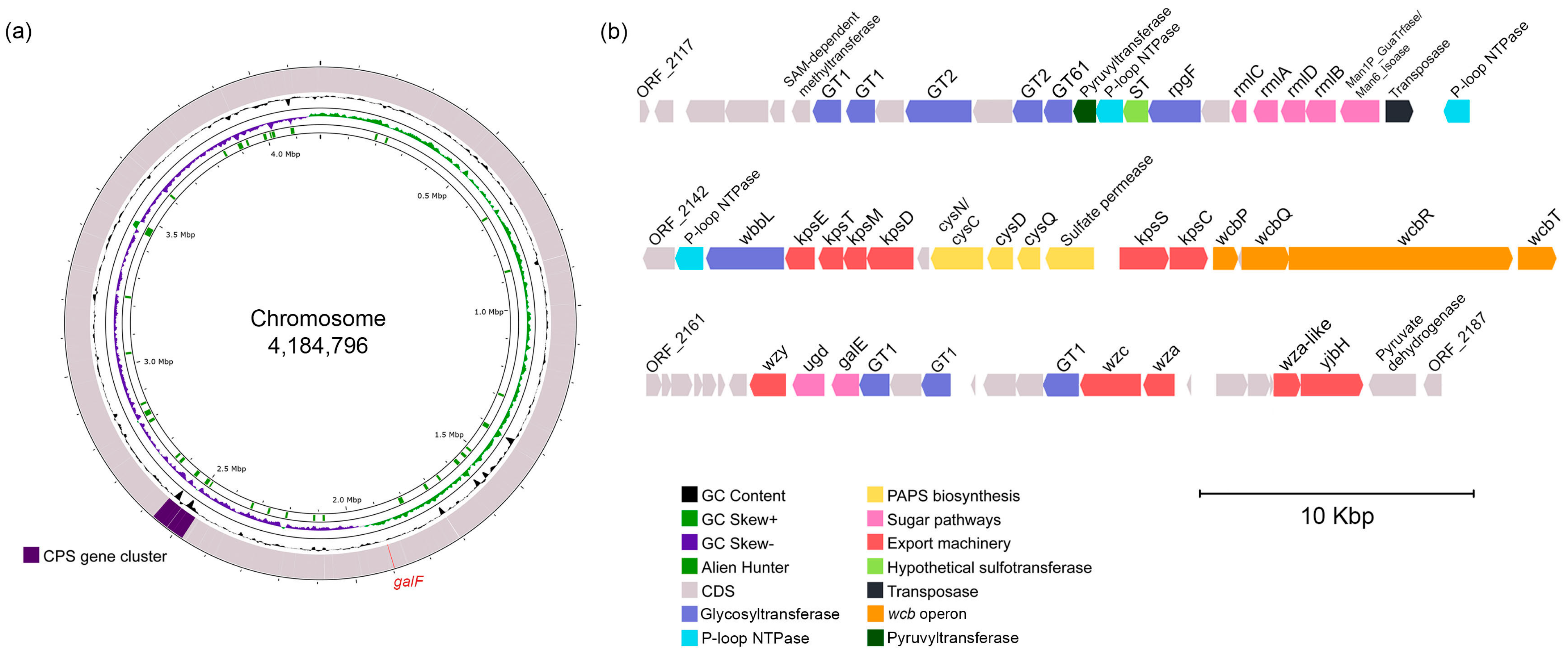
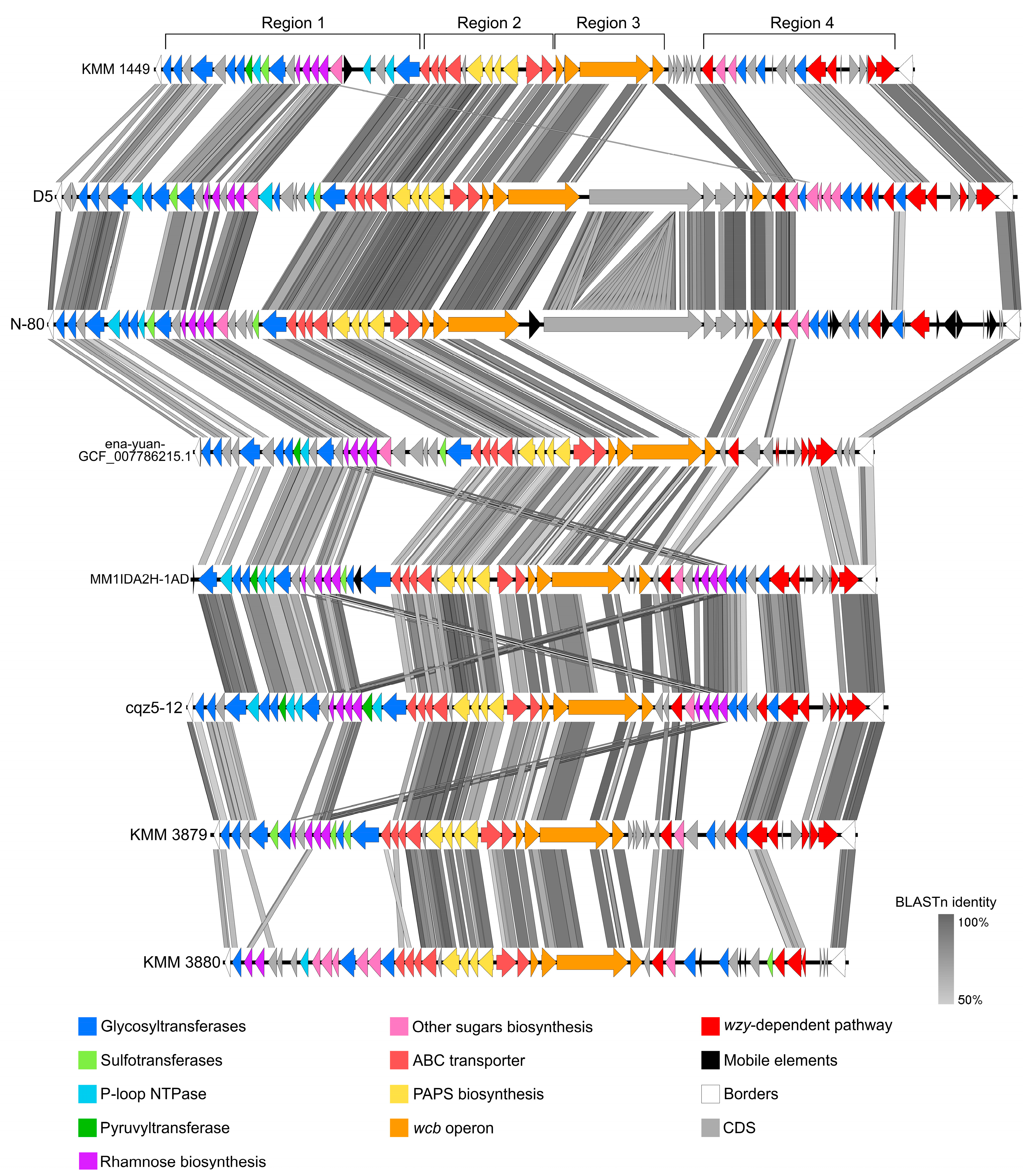
2.4. Comparative Analysis of CPS Gene Clusters Between Cobetia Strains
3. Materials and Methods
3.1. Isolation and Purification of CPS
3.2. Compositional Analysis of CPS Samples
3.3. Dephosphorylation of CPS
3.4. NMR and FTIR Spectroscopy
3.5. Whole-Genome Sequencing, Assembly, and Annotation
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Hoshino, T.; Doi, H.; Uramoto, G.I.; Wömer, L.; Adhikari, R.R.; Xiao, N. Global diversity of microbial communities in marine sediment. Proc. Natl. Acad. Sci. USA 2020, 117, 27587–27597. [Google Scholar] [CrossRef] [PubMed]
- Elsakhawy, T.A.; Sherief, F.A.; Abd-EL-Kodoos, R.Y. Marine microbial polysaccharides: Environmental role and applications (an overview). Env. Biodivers. Soil Secur. 2017, 1, 61–70. [Google Scholar] [CrossRef]
- Casillo, A.; Lanzetta, R.; Parrilli, M.; Corsaro, M.M. Exopolysaccharides from marine and marine extremophilic bacteria: Structures, properties, ecological roles, and applications. Mar. Drugs 2018, 16, 69. [Google Scholar] [CrossRef] [PubMed]
- Herget, S.; Toukach, P.V.; Ranzinger, R.; Hull, W.E.; Knirel, Y.A.; Von der Lieth, C.-W. Statistical analysis of the Bacterial Carbohydrate Structure Data Base (BCSDB): Characteristics and diversity of bacterial carbohydrates in comparison with mammalian glycans. BMC Struct. Biol. 2008, 8, 35. [Google Scholar] [CrossRef] [PubMed]
- Poli, A.; Anzelmo, G.; Nicolaus, B. Bacterial exopolysaccharides from extreme marine habitats: Production, characterization, and biological activities. Mar. Drugs 2010, 8, 1779–1802. [Google Scholar] [CrossRef] [PubMed]
- Laurienzo, P. Marine polysaccharides in pharmaceutical applications: An overview. Mar. Drugs 2010, 8, 2435–2465. [Google Scholar] [CrossRef] [PubMed]
- Rühmann, B.; Schmid, J.; Sieber, V. Methods to identify the unexplored diversity of microbial exopolysaccharides. Front. Microbiol. 2015, 6, 565. [Google Scholar] [CrossRef]
- Nedashkovskaya, O.; Balabanova, L.; Otstavnykh, N.; Zhukova, N.; Detkova, E.; Seitkalieva, A.; Bystritskaya, E.; Noskova, Y.; Tekutyeva, L.; Isaeva, M. In-Depth Genome Characterization and Pan-Genome Analysis of Strain KMM 296: Proposal for the Reclassification of Cobetia litoralis and Cobetia pacifica as Later Heterotypic Synonyms of Cobetia amphilecti and Cobetia marina, and Emended Description of the Species Cobetia amphilecti and Cobetia marina. Biomolecules 2024, 14, 196. [Google Scholar] [CrossRef]
- Lelchat, F.; Cérantola, S.; Brandily, C.; Colliec-Jouault, S.; Baudoux, A.-C.; Ojima, T.; Boisset, C. The marine bacteria Cobetia marina DSMZ 4741 synthesizes an unexpected K-antigen-like exopolysaccharide. Carbohydr. Polym. 2015, 124, 347–356. [Google Scholar] [CrossRef] [PubMed]
- Kokoulin, M.S.; Kalinovsky, A.I.; Komandrova, N.A.; Tomshich, S.V.; Romanenko, L.A.; Vaskovsky, V.E. The sulfated O-specific polysaccharide from the marine bacterium Cobetia pacifica KMM 3879T. Carbohydr. Res. 2014, 387, 4–9. [Google Scholar] [CrossRef]
- Kokoulin, M.S.; Kalinovsky, A.I.; Komandrova, N.A.; Tomshich, S.V.; Romanenko, L.A.; Vaskovsky, V.E. The new sulfated O-specific polysaccharide from marine bacterium Cobetia pacifica KMM 3878, containing 3,4-O-[(S)-1-carboxyethylidene]-D-galactose and 2,3-O-disulfate-D-galactose. Carbohydr. Res. 2014, 397, 46–51. [Google Scholar] [CrossRef] [PubMed]
- Kokoulin, M.S.; Kuzmich, A.S.; Kalinovsky, A.I.; Tomshich, S.V.; Romanenko, L.A.; Mikhailov, V.V.; Komandrova, N.A. Structure and anticancer activity of sulfated O-polysaccharide from marine bacterium Cobetia litoralis KMM 3880T. Carbohydr. Polym. 2016, 154, 55–61. [Google Scholar] [CrossRef] [PubMed]
- Fedorov, S.N.; Ermakova, S.P.; Zvyagintseva, T.N.; Stonik, V.A. Anticancer and cancer preventive properties of marine polysaccharides: Some results and prospects. Mar. Drugs 2013, 11, 4876–4901. [Google Scholar] [CrossRef]
- Kokoulin, M.S.; Kuzmich, A.S.; Filshtein, A.P.; Prassolov, V.S.; Romanenko, L.A. Capsular polysaccharide from the marine bacterium Cobetia marina induces apoptosis via both caspase-dependent and mitochondria-mediated pathways in HL-60 cells. Carbohydr. Polym. 2025, 347, 122791. [Google Scholar] [CrossRef] [PubMed]
- Kokoulin, M.S.; Sigida, E.N.; Kuzmich, A.S.; Ibrahim, I.M.; Fedonenko, Y.P.; Konnova, S.A. Structure and antiproliferative activity of the polysaccharide from Halomonas aquamarina related to Cobetia pacifica. Carbohydr. Polym. 2022, 298, 120125. [Google Scholar] [CrossRef]
- Leontein, K.; Lindberg, B.; Lönngren, J. Assignment of absolute configuration of sugars by G.L.C. of their acetylated glycosides formed from chiral alcohols. Carbohydr. Res. 1978, 62, 359–362. [Google Scholar] [CrossRef]
- Garegg, P.J.; Jansson, P.-E.; Lindberg, B.; Lindh, F.; Lönngren, J.; Kvarnstrom, I.; Nimmich, W. Configuration of the acetal carbon atom of pyruvic acid acetals in some bacterial polysaccharides. Carbohydr. Res. 1980, 78, 127–132. [Google Scholar] [CrossRef]
- Shashkov, A.S.; Tul’skaya, E.M.; Evtushenko, L.I.; Naumova, I.B. A teichoic acid of Nocardioides albus VKM Ac-805T cell walls. Biochemistry 1999, 64, 1305–1309. [Google Scholar] [PubMed]
- Lefort, V.; Desper, R.; Gascuel, O. FastME 2.0: A comprehensive, accurate and fast distance-based phylogeny inference program. Mol. Biol. Evol. 2015, 32, 2798–2800. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.; Oh, H.S.; Park, S.C.; Chun, J. Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int. J. Syst. Evol. Microbiol. 2014, 64, 346–351. [Google Scholar] [CrossRef] [PubMed]
- Richter, M.; Rosselló-Móra, R. Shifting the genomic gold standard for the prokaryotic species definition. Proc. Natl. Acad. Sci. USA 2009, 106, 19126–19131. [Google Scholar] [CrossRef] [PubMed]
- Clarke, B.R.; Pearce, R.; Roberts, I.S. Genetic organization of the Escherichia coli K10 capsule gene cluster: Identification and characterization of two conserved regions in group III capsule gene clusters encoding polysaccharide transport functions. J. Bacteriol. 1999, 181, 2279–2285. [Google Scholar] [CrossRef]
- Hong, Y.; Qin, J.; Forga, X.B.; Totsika, M. Extensive diversity in Escherichia coli group 3 capsules is driven by recombination and plasmid transfer from multiple species. Microbiol. Spectr. 2023, 11, e0143223. [Google Scholar] [CrossRef] [PubMed]
- Grant, J.R.; Enns, E.; Marinier, E.; Mandal, A.; Herman, E.K.; Chen, C.-Y.; Graham, M.; Van Domselaar, G.; Stothard, P. Proksee: In-depth characterization and visualization of bacterial genomes. Nucleic Acids Res. 2023, 51, W484–W492. [Google Scholar] [CrossRef] [PubMed]
- Kanehisa, M.; Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000, 28, 27–30. [Google Scholar] [CrossRef]
- Moriya, Y.; Itoh, M.; Okuda, S.; Yoshizawa, A.; Kanehisa, M. KAAS: An automatic genome annotation and pathway reconstruction server. Nucleic Acids Res. 2007, 35, W182–W185. [Google Scholar] [CrossRef] [PubMed]
- Willis, L.M.; Whitfield, C. KpsC and KpsS are retaining 3-deoxy-D-manno-oct-2-ulosonic acid (Kdo) transferases involved in synthesis of bacterial capsules. Proc. Natl. Acad. Sci. USA 2013, 110, 20753–20758. [Google Scholar] [CrossRef] [PubMed]
- Gay, S.C.; Fribourgh, J.L.; Donohoue, P.D.; Segel, I.H.; Fisher, A.J. Kinetic properties of ATP sulfurylase and APS kinase from Thiabacillus denitrificans. Arch. Biochem. Biophys. 2009, 489, 110–117. [Google Scholar] [CrossRef] [PubMed]
- Landi, S.; Esposito, S. Bioinformatic characterization of sulfotransferase provides new insights for the exploitation of sulfated polysaccharides in Caulerpa. Int. J. Mol. Sci. 2020, 21, 6681. [Google Scholar] [CrossRef] [PubMed]
- Mistry, R.; Byrne, D.P.; Starns, D.; Barsukov, I.L.; Yates, E.A.; Fernig, D.G. Polysaccharide sulfotransferases: The identification of putative sequences and respective functional characterization. Essays Biochem. 2024, 68, 431–447. [Google Scholar] [PubMed]
- Cuccui, J.; Milne, T.S.; Harmer, N.; George, A.J.; Harding, S.V.; Dean, R.E.; Scott, A.E.; Sarkar-Tyson, M.; Wren, B.W.; Titball, R.W.; et al. Characterization of the Burkholderia pseudomallei K96243 capsular polysaccharide I coding region. Infect Immun. 2012, 80, 1209–1218. [Google Scholar] [CrossRef]
- Warawa, J.M.; Long, D.; Rosenke, R.; Gardner, D.; Gherardini, F.C. Role for the Burkholderia pseudomallei capsular polysaccharide encoded by the wcb operon in acute disseminated melioidosis. Infect Immun. 2009, 77, 5252–5261. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Zakharova, I.B.; Vasilyeva, K.V.; Bui, L.A.T.; Victorov, A.D.; Shpak, I.M.; Viktorov, D.V. Genetic and morphological diversity of the Vietnamese population of Burkholderia thailandensis. Mol. Genet. Microbiol. Virol. 2022, 37, 34–46. [Google Scholar] [CrossRef]
- Edgar, R.J.; van Hensbergen, V.P.; Ruda, A.; Turner, A.G.; Deng, P.; Le Breton, Y.; El-Sayed, N.M.; Belew, A.T.; McIver, K.S.; McEwan, A.G.; et al. Discovery of glycerol phosphate modification on streptococcal rhamnose polysaccharides. Nat. Chem. Biol. 2019, 15, 463–471. [Google Scholar] [CrossRef] [PubMed]
- Ren, K.; Li, Y.; Shi, F.; Wang, X. Separation of lipopolysaccharides containing different fatty acid chains using hydrophobic interaction chromatography. Anal. Methods 2012, 4, 838. [Google Scholar] [CrossRef]
- Kolmogorov, M.; Yuan, J.; Lin, Y.; Pevzner, P.A. Assembly of long, error-prone reads using repeat graphs. Nat. Biotechnol. 2019, 37, 540–546. [Google Scholar] [CrossRef]
- Parks, D.H.; Imelfort, M.; Skennerton, C.T.; Hugenholtz, P.; Tyson, G.W. CheckM: Assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 2015, 25, 1043–1055. [Google Scholar] [CrossRef] [PubMed]
- Tatusova, T.; DiCuccio, M.; Badretdin, A.; Chetvernin, V.; Nawrocki, E.P.; Zaslavsky, L.; Lomsadze, A.; Pruitt, K.D.; Borodovsky, M.; Ostell, J. NCBI prokaryotic genome annotation pipeline. Nucleic Acids Res. 2016, 44, 6614–6624. [Google Scholar] [CrossRef] [PubMed]
- Aziz, R.K.; Bartels, D.; Best, A.A.; DeJongh, M.; Disz, T.; Edwards, R.A.; Formsma, K.; Gerdes, S.; Glass, E.M.; Kubal, M.; et al. The RAST server: Rapid annotations using subsystems technology. BMC Genom. 2008, 9, 75. [Google Scholar] [CrossRef] [PubMed]
- Seemann, T. Prokka: Rapid prokaryotic genome annotation. Bioinformatics 2014, 30, 2068–2069. [Google Scholar] [CrossRef] [PubMed]
- Cantalapiedra, C.P.; Hernández-Plaza, A.; Letunic, I.; Bork, P.; Huerta-Cepas, J. eggNOG-mapper v2: Functional annotation, orthology assignments and domain prediction at the metagenomic scale. Mol. Biol. Evol. 2021, 38, 5825–5829. [Google Scholar] [CrossRef] [PubMed]
- Paysan-Lafosse, T.; Blum, M.; Chuguransky, S.; Grego, T.; Lázaro Pinto, B.; Salazar, G.A.; Bileschi, M.L.; Bork, P.; Bridge, A.; Colwell, L.; et al. InterPro in 2022. Nucleic Acids Res. 2023, 51, D418–D427. [Google Scholar] [CrossRef]
- Coudert, E.; Gehant, S.; de Castro, E.; Pozzato, M.; Baratin, D.; Neto, T.; Sigrist, C.J.A.; Redaschi, N.; Bridge, A.; UniProt Consortium. Annotation of biologically relevant ligands in UniProtKB using ChEBI. Bioinformatics 2023, 39, btac793. [Google Scholar] [CrossRef] [PubMed]
- Blin, K.; Shaw, S.; Augustijn, H.E.; Reitz, Z.L.; Biermann, F.; Alanjary, M.; Fetter, A.; Terlouw, B.R.; Metcalf, W.W.; Helfrich, E.J.N.; et al. antiSMASH 7.0: New and improved predictions for detection, regulation, chemical structures and visualization. Nucleic Acids Res. 2023, 51, W46–W50. [Google Scholar] [CrossRef] [PubMed]
- Chalita, M.; Kim, Y.O.; Park, S.; Oh, H.S.; Cho, J.H.; Moon, J.; Baek, N.; Moon, C.; Lee, K.; Yang, J.; et al. EzBioCloud: A genome-driven database and platform for microbiome identification and discovery. Int. J. Syst. Evol. Microbiol. 2024, 74, 006421. [Google Scholar] [CrossRef] [PubMed]
- Meier-Kolthoff, J.P.; Göker, M. TYGS is an automated high-throughput platform for state-of-the-art genome-based taxonomy. Nat. Commun. 2019, 10, 2182. [Google Scholar] [CrossRef]
- Sullivan, M.J.; Petty, N.K.; Beatson, S.A. Easyfig: A genome comparison visualizer. Bioinformatics 2011, 27, 1009–1010. [Google Scholar] [CrossRef]
- Matsuda, M.; Yamori, T.; Naitoh, M.; Okutani, K. Structural revision of sulfated polysaccharide B-1 isolated from a marine Pseudomonas species and its cytotoxic activity against human cancer cell lines. Mar. Biotechnol. 2003, 5, 13–19. [Google Scholar] [CrossRef]
- Kokoulin, M.S.; Kuzmich, A.S.; Romanenko, L.A.; Chikalovets, I.V. Sulfated capsular polysaccharide from the marine bacterium Kangiella japonica inhibits T-47D cells growth in vitro. Carbohydr Polym. 2022, 290, 119477. [Google Scholar] [CrossRef] [PubMed]
- Kiselevskiy, M.V.; Anisimova, N.Y.; Ustyuzhanina, N.E.; Vinnitskiy, D.Z.; Tokatly, A.I.; Reshetnikova, V.V.; Chikileva, I.O.; Shubina, I.Z.; Kirgizov, K.I.; Nifantiev, N.E. Perspectives for the use of fucoidans in clinical oncology. Int. J. Mol. Sci. 2022, 23, 11821. [Google Scholar] [CrossRef] [PubMed]
- Toukach, P. Carbohydrate structure database: Current state and recent developments. Anal. Bioanal. Chem. 2024, 1–10. [Google Scholar] [CrossRef] [PubMed]
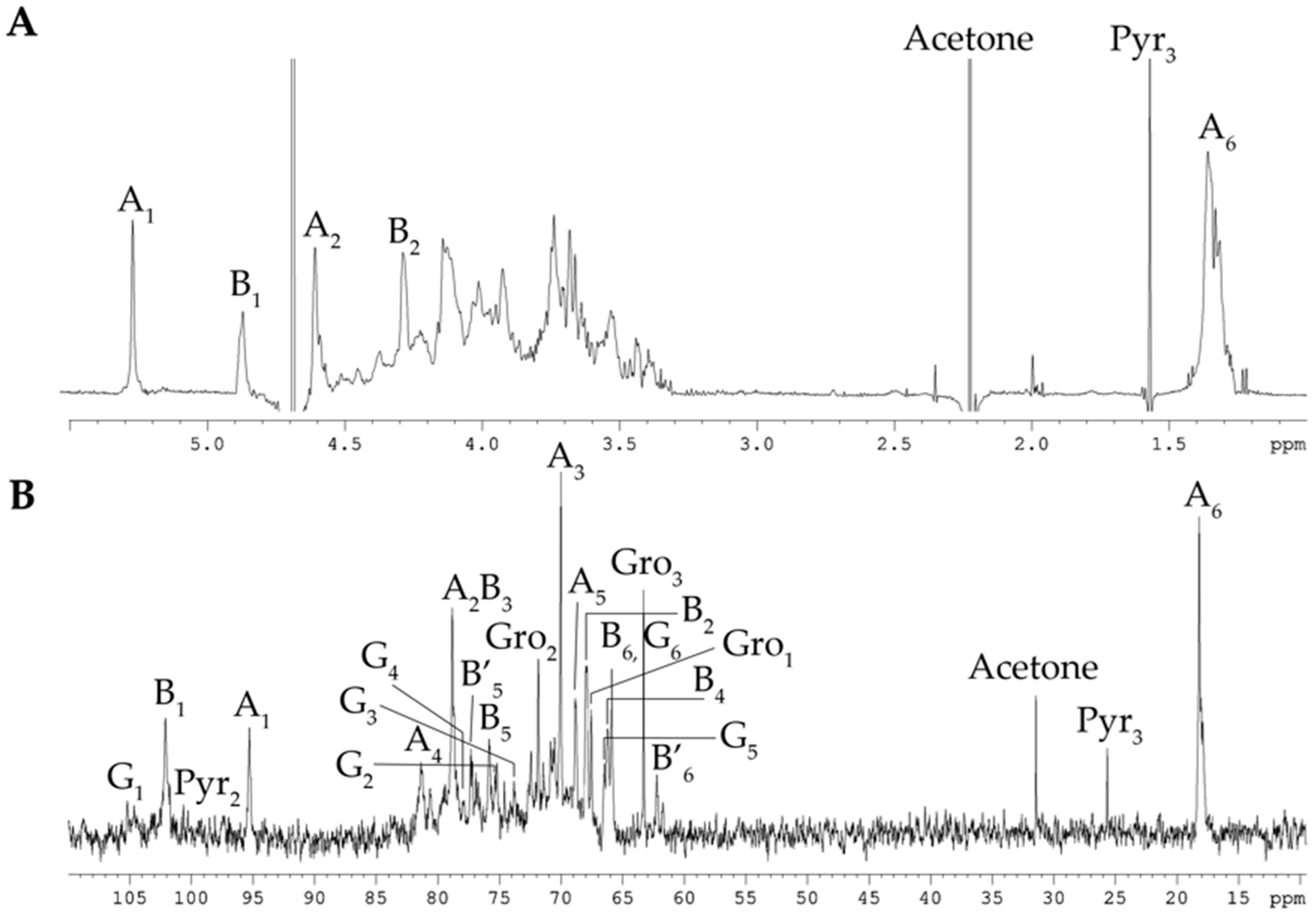
| Sugar Residue | H-1 | H-2 | H-3 | H-4 | H-5 | H-6a,b |
|---|---|---|---|---|---|---|
| C-1 | C-2 | C-3 | C-4 | C-5 | C-6 | |
| →4)-α-L-Rhap2S-(1→ | 5.27 | 4.61 | 4.13 | 3.68 | 4.02 | 1.34 |
| A | 95.3 | 78.9 | 70.2 | 80.6 | 68.7 | 18.1 |
| →4)-α-L-Rhap-(1→ | 4.97 | 4.01 | 3.98 | 3.68 | 4.02 | 1.34 |
| A′ | 97.5 | 71.8 | 71.4 | 80.6 | 68.7 | 18.1 |
| α-L-Rhap-(1→ | 5.29 | 4.13 | 3.93 | 3.46 | 3.91 | 1.28 |
| tA′ | 95.3 | 81.6 | 71.2 | 73.2 | 71.2 | 18.1 |
| →3)-β-D-Manp6PGro-(1→ | 4.88 | 4.28 | 3.71 | 3.67 | 3.53 | 4.14, 4.09 |
| B | 101.8 | 68.0 | 78.8 | 66.2 | 75.9 | 65.9 |
| Gro-(1→ | 3.97, 3.97 | 3.92 | 3.69, 3.63 | |||
| 67.7 | 71.9 | 63.3 | ||||
| →3)-β-D-Manp-(1→ | 4.88 | 4.28 | 3.71 | 3.67 | 3.39 | 3.94, 3.77 |
| B′ | 101.8 | 68.0 | 78.8 | 66.2 | 77.3 | 62.3 |
| β-D-Glcp4,6(S-Pyr)-(1→ | 4.67 | 3.42 | 3.69 | 3.45 | 3.53 | 4.13, 3.69 |
| G | 106.1 | 75.4 | 73.8 | 77.3 | 66.6 | 65.9 |
| S-Pyr | 1.57 | |||||
| 174.0 | 100.7 | 25.7 |
| Strain | Accession ID | Genome Size (bp) | Number of Contigs | G+C (mol%) | CPS/O-Antigen Length (bp) | Number of CDSs in Cluster |
|---|---|---|---|---|---|---|
| Cobetia marina KMM 1449 | GCF_045037775.1 | 4,184,796 | 1 | 62.5 | 87,222 | 65 |
| Cobetia sp. D5 | GCF_035666195.1 | 4,233,985 | 1 | 62.5 | 110,141 | 69 |
| Cobetia amphilecti N-80 | GCF_020217465.1 | 4,160,095 | 1 | 62.5 | 111,868 | 68 |
| Cobetia crustatorum ena-yuan-GCF_007786215.1 | GCF_963884875.1 | 4,215,468 | 163 | 57.5 | 78,218 | 56 |
| Cobetia marina MM1IDA2H-1AD | GCF_900119965.1 | 4,155,178 | 105 | 62 | 82,584 | 57 |
| Cobetia sp. cqz5-12 | GCF_016495405.1 | 4,209,007 | 1 | 62.5 | 78,267 | 55 |
| Cobetia marina NRIC 0814 (formerly C. pacifica KMM 3879T) | GCF_030010515.1 | 4,066,371 | 42 | 62.5 | 73,878 | 53 |
| Cobetia amphilecti NRIC 0813 (formerly C. litoralis KMM 3880T) | GCF_029846315.1 | 4,621,254 | 51 | 62.5 | 71,482 | 48 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kokoulin, M.S.; Savicheva, Y.V.; Filshtein, A.P.; Romanenko, L.A.; Isaeva, M.P. Structure of a Sulfated Capsular Polysaccharide from the Marine Bacterium Cobetia marina KMM 1449 and a Genomic Insight into Its Biosynthesis. Mar. Drugs 2025, 23, 29. https://doi.org/10.3390/md23010029
Kokoulin MS, Savicheva YV, Filshtein AP, Romanenko LA, Isaeva MP. Structure of a Sulfated Capsular Polysaccharide from the Marine Bacterium Cobetia marina KMM 1449 and a Genomic Insight into Its Biosynthesis. Marine Drugs. 2025; 23(1):29. https://doi.org/10.3390/md23010029
Chicago/Turabian StyleKokoulin, Maxim S., Yulia V. Savicheva, Alina P. Filshtein, Ludmila A. Romanenko, and Marina P. Isaeva. 2025. "Structure of a Sulfated Capsular Polysaccharide from the Marine Bacterium Cobetia marina KMM 1449 and a Genomic Insight into Its Biosynthesis" Marine Drugs 23, no. 1: 29. https://doi.org/10.3390/md23010029
APA StyleKokoulin, M. S., Savicheva, Y. V., Filshtein, A. P., Romanenko, L. A., & Isaeva, M. P. (2025). Structure of a Sulfated Capsular Polysaccharide from the Marine Bacterium Cobetia marina KMM 1449 and a Genomic Insight into Its Biosynthesis. Marine Drugs, 23(1), 29. https://doi.org/10.3390/md23010029






